Abstract
Eukaryotic genomes are packaged into high-order chromatin structures organized in discrete territories inside the cell nucleus, which is surrounded by the nuclear envelope acting as a barrier. This chromatin organization is complex and dynamic and, thus, determining the spatial and temporal distribution and folding of chromosomes within the nucleus is critical for understanding the role of chromatin topology in genome function. Primarily focusing on the regulation of gene expression, we review here how the genome of Drosophila melanogaster is organized into the cell nucleus, from small scale histone–DNA interactions to chromosome and lamina interactions in the nuclear space.
1. Introduction
In eukaryotic cells, nuclear organization refers to the spatial distribution of nuclear contents and components. The cell nucleus contains DNA, which is organized as multiple long linear molecules in a complex with a large variety of proteins, such as histones, to form chromatin and chromosomes. Thus, the eukaryotic genome is packaged into higher-order chromatin structures and organized in a manner that accommodates highly dynamic processes such as DNA replication, gene transcription, and DNA repair.
There are many different levels of nuclear organization and whether they affect gene function or just reflect this function is still unclear. Here, we focus on Drosophila melanogaster, a pre-eminent animal model system for genetic studies. Starting from basic DNA composition, we review what is currently known about the general structure of chromatin, chromosomes, and nuclear organization.
2. The Drosophila Genome
The first annotated whole genome sequence of the fruit fly Drosophila was published more than two decades ago, when it was estimated to contain around 13,600 genes [1,2]. The most recent version of the Drosophila genome identifies 13,969 protein-coding genes and 2545 long non-coding RNA genes, with a GC percentage of ~42% [3] (FlyBase r6.40, June 2021).
The Drosophila genome is divided into four chromosomes: the X and Y sex chromosomes, the autosomal chromosomes 2 and 3, and a tiny chromosome 4 containing no more than 100 genes and known as the “dot chromosome”. In Drosophila, like in other Diptera species, polytene chromosomes can be observed in the interphase nuclei of certain tissues such as the salivary glands. This highly specialized form of chromosomes develops by endoreduplication of the chromosomes of diploid nuclei, producing multiple chromatids of each chromosome. Polytene chromosomes have been very useful in cytogenetic studies due to their distinct patterns of bands and interbands showing different degrees of condensation, gene expression profiles, and protein composition [4,5].
3. Chromatin Composition and Structure
Eukaryotic DNA molecules, together with proteins and RNA, are packaged into a compact structure called chromatin. Different chromatin states have long been recognized, with chromatin classically divided into euchromatin (which decondenses regularly during the cell cycle, consists primarily of single-copy sequences, and is transcriptionally active) and heterochromatin (which is condensed throughout the cell cycle, consists mainly of repetitive sequences, and can silence gene expression) [6,7]. In Drosophila, heterochromatin comprises approximately a third of the genome and is organized primarily into pericentromeric and telomeric blocks [8]. Pericentromeric heterochromatin is mainly composed of repetitive sequences, including fragments of various transposable elements (TEs) and satellite DNAs (satDNA), which are large blocks of tandemly repeated DNA sequences [9]. Heterochromatin protein 1 (HP1a in Drosophila) is a conserved eukaryotic chromosomal protein that is associated with pericentromeric heterochromatin and mediates the concomitant gene silencing [10]. Despite its association with gene repression, it was reported recently that a significant part of the constitutive heterochromatin in Drosophila is, in fact, occupied by active genes [11]. Moreover, an RNAi screen conducted in flies revealed that heterochromatin is structurally complex and contains many dynamic smaller subdomains [12]. Beyond the binary classification of chromatin into euchromatin and heterochromatin, several groups have partitioned the Drosophila genome into different chromatin types or states based on a combinatorial signature of bound proteins, histone modifications and integrative analysis with other chromatin data [13] (discussed below).
3.1. Nucleosome Dynamics
The basic unit of chromatin is the nucleosome, which consists of an octamer composed of two copies of each of the core histones (H2A, H2B, H3, and H4) that is wrapped by 145–147 bp of DNA in a left-handed superhelical turn [14,15]. The core histones interact with DNA through the highly conserved histone-fold domain, while the N-terminal tail participates in nucleosome stabilization [15,16]. The different nucleosomes are separated by linker DNA and the resulting arrangement is an 11-nm chromatin fiber that resembles a beads-on-a-string structure [17]. Linker histones, such as H1, bind to DNA at the entry/exit site of the nucleosome, seal the structure, and protect an extra 20 bp of DNA [18,19,20]. The resulting structure is called a “chromatosome” [21]. Several studies have addressed the role of H1 in chromatin folding, showing that H1 promotes and stabilizes the compaction of nucleosomes into a 30-nm chromatin fiber [22,23]. However, the 30-nm fiber is only observed as short fragments in vivo, since nucleosomes are found in clutches of various sizes separated by nucleosome-depleted regions [24].
The organization of nucleosomes varies across the genome and plays a central role in controlling DNA accessibility. Nucleosome-depleted regions are characteristic of active chromatin sites and adjacent regions show a regular placement of nucleosomes. More irregular positions are commonly found elsewhere [25,26,27]. The determinants of nucleosome positioning were defined some years ago as a combination of DNA sequences, ATP-dependent chromatin remodeling enzymes, transcription factors (TFs), and elongating RNA polymerase II (RNAPII) [28]. A comparative analysis between Drosophila cell lines identified genomic regions that exhibited cell line-specific nucleosome enrichment or depletion. The same study revealed that nucleosomes were positioned in accordance with previously known DNA–nucleosome interactions, with helically repeating A/T di-nucleotide pairs arranged within nucleosomal DNA and AT-rich pentamers generally excluded from nucleosomal DNA [29].
Nucleosomes are highly dynamic structures and the partial unwrapping of DNA from the octamer leads to exposure of the different regions for protein recognition [30,31]. Nucleosome dynamics are controlled by a complex cooperation between different histones, histone post-translational modifications, nucleosome occupancy and positioning [32,33]. In recent years, the development of genome-wide mapping approaches has provided a vast amount of information regarding the genomic location of chromatin-associated proteins, such as histones, as well as of several of their modifications. A large number of these modifications have been implicated in the regulation of gene expression, as we will discuss below.
3.2. Core Histones and Their Variants
Drosophila contains five canonical histones (H1, H2A, H2B, H3, and H4), usually referred to as replication-coupled histones since they are mostly incorporated during DNA replication. The canonical histone genes are clustered into a highly repeated unit that contains one copy of each gene, although H4 is also encoded by another gene outside the cluster (H4r) [34].
In addition to the canonical histones, the fly genome encodes four histone variants (BigH1, H2Av, H3.3, and cenH3), with H2B and H4 being the only histones lacking variants. Histone variants confer different structural properties and carry out specialized functions in numerous processes [35]. The unique Drosophila H2A variant, H2Av, combines the features of the H2A.X and H2A.Z eukaryotic variants and has been linked to transcription, DNA repair, and heterochromatin [36]. H2Av may also have a role in chromosome organization, since the depletion of the machinery responsible for its incorporation results in the alteration of chromosome structure in salivary glands and S2 cells [37,38]. H2Av is broadly distributed in the Drosophila genome and nucleosomes with H2Av are particularly enriched downstream of the transcription start site (TSS) of active genes, which correlates positively with transcription levels [25,39]. It has been proposed that H2Av may facilitate the progression of RNAPII, since a reduction of H2Av levels results in an increase in RNAPII stalling [40]. In addition, H2Av has been implicated in gene silencing through the Polycomb group (PcG) of proteins. H2Av seems to participate in Polycomb (Pc) recruitment since Pc sites are lost in polytene chromosomes from H2Av mutants [41]. Moreover, H2Av is also found in heterochromatin [25,42] and HP1a binding may depend on the presence of H2Av [41].
The H3 replacement variant H3.3 is encoded by two genes, H3.3A and H3.3B, which are ubiquitously expressed throughout all the tissues and developmental stages [43]. H3.3 is usually enriched in active chromatin, such as active promoters and gene bodies of transcribed genes [44,45], and in sites with high nucleosome turnover rates [46]. However, clonally removing H3.3 in cells from the Drosophila wing disc does not affect gene expression [47]. Furthermore, H3.3 null mutant flies are viable and show no phenotypic alterations, except for infertility [47]. Interestingly, regions of H3.3 enrichment are generally depleted of H1 and knocking down H3.3 leads to increased H1 association at these sites [48].
3.3. Linker Histones
Drosophila contains only one somatic H1 [49] and one embryonic and germline-specific variant, BigH1 [50]. H1 is broadly distributed throughout the genome [48,51]. Its loss results in the misexpression of only a small subset of genes [52,53], mainly those located in heterochromatin [54,55]. The changes in gene expression include the upregulation of TEs [52,53,56]. In mammals, linker histones are also widely distributed [57,58,59,60] and, similar to flies, only a small number of genes are affected in triple knock-out mouse embryonic stem cells (ESCs) with 50% total H1 depletion [61] or in human cells with reduced H1 levels [59,62,63]. Moreover, these changes in gene expression mainly affect heterochromatic regions [57,59,62,64].
BigH1 is expressed in both male and female germlines with similar expression patterns [50,65,66]. While the function in the female germline is not known, in males, BigH1 is important for germ stem cell maintenance and spermatocyte differentiation [65]. BigH1 is retained in precellular embryos, where it is important in maintaining the silencing of the zygotic genome [50]. Recent studies have focused on the different properties of the two linker histones and have found that BigH1 has a greater repression ability than the somatic H1 due to its higher content in acidic residues [66]. Moreover, nucleosomes containing BigH1 are more stable, but show higher mobility than H1-nucleosomes [67].
4. Covalent Modifications of Chromatin
Covalent modifications play an essential role in the nucleosome–nucleosome interactions that dictate chromatin folding and dynamics. These modifications, occurring both in DNA and histones, have the potential to form a complex combinatorial regulatory system and are fundamental in regulating all processes that use DNA as a template, such as transcription, DNA repair, and replication.
4.1. DNA Methylation
Methylation of the carbon C5 of cytosine to form 5-methylcytosine (5mC) is probably the best-known modification of DNA in eukaryotes. Despite the general role 5mC plays in the repression of vertebrate gene expression, the situation may be different for invertebrates. There is evidence in favor of cytosine C5 methylation in Drosophila, although its source is still elusive [68]. Although 5mC is rare, methylation on N6 adenine (6mA) seems prevalent in Caenorhabditis elegans and Drosophila [69,70]. Recent studies have confirmed that NMAD-1 (MT-A70 family) and DMAD (DNA 6mA demethylase, TET ortholog) are 6mA demethylases in C. elegans and Drosophila respectively [71]. The Drosophila DMAD regulates 6mA levels during embryo development and oogenesis. DNA immunoprecipitation studies in ovaries from DMAD mutants show that 6mA is enriched in transposon regions and seems to promote their expression [70]. More recently, He and coworkers (2019) have shown that, in Drosophila embryos, 6 mA is found not only in transposon regions but also in zygotic genes. This modification can be read by the TF Jumu, which controls the activation of the zygotic genome (ZGA) through the regulation of zelda, among other genes [72].
4.2. Histone Modifications
Several residues in histones are susceptible to modification. The most well studied modifications are the ones occurring at the N-terminal tails of the core histones, although modifications of the globular domains have been gaining more attention [73,74]. The N-terminal tails of the core histones protrude outside the nucleosome and contact with adjacent nucleosomes. Thus, modifications in this region can directly affect nucleosome–nucleosome interactions and alter the chromatin structure. This is the case for H4K16 and H4K20. While acetylation of H4K16 reduces the level of compaction in vitro [75,76], the di- and tri-methylation of H4K20 has the opposite effect and enhances chromatin condensation in vitro [77]. However, modifications at the N-terminal tails also mediate the recruitment of effector proteins [78]. Recognition of these modifications is achieved through specialized domains present in “reader” proteins, which can be remodeling complexes, other modifying enzymes or scaffolds of the transcription machinery [78]. Some of the effector proteins and the enzymes that catalyze or remove these modifications are part of the PcG or Trithorax group (TrxG), which are chromatin-modifying complexes implicated in the maintenance of repressed or active gene expression states [79].
There are some modifications that correlate with gene activity [80]. The clearest example is the acetylation of histones H3 and H4, which is associated with active transcription. Acetylation of lysine (K) partially neutralizes the positive charge of histones, thus weakening the interaction between histones and DNA [81]. Methylation is, by far, more complex, and its correlation with gene expression depends not only on which amino acid is modified, but also on the degree of methylation (mono, di or tri). The hypothesis of the “histone code” proposes that the combination of different modifications is important in regulating gene expression and other DNA processes [82]. Recent studies, however, challenge the instructive role of histone modifications and suggest that those modifications traditionally associated with active genes do not directly trigger transcriptional activation [83]. Indeed, typical histone modifications only have a few roles in regulating transcription [84]. A study using different developmental time points showed that the transcription of genes temporally regulated during fly and worm development occurred in the absence of canonically active histone modifications [85]. Similarly, another study using genetic approaches and mutant derivatives found that transcriptional regulation can occur in the complete absence of H3K4 methylation [86]. Likewise, the depletion of H3K27ac in mouse ESCs does not alter chromatin accessibility or transcription, indicating that this modification is dispensable for enhancer activity in mouse ESCs [87]. These modifications might instead be necessary for sustained transcription, since there is a correlation between the amount of these modifications and the stability of expression [85].
Another example of transcription without the typical active histone modifications is the first wave of transcription of the ZGA, which is characterized by the enrichment of H4K8ac, H3K18ac, and H3K27ac in active genes [88]. Other modifications commonly associated with active genes, such as H3K4me3, H3K9ac, H3K36me3, and H3K4me1, do not become enriched until mitotic cycle 14 [88,89], indicating again that these modifications are not required for transcriptional competence, at least during the first wave of transcription. Recently, H3K14ac was shown to be important for the transcription of active genes that lack H3K9ac, H3K27ac, and H3K4me3 during Drosophila embryo development and in imaginal wing discs [90]. Moreover, modifications in the globular domain may have a role in regulating gene expression and other processes, since mutations of the H3 residues K56, K115, K122, T80, and T118 induce lethality at different developmental stages in Drosophila [91].
5. Functional Organization of the Genome
5.1. Segmentation of the Genome into Chromatin States
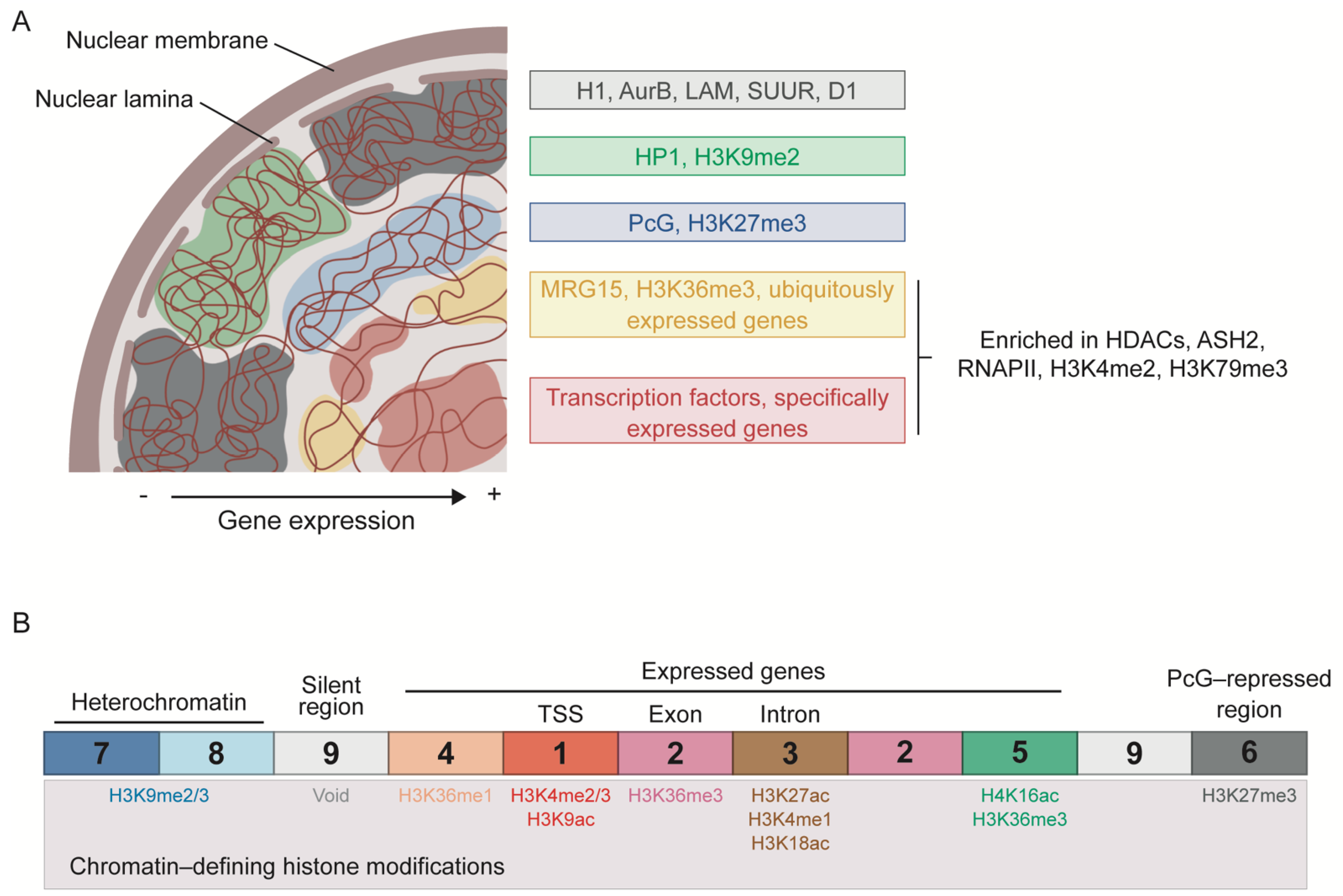
As mentioned above, several groups have attempted to classify chromatin into different types. Filion et al. (2010) proposed the segmentation of the genome of Drosophila cells into five types of chromatin based on genome-wide binding maps of selected chromatin components, with histone modification profiles used for independent validation (Figure 1A). These chromatin states, labeled as GREEN, BLUE, BLACK, RED, and YELLOW, are distributed throughout the genome in discrete domains with a length usually ranging from ~1 to 52 kb. The GREEN and BLUE chromatin types correspond to classic heterochromatic regions and are characterized by the binding of HP1 and HP1-interacting proteins or of PcG proteins, respectively. While GREEN chromatin is marked with H3K9me2 and usually corresponds to pericentromeric regions, BLUE chromatin is highly enriched in H3K27me3 and developmentally regulated genes. The last type of silent chromatin is BLACK chromatin, which covers about half of the genome and tends to have longer domains. BLACK chromatin is thus the predominant type and is characterized by being poor in genes and producing very low levels of mRNA. Interestingly, the genes within this type of chromatin can become active in specific cell types or tissues, suggesting that BLACK domains can be remodeled into a different chromatin type during development. Moreover, some of the proteins that mark this type of chromatin are histone H1, auroraB (AurB), Suppressor of Underreplication (SUUR), the AT-hook protein D1, and lamin (LAM), indicating a role of the nuclear lamina in the regulation of BLACK chromatin. The two remaining types of chromatin, RED and YELLOW, correspond to active regions and are characterized by the extensive binding of histone deacetylases (HDACs) and ASH2, as well as enrichment in H3K4me2, H3K79me3, and RNAPII. RED chromatin is also bound by several other proteins, including TFs such as the GAGA factor (GAF) and Jun-related antigen (JRA). Instead, YELLOW chromatin is uniquely marked by H3K36me3 and its reader protein MRG15. The genes located in RED and YELLOW chromatin are also different: the genes in RED chromatin have specific expression patterns and functions, such as signaling and TF activity, whereas those in YELLOW chromatin are ubiquitously expressed and have more general functions, such as DNA repair and metabolism [13].
Figure 1.
Classification of the chromatin landscape in Drosophila. (A) Chromatin segmentation into 5 types according to combinatorial protein binding. BLACK, GREEN, and BLUE chromatin types correspond to repressed and silenced domains, whereas YELLOW and RED chromatin types represent active regions. The more repressed regions tend to localize at the periphery of the nucleus, with BLACK and GREEN chromatin interacting with the nuclear lamina. Only the most characteristic components of each chromatin type are indicated. Based on Filion et al. [13]. (B) Division of chromatin into 9 states attending to histone modification patterns. States 1, 2, and 5 are associated with actively transcribed genes; states 3 and 4 with putative regulatory regions; state 6 with PcG-repressed regions; states 7 and 8 with heterochromatin; and state 9 corresponds to silent regions. The most enriched histone modifications are indicated for each chromatin state. Based on Karchkenko et al. [92].
Following a similar approach, Kharchenko et al. (2011) used histone modifications to determine nine distinct chromatin states in the Drosophila genome (Figure 1B). To functionally characterize these states, the authors integrate data from non-histone proteins, chromosome accessibility, transcription analyses, and short RNA production. In the 9-state model, transcriptionally active regions fall into more than two chromatin states, some of which can be observed at different regions of a particular gene. State 1 (red) is characterized by the enrichment of H3K4me3/me2 and H3K9ac, and is found at active promoters and TSSs. State 2 (purple) contains high levels of H3K36me3, an elongation mark enriched towards the 3′ end of the genes. State 3 (brown) is usually found within intronic regions and is enriched in H3K27ac, H3K4me1 and H3K18ac. Open chromatin regions are mainly associated with states 1 and 3, which are bound by different components of chromatin remodeling factors, such as NURF301 and MRG15 in the case of state 1, or SPT16 and dMI-2 in state 3. The authors propose a regulatory role for the state 3 domains, since they are also enriched in dCBP/p300 and almost half of them are bound by GAF and developmental TFs. State 4 (coral) resembles state 3, but lacks H3K27ac and is also marked by the presence of H3K36me1. Chromosome X is particularly enriched in state 5 (green), which is defined by high levels of H4K16ac and the modifications also present in state 2, probably reflecting a distinct mechanism of transcriptional regulation required for dosage compensation in male cells. State 6 (dark grey) is enriched in H3K27me3 and corresponds to PcG-repressed regions. Heterochromatic regions are depicted by state 7 (dark blue) and 8 (light blue), which are characterized by an enrichment in H3K9me2/m3, although levels are higher in state 7. Finally, the authors consider a last chromatin state characterized by the presence of low levels of the histone modifications considered in the study, the state 9 (light grey) [92].
These different types of chromatin have been related to physical domains of chromosome folding. For example, PcG-bound chromatin (BLUE chromatin or state 6) has been shown to form small subnuclear structures called PcG bodies or PcG-repressed domains [93,94]. As we discuss below, there is a strong link between chromatin activity and chromosome architecture.
5.2. 3D Organization of the Genome
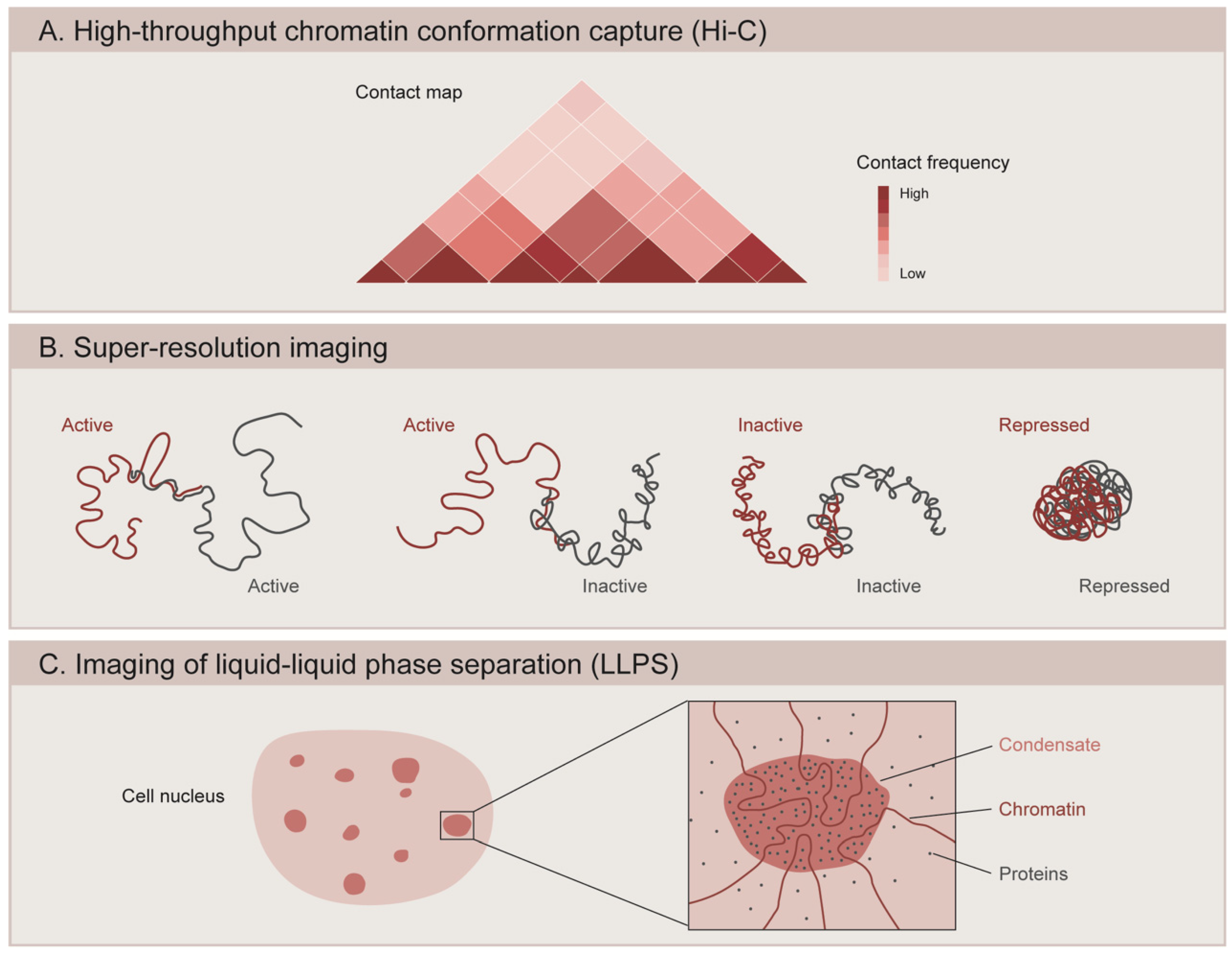
Technical advances such as the high-throughput derivative of chromosome conformation capture (Hi-C) have enabled the analysis of the three-dimensional architecture of genomes (Figure 2A) [95,96,97]. Chromatin interaction maps generated by Hi-C assays in Drosophila, mice, and humans have revealed that the genome is composed of several layers of structure that are organized in a hierarchical manner [98,99,100,101]. At the higher level of genome topology, chromatin is partitioned into two multimegabase compartments with distinct transcriptional activity: an active compartment (A) that is dense in expressed genes and correlates with histone modifications generally associated with active transcription, and an inactive compartment (B) that is gene-poor and heterochromatic [97,100]. The genome is further organized into sub-megabase domains called topologically associating domains (TADs), which are defined by a higher interaction frequency within the region than with those located outside of the TAD [98,102,103]. In mammals, TADs are likely to be formed from the active extrusion of chromatin loops mediated by the cohesin complex [104,105], and are insulated at the borders by the architectural chromatin protein CCCTC-binding factor (CTCF) [103,106]. Depletion of cohesin in mammalian cells results in the loss of the majority of TADs [107], while deleting CTCF sites or inverting their orientation reduces TAD insulation and facilitates crosstalk between adjacent TADs [108,109,110].
Figure 2.
Methods to study the 3D organization of the genome. (A) High-throughput chromatin conformation capture (Hi-C) generates contact maps that represent the interaction frequency between genomic loci. Studies using Hi-C have revealed that chromatin is organized into topologically associating domains (TADs). (B) Super-resolution microscopy is used to image the spatial organization of different chromatin domains: transcriptionally active (Active), inactive (Inactive), and Polycomb-repressed (Repressed). While active and inactive regions can partially intermix with one another, repressed domains show a more compact configuration and do not overlap with other neighboring domains. (C) Microscopy-based methods are used to assess the ability of chromatin components to form condensates through liquid–liquid phase separation (LLPS). Condensate formation is mediated by the biochemical properties of the macromolecules and their interactions.
In Drosophila, high-resolution Hi-C data from Rowley et al. (2017) suggested that TADs and compartments occur at a much lower scale than previously proposed. According to that study, the main topological features are compartmental domains, which represent small discrete regions of ~10 kb that preferentially interact within themselves and correlate with transcriptional activity states [111], thus indicating that Drosophila TADs actually correspond to smaller A/B compartments. This is consistent with the segmentation of the fly genome into the domains of the particular chromatin types mentioned above [13]. Thus, A compartmental domains would correspond to RED and YELLOW chromatin, while B compartmental domains would include BLUE, GREEN and BLACK chromatin types. These compartmental domains can be accurately modeled using only transcriptional data, while both CTCF and transcription-based simulations are required to generate an accurate human Hi-C map [111]. Significantly, the experimental inhibition of transcription in Drosophila cells by triptolide or heat shock results in a decrease in domain architecture that is more pronounced when RNAPII binding is also depleted [111]. In another study from Hug et al. (2017), Hi-C maps generated during fly embryogenesis showed that most TADs are formed concomitantly with the start of transcription. Inhibiting transcription before ZGA results in reduced contact density within domains and a significant loss of interdomain insulation, although TADs are not entirely eliminated [112].
Unlike in mammals, the majority of Drosophila domain borders coincide with active promoters or active gene minidomains rather than with CTCF sites [111,113]. Accordingly, domain boundaries are mostly unaffected by the loss of CTCF, as has been observed in Drosophila neurons [114]. CTCF mutant flies can progress through embryogenesis and larval stages, although they display strong homeotic defects [115]. Instead, many other insulator proteins have been described in Drosophila; however, their role in shaping chromatin domains is still unclear [116]. In recent studies using Hi-C data from Drosophila cell lines, significantly enriched DNA motifs have been identified at domain boundaries, including the motifs for the Suppressor of Hairy wing [Su(Hw)], the Boundary Element Associated Factor (BEAF-32), and the Motif-1 binding protein (M1BP) [113,117]. Interestingly, loss of BEAF-32 has no major effect on chromosome structure, while depletion of M1BP results in cell arrest and dramatic genome reorganization [113]. Another study revealed an essential role for the pioneer TF Zelda in establishing insulation at domain borders, at least partially, during embryogenesis [112]. That study showed that Zelda-depleted embryos display major chromatin conformation defects, although they are mostly locus-specific, i.e., at former Zelda-bound domain boundaries [112]. Furthermore, Zelda mutant embryos show loss of long-range gene interactions [118], suggesting a role of this pioneer factor in chromatin looping during Drosophila embryogenesis. Significantly, Zelda-dependent sites that fail to interact in mutant embryos are associated with the specific depletion of RNAPII binding [118]. Overall, these results suggest that compartmentalization in the fly genome is mostly driven by transcriptional activity and/or RNAPII presence, while CTCF looping is also required for domain formation in mammals. A recent study has proposed that mammalian TADs are subdivided into chromatin nanodomains of similar size to Drosophila TADs. These nanodomains, like Drosophila TADs, are mostly unaffected by CTCF or cohesin depletion [119].
The recent application of super-resolution microscopy coupled to DNA labeling has revealed that Drosophila TADs correlate well with epigenetic states, usually classified into transcriptionally active (associated with H3K4me3, H3K36me3 and acetylated histones), PcG-repressed (enriched in PcG proteins and H3K27me3), and inactive (devoid of specific marks) (Figure 2B). Using a combination of fluorescent in situ hybridization (FISH) and 3D-structured illumination microscopy (3D-SIM), Szabo and colleagues (2018) showed that the chromatin fiber is segmented into globular domains defined as nanocompartments [120]. These structures correspond to PcG or inactive TADs, and are interspersed by regions of less condensed active chromatin [120]. Similarly, 3D stochastic optical reconstruction microscopy (3D-STORM) of labeled chromatin revealed distinct spatial organization for each chromatin state, where PcG-repressed chromatin forms small compact domains that are distinct from the transcriptionally inactive regions [94]. Interestingly, active and inactive domains show less condensation and partial intermixing with each other, while PcG-repressed domains exhibit more compact packaging and a stronger tendency to spatially exclude neighboring active chromatin [94]. These PcG domains depend on Polycomb repressive complex 1 (PRC1), since knockdown of the Polyhomeotic (Ph) component of PRC1 leads to domain loss and an increased expression of PcG-target genes [94]. Similarly, genome editing of Polycomb responsive elements (PRE) caused the aberrant formation of TADs and a decreased silencing during Drosophila embryogenesis [118]. In mammals, PcG-bound chromatin also forms small dense domains in which developmentally regulated genes targeted by PRC1 are corepressed [121]. These domains are functionally lost upon the activation of PRC1-target genes or upon the PRC1 unbinding of target genes during cell fate specification [121]. The function of Ph in domain formation is also conserved for the mammalian ortholog Phc1, since knocking out Phc1 in mouse ESCs leads to domain loss and subsequent gene de-repression [121]. A recent study used Optical Reconstruction of Chromatin Architecture (ORCA), a method that combines high resolution microscopy with Oligopaint and RNA-FISH, to observe the 3D chromatin organization of the bithorax-complex (BX-C) inside the nuclei of individual cells from Drosophila embryos. The study revealed segment-specific organization of the BX-C and identified TADs exclusive to each body segment [122]. The boundaries of these segment-specific TADs often coincide with changes in chromatin states, such as at the edge of a H3K27me3 domain or at the border between transcriptionally active and inactive regions. However, the same study found adjacent TADs with the same epigenetic states that were independent of H3K27me3. The borders of these TADs might be determined by cohesins and CTCF, since the deletion of border regions marked by these proteins results in TADs fusion. Nevertheless, these deletions also include active genes, which could influence the structure of TADs [122]. Finally, another study showed that the boundaries of most Polycomb domains in Drosophila consist of active regions and that actively transcribed genes can stop the spreading of H3K27me3, since ectopic addition of a transcriptional terminator caused the extension of the PcG domain [123].
The results discussed above indicate that the fly genome is divided into functional domains in which regulatory contacts are spatially restricted. However, this model requires TADs to be physical units when they could instead represent statistical frequencies of chromatin interactions within cell populations [124]. Single-cell Hi-C (scHi-C) has been recently introduced to overcome the limitations of bulk Hi-C, but have shown contrasting results in this respect. The contact maps generated for single Drosophila nuclei bore striking resemblance to the TAD profile in bulk Hi-C, with over 40% of TAD boundaries conserved between individual cells [125]. However, the long-range interactions within and between TADs were very heterogeneous, suggesting substantial stochasticity in the folding of chromatin into domains [125]. Another study using human cell lines showed high cell-to-cell variations in contact patterns, where single-cell contact clusters did not match population TADs [126]. These scHi-C maps averaged into TADs when they were pooled together, suggesting that TADs merely reflect the tendencies of measured interactions within a population of cells [126]. Using 3D-SIM and DNA-labeling in Drosophila, contact probabilities within TADs were shown to be higher for all cells compared to contacts between neighboring TADs [127]. This preferent confinement of interactions within domains indicates the presence of TADs in individual cells. Furthermore, Cattoni and colleagues (2017) showed that active and repressive chromatin form discrete nanocompartments at the single-cell level, whereas stable looping between TAD borders is infrequent. This suggests that TAD assembly in flies is not the result of long-term stable interactions, but rather can be explained by the stochastic contacts between regions of similar chromatin types [127]. In agreement with this, Szabo and colleagues (2018) observed that TADs are generally consistent between cells, despite variable intra- and interdomain contacts, further suggesting that TADs are true physical entities of the fly genome [120]. Likewise, super-resolution microscopy of mouse ESCs showed significant variability at the TAD scale between individual cells, although chromatin contacts were more frequent within TADs than between adjacent TADs [119]. Further evidence in favor of TADs is the observation that bands from Drosophila polytene chromosomes strongly correlate with TADs, whereas the decondensed polytene regions, named interbands, mostly correspond to inter-TADs or TAD borders [128,129]. This finding reveals a direct relationship between sequencing-inferred chromatin structure and chromosome condensation observed by light microscopy [128,129], suggesting that TADs are a stable and conserved unit of chromosome folding.
5.3. Regulation of Gene Expression by Chromatin Organization
While there is an increasing amount of evidence showing that transcriptional activity plays a prominent role in chromatin organization [111,130,131,132], it is still rather controversial how—and if—genome topology modulates gene expression [133]. In a recent study, the simultaneous analysis of enhancer–promoter interactions and transcription in Drosophila embryos showed that the sustained proximity of the enhancer to its target gene was required for activation [131]. Moreover, the Drosophila Hox gene clusters Antennapedia and Bithorax, which are separated by 10 Mb in the linear genome, are thought to be corepressed by a PcG-mediated physical interaction, with mutations in one complex resulting in the de-repression of the genes in the other complex [93]. Interestingly, the absence of either one of the two PRC1 subunits Ph or Pc affects chromatin organization prior to ectopic Hox gene transcription, suggesting that PRC1 maintains gene silencing by compacting chromatin into domains [134]. In agreement with these results, disruption of the TAD structure has been found to cause misexpression in some cases. For instance, the deletion or inversion of the TAD boundary at the Epha4 locus causes ectopic interactions between promoters and enhancers, leading to aberrant gene expression and a pathogenic phenotype in mammalian limbs [135]. Furthermore, deletion of domain boundaries in the Notch locus of Drosophila results in TAD fusion and transcriptional changes, together with a decreased binding of RNAPII [136].
However, there is also significant evidence questioning the role of the chromatin structure modulating gene expression [133,137]. In Drosophila, the analysis of gene expression in balancers (highly rearranged chromosomes) showed that disruption of TADs has little effect on gene activity and only a subset of genes is sensitive to structural alterations [138]. Similarly, a recent study using Drosophila dorsoventral patterning as a model system indicated that genome topology and transcription are independent [139]. Following fly embryonic development, the study demonstrated that chromatin conformation is generally maintained across cell types, even if transcriptional profiles change dramatically [139]. Likewise, single-cell spatial genomics in Drosophila embryos revealed that developmentally relevant enhancer–promoter interactions appear before TAD formation and remain invariant during cell fate specification [140]. Finally, sonic hedgehog (Shh) gene expression during mouse limb development was shown to be robust to perturbations in the TAD structure and was only altered when the Shh limb enhancers were deleted [141]. Altogether, these results suggest that chromatin organization can play a role in regulating gene expression, but it is most likely to be one of many modulators and not the main regulatory factor.
5.4. Nuclear Lamina and Pericentromeric Heterochromatin
The functional organization of the genome is also influenced by the existing contacts between the chromatin and the nuclear lamina (NL) (Figure 1A) [142]. The NL is a dense meshwork composed of A- and B-type lamins and lamin-associated proteins that, together with the outer and inner membranes and the nuclear pore complexes, form the nuclear envelope. The NL functions as a support for multiple chromatin anchoring sites and it is suggested that NL proteins tether heterochromatin to the nuclear periphery. In fact, Drosophila genes that interact with B-type lamin are transcriptionally silent and late replicating [143]. During the differentiation of mouse ESCs, genes that move away from the lamina are activated, whereas the others remain inactive and become activated in the next differentiation step [144].
Lamin B receptor, located at the inner nuclear membrane, provides a tethering mechanism for heterochromatin by binding to HP1a, which is preferentially located in the H3K9me3-rich heterochromatic region [145,146,147]. Hi-C data have also confirmed that the inactive chromatin compartment is strongly enriched in NL contacts [148,149]. In addition, transcriptional repressors, such as HDACs, are known to bind to lamina proteins [150,151,152], raising the question of whether the contacts with the NL make the chromatin in lamina-associated domains (LADs) compact and inactive. Evidence indicates that the NL contacts can play a role in gene repression. In vitro experiments on Drosophila cell lines have demonstrated that knocking down lamin proteins decreases the compactness of inactive chromatin domains, increases the accessibility of the promoters located in heterochromatic regions, enhances the levels of histone H3 acetylation, and increases gene expression [144,153,154,155]. Similarly, in mouse ESCs, the expression of reporter genes inserted into LADs is remarkably lower when compared to inter-LADs, which is only partially explained by chromatin compaction, therefore pointing towards a repressive environment near the NL where transcription is less permissive [156]. Additionally, several studies on cell differentiation have revealed that the activation of tissue-specific gene expression is associated with translocations of loci from the NL to the nuclear interior [144,149,157,158]. For instance, the ectopic release of cardiac genes from the nuclear periphery in mouse ESCs leads to a premature myogenesis, providing a clear example of the relevance of gene positioning in the nuclear periphery during organogenesis [159]. Together, these findings indicate that the NL is essential for the positioning of the heterochromatin in the nuclear periphery, establishing a transcriptionally repressed domain near the nuclear envelope and a transcriptionally active domain in the nuclear interior [160].
Apart from the heterochromatin contacts with the NL, interactions between different heterochromatic regions, most likely mediated by the affinity between repetitive elements [161,162] or heterochromatin-associated proteins [163,164], could be essential for heterochromatin compartmentalization within the nucleus [165]. Contacts between pericentromeric heterochromatin (PCH) regions, which are located near the centromere and are enriched in HP1a and repetitive elements [166,167], have been proposed to contribute to the global genome architecture in the nucleus [168]. In Drosophila, the PCH regions from the four chromosomes cluster in the 3D nucleus, forming membraneless structures called chromocenters that were first described in the 1970s [169,170,171]. Although recent studies have found HP1a to be crucial for PCH clustering in Drosophila embryos [172], the mechanism driving this clustering in differentiated cells in mice does not depend on HP1a, indicating an alternative mechanism for PCH coalescence [173]. However, PCH contacts are not only restricted to other PCH regions. It was recently found that PCH can interact with euchromatic regions enriched in H3K9me2/3. Genes located in PCH-contacting euchromatin show higher expression than those located in euchromatin not contacting PCH [168]. Since HP1a is the reader of H3K9me2/3, the most probable scenario is that PCH-euchromatic interactions are driven by HP1a.
5.5. Nuclear Pore Complexes
Besides the nuclear lamina, chromatin can also interact with the nuclear pore complexes (NPCs) located in the nuclear envelope and whose canonical function is to mediate nucleocytoplasmic transport. NPCs are composed of multiple nucleoporins (Nups), for which ~30 different subtypes have been described [174]. Depending on the Nup subtype, the association to the NPC can be stable or dynamic, as certain Nups can be found freely within the nucleoplasm. In Drosophila, as well as in mammals, chromatin binding maps show differential binding preferences for different Nup subtypes. For instance, Nup153 and Megator (Mtor) are known to bind to the fly genome in continuous domains enriched in marks of active transcription [175]. In fact, the depletion of Nup153 results in the altered expression of thousands of genes, suggesting a role in transcriptional regulation [175]. However, not all Nup subtypes bind preferentially to active regions. An interesting example is the antagonistic binding preferences of core nuclear pore proteins Nup93 and Nup107 in Drosophila cells; while Nup107 is mainly found at active genes, Nup93 is located preferentially at repressed chromatin regions, bound by PcG proteins [176]. Indeed, the depletion of Nup93 leads to de-clustering of distant Polycomb regions that previously coalesced, and to de-repression of PcG target genes [176]. These findings point towards Nup93 as a key mediator of long-range Polycomb interactions, as well as a necessary factor for proper silencing of PcG-associated target genes.
Dynamic Nups can also interact with the genome within the nucleoplasm, outside the NPC. This is the case of Nup98, which binds genes both at the NPC and within the nucleoplasm [177,178]. The association involving nucleoplasmic Nup98 correlates with a higher degree of target gene activation compared to NPC-bound Nup98. The expression of these target genes depends on the levels of Nup98, as their expression increases upon the ectopic overexpression of Nup98, and decreases upon Nup98 depletion [178]. In line with these findings, the interacting partners of Nup98 include several proteins implicated in gene activation, including Thritorax (Trx), suggesting a role of Nup98 in maintaining active gene expression [179].
NPCs also seem to play a role in genome architecture. Multiple studies have demonstrated that NPC components contribute to the formation of long-range genomic contacts [180,181,182]. In Drosophila, as well as in mammals, several Nups are found targeting a subset of promoters and enhancers. These contacts are preferentially located in NPC-bound Nups rather than nucleoplasmic Nups, and occur regardless of the transcriptional state of the genes, suggesting that NPCs can target silent or poised genes and enhancers [180,181]. Particularly, Nup98 was found to bind promoters and enhancers of ecdysone-inducible genes and, upon its depletion, the enhancer–promoter loops induced by ecdysone were destabilized [180]. Moreover, NPC-bound Nup98 was also observed to bind at a subset of TAD boundary regions, and was identified as a physical interactor to insulator proteins such as CTCF [113]. Altogether, these findings suggest that NPCs not only bind to specific regions in the genome, but may also participate in the formation or maintenance of promoter-enhancer loops as well as in the formation of TAD borders.
6. Liquid–Liquid Phase Separation
Functional compartmentalization of the cell nucleus plays an important role in the regulation of genome activity. Recent evidence suggests that liquid–liquid phase separation (LLPS) underlies the formation of membraneless compartments in the nucleus [183]. These membraneless compartments are formed as a result of distinct biochemical properties that segregate macromolecules into a concentrated liquid phase and a dilute phase, referred to as condensates [184]. This non-covalent form of fluid compartmentalization is triggered by weak multivalent interactions between proteins, RNA, and DNA. Specifically, proteins are thought to mediate phase separation through interactions between domains that are called low-complexity domains (LDRs) or intrinsically disordered domains (IDRs) [184,185]. Recent studies suggest that chromatin compartments might be formed and organized through LLPS (Figure 2C). In Drosophila, HP1a undergoes liquid phase separation in vitro and can form condensates suggested to facilitate heterochromatin formation in early embryos [164]. Indeed, partial knock-down of HP1a causes major alterations in chromatin organization in Drosophila embryos, such as a reduced contact frequency within heterochromatic regions and increased crosstalk between active and inactive domains [172]. HP1a depletion in differentiated cells does not affect genome structure, suggesting that HP1a is required for establishing 3D structure in early embryos, but not for the maintenance of compartmentalization during cell differentiation [172]. The human counterpart, HP1α, is also capable of compacting heterochromatin through phase separation in an in vitro model system [163]. However, a recent study using mouse fibroblasts showed that HP1α has a weak capacity to form liquid droplets in living cells and that the compaction and maintenance of heterochromatin foci occur independently of HP1α [173].
Other proteins have been suggested to facilitate chromatin condensation through LLPS, including histones and transcriptional regulators. The linker histone H1 was recently shown to promote the phase separation of reconstituted chromatin into denser and less dynamic droplets [186]. On the other hand, acetylation of histone tails gradually reduces chromatin droplet density until its dissolution. This acetylated chromatin, which is unable to undergo phase separation by itself, undergoes LLPS after the addition of multi-bromodomain proteins in vitro [186]. Similarly, the Chromobox 2 (CBX2) component of PRC1 was recently proposed to mediate the phase separation of PcG-repressed chromatin in mice. In an in vitro model, reconstituted PRC1 can undergo phase separation into droplets, but fails to do so when CBX2 is mutated in its LDR [187]. The same mutations in CBX2 had been previously shown to decrease chromatin compaction and transcriptional repression in mouse cells [188]. Finally, superenhancers, which are clusters of enhancers that are able to recruit high levels of transcriptional regulators and can strongly activate gene expression [189,190], are also thought to assemble through LLPS [191]. Indeed, recent studies suggest that TFs and coactivators can form phase-separated condensates, in which the transcription machinery is highly concentrated and facilitates the expression of genes in both flies and mammals [191,192,193]. Interestingly, the carboxy-terminal domain of RNAPII, an LDR, can undergo phase separation in vitro even in the absence of other proteins [194]. Moreover, the activation domains of several TFs can form liquid droplets in vitro with the coactivator mediator subunit MED1, which results in gene activation [195]. Altogether, the evidence suggests that chromatin has an intrinsic capacity to undergo phase separation into functionally distinct, but physically adjacent domains. However, the precise contribution of LLPS to the formation of chromatin domains and genome function is still unclear.
7. Conclusions and Future Perspectives
In this review, we aimed to highlight what is known about the genome of Drosophila melanogaster regarding chromatin composition and 3D organization, mostly related to the regulation of gene expression. As we have discussed, the role that chromatin modifications play in transcription is still unclear, and new studies are now focusing on the effects of local chromatin environment and genome folding. In spite of several open questions, the 3D organization of the genome in nuclear compartments is increasingly recognized as a major feature of gene regulation. How these nuclear compartments are formed and what is their importance regarding gene function remains to be elucidated.
Over the last decade, numerous studies have used chromosome conformation capture techniques to analyze the 3D architecture of genomes [95]. In Drosophila, these studies have revealed that the genome is organized into physical domains of particular chromatin states named TADs. However, Hi-C experiments reflect averaged information coming from cell populations, and thus cannot account for individual heterogeneity between cells, cell-types or tissues. To overcome this limitation, new methodologies have been developed, including scHi-C [126] and microscopy-based techniques, which couple super-resolution imaging with DNA labeling [120,122,196,197]. These emerging microscopy methods not only enable the visualization of the 3D genome organization at the single-cell level, but also maintain the spatial information within tissues or organisms. Furthermore, they allow the incorporation of RNA probes to distinguish cell types and to link chromatin structure to gene expression [122,196,197]. Other technical approaches to study the relationship between genome organization and gene expression include gene silencing by RNA interference (RNAi) and genomic engineering with CRISPR/Cas9 technology [198]. In Drosophila, CRISPR/Cas9 screens have recently been used to identify factors involved in genome architecture, including chromatin-binding proteins and functional cis-regulatory elements, such as enhancers and boundary elements or insulators [198]. Furthermore, the development of new methods based on a nuclease-dead Cas9 (dCas9) coupled with live imaging is allowing researchers to study the 4D genome, changes in 3D chromatin structure over time [199]. In the CRISPR/dCas9 system, there is specific recruitment of a fluorescently labelled dCas9 to the genomic region of interest, which allows the tracking of the contact and folding dynamics of this region by live microscopy [199].
Sequencing-based technologies rely on the available version of the genome and, in the case of Drosophila, the assembly of heterochromatic regions is still incomplete. New technologies based on long-read sequencing will improve the current genome assembly and thus increase the quality of Hi-C maps. In the field of microscopy, one of the major limitations is the number of loci that can be probed simultaneously. Recent advances including ORCA [122], Hi-M [197], and OligoFISSEQ [200] allow the sequential labeling and imaging of multiple genomic sequences, but still require long acquisition times and are very expensive. Altogether, the development of these techniques is encouraging, and further optimization will allow us to resolve the chromatin dynamics of whole genomes and to uncover the degree of heterogeneity inside cell populations.
Funding
This work was supported by grants PGC2018-099763-B100 and 2017SGR1455 from AGAUR (Generalitat de Catalunya) to M.C., C.C.-R. holds a predoctoral FPI contract from the Spanish Government (Ministerio de Ciencia e Innovación) and P.L.-G. is a predoctoral fellow of FI, Generalitat de Catalunya.
Institutional Review Board Statement
Not applicable.
Acknowledgments
We thank Jordi Bernués for critical reading of the manuscript and we apologize to colleagues whose work could not be cited here due to space limitations. Figures were created using Illustrator.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Adams, M.D.; Celniker, S.E.; Holt, R.A.; Evans, C.A.; Gocayne, J.D.; Amanatides, P.G.; Scherer, S.E.; Li, P.W.; Hoskins, R.A.; Galle, R.F.; et al. The Genome Sequence of Drosophila melanogaster. Science 2000, 287, 2185–2195. [Google Scholar] [CrossRef]
- Myers, E.W. A Whole-Genome Assembly of Drosophila. Science 2000, 287, 2196–2204. [Google Scholar] [CrossRef]
- Hoskins, R.A.; Carlson, J.W.; Wan, K.H.; Park, S.; Mendez, I.; Galle, S.E.; Booth, B.W.; Pfeiffer, B.D.; George, R.A.; Svirskas, R.; et al. The Release 6 reference sequence of the Drosophila melanogaster genome. Genome Res. 2015, 25, 445–458. [Google Scholar] [CrossRef]
- Zhimulev, I.F.; Belyaeva, E.S.; Vatolina, T.Y.; Demakov, S.A. Banding patterns in Drosophila melanogaster polytene chromosomes correlate with DNA-binding protein occupancy. BioEssays 2012, 34, 498–508. [Google Scholar] [CrossRef]
- Zykova, T.Y.; Levitsky, V.G.; Belyaeva, E.S.; Zhimulev, I.F. Polytene Chromosomes—A Portrait of Functional Organization of the Drosophila Genome. Curr. Genom. 2018, 19, 179–191. [Google Scholar] [CrossRef]
- Grewal, S.I.S.; Elgin, S.C.R. Heterochromatin: New possibilities for the inheritance of structure. Curr. Opin. Genet. Dev. 2002, 12, 178–187. [Google Scholar] [CrossRef]
- Craig, J.M. Heterochromatin? many flavours, common themes. BioEssays 2005, 27, 17–28. [Google Scholar] [CrossRef]
- Pimpinelli, S.; Wakimoto, B.T. Expanding the boundaries of heterochromatin. Genetica 2003, 117, 111–116. [Google Scholar] [CrossRef]
- Pimpinelli, S.; Berloco, M.; Fanti, L.; Dimitri, P.; Bonaccorsi, S.; Marchetti, E.; Caizzi, R.; Caggese, C.; Gatti, M. Transposable elements are stable structural components of Drosophila melanogaster heterochromatin. Proc. Natl. Acad. Sci. USA 1995, 92, 3804–3808. [Google Scholar] [CrossRef]
- Eissenberg, J.C.; Elgin, S.C.R. HP1a: A structural chromosomal protein regulating transcription. Trends Genet. 2014, 30, 103–110. [Google Scholar] [CrossRef]
- Marsano, R.M.; Giordano, E.; Messina, G.; Dimitri, P. A New Portrait of Constitutive Heterochromatin: Lessons from Drosophila melanogaster. Trends Genet. 2019, 35, 615–631. [Google Scholar] [CrossRef]
- Swenson, J.M.; Colmenares, S.U.; Strom, A.R.; Costes, S.V.; Karpen, G.H. The composition and organization of Drosophila heterochromatin are heterogeneous and dynamic. eLife 2016, 5, e16096. [Google Scholar] [CrossRef]
- Filion, G.J.; van Bemmel, J.G.; Braunschweig, U.; Talhout, W.; Kind, J.; Ward, L.D.; Brugman, W.; de Castro, I.J.; Kerkhoven, R.M.; Bussemaker, H.J.; et al. Systematic Protein Location Mapping Reveals Five Principal Chromatin Types in Drosophila Cells. Cell 2010, 143, 212–224. [Google Scholar] [CrossRef]
- Kornberg, R.D. Chromatin Structure: A Repeating Unit of Histones and DNA. Science 1974, 184, 868–871. [Google Scholar] [CrossRef]
- Luger, K.; Mäder, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef]
- Iwasaki, W.; Miya, Y.; Horikoshi, N.; Osakabe, A.; Taguchi, H.; Tachiwana, H.; Shibata, T.; Kagawa, W.; Kurumizaka, H. Contribution of histone N-terminal tails to the structure and stability of nucleosomes. FEBS Open Bio 2013, 3, 363–369. [Google Scholar] [CrossRef]
- Cutter, A.R.; Hayes, J.J. A brief review of nucleosome structure. FEBS Lett. 2015, 589, 2914–2922. [Google Scholar] [CrossRef]
- Allan, J.; Hartman, P.G.; Crane-Robinson, C.; Aviles, F.X. The structure of histone H1 and its location in chromatin. Nature 1980, 288, 675–679. [Google Scholar] [CrossRef]
- Thoma, F.; Koller, T. Influence of histone H1 on chromatin structure. Cell 1977, 12, 101–107. [Google Scholar] [CrossRef]
- Thoma, F.; Koller, T.; Klug, A. Involvement of histone H1 in the organization of the nucleosome and of the salt-dependent superstructures of chromatin. J. Cell Biol. 1979, 83, 403–427. [Google Scholar] [CrossRef]
- Simpson, R.T. Structure of the chromatosome, a chromatin particle containing 160 base pairs of DNA and all the histones. Biochemistry 1978, 17, 5524–5531. [Google Scholar] [CrossRef]
- Bednar, J.; Garcia-Saez, I.; Boopathi, R.; Cutter, A.R.; Papai, G.; Reymer, A.; Syed, S.H.; Lone, I.N.; Tonchev, O.; Crucifix, C.; et al. Structure and Dynamics of a 197 bp Nucleosome in Complex with Linker Histone H1. Mol. Cell 2017, 66, 384–397.e8. [Google Scholar] [CrossRef]
- Cutter, A.R.; Hayes, J.J. Linker histones: Novel insights into structure-specific recognition of the nucleosome. Biochem. Cell Biol. 2017, 95, 171–178. [Google Scholar] [CrossRef]
- Ricci, M.A.; Manzo, C.; García-Parajo, M.F.; Lakadamyali, M.; Cosma, M.P. Chromatin Fibers Are Formed by Heterogeneous Groups of Nucleosomes In Vivo. Cell 2015, 160, 1145–1158. [Google Scholar] [CrossRef]
- Mavrich, T.N.; Ioshikhes, I.P.; Venters, B.J.; Jiang, C.; Tomsho, L.P.; Qi, J.; Schuster, S.C.; Albert, I.; Pugh, B.F. A barrier nucleosome model for statistical positioning of nucleosomes throughout the yeast genome. Genome Res. 2008, 18, 1073–1083. [Google Scholar] [CrossRef]
- Kaplan, N.; Moore, I.K.; Fondufe-Mittendorf, Y.; Gossett, A.J.; Tillo, D.; Field, Y.; LeProust, E.M.; Hughes, T.R.; Lieb, J.D.; Widom, J.; et al. The DNA-encoded nucleosome organization of a eukaryotic genome. Nature 2009, 458, 362–366. [Google Scholar] [CrossRef]
- Yuan, G.C.; Liu, Y.-J.; Dion, M.F.; Slack, M.D.; Wu, L.F.; Altschuler, S.J.; Rando, O.J. Genome-Scale Identification of Nucleosome Positions in S. cerevisiae. Science 2005, 309, 626–630. [Google Scholar] [CrossRef]
- Struhl, K.; Segal, E. Determinants of nucleosome positioning. Nat. Struct. Mol. Biol. 2013, 20, 267–273. [Google Scholar] [CrossRef]
- Martin, R.L.; Maiorano, J.; Beitel, G.J.; Marko, J.F.; McVicker, G.; Fondufe-Mittendorf, Y.N. A comparison of nucleosome organization in Drosophila cell lines. PLoS ONE 2017, 12, e0178590. [Google Scholar] [CrossRef]
- Li, G.; Levitus, M.; Bustamante, C.; Widom, J. Rapid spontaneous accessibility of nucleosomal DNA. Nat. Struct. Mol. Biol. 2005, 12, 46–53. [Google Scholar] [CrossRef]
- Poirier, M.G.; Oh, E.; Tims, H.S.; Widom, J. Dynamics and function of compact nucleosome arrays. Nat. Struct. Mol. Biol. 2009, 16, 938–944. [Google Scholar] [CrossRef]
- Zhou, K.; Gaullier, G.; Luger, K. Nucleosome structure and dynamics are coming of age. Nat. Struct. Mol. Biol. 2019, 26, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Lai, W.K.M.; Pugh, B.F. Understanding nucleosome dynamics and their links to gene expression and DNA replication. Nat. Rev. Mol. Cell Biol. 2017, 18, 548–562. [Google Scholar] [CrossRef] [PubMed]
- Horard, B.; Loppin, B. Histone storage and deposition in the early Drosophila embryo. Chromosoma 2015, 124, 163–175. [Google Scholar] [CrossRef]
- Talbert, P.B.; Henikoff, S. Histone variants at a glance. J. Cell Sci. 2021, 134, jcs244749. [Google Scholar] [CrossRef] [PubMed]
- Baldi, S.; Becker, P.B. The variant histone H2A.V of Drosophila—Three roles, two guises. Chromosoma 2013, 122, 245–258. [Google Scholar] [CrossRef]
- Prozzillo, Y.; Cuticone, S.; Ferreri, D.; Fattorini, G.; Messina, G.; Dimitri, P. In Vivo Silencing of Genes Coding for dTip60 Chromatin Remodeling Complex Subunits Affects Polytene Chromosome Organization and Proper Development in Drosophila melanogaster. Int. J. Mol. Sci. 2021, 22, 4525. [Google Scholar] [CrossRef]
- Messina, G.; Damia, E.; Fanti, L.; Atterrato, M.T.; Celauro, E.; Mariotti, F.R.; Accardo, M.C.; Walther, M.; Vernì, F.; Picchioni, D.; et al. Yeti, a Drosophila melanogaster essential gene, encodes a protein required for chromatin organization. J. Cell Sci. 2014, 127, 2577–2588. [Google Scholar] [CrossRef]
- Weber, C.M.; Henikoff, J.G.; Henikoff, S. H2A.Z nucleosomes enriched over active genes are homotypic. Nat. Struct. Mol. Biol. 2010, 17, 1500–1507. [Google Scholar] [CrossRef]
- Weber, C.M.; Ramachandran, S.; Henikoff, S. Nucleosomes are context-specific, H2A.Z-Modulated barriers to RNA polymerase. Mol. Cell 2014, 53, 819–830. [Google Scholar] [CrossRef]
- Swaminathan, J.; Baxter, E.M.; Corces, V.G. The role of histone H2Av variant replacement and histone H4 acetylation in the establishment of Drosophila heterochromatin. Genes Dev. 2005, 19, 65–76. [Google Scholar] [CrossRef]
- Leach, T.J.; Mazzeo, M.; Chotkowski, H.L.; Madigan, J.P.; Wotring, M.G.; Glaser, R.L. Histone H2A.Z is widely but nonrandomly distributed in chromosomes of Drosophila melanogaster. J. Biol. Chem. 2000, 275, 23267–23272. [Google Scholar] [CrossRef]
- Akhmanova, A.S.; Bindels, P.C.T.; Xu, J.; Miedema, K.; Kremer, H.; Hennig, W.; Xu, J.; Hennig, W. Structure and expression of histone H3.3 genes in Drosophila melanogaster and Drosophila hydei. Genome 1995, 38, 586–600. [Google Scholar] [CrossRef]
- Ahmad, K.; Henikoff, S. The Histone Variant H3.3 Marks Active Chromatin by Replication-Independent Nucleosome Assembly. Mol. Cell 2002, 9, 1191–1200. [Google Scholar] [CrossRef]
- Goldberg, A.D.; Banaszynski, L.A.; Noh, K.-M.; Lewis, P.W.; Elsaesser, S.J.; Stadler, S.; Dewell, S.; Law, M.; Guo, X.; Li, X.; et al. Distinct Factors Control Histone Variant H3.3 Localization at Specific Genomic Regions. Cell 2010, 140, 678–691. [Google Scholar] [CrossRef]
- Mito, Y.; Henikoff, J.G.; Henikoff, S. Histone Replacement Marks the Boundaries of cis-Regulatory Domains. Science 2007, 315, 1408–1411. [Google Scholar] [CrossRef]
- Hödl, M.; Basler, K. Transcription in the Absence of Histone H3.3. Curr. Biol. 2009, 19, 1221–1226. [Google Scholar] [CrossRef]
- Braunschweig, U.; Hogan, G.J.; Pagie, L.; van Steensel, B. Histone H1 binding is inhibited by histone variant H3.3. EMBO J. 2009, 28, 3635–3645. [Google Scholar] [CrossRef] [PubMed]
- Nagel, S.; Grossbach, U. Histone H1 Genes and Histone Gene Clusters in the Genus Drosophila. J. Mol. Evol. 2000, 51, 286–298. [Google Scholar] [CrossRef]
- Pérez-Montero, S.; Carbonell, A.; Morán, T.; Vaquero, A.; Azorín, F. The embryonic linker histone H1 variant of Drosophila, dBigH1, regulates zygotic genome activation. Dev. Cell 2013, 26, 578–590. [Google Scholar] [CrossRef]
- Nalabothula, N.; McVicker, G.; Maiorano, J.; Martin, R.; Pritchard, J.K.; Fondufe-Mittendorf, Y.N. The chromatin architectural proteins HMGD1 and H1 bind reciprocally and have opposite effects on chromatin structure and gene regulation. BMC Genomics 2014, 15, 92. [Google Scholar] [CrossRef]
- Lu, X.; Wontakal, S.N.; Kavi, H.; Kim, B.J.; Guzzardo, P.M.; Emelyanov, A.V.; Xu, N.; Hannon, G.J.; Zavadil, J.; Fyodorov, D.V.; et al. Drosophila H1 Regulates the Genetic Activity of Heterochromatin by Recruitment of Su(var)3-9. Science 2013, 340, 78–81. [Google Scholar] [CrossRef]
- Vujatovic, O.; Zaragoza, K.; Vaquero, A.; Reina, O.; Bernués, J.; Azorín, F. Drosophila melanogaster linker histone dH1 is required for transposon silencing and to preserve genome integrity. Nucleic Acids Res. 2012, 40, 5402–5414. [Google Scholar] [CrossRef]
- Lu, X.; Wontakal, S.N.; Emelyanov, A.V.; Morcillo, P.; Konev, A.Y.; Fyodorov, D.V.; Skoultchi, A.I. Linker histone H1 is essential for Drosophila development, the establishment of pericentric heterochromatin, and a normal polytene chromosome structure. Genes Dev. 2009, 23, 452–465. [Google Scholar] [CrossRef]
- Siriaco, G.; Deuring, R.; Chioda, M.; Becker, P.B.; Tamkun, J.W. Drosophila ISWI Regulates the Association of Histone H1 with Interphase Chromosomes in Vivo. Genetics 2009, 182, 661–669. [Google Scholar] [CrossRef]
- Bayona-Feliu, A.; Casas-Lamesa, A.; Reina, O.; Bernués, J.; Azorín, F. Linker histone H1 prevents R-loop accumulation and genome instability in heterochromatin. Nat. Commun. 2017, 8, 283. [Google Scholar] [CrossRef]
- Cao, K.; Lailler, N.; Zhang, Y.; Kumar, A.; Uppal, K.; Liu, Z.; Lee, E.K.; Wu, H.; Medrzycki, M.; Pan, C.; et al. High-Resolution Mapping of H1 Linker Histone Variants in Embryonic Stem Cells. PLoS Genet. 2013, 9, e1003417. [Google Scholar] [CrossRef]
- Izzo, A.; Kamieniarz-Gdula, K.; Ramírez, F.; Noureen, N.; Kind, J.; Manke, T.; van Steensel, B.; Schneider, R. The Genomic Landscape of the Somatic Linker Histone Subtypes H1.1 to H1.5 in Human Cells. Cell Rep. 2013, 3, 2142–2154. [Google Scholar] [CrossRef]
- Mayor, R.; Izquierdo-Bouldstridge, A.; Millán-Ariño, L.; Bustillos, A.; Sampaio, C.; Luque, N.; Jordan, A. Genome Distribution of Replication-independent Histone H1 Variants Shows H1.0 Associated with Nucleolar Domains and H1X Associated with RNA Polymerase II-enriched Regions. J. Biol. Chem. 2015, 290, 7474–7491. [Google Scholar] [CrossRef]
- Millán-Ariño, L.; Islam, A.B.M.M.K.; Izquierdo-Bouldstridge, A.; Mayor, R.; Terme, J.-M.; Luque, N.; Sancho, M.; López-Bigas, N.; Jordan, A. Mapping of six somatic linker histone H1 variants in human breast cancer cells uncovers specific features of H1.2. Nucleic Acids Res. 2014, 42, 4474–4493. [Google Scholar] [CrossRef]
- Fan, Y.; Nikitina, T.; Zhao, J.; Fleury, T.J.; Bhattacharyya, R.; Bouhassira, E.E.; Stein, A.; Woodcock, C.L.; Skoultchi, A.I. Histone H1 Depletion in Mammals Alters Global Chromatin Structure but Causes Specific Changes in Gene Regulation. Cell 2005, 123, 1199–1212. [Google Scholar] [CrossRef]
- Izquierdo-Bouldstridge, A.; Bustillos, A.; Bonet-Costa, C.; Aribau-Miralbés, P.; García-Gomis, D.; Dabad, M.; Esteve-Codina, A.; Pascual-Reguant, L.; Peiró, S.; Esteller, M.; et al. Histone H1 depletion triggers an interferon response in cancer cells via activation of heterochromatic repeats. Nucleic Acids Res. 2017, 45, 11622–11642. [Google Scholar] [CrossRef]
- Sancho, M.; Diani, E.; Beato, M.; Jordan, A. Depletion of Human Histone H1 Variants Uncovers Specific Roles in Gene Expression and Cell Growth. PLoS Genet. 2008, 4, e1000227. [Google Scholar] [CrossRef]
- Healton, S.E.; Pinto, H.D.; Mishra, L.N.; Hamilton, G.A.; Wheat, J.C.; Swist-Rosowska, K.; Shukeir, N.; Dou, Y.; Steidl, U.; Jenuwein, T.; et al. H1 linker histones silence repetitive elements by promoting both histone H3K9 methylation and chromatin compaction. Proc. Natl. Acad. Sci. USA 2020, 117, 14251–14258. [Google Scholar] [CrossRef]
- Carbonell, A.; Pérez-Montero, S.; Climent-Cantó, P.; Reina, O.; Azorín, F. The Germline Linker Histone dBigH1 and the Translational Regulator Bam Form a Repressor Loop Essential for Male Germ Stem Cell Differentiation. Cell Rep. 2017, 21, 3178–3189. [Google Scholar] [CrossRef]
- Climent-Cantó, P.; Carbonell, A.; Tamirisa, S.; Henn, L.; Pérez-Montero, S.; Boros, I.M.; Azorín, F. The tumour suppressor brain tumour (Brat) regulates linker histone dBigH1 expression in the Drosophila female germline and the early embryo. Open Biol. 2021, 11, 200408. [Google Scholar] [CrossRef]
- Henn, L.; Szabó, A.; Imre, L.; Román, Á.; Ábrahám, A.; Vedelek, B.; Nánási, P.; Boros, I.M. Alternative linker histone permits fast paced nuclear divisions in early Drosophila embryo. Nucleic Acids Res. 2020, 48, 9007–9018. [Google Scholar] [CrossRef]
- Dunwell, T.L.; Gerd, P.P. Drosophila genomic methylation: New evidence and new questions. Epigenomics 2014, 6, 459–461. [Google Scholar] [CrossRef]
- Greer, E.L.; Blanco, M.A.; Gu, L.; Sendinc, E.; Liu, J.; Aristizábal-Corrales, D.; Hsu, C.-H.; Aravind, L.; He, C.; Shi, Y. DNA Methylation on N6-Adenine in C. elegans. Cell 2015, 161, 868–878. [Google Scholar] [CrossRef]
- Zhang, G.; Huang, H.; Liu, D.; Cheng, Y.; Liu, X.; Zhang, W.; Yin, R.; Zhang, D.; Zhang, P.; Liu, J.; et al. N6-methyladenine DNA modification in Drosophila. Cell 2015, 161, 893–906. [Google Scholar] [CrossRef]
- Yu, G.; Wu, Q.; Gao, Y.; Chen, M.; Yang, M. The Epigenetics of Aging in Invertebrates. Int. J. Mol. Sci. 2019, 20, 4535. [Google Scholar] [CrossRef] [PubMed]
- He, S.; Zhang, G.; Wang, J.; Gao, Y.; Sun, R.; Cao, Z.; Chen, Z.; Zheng, X.; Yuan, J.; Luo, Y.; et al. 6mA-DNA-binding factor Jumu controls maternal-to-zygotic transition upstream of Zelda. Nat. Commun. 2019, 10, 2219. [Google Scholar] [CrossRef]
- Lawrence, M.; Daujat, S.; Schneider, R. Lateral Thinking: How Histone Modifications Regulate Gene Expression. Trends Genet. 2016, 32, 42–56. [Google Scholar] [CrossRef]
- Tessarz, P.; Kouzarides, T. Histone core modifications regulating nucleosome structure and dynamics. Nat. Rev. Mol. Cell Biol. 2014, 15, 703–708. [Google Scholar] [CrossRef]
- Akhtar, A.; Becker, P.B. Activation of Transcription through Histone H4 Acetylation by MOF, an Acetyltransferase Essential for Dosage Compensation in Drosophila. Mol. Cell 2000, 5, 367–375. [Google Scholar] [CrossRef]
- Shogren-Knaak, M.; Haruhiko, I.; Sun, J.-M.; Pazin, M.; Davie, J.R.; Peterson, C.L. Histone H4-K16 Acetylation Controls Chromatin Structure and Protein Interactions. Science 2006, 311, 844–847. [Google Scholar] [CrossRef]
- Lu, X.; Simon, M.D.; Chodaparambil, J.V.; Hansen, J.C.; Shokat, K.M.; Luger, K. The effect of H3K79 dimethylation and H4K20 trimethylation on nucleosome and chromatin structure. Nat. Struct. Mol. Biol. 2008, 15, 1122–1124. [Google Scholar] [CrossRef]
- Bannister, A.J.; Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 2011, 21, 381–395. [Google Scholar] [CrossRef]
- Schuettengruber, B.; Bourbon, H.-M.; Di Croce, L.; Cavalli, G. Genome Regulation by Polycomb and Trithorax: 70 Years and Counting. Cell 2017, 171, 34–57. [Google Scholar] [CrossRef]
- Zhang, T.; Cooper, S.; Brockdorff, N. The interplay of histone modifications—Writers that read. EMBO Rep. 2015, 16, 1467–1481. [Google Scholar] [CrossRef]
- Verdin, E.; Ott, M. 50 years of protein acetylation: From gene regulation to epigenetics, metabolism and beyond. Nat. Rev. Mol. Cell Biol. 2015, 16, 258–264. [Google Scholar] [CrossRef]
- Strahl, B.D.; Allis, C.D. The language of covalent histone modifications. Nature 2000, 403, 41–45. [Google Scholar] [CrossRef] [PubMed]
- Morgan, M.A.J.; Shilatifard, A. Reevaluating the roles of histone-modifying enzymes and their associated chromatin modifications in transcriptional regulation. Nat. Genet. 2020, 52, 1271–1281. [Google Scholar] [CrossRef] [PubMed]
- Talbert, P.B.; Henikoff, S. The Yin and Yang of Histone Marks in Transcription. Annu. Rev. Genom. Hum. Genet. 2021, 22, 147–170. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Lluch, S.; Blanco, E.; Tilgner, H.; Curado, J.; Ruiz-Romero, M.; Corominas, M.; Guigó, R. Absence of canonical marks of active chromatin in developmentally regulated genes. Nat. Genet. 2015, 47, 1158–1167. [Google Scholar] [CrossRef]
- Hödl, M.; Basler, K. Transcription in the Absence of Histone H3.2 and H3K4 Methylation. Curr. Biol. 2012, 22, 2253–2257. [Google Scholar] [CrossRef]
- Zhang, T.; Zhang, Z.; Dong, Q.; Xiong, J.; Zhu, B. Histone H3K27 acetylation is dispensable for enhancer activity in mouse embryonic stem cells. Genome Biol. 2020, 21, 45. [Google Scholar] [CrossRef]
- Li, X.-Y.; Harrison, M.M.; Villalta, J.E.; Kaplan, T.; Eisen, M.B. Establishment of regions of genomic activity during the Drosophila maternal to zygotic transition. eLife 2014, 3, e03737. [Google Scholar] [CrossRef]
- Chen, K.; Johnston, J.; Shao, W.; Meier, S.; Staber, C.; Zeitlinger, J. A global change in RNA polymerase II pausing during the Drosophila midblastula transition. eLife 2013, 2, e00861. [Google Scholar] [CrossRef] [PubMed]
- Regadas, I.; Dahlberg, O.; Vaid, R.; Ho, O.; Belikov, S.; Dixit, G.; Deindl, S.; Wen, J.; Mannervik, M. A unique histone 3 lysine 14 chromatin signature underlies tissue-specific gene regulation. Mol. Cell 2021, 81, 1766–1780.e10. [Google Scholar] [CrossRef]
- Graves, H.K.; Wang, P.; Lagarde, M.; Chen, Z.; Tyler, J.K. Mutations that prevent or mimic persistent post-translational modifications of the histone H3 globular domain cause lethality and growth defects in Drosophila. Epigenet. Chromatin 2016, 9, 9. [Google Scholar] [CrossRef] [PubMed]
- Kharchenko, P.V.; Alekseyenko, A.A.; Schwartz, Y.B.; Minoda, A.; Riddle, N.C.; Ernst, J.; Sabo, P.J.; Larschan, E.; Gorchakov, A.A.; Gu, T.; et al. Comprehensive analysis of the chromatin landscape in Drosophila melanogaster. Nature 2011, 471, 480–485. [Google Scholar] [CrossRef]
- Bantignies, F.; Roure, V.; Comet, I.; Leblanc, B.; Schuettengruber, B.; Bonnet, J.; Tixier, V.; Mas, A.; Cavalli, G. Polycomb-Dependent Regulatory Contacts between Distant Hox Loci in Drosophila. Cell 2011, 144, 214–226. [Google Scholar] [CrossRef] [PubMed]
- Boettiger, A.N.; Bintu, B.; Moffitt, J.R.; Wang, S.; Beliveau, B.J.; Fudenberg, G.; Imakaev, M.; Mirny, L.A.; Wu, C.; Zhuang, X. Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature 2016, 529, 418–422. [Google Scholar] [CrossRef] [PubMed]
- Kempfer, R.; Pombo, A. Methods for mapping 3D chromosome architecture. Nat. Rev. Genet. 2020, 21, 207–226. [Google Scholar] [CrossRef]
- Dekker, J.; Marti-Renom, M.A.; Mirny, L.A. Exploring the three-dimensional organization of genomes: Interpreting chromatin interaction data. Nat. Rev. Genet. 2013, 14, 390–403. [Google Scholar] [CrossRef]
- Lieberman-Aiden, E.; van Berkum, N.L.; Williams, L.; Imakaev, M.; Ragoczy, T.; Telling, A.; Amit, I.; Lajoie, B.R.; Sabo, P.J.; Dorschner, M.O.; et al. Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. Science 2009, 326, 289–293. [Google Scholar] [CrossRef]
- Sexton, T.; Yaffe, E.; Kenigsberg, E.; Bantignies, F.; Leblanc, B.; Hoichman, M.; Parrinello, H.; Tanay, A.; Cavalli, G. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 2012, 148, 458–472. [Google Scholar] [CrossRef]
- Bonev, B.; Cavalli, G. Organization and function of the 3D genome. Nat. Rev. Genet. 2016, 17, 661–678. [Google Scholar] [CrossRef]
- Rao, S.S.P.; Huntley, M.H.; Durand, N.C.; Stamenova, E.K.; Bochkov, I.D.; Robinson, J.T.; Sanborn, A.L.; Machol, I.; Omer, A.D.; Lander, E.S.; et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 2014, 159, 1665–1680. [Google Scholar] [CrossRef]
- Misteli, T. The Self-Organizing Genome: Principles of Genome Architecture and Function. Cell 2020, 183, 28–45. [Google Scholar] [CrossRef] [PubMed]
- Nora, E.P.; Lajoie, B.R.; Schulz, E.G.; Giorgetti, L.; Okamoto, I.; Servant, N.; Piolot, T.; van Berkum, N.L.; Meisig, J.; Sedat, J.; et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 2012, 485, 381–385. [Google Scholar] [CrossRef] [PubMed]
- Dixon, J.R.; Selvaraj, S.; Yue, F.; Kim, A.; Li, Y.; Shen, Y.; Hu, M.; Liu, J.S.; Ren, B. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 2012, 485, 376–380. [Google Scholar] [CrossRef] [PubMed]
- Wutz, G.; Várnai, C.; Nagasaka, K.; Cisneros, D.A.; Stocsits, R.R.; Tang, W.; Schoenfelder, S.; Jessberger, G.; Muhar, M.; Hossain, M.J.; et al. Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 2017, 36, 3573–3599. [Google Scholar] [CrossRef] [PubMed]
- Davidson, I.F.; Bauer, B.; Goetz, D.; Tang, W.; Wutz, G.; Peters, J. DNA loop extrusion by human cohesin. Science 2019, 366, 1338–1345. [Google Scholar] [CrossRef]
- Fudenberg, G.; Imakaev, M.; Lu, C.; Goloborodko, A.; Abdennur, N.; Mirny, L.A. Formation of Chromosomal Domains by Loop Extrusion. Cell Rep. 2016, 15, 2038–2049. [Google Scholar] [CrossRef]
- Rao, S.S.P.; Huang, S.-C.; Glenn St Hilaire, B.; Engreitz, J.M.; Perez, E.M.; Kieffer-Kwon, K.-R.; Sanborn, A.L.; Johnstone, S.E.; Bascom, G.D.; Bochkov, I.D.; et al. Cohesin Loss Eliminates All Loop Domains. Cell 2017, 171, 305–320.e24. [Google Scholar] [CrossRef]
- de Wit, E.; Vos, E.S.M.; Holwerda, S.J.B.; Valdes-Quezada, C.; Verstegen, M.J.A.M.; Teunissen, H.; Splinter, E.; Wijchers, P.J.; Krijger, P.H.L.; de Laat, W. CTCF Binding Polarity Determines Chromatin Looping. Mol. Cell 2015, 60, 676–684. [Google Scholar] [CrossRef]
- Hanssen, L.L.P.; Kassouf, M.T.; Oudelaar, A.M.; Biggs, D.; Preece, C.; Downes, D.J.; Gosden, M.; Sharpe, J.A.; Sloane-Stanley, J.A.; Hughes, J.R.; et al. Tissue-specific CTCF–cohesin-mediated chromatin architecture delimits enhancer interactions and function in vivo. Nat. Cell Biol. 2017, 19, 952–961. [Google Scholar] [CrossRef]
- Nora, E.P.; Goloborodko, A.; Valton, A.-L.; Gibcus, J.H.; Uebersohn, A.; Abdennur, N.; Dekker, J.; Mirny, L.A.; Bruneau, B.G. Targeted Degradation of CTCF Decouples Local Insulation of Chromosome Domains from Genomic Compartmentalization. Cell 2017, 169, 930–944. [Google Scholar] [CrossRef]
- Rowley, M.J.; Nichols, M.H.; Lyu, X.; Ando-Kuri, M.; Rivera, I.S.M.; Hermetz, K.; Wang, P.; Ruan, Y.; Corces, V.G. Evolutionarily Conserved Principles Predict 3D Chromatin Organization. Mol. Cell 2017, 67, 837–852. [Google Scholar] [CrossRef]
- Hug, C.B.; Grimaldi, A.G.; Kruse, K.; Vaquerizas, J.M. Chromatin Architecture Emerges during Zygotic Genome Activation Independent of Transcription. Cell 2017, 169, 216–228. [Google Scholar] [CrossRef] [PubMed]
- Ramírez, F.; Bhardwaj, V.; Arrigoni, L.; Lam, K.C.; Grüning, B.A.; Villaveces, J.; Habermann, B.; Akhtar, A.; Manke, T. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun. 2018, 9, 189. [Google Scholar] [CrossRef] [PubMed]
- Kaushal, A.; Mohana, G.; Dorier, J.; Özdemir, I.; Omer, A.; Cousin, P.; Semenova, A.; Taschner, M.; Dergai, O.; Marzetta, F.; et al. CTCF loss has limited effects on global genome architecture in Drosophila despite critical regulatory functions. Nat. Commun. 2021, 12, 1011. [Google Scholar] [CrossRef] [PubMed]
- Gambetta, M.C.; Furlong, E.E.M. The Insulator Protein CTCF Is Required for Correct Hox Gene Expression, but Not for Embryonic Development in Drosophila. Genetics 2018, 210, 129–136. [Google Scholar] [CrossRef]
- Schwartz, Y.B.; Cavalli, G. Three-dimensional genome organization and function in Drosophila. Genetics 2017, 205, 5–24. [Google Scholar] [CrossRef]
- Wang, Q.; Sun, Q.; Czajkowsky, D.M.; Shao, Z. Sub-kb Hi-C in D. melanogaster reveals conserved characteristics of TADs between insect and mammalian cells. Nat. Commun. 2018, 9, 188. [Google Scholar] [CrossRef]
- Ogiyama, Y.; Schuettengruber, B.; Papadopoulos, G.L.; Chang, J.-M.; Cavalli, G. Polycomb-Dependent Chromatin Looping Contributes to Gene Silencing during Drosophila Development. Mol. Cell 2018, 71, 73–88. [Google Scholar] [CrossRef]
- Szabo, Q.; Donjon, A.; Jerković, I.; Papadopoulos, G.L.; Cheutin, T.; Bonev, B.; Nora, E.P.; Bruneau, B.G.; Bantignies, F.; Cavalli, G. Regulation of single-cell genome organization into TADs and chromatin nanodomains. Nat. Genet. 2020, 52, 1151–1157. [Google Scholar] [CrossRef]
- Szabo, Q.; Jost, D.; Chang, J.-M.; Cattoni, D.I.; Papadopoulos, G.L.; Bonev, B.; Sexton, T.; Gurgo, J.; Jacquier, C.; Nollmann, M.; et al. TADs are 3D structural units of higher-order chromosome organization in Drosophila. Sci. Adv. 2018, 4, eaar8082. [Google Scholar] [CrossRef]
- Kundu, S.; Ji, F.; Sunwoo, H.; Jain, G.; Lee, J.T.; Sadreyev, R.I.; Dekker, J.; Kingston, R.E. Polycomb Repressive Complex 1 Generates Discrete Compacted Domains that Change during Differentiation. Mol. Cell 2017, 65, 432–446. [Google Scholar] [CrossRef]
- Mateo, L.J.; Murphy, S.E.; Hafner, A.; Cinquini, I.S.; Walker, C.A.; Boettiger, A.N. Visualizing DNA folding and RNA in embryos at single-cell resolution. Nature 2019, 568, 49–54. [Google Scholar] [CrossRef]
- De, S.; Gehred, N.D.; Fujioka, M.; Chan, F.W.; Jaynes, J.B.; Kassis, J.A. Defining the Boundaries of Polycomb Domains in Drosophila. Genetics 2020, 216, 689–700. [Google Scholar] [CrossRef]
- Szabo, Q.; Bantignies, F.; Cavalli, G. Principles of genome folding into topologically associating domains. Sci. Adv. 2019, 5, eaaw1668. [Google Scholar] [CrossRef]
- Ulianov, S.V.; Razin, S.V. The two waves in single-cell 3D genomics. Semin. Cell Dev. Biol. 2021. [Google Scholar] [CrossRef]
- Flyamer, I.M.; Gassler, J.; Imakaev, M.; Brandão, H.B.; Ulianov, S.V.; Abdennur, N.; Razin, S.V.; Mirny, L.A.; Tachibana-Konwalski, K. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 2017, 544, 110–114. [Google Scholar] [CrossRef]
- Cattoni, D.I.; Cardozo Gizzi, A.M.; Georgieva, M.; Di Stefano, M.; Valeri, A.; Chamousset, D.; Houbron, C.; Déjardin, S.; Fiche, J.-B.; González, I.; et al. Single-cell absolute contact probability detection reveals chromosomes are organized by multiple low-frequency yet specific interactions. Nat. Commun. 2017, 8, 1753. [Google Scholar] [CrossRef] [PubMed]
- Eagen, K.P.; Hartl, T.A.; Kornberg, R.D. Stable Chromosome Condensation Revealed by Chromosome Conformation Capture. Cell 2015, 163, 934–946. [Google Scholar] [CrossRef]
- Ulianov, S.V.; Khrameeva, E.E.; Gavrilov, A.A.; Flyamer, I.M.; Kos, P.; Mikhaleva, E.A.; Penin, A.A.; Logacheva, M.D.; Imakaev, M.V.; Chertovich, A.; et al. Active chromatin and transcription play a key role in chromosome partitioning into topologically associating domains. Genome Res. 2016, 26, 70–84. [Google Scholar] [CrossRef] [PubMed]
- Hou, C.; Li, L.; Qin, Z.S.; Corces, V.G. Gene Density, Transcription, and Insulators Contribute to the Partition of the Drosophila Genome into Physical Domains. Mol. Cell 2012, 48, 471–484. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Levo, M.; Barinov, L.; Fujioka, M.; Jaynes, J.B.; Gregor, T. Dynamic interplay between enhancer–promoter topology and gene activity. Nat. Genet. 2018, 50, 1296–1303. [Google Scholar] [CrossRef]
- van Steensel, B.; Furlong, E.E.M. The role of transcription in shaping the spatial organization of the genome. Nat. Rev. Mol. Cell Biol. 2019, 20, 327–337. [Google Scholar] [CrossRef]
- Misteli, T.; Finn, E.H. Chromatin architecture is a flexible foundation for gene expression. Nat. Genet. 2021, 53, 426–427. [Google Scholar] [CrossRef]
- Cheutin, T.; Cavalli, G. Loss of PRC1 induces higher-order opening of Hox loci independently of transcription during Drosophila embryogenesis. Nat. Commun. 2018, 9, 3898. [Google Scholar] [CrossRef]
- Lupiáñez, D.G.; Kraft, K.; Heinrich, V.; Krawitz, P.; Brancati, F.; Klopocki, E.; Horn, D.; Kayserili, H.; Opitz, J.M.; Laxova, R.; et al. Disruptions of Topological Chromatin Domains Cause Pathogenic Rewiring of Gene-Enhancer Interactions. Cell 2015, 161, 1012–1025. [Google Scholar] [CrossRef]
- Arzate-Mejía, R.G.; Josué Cerecedo-Castillo, A.; Guerrero, G.; Furlan-Magaril, M.; Recillas-Targa, F. In situ dissection of domain boundaries affect genome topology and gene transcription in Drosophila. Nat. Commun. 2020, 11, 894. [Google Scholar] [CrossRef] [PubMed]
- Cavalheiro, G.R.; Pollex, T.; Furlong, E.E. To loop or not to loop: What is the role of TADs in enhancer function and gene regulation? Curr. Opin. Genet. Dev. 2021, 67, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Ghavi-Helm, Y.; Jankowski, A.; Meiers, S.; Viales, R.R.; Korbel, J.O.; Furlong, E.E.M. Highly rearranged chromosomes reveal uncoupling between genome topology and gene expression. Nat. Genet. 2019, 51, 1272–1282. [Google Scholar] [CrossRef]
- Ing-Simmons, E.; Vaid, R.; Bing, X.Y.; Levine, M.; Mannervik, M.; Vaquerizas, J.M. Independence of chromatin conformation and gene regulation during Drosophila dorsoventral patterning. Nat. Genet. 2021, 53, 487–499. [Google Scholar] [CrossRef] [PubMed]
- Espinola, S.M.; Götz, M.; Bellec, M.; Messina, O.; Fiche, J.-B.; Houbron, C.; Dejean, M.; Reim, I.; Cardozo Gizzi, A.M.; Lagha, M.; et al. Cis-regulatory chromatin loops arise before TADs and gene activation, and are independent of cell fate during early Drosophila development. Nat. Genet. 2021, 53, 477–486. [Google Scholar] [CrossRef]
- Williamson, I.; Kane, L.; Devenney, P.S.; Flyamer, I.M.; Anderson, E.; Kilanowski, F.; Hill, R.E.; Bickmore, W.A.; Lettice, L.A. Developmentally regulated Shh expression is robust to TAD perturbations. Development 2019, 146, dev179523. [Google Scholar] [CrossRef]
- Shevelyov, Y.Y.; Ulianov, S.V. The Nuclear Lamina as an Organizer of Chromosome Architecture. Cells 2019, 8, 136. [Google Scholar] [CrossRef] [PubMed]
- Pickersgill, H.; Kalverda, B.; de Wit, E.; Talhout, W.; Fornerod, M.; van Steensel, B. Characterization of the Drosophila melanogaster genome at the nuclear lamina. Nat. Genet. 2006, 38, 1005–1014. [Google Scholar] [CrossRef] [PubMed]
- Peric-Hupkes, D.; Meuleman, W.; Pagie, L.; Bruggeman, S.W.M.; Solovei, I.; Brugman, W.; Gräf, S.; Flicek, P.; Kerkhoven, R.M.; van Lohuizen, M.; et al. Molecular Maps of the Reorganization of Genome-Nuclear Lamina Interactions during Differentiation. Mol. Cell 2010, 38, 603–613. [Google Scholar] [CrossRef] [PubMed]
- Nikolakaki, E.; Mylonis, I.; Giannakouros, T. Lamin B Receptor: Interplay between Structure, Function and Localization. Cells 2017, 6, 28. [Google Scholar] [CrossRef] [PubMed]
- Pindyurin, A.V.; Ilyin, A.A.; Ivankin, A.V.; Tselebrovsky, M.V.; Nenasheva, V.V.; Mikhaleva, E.A.; Pagie, L.; van Steensel, B.; Shevelyov, Y.Y. The large fraction of heterochromatin in Drosophila neurons is bound by both B-type lamin and HP1a. Epigenet. Chromatin 2018, 11, 65. [Google Scholar] [CrossRef] [PubMed]
- Solovei, I.; Wang, A.S.; Thanisch, K.; Schmidt, C.S.; Krebs, S.; Zwerger, M.; Cohen, T.V.; Devys, D.; Foisner, R.; Peichl, L.; et al. LBR and Lamin A/C Sequentially Tether Peripheral Heterochromatin and Inversely Regulate Differentiation. Cell 2013, 152, 584–598. [Google Scholar] [CrossRef]
- Zhu, J.; Adli, M.; Zou, J.Y.; Verstappen, G.; Coyne, M.; Zhang, X.; Durham, T.; Miri, M.; Deshpande, V.; De Jager, P.L.; et al. Genome-wide Chromatin State Transitions Associated with Developmental and Environmental Cues. Cell 2013, 152, 642–654. [Google Scholar] [CrossRef]
- Kind, J.; Pagie, L.; de Vries, S.S.; Nahidiazar, L.; Dey, S.S.; Bienko, M.; Zhan, Y.; Lajoie, B.; de Graaf, C.A.; Amendola, M.; et al. Genome-wide Maps of Nuclear Lamina Interactions in Single Human Cells. Cell 2015, 163, 134–147. [Google Scholar] [CrossRef]
- Briand, N.; Collas, P. Lamina-associated domains: Peripheral matters and internal affairs. Genome Biol. 2020, 21, 85. [Google Scholar] [CrossRef]
- Somech, R.; Shaklai, S.; Geller, O.; Amariglio, N.; Simon, A.J.; Rechavi, G.; Gal-Yam, E.N. The nuclear-envelope protein and transcriptional repressor LAP2β interacts with HDAC3 at the nuclear periphery, and induces histone H4 deacetylation. J. Cell Sci. 2005, 118, 4017–4025. [Google Scholar] [CrossRef]
- Demmerle, J.; Koch, A.J.; Holaska, J.M. The Nuclear Envelope Protein Emerin Binds Directly to Histone Deacetylase 3 (HDAC3) and Activates HDAC3 Activity. J. Biol. Chem. 2012, 287, 22080–22088. [Google Scholar] [CrossRef] [PubMed]
- Milon, B.C.; Cheng, H.; Tselebrovsky, M.V.; Lavrov, S.A.; Nenasheva, V.V.; Mikhaleva, E.A.; Shevelyov, Y.Y.; Nurminsky, D.I. Role of Histone Deacetylases in Gene Regulation at Nuclear Lamina. PLoS ONE 2012, 7, e49692. [Google Scholar] [CrossRef] [PubMed]
- Verboon, J.M.; Rincon-Arano, H.; Werwie, T.R.; Delrow, J.J.; Scalzo, D.; Nandakumar, V.; Groudine, M.; Parkhurst, S.M. Wash Interacts with Lamin and Affects Global Nuclear Organization. Curr. Biol. 2015, 25, 804–810. [Google Scholar] [CrossRef] [PubMed]
- Ulianov, S.V.; Doronin, S.A.; Khrameeva, E.E.; Kos, P.I.; Luzhin, A.V.; Starikov, S.S.; Galitsyna, A.A.; Nenasheva, V.V.; Ilyin, A.A.; Flyamer, I.M.; et al. Nuclear lamina integrity is required for proper spatial organization of chromatin in Drosophila. Nat. Commun. 2019, 10, 1176. [Google Scholar] [CrossRef]
- Akhtar, W.; de Jong, J.; Pindyurin, A.V.; Pagie, L.; Meuleman, W.; de Ridder, J.; Berns, A.; Wessels, L.F.A.; van Lohuizen, M.; van Steensel, B. Chromatin Position Effects Assayed by Thousands of Reporters Integrated in Parallel. Cell 2013, 154, 914–927. [Google Scholar] [CrossRef]
- Shevelyov, Y.Y.; Lavrov, S.A.; Mikhaylova, L.M.; Nurminsky, I.D.; Kulathinal, R.J.; Egorova, K.S.; Rozovsky, Y.M.; Nurminsky, D.I. The B-type lamin is required for somatic repression of testis-specific gene clusters. Proc. Natl. Acad. Sci. USA 2009, 106, 3282–3287. [Google Scholar] [CrossRef]
- Kind, J.; Pagie, L.; Ortabozkoyun, H.; Boyle, S.; de Vries, S.S.; Janssen, H.; Amendola, M.; Nolen, L.D.; Bickmore, W.A.; van Steensel, B. Single-Cell Dynamics of Genome-Nuclear Lamina Interactions. Cell 2013, 153, 178–192. [Google Scholar] [CrossRef]
- Poleshko, A.; Shah, P.P.; Gupta, M.; Babu, A.; Morley, M.P.; Manderfield, L.J.; Ifkovits, J.L.; Calderon, D.; Aghajanian, H.; Sierra-Pagán, J.E.; et al. Genome-Nuclear Lamina Interactions Regulate Cardiac Stem Cell Lineage Restriction. Cell 2017, 171, 573–587.e14. [Google Scholar] [CrossRef]
- van Steensel, B.; Belmont, A.S. Lamina-Associated Domains: Links with Chromosome Architecture, Heterochromatin, and Gene Repression. Cell 2017, 169, 780–791. [Google Scholar] [CrossRef]
- Tang, S.-J. Chromatin Organization by Repetitive Elements (CORE): A Genomic Principle for the Higher-Order Structure of Chromosomes. Genes 2011, 2, 502–515. [Google Scholar] [CrossRef]
- van de Werken, H.J.G.; Haan, J.C.; Feodorova, Y.; Bijos, D.; Weuts, A.; Theunis, K.; Holwerda, S.J.B.; Meuleman, W.; Pagie, L.; Thanisch, K.; et al. Small chromosomal regions position themselves autonomously according to their chromatin class. Genome Res. 2017, 27, 922–933. [Google Scholar] [CrossRef] [PubMed]
- Larson, A.G.; Elnatan, D.; Keenen, M.M.; Trnka, M.J.; Johnston, J.B.; Burlingame, A.L.; Agard, D.A.; Redding, S.; Narlikar, G.J. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 2017, 547, 236–240. [Google Scholar] [CrossRef]
- Strom, A.R.; Emelyanov, A.V.; Mir, M.; Fyodorov, D.V.; Darzacq, X.; Karpen, G.H. Phase separation drives heterochromatin domain formation. Nature 2017, 547, 241–245. [Google Scholar] [CrossRef] [PubMed]
- Falk, M.; Feodorova, Y.; Naumova, N.; Imakaev, M.; Lajoie, B.R.; Leonhardt, H.; Joffe, B.; Dekker, J.; Fudenberg, G.; Solovei, I.; et al. Heterochromatin drives compartmentalization of inverted and conventional nuclei. Nature 2019, 570, 395–399. [Google Scholar] [CrossRef] [PubMed]
- Riddle, N.C.; Minoda, A.; Kharchenko, P.V.; Alekseyenko, A.A.; Schwartz, Y.B.; Tolstorukov, M.Y.; Gorchakov, A.A.; Jaffe, J.D.; Kennedy, C.; Linder-Basso, D.; et al. Plasticity in patterns of histone modifications and chromosomal proteins in Drosophila heterochromatin. Genome Res. 2011, 21, 147–163. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.D.; Shu, S.; Mungall, C.J.; Karpen, G.H. The Release 5.1 Annotation of Drosophila melanogaster Heterochromatin. Science 2007, 316, 1586–1591. [Google Scholar] [CrossRef]
- Lee, Y.C.G.; Ogiyama, Y.; Martins, N.M.C.; Beliveau, B.J.; Acevedo, D.; Wu, C.-T.; Cavalli, G.; Karpen, G.H. Pericentromeric heterochromatin is hierarchically organized and spatially contacts H3K9me2 islands in euchromatin. PLoS Genet. 2020, 16, e1008673. [Google Scholar] [CrossRef]
- Jagannathan, M.; Cummings, R.; Yamashita, Y.M. The modular mechanism of chromocenter formation in Drosophila. eLife 2019, 8, e43938. [Google Scholar] [CrossRef]
- Zhang, P.; Spradling, A.C. The Drosophila salivary gland chromocenter contains highly polytenized subdomains of mitotic heterochromatin. Genetics 1995, 139, 659–670. [Google Scholar] [CrossRef]
- Jones, K.W. Chromosomal and Nuclear Location of Mouse Satellite DNA in Individual Cells. Nature 1970, 225, 912–915. [Google Scholar] [CrossRef] [PubMed]
- Zenk, F.; Zhan, Y.; Kos, P.; Löser, E.; Atinbayeva, N.; Schächtle, M.; Tiana, G.; Giorgetti, L.; Iovino, N. HP1 drives de novo 3D genome reorganization in early Drosophila embryos. Nature 2021, 593, 289–293. [Google Scholar] [CrossRef] [PubMed]
- Erdel, F.; Rademacher, A.; Vlijm, R.; Tünnermann, J.; Frank, L.; Weinmann, R.; Schweigert, E.; Yserentant, K.; Hummert, J.; Bauer, C.; et al. Mouse Heterochromatin Adopts Digital Compaction States without Showing Hallmarks of HP1-Driven Liquid-Liquid Phase Separation. Mol. Cell 2020, 78, 236–249.e7. [Google Scholar] [CrossRef] [PubMed]
- Beck, M.; Hurt, E. The nuclear pore complex: Understanding its function through structural insight. Nat. Rev. Mol. Cell Biol. 2017, 18, 73–89. [Google Scholar] [CrossRef]
- Vaquerizas, J.M.; Suyama, R.; Kind, J.; Miura, K.; Luscombe, N.M.; Akhtar, A. Nuclear Pore Proteins Nup153 and Megator Define Transcriptionally Active Regions in the Drosophila Genome. PLoS Genet. 2010, 6, e1000846. [Google Scholar] [CrossRef] [PubMed]
- Gozalo, A.; Duke, A.; Lan, Y.; Pascual-Garcia, P.; Talamas, J.A.; Nguyen, S.C.; Shah, P.P.; Jain, R.; Joyce, E.F.; Capelson, M. Core Components of the Nuclear Pore Bind Distinct States of Chromatin and Contribute to Polycomb Repression. Mol. Cell 2020, 77, 67–81.e7. [Google Scholar] [CrossRef]
- Capelson, M.; Liang, Y.; Schulte, R.; Mair, W.; Wagner, U.; Hetzer, M.W. Chromatin-Bound Nuclear Pore Components Regulate Gene Expression in Higher Eukaryotes. Cell 2010, 140, 372–383. [Google Scholar] [CrossRef]
- Kalverda, B.; Pickersgill, H.; Shloma, V.V.; Fornerod, M. Nucleoporins Directly Stimulate Expression of Developmental and Cell-Cycle Genes Inside the Nucleoplasm. Cell 2010, 140, 360–371. [Google Scholar] [CrossRef]
- Pascual-Garcia, P.; Jeong, J.; Capelson, M. Nucleoporin Nup98 Associates with Trx/MLL and NSL Histone-Modifying Complexes and Regulates Hox Gene Expression. Cell Rep. 2014, 9, 433–442. [Google Scholar] [CrossRef]
- Pascual-Garcia, P.; Debo, B.; Aleman, J.R.; Talamas, J.A.; Lan, Y.; Nguyen, N.H.; Won, K.J.; Capelson, M. Metazoan Nuclear Pores Provide a Scaffold for Poised Genes and Mediate Induced Enhancer-Promoter Contacts. Mol. Cell 2017, 66, 63–76. [Google Scholar] [CrossRef]
- Ibarra, A.; Benner, C.; Tyagi, S.; Cool, J.; Hetzer, M.W. Nucleoporin-mediated regulation of cell identity genes. Genes Dev. 2016, 30, 2253–2258. [Google Scholar] [CrossRef]
- Tan-Wong, S.M.; Wijayatilake, H.D.; Proudfoot, N.J. Gene loops function to maintain transcriptional memory through interaction with the nuclear pore complex. Genes Dev. 2009, 23, 2610–2624. [Google Scholar] [CrossRef]
- Strom, A.R.; Brangwynne, C.P. The liquid nucleome—Phase transitions in the nucleus at a glance. J. Cell Sci. 2019, 132, jcs235093. [Google Scholar] [CrossRef]
- Banani, S.F.; Lee, H.O.; Hyman, A.A.; Rosen, M.K. Biomolecular condensates: Organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 2017, 18, 285–298. [Google Scholar] [CrossRef]
- King, J.T.; Shakya, A. Phase separation of DNA: From past to present. Biophys. J. 2021, 120, 1139–1149. [Google Scholar] [CrossRef]
- Gibson, B.A.; Doolittle, L.K.; Schneider, M.W.G.; Jensen, L.E.; Gamarra, N.; Henry, L.; Gerlich, D.W.; Redding, S.; Rosen, M.K. Organization of Chromatin by Intrinsic and Regulated Phase Separation. Cell 2019, 179, 470–484.e21. [Google Scholar] [CrossRef] [PubMed]
- Plys, A.J.; Davis, C.P.; Kim, J.; Rizki, G.; Keenen, M.M.; Marr, S.K.; Kingston, R.E. Phase separation of polycomb-repressive complex 1 is governed by a charged disordered region of CBX2. Genes Dev. 2019, 33, 799–813. [Google Scholar] [CrossRef] [PubMed]
- Lau, M.S.; Schwartz, M.G.; Kundu, S.; Savol, A.J.; Wang, P.I.; Marr, S.K.; Grau, D.J.; Schorderet, P.; Sadreyev, R.I.; Tabin, C.J.; et al. Mutation of a nucleosome compaction region disrupts Polycomb-mediated axial patterning. Science 2017, 355, 1081–1084. [Google Scholar] [CrossRef]
- Whyte, W.A.; Orlando, D.A.; Hnisz, D.; Abraham, B.J.; Lin, C.Y.; Kagey, M.H.; Rahl, P.B.; Lee, T.I.; Young, R.A. Master Transcription Factors and Mediator Establish Super-Enhancers at Key Cell Identity Genes. Cell 2013, 153, 307–319. [Google Scholar] [CrossRef] [PubMed]
- Pott, S.; Lieb, J.D. What are super-enhancers? Nat. Genet. 2015, 47, 8–12. [Google Scholar] [CrossRef]
- Sabari, B.R.; Dall’Agnese, A.; Boija, A.; Klein, I.A.; Coffey, E.L.; Shrinivas, K.; Abraham, B.J.; Hannett, N.M.; Zamudio, A.V.; Manteiga, J.C.; et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 2018, 361, eaar3958. [Google Scholar] [CrossRef]
- Cho, W.-K.; Spille, J.-H.; Hecht, M.; Lee, C.; Li, C.; Grube, V.; Cisse, I.I. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 2018, 361, 412–415. [Google Scholar] [CrossRef] [PubMed]
- Mir, M.; Stadler, M.R.; Ortiz, S.A.; Hannon, C.E.; Harrison, M.M.; Darzacq, X.; Eisen, M.B. Dynamic multifactor hubs interact transiently with sites of active transcription in Drosophila embryos. eLife 2018, 7, e40497. [Google Scholar] [CrossRef] [PubMed]
- Boehning, M.; Dugast-Darzacq, C.; Rankovic, M.; Hansen, A.S.; Yu, T.; Marie-Nelly, H.; McSwiggen, D.T.; Kokic, G.; Dailey, G.M.; Cramer, P.; et al. RNA polymerase II clustering through carboxy-terminal domain phase separation. Nat. Struct. Mol. Biol. 2018, 25, 833–840. [Google Scholar] [CrossRef] [PubMed]
- Boija, A.; Klein, I.A.; Sabari, B.R.; Dall’Agnese, A.; Coffey, E.L.; Zamudio, A.V.; Li, C.H.; Shrinivas, K.; Manteiga, J.C.; Hannett, N.M.; et al. Transcription Factors Activate Genes through the Phase-Separation Capacity of Their Activation Domains. Cell 2018, 175, 1842–1855. [Google Scholar] [CrossRef] [PubMed]
- Cardozo Gizzi, A.M.; Cattoni, D.I.; Fiche, J.-B.; Espinola, S.M.; Gurgo, J.; Messina, O.; Houbron, C.; Ogiyama, Y.; Papadopoulos, G.L.; Cavalli, G.; et al. Microscopy-Based Chromosome Conformation Capture Enables Simultaneous Visualization of Genome Organization and Transcription in Intact Organisms. Mol. Cell 2019, 74, 212–222. [Google Scholar] [CrossRef] [PubMed]
- Cardozo Gizzi, A.M.; Espinola, S.M.; Gurgo, J.; Houbron, C.; Fiche, J.-B.; Cattoni, D.I.; Nollmann, M. Direct and simultaneous observation of transcription and chromosome architecture in single cells with Hi-M. Nat. Protoc. 2020, 15, 840–876. [Google Scholar] [CrossRef]
- Askjaer, P.; Harr, J.C. Genetic approaches to revealing the principles of nuclear architecture. Curr. Opin. Genet. Dev. 2021, 67, 52–60. [Google Scholar] [CrossRef]
- Jerkovic, I.; Cavalli, G. Understanding 3D genome organization by multidisciplinary methods. Nat. Rev. Mol. Cell Biol. 2021, 22, 511–528. [Google Scholar] [CrossRef]
- Nguyen, H.Q.; Chattoraj, S.; Castillo, D.; Nguyen, S.C.; Nir, G.; Lioutas, A.; Hershberg, E.A.; Martins, N.M.C.; Reginato, P.L.; Hannan, M.; et al. 3D mapping and accelerated super-resolution imaging of the human genome using in situ sequencing. Nat. Methods 2020, 17, 822–832. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).