Collisions of Cortical Microtubules with Membrane Associated Myosin VIII Tail
Abstract
1. Introduction
2. Materials and Methods
2.1. Plants, Plasmids and Transient Expression
2.2. Microscopy and Image Analysis
2.3. The Simulation
3. Results
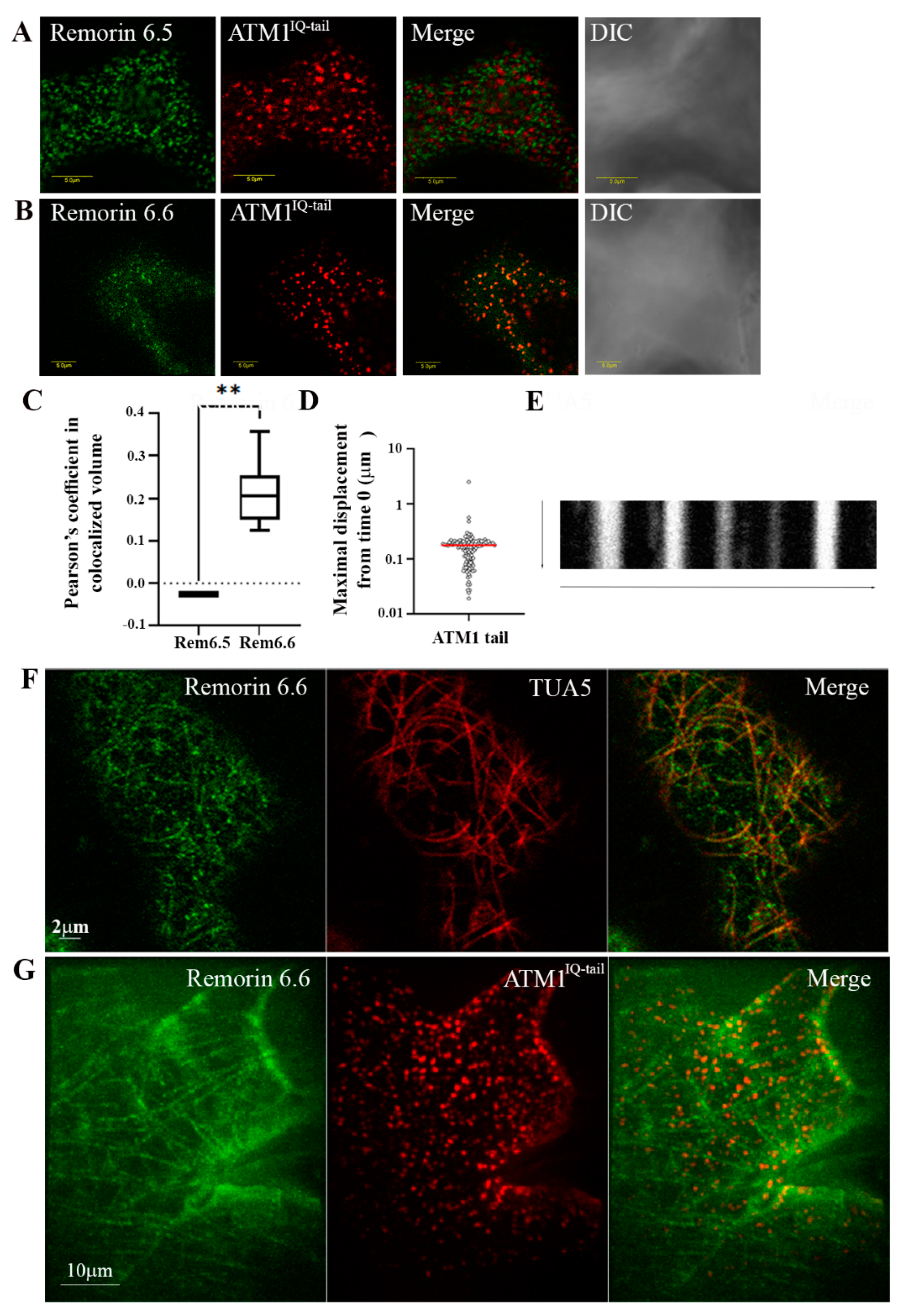
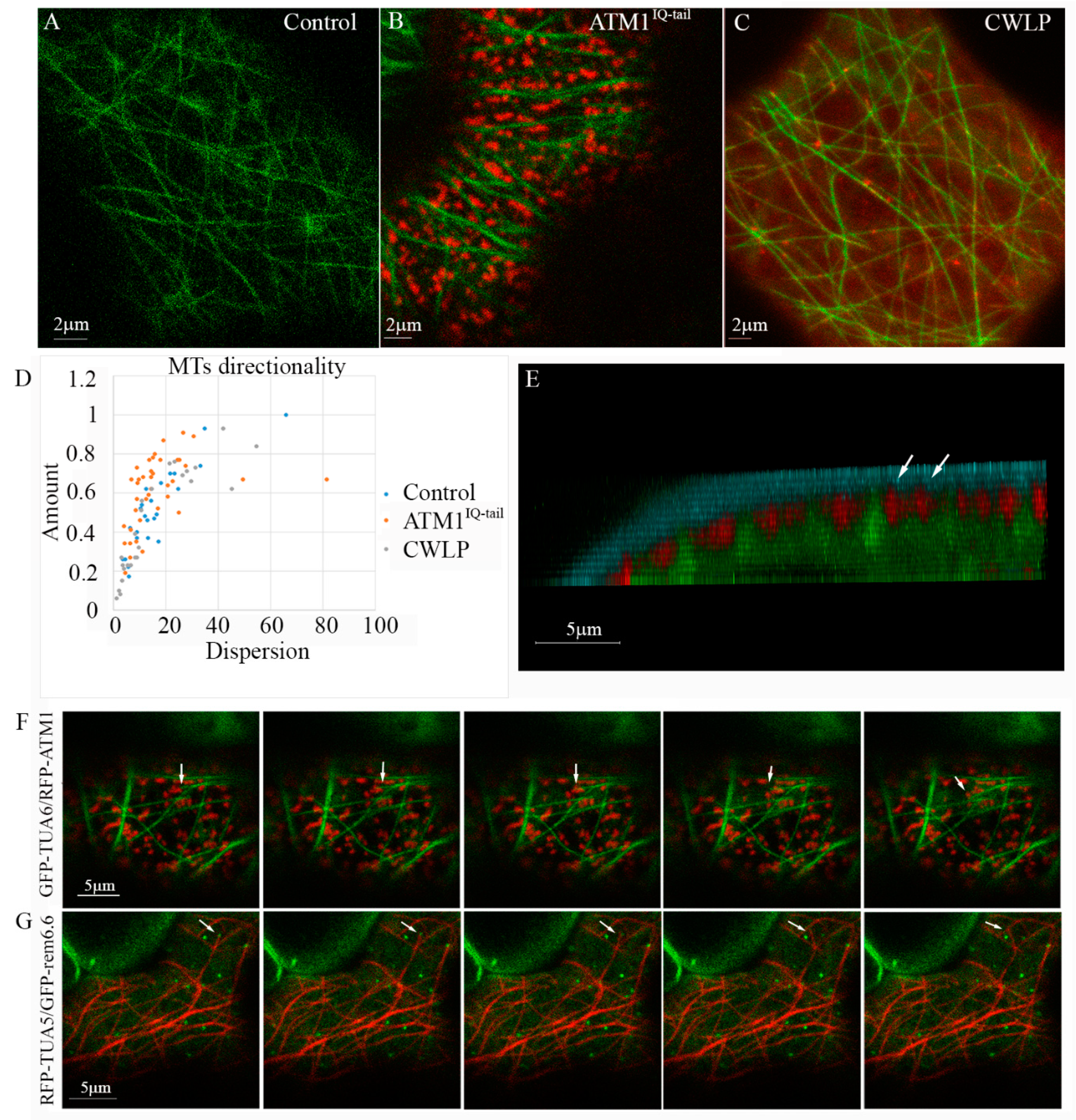
3.1. Coordinated Positioning of Cortical MTs and ATM1IQ-tail
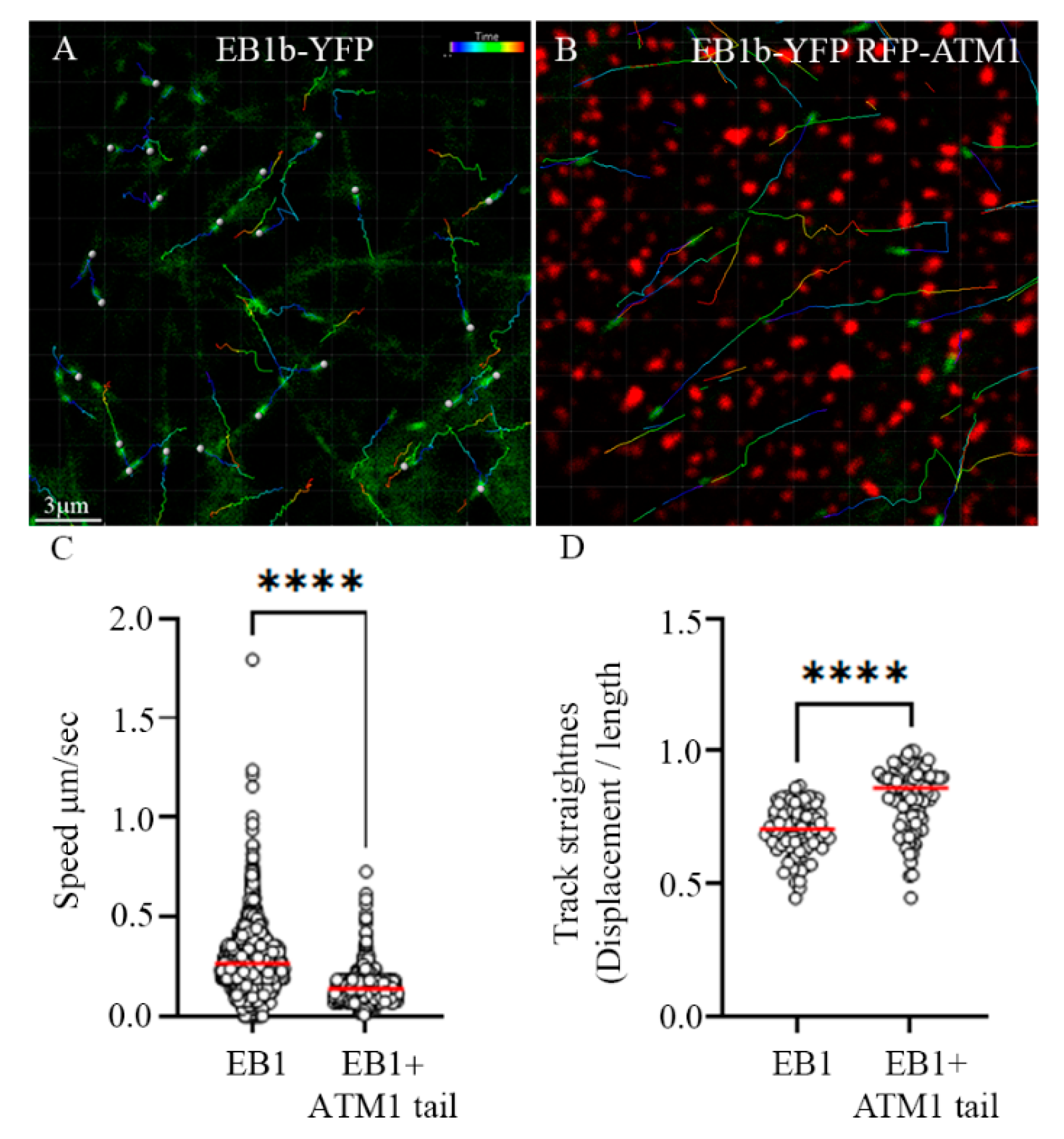
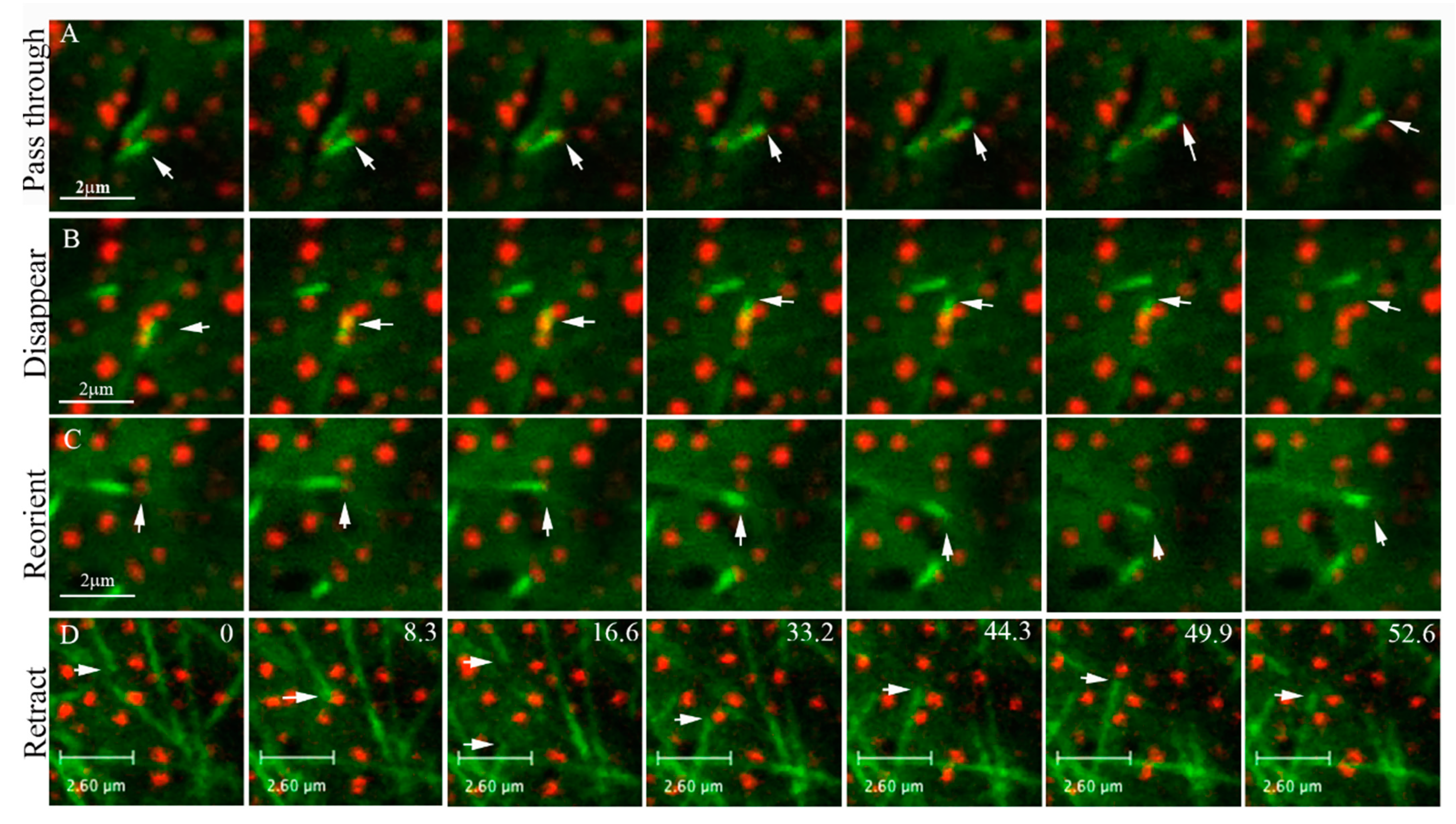
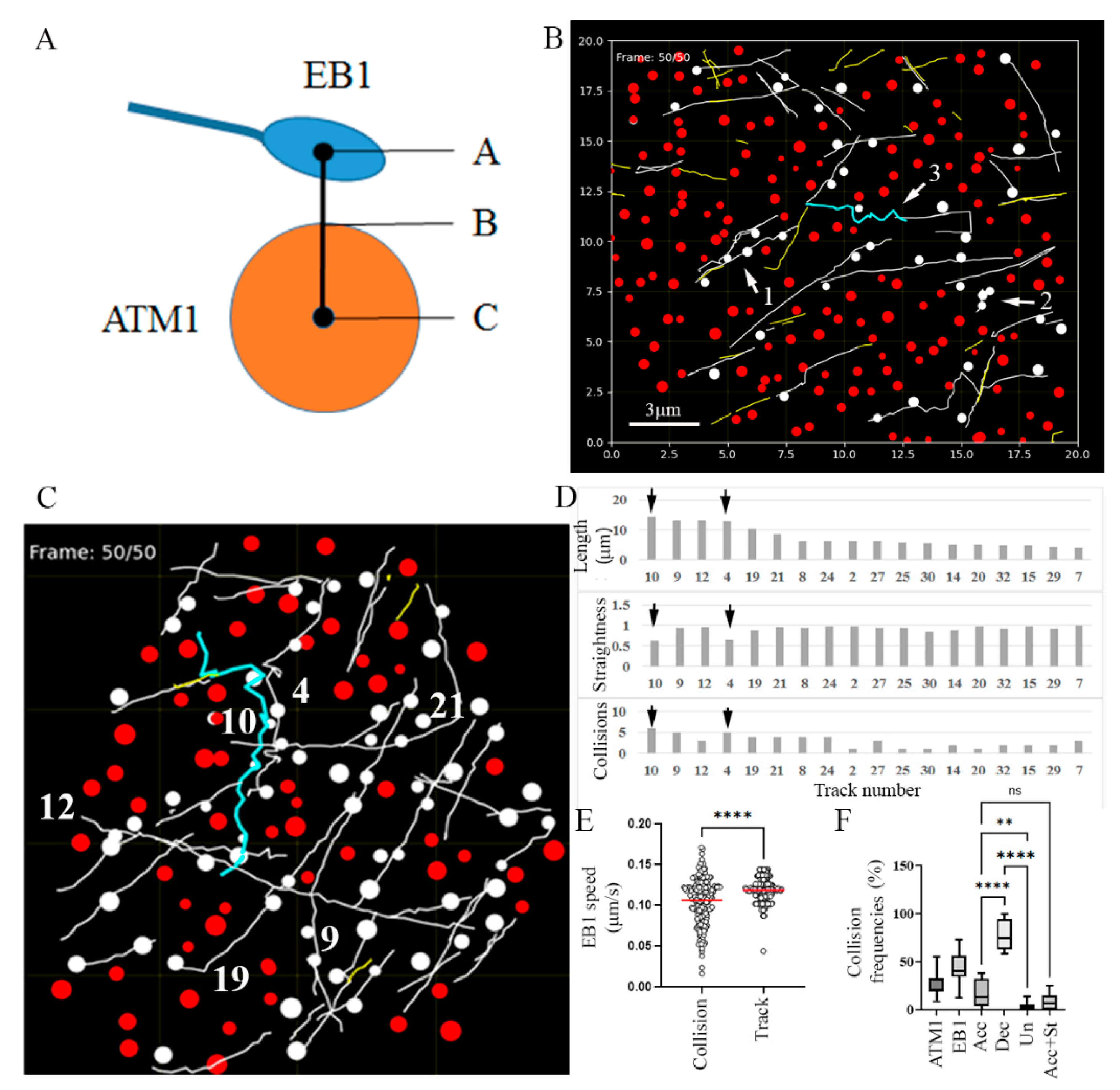
3.2. Collision of MTs with ATM1IQ-tail Clusters Can Result in Cross Over, Reduced Elongation Rate, Detachment and Reorientation or Retraction
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wasteneys, G.O. Microtubule organization in the green kingdom: Chaos or self-order? J. Cell Sci. 2002, 115, 1345–1354. [Google Scholar] [CrossRef]
- Elliott, A.; Shaw, S.L. Update: Plant Cortical Microtubule Arrays. Plant Physiol. 2018, 176, 94–105. [Google Scholar] [CrossRef] [PubMed]
- Hardham, A.R.; Gunning, B.E. Structure of cortical microtubule arrays in plant cells. J. Cell Biol. 1978, 77, 14–34. [Google Scholar] [CrossRef] [PubMed]
- Ledbetter, M.C.; Porter, K.R. A “microtubule” in plant cell fine structure. J. Cell Biol. 1963, 19, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.; Calder, G.M.; Doonan, J.H.; Lloyd, C.W. EB1 reveals mobile microtubule nucleation sites in Arabidopsis. Nat. Cell Biol. 2003, 5, 967–971. [Google Scholar] [CrossRef] [PubMed]
- Shaw, S.L.; Kamyar, R.; Ehrhardt, D.W. Sustained microtubule treadmilling in Arabidopsis cortical arrays. Science 2003, 300, 1715–1718. [Google Scholar] [CrossRef] [PubMed]
- Crowell, E.F.; Bischoff, V.; Desprez, T.; Rolland, A.; Stierhof, Y.D.; Schumacher, K.; Gonneau, M.; Hofte, H.; Vernhettes, S. Pausing of Golgi bodies on microtubules regulates secretion of cellulose synthase complexes in Arabidopsis. Plant Cell 2009, 21, 1141–1154. [Google Scholar] [CrossRef]
- Gutierrez, R.; Lindeboom, J.J.; Paredez, A.R.; Emons, A.M.; Ehrhardt, D.W. Arabidopsis cortical microtubules position cellulose synthase delivery to the plasma membrane and interact with cellulose synthase trafficking compartments. Nat. Cell Biol. 2009, 11, 797–806. [Google Scholar] [CrossRef]
- Paredez, A.R.; Somerville, C.R.; Ehrhardt, D.W. Visualization of Cellulose Synthase Demonstrates Functional Association with Microtubules. Science 2006, 312, 1491–1495. [Google Scholar] [CrossRef]
- Ehrhardt, D.W.; Shaw, S.L. Microtubule dynamics and organization in the plant cortical array. Ann. Rev. Plant Biol. 2006, 57, 859–875. [Google Scholar] [CrossRef]
- Dixit, R.; Cyr, R. Encounters between dynamic cortical microtubules promote ordering of the cortical array through angle-dependent modifications of microtubule behavior. Plant Cell 2004, 16, 3274–3284. [Google Scholar] [CrossRef]
- Wightman, R.; Turner, S.R. Severing at sites of microtubule crossover contributes to microtubule alignment in cortical arrays. Plant J. 2007, 52, 742–751. [Google Scholar] [CrossRef] [PubMed]
- Lindeboom, J.J.; Nakamura, M.; Hibbel, A.; Shundyak, K.; Gutierrez, R.; Ketelaar, T.; Emons, A.M.; Mulder, B.M.; Kirik, V.; Ehrhardt, D.W. A Mechanism for Reorientation of Cortical Microtubule Arrays Driven by Microtubule Severing. Science 2013, 342, 1245533. [Google Scholar] [CrossRef]
- Zhang, Q.; Fishel, E.; Bertroche, T.; Dixit, R. Microtubule severing at crossover sites by katanin generates ordered cortical microtubule arrays in Arabidopsis. Curr. Biol. 2013, 23, 2191–2195. [Google Scholar] [CrossRef] [PubMed]
- Gardiner, J.C.; Harper, J.D.; Weerakoon, N.D.; Collings, D.A.; Ritchie, S.; Gilroy, S.; Cyr, R.J.; Marc, J. A 90-kD phospholipase D from tobacco binds to microtubules and the plasma membrane. Plant Cell 2001, 13, 2143–2158. [Google Scholar] [CrossRef]
- Kesten, C.; Wallmann, A.; Schneider, R.; McFarlane, H.E.; Diehl, A.; Khan, G.A.; van Rossum, B.-J.; Lampugnani, E.R.; Szymanski, W.G.; Cremer, N.; et al. The companion of cellulose synthase 1 confers salt tolerance through a Tau-like mechanism in plants. Nat. Commun. 2019, 10, 857. [Google Scholar] [CrossRef]
- Danek, M.; Angelini, J.; Malinska, K.; Andrejch, J.; Amlerova, Z.; Kocourkova, D.; Brouzdova, J.; Valentova, O.; Martinec, J.; Petrasek, J. Cell wall contributes to the stability of plasma membrane nanodomain organization of Arabidopsis thaliana FLOTILLIN2 and HYPERSENSITIVE INDUCED REACTION1 proteins. Plant J. 2020, 101, 619–636. [Google Scholar] [CrossRef]
- Martiniere, A.; Gayral, P.; Hawes, C.; Runions, J. Building bridges: Formin1 of Arabidopsis forms a connection between the cell wall and the actin cytoskeleton. Plant J. 2011, 66, 354–365. [Google Scholar] [CrossRef]
- Oda, Y.; Fukuda, H. Initiation of cell wall pattern by a Rho- and microtubule-driven symmetry breaking. Science 2012, 337, 1333–1336. [Google Scholar] [CrossRef]
- Kusumi, A.; Fujiwara, T.K.; Chadda, R.; Xie, M.; Tsunoyama, T.A.; Kalay, Z.; Kasai, R.S.; Suzuki, K.G. Dynamic organizing principles of the plasma membrane that regulate signal transduction: Commemorating the fortieth anniversary of Singer and Nicolson’s fluid-mosaic model. Ann. Rev. Cell Dev. Biol. 2012, 28, 215–250. [Google Scholar] [CrossRef] [PubMed]
- Grecco, H.E.; Schmick, M.; Bastiaens, P.I. Signaling from the living plasma membrane. Cell 2011, 144, 897–909. [Google Scholar] [CrossRef]
- Avisar, D.; Abu-Abied, M.; Belausov, E.; Sadot, E.; Hawes, C.; Sparkes, I.A. A comparative study of the involvement of 17 Arabidopsis myosin family members on the motility of Golgi and other organelles. Plant Physiol. 2009, 150, 700–709. [Google Scholar] [CrossRef]
- Golomb, L.; Abu-Abied, M.; Belausov, E.; Sadot, E. Different subcellular localizations and functions of Arabidopsis myosin VIII. BMC Plant Biol. 2008, 8, 3. [Google Scholar] [CrossRef]
- Haraguchi, T.; Tominaga, M.; Matsumoto, R.; Sato, K.; Nakano, A.; Yamamoto, K.; Ito, K. Molecular Characterization and Subcellular Localization of Arabidopsis class VIII myosin, ATM1. J. Biol. Chem. 2014, 289, 12343–12355. [Google Scholar] [CrossRef]
- Wu, S.Z.; Bezanilla, M. Myosin VIII associates with microtubule ends and together with actin plays a role in guiding plant cell division. eLife 2014, 3, 1–20. [Google Scholar] [CrossRef]
- Reichelt, S.; Knight, A.E.; Hodge, T.P.; Baluška, F.; Samaj, J.; Volkmann, D.; Kendrick-Jones, J. Characterization of the unconventional myosin VIII in plant cells and its localization at the post-cytokinetic cell wall. Plant J. 1999, 19, 555–567. [Google Scholar] [CrossRef]
- Volkmann, D.; Mori, T.; Tirlapur, U.K.; König, K.; Fujiwara, T.; Kendrick-Jones, J.; Baluška, F. Unconventional myosins of the plant-specific class VIII: Endocytosis, cytokinesis, plasmodesmata/pit-fields, and cell-to-cell coupling. Cell Biol. Int. 2003, 27, 289–291. [Google Scholar] [CrossRef]
- Van Damme, D.; Bouget, F.Y.; Van Poucke, K.; Inze, D.; Geelen, D. Molecular dissection of plant cytokinesis and phragmoplast structure: A survey of GFP-tagged proteins. Plant J. 2004, 40, 386–398. [Google Scholar] [CrossRef]
- Baluška, F.; Barlow, P.W.; Volkmann, D. Actin and Myosin VIII in developing root cells. In Actin: A Dynamic Framework for Multiple Plant Cell Functions; Staiger, C.J., Baluska, F., Volkmann, D., Barlow, P.W., Eds.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2000; pp. 457–476. [Google Scholar]
- Baluška, F.; Cvrčková, F.; Kendrick-Jones, J.; Volkmann, D. Sink plasmodesmata as gateways for phloem unloading. Myosin VIII and calreticulin as molecular determinants of sink strength? Plant Physiol. 2001, 126, 39–46. [Google Scholar] [CrossRef]
- Avisar, D.; Prokhnevsky, A.I.; Dolja, V.V. Class VIII myosins are required for plasmodesmatal localization of a closterovirus Hsp70 homolog. J. Virol. 2008, 82, 2836–2843. [Google Scholar] [CrossRef][Green Version]
- Yuan, Z.; Chen, H.; Chen, Q.; Omura, T.; Xie, L.; Wu, Z.; Wei, T. The early secretory pathway and an actin-myosin VIII motility system are required for plasmodesmatal localization of the NSvc4 protein of Rice stripe virus. Virus Res. 2011, 159, 62–68. [Google Scholar] [CrossRef] [PubMed]
- Henn, A.; Sadot, E. The unique enzymatic and mechanistic properties of plant myosins. Curr. Opin. Plant Biol. 2014, 22, 65–70. [Google Scholar] [CrossRef] [PubMed]
- Rula, S.; Suwa, T.; Kijima, S.T.; Haraguchi, T.; Wakatsuki, S.; Sato, N.; Duan, Z.; Tominaga, M.; Uyeda, T.Q.P.; Ito, K. Measurement of enzymatic and motile activities of Arabidopsis myosins by using Arabidopsis actins. Biochem. Biophys. Res. Commun. 2018, 495, 2145–2151. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.Z.; Bezanilla, M. Actin and microtubule cross talk mediates persistent polarized growth. J. Cell Biol. 2018, 217, 3531–3544. [Google Scholar] [CrossRef]
- Stein, H.; Honig, A.; Miller, G.; Erster, O.; Eilenberg, H.; Csonka, L.N.; Szabados, L.; Koncz, C.; Zilberstein, A. Elevation of free proline and proline-rich protein levels by simultaneous manipulations of proline biosynthesis and degradation in plants. Plant Sci. 2011, 181, 140–150. [Google Scholar] [CrossRef]
- Jarsch, I.K.; Konrad, S.S.; Stratil, T.F.; Urbanus, S.L.; Szymanski, W.; Braun, P.; Braun, K.H.; Ott, T. Plasma Membranes Are Subcompartmentalized into a Plethora of Coexisting and Diverse Microdomains in Arabidopsis and Nicotiana benthamiana. Plant Cell 2014, 26, 1698–1711. [Google Scholar] [CrossRef]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef]
- Seetharaman, S.; Etienne-Manneville, S. Microtubules at focal adhesions—A double-edged sword. J. Cell Sci. 2019, 132, jcs232843. [Google Scholar] [CrossRef]
- Krylyshkina, O.; Anderson, K.I.; Kaverina, I.; Upmann, I.; Manstein, D.J.; Small, J.V.; Toomre, D.K. Nanometer targeting of microtubules to focal adhesions. J. Cell Biol. 2003, 161, 853–859. [Google Scholar] [CrossRef]
- Konrad, S.S.; Popp, C.; Stratil, T.F.; Jarsch, I.K.; Thallmair, V.; Folgmann, J.; Marin, M.; Ott, T. S-acylation anchors remorin proteins to the plasma membrane but does not primarily determine their localization in membrane microdomains. New Phytol. 2014, 203, 758–769. [Google Scholar] [CrossRef]
- Gillespie, T.; Boevink, P.; Haupt, S.; Roberts, A.G.; Toth, R.; Valentine, T.; Chapman, S.; Oparka, K.J. Functional analysis of a DNA-shuffled movement protein reveals that microtubules are dispensable for the cell-to-cell movement of tobacco mosaic virus. Plant Cell 2002, 14, 1207–1222. [Google Scholar] [CrossRef]
- Chan, J.; Calder, G.; Fox, S.; Lloyd, C. Localization of the microtubule end binding protein EB1 reveals alternative pathways of spindle development in Arabidopsis suspension cells. Plant Cell 2005, 17, 1737–1748. [Google Scholar] [CrossRef] [PubMed]
- Maurer, S.P.; Cade, N.I.; Bohner, G.; Gustafsson, N.; Boutant, E.; Surrey, T. EB1 accelerates two conformational transitions important for microtubule maturation and dynamics. Curr. Biol. 2014, 24, 372–384. [Google Scholar] [CrossRef]
- Demir, F.; Horntrich, C.; Blachutzik, J.O.; Scherzer, S.; Reinders, Y.; Kierszniowska, S.; Schulze, W.X.; Harms, G.S.; Hedrich, R.; Geiger, D.; et al. Arabidopsis nanodomain-delimited ABA signaling pathway regulates the anion channel SLAH3. Proc. Natl. Acad. Sci. USA 2013, 110, 8296–8301. [Google Scholar] [CrossRef] [PubMed]
- Haney, C.H.; Long, S.R. Plant flotillins are required for infection by nitrogen-fixing bacteria. Proc. Natl. Acad. Sci. USA 2010, 107, 478–483. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Liu, P.; Wan, Y.; Chen, T.; Wang, Q.; Mettbach, U.; Baluska, F.; Samaj, J.; Fang, X.; Lucas, W.J.; et al. A membrane microdomain-associated protein, Arabidopsis Flot1, is involved in a clathrin-independent endocytic pathway and is required for seedling development. Plant Cell 2012, 24, 2105–2122. [Google Scholar] [CrossRef]
- Raffaele, S.; Bayer, E.; Lafarge, D.; Cluzet, S.; German Retana, S.; Boubekeur, T.; Leborgne-Castel, N.; Carde, J.P.; Lherminier, J.; Noirot, E.; et al. Remorin, a solanaceae protein resident in membrane rafts and plasmodesmata, impairs potato virus X movement. Plant Cell 2009, 21, 1541–1555. [Google Scholar] [CrossRef] [PubMed]
- Gouguet, P.; Gronnier, J.; Legrand, A.; Perraki, A.; Jolivet, M.D.; Deroubaix, A.F.; German-Retana, S.; Boudsocq, M.; Habenstein, B.; Mongrand, S.; et al. Connecting the dots: From nanodomains to physiological functions of REMORINs. Plant Physiol. 2021, 185, 632–649. [Google Scholar] [CrossRef] [PubMed]
- Bürstenbinder, K.; Möller, B.; Plötner, R.; Stamm, G.; Hause, G.; Mitra, D.; Abel, S. The IQD Family of Calmodulin-Binding Proteins Links Calcium Signaling to Microtubules, Membrane Subdomains, and the Nucleus. Plant Physiol. 2017, 173, 1692–1708. [Google Scholar] [CrossRef]
- Phillips, R.; Ursell, T.; Wiggins, P.; Sens, P. Emerging roles for lipids in shaping membrane-protein function. Nature 2009, 459, 379–385. [Google Scholar] [CrossRef]
- Ambrose, J.C.; Wasteneys, G.O. CLASP modulates microtubule-cortex interaction during self-organization of acentrosomal microtubules. Mol. Biol. Cell 2008, 19, 4730–4737. [Google Scholar] [CrossRef]
- Allard, J.F.; Wasteneys, G.O.; Cytrynbaum, E.N. Mechanisms of self-organization of cortical microtubules in plants revealed by computational simulations. Mol. Biol. Cell 2010, 21, 278–286. [Google Scholar] [CrossRef]
- Bidhendi, A.J.; Altartouri, B.; Gosselin, F.P.; Geitmann, A. Mechanical Stress Initiates and Sustains the Morphogenesis of Wavy Leaf Epidermal Cells. Cell Rep. 2019, 28, 1237–1250. [Google Scholar] [CrossRef]
- Hamant, O.; Inoue, D.; Bouchez, D.; Dumais, J.; Mjolsness, E. Are microtubules tension sensors? Nat. Commun. 2019, 10, 2360. [Google Scholar] [CrossRef] [PubMed]
- Inoue, D.; Nitta, T.; Kabir, A.M.R.; Sada, K.; Gong, J.P.; Konagaya, A.; Kakugo, A. Sensing surface mechanical deformation using active probes driven by motor proteins. Nat. Commun. 2016, 7, 12557. [Google Scholar] [CrossRef] [PubMed]
- Geiger, B.; Avnur, Z.; Rinnerthaler, G.; Hinssen, H.; Small, V.J. Microfilament-organizing centers in areas of cell contact: Cytoskeletal interactions during cell attachment and locomotion. J. Cell Biol. 1984, 99, 83s–91s. [Google Scholar] [CrossRef]
- Rinnerthaler, G.; Geiger, B.; Small, J.V. Contact formation during fibroblast locomotion: Involvement of membrane ruffles and microtubules. J. Cell Biol. 1988, 106, 747–760. [Google Scholar] [CrossRef]
- Efimov, A.; Schiefermeier, N.; Grigoriev, I.; Ohi, R.; Brown, M.C.; Turner, C.E.; Small, J.V.; Kaverina, I. Paxillin-dependent stimulation of microtubule catastrophes at focal adhesion sites. J. Cell Sci. 2008, 121, 196–220. [Google Scholar] [CrossRef]
- Zamir, E.; Geiger, B. Molecular complexity and dynamics of cell-matrix adhesions. Cell Sci. 2001, 114, 3583–3590. [Google Scholar] [CrossRef]
- Byron, A.; Askari, J.A.; Humphries, J.D.; Jacquemet, G.; Koper, E.J.; Warwood, S.; Choi, C.K.; Stroud, M.J.; Chen, C.S.; Knight, D.; et al. A proteomic approach reveals integrin activation state-dependent control of microtubule cortical targeting. Nat. Commun. 2015, 6, 6135. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bar-Sinai, S.; Belausov, E.; Dwivedi, V.; Sadot, E. Collisions of Cortical Microtubules with Membrane Associated Myosin VIII Tail. Cells 2022, 11, 145. https://doi.org/10.3390/cells11010145
Bar-Sinai S, Belausov E, Dwivedi V, Sadot E. Collisions of Cortical Microtubules with Membrane Associated Myosin VIII Tail. Cells. 2022; 11(1):145. https://doi.org/10.3390/cells11010145
Chicago/Turabian StyleBar-Sinai, Sefi, Eduard Belausov, Vikas Dwivedi, and Einat Sadot. 2022. "Collisions of Cortical Microtubules with Membrane Associated Myosin VIII Tail" Cells 11, no. 1: 145. https://doi.org/10.3390/cells11010145
APA StyleBar-Sinai, S., Belausov, E., Dwivedi, V., & Sadot, E. (2022). Collisions of Cortical Microtubules with Membrane Associated Myosin VIII Tail. Cells, 11(1), 145. https://doi.org/10.3390/cells11010145






