Untargeted Multimodal Metabolomics Investigation of the Haemonchus contortus Exsheathment Secretome
Abstract
:1. Introduction
2. Materials and Methods
2.1. Production and Procurement of H. contortus
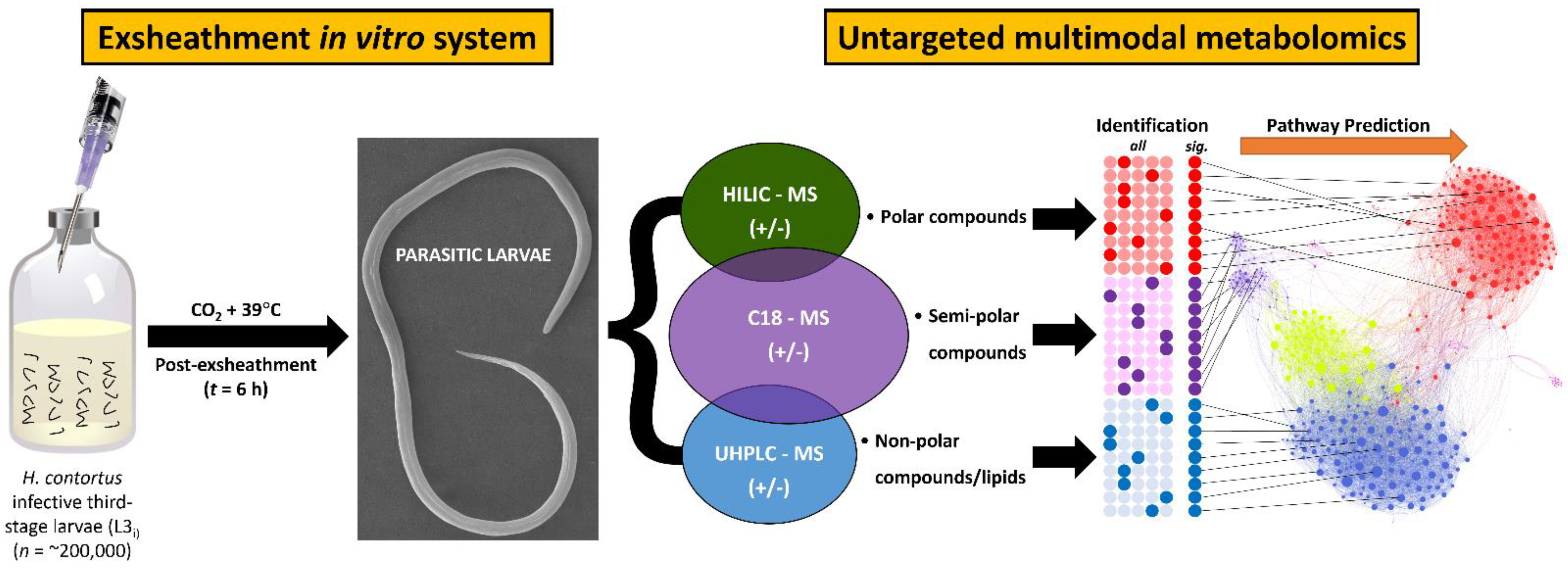
2.2. In Vitro Larval Exsheathment Culture Assay
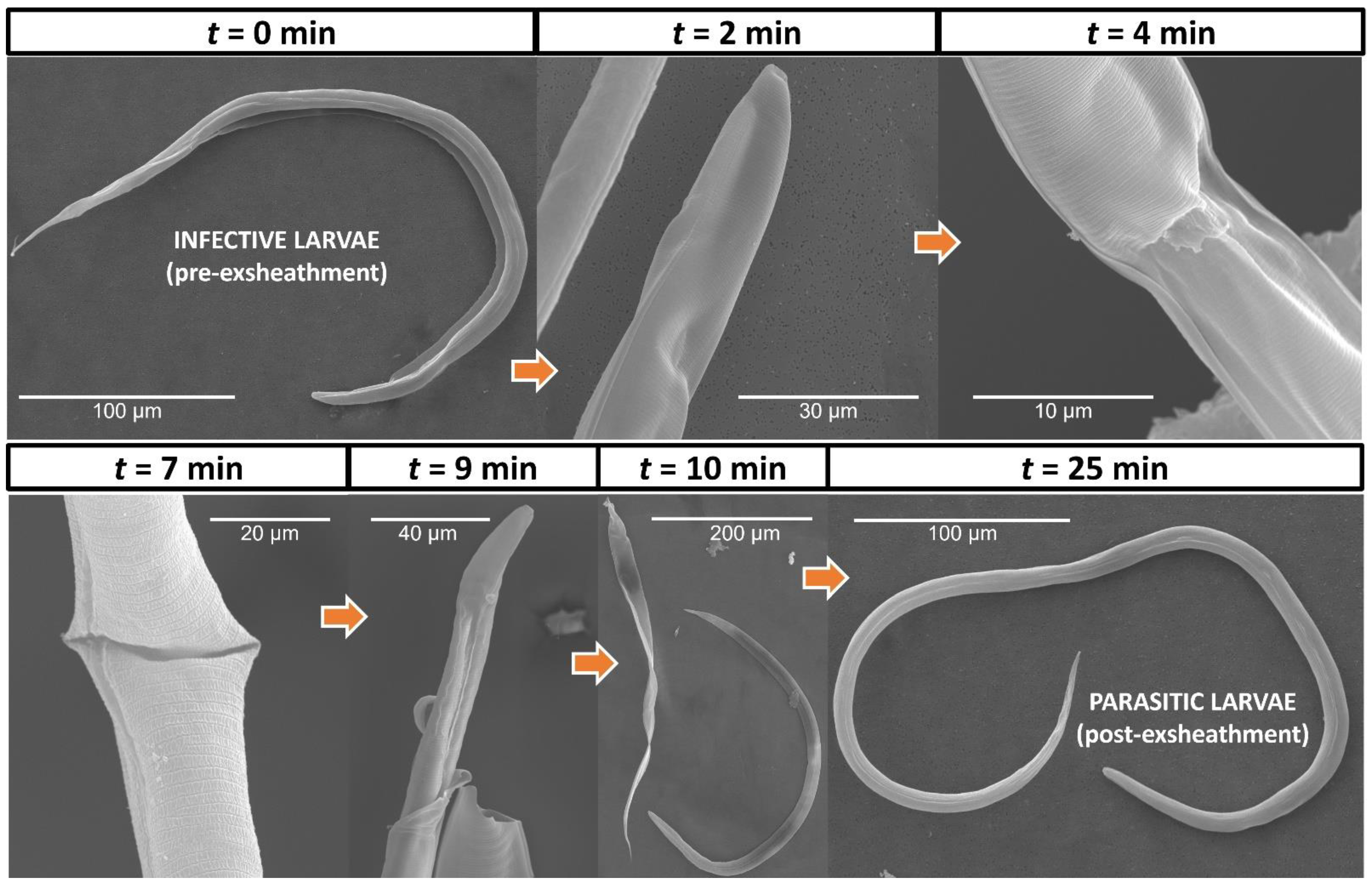
2.3. Electron Microscopy of H. contortus Larval Exsheathment
2.4. Metabolomics Analysis
2.4.1. Metabolomics Conditions and Analytical Procedures
2.4.2. Chromatography and Mass Spectrometry Spectral Acquisition
2.4.3. Data Processing, Peak Detection and Statistical Analysis
2.4.4. Metabolite Annotation and Identification
3. Results
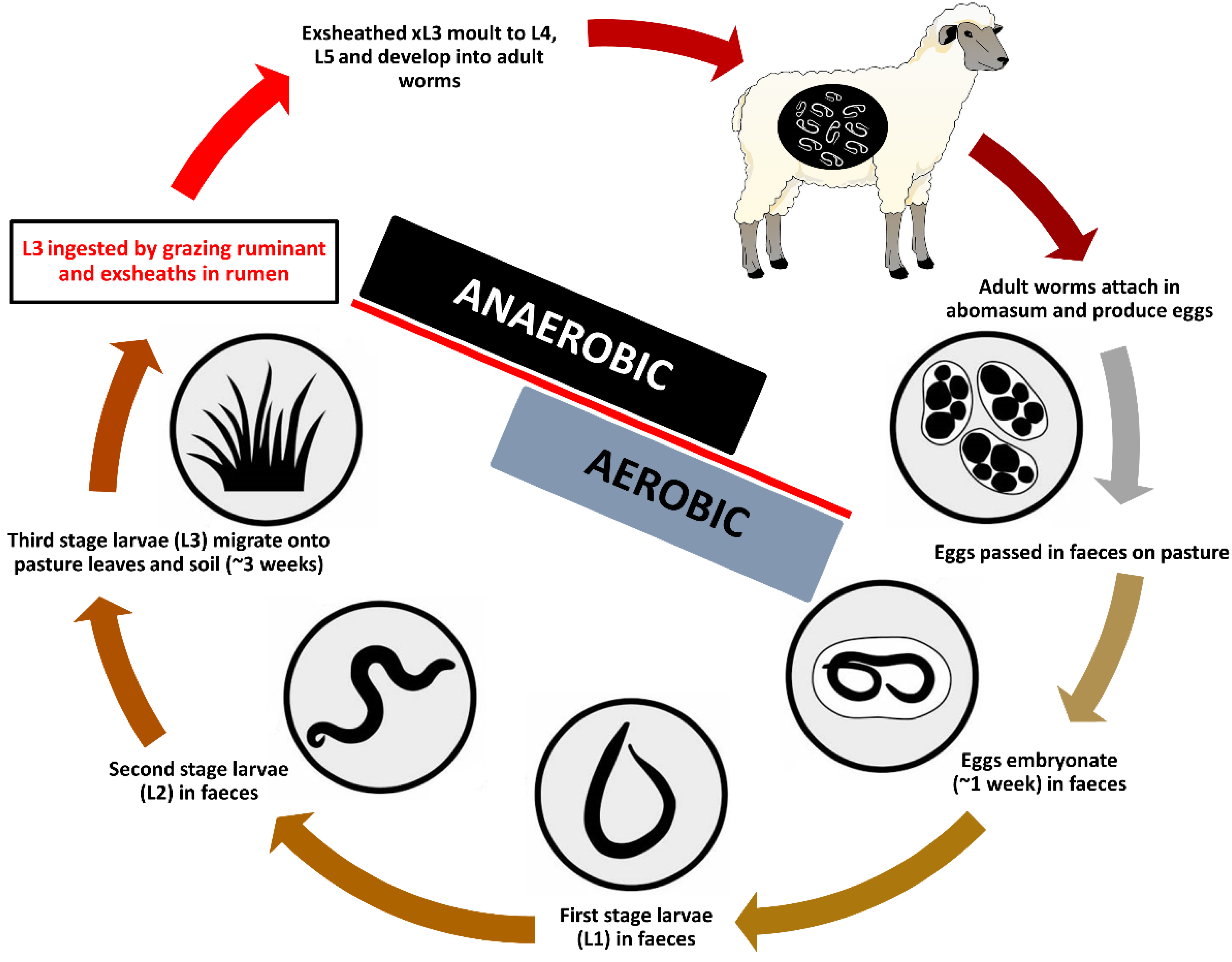
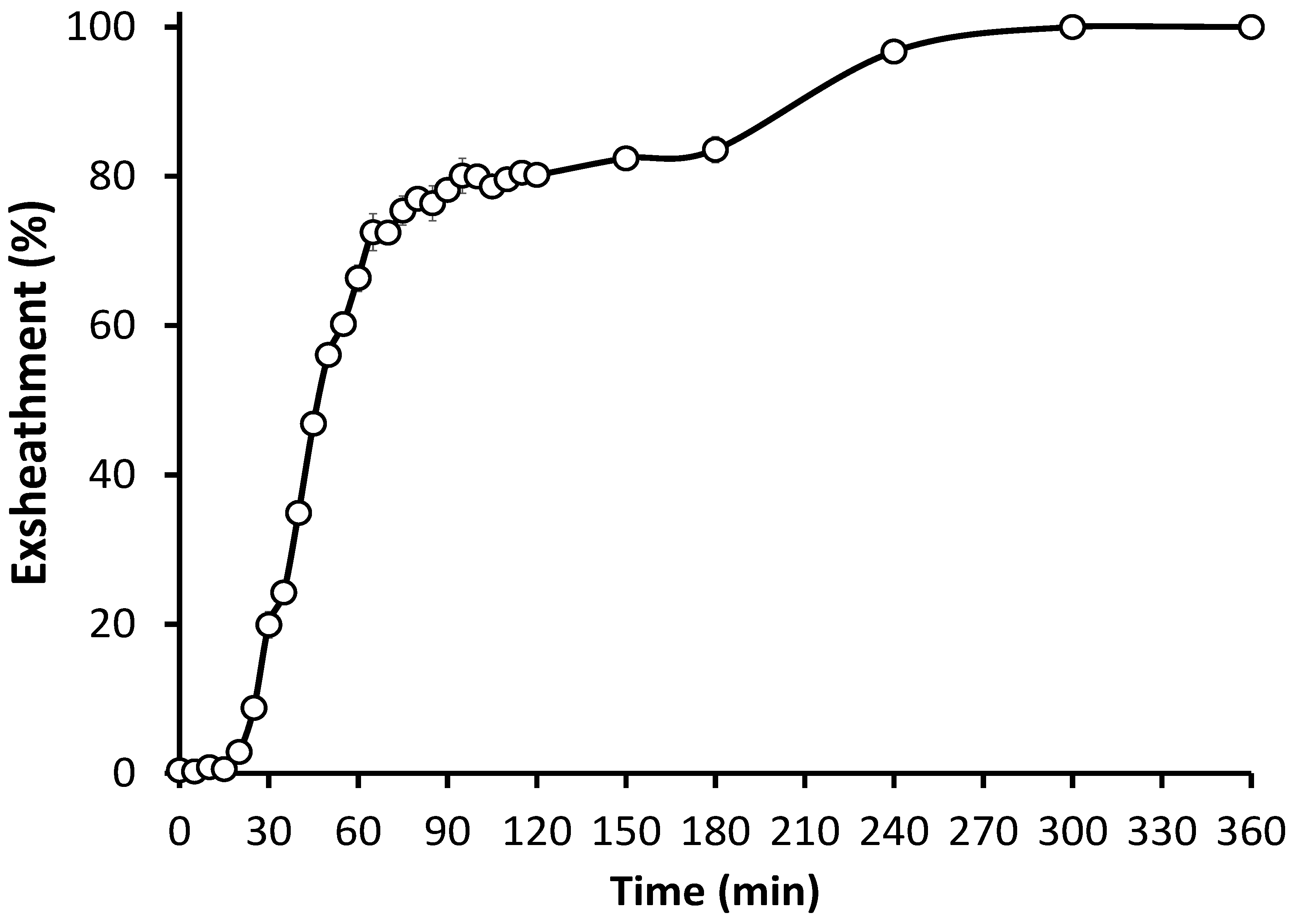
3.1. In Vitro Culture Technique Effectively Induces Third-Stage Larval Exsheathment in H. contortus
3.2. Identification of Exsheathment Associated Metabolites
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jourdan, P.M.; Lamberton, P.H.; Fenwick, A.; Addiss, D.G. Soil-transmitted helminth infections. Lancet 2018, 391, 252–265. [Google Scholar] [CrossRef]
- Charlier, J.; van der Voort, M.; Kenyon, F.; Skuce, P.; Vercruysse, J. Chasing helminths and their economic impact on farmed ruminants. Trends Parasitol. 2014, 30, 361–367. [Google Scholar] [CrossRef] [PubMed]
- Evans, T.; Chapple, N. The animal health market. Nat. Rev. Drug Discov. 2002, 1, 937–938. [Google Scholar] [CrossRef] [PubMed]
- Besier, R.; Kahn, L.; Sargison, N.; Van Wyk, J. Diagnosis, treatment and management of Haemonchus contortus in small ruminants. Adv. Parasitol. 2016, 93, 181–238. [Google Scholar] [PubMed]
- Kaplan, R.M.; Vidyashankar, A.N. An inconvenient truth: Global worming and anthelmintic resistance. Vet. Parasitol. 2012, 186, 70–78. [Google Scholar] [CrossRef] [PubMed]
- Geary, T.G.; Sakanari, J.A.; Caffrey, C.R. Anthelmintic drug discovery: Into the future. J. Parasitol. 2015, 101, 125–133. [Google Scholar] [CrossRef]
- Kotze, A.; Prichard, R. Anthelmintic resistance in Haemonchus contortus: History, mechanisms and diagnosis. Adv. Parasitol. 2016, 93, 397–428. [Google Scholar]
- Lee, D.L. Cuticle, moulting and exsheathment. In The Biology of Nematodes; Taylor and Francis: London, UK, 2002; pp. 171–209. [Google Scholar] [CrossRef]
- Rogers, W. The physiology of infective processes of nematode parasites; the stimulus from the animal host. Proc. R. Soc. London. Ser. B Biol. Sci. 1960, 152, 367–386. [Google Scholar]
- Sommerville, R. The exsheathing mechanism of nematode infective larvae. Exp. Parasitol. 1957, 6, 18–30. [Google Scholar] [CrossRef]
- Taylor, A.; Whitlock, J. The exsheathing stimulus for infective larvae of Haemonchus contortus. Cornell Vet. 1960, 50, 339–344. [Google Scholar]
- Rogers, W.; Sommerville, R. The physiology of the second ecdysis of parasitic nematodes. Parasitology 1960, 50, 329–348. [Google Scholar] [CrossRef]
- Silverman, P.H.; Podger, K.R. In vitro exsheathment of some nematode infective larvae. Exp. Parasitol. 1964, 15, 314–324. [Google Scholar] [CrossRef]
- Rogers, W.; Sommerville, R. Physiology of exsheathment in nematodes and its relation to parasitism. Nature 1957, 179, 619–621. [Google Scholar] [CrossRef] [PubMed]
- Petronijevic, T.; Rogers, W. Undissociated bases as the stimulus for the development of early parasitic stages of nematodes. Int. J. Parasitol. 1987, 17, 911–915. [Google Scholar] [CrossRef]
- Petronijevic, T.; Rogers, W.; Sommerville, R. Organic and inorganic acids as the stimulus for exsheathment of infective juveniles of nematodes. Int. J. Parasitol. 1986, 16, 163–168. [Google Scholar] [CrossRef]
- Petronijevic, T.; Rogers, W.; Sommerville, R. Carbonic acid as the host signal for the development of parasitic stages of nematodes. Int. J. Parasitol. 1985, 15, 661–667. [Google Scholar] [CrossRef]
- Petronijevic, T.; Rogers, W. The physiology of infection with nematodes: The role of intracellular pH in the development of the early parasitic stage. Comp. Biochem. Physiol. Part A Physiol. 1987, 88, 207–212. [Google Scholar] [CrossRef]
- Bekelaar, K.; Waghorn, T.; Tavendale, M.; McKenzie, C.; Leathwick, D. Heat shock, but not temperature, is a biological trigger for the exsheathment of third-stage larvae of Haemonchus contortus. Parasitol. Res. 2018, 117, 2395–2402. [Google Scholar] [CrossRef]
- Ma, G.; Wang, T.; Korhonen, P.K.; Hofmann, A.; Sternberg, P.W.; Young, N.D.; Gasser, R.B. Elucidating the molecular and developmental biology of parasitic nematodes: Moving to a multiomics paradigm. In Advances in Parasitology; Elsevier: Amsterdam, The Netherlands, 2020; Volume 108, pp. 175–229. [Google Scholar]
- Borges, D.G.L.; Echeverria, J.T.; de Oliveira, T.L.; Heckler, R.P.; de Freitas, M.G.; Damasceno-Junior, G.A.; Carollo, C.A.; Borges, F.d.A. Discovery of potential ovicidal natural products using metabolomics. PLoS ONE 2019, 14, e0211237. [Google Scholar] [CrossRef]
- Watts, J.L.; Ristow, M. Lipid and carbohydrate metabolism in Caenorhabditis elegans. Genetics 2017, 207, 413–446. [Google Scholar]
- Wang, T.; Nie, S.; Reid, G.E.; Gasser, R.B. Helminth lipidomics–technical aspects and future prospects. Curr. Res. Parasitol. Vector Borne Dis. 2021, 1, 100018. [Google Scholar] [CrossRef] [PubMed]
- Van Assche, R.; Broeckx, V.; Boonen, K.; Maes, E.; De Haes, W.; Schoofs, L.; Temmerman, L. Integrating-omics: Systems biology as explored through C. elegans research. J. Mol. Biol. 2015, 427, 3441–3451. [Google Scholar] [CrossRef] [PubMed]
- Rougvie, A.E.; Moss, E.G. Developmental transitions in C. elegans larval stages. Curr. Top. Dev. Biol. 2013, 105, 153–180. [Google Scholar] [PubMed]
- Faunes, F.; Larraín, J. Conservation in the involvement of heterochronic genes and hormones during developmental transitions. Dev. Biol. 2016, 416, 3–17. [Google Scholar] [CrossRef] [PubMed]
- Mahanti, P.; Bose, N.; Bethke, A.; Judkins, J.C.; Wollam, J.; Dumas, K.J.; Zimmerman, A.M.; Campbell, S.L.; Hu, P.J.; Antebi, A. Comparative metabolomics reveals endogenous ligands of DAF-12, a nuclear hormone receptor, regulating C. elegans development and lifespan. Cell Metab. 2014, 19, 73–83. [Google Scholar] [CrossRef] [PubMed]
- Salzer, L.; Witting, M. Quo Vadis Caenorhabditis elegans Metabolomics—a review of current methods and applications to explore metabolism in the nematode. Metabolites 2021, 11, 284. [Google Scholar] [CrossRef]
- Stoltzfus, J.D.; Pilgrim, A.A.; De’Broski, R.H. Perusal of parasitic nematode ‘omics in the post-genomic era. Mol. Biochem. Parasitol. 2017, 215, 11–22. [Google Scholar] [CrossRef]
- Nikolaou, S.; Gasser, R. Prospects for exploring molecular developmental processes in Haemonchus contortus. Int. J. Parasitol. 2006, 36, 859–868. [Google Scholar] [CrossRef]
- Ma, G.; Wang, T.; Korhonen, P.K.; Young, N.D.; Nie, S.; Ang, C.-S.; Williamson, N.A.; Reid, G.E.; Gasser, R.B. Dafachronic acid promotes larval development in Haemonchus contortus by modulating dauer signalling and lipid metabolism. PLoS Pathog. 2019, 15, e1007960. [Google Scholar] [CrossRef]
- Wang, T.; Nie, S.; Ma, G.; Korhonen, P.K.; Koehler, A.V.; Ang, C.-S.; Reid, G.E.; Williamson, N.A.; Gasser, R.B. The developmental lipidome of Haemonchus contortus. Int. J. Parasitol. 2018, 48, 887–895. [Google Scholar] [CrossRef]
- Wang, T.; Ma, G.; Ang, C.-S.; Korhonen, P.K.; Stroehlein, A.J.; Young, N.D.; Hofmann, A.; Chang, B.C.; Williamson, N.A.; Gasser, R.B. The developmental phosphoproteome of Haemonchus contortus. J. Proteom. 2020, 213, 103615. [Google Scholar] [CrossRef] [PubMed]
- Viglierchio, D.; Schmitt, R.V. On the methodology of nematode extraction from field samples: Baermann funnel modifications. J. Nematol. 1983, 15, 438. [Google Scholar] [PubMed]
- Bisset, S.; Knight, J.; Bouchet, C. A multiplex PCR-based method to identify strongylid parasite larvae recovered from ovine faecal cultures and/or pasture samples. Vet. Parasitol. 2014, 200, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Palevich, N.; Maclean, P.H.; Baten, A.; Scott, R.W.; Leathwick, D.M. The genome sequence of the anthelmintic-susceptible New Zealand Haemonchus contortus. Genome Biol. Evol. 2019, 11, 1965–1970. [Google Scholar] [CrossRef]
- Palevich, N.; Maclean, P.H.; Choi, Y.-J.; Mitreva, M. Characterization of the Complete Mitochondrial Genomes of Two Sibling Species of Parasitic Roundworms, Haemonchus contortus and Teladorsagia circumcincta. Front. Genet. 2020, 11, 573395. [Google Scholar] [CrossRef]
- Palevich, N.; Maclean, P.; Baten, A.; Scott, R.; Leathwick, D.M. The complete mitochondrial genome of the New Zealand parasitic roundworm Haemonchus contortus (Trichostrongyloidea: Haemonchidae) field strain NZ_Hco_NP. Mitochondrial Dna Part B 2019, 4, 2208–2210. [Google Scholar] [CrossRef] [PubMed]
- Palevich, N.; Kelly, W.J.; Ganesh, S.; Rakonjac, J.; Attwood, G.T. Butyrivibrio hungatei MB2003 competes effectively for soluble sugars released by Butyrivibrio proteoclasticus B316T during growth on xylan or pectin. Appl. Environ. Microbiol. 2019, 85, e02056-18. [Google Scholar] [CrossRef]
- Palevich, N.; Kelly, W.J.; Leahy, S.C.; Altermann, E.; Rakonjac, J.; Attwood, G.T. The complete genome sequence of the rumen bacterium Butyrivibrio hungatei MB2003. Stand. Genom. Sci. 2017, 12, 72. [Google Scholar] [CrossRef]
- Palevich, N.; Kelly, W.J.; Leahy, S.C.; Denman, S.; Altermann, E.; Rakonjac, J.; Attwood, G.T. Comparative genomics of rumen Butyrivibrio spp. uncovers a continuum of polysaccharide-degrading capabilities. Appl. Environ. Microbiol. 2019, 86, e01993-19. [Google Scholar] [CrossRef]
- Palevich, N.; Maclean, P.H.; Kelly, W.J.; Leahy, S.C.; Rakonjac, J.; Attwood, G.T. Complete genome sequence of the polysaccharide-degrading rumen bacterium Pseudobutyrivibrio xylanivorans MA3014 reveals an incomplete glycolytic pathway. Genome Biol. Evol. 2020, 12, 1566–1572. [Google Scholar] [CrossRef]
- Seshadri, R.; Leahy, S.C.; Attwood, G.T.; Teh, K.H.; Lambie, S.C.; Cookson, A.L.; Eloe-Fadrosh, E.A.; Pavlopoulos, G.A.; Hadjithomas, M.; Varghese, N.J. Cultivation and sequencing of rumen microbiome members from the Hungate1000 Collection. Nat. Biotechnol. 2018, 36, 359–367. [Google Scholar] [CrossRef] [PubMed]
- Pahalagedara, A.S.N.W.; Flint, S.; Palmer, J.; Subbaraj, A.; Brightwell, G.; Gupta, T.B. Antimicrobial activity of soil Clostridium enriched conditioned media against Bacillus mycoides, Bacillus cereus, and Pseudomonas aeruginosa. Front. Microbiol. 2020, 11, 3113. [Google Scholar] [CrossRef] [PubMed]
- Fraser, K.; Harrison, S.J.; Lane, G.A.; Otter, D.E.; Hemar, Y.; Quek, S.-Y.; Rasmussen, S. Non-targeted analysis of tea by hydrophilic interaction liquid chromatography and high resolution mass spectrometry. Food Chem. 2012, 134, 1616–1623. [Google Scholar] [CrossRef]
- Subbaraj, A.K.; Huege, J.; Fraser, K.; Cao, M.; Rasmussen, S.; Faville, M.; Harrison, S.J.; Jones, C.S. A large-scale metabolomics study to harness chemical diversity and explore biochemical mechanisms in ryegrass. Commun. Biol. 2019, 2, 87. [Google Scholar] [CrossRef]
- Holman, J.D.; Tabb, D.L.; Mallick, P. Employing ProteoWizard to convert raw mass spectrometry data. Curr. Protoc. Bioinform. 2014, 46, 13.24.1–13.24.9. [Google Scholar] [CrossRef]
- Dunn, W.B.; Wilson, I.D.; Nicholls, A.W.; Broadhurst, D. The importance of experimental design and QC samples in large-scale and MS-driven untargeted metabolomic studies of humans. Bioanalysis 2012, 4, 2249–2264. [Google Scholar] [CrossRef]
- Smith, C.A.; Want, E.J.; O’Maille, G.; Abagyan, R.; Siuzdak, G. XCMS: Processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 2006, 78, 779–787. [Google Scholar] [CrossRef]
- Cao, M.; Fraser, K.; Jones, C.; Stewart, A.; Lyons, T.; Faville, M.; Barrett, B. Untargeted metabotyping Lolium perenne reveals population-level variation in plant flavonoids and alkaloids. Front. Plant Sci. 2017, 8, 133. [Google Scholar] [CrossRef]
- Cao, M.; Fraser, K.; Huege, J.; Featonby, T.; Rasmussen, S.; Jones, C. Predicting retention time in hydrophilic interaction liquid chromatography mass spectrometry and its use for peak annotation in metabolomics. Metabolomics 2015, 11, 696–706. [Google Scholar] [CrossRef]
- Cao, M.; Fraser, K.; Rasmussen, S. Computational analyses of spectral trees from electrospray multi-stage mass spectrometry to aid metabolite identification. Metabolites 2013, 3, 1036–1050. [Google Scholar] [CrossRef]
- Campbell, L.R.; Gaugler, R. Mechanisms for exsheathment of entomopathogenic nematodes. Int. J. Parasitol. 1991, 21, 219–224. [Google Scholar] [CrossRef]
- Lapage, G. The second ecdysis of infective nematode larvae. Parasitology 1935, 27, 186–206. [Google Scholar] [CrossRef]
- Van Wyk, J. Viability of nematode larvae after exsheathment with sodium hypochlorite. Parasitol. Today (Pers. Ed.) 1998, 14, 474–475. [Google Scholar] [CrossRef]
- Ma, G.; Gasser, R.B.; Wang, T.; Korhonen, P.K.; Young, N.D. Toward integrative ‘omics of the barber’s pole worm and related parasitic nematodes. Infect. Genet. Evol. 2020, 85, 104500. [Google Scholar] [CrossRef] [PubMed]
- Cantacessi, C.; Campbell, B.E.; Young, N.D.; Jex, A.R.; Hall, R.S.; Presidente, P.J.A.; Zawadzki, J.L.; Zhong, W.; Aleman-Meza, B.; Loukas, A.; et al. Differences in transcription between free-living and CO2-activated third-stage larvae of Haemonchus contortus. Bmc Genom. 2010, 11, 266. [Google Scholar] [CrossRef] [PubMed]
- Jasmer, D.P.; Rosa, B.A.; Mitreva, M. Cell death and transcriptional responses induced in larvae of the nematode Haemonchus contortus by toxins/toxicants with broad phylogenetic efficacy. Pharmaceuticals 2021, 14, 598. [Google Scholar] [CrossRef] [PubMed]
- Ma, G.; Wang, T.; Korhonen, P.K.; Ang, C.-S.; Williamson, N.A.; Young, N.D.; Stroehlein, A.J.; Hall, R.S.; Koehler, A.V.; Hofmann, A.; et al. Molecular alterations during larval development of Haemonchus contortus in vitro are under tight post-transcriptional control. Int. J. Parasitol. 2018, 48, 763–772. [Google Scholar] [CrossRef]
- Zuzarte-Luís, V.; Mota, M.M. Parasite sensing of host nutrients and environmental cues. Cell Host Microbe 2018, 23, 749–758. [Google Scholar] [CrossRef] [PubMed]
- El-Ashram, S.; Suo, X. Exploring the microbial community (microflora) associated with ovine Haemonchus contortus (macroflora) field strains. Sci. Rep. 2017, 7, 70. [Google Scholar] [CrossRef] [PubMed]
- Mafuna, T.; Soma, P.; Tsotetsi-Khambule, A.; Hefer, C.; Muchadeyi, F.; Thekisoe, O.; Pierneef, R. Bacterial profiling of Haemonchus contortus gut microbiome infecting Dohne Merino sheep in South Africa. Sci. Rep. 2021, 11, 5905. [Google Scholar] [CrossRef] [PubMed]
- Hogan, G.; Walker, S.; Turnbull, F.; Curiao, T.; Morrison, A.A.; Flores, Y.; Andrews, L.; Claesson, M.J.; Tangney, M.; Bartley, D.J. Microbiome analysis as a platform R&D tool for parasitic nematode disease management. ISME J. 2019, 13, 2664–2680. [Google Scholar] [PubMed]
- Wu, Z.; Wang, L.; Li, J.; Wang, L.; Wu, Z.; Sun, X. Extracellular vesicle-mediated communication within host-parasite interactions. Front. Immunol. 2019, 9, 3066. [Google Scholar] [CrossRef] [PubMed]
- Hobson, P.N.; Stewart, C.S. The Rumen Microbial Ecosystem; Springer Science & Business Media: Berlin, Germany, 2012; Available online: https://link.springer.com/book/10.1007/978-94-009-1453-7 (accessed on 20 April 2022).
- Harder, A. The biochemistry of Haemonchus contortus and other parasitic nematodes. Adv. Parasitol. 2016, 93, 69–94. [Google Scholar] [PubMed]
- Mansfeld, J.; Urban, N.; Priebe, S.; Groth, M.; Frahm, C.; Hartmann, N.; Gebauer, J.; Ravichandran, M.; Dommaschk, A.; Schmeisser, S. Branched-chain amino acid catabolism is a conserved regulator of physiological ageing. Nat. Commun. 2015, 6, 10043. [Google Scholar] [CrossRef] [PubMed]
- Fuchs, S.; Bundy, J.G.; Davies, S.K.; Viney, J.M.; Swire, J.S.; Leroi, A.M. A metabolic signature of long life in Caenorhabditis elegans. Bmc Biol. 2010, 8, 14. [Google Scholar] [CrossRef] [PubMed]
- Calvo, A.C.; Pey, A.L.; Ying, M.; Loer, C.M.; Martinez, A. Anabolic function of phenylalanine hydroxylase in Caenorhabditis elegans. FASEB J. 2008, 22, 3046–3058. [Google Scholar] [CrossRef]
- Walker, J.; Barrett, J. Parasite sulphur amino acid metabolism. Int. J. Parasitol. 1997, 27, 883–897. [Google Scholar] [CrossRef]
- Yeshi, K.; Creek, D.J.; Anderson, D.; Ritmejerytė, E.; Becker, L.; Loukas, A.; Wangchuk, P. Metabolomes and lipidomes of the infective stages of the gastrointestinal nematodes, Nippostrongylus brasiliensis and Trichuris muris. Metabolites 2020, 10, 446. [Google Scholar] [CrossRef]
- Schwarz, E.M.; Korhonen, P.K.; Campbell, B.E.; Young, N.D.; Jex, A.R.; Jabbar, A.; Hall, R.S.; Mondal, A.; Howe, A.C.; Pell, J. The genome and developmental transcriptome of the strongylid nematode Haemonchus contortus. Genome Biol. 2013, 14, R89. [Google Scholar] [CrossRef]
- Laing, R.; Kikuchi, T.; Martinelli, A.; Tsai, I.J.; Beech, R.N.; Redman, E.; Holroyd, N.; Bartley, D.J.; Beasley, H.; Britton, C. The genome and transcriptome of Haemonchus contortus, a key model parasite for drug and vaccine discovery. Genome Biol. 2013, 14, R88. [Google Scholar] [CrossRef]
- Wyss, M.; Kaddurah-Daouk, R. Creatine and creatinine metabolism. Physiol. Rev. 2000, 80, 1107–1213. [Google Scholar] [CrossRef]
- Murray, P.; Hayward, S.A.; Govan, G.G.; Gracey, A.Y.; Cossins, A.R. An explicit test of the phospholipid saturation hypothesis of acquired cold tolerance in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 2007, 104, 5489–5494. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.K.; Li, Z.; Lu, A.Y.; Sun, F.; Chen, S.; Rothe, M.; Menzel, R.; Sun, F.; Horvitz, H.R. Acyl-CoA dehydrogenase drives heat adaptation by sequestering fatty acids. Cell 2015, 161, 1152–1163. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, S.; Svahn, S.L.; Johansson, M.E. Effects of omega-3 fatty acids on immune cells. Int. J. Mol. Sci. 2019, 20, 5028. [Google Scholar] [CrossRef] [PubMed]
- Tzeng, H.-T.; Chyuan, I.-T.; Chen, W.-Y. Shaping of innate immune response by fatty acid metabolite palmitate. Cells 2019, 8, 1633. [Google Scholar] [CrossRef]
- Calder, P. N-3 polyunsaturated fatty acids, inflammation and immunity: Pouring oil on troubled waters or another fishy tale? Nutr. Res. 2001, 21, 309–341. [Google Scholar] [CrossRef]
- Tebbey, P.; Buttke, T. Molecular basis for the immunosuppressive action of stearic acid on T cells. Immunology 1990, 70, 379. [Google Scholar] [PubMed]
| Metabolite Class | Molecular Species | Platform | Isotopic Peak (m/z_rt) | Ion Type | Calc Mass | Rt * (min) | C. elegans KEGG Pathway ID |
|---|---|---|---|---|---|---|---|
| C18 and HILIC | |||||||
| Amino acids | Lysine | HP | 147.1128_15.79 | [M+H]+ | 147.1128 | 15.8 | Lysine biosynthesis (cel00300) and degradation (cel00310) |
| HN | 145.0982_15.78 | [M−H]− | 145.0972 | 15.8 | |||
| Glycine | HP | 76.0396_13.21 | [M+H]+ | 76.0398 | 13.2 | cel00260-Glycine, serine, and threonine metabolism | |
| Threonine | HP | 120.0654_12.76 | [M+H]+ | 120.0655 | 12.7 | ||
| Methionine | HP | 150.0583_10.57 | [M+H]+ | 150.0583 | 10.5 | cel00270-Cysteine and methionine metabolism | |
| CP | 150.0595_1.10 | [M+H]+ | 150.0583 | 1.08 | |||
| Alanine | HP | 90.0546_12.54 | [M+H]+ | 90.0549 | 12.5 | cel00250-Alanine, aspartate and glutamate metabolism | |
| Glutamic acid | HN | 146.0465_13.2 | [M−H]− | 146.0448 | 13.3 | ||
| Isoleucine | HP | 132.1030_10.12 | [M+H]+ | 132.1019 | 10.2 | Valine, leucine, and isoleucine degradation (cel00280) and biosynthesis (cel00290) | |
| CP | 132.1022_1.87 | [M+H]+ | 132.1019 | 1.76 | |||
| Leucine | HP | 132.1025_9.84 | [M+H]+ | 132.1019 | 9.8 | ||
| CP | 132.1027_1.73 | [M+H]+ | 132.1019 | 1.75 | |||
| Tryptophan | HP | 205.0969_10.45 | [M+H]+ | 205.0972 | 10.5 | cel00380-Tryptophan metabolism | |
| Phenylalanine | HP | 166.0877_9.80 | [M+H]+ | 166.0863 | 9.8 | cel00360-Phenylalanine metabolism | |
| Purines | Xanthine | HN | 151.0258_8.62 | [M−H]− | 151.0251 | 8.3 * | cel00230-Purine metabolism |
| CP | 153.0419_1.39 | [M+H]+ | 153.0407 | ||||
| Hypoxanthine | HP | 137.0470_8.24 | [M+H]+ | 137.0458 | 7.9 * | ||
| HN | 135.0309_8.24 | [M−H]− | 135.0301 | ||||
| CP | 137.0473_1.2 | [M+H]+ | 137.0458 | 1.2 | |||
| Organic compounds | Piperidine | HP | 86.0963_9.84 | [M+H]+ | 86.097 | 10.3 ** | cel00310-Lysine degradation |
| Carnitine | HP | 162.1125_10.46 | [M+H]+ | 162.1125 | 10.6 ** | cel01212-Fatty acid metabolism | |
| CP | 162.1131_0.67 | [M+H]+ | 162.1125 | ||||
| LIPIDS | |||||||
| LPG | Palmitic acid (16:0) | LN | 255.2327_3.5 | [M−H]− | 255.2319 | [52] | cel00061-Fatty acid biosynthesis |
| Stearic acid (18:0) | LN | 283.2641_4.18 | [M−H]− | 283.2632 | |||
| DG | DG(32:0) | LN | 603.4765_7.85 | [M+Cl]− | 603.475 | cel00561-Glycerolipid metabolism | |
| DG(34:0) | LN | 631.5075_8.56 | [M+Cl]− | 631.5063 | |||
| DG(36:0) | LN | 659.5388_9.23 | [M+Cl]− | 659.5376 | |||
| DG(18:0/18:0) MS2 | LP | 642.6033_9.98 | [M+NH4]+ | 642.6031 | |||
| DG(16:0/18:0) MS2 | LP | 614.5719_8.55 | [M+NH4]+ | 614.5718 | |||
| GP | PKHdiA-PS | LN | 686.2696_6.46 | [M+Cl]− | 686.2703 | cel00564-Glycerophospholipid metabolism | |
| OHHdiA-PS | LN | 714.3014_7.23 | [M+Cl]− | 714.3016 | |||
| FA | hydroxy-eicosenoic acid | LP | 344.3161_2.28 | [M+NH4]+ | 344.3159 | cel00590-Arachidonic acid metabolism | |
| 2-oxo-docosanoic acid | LP | 372.3473_2.88 | [M+NH4]+ | 372.3472 | cel01040-Biosynthesis of unsaturated fatty acids | ||
| TG * | TG(16:0/18:0/18:0) MS2 | LP | 880.8331_12.57 | [M+NH4]+ | 880.8328 | cel00561-Glycerolipid metabolism |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Palevich, N.; Maclean, P.H.; Candy, P.M.; Taylor, W.; Mladineo, I.; Cao, M. Untargeted Multimodal Metabolomics Investigation of the Haemonchus contortus Exsheathment Secretome. Cells 2022, 11, 2525. https://doi.org/10.3390/cells11162525
Palevich N, Maclean PH, Candy PM, Taylor W, Mladineo I, Cao M. Untargeted Multimodal Metabolomics Investigation of the Haemonchus contortus Exsheathment Secretome. Cells. 2022; 11(16):2525. https://doi.org/10.3390/cells11162525
Chicago/Turabian StylePalevich, Nikola, Paul H. Maclean, Paul M. Candy, Wendy Taylor, Ivona Mladineo, and Mingshu Cao. 2022. "Untargeted Multimodal Metabolomics Investigation of the Haemonchus contortus Exsheathment Secretome" Cells 11, no. 16: 2525. https://doi.org/10.3390/cells11162525
APA StylePalevich, N., Maclean, P. H., Candy, P. M., Taylor, W., Mladineo, I., & Cao, M. (2022). Untargeted Multimodal Metabolomics Investigation of the Haemonchus contortus Exsheathment Secretome. Cells, 11(16), 2525. https://doi.org/10.3390/cells11162525







