Oncogenic FGFR Fusions Produce Centrosome and Cilia Defects by Ectopic Signaling
Abstract
1. Primary Cilium and Its Role in Cancer Development
2. FGFR Regulates Cilia Motility and Signaling during Morphogenesis
3. Aberrant FGFR Signaling Affects Primary Cilia
4. FGFR Gene Fusions in Cancer
4.1. FGFR3-TACC3
4.2. FGFR1-TACC1
4.3. FGFR2-BICC1
4.4. FGFR2-NDC80
4.5. FGFR2-CIT
4.6. FGFR2-OFD1
4.7. FOP-FGFR1
4.8. CEP110-FGFR1
5. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Reiter, J.F.; Leroux, M.R. Genes and molecular pathways underpinning ciliopathies. Nat. Rev. Mol. Cell Biol. 2017, 18, 533–547. [Google Scholar] [CrossRef] [PubMed]
- Kopinke, D.; Norris, A.M.; Mukhopadhyay, S. Developmental and regenerative paradigms of cilia regulated hedgehog signaling. Semin. Cell Dev. Biol. 2021, 110, 89–103. [Google Scholar] [CrossRef] [PubMed]
- Ford, M.J.; Yeyati, P.L.; Mali, G.R.; Keighren, M.A.; Waddell, S.H.; Mjoseng, H.K.; Douglas, A.T.; Hall, E.A.; Sakaue-Sawano, A.; Miyawaki, A.; et al. A Cell/Cilia Cycle Biosensor for Single-Cell Kinetics Reveals Persistence of Cilia after G1/S Transition Is a General Property in Cells and Mice. Dev. Cell 2018, 47, 509–523.e5. [Google Scholar] [CrossRef] [PubMed]
- Paridaen, J.T.M.L.; Wilsch-Bräuninger, M.; Huttner, W.B. Asymmetric Inheritance of Centrosome-Associated Primary Cilium Membrane Directs Ciliogenesis after Cell Division. Cell 2013, 155, 333–344. [Google Scholar] [CrossRef] [PubMed]
- Plotnikova, O.V.; Pugacheva, E.N.; Golemis, E.A. Primary Cilia and the Cell Cycle, 1st ed.; Elsevier: Amsterdam, The Netherlands, 2009; Volume 94, ISBN 9780123750242. [Google Scholar]
- Ke, Y.-N.; Yang, W.-X. Primary cilium: An elaborate structure that blocks cell division? Gene 2014, 547, 175–185. [Google Scholar] [CrossRef] [PubMed]
- Pugacheva, E.N.; Jablonski, S.A.; Hartman, T.R.; Henske, E.P.; Golemis, E.A. HEF1-Dependent Aurora A Activation Induces Disassembly of the Primary Cilium. Cell 2007, 129, 1351–1363. [Google Scholar] [CrossRef] [PubMed]
- Inoko, A.; Matsuyama, M.; Goto, H.; Ohmuro-Matsuyama, Y.; Hayashi, Y.; Enomoto, M.; Ibi, M.; Urano, T.; Yonemura, S.; Kiyono, T.; et al. Trichoplein and Aurora A block aberrant primary cilia assembly in proliferating cells. J. Cell Biol. 2012, 197, 391–405. [Google Scholar] [CrossRef]
- Wang, G.; Chen, Q.; Zhang, X.; Zhang, B.; Zhuo, X.; Liu, J.; Jiang, Q.; Zhang, C. PCM1 recruits Plk1 to the pericentriolar matrix to promote primary cilia disassembly before mitotic entry. J. Cell Sci. 2013, 126, 1355–1365. [Google Scholar] [CrossRef] [PubMed]
- Cappello, P.; Blaser, H.; Gorrini, C.; Lin, D.C.C.; Elia, A.J.; Wakeham, A.; Haider, S.; Boutros, P.C.; Mason, J.M.; Miller, N.A.; et al. Role of Nek2 on centrosome duplication and aneuploidy in breast cancer cells. Oncogene 2014, 33, 2375–2384. [Google Scholar] [CrossRef]
- Dere, R.; Perkins, A.L.; Bawa-Khalfe, T.; Jonasch, D.; Walker, C.L. β-Catenin links von Hippel-Lindau to Aurora kinase A and loss of primary cilia in renal cell carcinoma. J. Am. Soc. Nephrol. 2015, 26, 553–564. [Google Scholar] [CrossRef]
- Egeberg, D.L.; Lethan, M.; Manguso, R.; Schneider, L.; Awan, A.; Jørgensen, T.S.; Byskov, A.G.; Pedersen, L.B.; Christensen, S.T. Primary cilia and aberrant cell signaling in epithelial ovarian cancer. Cilia 2012, 1, 15. [Google Scholar] [CrossRef]
- Sarkisian, M.R.; Li, W.; Di Cunto, F.; D’Mello, S.R.; LoTurco, J.J. Citron-kinase, a protein essential to cytokinesis in neuronal progenitors, is deleted in the flathead mutant rat. J. Neurosci. 2002, 22, 1–5. [Google Scholar] [CrossRef]
- Miyamoto, T.; Hosoba, K.; Ochiai, H.; Royba, E.; Izumi, H.; Sakuma, T.; Yamamoto, T.; Dynlacht, B.D.; Matsuura, S. The Microtubule-Depolymerizing activity of a mitotic kinesin protein KIF2A drives primary cilia disassembly coupled with cell proliferation. Cell Rep. 2015, 10, 664–673. [Google Scholar] [CrossRef]
- Michaud, E.J.; Yoder, B.K. The primary cilium in cell signaling and cancer. Cancer Res. 2006, 66, 6463–6467. [Google Scholar] [CrossRef]
- Frett, B.; Brown, R.V.; Ma, M.; Hu, W.; Han, H.; Li, H.Y. Therapeutic melting pot of never in mitosis gene a related kinase 2 (Nek2): A perspective on Nek2 as an oncology target and recent advancements in Nek2 small molecule inhibition. J. Med. Chem. 2014, 57, 5835–5844. [Google Scholar] [CrossRef]
- Gradilone, S.A.; Habringer, S.; Masyuk, T.V.; Howard, B.N.; Masyuk, A.I.; Larusso, N.F. HDAC6 is overexpressed in cystic cholangiocytes and its inhibition reduces cystogenesis. Am. J. Pathol. 2014, 184, 600–608. [Google Scholar] [CrossRef]
- Lorenzo Pisarello, M.; Masyuk, T.V.; Gradilone, S.A.; Masyuk, A.I.; Ding, J.F.; Lee, P.Y.; LaRusso, N.F. Combination of a Histone Deacetylase 6 Inhibitor and a Somatostatin Receptor Agonist Synergistically Reduces Hepatorenal Cystogenesis in an Animal Model of Polycystic Liver Disease. Am. J. Pathol. 2018, 188, 981–994. [Google Scholar] [CrossRef]
- Sarkisian, M.R.; Siebzehnrubl, D.; Hoang-Minh, L.; Deleyrolle, L.; Silver, D.J.; Siebzehnrubl, F.A.; Guadiana, S.M.; Srivinasan, G.; Semple-Rowland, S.; Harrison, J.K.; et al. Detection of primary cilia in human glioblastoma. J. Neurooncol. 2014, 117, 15–24. [Google Scholar] [CrossRef]
- Gradilone, S.A.; Radtke, B.N.; Bogert, P.S.; Huang, B.Q.; Gajdos, G.B.; LaRusso, N.F. HDAC6 inhibition restores ciliary expression and decreases tumor growth. Cancer Res. 2013, 73, 2259–2270. [Google Scholar] [CrossRef]
- Xiang, W.; Guo, F.; Cheng, W.; Zhang, J.; Huang, J.; Wang, R.; Ma, Z.; Xu, K. HDAC6 inhibition suppresses chondrosarcoma by restoring the expression of primary cilia. Oncol. Rep. 2017, 38, 229–236. [Google Scholar] [CrossRef]
- Higgins, M.; Obaidi, I.; McMorrow, T. Primary cilia and their role in cancer (Review). Oncol. Lett. 2019, 17, 3041–3047. [Google Scholar] [CrossRef]
- Peixoto, E.; Richard, S.; Pant, K.; Biswas, A.; Gradilone, S.A. The primary cilium: Its role as a tumor suppressor organelle. Biochem. Pharmacol. 2020, 175, 113906. [Google Scholar] [CrossRef]
- Kiseleva, A.A.; Nikonova, A.S.; Golemis, E.A. Patterns of Ciliation and Ciliary Signaling in Cancer. Rev. Physiol. Biochem. Pharmacol. 2020. [Google Scholar] [CrossRef]
- Sabanovic, B.; Giulietti, M.; Piva, F. Role of primary cilium in pancreatic ductal adenocarcinoma (Review). Int. J. Oncol. 2020, 57, 1095–1102. [Google Scholar] [CrossRef]
- Moser, J.J.; Fritzler, M.J.; Rattner, J.B. Primary ciliogenesis defects are associated with human astrocytoma/glioblastoma cells. BMC Cancer 2009, 9, 1–12. [Google Scholar] [CrossRef]
- Seeley, E.S.; Carrière, C.; Goetze, T.; Longnecker, D.S.; Korc, M. Pancreatic cancer and precursor pancreatic intraepithelial neoplasia lesions are devoid of primary cilia. Cancer Res. 2009, 69, 422–430. [Google Scholar] [CrossRef]
- Tian, H.; Callahan, C.A.; Dupree, K.J.; Darbonne, W.C.; Ahn, C.P.; Scales, S.J.; De Sauvage, F.J. Hedgehog signaling is restricted to the stromal compartment during pancreatic carcinogenesis. Proc. Natl. Acad. Sci. USA 2009, 106, 4254–4259. [Google Scholar] [CrossRef]
- Yuan, K.; Frolova, N.; Xie, Y.; Wang, D.; Cook, L.; Kwon, Y.J.; Steg, A.D.; Serra, R.; Frost, A.R. Primary cilia are decreased in breast cancer: Analysis of a collection of human breast cancer cell lines and tissues. J. Histochem. Cytochem. 2010, 58, 857–870. [Google Scholar] [CrossRef]
- Nobutani, K.; Shimono, Y.; Yoshida, M.; Mizutani, K.; Minami, A.; Kono, S.; Mukohara, T.; Yamasaki, T.; Itoh, T.; Takao, S.; et al. Absence of primary cilia in cell cycle-arrested human breast cancer cells. Genes Cells 2014, 19, 141–152. [Google Scholar] [CrossRef]
- Menzl, I.; Lebeau, L.; Pandey, R.; Hassounah, N.B.; Li, F.W.; Nagle, R.; Weihs, K.; McDermott, K.M. Loss of primary cilia occurs early in breast cancer development. Cilia 2014, 3, 7. [Google Scholar] [CrossRef]
- Hassounah, N.B.; Nunez, M.; Fordyce, C.; Roe, D.; Nagle, R.; Bunch, T.; McDermott, K.M. Inhibition of ciliogenesis promotes Hedgehog signaling, tumorigenesis, and metastasis in breast cancer. Mol. Cancer Res. 2017, 15, 1421–1430. [Google Scholar] [CrossRef] [PubMed]
- Hassounah, N.B.; Nagle, R.; Saboda, K.; Roe, D.J.; Dalkin, B.L.; McDermott, K.M. Primary Cilia Are Lost in Preinvasive and Invasive Prostate Cancer. PLoS ONE 2013, 8, e68521. [Google Scholar] [CrossRef] [PubMed]
- Fu, W.; Asp, P.; Canter, B.; Dynlacht, B.D. Primary cilia control hedgehog signaling during muscle differentiation and are deregulated in rhabdomyosarcoma. Proc. Natl. Acad. Sci. USA 2014, 111, 9151–9156. [Google Scholar] [CrossRef] [PubMed]
- Ho, L.; Ali, S.A.; Al-Jazrawe, M.; Kandel, R.; Wunder, J.S.; Alman, B.A. Primary cilia attenuate hedgehog signalling in neoplastic chondrocytes. Oncogene 2013, 32, 5388–5396. [Google Scholar] [CrossRef]
- Jackman, W.R.; Yoo, J.J.; Stock, D.W. Hedgehog signaling is required at multiple stages of zebrafish tooth development. BMC Dev. Biol. 2010, 10. [Google Scholar] [CrossRef]
- Cobourne, M.T.; Sharpe, P.T. Sonic Hedgehog Signaling and the Developing Tooth. Curr. Top. Dev. Biol. 2004, 65, 255–287. [Google Scholar] [CrossRef]
- Rallis, A.; Navarro, J.A.; Rass, M.; Hu, A.; Birman, S.; Schneuwly, S.; Thérond, P.P. Hedgehog Signaling Modulates Glial Proteostasis and Lifespan. Cell Rep. 2020, 30, 2627–2643.e5. [Google Scholar] [CrossRef]
- Petrova, R.; Joyner, A.L. Roles for Hedgehog signaling in adult organ homeostasis and repair. Development 2014, 141, 3445–3457. [Google Scholar] [CrossRef]
- Büller, N.V.J.A.; Rosekrans, S.L.; Westerlund, J.; van den Brink, G.R. Hedgehog signaling and maintenance of homeostasis in the intestinal epithelium. Physiology 2012, 27, 148–155. [Google Scholar] [CrossRef][Green Version]
- Heemskerk, J.; DiNardo, S. Drosophila hedgehog acts as a morphogen in cellular patterning. Cell 1994, 76, 449–460. [Google Scholar] [CrossRef]
- Chapouly, C.; Guimbal, S.; Hollier, P.L.; Renault, M.A. Role of hedgehog signaling in vasculature development, differentiation, and maintenance. Int. J. Mol. Sci. 2019, 20, 3076. [Google Scholar] [CrossRef]
- Ehlen, H.W.A.; Buelens, L.A.; Vortkamp, A. Hedgehog signaling in skeletal develoment. Birth Defects Res. Part C Embryo Today Rev. 2006, 78, 267–279. [Google Scholar] [CrossRef]
- Hebrok, M. Hedgehog signaling in pancreas development. Mech. Dev. 2003, 120, 45–57. [Google Scholar] [CrossRef]
- Katoh, Y.; Katoh, M. Hedgehog Target Genes: Mechanisms of Carcinogenesis Induced by Aberrant Hedgehog Signaling Activation. Curr. Mol. Med. 2009, 9, 873–886. [Google Scholar] [CrossRef]
- Jeng, K.S.; Chang, C.F.; Lin, S.S. Sonic hedgehog signaling in organogenesis, tumors, and tumor microenvironments. Int. J. Mol. Sci. 2020, 21, 758. [Google Scholar] [CrossRef]
- Haycraft, C.J.; Banizs, B.; Aydin-Son, Y.; Zhang, Q.; Michaud, E.J.; Yoder, B.K. Gli2 and Gli3 localize to cilia and require the intraflagellar transport protein polaris for processing and function. PLoS Genet. 2005, 1, e10053. [Google Scholar] [CrossRef]
- Tukachinsky, H.; Lopez, L.V.; Salic, A. A mechanism for vertebrate Hedgehog signaling: Recruitment to cilia and dissociation of SuFu-Gli protein complexes. J. Cell Biol. 2010, 191, 415–428. [Google Scholar] [CrossRef]
- Dai, P.; Akimaru, H.; Tanaka, Y.; Maekawa, T.; Nakafuku, M.; Ishii, S. Sonic hedgehog-induced activation of the Gli1 promoter is mediated by GLI3. J. Biol. Chem. 1999, 274, 8143–8152. [Google Scholar] [CrossRef]
- Su, Y.; Ospina, J.K.; Zhang, J.; Michelson, A.P.; Schoen, A.M.; Zhu, A.J. Sequential phosphorylation of smoothened transduces graded hedgehog signaling. Sci. Signal. 2011, 4, 1–15. [Google Scholar] [CrossRef]
- Rohatgi, R.; Milenkovic, L.; Scott, M.P. Patched1 Regulates Hedgehog Signaling at the Primary Cilium. Science 2007, 317, 372–376. [Google Scholar] [CrossRef]
- Han, Y.G.; Kim, H.J.; Dlugosz, A.A.; Ellison, D.W.; Gilbertson, R.J.; Alvarez-Buylla, A. Dual and opposing roles of primary cilia in medulloblastoma development. Nat. Med. 2009, 15, 1062–1065. [Google Scholar] [CrossRef] [PubMed]
- Barakat, M.T.; Humke, E.W.; Scott, M.P. Kif3a is necessary for initiation and maintenance of medulloblastoma. Carcinogenesis 2013, 34, 1382–1392. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.Y.; Seol, A.D.; So, P.-L.; Ermilov, A.N.; Bichakjian, C.K.; Epstein, E.H.; Dlugosz, A.A.; Reiter, J.F. Primary cilia can both mediate and suppress Hedgehog pathway–dependent tumorigenesis. Nat. Med. 2009, 15, 1055–1061. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Grausam, K.B.; Wang, J.; Lun, M.P.; Ohli, J.; Lidov, H.G.W.; Calicchio, M.L.; Zeng, E.; Salisbury, J.L.; Wechsler-Reya, R.J.; et al. Sonic Hedgehog promotes proliferation of Notch-dependent monociliated choroid plexus tumour cells. Nat. Cell Biol. 2016, 18, 418–430. [Google Scholar] [CrossRef]
- Guen, V.J.; Chavarria, T.E.; Kröger, C.; Ye, X.; Weinberg, R.A.; Lees, J.A. EMT programs promote basal mammary stem cell and tumor-initiating cell stemness by inducing primary ciliogenesis and Hedgehog signaling. Proc. Natl. Acad. Sci. USA 2017, 114, E10532–E10539. [Google Scholar] [CrossRef]
- Lee, P.L.; Johnson, D.E.; Cousens, L.S.; Fried, V.A.; Williams, L.T. Purification and complementary DNA cloning of a receptor for basic fibroblast growth factor. Science 1989, 245, 57–60. [Google Scholar] [CrossRef]
- Kornbluth, S.; Paulson, K.E.; Hanafusa, H. Novel tyrosine kinase identified by phosphotyrosine antibody screening of cDNA libraries. Mol. Cell. Biol. 1988, 8, 5541–5544. [Google Scholar] [CrossRef]
- Keegan, K.; Johnson, D.E.; Williams, L.T.; Hayman, M.J. Isolation of an additional member of the fibroblast growth factor receptor family, FGFR-3. Proc. Natl. Acad. Sci. USA 1991, 88, 1095–1099. [Google Scholar] [CrossRef]
- Partanen, J.; Makela, T.P.; Eerola, E.; Korhonen, J.; Hirvonen, H.; Claesson-Welsh, L.; Alitalo, K. FGFR-4, a novel acidic fibroblast growth factor receptor with a distinct expression pattern. EMBO J. 1991, 10, 1347–1354. [Google Scholar] [CrossRef]
- Ornitz, D.M.; Xu, J.; Colvin, J.S.; McEwen, D.G.; MacArthur, C.A.; Coulier, F.; Gao, G.; Goldfarb, M. Receptor specificity of the fibroblast growth factor family. J. Biol. Chem. 1996, 271, 15292–15297. [Google Scholar] [CrossRef]
- Ornitz, D.M.; Itoh, N. Fibroblast growth factors. Genome Biol. 2001, 2, reviews3005.1. [Google Scholar] [CrossRef]
- Ornitz, D.M.; Marie, P.J. Fibroblast growth factor signaling in skeletal development and disease. Genes Dev. 2015, 29, 1463–1486. [Google Scholar] [CrossRef]
- Turner, N.; Grose, R. Fibroblast growth factor signalling: From development to cancer. Nat. Rev. Cancer 2010, 10, 116–129. [Google Scholar] [CrossRef]
- Plotnikov, A.N.; Schlessinger, J.; Hubbard, S.R.; Mohammadi, M. Structural Basis for FGF Receptor Dimerization and Activation et al Dimerization of the extracellular domains leads to juxtaposition of the cytoplasmic domains and. Cell 1999, 98, 641–650. [Google Scholar] [CrossRef]
- Eswarakumar, V.P.; Lax, I.; Schlessinger, J. Cellular signaling by fibroblast growth factor receptors. Cytokine Growth Factor Rev. 2005, 16, 139–149. [Google Scholar] [CrossRef]
- Schlessinger, J.; Plotnikov, A.N.; Ibrahimi, O.A.; Eliseenkova, A.V.; Yeh, B.K.; Yayon, A.; Linhardt, R.J.; Mohammadi, M. Crystal Structure of a Ternary FGF-FGFR-Heparin Complex Reveals a Dual Role for Heparin in FGFR Binding and Dimerization. Mol. Cell 2000, 6, 743–750. [Google Scholar] [CrossRef]
- Luo, Y.; Ye, S.; Kan, M.; McKeehan, W.L. Control of Fibroblast Growth Factor (FGF) 7- and FGF1-induced mitogenesis and downstream signaling by distinct heparin octasaccharide motifs. J. Biol. Chem. 2006, 281, 21052–21061. [Google Scholar] [CrossRef]
- Goetz, R.; Beenken, A.; Ibrahimi, O.A.; Kalinina, J.; Olsen, S.K.; Eliseenkova, A.V.; Xu, C.; Neubert, T.A.; Zhang, F.; Linhardt, R.J.; et al. Molecular Insights into the Klotho-Dependent, Endocrine Mode of Action of Fibroblast Growth Factor 19 Subfamily Members. Mol. Cell. Biol. 2007, 27, 3417–3428. [Google Scholar] [CrossRef]
- Goetz, R.; Ohnishi, M.; Ding, X.; Kurosu, H.; Wang, L.; Akiyoshi, J.; Ma, J.; Gai, W.; Sidis, Y.; Pitteloud, N.; et al. Klotho Coreceptors Inhibit Signaling by Paracrine Fibroblast Growth Factor 8 Subfamily Ligands. Mol. Cell. Biol. 2012, 32, 1944–1954. [Google Scholar] [CrossRef]
- Lin, B.C.; Wang, M.; Blackmore, C.; Desnoyers, L.R. Liver-specific activities of FGF19 require klotho beta. J. Biol. Chem. 2007, 282, 27277–27284. [Google Scholar] [CrossRef]
- Quarto, N.; Amalric, F. Heparan sulfate proteoglycans as transducers of FGF-2 signalling. J. Cell Sci. 1994, 107, 3201–3212. [Google Scholar] [CrossRef]
- Zhang, Z.; Coomans, C.; David, G. Membrane heparan sulfate proteoglycan-supported FGF2-FGFR1 signaling: Evidence in support of the “cooperative end structures” model. J. Biol. Chem. 2001, 276, 41921–41929. [Google Scholar] [CrossRef]
- Ornitz, D.M.; Yayon, A.; Flanagan, J.G.; Svahn, C.M.; Levi, E.; Leder, P. Heparin is required for cell-free binding of basic fibroblast growth factor to a soluble receptor and for mitogenesis in whole cells. Mol. Cell. Biol. 1992, 12, 240–247. [Google Scholar] [CrossRef]
- Spivak-Kroizman, T.; Lemmon, M.A.; Dikic, I.; Ladbury, J.E.; Pinchasi, D.; Huang, J.; Jaye, M.; Crumley, G.; Schlessinger, J.; Lax, I. Heparin-induced oligomerization of FGF molecules is responsible for FGF receptor dimerization, activation, and cell proliferation. Cell 1994, 79, 1015–1024. [Google Scholar] [CrossRef]
- Rapraeger, A.C.; Krufka, A.; Olwin, B.B. Requirement of heparan sulfate for bFGF-mediated fibroblast growth and myoblast differentiation. Science 1991, 252, 1705–1708. [Google Scholar] [CrossRef]
- Kuro-o, M. The Klotho proteins in health and disease. Nat. Rev. Nephrol. 2019, 15, 27–44. [Google Scholar] [CrossRef]
- Hu, M.C.; Shiizaki, K.; Kuro-O, M.; Moe, O.W. Fibroblast growth factor 23 and klotho: Physiology and pathophysiology of an endocrine network of mineral metabolism. Annu. Rev. Physiol. 2013, 75, 503–533. [Google Scholar] [CrossRef]
- Tacer, K.F.; Bookout, A.L.; Ding, X.; Kurosu, H.; John, G.B.; Wang, L.; Goetz, R.; Mohammadi, M.; Kuro-o, M.; Mangelsdorf, D.J.; et al. Research resource: Comprehensive expression atlas of the fibroblast growth factor system in adult mouse. Mol. Endocrinol. 2010, 24, 2050–2064. [Google Scholar] [CrossRef]
- Arman, E.; Haffner-Krausz, R.; Gorivodsky, M.; Lonai, P. Fgfr2 is required for limb outgrowth and lung-branching morphogenesis. Proc. Natl. Acad. Sci. USA 1999, 96, 11895–11899. [Google Scholar] [CrossRef]
- Danopoulos, S.; Thornton, M.E.; Grubbs, B.H.; Frey, M.R.; Warburton, D.; Bellusci, S.; Al Alam, D. Discordant roles for FGF ligands in lung branching morphogenesis between human and mouse. J. Pathol. 2019, 247, 254–265. [Google Scholar] [CrossRef] [PubMed]
- McDougall, K.; Kubu, C.; Verdi, J.M.; Meakin, S.O. Developmental expression patterns of the signaling adapters FRS-2 and FRS-3 during early embryogenesis. Mech. Dev. 2001, 103, 145–148. [Google Scholar] [CrossRef]
- Weinstein, M.; Xu, X.; Ohyama, K.; Deng, C.X. FGFR-3 and FGFR-4 function cooperatively to direct alveogenesis in the murine lung. Development 1998, 125, 3615–3623. [Google Scholar] [CrossRef] [PubMed]
- Dudley, A.T.; Godin, R.E.; Robertson, E.J. Interaction between FGF and BMP signaling pathways regulates development of metanephric mesenchyme. Genes Dev. 1999, 13, 1601–1613. [Google Scholar] [CrossRef] [PubMed]
- Kastner, S.; Elias, M.C.; Rivera, A.J.; Yablonka-Reuveni, Z. Gene expression patterns of the fibroblast growth factors and their receptors during myogenesis of rat satellite cells. J. Histochem. Cytochem. 2000, 48, 1079–1096. [Google Scholar] [CrossRef]
- Walker, K.A.; Sims-Lucas, S.; Bates, C.M. Fibroblast growth factor receptor signaling in kidney and lower urinary tract development. Pediatr. Nephrol. 2016, 31, 885–895. [Google Scholar] [CrossRef]
- Colvin, J.S.; Bohne, B.A.; Harding, G.W.; McEwen, D.G.; Ornitz, D.M. Skeletal overgrowth and deafness in mice lacking fibroblast growth factor receptor 3. Nat. Genet. 1996, 12, 390–397. [Google Scholar] [CrossRef]
- Shimada, T.; Kakitani, M.; Yamazaki, Y.; Hasegawa, H.; Takeuchi, Y.; Fujita, T.; Fukumoto, S.; Tomizuka, K.; Yamashita, T. Targeted ablation of Fgf23 demonstrates an essential physiological role of FGF23 in phosphate and vitamin D metabolism. J. Clin. Invest. 2004, 113, 561–568. [Google Scholar] [CrossRef]
- Zhou, M.; Luo, J.; Chen, M.; Yang, H.; Learned, R.M.; DePaoli, A.M.; Tian, H.; Ling, L. Mouse species-specific control of hepatocarcinogenesis and metabolism by FGF19/FGF15. J. Hepatol. 2017, 66, 1182–1192. [Google Scholar] [CrossRef]
- Tomlinson, E.; Fu, L.; John, L.; Hultgren, B.; Huang, X.; Renz, M.; Stephan, J.P.; Tsai, S.P.; Powell-Braxton, L.; French, D.; et al. Transgenic mice expressing human fibroblast growth factor-19 display increased metabolic rate and decreased adiposity. Endocrinology 2002, 143, 1741–1747. [Google Scholar] [CrossRef]
- Potthoff, M.J.; Boney-Montoya, J.; Choi, M.; He, T.; Sunny, N.E.; Satapati, S.; Suino-Powell, K.; Xu, H.E.; Gerard, R.D.; Finck, B.N.; et al. FGF15/19 regulates hepatic glucose metabolism by inhibiting the CREB-PGC-1α pathway. Cell Metab. 2011, 13, 729–738. [Google Scholar] [CrossRef]
- Quarles, L.D. Skeletal secretion of FGF-23 regulates phosphate and vitamin D metabolism. Nat. Rev. Endocrinol. 2012, 8, 276–286. [Google Scholar] [CrossRef]
- Xie, Y.; Su, N.; Yang, J.; Tan, Q.; Huang, S.; Jin, M.; Ni, Z.; Zhang, B.; Zhang, D.; Luo, F.; et al. FGF/FGFR signaling in health and disease. Signal Transduct. Target. Ther. 2020, 5. [Google Scholar] [CrossRef]
- Floss, T.; Arnold, H.H.; Braun, T. A role for FGF-6 in skeletal muscle regeneration. Genes Dev. 1997, 11, 2040–2051. [Google Scholar] [CrossRef]
- Schmid, G.J.; Kobayashi, C.; Sandell, L.J.; Ornitz, D.M. Fibroblast growth factor expression during skeletal fracture healing in mice. Dev. Dyn. 2009, 238, 766–774. [Google Scholar] [CrossRef]
- Nakajima, A.; Nakajima, F.; Shimizu, S.; Ogasawara, A.; Wanaka, A.; Moriya, H.; Einhorn, T.A.; Yamazaki, M. Spatial and temporal gene expression for fibroblast growth factor type I receptor (FGFR1) during fracture healing in the rat. Bone 2001, 29, 458–466. [Google Scholar] [CrossRef]
- Goebel, S.; Lienau, J.; Rammoser, U.; Seefried, L.; Wintgens, K.F.; Seufert, J.; Duda, G.; Jakob, F.; Ebert, R. FGF23 is a putative marker for bone healing and regeneration. J. Orthop. Res. 2009, 27, 1141–1146. [Google Scholar] [CrossRef]
- Hurley, M.M.; Adams, D.J.; Wang, L.; Jiang, X.; Burt, P.M.; Du, E.; Xiao, L. Accelerated fracture healing in transgenic mice overexpressing an anabolic isoform of fibroblast growth factor 2. J. Cell. Biochem. 2016, 117, 599–611. [Google Scholar] [CrossRef]
- Kunova Bosakova, M.; Varecha, M.; Hampl, M.; Duran, I.; Nita, A.; Buchtova, M.; Dosedelova, H.; Machat, R.; Xie, Y.; Ni, Z.; et al. Regulation of ciliary function by fibroblast growth factor signaling identifies FGFR3-related disorders achondroplasia and thanatophoric dysplasia as ciliopathies. Hum. Mol. Genet. 2018, 27, 1093–1105. [Google Scholar] [CrossRef]
- Martin, L.; Kaci, N.; Estibals, V.; Goudin, N.; Garfa-Traore, M.; Benoist-Lasselin, C.; Dambroise, E.; Legeai-Mallet, L. Constitutively-active FGFR3 disrupts primary cilium length and IFT20 trafficking in various chondrocyte models of achondroplasia. Hum. Mol. Genet. 2018, 27, 1–13. [Google Scholar] [CrossRef]
- Kunova Bosakova, M.; Nita, A.; Gregor, T.; Varecha, M.; Gudernova, I.; Fafilek, B.; Barta, T.; Basheer, N.; Abraham, S.P.; Balek, L.; et al. Fibroblast growth factor receptor influences primary cilium length through an interaction with intestinal cell kinase. Proc. Natl. Acad. Sci. USA 2019, 116, 4316–4325. [Google Scholar] [CrossRef]
- Katoh, M.; Nakagama, H. FGF Receptors: Cancer Biology and Therapeutics. Med. Res. Rev. 2014, 34, 280–300. [Google Scholar] [CrossRef] [PubMed]
- Katoh, M. Fibroblast growth factor receptors as treatment targets in clinical oncology. Nat. Rev. Clin. Oncol. 2019, 16, 105–122. [Google Scholar] [CrossRef] [PubMed]
- Dutt, A.; Salvesen, H.B.; Chen, T.H.; Ramos, A.H.; Onofrio, R.C.; Hatton, C.; Nicoletti, R.; Winckler, W.; Grewal, R.; Hanna, M.; et al. Drug-sensitive FGFR2 mutations in endometrial carcinoma. Proc. Natl. Acad. Sci. USA 2008, 105, 8713–8717. [Google Scholar] [CrossRef] [PubMed]
- Thomas, A.; Lee, J.H.; Abdullaev, Z.; Park, K.S.; Pineda, M.; Saidkhodjaeva, L.; Miettinen, M.; Wang, Y.; Pack, S.D.; Giaccone, G. Characterization of fibroblast growth factor receptor 1 in small-cell lung cancer. J. Thorac. Oncol. 2014, 9, 567–571. [Google Scholar] [CrossRef]
- Rosty, C.; Aubriot, M.H.; Cappellen, D.; Bourdin, J.; Cartier, I.; Thiery, J.P.; Sastre-Garau, X.; Radvanyi, F. Clinical and biological characteristics of cervical neoplasias with FGFR3 mutation. Mol. Cancer 2005, 4, 2–9. [Google Scholar] [CrossRef]
- Neugebauer, J.M.; Amack, J.D.; Peterson, A.G.; Bisgrove, B.W.; Yost, H.J. FGF signalling during embryo development regulates cilia length in diverse epithelia. Nature 2009, 458, 651–654. [Google Scholar] [CrossRef]
- Essner, J.J.; Amack, J.D.; Nyholm, M.K.; Harris, E.B.; Yost, H.J. Kupffer’s vesicle is a ciliated organ of asymmetry in the zebrafish embryo that initiates left-right development of the brain, heart and gut. Development 2005, 132, 1247–1260. [Google Scholar] [CrossRef]
- Yamauchi, H.; Miyakawa, N.; Miyake, A.; Itoh, N. Fgf4 is required for left-right patterning of visceral organs in zebrafish. Dev. Biol. 2009, 332, 177–185. [Google Scholar] [CrossRef]
- Liu, D.-W.W.; Hsu, C.-H.H.; Tsai, S.-M.M.; Hsiao, C.-D.; Wang, W.-P.P. A Variant of Fibroblast Growth Factor Receptor 2 (Fgfr2) Regulates Left-Right Asymmetry in Zebrafish. PLoS ONE 2011, 6, e21793. [Google Scholar] [CrossRef]
- Sempou, E.; Lakhani, O.A.; Amalraj, S.; Khokha, M.K. Candidate Heterotaxy Gene FGFR4 Is Essential for Patterning of the Left-Right Organizer in Xenopus. Front. Physiol. 2018, 9, 1–9. [Google Scholar] [CrossRef]
- Hong, S.K.; Dawid, I.B. FGF-dependent left-right asymmetry patterning in zebrafish is mediated by Ier2 and Fibp1. Proc. Natl. Acad. Sci. USA 2009, 106, 2230–2235. [Google Scholar] [CrossRef]
- Neugebauer, J.M.; Cadwallader, A.B.; Amack, J.D.; Bisgrove, B.W.; Joseph Yost, H. Differential roles for 3-OSTs in the regulation of cilia length and motility. Development 2013, 140, 3892–3902. [Google Scholar] [CrossRef][Green Version]
- Caron, A.; Xu, X.; Lin, X. Wnt/β-catenin signaling directly regulates Foxj1 expression and ciliogenesis in zebrafish Kupffer’s vesicle. Development 2012, 139, 514–524. [Google Scholar] [CrossRef]
- Bonnafe, E.; Touka, M.; AitLounis, A.; Baas, D.; Barras, E.; Ucla, C.; Moreau, A.; Flamant, F.; Dubruille, R.; Couble, P.; et al. The Transcription Factor RFX3 Directs Nodal Cilium Development and Left-Right Asymmetry Specification. Mol. Cell. Biol. 2004, 24, 4417–4427. [Google Scholar] [CrossRef]
- Bisgrove, B.W.; Snarr, B.S.; Emrazian, A.; Yost, H.J. Polaris and Polycystin-2 in dorsal forerunner cells and Kupffer’s vesicle are required for specification of the zebrafish left-right axis. Dev. Biol. 2005, 287, 274–288. [Google Scholar] [CrossRef]
- Honda, A.; Kita, T.; Seshadri, S.V.; Misaki, K.; Ahmed, Z.; Ladbury, J.E.; Richardson, G.P.; Yonemura, S.; Ladher, R.K. FGFR1-mediated protocadherin-15 loading mediates cargo specificity during intraflagellar transport in inner ear hair-cell kinocilia. Proc. Natl. Acad. Sci. USA 2018, 115, 8388–8393. [Google Scholar] [CrossRef]
- Yuan, X.; Liu, M.; Cao, X.; Yang, S. Ciliary IFT80 regulates dental pulp stem cells differentiation by FGF/FGFR1 and Hh/BMP2 signaling. Int. J. Biol. Sci. 2019, 15, 2087–2099. [Google Scholar] [CrossRef]
- Taylor, S.P.; Bosakova, M.K.; Varecha, M.; Balek, L.; Barta, T.; Trantirek, L.; Jelinkova, I.; Duran, I.; Vesela, I.; Forlenza, K.N.; et al. An inactivating mutation in intestinal cell kinase, ICK, impairs hedgehog signalling and causes short rib-polydactyly syndrome. Hum. Mol. Genet. 2016, 25, 3998–4011. [Google Scholar] [CrossRef]
- Moon, H.; Song, J.; Shin, J.O.; Lee, H.; Kim, H.K.; Eggenschwiller, J.T.; Bok, J.; Ko, H.W. Intestinal cell kinase, aprotein associated with endocrine-cerebro-osteodysplasia syndrome is a key regulator of cilia length and Hedgehog signaling. Proc. Natl. Acad. Sci. USA 2014, 111, 8541–8546. [Google Scholar] [CrossRef]
- Chaya, T.; Omori, Y.; Kuwahara, R.; Furukawa, T. ICK is essential for cell type-specific ciliogenesis and the regulation of ciliary transport. EMBO J. 2014, 33, 1227–1242. [Google Scholar] [CrossRef]
- Tong, Y.; Park, S.H.; Wu, D.; Xu, W.; Guillot, S.J.; Jin, L.; Li, X.; Wang, Y.; Lin, C.-S.; Fu, Z. An essential role of intestinal cell kinase in lung development is linked to the perinatal lethality of human ECO syndrome. FEBS Lett. 2017, 591, 1247–1257. [Google Scholar] [CrossRef]
- Ding, M.; Jin, L.; Xie, L.; Park, S.H.; Tong, Y.; Wu, D.; Chhabra, A.B.; Fu, Z.; Li, X. A Murine Model for Human ECO Syndrome Reveals a Critical Role of Intestinal Cell Kinase in Skeletal Development. Calcif. Tissue Int. 2018, 102, 348–357. [Google Scholar] [CrossRef]
- Okamoto, S.; Chaya, T.; Omori, Y.; Kuwahara, R.; Kubo, S.; Sakaguchi, H.; Furukawa, T. Ick ciliary kinase is essential for planar cell polarity formation in inner ear hair cells and hearing function. J. Neurosci. 2017, 37, 2073–2085. [Google Scholar] [CrossRef]
- Berman, S.A.; Wilson, N.F.; Haas, N.A.; Lefebvre, P.A. A Novel MAP Kinase Regulates Flagellar Length in Chlamydomonas. Curr. Biol. 2003, 13, 1145–1149. [Google Scholar] [CrossRef]
- Burghoorn, J.; Dekkers, M.P.J.; Rademakers, S.; De Jong, T.; Willemsen, R.; Jansen, G. Mutation of the MAP kinase DYF-5 affects docking and undocking of kinesin-2 motors and reduces their speed in the cilia of Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 2007, 104, 7157–7162. [Google Scholar] [CrossRef]
- Rousseau, F.; El Ghouzzi, V.; Delezoide, A.L.; Legeai-Mallet, L.; Le Merrer, M.; Munnich, A.; Bonaventure, J. Missense FGFR3 mutations create cysteine residues in thanatophoric dwarfism type I (TD1). Hum. Mol. Genet. 1996, 5, 509–512. [Google Scholar] [CrossRef]
- Tavormina, P.L.; Shiang, R.; Thompson, L.M.; Zhu, Y.Z.; Wilkin, D.J.; Lachman, R.S.; Wilcox, W.R.; Rimoin, D.L.; Cohn, D.H.; Wasmuth, J.J. Thanatophoric dysplasia (types I and II) caused by distinct mutations in fibroblast growth factor receptor 3. Nat. Genet. 1995, 9, 321–328. [Google Scholar] [CrossRef]
- Tavormina, P.L.; Rimoin, D.L.; Cohn, D.H.; Zhu, Y.Z.; Shiang, R.; Wasmuth, J.J. Another mutation that results in the substitution of an unpaired cysteine residue in the extracellular domain of FGFR3 in thanatophoric dysplasia type I. Hum. Mol. Genet. 1995, 4, 2175–2177. [Google Scholar] [CrossRef]
- Shiang, R.; Thompson, L.M.; Zhu, Y.Z.; Church, D.M.; Fielder, T.J.; Bocian, M.; Winokur, S.T.; Wasmuth, J.J. Mutations in the transmembrane domain of FGFR3 cause the most common genetic form of dwarfism, achondroplasia. Cell 1994, 78, 335–342. [Google Scholar] [CrossRef]
- Bellus, G.A.; Hefferon, T.W.; De Luna, R.I.O.; Hecht, J.T.; Horton, W.A.; Machado, M.; Kaitila, I.; McIntosh, I.; Francomano, C.A. Achondroplasia is defined by recurrent G380R mutations of FGFR3. Am. J. Hum. Genet. 1995, 56, 368–373. [Google Scholar]
- Van Rhijn, B.W.G.; van Tilborg, A.A.G.; Lurkin, I.; Bonaventure, J.; de Vries, A.; Thiery, J.P.; van der Kwast, T.H.; Zwarthoff, E.C. Novel fibroblast growth factor receptor 3 (FGFR3) mutations in bladder cancer previously identified in non-lethal skeletal disorders. Eur. J. Hum. Genet. 2002, 10, 819–824. [Google Scholar] [CrossRef] [PubMed]
- Gallo, L.H.; Nelson, K.N.; Meyer, A.N.; Donoghue, D.J. Functions of Fibroblast Growth Factor Receptors in cancer defined by novel translocations and mutations. Cytokine Growth Factor Rev. 2015, 26, 425–449. [Google Scholar] [CrossRef] [PubMed]
- Bernard-Pierrot, I.; Brams, A.; Dunois-Lardé, C.; Caillault, A.; Diez de Medina, S.G.; Cappellen, D.; Graff, G.; Thiery, J.P.; Chopin, D.; Ricol, D.; et al. Oncogenic properties of the mutated forms of fibroblast growth factor receptor 3b. Carcinogenesis 2006, 27, 740–747. [Google Scholar] [CrossRef] [PubMed]
- Greulich, H.; Pollock, P.M. Targeting mutant fibroblast growth factor receptors in cancer. Trends Mol. Med. 2011, 17, 283–292. [Google Scholar] [CrossRef]
- Shinmura, K.; Kato, H.; Matsuura, S.; Inoue, Y.; Igarashi, H.; Nagura, K.; Nakamura, S.; Maruyama, K.; Tajima, M.; Funai, K.; et al. A novel somatic FGFR3 mutation in primary lung cancer. Oncol. Rep. 2014, 31, 1219–1224. [Google Scholar] [CrossRef]
- Naski, M.C.; Colvin, J.S.; Coffin, J.D.; Ornitz, D.M. Repression of hedgehog signaling and BMP4 expression in growth plate cartilage by fibroblast growth factor receptor 3. Development 1998, 125, 4977–4988. [Google Scholar] [CrossRef]
- Chen, J.; Chien, K.R. Complexity in simplicity: Monogenic disorders and c complex cardiomyopathies. J. Clin. Invest. 1999, 103, 1483–1485. [Google Scholar] [CrossRef]
- Chen, L.; Li, C.; Qiao, W.; Xu, X.; Deng, C. A Ser365→Cys mutation of fibroblast growth factor receptor 3 in mouse downregulates lhh/PTHrP signals and causes severe achondroplasia. Hum. Mol. Genet. 2001, 10, 457–465. [Google Scholar] [CrossRef]
- Wren, K.N.; Craft, J.M.; Tritschler, D.; Schauer, A.; Patel, D.K.; Smith, E.F.; Porter, M.E.; Kner, P.; Lechtreck, K.F. A Differential Cargo-Loading Model of Ciliary Length Regulation by IFT. Curr. Biol. 2013, 23, 2463–2471. [Google Scholar] [CrossRef]
- Zhou, S.; Xie, Y.; Tang, J.; Huang, J.; Huang, Q.; Xu, W.; Wang, Z.; Luo, F.; Wang, Q.; Chen, H.; et al. FGFR3 Deficiency Causes Multiple Chondroma-like Lesions by Upregulating Hedgehog Signaling. PLoS Genet. 2015, 11, e1005214. [Google Scholar] [CrossRef]
- Du, E.; Lu, C.; Sheng, F.; Li, C.; Li, H.; Ding, N.; Chen, Y.; Zhang, T.; Yang, K.; Xu, Y. Analysis of potential genes associated with primary cilia in bladder cancer. Cancer Manag. Res. 2018, 10, 3047–3056. [Google Scholar] [CrossRef]
- Lee, K.H.S.; Johmura, Y.; Yu, L.R.; Park, J.E.; Gao, Y.; Bang, J.K.; Zhou, M.; Veenstra, T.D.; Yeon Kim, B.; Lee, K.H.S. Identification of a novel Wnt5a-CK1ε-Dvl2-Plk1-mediated primary cilia disassembly pathway. EMBO J. 2012, 31, 3104–3117. [Google Scholar] [CrossRef]
- Hsu, Y.C.; Kao, C.Y.; Chung, Y.F.; Lee, D.C.; Liu, J.W.; Chiu, I.M. Activation of Aurora A kinase through the FGF1/FGFR signaling axis sustains the stem cell characteristics of glioblastoma cells. Exp. Cell Res. 2016, 344, 153–166. [Google Scholar] [CrossRef]
- Li, X.; Martinez-Ledesma, E.; Zhang, C.; Gao, F.; Zheng, S.; Ding, J.; Wu, S.; Nguyen, N.; Clifford, S.C.; Wen, P.Y.; et al. TIE2–FGFR1 interaction induces adaptive PI3K inhibitor resistance by upregulating Aurora A/PlK1/CDK1 signaling in glioblastoma. Cancer Res. 2019, 79, 5088–5101. [Google Scholar] [CrossRef]
- Montaudon, E.; Nikitorowicz-Buniak, J.; Sourd, L.; Morisset, L.; El Botty, R.; Huguet, L.; Dahmani, A.; Painsec, P.; Nemati, F.; Vacher, S.; et al. PLK1 inhibition exhibits strong anti-tumoral activity in CCND1-driven breast cancer metastases with acquired palbociclib resistance. Nat. Commun. 2020, 11. [Google Scholar] [CrossRef]
- Sánchez, I.; Dynlacht, B.D. Cilium assembly and disassembly. Nat. Cell Biol. 2016, 18, 711–717. [Google Scholar] [CrossRef]
- Werner, S.; Pimenta-Marques, A.; Bettencourt-Dias, M. Maintaining centrosomes and cilia. J. Cell Sci. 2017, 130, 3789–3800. [Google Scholar] [CrossRef]
- Wu, Y.M.; Su, F.; Kalyana-Sundaram, S.; Khazanov, N.; Ateeq, B.; Cao, X.; Lonigro, R.J.; Vats, P.; Wang, R.; Lin, S.F.; et al. Identification of targetable FGFR gene fusions in diverse cancers. Cancer Discov. 2013, 3, 636–647. [Google Scholar] [CrossRef]
- L’Hôte, C.G.M.; Knowles, M.A. Cell responses to FGFR3 signalling: Growth, differentiation and apoptosis. Exp. Cell Res. 2005, 304, 417–431. [Google Scholar] [CrossRef]
- Pepper, M.S.; Ferrara, N.; Orci, L.; Montesano, R. Potent synergism between vascular endothelial growth factor and basic fibroblast growth factor in the induction of angiogenesis in vitro. Biochem. Biophys. Res. Commun. 1992, 189, 824–831. [Google Scholar] [CrossRef]
- Lieu, C.; Heymach, J.; Overman, M.; Tran, H.; Kopetz, S. Beyond VEGF: Inhibition of the fibroblast growth factor pathway and antiangiogenesis. Clin. Cancer Res. 2011, 17, 6130–6139. [Google Scholar] [CrossRef]
- Helsten, T.; Elkin, S.; Arthur, E.; Tomson, B.N.; Carter, J.; Kurzrock, R. The FGFR landscape in cancer: Analysis of 4,853 tumors by next-generation sequencing. Clin. Cancer Res. 2016, 22, 259–267. [Google Scholar] [CrossRef]
- Dienstmann, R.; Rodon, J.; Prat, A.; Perez-Garcia, J.; Adamo, B.; Felip, E.; Cortes, J.; Iafrate, A.J.; Nuciforo, P.; Tabernero, J. Genomic aberrations in the FGFR pathway: Opportunities for targeted therapies in solid tumors. Ann. Oncol. 2014, 25, 552–563. [Google Scholar] [CrossRef]
- Kim, S.; Dubrovska, A.; Salamone, R.J.; Walker, J.R.; Grandinetti, K.B.; Bonamy, G.M.C.; Orth, A.P.; Elliott, J.; Porta, D.G.; Garcia-Echeverria, C.; et al. FGFR2 Promotes Breast Tumorigenicity through Maintenance of Breast Tumor-Initiating Cells. PLoS ONE 2013, 8, e51671. [Google Scholar] [CrossRef]
- Tsimafeyeu, I.; Demidov, L.; Stepanova, E.; Wynn, N.; Ta, H. Overexpression of fibroblast growth factor receptors FGFR1 and FGFR2 in renal cell carcinoma. Scand. J. Urol. Nephrol. 2011, 45, 190–195. [Google Scholar] [CrossRef]
- Giri, D.; Ropiquet, F.; Ittmann, M. Alterations in expression of basic fibroblast growth factor (FGF) 2 and its receptor FGFR-1 in human prostate cancer. Clin. Cancer Res. 1999, 5, 1063–1071. [Google Scholar]
- Di Martino, E.; Tomlinson, D.C.; Knowles, M.A. A decade of FGF receptor research in bladder cancer: Past, present, and future challenges. Adv. Urol. 2012, 2012. [Google Scholar] [CrossRef]
- Byron, S.A.; Gartside, M.; Powell, M.A.; Wellens, C.L.; Gao, F.; Mutch, D.G.; Goodfellow, P.J.; Pollock, P.M. Fgfr2 point mutations in 466 endometrioid endometrial tumors: Relationship with msi, kras, pik3ca, ctnnb1 mutations and clinicopathological features. PLoS ONE 2012, 7, e30801. [Google Scholar] [CrossRef]
- Cappellen, D.; De Oliveira, C.; Ricol, D.; de Medina, S.; Bourdin, J.; Sastre-Garau, X.; Chopin, D.; Thiery, J.P.; Radvanyi, F. Frequent activating mutations of FGFR3 in human bladder and cervix carcinomas. Nat. Genet. 1999, 23, 18–20. [Google Scholar] [CrossRef]
- Ahmed, Z.; Schüller, A.C.; Suhling, K.; Tregidgo, C.; Ladbury, J.E. Extracellular point mutations in FGFR2 elicit unexpected changes in intracellular signalling. Biochem. J. 2008, 413, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Neilson, K.M.; Friesel, R. Ligand-independent activation of fibroblast growth factor receptors by point mutations in the extracellular, transmembrane, and kinase domains. J. Biol. Chem. 1996, 271, 25049–25057. [Google Scholar] [CrossRef] [PubMed]
- Krook, M.A.; Reeser, J.W.; Ernst, G.; Barker, H.; Wilberding, M.; Li, G.; Chen, H.Z.; Roychowdhury, S. Fibroblast growth factor receptors in cancer: Genetic alterations, diagnostics, therapeutic targets and mechanisms of resistance. Br. J. Cancer 2021, 124, 880–892. [Google Scholar] [CrossRef] [PubMed]
- Ibrahimi, O.A.; Yeh, B.K.; Eliseenkova, A.V.; Zhang, F.; Olsen, S.K.; Igarashi, M.; Aaronson, S.A.; Linhardt, R.J.; Mohammadi, M. Analysis of Mutations in Fibroblast Growth Factor (FGF) and a Pathogenic Mutation in FGF Receptor (FGFR) Provides Direct Evidence for the Symmetric Two-End Model for FGFR Dimerization. Mol. Cell. Biol. 2005, 25, 671–684. [Google Scholar] [CrossRef] [PubMed]
- Webster, M.K.; D’Avis, P.Y.; Robertson, S.C.; Donoghue, D.J. Profound ligand-independent kinase activation of fibroblast growth factor receptor 3 by the activation loop mutation responsible for a lethal skeletal dysplasia, thanatophoric dysplasia type II. Mol. Cell. Biol. 1996, 16, 4081–4087. [Google Scholar] [CrossRef]
- Naski, M.C.; Wang, Q.; Xu, J.; Ornitz, D.M. Graded activation of fibroblast growth factor receptor 3 by mutations causing achondroplasia and thanatophoric dysplasia. Nat. Genet. 1996, 13, 233–237. [Google Scholar] [CrossRef]
- Bellus, G.A.; Bamshad, M.J.; Przylepa, K.A.; Dorst, J.; Lee, R.R.; Hurko, O.; Jabs, E.W.; Curry, C.J.R.; Wilcox, W.R.; Lachman, R.S.; et al. Severe achondroplasia with developmental delay and Acanthosis nigricans (SADDAN): Phenotypic analysis of a new skeletal dysplasia caused by a Lys650Met mutation in fibroblast growth factor receptor 3. Am. J. Med. Genet. 1999, 85, 53–65. [Google Scholar] [CrossRef]
- Foth, M.; Ahmad, I.; Van Rhijn, B.W.G.; Van Der Kwast, T.; Bergman, A.M.; King, L.; Ridgway, R.; Leung, H.Y.; Fraser, S.; Sansom, O.J.; et al. Fibroblast growth factor receptor 3 activation plays a causative role in urothelial cancer pathogenesis in cooperation with Pten loss in mice. J. Pathol. 2014, 233, 148–158. [Google Scholar] [CrossRef]
- Ahmad, I.; Singh, L.B.; Foth, M.; Morris, C.A.; Taketo, M.M.; Wu, X.R.; Leung, H.Y.; Sansom, O.J.; Iwata, T. K-Ras and β-catenin mutations cooperate with Fgfr3 mutations in mice to promote tumorigenesis in the skin and lung, but not in the bladder. DMM Dis. Model. Mech. 2011, 4, 548–555. [Google Scholar] [CrossRef]
- Mertens, F.; Johansson, B.; Fioretos, T.; Mitelman, F. The emerging complexity of gene fusions in cancer. Nat. Rev. Cancer 2015, 15, 371–381. [Google Scholar] [CrossRef]
- Schram, A.M.; Chang, M.T.; Jonsson, P.; Drilon, A. Fusions in solid tumours: Diagnostic strategies, targeted therapy, and acquired resistance. Nat. Rev. Clin. Oncol. 2017, 14, 735–748. [Google Scholar] [CrossRef]
- De Luca, A.; Abate, R.E.; Rachiglio, A.M.; Maiello, M.R.; Esposito, C.; Schettino, C.; Izzo, F.; Nasti, G.; Normanno, N. FGFR fusions in cancer: From diagnostic approaches to therapeutic intervention. Int. J. Mol. Sci. 2020, 21, 6856. [Google Scholar] [CrossRef]
- Katoh, M. FGFR inhibitors: Effects on cancer cells, tumor microenvironment and whole-body homeostasis (Review). Int. J. Mol. Med. 2016, 38, 3–15. [Google Scholar] [CrossRef]
- Wang, Y.; Ding, X.; Wang, S.; Moser, C.D.; Shaleh, H.M.; Mohamed, E.A.; Chaiteerakij, R.; Allotey, L.K.; Chen, G.; Miyabe, K.; et al. Antitumor effect of FGFR inhibitors on a novel cholangiocarcinoma patient derived xenograft mouse model endogenously expressing an FGFR2-CCDC6 fusion protein. Cancer Lett. 2016, 380, 163–173. [Google Scholar] [CrossRef]
- Arai, Y.; Totoki, Y.; Hosoda, F.; Shirota, T.; Hama, N.; Nakamura, H.; Ojima, H.; Furuta, K.; Shimada, K.; Okusaka, T.; et al. Fibroblast growth factor receptor 2 tyrosine kinase fusions define a unique molecular subtype of cholangiocarcinoma. Hepatology 2014, 59, 1427–1434. [Google Scholar] [CrossRef]
- Ochiai, M.; Yoshihara, Y.; Maru, Y.; Tetsuya, M.; Izumiya, M.; Imai, T.; Hippo, Y. Kras-driven heterotopic tumor development from hepatobiliary organoids. Carcinogenesis 2019. [Google Scholar] [CrossRef]
- Sia, D.; Losic, B.; Moeini, A.; Cabellos, L.; Hao, K.; Revill, K.; Bonal, D.; Miltiadous, O.; Zhang, Z.; Hoshida, Y.; et al. Massive parallel sequencing uncovers actionable FGFR2-PPHLN1 fusion and ARAF mutations in intrahepatic cholangiocarcinoma. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef]
- Li, F.; Meyer, A.N.; Peiris, M.N.; Nelson, K.N.; Donoghue, D.J. Oncogenic fusion protein FGFR2-PPHLN1: Requirements for biological activation, and efficacy of inhibitors. Transl. Oncol. 2020, 13, 100853. [Google Scholar] [CrossRef]
- Williams, S.V.; Hurst, C.D.; Knowles, M.A. Oncogenic FGFR3 gene fusions in bladder cancer. Hum. Mol. Genet. 2013, 22, 795–803. [Google Scholar] [CrossRef]
- Nakanishi, Y.; Akiyama, N.; Tsukaguchi, T.; Fujii, T.; Satoh, Y.; Ishii, N.; Aoki, M. Mechanism of Oncogenic Signal Activation by the Novel Fusion Kinase FGFR3-BAIAP2L1. Mol. Cancer Ther. 2015, 14, 704–712. [Google Scholar] [CrossRef]
- Ren, M.; Cowell, J.K. Constitutive Notch pathway activation in murine ZMYM2-FGFR1-induced T-cell lymphomas associated with atypical myeloproliferative disease. Blood 2011, 117, 6837–6847. [Google Scholar] [CrossRef]
- Agerstam, H.; Järås, M.; Andersson, A.; Johnels, P.; Hansen, N.; Lassen, C.; Rissler, M.; Gisselsson, D.; Olofsson, T.; Richter, J.; et al. Modeling the human 8p11-myeloproliferative syndrome in immunodeficient mice. Blood 2010, 116, 2103–2111. [Google Scholar] [CrossRef] [PubMed]
- Chase, A.; Grand, F.H.; Cross, N.C.P.P. Activity of TKI258 against primary cells and cell lines with FGFR1 fusion genes associated with the 8p11 myeloproliferative syndrome. Blood 2007, 110, 3729–3734. [Google Scholar] [CrossRef] [PubMed]
- Peiris, M.N.; Meyer, A.N.; Nelson, K.N.; Bisom-Rapp, E.W.; Donoghue, D.J. Oncogenic fusion protein BCR-FGFR1 requires the breakpoint cluster region-mediated oligomerization and chaperonin Hsp90 for activation. Haematologica 2020, 105, 1262–1273. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, D.M.; Mirante, T.; Mignogna, C.; Scrima, M.; Migliozzi, S.; Rocco, G.; Franco, R.; Corcione, F.; Viglietto, G.; Malanga, D.; et al. Simultaneous identification of clinically relevant single nucleotide variants, copy number alterations and gene fusions in solid tumors by targeted next-generation sequencing. Oncotarget 2018, 9, 22749–22768. [Google Scholar] [CrossRef][Green Version]
- Di Stefano, A.L.; Fucci, A.; Frattini, V.; Labussiere, M.; Mokhtari, K.; Zoppoli, P.; Marie, Y.; Bruno, A.; Boisselier, B.; Giry, M.; et al. Detection, characterization, and inhibition of FGFR-TACC fusions in IDH wild-type glioma. Clin. Cancer Res. 2015, 21, 3307–3317. [Google Scholar] [CrossRef]
- Singh, D.; Chan, J.M.; Zoppoli, P.; Niola, F.; Sullivan, R.; Castano, A.; Liu, E.M.; Reichel, J.; Porrati, P.; Pellegatta, S.; et al. Transforming fusions of FGFR and TACC genes in human glioblastoma. Science 2012, 337, 1231–1235. [Google Scholar] [CrossRef]
- Costa, R.; Carneiro, B.A.; Taxter, T.; Tavora, F.A.; Kalyan, A.; Pai, S.A.; Chae, Y.K.; Giles, F.J. FGFR3-TACC3 fusion in solid tumors: Mini review. Oncotarget 2016, 7, 55924–55938. [Google Scholar] [CrossRef]
- Weinstein, J.N.; Akbani, R.; Broom, B.M.; Wang, W.; Verhaak, R.G.W.; McConkey, D.; Lerner, S.; Morgan, M.; Creighton, C.J.; Smith, C.; et al. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 2014, 507, 315–322. [Google Scholar] [CrossRef]
- Guo, G.; Sun, X.; Chen, C.; Wu, S.; Huang, P.; Li, Z.; Dean, M.; Huang, Y.; Jia, W.; Zhou, Q.; et al. Whole-genome and whole-exome sequencing of bladder cancer identifies frequent alterations in genes involved in sister chromatid cohesion and segregation. Nat. Genet. 2013, 45, 1459–1463. [Google Scholar] [CrossRef]
- Bao, Z.S.; Chen, H.M.; Yang, M.Y.; Zhang, C.B.; Yu, K.; Ye, W.L.; Hu, B.Q.; Yan, W.; Zhang, W.; Akers, J.; et al. RNA-seq of 272 gliomas revealed a novel, recurrent PTPRZ1-MET fusion transcript in secondary glioblastomas. Genome Res. 2014, 24, 1765–1773. [Google Scholar] [CrossRef]
- Wang, R.; Wang, L.; Li, Y.; Hu, H.; Shen, L.; Shen, X.; Pan, Y.; Ye, T.; Zhang, Y.; Luo, X.; et al. FGFR1/3 tyrosine kinase fusions define a unique molecular subtype of non-small cell lung cancer. Clin. Cancer Res. 2014, 20, 4107–4114. [Google Scholar] [CrossRef]
- Kim, Y.; Hammerman, P.S.; Kim, J.; Yoon, J.A.; Lee, Y.; Sun, J.M.; Wilkerson, M.D.; Pedamallu, C.S.; Cibulskis, K.; Yoo, Y.K.; et al. Integrative and comparative genomic analysis of lung squamous cell carcinomas in East Asian patients. J. Clin. Oncol. 2014, 32, 121–128. [Google Scholar] [CrossRef]
- Javle, M.; Rashid, A.; Churi, C.; Kar, S.; Zuo, M.; Eterovic, A.K.; Nogueras-Gonzalez, G.M.; Janku, F.; Shroff, R.T.; Aloia, T.A.; et al. Molecular characterization of gallbladder cancer using somatic mutation profiling. Hum. Pathol. 2014, 45, 701–708. [Google Scholar] [CrossRef]
- Yuan, L.; Liu, Z.H.; Lin, Z.R.; Xu, L.H.; Zhong, Q.; Zeng, M.S. Recurrent FGFR3-TACC3 fusion gene in nasopharyngeal carcinoma. Cancer Biol. Ther. 2014, 15, 1613–1621. [Google Scholar] [CrossRef]
- Carneiro, B.A.; Elvin, J.A.; Kamath, S.D.; Ali, S.M.; Paintal, A.S.; Restrepo, A.; Berry, E.; Giles, F.J.; Johnson, M.L. FGFR3-TACC3: A novel gene fusion in cervical cancer. Gynecol. Oncol. Rep. 2015, 13, 53–56. [Google Scholar] [CrossRef]
- Lawrence, M.S.; Sougnez, C.; Lichtenstein, L.; Cibulskis, K.; Lander, E.; Gabriel, S.B.; Getz, G.; Ally, A.; Balasundaram, M.; Birol, I.; et al. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 2015, 517, 576–582. [Google Scholar] [CrossRef]
- Majewski, I.J.; Mittempergher, L.; Davidson, N.M.; Bosma, A.; Willems, S.M.; Horlings, H.M.; De Rink, I.; Greger, L.; Hooijer, G.K.J.; Peters, D.; et al. Identification of recurrent FGFR3 fusion genes in lung cancer through kinome-centred RNA sequencing. J. Pathol. 2013, 230, 270–276. [Google Scholar] [CrossRef]
- Mizukami, T.; Sakai, K.; Naruki, S.; Taniyama, T.; Horie, Y.; Izawa, N.; Tsuda, T.; Fujino, T.; Boku, N.; Yasuda, H.; et al. Identification of a FGFR3-TACC3 fusion in esophageal cancer. Ann. Oncol. 2017, 28, 437–438. [Google Scholar] [CrossRef]
- Capelletti, M.; Dodge, M.E.; Ercan, D.; Hammerman, P.S.; Park, S.-I.; Kim, J.; Sasaki, H.; Jablons, D.M.; Lipson, D.; Young, L.; et al. Identification of recurrent FGFR3-TACC3 fusion oncogenes from lung adenocarcinoma. Clin. Cancer Res. 2014, 20, 6551–6558. [Google Scholar] [CrossRef]
- Nelson, K.N.; Meyer, A.N.; Siari, A.; Campos, A.R.; Motamedchaboki, K.; Donoghue, D.J. Oncogenic gene fusion FGFR3-TACC3 Is regulated by tyrosine phosphorylation. Mol. Cancer Res. 2016, 14, 458–469. [Google Scholar] [CrossRef]
- Nelson, K.N.; Meyer, A.N.; Wang, C.G.; Donoghue, D.J. Oncogenic driver FGFR3-TACC3 is dependent on membrane trafficking and ERK signaling. Oncotarget 2018, 9, 34306–34319. [Google Scholar] [CrossRef] [PubMed]
- Parker, B.C.; Annala, M.J.; Cogdell, D.E.; Granberg, K.J.; Sun, Y.; Ji, P.; Li, X.; Gumin, J.; Zheng, H.; Hu, L.; et al. The tumorigenic FGFR3-TACC3 gene fusion escapes miR-99a regulation in glioblastoma. J. Clin. Invest. 2013, 123, 855–865. [Google Scholar] [CrossRef] [PubMed]
- Sievers, P.; Stichel, D.; Schrimpf, D.; Sahm, F.; Koelsche, C.; Reuss, D.E.; Wefers, A.K.; Reinhardt, A.; Huang, K.; Ebrahimi, A.; et al. FGFR1:TACC1 fusion is a frequent event in molecularly defined extraventricular neurocytoma. Acta Neuropathol. 2018, 136, 293–302. [Google Scholar] [CrossRef] [PubMed]
- Bale, T.A. FGFR- gene family alterations in low-grade neuroepithelial tumors. Acta Neuropathol. Commun. 2020, 8, 21. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Wu, G.; Miller, C.P.; Tatevossian, R.G.; Dalton, J.D.; Tang, B.; Orisme, W.; Punchihewa, C.; Parker, M.; Qaddoumi, I.; et al. Whole-genome sequencing identifies genetic alterations in pediatric low-grade gliomas. Nat. Genet. 2013, 45, 602–612. [Google Scholar] [CrossRef] [PubMed]
- Shi, E.; Chmielecki, J.; Tang, C.M.; Wang, K.; Heinrich, M.C.; Kang, G.; Corless, C.L.; Hong, D.; Fero, K.E.; Murphy, J.D.; et al. FGFR1 and NTRK3 actionable alterations in “Wild-Type” gastrointestinal stromal tumors. J. Transl. Med. 2016, 14, 1–11. [Google Scholar] [CrossRef]
- Lucas, C.H.G.; Gupta, R.; Doo, P.; Lee, J.C.; Cadwell, C.R.; Ramani, B.; Hofmann, J.W.; Sloan, E.A.; Kleinschmidt-Demasters, B.K.; Lee, H.S.; et al. Comprehensive analysis of diverse low-grade neuroepithelial tumors with FGFR1 alterations reveals a distinct molecular signature of rosette-forming glioneuronal tumor. Acta Neuropathol. Commun. 2020, 8, 1–17. [Google Scholar] [CrossRef]
- Sievers, P.; Schrimpf, D.; Stichel, D.; Reuss, D.E.; Hasselblatt, M.; Hagel, C.; Staszewski, O.; Hench, J.; Frank, S.; Brandner, S.; et al. Posterior fossa pilocytic astrocytomas with oligodendroglial features show frequent FGFR1 activation via fusion or mutation. Acta Neuropathol. 2020, 139, 403–406. [Google Scholar] [CrossRef]
- Daoud, E.V.; Patel, A.; Gagan, J.; Raisanen, J.M.; Snipes, G.J.; Mantilla, E.; Krothapally, R.; Hatanpaa, K.J.; Pan, E. Spinal Cord Pilocytic Astrocytoma With FGFR1-TACC1 Fusion and Anaplastic Transformation. J. Neuropathol. Exp. Neurol. 2021, 80, 283–285. [Google Scholar] [CrossRef]
- Devereaux, K.A.; Weiel, J.J.; Mills, A.M.; Kunder, C.A.; Longacre, T.A. Neurofibrosarcoma Revisited: An Institutional Case Series of Uterine Sarcomas Harboring Kinase-related Fusions With Report of a Novel FGFR1-TACC1 Fusion. Am. J. Surg. Pathol. 2021, 45, 638–652. [Google Scholar] [CrossRef]
- Borad, M.J.; Champion, M.D.; Egan, J.B.; Liang, W.S.; Fonseca, R.; Bryce, A.H.; McCullough, A.E.; Barrett, M.T.; Hunt, K.; Patel, M.D.; et al. Integrated Genomic Characterization Reveals Novel, Therapeutically Relevant Drug Targets in FGFR and EGFR Pathways in Sporadic Intrahepatic Cholangiocarcinoma. PLoS Genet. 2014, 10, e1004135. [Google Scholar] [CrossRef]
- Ying, X.; Tu, J.; Wang, W.; Li, X.; Xu, C.; Ji, J. FGFR2-BICC1: A subtype of FGFR2 oncogenic fusion variant in cholangiocarcinoma and the response to sorafenib. Onco. Targets. Ther. 2019, 12, 9303–9307. [Google Scholar] [CrossRef]
- Ross, J.S.; Wang, K.; Gay, L.; Al-Rohil, R.; Rand, J.V.; Jones, D.M.; Lee, H.J.; Sheehan, C.E.; Otto, G.A.; Palmer, G.; et al. New Routes to Targeted Therapy of Intrahepatic Cholangiocarcinomas Revealed by Next-Generation Sequencing. Oncologist 2014, 19, 235–242. [Google Scholar] [CrossRef]
- Mazzaferro, V.; El-Rayes, B.F.; Droz dit Busset, M.; Cotsoglou, C.; Harris, W.P.; Damjanov, N.; Masi, G.; Rimassa, L.; Personeni, N.; Braiteh, F.; et al. Derazantinib (ARQ 087) in advanced or inoperable FGFR2 gene fusion-positive intrahepatic cholangiocarcinoma. Br. J. Cancer 2019, 120, 165–171. [Google Scholar] [CrossRef]
- Scheiter, A.; Keil, F.; Lüke, F.; Grosse, J.; Verloh, N.; Opitz, S.; Schlosser, S.; Kandulski, A.; Pukrop, T.; Dietmaier, W.; et al. Identification and In-Depth Analysis of the Novel FGFR2-NDC80 Fusion in a Cholangiocarcinoma Patient: Implication for Therapy. Curr. Oncol. 2021, 28, 112. [Google Scholar] [CrossRef]
- Seo, J.S.; Ju, Y.S.; Lee, W.C.; Shin, J.Y.; Lee, J.K.; Bleazard, T.; Lee, J.; Jung, Y.J.; Kim, J.O.; Shin, J.Y.; et al. The transcriptional landscape and mutational profile of lung adenocarcinoma. Genome Res. 2012, 22, 2109–2119. [Google Scholar] [CrossRef]
- Qin, A.; Johnson, A.; Ross, J.S.; Miller, V.A.; Ali, S.M.; Schrock, A.B.; Gadgeel, S.M. Detection of Known and Novel FGFR Fusions in Non–Small Cell Lung Cancer by Comprehensive Genomic Profiling. J. Thorac. Oncol. 2019, 14, 54–62. [Google Scholar] [CrossRef]
- Tabernero, J.; Bahleda, R.; Dienstmann, R.; Infante, J.R.; Mita, A.; Italiano, A.; Calvo, E.; Moreno, V.; Adamo, B.; Gazzah, A.; et al. Phase I dose-escalation study of JNJ-42756493, an oral pan-fibroblast growth factor receptor inhibitor, in patients with advanced solid tumors. J. Clin. Oncol. 2015, 33, 3401–3408. [Google Scholar] [CrossRef]
- Popovici, C.; Zhang, B.; Grégoire, M.J.; Jonveaux, P.; Lafage-Pochitaloff, M.; Birnbaum, D.; Pébusque, M.J. The t(6;8)(q27;p11) translocation in a stem cell myeloproliferative disorder fuses a novel gene, FOP, to Fibroblast Growth Factor Receptor 1. Blood 1999, 93, 1381–1389. [Google Scholar] [CrossRef]
- Sohal, J.; Chase, A.; Mould, S.; Corcoran, M.; Oscier, D.; Iqbal, S.; Parker, S.; Welborn, J.; Harris, R.I.; Martinelli, G.; et al. Identification of four new translocations involvingFGFR1 in myeloid disorders. Genes, Chromosom. Cancer 2001, 32, 155–163. [Google Scholar] [CrossRef]
- Vannier, J.P.; Bizet, M.; Bastard, C.; Bernard, A.; Ducastelle, T.; Tron, P. Simultaneous occurrence of a T-cell lymphoma and a chronic myelogenous leukemia with an unusual karyotype. Leuk. Res. 1984, 8, 647–657. [Google Scholar] [CrossRef]
- Vizmanos, J.L.; Hernández, R.; Vidal, M.J.; Larráyoz, M.J.; Odero, M.D.; Marín, J.; Ardanaz, M.T.; Calasanz, M.J.; Cross, N.C.P. Clinical variability of patients with the t(6;8)(q27;p12) and FGFR1OP-FGFR1 fusion: Two further cases. Hematol. J. 2004, 5, 534–537. [Google Scholar] [CrossRef]
- Chaffanet, M.; Popovici, C.; Leroux, D.; Jacrot, M.; Adélaïde, J.; Dastugue, N.; Grégoire, M.J.; Hagemeijer, A.; Lafage-Pochitaloff, M.; Birnbaum, D.; et al. t(6;8), t(8;9) and t(8;13) translocations associated with stem cell myeloproliferative disorders have close or identical breakpoints in chromosome region 8p11-12. Oncogene 1998, 16, 945–949. [Google Scholar] [CrossRef][Green Version]
- Onozawa, M.; Ohmura, K.; Ibata, M.; Iwasaki, J.; Okada, K.; Kasahara, I.; Yamaguchi, K.; Kubota, K.; Fujisawa, S.; Shigematsu, A.; et al. The 8p11 myeloproliferative syndrome owing to rare FGFR1OP2-FGFR1 fusion. Eur. J. Haematol. 2011, 86, 347–349. [Google Scholar] [CrossRef]
- Macdonald, D.; Aguiar, R.C.; Mason, P.J.; Goldman, J.M.; Cross, N.C. A new myeloproliferative disorder associated with chromosomal translocations involving 8p11: A review. Leukemia 1995, 9, 1628–1630. [Google Scholar]
- Macdonald, D.; Reiter, A.; Cross, N.C.P. The 8p11 myeloproliferative syndrome: A distinct clinical entity caused by constitutive activation of FGFR1. Acta Haematol. 2002, 107, 101–107. [Google Scholar] [CrossRef]
- Friedhoff, F.; Rajendra, B.; Moody, R.; Alapatt, T. Novel reciprocal translocation between chromosomes 8 and 9 found in a patient with myeloproliferative disorder. Cancer Genet. Cytogenet. 1983, 9, 391–394. [Google Scholar] [CrossRef]
- Yamamoto, K.; Kawano, H.; Nishikawa, S.; Yakushijin, K.; Okamura, A.; Matsui, T. A biphenotypic transformation of 8p11 myeloproliferative syndrome with CEP1/FGFR1 fusion gene. Eur. J. Haematol. 2006, 77, 349–354. [Google Scholar] [CrossRef] [PubMed]
- Park, T.S.; Song, J.; Kim, J.S.; Yang, W.I.; Song, S.; Kim, S.J.; Suh, B.; Choi, J.R. 8p11 myeloproliferative syndrome preceded by t(8;9)(p11;q33), CEP110/FGFR1 fusion transcript: Morphologic, molecular, and cytogenetic characterization of myeloid neoplasms associated with eosinophilia and FGFR1 abnormality. Cancer Genet. Cytogenet. 2008, 181, 93–99. [Google Scholar] [CrossRef]
- Mozziconacci, M.J.; Carbuccia, N.; Prebet, T.; Charbonnier, A.; Murati, A.; Vey, N.; Chaffanet, M.; Birnbaum, D. Common features of myeloproliferative disorders with t(8;9)(p12;q33) and CEP110-FGFR1 fusion: Report of a new case and review of the literature. Leuk. Res. 2008, 32, 1304–1308. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Fu, W.; Yuan, Z.; Hou, J. Complete molecular remission after interferon alpha treatment in a case of 8p11 myeloproliferative syndrome. Leuk. Res. 2010, 34, e306–e307. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; He, Y.; Zhu, X.; Li, J.; He, H. Myeloproliferative disorders with t(8;9)(p12;q33): A case report and review of the literature. Pediatr. Hematol. Oncol. 2011, 28, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, S.; Ebihara, Y.; Mochizuki, S.; Kawakita, T.; Kato, S.; Ooi, J.; Takahashi, S.; Tojo, A.; Yusa, N.; Furukawa, Y.; et al. Quantitative polymerase chain reaction detection of CEP110-FGFR1 fusion gene in a patient with 8p11 myeloproliferative syndrome. Leuk. Lymphoma 2013, 54, 2068–2069. [Google Scholar] [CrossRef] [PubMed]
- Sarah, O.-O.; Anthony, A.; Titilope, A.; Alani, S. The 8p12 myeloproliferative syndrome. Niger. Med. J. 2014, 55, 176. [Google Scholar] [CrossRef]
- Wehrli, M.; Oppliger Leibundgut, E.; Gattiker, H.H.; Manz, M.G.; Müller, A.M.S.; Goede, J.S. Response to Tyrosine Kinase Inhibitors in Myeloproliferative Neoplasia with 8p11 Translocation and CEP110-FGFR1 Rearrangement. Oncologist 2017, 22, 480–483. [Google Scholar] [CrossRef]
- Sarthy, J.F.; Reddivalla, N.; Radhi, M.; Chastain, K. Pediatric 8p11 eosinophilic myeloproliferative syndrome (EMS): A case report and review of the literature. Pediatr. Blood Cancer 2017, 64, 1–8. [Google Scholar] [CrossRef]
- Chen, M.; Wang, K.; Cai, X.; Zhang, X.; Chao, H.; Chen, S.; Shen, H.; Wang, Q.; Zhang, R. Myeloid/lymphoid neoplasm with CEP110-FGFR1 fusion: An analysis of 16 cases show common features and poor prognosis. Hematology 2021, 26, 153–159. [Google Scholar] [CrossRef]
- Oscier, D.G.; Mufti, G.J.; Gardiner, A.; Hamblin, T.J. Reciprocal translocation between chromosomes 8 and 9 in atypical chronic myeloid leukaemia. J. Med. Genet. 1985, 22, 398–401. [Google Scholar] [CrossRef][Green Version]
- Yamamoto, S.; Otsu, M.; Matsuzaka, E.; Konishi, C.; Takagi, H.; Hanada, S.; Mochizuki, S.; Nakauchi, H.; Imai, K.; Tsuji, K.; et al. Screening of drugs to treat 8p11 myeloproliferative syndrome using patient-derived induced pluripotent stem cells with fusion gene CEP110-FGFR1. PLoS ONE 2015, 10, e0120841. [Google Scholar] [CrossRef]
- Brown, L.M.; Bartolo, R.C.; Davidson, N.M.; Schmidt, B.; Brooks, I.; Challis, J.; Petrovic, V.; Khuong-Quang, D.A.; Mechinaud, F.; Khaw, S.L.; et al. Targeted therapy and disease monitoring in CNTRL-FGFR1-driven leukaemia. Pediatr. Blood Cancer 2019, 66, 1–5. [Google Scholar] [CrossRef]
- Lewis, J.P.; Welborn, J.L.; Meyers, F.J.; Levy, N.B.; Roschak, T. Mast cell disease followed by leukemia with clonal evolution. Leuk. Res. 1987, 11, 769–773. [Google Scholar] [CrossRef]
- Jotterand Bellomo, M.; Mühlematter, D.; Wicht, M.; Delacrétaz, F.; Schmidt, P.M. t(8;9)(p11;q32) in atypical chronic myeloid leukaemia: A new cytogenetic-clinicopathologic association? Br. J. Haematol. 1992, 81, 307–308. [Google Scholar] [CrossRef]
- Van den Berg, H.; Kroes, W.; van der Schoot, C.E.; Dee, R.; Pals, S.T.; Bouts, T.H.; Slater, R.M. A young child with acquired t(8;9)(p11;q34): Additional proof that 8p11 is involved in mixed myeloid/T lymphoid malignancies. Leukemia 1996, 10, 1252–1253. [Google Scholar]
- Nakayama, H.; Inamitsu, T.; Ohga, S.; Kai, T.; Suda, M.; Matsuzaki, A.; Ueda, K. Chronic myelomonocytic leukaemia with t(8;9)(p11;q34) in childhood: An example of the 8p11 myeloproliferative disorder? Br. J. Haematol. 1996, 92, 692–695. [Google Scholar] [CrossRef]
- Vandergoten, P.; Janssen, M.; Madoe, V.; Vanstraelen, D. A myeloproliferative disorder with eosinophilia, a translocation t(8; 9)(p22;23) and a cerebellar degeneration in regression with interferon alpha therapy. Acta Haematol. 1998, 100, 8. [Google Scholar]
- Guasch, G.; Mack, G.J.; Popovici, C.; Dastugue, N.; Birnbaum, D.; Rattner, J.B.; Pébusque, M.J. FGFR1 is fused to the centrosome-associated protein CEP110 in the 8p12 stem cell myeloproliferative disorder with t(8;9)(p12;q33). Blood 2000, 95, 1788–1796. [Google Scholar] [CrossRef]
- Heiss, S.; Erdel, M.; Gunsilius, E.; Nachbaur, D.; Tzankov, A. Myelodysplastic/myeloproliferative disease with erythropoietic hyperplasia (erythroid preleukemia) and the unique translocation (8;9)(p23;p24): First description of a case. Hum. Pathol. 2005, 36, 1148–1151. [Google Scholar] [CrossRef]
- Chen, J.; Williams, I.R.; Lee, B.H.; Duclos, N.; Huntly, B.J.P.; Donoghue, D.J.; Gilliland, D.G. Constitutively activated FGFR3 mutants signal through PLCγ-dependent and -independent pathways for hematopoietic transformation. Blood 2005, 106, 328–337. [Google Scholar] [CrossRef]
- Yamaguchi, T.; Kakefuda, R.; Tajima, N.; Sowa, Y.; Sakai, T. Antitumor activities of JTP-74057 (GSK1120212), a novel MEK1/2 inhibitor, on colorectal cancer cell lines in vitro and in vivo. Int. J. Oncol. 2011, 39, 23–31. [Google Scholar] [CrossRef]
- Guagnano, V.; Furet, P.; Spanka, C.; Bordas, V.; Le Douget, M.; Stamm, C.; Brueggen, J.; Jensen, M.R.; Schnell, C.; Schmid, H.; et al. Discovery of 3-(2,6-Dichloro-3,5-dimethoxy-phenyl)-1-{6-[4-(4-ethyl- piperazin-1-yl)-phenylamino]-pyrimidin-4-yl}-1-methyl-urea (NVP-BGJ398), A potent and selective inhibitor of the fibroblast growth factor receptor family of receptor tyrosine kinase. J. Med. Chem. 2011, 54, 7066–7083. [Google Scholar] [CrossRef]
- Liu, P.C.C.; Koblish, H.; Wu, L.; Bowman, K.; Diamond, S.; DiMatteo, D.; Zhang, Y.; Hansbury, M.; Rupar, M.; Wen, X.; et al. INCB054828 (pemigatinib), a potent and selective inhibitor of fibroblast growth factor receptors 1, 2, and 3, displays activity against genetically defined tumor models. PLoS ONE 2020, 15, e231877. [Google Scholar] [CrossRef]
- Hood, F.E.; Williams, S.J.; Burgess, S.G.; Richards, M.W.; Roth, D.; Straube, A.; Pfuhl, M.; Bayliss, R.; Royle, S.J. Coordination of adjacent domains mediates TACC3-ch-TOG-clathrin assembly and mitotic spindle binding. J. Cell Biol. 2013, 202, 463–478. [Google Scholar] [CrossRef]
- Hood, F.E.; Royle, S.J. Pulling it together: The mitotic function of TACC3. Bioarchitecture 2011, 1, 105–109. [Google Scholar] [CrossRef]
- Nixon, F.M.; Gutiérrez-Caballero, C.; Hood, F.E.; Booth, D.G.; Prior, I.A.; Royle, S.J. The mesh is a network of microtubule connectors that stabilizes individual kinetochore fibers of the mitotic spindle. Elife 2015, 4, 1–21. [Google Scholar] [CrossRef]
- Sarkar, S.; Ryan, E.L.; Royle, S.J. FGFR3-TACC3 cancer gene fusions cause mitotic defects by removal of endogenous TACC3 from the mitotic spindle. Open Biol. 2017, 7. [Google Scholar] [CrossRef]
- Yao, R.; Natsume, Y.; Saiki, Y.; Shioya, H.; Takeuchi, K.; Yamori, T.; Toki, H.; Aoki, I.; Saga, T.; Noda, T. Disruption of Tacc3 function leads to in vivo tumor regression. Oncogene 2012, 31, 135–148. [Google Scholar] [CrossRef][Green Version]
- Yao, R.; Oyanagi, J.; Natsume, Y.; Kusama, D.; Kato, Y.; Nagayama, S.; Noda, T. Suppression of intestinal tumors by targeting the mitotic spindle of intestinal stem cells. Oncogene 2016, 35, 6109–6119. [Google Scholar] [CrossRef]
- Akbulut, O.; Lengerli, D.; Saatci, O.; Duman, E.; Seker, U.O.S.; Isik, A.; Akyol, A.; Caliskan, B.; Banoglu, E.; Sahin, O. A highly potent TACC3 inhibitor as a novel anticancer drug candidate. Mol. Cancer Ther. 2020, 19, 1243–1254. [Google Scholar] [CrossRef]
- Kinoshita, K.; Noetzel, T.L.; Pelletier, L.; Mechtler, K.; Drechsel, D.N.; Schwager, A.; Lee, M.; Raff, J.W.; Hyman, A.A. Aurora A phosphorylation of TACC3/maskin is required for centrosome-dependent microtubule assembly in mitosis. J. Cell Biol. 2005, 170, 1047–1055. [Google Scholar] [CrossRef]
- Adams, M.; Simms, R.J.; Abdelhamed, Z.; Dawe, H.R.; Szymanska, K.; Logan, C.V.; Wheway, G.; Pitt, E.; Gull, K.; Knowles, M.A.; et al. A meckelin-filamin a interaction mediates ciliogenesis. Hum. Mol. Genet. 2012, 21, 1272–1286. [Google Scholar] [CrossRef]
- Qie, Y.; Wang, L.; Du, E.; Chen, S.; Lu, C.; Ding, N.; Yang, K.; Xu, Y. TACC3 promotes prostate cancer cell proliferation and restrains primary cilium formation. Exp. Cell Res. 2020, 390. [Google Scholar] [CrossRef] [PubMed]
- Still, I.H.; Hamilton, M.; Vince, P.; Wolfman, A.; Cowell, J.K. Cloning of TACC1, an embryonically expressed, potentially transforming coiled coil containing gene, from the 8p11 breast cancer amplicon. Oncogene 1999, 18, 4032–4038. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Peset, I.; Vernos, I. The TACC proteins: TACC-ling microtubule dynamics and centrosome function. Trends Cell Biol. 2008, 18, 379–388. [Google Scholar] [CrossRef] [PubMed]
- Gergely, F.; Karlsson, C.; Still, I.; Cowell, J.; Kilmartin, J.; Raff, J.W. The TACC domain identifies a family of centrosomal proteins that can interact with microtubules. Proc. Natl. Acad. Sci. USA 2000, 97, 14352–14357. [Google Scholar] [CrossRef]
- Conte, N.; Delaval, B.; Ginestier, C.; Ferrand, A.; Isnardon, D.; Larroque, C.; Prigent, C.; Séraphin, B.; Jacquemier, J.; Birnbaum, D. TACC1-chTOG-Aurora A protein complex in breast cancer. Oncogene 2003, 22, 8102–8116. [Google Scholar] [CrossRef]
- Cristinziano, G.; Porru, M.; Lamberti, D.; Buglioni, S.; Rollo, F.; Amoreo, C.A.; Manni, I.; Giannarelli, D.; Cristofoletti, C.; Russo, G.; et al. FGFR2 fusion proteins drive oncogenic transformation of mouse liver organoids towards cholangiocarcinoma. J. Hepatol. 2021, 11, 3–5. [Google Scholar] [CrossRef]
- Parker, B.C.; Engels, M.; Annala, M.; Zhang, W. Emergence of FGFR family gene fusions as therapeutic targets in a wide spectrum of solid tumours. J. Pathol. 2014, 232, 4–15. [Google Scholar] [CrossRef]
- Bahleda, R.; Meric-Bernstam, F.; Goyal, L.; Tran, B.; He, Y.; Yamamiya, I.; Benhadji, K.A.; Matos, I.; Arkenau, H.-T. Phase I, first-in-human study of futibatinib, a highly selective, irreversible FGFR1–4 inhibitor in patients with advanced solid tumors. Ann. Oncol. 2020, 31, 1405–1412. [Google Scholar] [CrossRef]
- Goyal, L.; Saha, S.K.; Liu, L.Y.; Siravegna, G.; Leshchiner, I.; Ahronian, L.G.; Lennerz, J.K.; Vu, P.; Deshpande, V.; Kambadakone, A.; et al. Polyclonal Secondary FGFR2 Mutations Drive Acquired Resistance to FGFR Inhibition in Patients with FGFR2 Fusion–Positive Cholangiocarcinoma. Cancer Discov. 2017, 7, 252–263. [Google Scholar] [CrossRef]
- Mahone, M.; Saffman, E.E.; Lasko, P.F. Localized Bicaudal-C RNA encodes a protein containing a KH domain, the RNA binding motif of FMR1. EMBO J. 1995, 14, 2043–2055. [Google Scholar] [CrossRef]
- Park, S.; Blaser, S.; Marchal, M.A.; Houston, D.W.; Sheets, M.D. A gradient of maternal Bicaudal-C controls vertebrate embryogenesis via translational repression of mRNAs encoding cell fate regulators. Development 2016, 143, 864–871. [Google Scholar] [CrossRef]
- Wessely, O.; De Robertis, E.M. The Xenopus homologue of Bicaudal-C is a localized maternal mRNA that can induce endoderm formation. Development 2000, 127, 2053–2062. [Google Scholar] [CrossRef]
- Dowdle, M.E.; Park, S.; Blaser Imboden, S.; Fox, C.A.; Houston, D.W.; Sheets, M.D. A single KH domain in Bicaudal-C links mRNA binding and translational repression functions to maternal development. Development 2019, 146. [Google Scholar] [CrossRef]
- Saffman, E.E.; Styhler, S.; Rother, K.; Li, W.; Richard, S.; Lasko, P. Premature translation of oskar in oocytes lacking the RNA-binding protein bicaudal-C. Mol. Cell. Biol. 1998, 18, 4855–4862. [Google Scholar] [CrossRef]
- Tran, U.; Zakin, L.; Schweickert, A.; Agrawal, R.; Döger, R.; Blum, M.; De Robertis, E.M.; Wessely, O. The RNA-binding protein bicaudal C regulates polycystin 2 in the kidney by antagonizing miR-17 activity. Development 2010, 137, 1107–1116. [Google Scholar] [CrossRef]
- Cogswell, C.; Price, S.J.; Hou, X.; Guay-Woodford, L.M.; Flaherty, L.; Bryda, E.C. Positional cloning of jcpk/bpk locus of the mouse. Mamm. Genome 2003, 14, 242–249. [Google Scholar] [CrossRef]
- Lemaire, L.A.; Goulley, J.; Kim, Y.H.; Carat, S.; Jacquemin, P.; Rougemont, J.; Constam, D.B.; Grapin-Botton, A. Bicaudal C1 promotes pancreatic NEUROG3+ endocrine progenitor differentiation and ductal morphogenesis. Development 2015, 142, 858–870. [Google Scholar] [CrossRef]
- Maisonneuve, C.; Guilleret, I.; Vick, P.; Weber, T.; Andre, P.; Beyer, T.; Blum, M.; Constam, D.B. Bicaudal C, a novel regulator of Dvl signaling abutting RNA-processing bodies, controls cilia orientation and leftward flow. Development 2009, 136, 3019–3030. [Google Scholar] [CrossRef]
- Guay-Woodford, L.M.; Bryda, E.C.; Christine, B.; Lindsey, J.R.; Collier, W.R.; Avner, E.D.; D’Eustachio, P.; Flaherty, L. Evidence that two phenotypically distinct mouse PKD mutations, bpk and jcpk, are allelic. Kidney Int. 1996, 50, 1158–1165. [Google Scholar] [CrossRef][Green Version]
- Flaherty, L.; Bryda, E.C.; Collins, D.; Rudofsky, U.; Montogomery, J.C. New mouse model for polycystic kidney disease with both recessive and dominant gene effects. Kidney Int. 1995, 47, 552–558. [Google Scholar] [CrossRef]
- Bouvrette, D.J.; Sittaramane, V.; Heidel, J.R.; Chandrasekhar, A.; Bryda, E.C. Knockdown of bicaudal C in zebrafish (Danio rerio) causes cystic kidneys: A nonmammalian model of polycystic kidney disease. Comp. Med. 2010, 60, 96–106. [Google Scholar]
- Gamberi, C.; Hipfner, D.R.; Trudel, M.; Lubell, W.D. Bicaudal C mutation causes myc and TOR pathway up-regulation and polycystic kidney disease-like phenotypes in Drosophila. PLoS Genet. 2017, 13, e1006694. [Google Scholar] [CrossRef] [PubMed]
- Iaconis, D.; Monti, M.; Renda, M.; Van Koppen, A.; Tammaro, R.; Chiaravalli, M.; Cozzolino, F.; Pignata, P.; Crina, C.; Pucci, P.; et al. The centrosomal OFD1 protein interacts with the translation machinery and regulates the synthesis of specific targets. Sci. Rep. 2017, 7, 1224. [Google Scholar] [CrossRef] [PubMed]
- Rothé, B.; Gagnieux, C.; Leal-Esteban, L.C.; Constam, D.B. Role of the RNA-binding protein Bicaudal-C1 and interacting factors in cystic kidney diseases. Cell. Signal. 2020, 68, 109499. [Google Scholar] [CrossRef]
- Kraus, M.R.C.; Clauin, S.; Pfister, Y.; Di Maïo, M.; Ulinski, T.; Constam, D.; Bellanné-Chantelot, C.; Grapin-Botton, A. Two mutations in human BICC1 resulting in wnt pathway hyperactivity associated with cystic renal dysplasia. Hum. Mutat. 2012, 33, 86–90. [Google Scholar] [CrossRef]
- Mesner, L.D.; Ray, B.; Hsu, Y.H.; Manichaikul, A.; Lum, E.; Bryda, E.C.; Rich, S.S.; Rosen, C.J.; Criqui, M.H.; Allison, M.; et al. Bicc1 is a genetic determinant of osteoblastogenesis and bone mineral density. J. Clin. Invest. 2014, 124, 2736–2749. [Google Scholar] [CrossRef]
- Rothé, B.; Leal-Esteban, L.; Bernet, F.; Urfer, S.; Doerr, N.; Weimbs, T.; Iwaszkiewicz, J.; Constam, D.B. Bicc1 Polymerization Regulates the Localization and Silencing of Bound mRNA. Mol. Cell. Biol. 2015, 35, 3339–3353. [Google Scholar] [CrossRef] [PubMed]
- Bouvrette, D.J.; Price, S.J.; Bryda, E.C. K homology domains of the mouse polycystic kidney disease-related protein, Bicaudal-C (Bicc1), mediate RNA binding in vitro. Nephron. Exp. Nephrol. 2008, 108, e27–e34. [Google Scholar] [CrossRef]
- Wimbish, R.T.; DeLuca, J.G. Hec1/Ndc80 Tail Domain Function at the Kinetochore-Microtubule Interface. Front. Cell Dev. Biol. 2020, 8, 1–16. [Google Scholar] [CrossRef]
- Qu, Y.; Li, J.; Cai, Q.; Liu, B. Hec1/Ndc80 is overexpressed in human gastric cancer and regulates cell growth. J. Gastroenterol. 2014, 49, 408–418. [Google Scholar] [CrossRef]
- Wang, G.; Jiang, Q.; Zhang, C. The role of mitotic kinases in coupling the centrosome cycle with the assembly of the mitotic spindle. J. Cell Sci. 2014, 127, 4111–4122. [Google Scholar] [CrossRef]
- Bièche, I.; Vacher, S.; Lallemand, F.; Tozlu-Kara, S.; Bennani, H.; Beuzelin, M.; Driouch, K.; Rouleau, E.; Lerebours, F.; Ripoche, H.; et al. Expression analysis of mitotic spindle checkpoint genes in breast carcinoma: Role of NDC80/HEC1 in early breast tumorigenicity, and a two-gene signature for aneuploidy. Mol. Cancer 2011, 10, 1–18. [Google Scholar] [CrossRef]
- Hu, C.M.; Zhu, J.; Guo, X.E.; Chen, W.; Qiu, X.L.; Ngo, B.; Chien, R.; Wang, Y.V.; Tsai, C.Y.; Wu, G.; et al. Novel small molecules disrupting Hec1/Nek2 interaction ablate tumor progression by triggering Nek2 degradation through a death-trap mechanism. Oncogene 2015, 34, 1220–1230. [Google Scholar] [CrossRef]
- Diaz-Rodríguez, E.; Sotillo, R.; Schvartzman, J.M.; Benezra, R. Hec1 overexpression hyperactivates the mitotic checkpoint and induces tumor formation in vivo. Proc. Natl. Acad. Sci. USA 2008, 105, 16719–16724. [Google Scholar] [CrossRef]
- Leber, B.; Maier, B.; Fuchs, F.; Chi, J.; Riffel, P.; Anderhub, S.; Wagner, L.; Ho, A.D.; Salisbury, J.L.; Boutros, M.; et al. Proteins required for centrosome clustering in cancer cells. Sci. Transl. Med. 2010, 2. [Google Scholar] [CrossRef]
- Wu, G.; Qiu, X.L.; Zhou, L.; Zhu, J.; Chamberlin, R.; Lau, J.; Chen, P.L.; Lee, W.H. Small molecule targeting the Hec1/Nek2 mitotic pathway suppresses tumor cell growth in culture and in animal. Cancer Res. 2008, 68, 8393–8399. [Google Scholar] [CrossRef]
- Huang, L.Y.L.; Chang, C.C.; Lee, Y.S.; Huang, J.J.; Chuang, S.H.; Chang, J.M.; Kao, K.J.; Lau, G.M.G.; Tsai, P.Y.; Liu, C.W.; et al. Inhibition of Hec1 as a novel approach for treatment of primary liver cancer. Cancer Chemother. Pharmacol. 2014, 74, 511–520. [Google Scholar] [CrossRef]
- Hall, T.G.; Yu, Y.; Eathiraj, S.; Wang, Y.; Savage, R.E.; Lapierre, J.-M.; Schwartz, B.; Abbadessa, G. Preclinical Activity of ARQ 087, a Novel Inhibitor Targeting FGFR Dysregulation. PLoS ONE 2016, 11, e0162594. [Google Scholar] [CrossRef]
- Gai, M.; Bianchi, F.T.; Vagnoni, C.; Vernì, F.; Bonaccorsi, S.; Pasquero, S.; Berto, G.E.; Sgrò, F.; Chiotto, A.A.; Annaratone, L.; et al. ASPM and CITK regulate spindle orientation by affecting the dynamics of astral microtubules. EMBO Rep. 2017, 18, 1870. [Google Scholar] [CrossRef]
- Watanabe, S.; De Zan, T.; Ishizaki, T.; Narumiya, S. Citron kinase mediates transition from constriction to abscission through its coiled-coil domain. J. Cell Sci. 2013, 126, 1773–1784. [Google Scholar] [CrossRef]
- Bassi, Z.I.; Audusseau, M.; Riparbelli, M.G.; Callaini, G.; D’Avino, P.P. Citron kinase controls a molecular network required for midbody formation in cytokinesis. Proc. Natl. Acad. Sci. USA 2013, 110, 9782–9787. [Google Scholar] [CrossRef] [PubMed]
- Gruneberg, U.; Neef, R.; Li, X.; Chan, E.H.Y.; Chalamalasetty, R.B.; Nigg, E.A.; Barr, F.A. KIF14 and citron kinase act together to promote efficient cytokinesis. J. Cell Biol. 2006, 172, 363–372. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Zhu, X.; Xu, W.; Zhang, Y.; Chen, L.; Qiu, F.; Zhang, B.; Wu, L.; Peng, Z.; Tang, H. Up-regulation of CIT promotes the growth of colon cancer cells. Oncotarget 2017, 8, 71954–71964. [Google Scholar] [CrossRef]
- Fu, Y.; Huang, J.; Wang, K.S.; Zhang, X.; Han, Z.G. RNA interference targeting CITRON can significantly inhibit the proliferation of hepatocellular carcinoma cells. Mol. Biol. Rep. 2011, 38, 693–702. [Google Scholar] [CrossRef] [PubMed]
- Ehrlichova, M.; Mohelnikova-Duchonova, B.; Hrdy, J.; Brynychova, V.; Mrhalova, M.; Kodet, R.; Rob, L.; Pluta, M.; Gut, I.; Soucek, P.; et al. The association of taxane resistance genes with the clinical course of ovarian carcinoma. Genomics 2013, 102, 96–101. [Google Scholar] [CrossRef] [PubMed]
- Meng, D.; Yu, Q.; Feng, L.; Luo, M.; Shao, S.; Huang, S.; Wang, G.; Jing, X.; Tong, Z.; Zhao, X.; et al. Citron kinase (CIT-K) promotes aggressiveness and tumorigenesis of breast cancer cells in vitro and in vivo: Preliminary study of the underlying mechanism. Clin. Transl. Oncol. 2019, 21, 910–923. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Yan, H.; Yang, Y.; Wei, L.; Xia, S.; Xiu, Y. Down-regulation of CIT can inhibit the growth of human bladder cancer cells. Biomed. Pharmacother. 2020, 124, 109830. [Google Scholar] [CrossRef]
- Liu, J.; Dou, J.; Wang, W.; Liu, H.; Qin, Y.; Yang, Q.; Jiang, W.; Liang, Y.; Liu, Y.; He, J.; et al. High expression of citron kinase predicts poor prognosis of prostate cancer. Oncol. Lett. 2020, 19, 1815–1823. [Google Scholar] [CrossRef]
- Pallavicini, G.; Iegiani, G.; Berto, G.E.; Calamia, E.; Trevisiol, E.; Veltri, A.; Allis, S.; Di Cunto, F. CITK loss inhibits growth of group 3 and group 4 medulloblastoma cells and sensitizes them to DNA-damaging agents. Cancers 2020, 12, 542. [Google Scholar] [CrossRef]
- Di Cunto, F.; Imarisio, S.; Hirsch, E.; Broccoli, V.; Bulfone, A.; Migheli, A.; Atzori, C.; Turco, E.; Triolo, R.; Dotto, G.P.; et al. Defective neurogenesis in citron kinase knockout mice by altered cytokinesis and massive apoptosis. Neuron 2000, 28, 115–127. [Google Scholar] [CrossRef]
- Di Cunto, F.; Imarisio, S.; Camera, P.; Boitani, C.; Altruda, F.; Silengo, L. Essential role of citron kinase in cytokinesis of spermatogenic precursors. J. Cell Sci. 2002, 115, 4819–4826. [Google Scholar] [CrossRef] [PubMed]
- Mick, D.U.; Rodrigues, R.B.; Leib, R.D.; Adams, C.M.; Chien, A.S.; Gygi, S.P.; Nachury, M. V Proteomics of Primary Cilia by Proximity Labeling. Dev. Cell 2015, 35, 497–512. [Google Scholar] [CrossRef] [PubMed]
- Kuhns, S.; Schmidt, K.N.; Reymann, J.; Gilbert, D.F.; Neuner, A.; Hub, B.; Carvalho, R.; Wiedemann, P.; Zentgraf, H.; Erfle, H.; et al. The microtubule affinity regulating kinase MARK4 promotes axoneme extension during early ciliogenesis. J. Cell Biol. 2013, 200, 505–522. [Google Scholar] [CrossRef] [PubMed]
- Anastas, S.B.; Mueller, D.; Semple-Rowland, S.L.; Breunig, J.J.; Sarkisian, M.R. Failed cytokinesis of neural progenitors in citron kinase-deficient rats leads to multiciliated neurons. Cereb. Cortex 2011, 21, 338–344. [Google Scholar] [CrossRef]
- Karkera, J.D.; Cardona, G.M.; Bell, K.; Gaffney, D.; Portale, J.C.; Santiago-Walker, A.; Moy, C.H.; King, P.; Sharp, M.; Bahleda, R.; et al. Oncogenic characterization and pharmacologic sensitivity of activating Fibroblast Growth Factor Receptor (FGFR) genetic alterations to the selective FGFR inhibitor erdafitinib. Mol. Cancer Ther. 2017, 16, 1717–1726. [Google Scholar] [CrossRef]
- Romio, L.; Fry, A.M.; Winyard, P.J.D.; Malcolm, S.; Woolf, A.S.; Feather, S.A. OFD1 is a centrosomal/basal body protein expressed during mesenchymal-epithelial transition in human nephrogenesis. J. Am. Soc. Nephrol. 2004, 15, 2556–2568. [Google Scholar] [CrossRef]
- Ferrante, M.I.; Zullo, A.; Barra, A.; Bimonte, S.; Messaddeq, N.; Studer, M.; Dollé, P.; Franco, B. Oral-facial-digital type I protein is required for primary cilia formation and left-right axis specification. Nat. Genet. 2006, 38, 112–117. [Google Scholar] [CrossRef]
- Singla, V.; Romaguera-Ros, M.; Garcia-Verdugo, J.M.; Reiter, J.F. Ofd1, a Human Disease Gene, Regulates the Length and Distal Structure of Centrioles. Dev. Cell 2010, 18, 410–424. [Google Scholar] [CrossRef]
- Lopes, C.A.M.; Prosser, S.L.; Romio, L.; Hirst, R.A.; O’Callaghan, C.; Woolf, A.S.; Fry, A.M. Centriolar satellites are assembly points for proteins implicated in human ciliopathies, including oral-facial-digital syndrome 1. J. Cell Sci. 2011, 124, 600–612. [Google Scholar] [CrossRef]
- Chetty-John, S.; Piwnica-Worms, K.; Bryant, J.; Bernardini, I.; Fischer, R.E.; Heller, T.; Gahl, W.A.; Gunay-Aygun, M. Fibrocystic disease of liver and pancreas; under-recognized features of the X-linked ciliopathy oral-facial-digital syndrome type 1 (OFD I). Am. J. Med. Genet. 2010, 152, 2640–2645. [Google Scholar] [CrossRef]
- Thauvin-Robinet, C.; Cossée, M.; Cormier-Daire, V.; Van Maldergem, L.; Toutain, A.; Alembik, Y.; Bieth, E.; Layet, V.; Parent, P.; David, A.; et al. Clinical, molecular, and genotype-phenotype correlation studies from 25 cases of oral-facial-digital syndrome type 1: A French and Belgian collaborative study. J. Med. Genet. 2006, 43, 54–61. [Google Scholar] [CrossRef]
- Saal, S.; Faivre, L.; Aral, B.; Gigot, N.; Toutain, A.; Van Maldergem, L.; Destree, A.; Maystadt, I.; Cosyns, J.-P.P.; Jouk, P.-S.S.; et al. Renal insufficiency, a frequent complication with age in oral-facial-digital syndrome type I. Clin. Genet. 2010, 77, 258–265. [Google Scholar] [CrossRef]
- Marina, M.; Franco, B. The molecular basis of oral-facial-digital syndrome, type 1. Am. J. Med. Genet. Part C Semin. Med. Genet. 2009, 151, 318–325. [Google Scholar] [CrossRef]
- Zullo, A.; Iaconis, D.; Barra, A.; Cantone, A.; Messaddeq, N.; Capasso, G.; Dollé, P.; Igarashi, P.; Franco, B. Kidney-specific inactivation of Ofd1 leads to renal cystic disease associated with upregulation of the mTOR pathway. Hum. Mol. Genet. 2010, 19, 2792–2803. [Google Scholar] [CrossRef]
- Bimonte, S.; De Angelis, A.; Quagliata, L.; Giusti, F.; Tammaro, R.; Dallai, R.; Ascenzi, M.-G.G.; Diez-Roux, G.; Franco, B. Ofd1 is required in limb bud patterning and endochondral bone development. Dev. Biol. 2011, 349, 179–191. [Google Scholar] [CrossRef]
- Ferrante, M.I.; Romio, L.; Castro, S.; Collins, J.E.; Goulding, D.A.; Stemple, D.L.; Woolf, A.S.; Wilson, S.W. Convergent extension movements and ciliary function are mediated by ofd1, a zebrafish orthologue of the human oral-facial-digital type 1 syndrome gene. Hum. Mol. Genet. 2009, 18, 289–303. [Google Scholar] [CrossRef]
- Lelièvre, H.; Chevrier, V.; Tassin, A.-M.; Birnbaum, D. Myeloproliferative disorder FOP-FGFR1 fusion kinase recruits phosphoinositide-3 kinase and phospholipase Cgamma at the centrosome. Mol. Cancer 2008, 7, 30. [Google Scholar] [CrossRef]
- Delaval, B.; Létard, S.; Lelièvre, H.; Chevrier, V.; Daviet, L.; Dubreuil, P.; Birnbaum, D. Oncogenic tyrosine kinase of malignant hemopathy targets the centrosome. Cancer Res. 2005, 65, 7231–7240. [Google Scholar] [CrossRef][Green Version]
- Guasch, G.; Ollendorff, V.; Borg, J.-P.; Birnbaum, D.; Pébusque, M.-J. 8p12 Stem Cell Myeloproliferative Disorder: The FOP-Fibroblast Growth Factor Receptor 1 Fusion Protein of the t(6;8) Translocation Induces Cell Survival Mediated by Mitogen-Activated Protein Kinase and Phosphatidylinositol 3-Kinase/Akt/mTOR Pathways. Mol. Cell. Biol. 2001, 21, 8129–8142. [Google Scholar] [CrossRef]
- Guasch, G.; Delaval, B.; Arnoulet, C.; Xie, M.J.; Xerri, L.; Sainty, D.; Birnbaum, D.; Pébusque, M.J. FOP-FGFR1 tyrosine kinase, the product of a t(6;8) translocation, induces a fatal myeloproliferative disease in mice. Blood 2004, 103, 309–312. [Google Scholar] [CrossRef][Green Version]
- Mikolajka, A.; Yan, X.; Popowicz, G.M.; Smialowski, P.; Nigg, E.A.; Holak, T.A. Structure of the N-terminal Domain of the FOP (FGFR1OP) Protein and Implications for its Dimerization and Centrosomal Localization. J. Mol. Biol. 2006, 359, 863–875. [Google Scholar] [CrossRef]
- Hori, A.; Toda, T. Regulation of centriolar satellite integrity and its physiology. Cell. Mol. Life Sci. 2016, 74, 213–229. [Google Scholar] [CrossRef]
- Odabasi, E.; Gul, S.; Kavakli, I.H.; Firat-Karalar, E.N. Centriolar satellites are required for efficient ciliogenesis and ciliary content regulation. EMBO Rep. 2019, 20, 1–20. [Google Scholar] [CrossRef]
- Tollenaere, M.A.X.; Mailand, N.; Bekker-Jensen, S. Centriolar satellites: Key mediators of centrosome functions. Cell. Mol. Life Sci. 2015, 72, 11–23. [Google Scholar] [CrossRef]
- Bärenz, F.; Mayilo, D.; Gruss, O.J. Centriolar satellites: Busy orbits around the centrosome. Eur. J. Cell Biol. 2011, 90, 983–989. [Google Scholar] [CrossRef]
- Yan, X.; Habedanck, R.; Nigg, E.A. A Complex of Two Centrosomal Proteins, CAP350 and FOP, Cooperates with EB1 in Microtubule Anchoring. Mol. Biol. Cell 2006, 17, 634–644. [Google Scholar] [CrossRef]
- Mohammadi, M.; McMahon, G.; Sun, L.; Tang, C.; Hirth, P.; Yeh, B.K.; Hubbard, S.R.; Schlessinger, J. Structures of the tyrosine kinase domain of fibroblast growth factor receptor in complex with inhibitors. Science 1997, 276, 955–960. [Google Scholar] [CrossRef]
- Mohammadi, M.; Honegger, A.M.; Rotin, D.; Fischer, R.; Bellot, F.; Li, W.; Dionne, C.A.; Jaye, M.; Rubinstein, M.; Schlessinger, J. A tyrosine-phosphorylated carboxy-terminal peptide of the fibroblast growth factor receptor (Flg) is a binding site for the SH2 domain of phospholipase C-gamma 1. Mol. Cell. Biol. 1991, 11, 5068–5078. [Google Scholar] [CrossRef]
- Lee, J.Y.; Hong, W.J.; Majeti, R.; Stearns, T. Centrosome-kinase fusions promote oncogenic signaling and disrupt centrosome function in myeloproliferative neoplasms. PLoS ONE 2014, 9, e92641. [Google Scholar] [CrossRef]
- Lee, J.Y.; Stearns, T. FOP Is a Centriolar Satellite Protein Involved in Ciliogenesis. PLoS ONE 2013, 8, e58589. [Google Scholar] [CrossRef]
- Mojarad, B.A.; Gupta, G.D.; Hasegan, M.; Goudiam, O.; Basto, R.; Gingras, A.C.; Pelletier, L. CEP19 cooperates with FOP and CEP350 to drive early steps in the ciliogenesis programme. Open Biol. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Cabaud, O.; Roubin, R.; Comte, A.; Bascunana, V.; Sergé, A.; Sedjaï, F.; Birnbaum, D.; Rosnet, O.; Acquaviva, C. Mutation of FOP/FGFR1OP in mice recapitulates human short rib-polydactyly ciliopathy. Hum. Mol. Genet. 2018, 27, 3377–3391. [Google Scholar] [CrossRef] [PubMed]
- Bangs, F.; Anderson, K.V. Primary Cilia and Mammalian Hedgehog Signaling. Cold Spring Harb. Perspect. Biol. 2017, 9, a028175. [Google Scholar] [CrossRef] [PubMed]
- Satir, P.; Christensen, S.T. Overview of Structure and Function of Mammalian Cilia. Annu. Rev. Physiol. 2007, 69, 377–400. [Google Scholar] [CrossRef]
- Giehl, M.; Fabarius, A.; Frank, O.; Hochhaus, A.; Hafner, M.; Hehlmann, R.; Seifarth, W. Centrosome aberrations in chronic myeloid leukemia correlate with stage of disease and chromosomal instability. Leukemia 2005, 19, 1192–1197. [Google Scholar] [CrossRef]
- Ren, M.; Qin, H.; Kitamura, E.; Cowell, J.K. Dysregulated signaling pathways in the development of CNTRL-FGFR1-induced myeloid and lymphoid malignancies associated with FGFR1 in human and mouse models. Blood 2013, 122, 1007–1016. [Google Scholar] [CrossRef][Green Version]
- Ren, M.; Qin, H.; Ren, R.; Tidwell, J.; Cowell, J.K. Src activation plays an important key role in lymphomagenesis induced by FGFR1 fusion kinases. Cancer Res. 2011, 71, 7312–7322. [Google Scholar] [CrossRef]
- Arber, D.A.; Orazi, A.; Hasserjian, R.; Thiele, J.; Borowitz, M.J.; Le Beau, M.M.; Bloomfield, C.D.; Cazzola, M.; Vardiman, J.W. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood 2016, 127, 2391–2405. [Google Scholar] [CrossRef]
- Ren, M.; Qin, H.; Ren, R.; Cowell, J.K. Ponatinib suppresses the development of myeloid and lymphoid malignancies associated with FGFR1 abnormalities. Leukemia 2013, 27, 32–40. [Google Scholar] [CrossRef]
- Ou, Y.Y.; Mack, G.J.; Zhang, M.; Rattner, J.B. CEP110 and ninein are located in a specific domain of the centrosome associated with centrosome maturation. J. Cell Sci. 2002, 115, 1825–1835. [Google Scholar] [CrossRef]
- Kashihara, H.; Chiba, S.; Kanno, S.-I.; Suzuki, K.; Yano, T.; Tsukita, S. Cep128 associates with Odf2 to form the subdistal appendage of the centriole. Genes Cells 2019, 24, 231–243. [Google Scholar] [CrossRef]
- Sun, T.-Y.; Wang, H.-Y.; Kwon, J.-W.; Yuan, B.; Lee, I.-W.; Cui, X.-S.; Kim, N.-H. Centriolin, a centriole-appendage protein, regulates peripheral spindle migration and asymmetric division in mouse meiotic oocytes. Cell Cycle 2017, 16, 1774–1780. [Google Scholar] [CrossRef]
- Chen, C.-T.; Hehnly, H.; Yu, Q.; Farkas, D.; Zheng, G.; Redick, S.D.; Hung, H.-F.; Samtani, R.; Jurczyk, A.; Akbarian, S.; et al. A unique set of centrosome proteins requires pericentrin for spindle-pole localization and spindle orientation. Curr. Biol. 2014, 24, 2327–2334. [Google Scholar] [CrossRef]
- Lassman, A.B.; Sepúlveda-Sánchez, J.M.; Cloughesy, T.; Gil-Gil, J.M.; Puduvalli, V.K.; Raizer, J.; De Vos, F.Y.; Wen, P.Y.; Butowski, N.; Clement, P.; et al. OS10.6 Infigratinib (BGJ398) in patients with recurrent gliomas with fibroblast growth factor receptor (FGFR) alterations: A multicenter phase II study. Neuro. Oncol. 2019, 21, iii21–iii22. [Google Scholar] [CrossRef]
- Goyal, L.; Shi, L.; Liu, L.Y.; de la Cruz, F.F.; Lennerz, J.K.; Raghavan, S.; Leschiner, I.; Elagina, L.; Siravegna, G.; Ng, R.W.S.; et al. TAS-120 overcomes resistance to atp-competitive FGFR inhibitors in patients with FGFR2 fusion–positive intrahepatic cholangiocarcinoma. Cancer Discov. 2019, 9, 1064–1079. [Google Scholar] [CrossRef]
- Krzyscik, M.A.; Zakrzewska, M.; Otlewski, J. Site-Specific, Stoichiometric-Controlled, PEGylated Conjugates of Fibroblast Growth Factor 2 (FGF2) with Hydrophilic Auristatin y for Highly Selective Killing of Cancer Cells Overproducing Fibroblast Growth Factor Receptor 1 (FGFR1). Mol. Pharm. 2020, 17, 2734–2748. [Google Scholar] [CrossRef]
- Porębska, N.; Latko, M.; Kucińska, M.; Zakrzewska, M.; Otlewski, J.; Opaliński, Ł. Targeting Cellular Trafficking of Fibroblast Growth Factor Receptors as a Strategy for Selective Cancer Treatment. J. Clin. Med. 2018, 8, 7. [Google Scholar] [CrossRef] [PubMed]
- Canning, P.; Park, K.; Gonçalves, J.; Li, C.; Howard, C.J.; Sharpe, T.D.; Holt, L.J.; Pelletier, L.; Bullock, A.N.; Leroux, M.R. CDKL Family Kinases Have Evolved Distinct Structural Features and Ciliary Function. Cell Rep. 2018, 22, 885–894. [Google Scholar] [CrossRef]
- Harrington, K.J.; Hingorani, M.; Tanay, M.A.; Hickey, J.; Bhide, S.A.; Clarke, P.M.; Renouf, L.C.; Thway, K.; Sibtain, A.; McNeish, I.A.; et al. Phase I/II study of oncolytic HSVGM-CSFin combination with radiotherapy and cisplatin in untreated stage III/IV squamous cell cancer of the head and neck. Clin. Cancer Res. 2010, 16, 4005–4015. [Google Scholar] [CrossRef]
- Heo, J.; Reid, T.; Ruo, L.; Breitbach, C.J.; Rose, S.; Bloomston, M.; Cho, M.; Lim, H.Y.; Chung, H.C.; Kim, C.W.; et al. Randomized dose-finding clinical trial of oncolytic immunotherapeutic vaccinia JX-594 in liver cancer. Nat. Med. 2013, 19, 329–336. [Google Scholar] [CrossRef]
- Freytag, S.O.; Stricker, H.; Lu, M.; Elshaikh, M.; Aref, I.; Pradhan, D.; Levin, K.; Kim, J.H.; Peabody, J.; Siddiqui, F.; et al. Prospective randomized phase 2 trial of intensity modulated radiation therapy with or without oncolytic adenovirus-mediated cytotoxic gene therapy in intermediate-risk prostate cancer. Int. J. Radiat. Oncol. Biol. Phys. 2014, 89, 268–276. [Google Scholar] [CrossRef] [PubMed]
- Ronca, R.; Giacomini, A.; Di Salle, E.; Coltrini, D.; Pagano, K.; Ragona, L.; Matarazzo, S.; Rezzola, S.; Maiolo, D.; Torrella, R.; et al. Long-Pentraxin 3 Derivative as a Small-Molecule FGF Trap for Cancer Therapy. Cancer Cell 2015, 28, 225–239. [Google Scholar] [CrossRef] [PubMed]
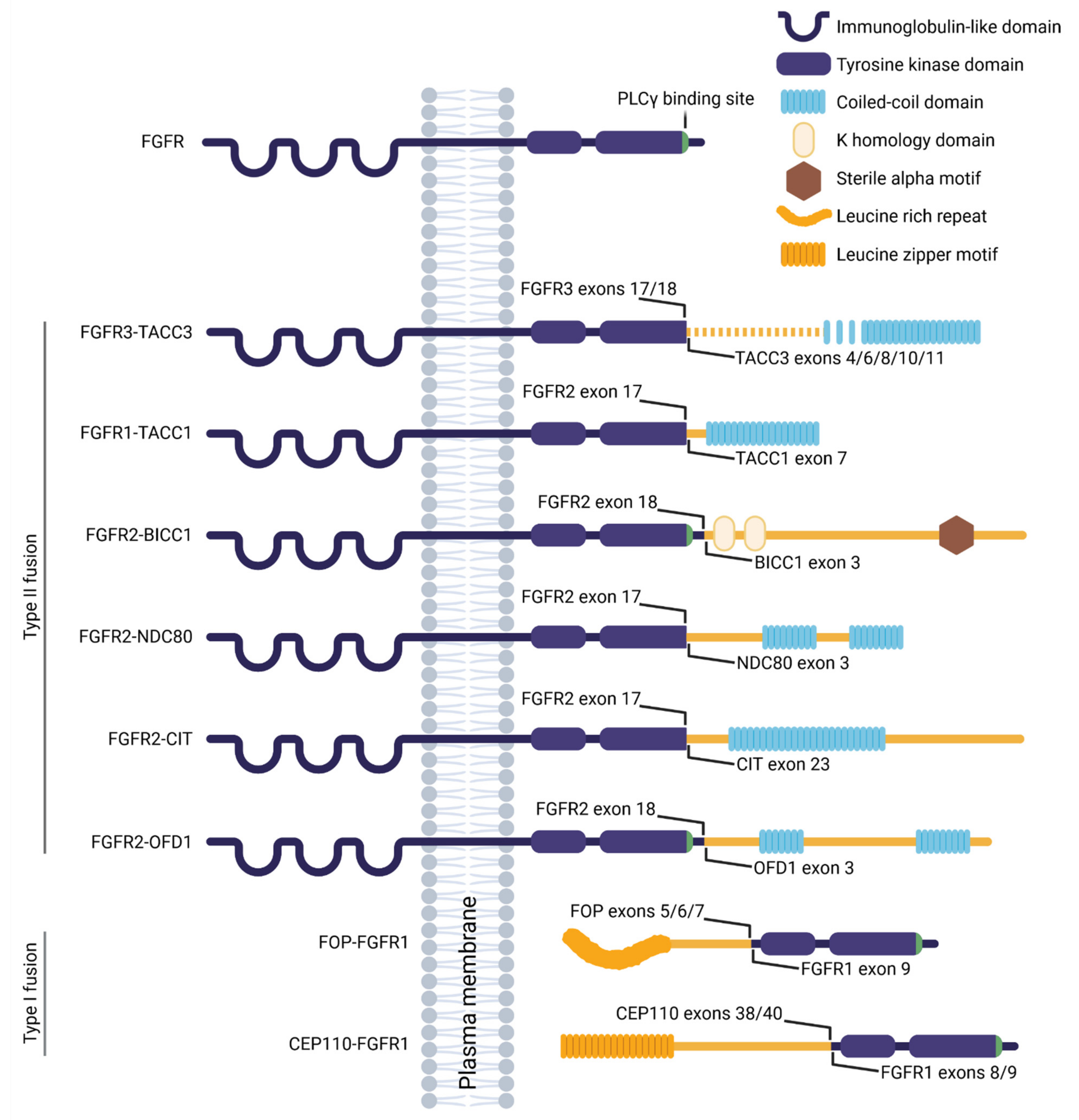
| Gene Fusion | Cancer Type (Cases) | Reference |
|---|---|---|
| FGFR3-TACC3 | glioblastoma (29); NSCLC (28); HNSCC (11); bladder cancer (10); urothelial carcinoma (4); nasopharyngeal carcinoma (4); LUSC (4); glioma (3); ESCC (2); cervical cancer (2); gallbladder cancer (2); oral cancer (1); renal cell cancer (1); endometrial adenocarcinoma (1) | [133,149,153,179,185,186,187,188,189,190,191,192,193,194,195,196,197,198,199,200] |
| FGFR1-TACC1 | low grade glioma (7); extraventricular neurocytoma (3); glioblastoma (2); spinal cord pilocytic astrocytoma (2); GIST (1) | [186,187,204,205,206,207,208,209,210,211] |
| FGFR2-BICC1 | cholangiocarcinoma (47); hepatocellular cancer (1); colorectal cancer (1) | [149,175,177,212,213,214,215] |
| FGFR2-NDC80 | cholangiocarcinoma (1) | [216] |
| FGFR2-CIT | NSCLC (3); cholangiocarcinoma (1) | [215,217,218] |
| FGFR2-OFD1 | thyroid cancer (1); endometrial cancer (1) | [149,219] |
| FOP-FGFR1 | AML (9); EMS (1) | [220,221,222,223,224,225,226,227] |
| CEP110-FGFR1 | EMS (9); MPD (5); aCML (3); AML (2); CMML (1); AMML (1) | [221,228,229,230,231,232,233,234,235,236,237,238,239,240,241,242,243,244,245,246,247,248] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nita, A.; Abraham, S.P.; Krejci, P.; Bosakova, M. Oncogenic FGFR Fusions Produce Centrosome and Cilia Defects by Ectopic Signaling. Cells 2021, 10, 1445. https://doi.org/10.3390/cells10061445
Nita A, Abraham SP, Krejci P, Bosakova M. Oncogenic FGFR Fusions Produce Centrosome and Cilia Defects by Ectopic Signaling. Cells. 2021; 10(6):1445. https://doi.org/10.3390/cells10061445
Chicago/Turabian StyleNita, Alexandru, Sara P. Abraham, Pavel Krejci, and Michaela Bosakova. 2021. "Oncogenic FGFR Fusions Produce Centrosome and Cilia Defects by Ectopic Signaling" Cells 10, no. 6: 1445. https://doi.org/10.3390/cells10061445
APA StyleNita, A., Abraham, S. P., Krejci, P., & Bosakova, M. (2021). Oncogenic FGFR Fusions Produce Centrosome and Cilia Defects by Ectopic Signaling. Cells, 10(6), 1445. https://doi.org/10.3390/cells10061445






