Abstract
The immune system has evolved to protect organisms from infections caused by bacteria, viruses, and parasitic pathogens. In addition, it provides regenerative capacities, tissue maintenance, and self/non-self recognition of foreign tissues. Phagocytosis and cytotoxicity are two prominent cellular immune activities positioned at the base of immune effector function in mammals. Although these immune mechanisms have diversified into a wide heterogeneous repertoire of effector cells, it appears that they share some common cellular and molecular features in all animals, but also some interesting convergent mechanisms. In this review, we will explore the current knowledge about the evolution of phagocytic and cytotoxic immune lineages against pathogens, in the clearance of damaged cells, for regeneration, for histocompatibility recognition, and in killing virally infected cells. To this end, we give different immune examples of multicellular organism models, ranging from the roots of bilateral organisms to chordate invertebrates, comparing to vertebrates’ lineages. In this review, we compare cellular lineage homologies at the cellular and molecular levels. We aim to highlight and discuss the diverse function plasticity within the evolved immune effector cells, and even suggest the costs and benefits that it may imply for organisms with the meaning of greater defense against pathogens but less ability to regenerate damaged tissues and organs.
1. Introduction
Over hundreds of millions of years, the development of defense mechanisms in multicellular organisms to fight the invasion of microbes and viruses has represented an essential cue for the evolution of life. Not surprisingly, immune systems have diversified into a highly complex set of cells and molecular mechanisms to target pathogens and also to allow regeneration, tissue repair and the recognition of foreign transplants.
Evolution of animal immune systems can be analyzed during a time span of at least 1000 million years. From the most primitive multicellular organisms (i.e., sponges) to protostomes organisms (i.e., Platyhelminthes, mollusks, nematodes and arthropods) to chordate invertebrates (i.e., Tunicata) defense and self/non-self-recognition responses rely completely on innate immunity. Jawless vertebrates such as hagfish and lampreys were the first to connect innate responses with some adaptive elements. Whereas all jawed vertebrates ranging from primitive sharks to mammals are equipped with innate and adaptive immunity [1,2].
In general, innate immunity is a fast and non-specific response associated with the presence of humoral and cellular elements [3]. By contrast, adaptive immunity uses the induction of specialized cells such as B and T lymphocytes and molecules including the major histocompatibility complex (MHC), B-cell receptors (BCR), T-cell receptors (TCR), immunoglobulins (Ig), and antibodies to confer immunological memory and very high specificity thus fighting against a previously recognized infection [4]. Both kinds of immune responses rely on two main cellular activities which are phagocytosis and cytotoxicity. These cellular immune mechanisms have been found at the earliest evolutionary stages of multicellular animals and diversified into a wide heterogeneous repertoire of effector cells through evolution.
Here, we will review how phagocytic and cytotoxic effector cell lineages have evolved against pathogens, in histocompatibility recognition and in killing virally infected cells, highlighting the common molecular features shared between the more evolved species with their early invertebrate ancestors. We emphasize on multicellular organisms, where we assume existence of specialized immune effector cells subpopulations. Finally, we aim to discuss the functional plasticity of these immune cells in the clearance of damaged cells, and their repercussions on regeneration and tissue repair capacities.
2. Phagocytosis in the Evolution of Immune Response
Phagocytosis is the process by which particles (>0.5 μm) are recognized, bound to a plasma-membrane envelope, and internalized into an organelle called “phagosome”. Two major classes of particles are engulfed by phagocytes: foreign microorganisms and “altered self” (apoptotic and necrotic cells) particles. Élie Metchnikoff was the first to realize the significance of phagocytosis for cellular immunity in processes such as the host response to injury and infection, inflammation, and tissue homeostasis [5,6,7]. Phagocytosis is highly conserved between species and has essential roles for the development and maintenance of multicellular organisms. The phagocytic process requires a coordinated sequence of events that begins with the engagement of plasma-membrane receptors, which allow the recognition of damaged or infected cells and different classes of pathogens. This leads to the activation of signaling pathways needed for the rearrangement of the cytoskeleton and the internalization of the particle. Once internalized, the phagosome dynamically undergoes fusion and fission events with intracellular secretory vesicles to finally mature into a phagolysosome. The hydrolytic activity of phagolysosomes results in the digestion of the internalized cargo [5].
In invertebrates, “professional phagocytes” have a macrophage-like phenotype and express receptors that recognize pathogen-associated molecules. Ligand-receptor binding induces different signaling cascades leading to the production of effector molecules to eliminate or inactivate the infection. These include oxygen-dependent (superoxide anion, hydrogen peroxide, hydroxyl radical) and oxygen-independent (lysozyme and defensins) molecules [8]. Phagocytic cell lineages and mechanisms to engulf and combat microbes and viruses have been diversified in vertebrates, but have several homologous molecules and cells with their lower ancestors [2,8].
2.1. Phagocytic Effector Cells
Amebocytes are the most primitive animal cells with phagocytic ability, these are found in sponges and cnidarians acting in the recognition of nutrients and foreign elements [9]. We have recently characterized two populations of phagocytes (granular spheroid and ameboid phenotype) from members of the Hexacorallia class, the coral Pocillopora damicornis and the sea anemone Nematostella vectensis, finding that they engulf bacteria, fungal antigens, carboxylated beads and self-damaged cells. We showed that this phagocytosis is different than pinocytosis which is performed by the majority of the cells [10]. Reticular cells, the phagocytic cells of free living planarians from the phylum Platyhelminthes, can migrate towards heat-shocked bacteria and phagocytose them. Their pseudopodia and the modes of their endocytosis can be morphologically compared to human neutrophils [11,12]. From the phylum Annelida, the earthworm Eisenia andrei has several subtypes of coelomocytes including eleocytes and granular amoebocytes [13]. In arthropods, several cell types have been characterized. For example, Drosophila melanogaster has hemolymph circulating cells collectively called hemocytes. Two types of hemocytes have been identified in the larval stage: Plasmatocytes, which phagocytose microorganisms and cell debris and secrete signaling molecules (i.e., Eiger, Upds, drosocin, defensins) with high similarity to mammalian macrophages and neutrophils, according to transcriptional profiles; and crystal cells, which activate the melanization cascade upon wounds or infections [14]. However, single-cell RNA sequencing studies showed that these cell types are more heterogeneous as 14 different functional clusters of hematocytes have been recently discovered, including lamellocytes, which are activated immune cells that only differentiate upon immune induction [15]. The tunicate Botryllus schlosseri has a myeloid lineage of phagocytic cells with high similarities and gene set homology to mammalian myeloid lineage [16], and also amoebocytes and large phagocytes suggested to be related to arthropods and echinoderms [17]. In bony fishes, the phagocytic armament includes macrophages, monocytes, dendritic cells, neutrophils, granulocytes, eosinophils, basophils and mast cells [18]. While in mammals, these professional phagocytes are able to differentiate into highly specialized subtypes of cells which exert different cytokine profiles, exclusive functions in regeneration and infection fighting (i.e., M1 and M2 macrophages), tissue specificity and also novel molecular mechanisms such as the antibody-dependent cellular phagocytosis (ADCP) for the clearance of microorganisms and immune complexes [19,20].
2.2. Receptors
Phagocytic cell lineages possess cell-surface receptors as the mechanism to respond and initiate the phagocytic process and the production of killing molecules. Primitive sponges possess LPS binding receptors and intracellular receptors “Nucleotide-binding domain and Leucine-rich Repeat” (NLRs) to bind fungal polysaccharides, bacteria and virus [21,22]. Toll-like receptors (TLRs) are ancient sensors and the best characterized in detecting and respond against invading pathogens. TLRs have been traced to several invertebrate species as their emergence predate the separation of bilaterians and cnidarians. In D. melanogaster, Toll receptors have a crucial role in immune defense. Flies deficient in Toll members are severely immunocompromised to response against fungal and bacterial infections [23]. Moreover, the Down syndrome cell-adhesion molecule (DsCAM) acts as a phagocytic cell-surface receptor that binds to bacteria such as Escherichia coli [24]. A new study in B. schlosseri has identified BsTLR1 as a member of the TLR family which is actively transcribed in phagocytes and morula cells as a mechanism of non-self recognition [25]. Interestingly, TLRs are highly diversified in urchins (253 genome sequences), more than in vertebrates [26]. Agnathans have seven identified TLRs, while in mammals this number is up to 13, which have high specificity in the ligands they recognize, suggesting a host–ligand coevolution [27]. Although TLRs have not been identified in planarians, extracellular leucine rich repeat (LRR)-motifs are coded in their genome [28]; however, the exact role LRRs play in the immune response is unknow. Other types of receptors are highly conserved through evolution. For example, the scavenger receptors Croquemort (Crq) and Peste of D. melanogaster have convergently evolved with their mammalian orthologues CD36 and SR-BI, respectively [29]. There are also three predicted CD36-family homologs in the genome of Caenorhabditis elegans [30]. They are CD36-like family members and major plasmatocyte markers that recognize apoptotic cells and bind to surface lipopetides of different Gram-negative and Gram-positive bacteria and fungi. Both are required to efficiently eliminate infections, and their absence causes poor phagocytic plasmatocytes and defects on phagosome maturation [31]. The Transformer (Trf) proteins also have important roles in the immunity of sea urchins by inducing the phagocytic activity of coelomocytes to directly engulf bacteria in an ADCP resembling matter [32,33]. SpTrf genes have two exons, recombination mechanisms between the gene elements of the second exon result in the fact that sea urchins express more than 260 different Trfs proteins, single one recombinant protein expressed in an individual cell, which act synergistically to detect a variety of pathogens thus providing an efficient immunological army [34,35]. Furthermore, no Trfs have been identified outside the echinoid lineage [36], indicating that there are several unique immune mechanisms that should be studied in sea urchins. Moreover, this discovery suggests that there are diverse mechanisms of immune receptor diversification, which are different from what we know in vertebrates.
2.3. Effector Molecules
After the accumulation of phagocytes in a localized area of physical and chemical insult, these release several molecules to allow recognition, pathogen killing, and inflammation. In invertebrates these molecules include oxidative killing (ROS and NOS species), agglutination, clotting, coagulation (i.e., fibrinogen-related peptides FREPS), melanization (cytotoxic quinones, melanin) and antimicrobial peptides. In the hexacorallian P. damicornis and N. vectensis, a high production of ROS is associated with phagocytic activity [10]. In the protochordate B. schlosseri the rhamnose-binding lectin (BsRBL) activate phagocytes and induces the synthesis and release of cytokines recognized by the antibodies anti-IL1α and anti-TNFα [37]. Mammalian phagocytes secrete a wide array of chemokines (CXC) and cytokines. Cytokines have been divided into different classes: interferons (IFNs), interleukins (ILs), tumor necrosis factors (TNFs) and transforming growth factors (TGFs). Studies have been conducted to search for homologs in lower species. Some worms and mollusks have shown to enhance phagocytosis in response to the cytokines TNF-α and IFN-γ, but no encoded genes have been characterized to explain their function [2]. Drosophila genome encodes different cytokine types with certain mammalian homology: Eiger is homolog to TNF, Upds is similar to type I cytokines and Spätzle is related to IL-17F. Flies with mutations in these genes are highly sensitive to bacterial infections and have less capacity to remove dead cells, showing its important role in innate immunity [38]. Transforming growth factor β (TGF-β) modulates inflammation and repair functions. The TGF-β family member, tgfβ-f, has been identified in B. schlosseri although its principal knowing function is related with germ cell differentiation [39]. Genes of all the major mammalian cytokines and CXC have been identified in fishes. These include pro-inflammatory cytokines IL-1 β, TNF-α and IL-6 and cytokines associated with the adaptive immunity such as IL-2, IFN-γ, IL-4/13 (homologue to IL-4 and IL13), IL-10, IL-17A/F, IL-21, IL-22 and TGF-β family members, providing strong protection against diverse viral infections (i.e., haemorrhaic virus) [40]. Although cytokines are highly conserved among vertebrates, distinct differences have been identified between classes. For example, IFN-γ is the only type II IFN identified in mammals, whereas multiple type II IFNs have been found in lower vertebrates. In contrast, type III IFNs appear to be unique to mammals. IL-1, IL-2 and IL-8 are the most conserved interleukins among vertebrates, while IL-4, IL-5, IL-10, IL-13, IL-20, IL-21 and others have been only characterized in mice and humans with very specific biological functions [41].
2.4. Signaling Pathways
The coordination of the phagocytic process in an integrated network of signaling cascades must have occurred since the most primitive forms of life (Figure 1). We have stated that TLRs are found in phagosomes of many invertebrates. The Toll pathway activates the nuclear factor-κB (NFκB) signaling, a master regulator of the phagolysosomal degradation of pathogens. Genome and transcriptome analysis have shown that homologs of mammalian NFκB transcription factor, its inhibitors (IκB) and activators are traced to sponges. In the demosponge Amphimedon queenslandica the Aq-NF-κB has been identified, this is structurally similar to NFκB p100/p105 among vertebrate subfamily transcription factors with DNA binding domains and a C-terminal that allows its translocation to the nucleus [42]. Homolog transcripts of upstream NFκB pathway components have been also identified in cnidarians. In Drosophila, Dorsal/DIF and Cactus has been described as homologs to NFκB and IκB, and several NFκB binding motifs were identified in promoters of immunoresponsive genes of phagocytic cells, suggesting its conserved role in the immune response (Figure 1) [43]. Furthermore, phagocytes of B. schlosseri activate Ras-like small GTPases, MAPKs and NFκB signaling networks as a response to the recognition of foreign cells (Figure 1) [44]. In mammalian macrophages, NFκB signaling is highly dynamic, specific, and strictly regulated acting in direct and indirect crosstalk with many other signaling pathways during an immune response, depending on the stimuli given by each interacting pathogen. This complexity allows macrophages and several other cell types to have diverse mechanisms of action thus mounting an effective immune response and in parallel protecting the tissues of possible damages caused by hyperinflammation (Figure 1) [45].
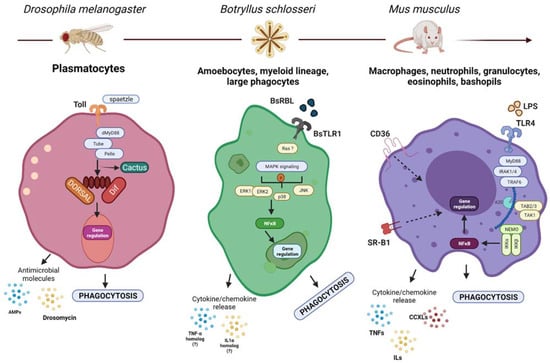
Figure 1.
Molecular mechanisms guiding phagocytosis in different organisms. Although the phagocytic lineages and their molecular mechanisms guiding the process of phagocytosis could be different, they share several basic mechanisms with their ancestral form. Shown here: (left) the Toll pathway in Drosophila is activated by spaetzle, leading to an intracellular signaling that results in the degradation of cactus and the concomitant release of DIF/DORSAL which translocate to the nucleus and activate the transcription of immune genes including AMPs and drosomycin, which secreted as a response to infection [46]. (center) BsTLR1, Ras, MAPK, NFκB and cytokine homologs have been associated with the modulation of phagocytosis in B. Schlosseri. (?)—Represents missing functional or molecular validation. Similarly (right), mammalian TLRs are activated by microbial- and self-derived products leading to the activation of intracellular signaling cascade that results in the translocation of NFκB to the nucleus to activate the transcription of different cytokines and chemokines which enhance the phagocytic activity of the cells [47]. Created with BioRender.com.
3. Cellular Cytotoxicity in the Evolution of Immune Response against Abnormal Self and Pathogens
Cytotoxicity (cell-mediated killing) is the process in which certain cells secrete “toxins” that lysate and neutralize unwanted target cells (i.e., pathogens, viral-infected cells, tumoral cells, foreign transplanted cells). Cooper has defined that through evolution, immune-mediated cytotoxicity could be classified into level I and level II cytotoxicity. Level I in mammals is mediated by macrophages and neutrophils which release toxic granules in the inflammatory area. They are considered the most basic manifestations of cytotoxicity and are present in invertebrates (sponges, coelenterates, sipunculids, annelids, mollusks, arthropods, echinoderms and protochordates). In contrast, level II or acquired cytotoxicity is mediated by cells that require a specific induction to be activated, through specific recognition, such as allogeneic responses, or induced by adaptive molecules. These include cytotoxic T lymphocytes (CTL), natural killer (NK), and antibody-dependent cell-mediated cytotoxicity (ADCC). It seems that these cells appear later in evolution [2,48], but the characterization of specific allogeneic responses suggests that they might be in more animal groups than what is currently known. Since the work of Cooper in 1980, NK cells were shown to have specificity and recognition of targets for cytotoxicity trough immunological synapse without the killing of adjacent cells [49,50], for that reason we moved NK-like recognition based cytotoxicity mechanism to level II cytotoxicity, compared to the original definition by Cooper [48].
All cytotoxic cells share certain characteristics such as (1) the ability to trigger cell death of target cells, (2) the presence of adhesion molecules allowing their adhesion to the target, (3) their activity is mediated by monokines and lymphokines [51].
3.1. Effector Cells
All animals have several immune cells with cytotoxic activity to lyse host and foreign cells. Regarding level I cytotoxicity, those cells have been found in different invertebrate organisms as probably the most ancient type of cytotoxic immune response. For example, killer cells from sipunculid worms exert cytotoxic effects on allogeneic erythrocytes, but not on erythrocytes from worms that reside closely [52]. In the freshwater pulmonated mollusk Planorbarius corneus, a class of hemocyte that has been morphologically characterized as round hemocytes (RH) exerts cytotoxic activity on the human erythroleukemia K562 cells thus participating in the graft rejection of allo- and xeno-grafts [51]. In arthropods, the amoeboid hemocytes of Limolus polyphemus are stimulated by bacterial lipopolysaccharide (LPS) endotoxins to secrete great amounts of clottable protein and to produce nitric oxide (NO) synthesis as a cytotoxic mechanism to protect the host from invading pathogens [53]. Additionally, in in vitro experiments, crayfish (Astacus astacus) granular and semi granular hemocytes with phenoloxidase and laccase activity display cytotoxic effects towards different mammalian tumor and non-tumor cells [54,55]. Coelomocytes from the sea urchin Arbacia punctulata exert cytotoxic activity against human and murine target cells. A particular population of these phagocytic cells is the one with the highest cytotoxic activity which are positive to the human NK markers CD14, CD56 and CD158b [56]. These cytotoxic cells have also found in Strongylocentrotus droebachiensis, S. pallidus and Echinus esculentus in which targeted-cell death is characterized by cell detachment, disintegration and the formation of a multinuclear non-cellular protoplasm [57].
At this point we do not think that phagocytosis of damaged self-cells should be considered as level 1 cytotoxicity. Although, we do think that where phagocytosis of damaged self-cells is found, it can be suggestive of potential level 1 cytotoxicity (as the mechanisms of altered-self recognition exist) and should be further investigated (Figure 2). For instance, the recent discovery of phagocytosis of heat-stressed cells in ex-vivo experiments in Hexacorallians [10].
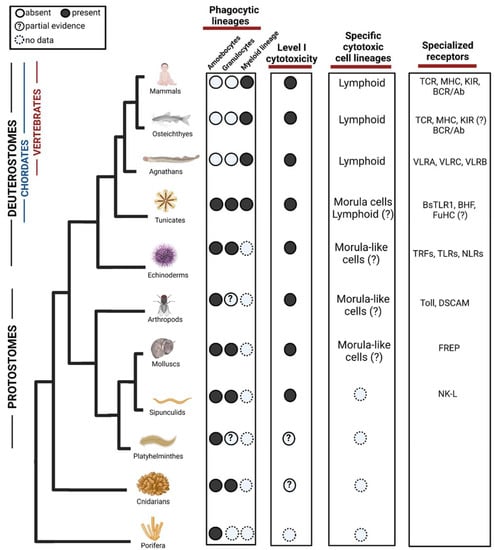
Figure 2.
Evolution of phagocytic and cytotoxic cell lineages in the immune system. Among phagocytic lineages, amoebocytes and granulocytes are present in invertebrates, while myeloid lineage seems to evolve in the ancestor of tunicates and vertebrates. Cells representing the Level I cytotoxicity exist in Protostomes but might appear in cnidarian. Among specific cytotoxic cell lineages, morula cells have been found in tunicates although partial evidence suggests that they may evolve in earlier ancestors. Lymphoid cells are found in agnathans and have diversified in vertebrates, there are suggestions of their form in Tunicates but not validated yet. In the third column, we show examples of diversification of specialized receptors acting in both phagocytosis and cytotoxicity. TLR = Toll-like receptors, TCR = T-cell receptor, MHC = Major histocompatibility complex, KIR = Killer Inhibitory Receptors, BCR = B cell receptor, VLR = Variable lymphocyte receptor, BHF = Botryllus histocompatibility factor, TRF = Transformer proteins, NLR = NOD like receptors, DSCAM = Down syndrome cell adhesion molecule, FREP = Fibrinogen-related proteins, NK-L = Natural killer-like receptor. ?—Represents missing functional or molecular validation. Created with BioRender.com. Data sources are summarized in Table 1.
For level II of specific cytotoxicity, in the tunicate B. schlosseri, morula cells (MC) have been characterized as cytotoxic cells. They contain phenoloxidase, accumulate in rejection points and morphologically resembles NK cells. While 15% of their genes are shared with cytotoxic lymphocytes of vertebrates, MC express 85% tunicate-specific gene repertoire [16,17], suggestive of convergent lineage more resembling other invertebrates phenoloxidase based immune cells (Figure 2). The cytotoxic activity of MCs is involved in a natural process of allorecognition that occurs when two B. schlosseri colonies with different genotypes touch [58]. This self–nonself recognition is regulated by a single polymorphic histocompatibility gene: the Botryllus Histocompatibility Factor (BHF) [59]. The sharing of at least one BHF allele is a determinant for the fusion or rejection of the colonies [16,17]. Interestingly, MC express high levels of the polymorphic gene FuHC which could be a base for BHF recognition mechanism [17,60] (Figure 2).
Transcriptome analysis and cellular characterization studies in the jawless vertebrates, lampreys, have proposed that two T-like cell populations expressing the variable lymphocyte receptors (VLRs) VLRA and VLRC, respectively, carries cytotoxic activity [61,62]. These cells express several genes including cell surface receptors (i.e., CD45, TCR, VpreB, paired-Ig-like receptors), cytokines (IL17) and transcription factors (i.e., GATA 2/3, c-Rel, BCL11b) resembling those that T lymphocytes use to migrate, proliferate and differentiate in jawed vertebrates [63]. Lymphocyte lineages have also been found in the hagfish, however these have not been fully characterized [61,64]. This shows a whole convergent parallel adaptive immune system of lymphocytes, which is based on LLRs instead of Ig superfamily like the jawed vertebrates (Figure 2).
In jawed vertebrates, the cytotoxic cells NK and CTLs belong to the lymphoid lineage. NK cells do not express antigen-specific receptors but are highly capable to recognize and kill tumor and viral-infected cells [40]. NK cell function has been recognized in representatives of all vertebrate classes [65]. Comparative studies using high-quality genome assemblies have shown that NK cells diversity in mammals has a big influence on species-specific immune responses and outcomes of pathogen infection [66]. NKT-like cells have been identified in fishes [65,67] the frog Xenopus laevis [68] and through the identification of CD1 molecules, it is suggested that this kind of cells are also present in reptiles and birds [65].
CTLs have three important elements to recognize antigens expressed by viral-infected cells or cancer tissue: (1) a diverse repertoire of Ig domain-based receptors, (2) T cell receptor (TCR) and (3) major histocompatibility complex (MHC). Genes of these elements and the presence of CTLs is a common feature found in all jawed vertebrates which include cartilaginous and bony fishes, amphibians, reptiles, birds, and mammals. Moreover, these cells were also found in the extinct placoderms [65,69]. In mammals, also activated macrophages could selectively lyse malignant cells in a contact-dependent and non-phagocytic process, having important roles in the restoration of tissue homeostasis [70]. Finally, ADCC is well characterized in mammals by eosinophilic and neutrophilic granulocytes, NK cells, and CD16+ blood monocytes [71]. Their main effects are associated with graft rejection, autoimmune diseases, tumor surveillance, antiviral and antiparasitic defense [71,72,73,74].
We think that in cases where allogeneic specificity is found in cytotoxicity, this should be considered and further investigated as potential for level II cytotoxicity. For instance, the example mentioned earlier of sipunculid worms showing different cytotoxic effects between levels of allogeneic proximity of the target erythrocytes is suggestive of specificity recognition by the cytotoxic effector cells [52]. Characterization of the cells and those recognition mechanisms would shed light if there were a presence of level II specific cytotoxicity (Figure 2). Moreover, receptor recombination as an example of Trfs in urchins could also suggest specific cytotoxicity, at this stage it was only tested against bacterial opsonization [32,33], but if it is effective against foreign cells or abnormal self, then could be considered as level II specific lysis.
3.2. Receptors
Killer cells possess diverse receptors for triggering their cytotoxic functions, these receptors seem to appear early in evolution although without high specificity. In mollusks, the Fibrinogen-related proteins (FREPs) bind to parasites and has been suggested as potential adaptive defense molecules [75,76,77]. Hemocyte membranes of the arthropod P. bicarinatus have trypsin-labile receptors and nonspecific cytotoxic cell receptors protein 1 (NCCRP-1) which appear to be mediators of allorecognition [78]. Additionally, haemocytes from Prodenia eridania express non-specific receptors for foreign particles [48]. NK cells from mammals have Killer Inhibitory Receptors (KIRs) which inhibit the activation of cytotoxic programs when recognize self-MHC, according to the “missing self” hypothesis NK cells kill in this manner allogeneic foreign cells that do not express self-MHC or that have altered expression of MHC molecules [79,80,81]. In humans, CD94, NKG2 and NKR-P1 are examples of these receptors, while rodents express a group of Ly49 receptors. Blood cells of the tunicate B. schlosseri also express a gene coding for a type II transmembrane protein with high similarity to vertebrates CD94 and NKR-P1, which is upregulated during allorecognition [82]. VLRA, VLRB and VLRC genes have been identified in lampreys and hagfish [61,62]. These receptors are highly diverse due to a somatic diversification strategy based on the insertion of LRR sequences in a single germline gene [83]. The locus VLRA has >500 LRR cassettes, while there are >800 LRR cassettes in the VLRB locus and >200 LRR cassettes in the VLRC locus [84,85]. This evolutionary mechanism to diversify lymphocyte receptors at a single cell level is similar, but not identical, to the multigene recombinatorial strategy of immunoglobulin gene segments and TCR molecules in jawed vertebrates [83,86].
In all jawed vertebrates, CTLs recognize peptides bound to MHC molecules by expressing the T cell receptor complex (TCR). Peptides are derived from viral or bacterial proteins during infection [87], and MHC I nonself recognition also occurs after allotransplantation [88]. In this sense, the expression of TCRs is clonotypic since one T cell expresses a single TCR specificity that is determined by both the antigen peptide and the MHC determinants. TCR diversity, which is generated by mechanisms of genomic rearrangement, and the fact that MHC molecules are highly polymorphic allow organisms to counteract pathogen evasion of T cell responses [89]. While TCR is the major CTL marker, the glycoproteins CD3, CD8 and CD57 are other important markers which bind MHC molecules [89]. The structure and function of immunoglobulin M (IgM), the classic TCRs (αβ chains) and MHC class II are conserved in jawed vertebrates from cartilaginous and bony fishes, to amphibians, reptiles, birds, and mammals. However, other immune receptors as γδ TCR, NKRs and nonclassical MHCs are present but with diverse function, which in most cases is still unexplored [65,90,91,92].
3.3. Effector Molecules
As in phagocytosis, the secretion of chemokines and effector molecules is important for the activation of cytotoxic programs which are an important immune army defense against pathogens and foreign transplanted cells. Between the most conserved cytotoxic components among invertebrate humoral defenses is lysozyme. This is an enzyme that digest components of the cell-wall from Gram-positive bacteria. It has been found in the soluble fraction of hemocytes preparations from mollusk, freshwater crayfish, freshwater snail, earthworms and ascidian. Two types of lysozymes are also found among vertebrate animals that are the Chicken (c)-type lysozyme and the goose (g)-type lysozyme [58]. Other soluble factors with antibacterial/antiviral activity (proteinases, pore forming in cell membranes or ion gradient disruptors) including astacidines, hemagglutinin and anti-LPS factor have been identified in the hemolymph of many invertebrates [93]. The production of reactive oxygen intermediates (O2, H2O2, HOCl) as well as reactive nitrogen intermediates (NO, N2O3, ONOO, NO2, NO3) has been found associated with cytotoxic activity in echinoderms, coelenterates, nematodes, annelids, insects, crustaceans and mollusks [94]. The production of Nitric Oxide (NO) has been detected in two parasitic and two free living flatworms, in these organisms the presence of phenoloxidase has also been detected although its function in their cytotoxic immune responses is still unknown [12]. In mammals, ROS/NOS-mediated cytotoxicity by macrophages, NK cells and monocytes has also been associated with the eradication of pathogens, viral infections, and antitumor response. Additionally, it was also associated with the modulation of inflammation during tissue repair and wound healing processes in vertebrate species including humans and rodents [95,96]. Vertebrate activated macrophages contain primary lysosomes with more than 40 enzymes with proteolytic activity including acid hydrolases, peroxidases and neutral proteases [97]. The most important cytokines produced by human NK cells are TNF-α, IFN-γ and IL-22. These promote the killing of intracellular pathogens and the skewing of T adaptive immune responses [98]. NK-like from mollusks also contain TNF-α, however its effects on cytotoxic killing remains to be studied [99]. Upon MHC recognition, CTLs generate typical lysosomal granules with hydrolytic and pore forming proteins (i.e., acid phosphatase, β-glucoronidase, acid esterase, serine proteases and perforin) in order to lyse their target cells [100].
3.4. Signaling Pathways
After the engagement of cytotoxic cell receptors that recognize target cells, phosphorylation cascades result in the activation of key signaling pathways for the execution of cytotoxic programs including the release of lysosomal granules. In NK cells, this signaling is primarily mediated by Src family kinases, MAPK activation mediated by PI3K and Rac-1 and ERK pathway. In ADCC, a Ras-dependent pathway also guides to ERK activation and thus to granule releasing. CTLs classically use the TCR signaling pathway triggered by ligation of peptide-MHC. This involves the recruitment of Ras, Raf-1 activation, phosphorylation of MEK-1 and ERK phosphorylation and activation [101,102]. Evidence suggests that the cytotoxicity of molluscan and Drosophila hemocytes relies in signal transduction pathways that are similar to vertebrate response including protein kinases A and C, focal adhesion kinases, Src, FAK and MAPK proteins, small G proteins (Ras), and second messengers as cAMP and Ca2+ [103]. In the protochordate B. schlosseri, immune signal transduction pathways include the activation of G-proteins, protein kinases A and C, PI3K and MAPKs, although these have been more related with phagocytic than with cytotoxic programs [104].
4. Perspective on the Effect of the Immune System in the Evolution of Regenerative Capacities
Regeneration is the capacity of tissues and organs to reconstitute their structure and function after injury. Regenerative potential has been lost through the evolution of animals. Basal invertebrates have extraordinary abilities to regenerate complex structures and even an entire organism. Among vertebrates, fishes and amphibians have a well-developed capacity to regenerate tail, jaws, spinal cord, lens, myocardium, and some gastrointestinal regions. Regeneration of complex structures in mammals is limited to antlers, earholes, nipples, digit tips and liver, instead they have remarkable capacities to renew and repair tissues and healing wounds (i.e., skin, bone, skeletal muscle, hematopoietic system) [121,122].
Several questions about the evolution of the regenerative potential have been discussed. Is it a normal development epiphenomenon that has been selectively lost? Has it lacked selective pressure to be preserved in vertebrate taxa as an adaptive trait? What physiological benefits predominate over the ability to regenerate during the evolution of life?
The immune system has pivotal roles in both antimicrobial/antiviral response, and tissue repair and regeneration. At the wound site, phagocytic and cytotoxic immune cells (i.e., neutrophils, macrophages, monocytes, lymphocytes) are recruited to clear debris and to secrete signaling molecules activating inflammation, proliferation, differentiation and extracellular matrix remodeling programs [123]. In this regard, the group of Anthony Mescher proposed in 2003 an immunologic hypothesis stating that the appearance of novel cell types (and their interactions) during the evolution of the adaptive immune system allowed vertebrates to have better response against pathogens in injured tissue but, these advantages implied the possible loosing of regenerative capacities. This has been reviewed in detailed in [124,125].
Mescher’s hypothesis relies on experimental studies in different models of regeneration: full-thickness injuries in fetal skin [126], transgenic mouse models deficient for specific immune cells [127], hind-limb amputation in urodeles and anurans [128]. The given evidence suggests that the profile of the infiltrated immune cells and their interactions within the injured tissue directly influences inflammation and scarring [125,129,130].
It has been proposed that inflammation and scarring prevent tissue/organ regeneration. This is supported by evidence showing that vertebrates with less well-developed adaptive immune system (such as teleost fish, larval anurans, urodele amphibians) show minimal inflammatory responses to injury and excellent regenerative potential [129]. Furthermore, the success of the regenerative process is also dependent on the timely resolution of inflammation, which requires changes in the macrophage populations and in the gene expression of other immune cells. The regeneration of larval Xenopus hindlimbs is reduced when this process is anticipated or delayed [131]. Moreover, scar-free repair of fetal skin wounds has been reported in diverse mammals. In humans, this occurs until the early second trimester of gestation, after which there is a switch to the scarring repair of the adult skin healing. A remarkable difference between both repairing processes relies in the development of an immune system that causes inflammation in wounded tissues, while in fetus there is a minimal acute inflammatory response which is quickly resolved [125]. In this sense, the genetic ablation of macrophages and neutrophils in the paw of PU.1 null mouse resulted in a scarless healing process [132,133]; while wounded fetal skin of IL-10 deficient mice (anti-inflammatory cytokine) form scar and increase the inflammatory infiltrate [134]. This regenerative potential in fetuses is, however, dependent on the type of injury since severe skin wounds produce high migration of macrophages, local cellular necrosis and scar formation [135,136]. Scarring is strikingly linked to inflammation; it is caused by immune cell interactions that produce several cytokines that activate mesenchymal cells and fibroblasts, promoting fibroplasia, contraction, and collagen production. According to the data, the strength of the inflammatory response correlates strongly with the amount of the scar that the tissue or organ will form, for more information review [137].
However, research in different regenerative models has shown that specific inflammatory components (cells, cytokines and chemokines) are central promoters of regeneration. This has been deeply reviewed in [138]. In the mice ear-hole model, during the early acute inflammatory response, the overexpression of inflammatory genes related to progenitor mast cells (i.e., MCP-7, CD34, CD14/CDw17) promotes the ear-hole closure [139,140]. A strong infiltration of leukocytes occurs during the skin regeneration in adult axolotls [141], and macrophage deletion reduces zebrafish embryonic and adult tail fin regeneration, which does not occur in their caudal fin [142,143,144]. The use of glucocorticoids to reduce inflammation at the moment of amputation can fully prevent regeneration of Xenopus limbs [145] evidencing that regeneration does not occur without inflammation.
All this evidence shows that inflammation is essential for triggering the process of regeneration; however, the timely resolution of the inflammatory phase will also determine the success of the generation of new and functional organs or tissues. More research into immune activities during various biological responses is needed to understand how they have evolved simultaneously in such a way in which the specialization of some responses (against infections) may have negative effects on capabilities that appear to be advantageous for life, such as regeneration.
5. Concluding Remarks
Researchers are still dealing with the understanding about the origin and phylogenetic evolution of both phagocytosis and cytotoxicity functions in the immune response. More integral analysis with evolutive perspectives are needed to provide a broad overview about the commonalities and divergences between immune systems of different organisms. For this, we require to establish simple animal models that allow the study of complex responses in the defense of an organism against a pathogen, in the recognition of foreign tissues or in the regeneration or repair of a damaged organ/tissue. With the available information, it is easier to speculate that complex immune system responses arose in ancestral organisms more than 550 million years ago, and that selective pressures influenced the diversification of effector cells and molecules but conserving essential activities as the basis to maintain homeostasis and life.
Beyond the basic research of the evolution and diversification of the immune system, there is a potential for discoveries which could be relevant to biomedical research and applications. Some very interesting and important discoveries came from comparative immunology research, such as: (I) the first discovery of immune “professional” cells and phagocytosis was done in starfish larvae [7]; (II) the antiviral mechanism of siRNA was elucidated in C. elegance [146], which led to the discovery of other silencing mechanisms conducted by microRNAs [147] and the widespread use of synthetic siRNAs for RNA interference in mammalian cells [148]; (III) Toll and TLR immune receptors were first discovered in D. melanogaster [149,150]; (IV) a new and convergent adaptive immunity, based on leucin reach repeats (LLR) was discovered in jawless vertebrates [83]; (V) stem cell competition in transplantation was first revealed in Tunicates [151], and later shown in mammalian development, aging, cancer and atherosclerosis (reviewed in [152]); (VI) CRISPR-Cas was discovered as an anti-phage mechanism in bacteria [153,154]. It is important to state that four of those discoveries (I–III, VI) were awarded the Nobel Prize due to their significance in biomedical research and medical applications [155,156,157,158]. Those examples show the potential of discoveries which could be revealed trough comparative immunology approach.
Author Contributions
Conceptualization, B.R. and E.A.M.-T.; writing—original draft preparation, E.A.M.-T. and B.R.; writing—review and editing, E.A.M.-T., B.R., E.S., A.O. and O.G.-Y.; supervision, B.R.; project administration, O.G.-Y.; funding acquisition, B.R. All authors have read and agreed to the published version of the manuscript.
Funding
The work of B.R. was supported by Israel Science Foundation (ISF) grant number 1416/19, and HFSP Research Grant, RGY0085/2019.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Rolff, J. Why did the acquired immune system of vertebrates evolve? Dev. Comp. Immunol. 2007, 31, 476–482. [Google Scholar] [CrossRef] [PubMed]
- Buchmann, K. Evolution of Innate Immunity: Clues from Invertebrates via Fish to Mammals. Front. Immunol. 2014, 5, 459. [Google Scholar] [CrossRef] [PubMed]
- Kimbrell, D.A.; Beutler, B. The evolution and genetics of innate immunity. Nat. Rev. Genet. 2001, 2, 256–267. [Google Scholar] [CrossRef]
- Netea, M.G.; Schlitzer, A.; Placek, K.; Joosten, L.A.B.; Schultze, J.L. Innate and Adaptive Immune Memory: An Evolutionary Continuum in the Host’s Response to Pathogens. Cell Host Microbe 2019, 25, 13–26. [Google Scholar] [CrossRef]
- Gordon, S. Phagocytosis: An Immunobiologic Process. Immunity 2016, 44, 463–475. [Google Scholar] [CrossRef]
- Gordon, S. Elie Metchnikoff, the Man and the Myth. J. Innate Immun. 2016, 8, 223–227. [Google Scholar] [CrossRef]
- Metschnikoff, E. Memoirs: The Ancestral History of the Inflammatory Process. J. Cell Sci. 1884, s2–s24, 112–117. [Google Scholar] [CrossRef]
- Schulenburg, H.; Kurz, C.L.; Ewbank, J.J. Evolution of the innate immune system: The worm perspective. Immunol. Rev. 2004, 198, 36–58. [Google Scholar] [CrossRef]
- Mukherjee, S.; Ray, M.; Ray, S. Phagocytic efficiency and cytotoxic responses of Indian freshwater sponge (Eunapius carteri) cells isolated by density gradient centrifugation and flow cytometry: A morphofunctional analysis. Zoology 2015, 118, 8–18. [Google Scholar] [CrossRef] [PubMed]
- Snyder, G.A.; Eliachar, S.; Connelly, M.T.; Talice, S.; Hadad, U.; Gershoni-Yahalom, O.; Browne, W.E.; Palmer, C.V.; Rosental, B.; Traylor-Knowles, N. Functional characterization of Hexacorallia phagocytic cells. Front. Immunol. 2021, 12, 2402. [Google Scholar]
- Morita, M. Phagocytic response of planarian reticular cells to heat-killed bacteria. Hydrobiologia 1991, 227, 193–199. [Google Scholar] [CrossRef]
- Maciel, E.I.; Oviedo, N.J. Platyhelminthes: Molecular Dissection of the Planarian Innate Immune System. In Advances in Comparative Immunology; Cooper, E.L., Ed.; Springer: Cham, Switzerland, 2018. [Google Scholar]
- Homa, J. Earthworm coelomocyte extracellular traps: Structural and functional similarities with neutrophil NETs. Cell Tissue Res. 2018, 371, 407–414. [Google Scholar] [CrossRef]
- Csordás, G.; Gábor, E.; Honti, V. There and back again: The mechanisms of differentiation and transdifferentiation in Drosophila blood cells. Dev. Biol. 2021, 469, 135–143. [Google Scholar] [CrossRef]
- Cattenoz, P.B.; Sakr, R.; Pavlidaki, A.; Delaporte, C.; Riba, A.; Molina, N.; Hariharan, N.; Mukherjee, T.; Giangrande, A. Temporal specificity and heterogeneity of Drosophila immune cells. EMBO J. 2020, 39, e104486. [Google Scholar] [CrossRef]
- Rosental, B.; Kowarsky, M.; Seita, J.; Corey, D.M.; Ishizuka, K.J.; Palmeri, K.J.; Chen, S.-Y.; Sinha, R.; Okamoto, J.; Mantalas, G.; et al. Complex mammalian-like haematopoietic system found in a colonial chordate. Nature 2018, 564, 425–429. [Google Scholar] [CrossRef]
- Rosental, B.; Raveh, T.; Voskoboynik, A.; Weissman, I.L. Evolutionary perspective on the hematopoietic system through a colonial chordate: Allogeneic immunity and hematopoiesis. Curr. Opin. Immunol. 2020, 62, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Grayfer, L.; Kerimoglu, B.; Yaparla, A.; Hodgkinson, J.W.; Xie, J.; Belosevic, M. Mechanisms of Fish Macrophage Antimicrobial Immunity. Front. Immunol. 2018, 9, 1105. [Google Scholar] [CrossRef]
- Vandendriessche, S.; Cambier, S.; Proost, P.; Marques, P.E. Complement Receptors and Their Role in Leukocyte Recruitment and Phagocytosis. Front. Cell Dev. Biol. 2021, 9, 624025. [Google Scholar] [CrossRef] [PubMed]
- Hirayama, D.; Iida, T.; Nakase, H. The Phagocytic Function of Macrophage-Enforcing Innate Immunity and Tissue Homeostasis. Int. J. Mol. Sci. 2017, 1, 92. [Google Scholar] [CrossRef]
- Yuen, B.; Bayes, J.M.; Degnan, S.M. The characterization of sponge NLRs provides insight into the origin and evolution of this innate immune gene family in animals. Mol. Biol. Evol. 2014, 31, 106–120. [Google Scholar] [CrossRef] [PubMed]
- Wiens, M.; Korzhev, M.; Krasko, A.; Thakur, N.L.; Perović-Ottstadt, S.; Breter, H.J.; Ushijima, H.; Diehl-Seifert, B.; Müller, I.M.; Müller, W.E. Innate immune defense of the sponge Suberites domuncula against bacteria involves a MyD88-dependent signaling pathway. Induction of a perforin-like molecule. J. Biol. Chem. 2005, 280, 27949–27959. [Google Scholar] [CrossRef] [PubMed]
- Narbonne-Reveau, K.; Charroux, B.; Royet, J. Lack of an antibacterial response defect in Drosophila Toll-9 mutant. PLoS ONE 2011, 6, e17470. [Google Scholar] [CrossRef] [PubMed]
- Stuart, L.M.; Ezekowitz, R.A. Phagocytosis and comparative innate immunity: Learning on the fly. Nat. Rev. Immunol. 2008, 8, 131–141. [Google Scholar] [CrossRef]
- Peronato, A.; Franchi, N.; Loriano, B. BsTLR1: A new member of the TLR family of recognition proteins from the colonial ascidian Botryllus schlosseri. Fish Shellfish Immunol. 2020, 106, 967–974. [Google Scholar] [CrossRef]
- Buckley, K.M.; Rast, J.P. Dynamic evolution of toll-like receptor multigene families in echinoderms. Front. Immunol. 2012, 3, 136. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Zhang, H.; Zhao, C.; Zhang, H. Evolutionary History of the Toll-Like Receptor Gene Family across Vertebrates. Genome Biol. Evol. 2020, 12, 3615–3634. [Google Scholar] [CrossRef]
- Peiris, T.H.; Hoyer, K.K.; Oviedo, N.J. Innate immune system and tissue regeneration in planarians: An area ripe for exploration. Semin. Immunol. 2014, 26, 295–302. [Google Scholar] [CrossRef]
- Guillou, A.; Troha, K.; Wang, H.; Franc, N.C.; Buchon, N. The Drosophila CD36 Homologue croquemort is required to Maintain Immune and Gut Homeostasis during Development and Aging. PLoS Pathog. 2016, 12, e1005961. [Google Scholar] [CrossRef]
- Means, T.K.; Mylonakis, E.; Tampakakis, E.; Colvin, R.A.; Seung, E.; Puckett, L.; Tai, M.F.; Stewart, C.R.; Pukkila-Worley, R.; Hickman, S.E.; et al. Evolutionarily conserved recognition and innate immunity to fungal pathogens by the scavenger receptors SCARF1 and CD36. J. Exp. Med. 2009, 206, 637–653. [Google Scholar] [CrossRef]
- Febbraio, M.; Hajjar, D.P.; Silverstein, R.L. CD36: A class B scavenger receptor involved in angiogenesis, atherosclerosis, inflammation, and lipid metabolism. J. Clin. Investig. 2001, 108, 785–791. [Google Scholar] [CrossRef] [PubMed]
- Yakovenko, I.; Donnyo, A.; Ioscovich, O.; Rosental, B.; Oren, M. The Diverse Transformer (Trf) Protein Family in the Sea Urchin Paracentrotus lividus Acts through a Collaboration between Cellular and Humoral Immune Effector Arms. Int. J. Mol. Sci. 2021, 22, 6639. [Google Scholar] [CrossRef]
- Chou, H.-Y.; Lun, C.M.; Smith, L.C. SpTransformer proteins from the purple sea urchin opsonize bacteria, augment phagocytosis, and retard bacterial growth. PLoS ONE 2018, 13, e0196890. [Google Scholar] [CrossRef]
- Oren, M.; Barela Hudgell, M.A.; D’Allura, B.; Agronin, J.; Gross, A.; Podini, D.; Smith, L.C. Short tandem repeats, segmental duplications, gene deletion, and genomic instability in a rapidly diversified immune gene family. BMC Genom. 2016, 17, 900. [Google Scholar] [CrossRef]
- Smith, L.C.; Lun, C.M. The SpTransformer Gene Family (Formerly Sp185/333) in the Purple Sea Urchin and the Functional Diversity of the Anti-Pathogen rSpTransformer-E1 Protein. Front. Immunol. 2017, 8, 725. [Google Scholar] [CrossRef] [PubMed]
- Majeske, A.J.; Oren, M.; Sacchi, S.; Smith, L.C. Single sea urchin phagocytes express messages of a single sequence from the diverse Sp185/333 gene family in response to bacterial challenge. J. Immunol. 2014, 193, 5678–5688. [Google Scholar] [CrossRef]
- Franchi, N.; Schiavon, F.; Carletto, M.; Gasparini, F.; Bertoloni, G.; Tosatto, S.C.; Ballarin, L. Immune roles of a rhamnose-binding lectin in the colonial ascidian Botryllus schlosseri. Immunobiology 2011, 216, 725–736. [Google Scholar] [CrossRef]
- Vanha-Aho, L.M.; Valanne, S.; Rämet, M. Cytokines in Drosophila immunity. Immunol. Lett. 2016, 170, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Langenbacher, A.D.; De Tomaso, A.W. Temporally and spatially dynamic germ cell niches in Botryllus schlosseri revealed by expression of a TGF-beta family ligand and vasa. EvoDevo 2016, 7, 9. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.; Secombes, C.J. The Function of Fish Cytokines. Biology 2016, 5, 23. [Google Scholar] [CrossRef]
- Savan, R.; Ravichandran, S.; Collins, J.R.; Sakai, M.; Young, H.A. Structural conservation of interferon gamma among vertebrates. Cytokine Growth Factor Rev. 2009, 20, 115–124. [Google Scholar] [CrossRef]
- Williams, L.M.; Inge, M.M.; Mansfield, K.M.; Rasmussen, A.; Afghani, J.; Agrba, M.; Albert, C.; Andersson, C.; Babaei, M.; Babaei, M.; et al. Transcription factor NF-κB in a basal metazoan, the sponge, has conserved and unique sequences, activities, and regulation. Dev. Comp. Immunol. 2020, 104, 103559. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, M.A.; Fontenele, M.; Lim, B.; Bisch, P.M.; Shvartsman, S.Y.; Araujo, H.M. A novel function for the IκB inhibitor Cactus in promoting Dorsal nuclear localization and activity in the Drosophila embryo. Development 2017, 144, 2907–2913. [Google Scholar]
- Franchi, N.; Schiavon, F.; Betti, M.; Canesi, L.; Ballarin, L. Insight on signal transduction pathways involved in phagocytosis in the colonial ascidian Botryllus schlosseri. J. Invertebr. Pathol. 2013, 112, 260–266. [Google Scholar] [CrossRef] [PubMed]
- Dorrington, M.G.; Fraser, I.D.C. NF-κB Signaling in Macrophages: Dynamics, Crosstalk, and Signal Integration. Front. Immunol. 2019, 10, 705. [Google Scholar] [CrossRef] [PubMed]
- Valanne, S.; Wang, J.H.; Rämet, M. The Drosophila Toll signaling pathway. J. Immunol. 2011, 186, 649–656. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, L.A.J.; Golenbock, D.; Bowie, A.G. The history of Toll-like receptors—Redefining innate immunity. Nat. Rev. Immunol. 2013, 13, 453–460. [Google Scholar] [CrossRef]
- Cooper, E.L. Phylogeny of cytotoxicity. Endeavour 1980, 4, 160–165. [Google Scholar] [CrossRef]
- Colantonio, A.D.; Bimber, B.N.; Neidermyer, W.J., Jr.; Reeves, R.K.; Alter, G.; Altfeld, M.; Johnson, R.P.; Carrington, M.; O’Connor, D.H.; Evans, D.T. KIR polymorphisms modulate peptide-dependent binding to an MHC class I ligand with a Bw6 motif. PLoS Pathog. 2011, 7, e1001316. [Google Scholar] [CrossRef]
- Rosental, B.; Appel, M.Y.; Yossef, R.; Hadad, U.; Brusilovsky, M.; Porgador, A. The effect of chemotherapy/radiotherapy on cancerous pattern recognition by NK cells. Curr. Med. Chem. 2012, 19, 1780–1791. [Google Scholar] [CrossRef]
- Franceschi, C.; Cossarizza, A.; Ortolani, C.; Monti, D.; Ottaviani, E. Natural cytotoxicity in a freshwater pulmonate mollusc: An unorthodox comparative approach. Adv. Neuroimmunol. 1991, 1, 99–113. [Google Scholar] [CrossRef]
- Boiledieu, D.; Valembois, P. Natural cytotoxic activity of sipunculid leukocytes on allogenic and xenogenic erythrocytes. Dev. Comp. Immunol. 1977, 1, 207–216. [Google Scholar] [CrossRef]
- Radomski, M.W.; Martin, J.F.; Moncada, S. Synthesis of Nitric Oxide by the Haemocytes of the American Horseshoe Crab (Limulus polyphemus). Philos. Trans. Biol. Sci. 1991, 334, 129–133. [Google Scholar]
- Söderhäll, K.; Wingren, A.; Johansson, M.W.; Bertheussen, K. The cytotoxic reaction of hemocytes from the freshwater crayfish, Astacus astacus. Cell. Immunol. 1985, 94, 326–332. [Google Scholar] [CrossRef]
- Cárdenas, W.; Dankert, J.R. Cresolase, catecholase and laccase activities in haemocytes of the red swamp crayfish. Fish Shellfish Immunol. 2000, 10, 33–46. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.; Zhang, H.; Beck, G. Phylogeny of natural cytotoxicity: Cytotoxic activity of coelomocytes of the purple sea urchin, Arbacia punctulata. J. Exp. Zool. 2001, 290, 741–750. [Google Scholar] [CrossRef] [PubMed]
- Gross, P.S.; Al-Sharif, W.Z.; Clow, L.A.; Smith, L.C. Echinoderm immunity and the evolution of the complement system. Dev. Comp. Immunol. 1999, 23, 429–442. [Google Scholar] [CrossRef]
- Corey, D.M.; Rosental, B.; Kowarsky, M.; Sinha, R.; Ishizuka, K.J.; Palmeri, K.J.; Quake, S.R.; Voskoboynik, A.; Weissman, I.L. Developmental cell death programs license cytotoxic cells to eliminate histocompatible partners. Proc. Natl. Acad. Sci. USA 2016, 113, 6520–6525. [Google Scholar] [CrossRef]
- Voskoboynik, A.; Newman, A.M.; Corey, D.M.; Sahoo, D.; Pushkarev, D.; Neff, N.F.; Passarelli, B.; Koh, W.; Ishizuka, K.J.; Palmeri, K.J.; et al. Identification of a colonial chordate histocompatibility gene. Science 2013, 341, 384–387. [Google Scholar] [CrossRef]
- De Tomaso, A.W.; Nyholm, S.V.; Palmeri, K.J.; Ishizuka, K.J.; Ludington, W.B.; Mitchel, K.; Weissman, I.L. Isolation and characterization of a protochordate histocompatibility locus. Nature 2005, 438, 454–459. [Google Scholar] [CrossRef]
- Das, S.; Li, J.; Hirano, M.; Sutoh, Y.; Herrin, B.R.; Cooper, M.D. Evolution of two prototypic T cell lineages. Cell. Immunol. 2015, 296, 87–94. [Google Scholar] [CrossRef]
- Mayer, W.E.; Uinuk-ool, T.; Tichy, H.; Gartland, L.A.; Klein, J.; Cooper, M.D. Isolation and characterization of lymphocyte-like cells from a lamprey. Proc. Natl. Acad. Sci. USA 2002, 99, 14350–14355. [Google Scholar] [CrossRef]
- Uinuk-ool, T.; Mayer, W.E.; Sato, A.; Dongak, R.; Cooper, M.D.; Klein, J. Lamprey lymphocyte-like cells express homologs of genes involved in immunologically relevant activities of mammalian lymphocytes. Proc. Natl. Acad. Sci. USA 2002, 99, 14356–14361. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Das, S.; Herrin, B.R.; Hirano, M.; Cooper, M.D. Definition of a third VLR gene in hagfish. Proc. Natl. Acad. Sci. USA 2013, 110, 15013–15018. [Google Scholar] [CrossRef]
- Flajnik, M.F. A cold-blooded view of adaptive immunity. Nat. Rev. Immunol. 2018, 18, 438–453. [Google Scholar] [CrossRef]
- Schwartz, J.C.; Gibson, M.S.; Heimeier, D.; Koren, S.; Phillippy, A.M.; Bickhart, D.M.; Smith, T.P.; Medrano, J.F.; Hammond, J.A. The evolution of the natural killer complex; a comparison between mammals using new high-quality genome assemblies and targeted annotation. Immunogenetics 2017, 69, 255–269. [Google Scholar] [CrossRef]
- Shen, L.; Stuge, T.B.; Zhou, H.; Khayat, M.; Barker, K.S.; Quiniou, S.M.; Wilson, M.; Bengtén, E.; Chinchar, V.G.; Clem, L.W.; et al. Channel catfish cytotoxic cells: A mini-review. Dev. Comp. Immunol. 2002, 26, 141–149. [Google Scholar] [CrossRef]
- Morales, H.D.; Robert, J. Characterization of primary and memory CD8 T-cell responses against ranavirus (FV3) in Xenopus laevis. J. Virol. 2007, 81, 2240–2248. [Google Scholar] [CrossRef] [PubMed]
- Hirano, M.; Das, S.; Guo, P.; Cooper, M.D. The evolution of adaptive immunity in vertebrates. Adv. Immunol. 2011, 109, 125–157. [Google Scholar] [PubMed]
- Butenko, S.; Satyanarayanan, S.K.; Assi, S.; Schif-Zuck, S.; Barkan, D.; Sher, N.; Ariel, A. Transcriptomic Analysis of Monocyte-Derived Non-Phagocytic Macrophages Favors a Role in Limiting Tissue Repair and Fibrosis. Front. Immunol. 2020, 11, 405. [Google Scholar] [CrossRef] [PubMed]
- Geffner, J. Antibody-Dependent Cellular Cytotoxicity. In Encyclopedia of Immunotoxicology; Vohr, H.-W., Ed.; Springer: Berlin/Heidelberg, Germany, 2005; pp. 1–5. [Google Scholar]
- Hanton, G.; Pastoret, P.P. The reaction of antibody-dependent cell-mediated cytotoxicity (ADCC). Ann. Vet. Res. 1984, 15, 443–454. [Google Scholar]
- Torben, W.; Ahmad, G.; Zhang, W.; Nash, S.; Le, L.; Karmakar, S.; Siddiqui, A.A. Role of antibody dependent cell mediated cytotoxicity (ADCC) in Sm-p80-mediated protection against Schistosoma mansoni. Vaccine 2012, 30, 6753–6758. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sun, Y.; Asmal, M.; Lane, S.; Permar, S.R.; Schmidt, S.D.; Mascola, J.R.; Letvin, N.L. Antibody-Dependent Cell-Mediated Cytotoxicity in Simian Immunodeficiency Virus-Infected Rhesus Monkeys. J. Virol. 2011, 85, 6906–6912. [Google Scholar] [CrossRef]
- Flajnik, M.F.; Du Pasquier, L. Evolution of innate and adaptive immunity: Can we draw a line? Trends Immunol. 2004, 25, 640–644. [Google Scholar] [CrossRef]
- Schultz, J.H.; Bu, L.; Adema, C.M. Comparative immunological study of the snail Physella acuta (Hygrophila, Pulmonata) reveals shared and unique aspects of gastropod immunobiology. Mol. Immunol. 2018, 101, 108–119. [Google Scholar] [CrossRef]
- Loker, E.S.; Adema, C.M.; Zhang, S.M.; Kepler, T.B. Invertebrate immune systems--not homogeneous, not simple, not well understood. Immunol. Rev. 2004, 198, 10–24. [Google Scholar] [CrossRef] [PubMed]
- Cárdenas, W.; Dankert, J.R.; Jenkins, J.A. Flow cytometric analysis of crayfish haemocytes activated by lipopolysaccharides. Fish Shellfish Immunol. 2004, 17, 223–233. [Google Scholar] [CrossRef]
- Ljunggren, H.-G.; Kärre, K. In search of the ‘missing self’: MHC molecules and NK cell recognition. Immunol. Today 1990, 11, 237–244. [Google Scholar] [CrossRef]
- Raulet, D.H. Missing self recognition and self tolerance of natural killer (NK) cells. Semin. Immunol. 2006, 18, 145–150. [Google Scholar] [CrossRef] [PubMed]
- Borrego, F. The First Molecular Basis of the “Missing Self” Hypothesis. J. Immunol. 2006, 177, 5759–5760. [Google Scholar] [CrossRef]
- Khalturin, K.; Becker, M.; Rinkevich, B.; Bosch, T.C. Urochordates and the origin of natural killer cells: Identification of a CD94/NKR-P1-related receptor in blood cells of Botryllus. Proc. Natl. Acad. Sci. USA 2003, 100, 622–627. [Google Scholar] [CrossRef]
- Pancer, Z.; Amemiya, C.T.; Ehrhardt, G.R.A.; Ceitlin, J.; Larry Gartland, G.; Cooper, M.D. Somatic diversification of variable lymphocyte receptors in the agnathan sea lamprey. Nature 2004, 430, 174–180. [Google Scholar] [CrossRef]
- Kishishita, N.; Nagawa, F. Evolution of adaptive immunity: Implications of a third lymphocyte lineage in lampreys. BioEssays News Rev. Mol. Cell. Dev. Biol. 2014, 36, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Hirano, M.; Aghaallaei, N.; Bajoghli, B.; Boehm, T.; Cooper, M.D. Organization of lamprey variable lymphocyte receptor C locus and repertoire development. Proc. Natl. Acad. Sci. USA 2013, 110, 6043–6048. [Google Scholar] [CrossRef]
- Kasahara, M. Variable Lymphocyte Receptors: A Current Overview. Results Probl. Cell Differ. 2015, 57, 175–192. [Google Scholar]
- Croft, N.P.; Smith, S.A.; Pickering, J.; Sidney, J.; Peters, B.; Faridi, P.; Witney, M.J.; Sebastian, P.; Flesch, I.E.A.; Heading, S.L.; et al. Most viral peptides displayed by class I MHC on infected cells are immunogenic. Proc. Natl. Acad. Sci. USA 2019, 116, 3112–3117. [Google Scholar] [CrossRef]
- Marino, J.; Paster, J.; Benichou, G. Allorecognition by T Lymphocytes and Allograft Rejection. Front. Immunol. 2016, 7, 582. [Google Scholar] [CrossRef] [PubMed]
- Bosselut, R. T cell antigen recognition: Evolution-driven affinities. Proc. Natl. Acad. Sci. USA 2019, 116, 21969–21971. [Google Scholar] [CrossRef] [PubMed]
- Kasahara, M.; Flajnik, M.F. Origin and evolution of the specialized forms of proteasomes involved in antigen presentation. Immunogenetics 2019, 71, 251–261. [Google Scholar] [CrossRef]
- Castro, R.; Bernard, D.; Lefranc, M.P.; Six, A.; Benmansour, A.; Boudinot, P. T cell diversity and TcR repertoires in teleost fish. Fish Shellfish Immunol. 2011, 31, 644–654. [Google Scholar] [CrossRef]
- Foulkrod, A.M.; Appasamy, P.M. Expression of TCR genes in adult and larval Xenopus laevis. Dev. Comp. Immunol. 2019, 96, 78–82. [Google Scholar] [CrossRef]
- Jiravanichpaisal, P.; Lee, S.Y.; Kim, Y.A.; Andrén, T.; Söderhäll, I. Antibacterial peptides in hemocytes and hematopoietic tissue from freshwater crayfish Pacifastacus leniusculus: Characterization and expression pattern. Dev. Comp. Immunol. 2007, 31, 441–455. [Google Scholar] [CrossRef] [PubMed]
- Nappi, A.J.; Ottaviani, E. Cytotoxicity and cytotoxic molecules in invertebrates. BioEssays News Rev. Mol. Cell. Dev. Biol. 2000, 22, 469–480. [Google Scholar] [CrossRef]
- Wink, D.A.; Hines, H.B.; Cheng, R.Y.; Switzer, C.H.; Flores-Santana, W.; Vitek, M.P.; Ridnour, L.A.; Colton, C.A. Nitric oxide and redox mechanisms in the immune response. J. Leukoc. Biol. 2011, 89, 873–891. [Google Scholar] [CrossRef] [PubMed]
- Lisi, F.; Zelikin, A.N.; Chandrawati, R. Nitric Oxide to Fight Viral Infections. Adv. Sci. 2021, 8, 2003895. [Google Scholar] [CrossRef] [PubMed]
- Weber, K.; Schilling, J.D. Distinct lysosome phenotypes influence inflammatory function in peritoneal and bone marrow-derived macrophages. Int. J. Inflamm. 2014, 2014, 154936. [Google Scholar] [CrossRef]
- Fauriat, C.; Long, E.O.; Ljunggren, H.G.; Bryceson, Y.T. Regulation of human NK-cell cytokine and chemokine production by target cell recognition. Blood 2010, 115, 2167–2176. [Google Scholar] [CrossRef]
- Schultz, J.H.; Adema, C.M. Comparative immunogenomics of molluscs. Dev. Comp. Immunol. 2017, 75, 3–15. [Google Scholar] [CrossRef]
- Peters, P.J.; Borst, J.; Oorschot, V.; Fukuda, M.; Krähenbühl, O.; Tschopp, J.; Slot, J.W.; Geuze, H.J. Cytotoxic T lymphocyte granules are secretory lysosomes, containing both perforin and granzymes. J. Exp. Med. 1991, 173, 1099–1109. [Google Scholar] [CrossRef]
- Perussia, B. Signaling for cytotoxicity. Nat. Immunol. 2000, 1, 372–374. [Google Scholar] [CrossRef]
- Kabanova, A.; Zurli, V.; Baldari, C.T. Signals Controlling Lytic Granule Polarization at the Cytotoxic Immune Synapse. Front. Immunol. 2018, 9, 307. [Google Scholar] [CrossRef]
- Humphries, J.E.; Yoshino, T.P. Cellular receptors and signal transduction in molluscan hemocytes: Connections with the innate immune system of vertebrates. Integr. Comp. Biol. 2003, 43, 305–312. [Google Scholar] [CrossRef] [PubMed]
- Franchi, N.; Ballarin, L. Immunity in Protochordates: The Tunicate Perspective. Front. Immunol. 2017, 8, 674. [Google Scholar] [CrossRef] [PubMed]
- Wolff, L.; Humeniuk, R. Concise review: Erythroid versus myeloid lineage commitment: Regulating the master regulators. Stem cells 2013, 31, 1237–1244. [Google Scholar] [CrossRef]
- Potts, K.S.; Bowman, T.V. Modeling Myeloid Malignancies Using Zebrafish. Front. Oncol. 2017, 7, 297. [Google Scholar] [CrossRef] [PubMed]
- Wei, S.; Zhou, J.M.; Chen, X.; Shah, R.N.; Liu, J.; Orcutt, T.M.; Traver, D.; Djeu, J.Y.; Litman, G.W.; Yoder, J.A. The zebrafish activating immune receptor Nitr9 signals via Dap12. Immunogenetics 2007, 59, 813–821. [Google Scholar] [CrossRef] [PubMed]
- Moss, L.D.; Monette, M.M.; Jaso-Friedmann, L.; Leary, J.H., 3rd; Dougan, S.T.; Krunkosky, T.; Evans, D.L. Identification of phagocytic cells, NK-like cytotoxic cell activity and the production of cellular exudates in the coelomic cavity of adult zebrafish. Dev. Comp. Immunol. 2009, 33, 1077–1087. [Google Scholar] [CrossRef] [PubMed]
- Tang, Q.; Iyer, S.; Lobbardi, R.; Moore, J.C.; Chen, H.; Lareau, C.; Hebert, C.; Shaw, M.L.; Neftel, C.; Suva, M.L.; et al. Dissecting hematopoietic and renal cell heterogeneity in adult zebrafish at single-cell resolution using RNA sequencing. J. Exp. Med. 2017, 214, 2875–2887. [Google Scholar] [CrossRef] [PubMed]
- Han, Q.; Das, S.; Hirano, M.; Holland, S.J.; McCurley, N.; Guo, P.; Rosenberg, C.S.; Boehm, T.; Cooper, M.D. Characterization of Lamprey IL-17 Family Members and Their Receptors. J. Immunol. 2015, 195, 5440–5451. [Google Scholar] [CrossRef] [PubMed]
- Hirano, M.; Guo, P.; McCurley, N.; Schorpp, M.; Das, S.; Boehm, T.; Cooper, M.D. Evolutionary implications of a third lymphocyte lineage in lampreys. Nature 2013, 501, 435–438. [Google Scholar] [CrossRef]
- Nyholm, S.V.; Passegue, E.; Ludington, W.B.; Voskoboynik, A.; Mitchel, K.; Weissman, I.L.; De Tomaso, A.W. Fester, a candidate allorecognition receptor from a primitive chordate. Immunity 2006, 25, 163–173. [Google Scholar] [CrossRef]
- Arizza, V.; Giaramita, F.T.; Parrinello, D.; Cammarata, M.; Parrinello, N. Cell cooperation in coelomocyte cytotoxic activity of Paracentrotus lividus coelomocytes. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2007, 147, 389–394. [Google Scholar] [CrossRef]
- Muñoz-Chápuli, R.; Carmona, R.; Guadix, J.A.; Macías, D.; Pérez-Pomares, J.M. The origin of the endothelial cells: An evo-devo approach for the invertebrate/vertebrate transition of the circulatory system. Evol. Dev. 2005, 7, 351–358. [Google Scholar] [CrossRef]
- Melcarne, C.; Lemaitre, B.; Kurant, E. Phagocytosis in Drosophila: From molecules and cellular machinery to physiology. Insect Biochem. Mol. Biol. 2019, 109, 1–12. [Google Scholar] [CrossRef]
- Lanot, R.; Zachary, D.; Holder, F.; Meister, M. Postembryonic Hematopoiesis in Drosophila. Dev. Biol. 2001, 230, 243–257. [Google Scholar] [CrossRef] [PubMed]
- Meister, M.; Lagueux, M. Drosophila blood cells. Cell. Microbiol. 2003, 5, 573–580. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, K.; Nomoto, A.M.; Nishijima, M.; Maruyama, T. Morphological and Functional Characterization of Hemocytes in the Giant ClamTridacna crocea. J. Invertebr. Pathol. 1997, 69, 105–111. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, M.S.; Lopes, K.A.R.; Leite, P.M.S.C.M.; Morais, F.V.; de Campos Velho, N.M.R. Physiological evaluation of the behavior and epidermis of freshwater planarians (Girardia tigrina and Girardia sp.) exposed to stressors. Biol. Open 2018, 7, 6. [Google Scholar] [CrossRef] [PubMed]
- Pita, L.; Hoeppner, M.P.; Ribes, M.; Hentschel, U. Differential expression of immune receptors in two marine sponges upon exposure to microbial-associated molecular patterns. Sci. Rep. 2018, 8, 16081. [Google Scholar] [CrossRef] [PubMed]
- Iismaa, S.E.; Kaidonis, X.; Nicks, A.M.; Bogush, N.; Kikuchi, K.; Naqvi, N.; Harvey, R.P.; Husain, A.; Graham, R.M. Comparative regenerative mechanisms across different mammalian tissues. NPJ Regen. Med. 2018, 3, 6. [Google Scholar] [CrossRef]
- Abarca-Buis, R.F.; Mandujano-Tinoco, E.A.; Cabrera-Wrooman, A.; Krötzsch, E. The complexity of TGFβ/activin signaling in regeneration. J. Cell Commun. Signal. 2021, 15, 7–23. [Google Scholar] [CrossRef]
- Abnave, P.; Ghigo, E. Role of the immune system in regeneration and its dynamic interplay with adult stem cells. Semin. Cell Dev. Biol. 2019, 87, 160–168. [Google Scholar] [CrossRef]
- Harty, M.; Neff, A.W.; King, M.W.; Mescher, A.L. Regeneration or scarring: An immunologic perspective. Dev. Dyn. Off. Publ. Am. Assoc. Anat. 2003, 226, 268–279. [Google Scholar] [CrossRef] [PubMed]
- Mescher, A.L.; Neff, A.W. Regenerative capacity and the developing immune system. Adv. Biochem. Eng. Biotechnol. 2005, 93, 39–66. [Google Scholar]
- Adolph, V.R.; DiSanto, S.K.; Bleacher, J.C.; Dillon, P.W.; Krummel, T.M. The potential role of the lymphocyte in fetal wound healing. J. Pediatric Surg. 1993, 28, 1316–1320. [Google Scholar] [CrossRef]
- Jameson, J.; Ugarte, K.; Chen, N.; Yachi, P.; Fuchs, E.; Boismenu, R.; Havran, W.L. A role for skin gammadelta T cells in wound repair. Science 2002, 296, 747–749. [Google Scholar] [CrossRef]
- Kimura, Y.; Madhavan, M.; Call, M.K.; Santiago, W.; Tsonis, P.A.; Lambris, J.D.; Del Rio-Tsonis, K. Expression of Complement 3 and Complement 5 in Newt Limb and Lens Regeneration. J. Immunol. 2003, 170, 2331–2339. [Google Scholar] [CrossRef] [PubMed]
- Godwin, J.W.; Rosenthal, N. Scar-free wound healing and regeneration in amphibians: Immunological influences on regenerative success. Differ. Res. Biol. Divers. 2014, 87, 66–75. [Google Scholar] [CrossRef] [PubMed]
- Abarca-Buis, R.F.; Martínez-Jiménez, A.; Vera-Gómez, E.; Contreras-Figueroa, M.E.; Garciadiego-Cázares, D.; Paus, R.; Robles-Tenorio, A.; Krötzsch, E. Mechanisms of epithelial thickening due to IL-1 signalling blockade and TNF-α administration differ during wound repair and regeneration. Differ. Res. Biol. Divers. 2018, 99, 10–20. [Google Scholar] [CrossRef] [PubMed]
- Mescher, A.L.; Neff, A.W.; King, M.W. Changes in the inflammatory response to injury and its resolution during the loss of regenerative capacity in developing Xenopus limbs. PLoS ONE 2013, 8, e80477. [Google Scholar] [CrossRef] [PubMed]
- Martin, P.; D’Souza, D.; Martin, J.; Grose, R.; Cooper, L.; Maki, R.; McKercher, S.R. Wound Healing in the PU.1 Null Mouse—Tissue Repair is Not Dependent on Inflammatory Cells. Curr. Biol. 2003, 13, 1122–1128. [Google Scholar] [CrossRef]
- Redd, M.J.; Cooper, L.; Wood, W.; Stramer, B.; Martin, P. Wound healing and inflammation: Embryos reveal the way to perfect repair. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2004, 359, 777–784. [Google Scholar] [CrossRef]
- Liechty, K.W.; Kim, H.B.; Adzick, N.S.; Crombleholme, T.M. Fetal wound repair results in scar formation in interleukin-10-deficient mice in a syngeneic murine model of scarless fetal wound repair. J. Pediatr. Surg. 2000, 35, 866–872, discussion 872–873. [Google Scholar] [CrossRef] [PubMed]
- Hopkinson-Woolley, J.; Hughes, D.; Gordon, S.; Martin, P. Macrophage recruitment during limb development and wound healing in the embryonic and foetal mouse. J. Cell Sci. 1994, 107, 1159–1167. [Google Scholar] [CrossRef] [PubMed]
- Yates, C.C.; Hebda, P.; Wells, A. Skin Wound Healing and Scarring: Fetal Wounds and Regenerative Restitution. Birth Defects Res. Part C Embryo Today Rev. 2012, 96, 325–333. [Google Scholar] [CrossRef] [PubMed]
- Wilgus, T.A. Inflammation as an orchestrator of cutaneous scar formation: A review of the literature. Plast. Aesthetic Res. 2020, 7, 54. [Google Scholar] [CrossRef]
- Mescher, A.L.; Neff, A.W.; King, M.W. Inflammation and immunity in organ regeneration. Dev. Comp. Immunol. 2017, 66, 98–110. [Google Scholar] [CrossRef]
- Gourevitch, D.; Kossenkov, A.V.; Zhang, Y.; Clark, L.; Chang, C.; Showe, L.C.; Heber-Katz, E. Inflammation and Its Correlates in Regenerative Wound Healing: An Alternate Perspective. Adv. Wound Care 2014, 3, 592–603. [Google Scholar] [CrossRef] [PubMed]
- Canhamero, T.; Garcia, L.V.; De Franco, M. Acute Inflammation Loci are Involved in Wound Healing in the Mouse Ear Punch Model. Adv. Wound Care 2014, 3, 582–591. [Google Scholar] [CrossRef] [PubMed]
- Seifert, A.W.; Monaghan, J.R.; Voss, S.R.; Maden, M. Skin regeneration in adult axolotls: A blueprint for scar-free healing in vertebrates. PLoS ONE 2012, 7, e32875. [Google Scholar] [CrossRef]
- Mathew, L.K.; Sengupta, S.; Kawakami, A.; Andreasen, E.A.; Löhr, C.V.; Loynes, C.A.; Renshaw, S.A.; Peterson, R.T.; Tanguay, R.L. Unraveling tissue regeneration pathways using chemical genetics. J. Biol. Chem. 2007, 282, 35202–35210. [Google Scholar] [CrossRef]
- Li, L.; Yan, B.; Shi, Y.Q.; Zhang, W.Q.; Wen, Z.L. Live imaging reveals differing roles of macrophages and neutrophils during zebrafish tail fin regeneration. J. Biol. Chem. 2012, 287, 25353–25360. [Google Scholar] [CrossRef]
- Petrie, T.A.; Strand, N.S.; Yang, C.T.; Rabinowitz, J.S.; Moon, R.T. Macrophages modulate adult zebrafish tail fin regeneration. Development 2014, 141, 2581–2591. [Google Scholar] [CrossRef]
- King, M.W.; Neff, A.W.; Mescher, A.L. The Developing Xenopus Limb as a Model for Studies on the Balance between Inflammation and Regeneration. Anat. Rec. 2012, 295, 1552–1561. [Google Scholar] [CrossRef] [PubMed]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Elbashir, S.M.; Harborth, J.; Lendeckel, W.; Yalcin, A.; Weber, K.; Tuschl, T. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 2001, 411, 494–498. [Google Scholar] [CrossRef] [PubMed]
- Janeway, C.A., Jr. Approaching the asymptote? Evolution and revolution in immunology. Cold Spring Harb. Symp. Quant. Biol. 1989, 54, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Lemaitre, B.; Nicolas, E.; Michaut, L.; Reichhart, J.M.; Hoffmann, J.A. The dorsoventral regulatory gene cassette spätzle/Toll/cactus controls the potent antifungal response in Drosophila adults. Cell 1996, 86, 973–983. [Google Scholar] [CrossRef]
- Laird, D.J.; De Tomaso, A.W.; Weissman, I.L. Stem Cells are Units of Natural Selection in a Colonial Ascidian. Cell 2005, 123, 1351–1360. [Google Scholar] [CrossRef] [PubMed]
- Weissman, I.L. Stem cells are units of natural selection for tissue formation, for germline development, and in cancer development. Proc. Natl. Acad. Sci. USA 2015, 112, 8922–8928. [Google Scholar] [CrossRef]
- Meselson, M.; Yuan, R. DNA Restriction Enzyme from E. coli. Nature 1968, 217, 1110–1114. [Google Scholar] [CrossRef] [PubMed]
- Mojica, F.J.; Díez-Villaseñor, C.; García-Martínez, J.; Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 2005, 60, 174–182. [Google Scholar] [CrossRef] [PubMed]
- Lokaj, J.; John, C. Ilya Ilich Metchnikov and Paul Ehrlich: 1908 Nobel Prize winners for their research on immunity. Epidemiol. Mikrobiol. Imunol. Cas. Spol. Epidemiol. Mikrobiol. Ceske Lek. Spol. J. E. Purkyne 2008, 57, 119–124. [Google Scholar]
- Bernards, R. The Nobel Prize in Physiology or Medicine for 2006 for the discovery of RNA interference. Ned. Tijdschr. Geneeskd. 2006, 150, 2849–2853. [Google Scholar] [PubMed]
- Volchenkov, R.; Sprater, F.; Vogelsang, P.; Appel, S. The 2011 Nobel Prize in Physiology or Medicine. Scand. J. Immunol. 2012, 75, 1–4. [Google Scholar] [CrossRef]
- Westermann, L.; Neubauer, B.; Köttgen, M. Nobel Prize 2020 in Chemistry honors CRISPR: A tool for rewriting the code of life. Pflug. Arch. Eur. J. Physiol. 2021, 473, 1–2. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
