Chromothripsis—Explosion in Genetic Science
Abstract
1. Introduction
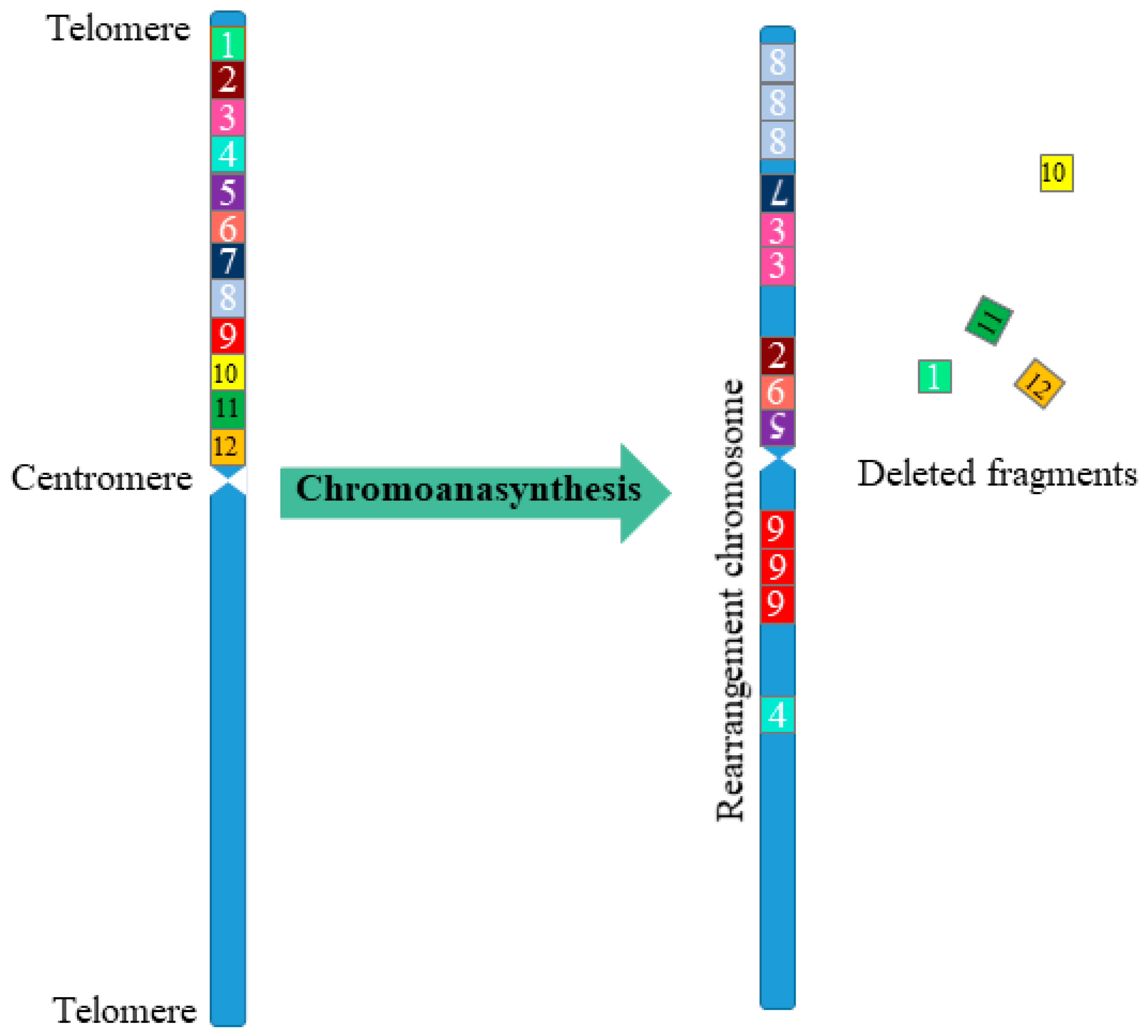
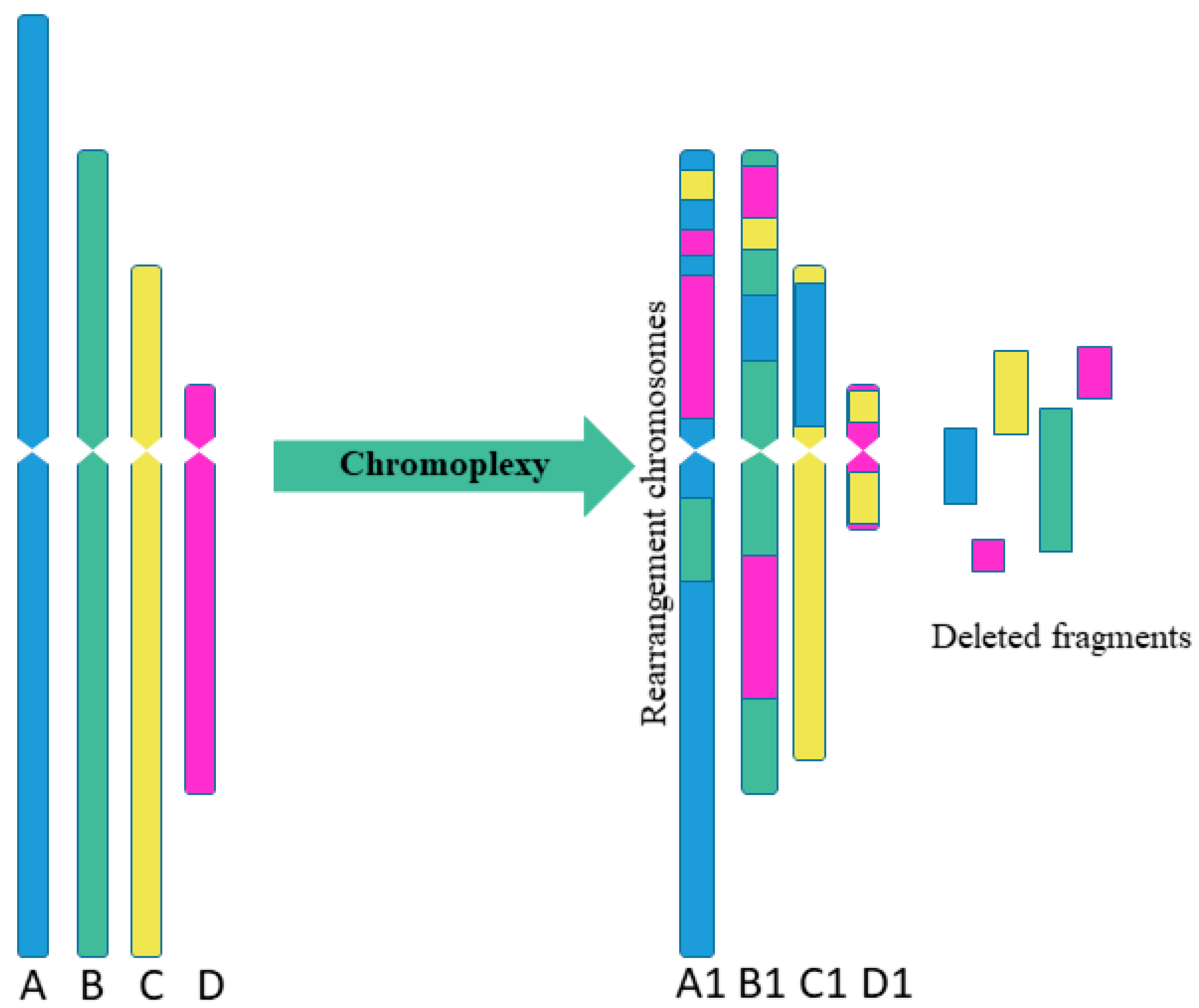
2. Types of Multiple Complex Chromosomal Rearrangements
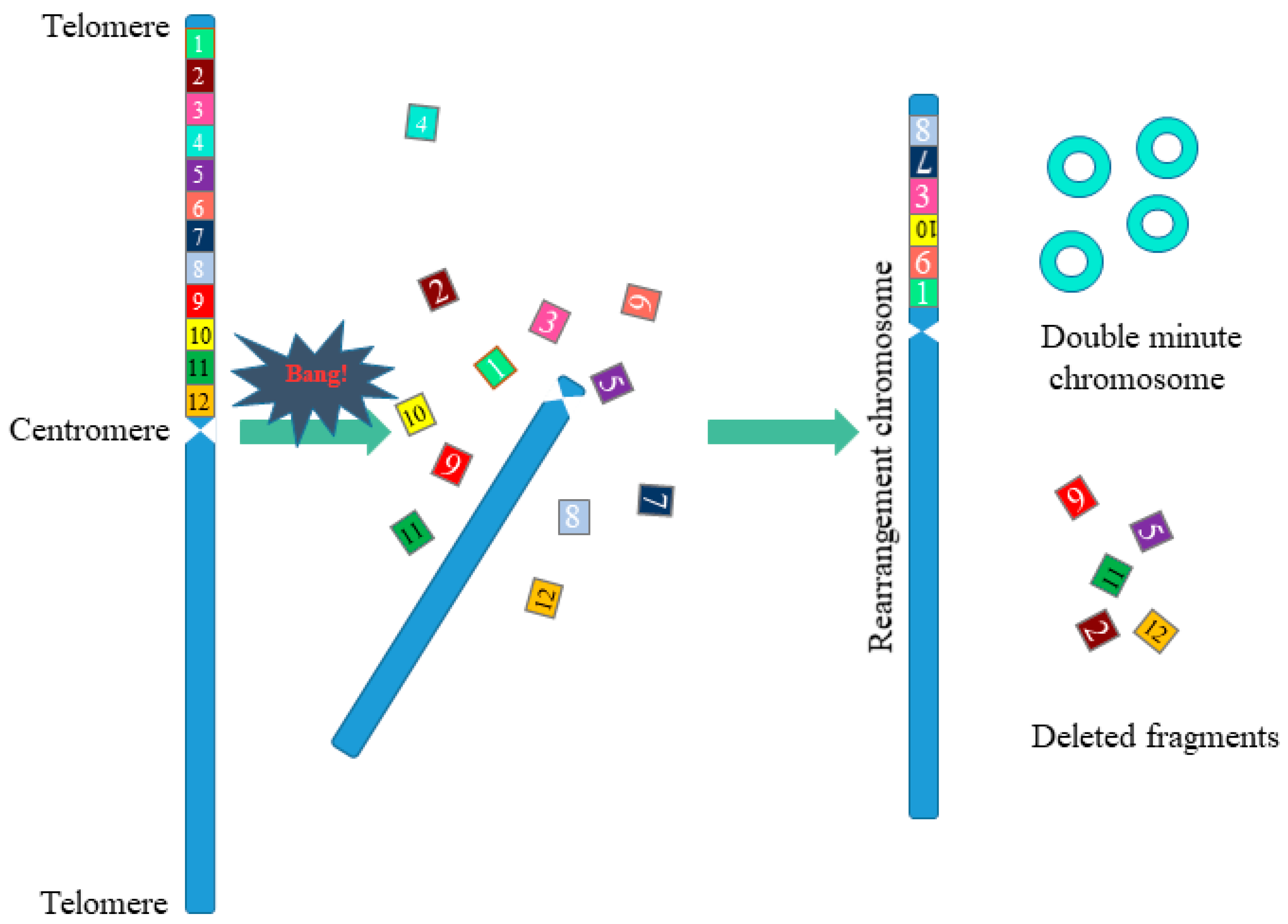
3. Chromothripsis
- The genomes of cells with chromothripsis are characterized by clustered DNA breaks. Such clustered regions have multiple DNA breaks in close proximity to each other. These clustered regions are surrounded by large regions of DNA not affected by rearrangements.
- As a result of chromothripsis, part of the chromosome fragments can be deleted.
- Chromothripsis usually occurs on one parental copy (haplotype) of chromosomes.
- Fragments of chromosomes obtained as a result of chromothripsis are combined in random order and have a random orientation.
- After chromothripsis in a new rearranged chromosome, each fragment is either retained or lost.
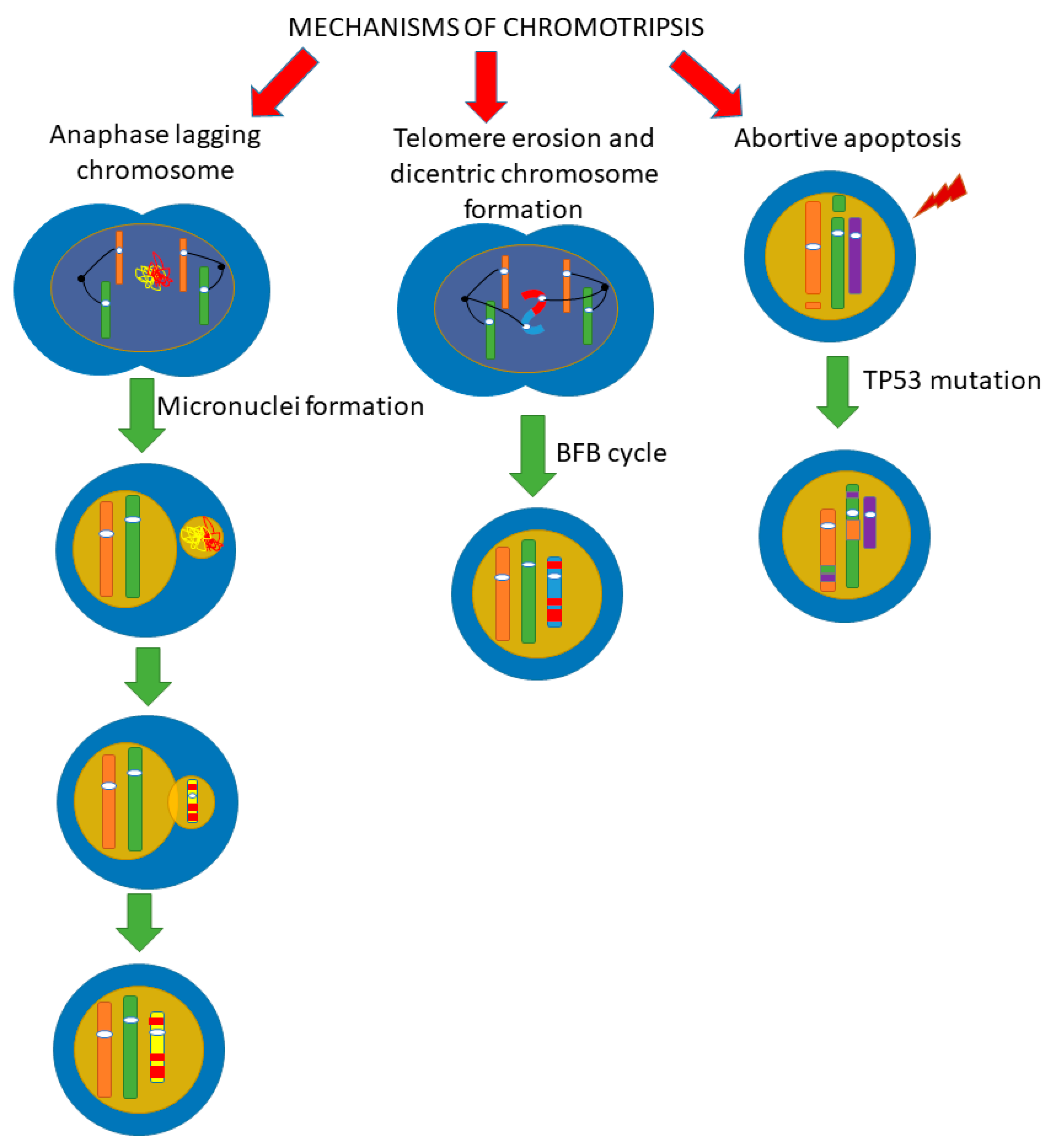
4. Mechanisms of Chromothripsis
4.1. Fragmentation of Chromosomes in Micronuclei
4.2. Telomere Erosion and Dicentric Chromosome Formation
4.3. Abortive Apoptosis
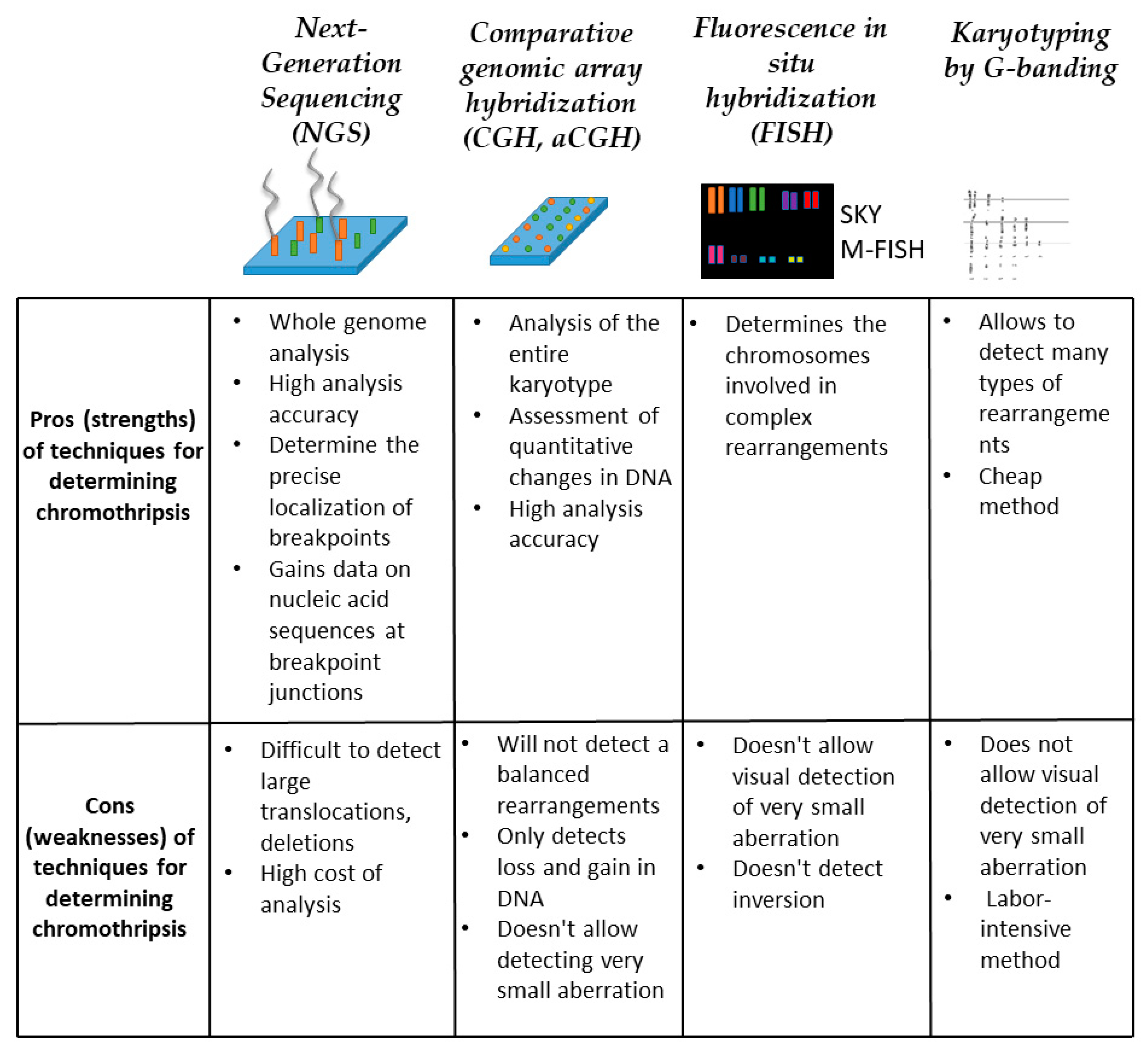
5. Chromothripsis Detection Methods
5.1. Next-Generation Sequencing (NGS)
5.2. Comparative Genomic Array Hybridization (CGH, aCGH)
5.3. Fluorescence In Situ Hybridization (FISH)
5.4. Karyotyping by G-Banding
6. Prevalence of Chromothripsis
7. Contribution of Chromothripsis to Oncogenesis Process
8. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Shilina, M.; Grinchuk, T.; Anatskaya, O.; Vinogradov, A.; Alekseenko, L.; Elmuratov, A.; Nikolsky, N. Cytogenetic and Transcriptomic Analysis of Human Endometrial MSC Retaining Proliferative Activity after Sublethal Heat Shock. Cells 2018, 7, 184. [Google Scholar] [CrossRef]
- Shilina, M.; Nikolsky, N.; Alekseenko, L.; Grinchuk, T. Genetic Stability and Aging. J. Gerontol. Geriatr. Res. 2016, 5, 6. [Google Scholar] [CrossRef]
- Shilina, M.; Kovaleva, Z.; Nikolsky, N.; Grinchuk, T. Genetic stability of human mesenchymal stem cells exposed to X-rays or heat shock in culture. RAD Conf. Proc. 2017, 2, 155–158. [Google Scholar] [CrossRef]
- Alekseenko, L.; Shilina, M.; Kozhukharova, I.; Lyublinskaya, O.; Fridlyanskaya, I.; Nikolsky, N.; Grinchuk, T. Impact of Polyallylamine Hydrochloride on Gene Expression and Karyotypic Stability of Multidrug Resistant Transformed Cells. Cells 2020, 9, 2332. [Google Scholar] [CrossRef] [PubMed]
- Vig, B.K.; Willcourt, M. Decondensation of pericentric heterochromatin alters the sequence of centromere separation in mouse cells. Chromosoma 1998, 107, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Heng, H.H.Q.; Liu, G.; Stevens, J.B.; Abdallah, B.Y.; Horne, S.D.; Ye, K.J.; Bremer, S.W.; Chowdhury, S.K.; Ye, C.J. Karyotype heterogeneity and unclassified chromosomal abnormalities. Cytogenet. Genome Res. 2013, 139, 144–157. [Google Scholar] [CrossRef]
- Vogelstein, B.; Papadopoulos, N.; Velculescu, V.E.; Zhou, S.; Diaz, L.A.; Kinzler, K.W. Cancer genome landscapes. Science 2013, 339, 1546–1558. [Google Scholar] [CrossRef]
- Stephens, P.J.; Greenman, C.D.; Fu, B.; Yang, F.; Bignell, G.R.; Mudie, L.J.; Pleasance, E.D.; Lau, K.W.; Beare, D.; Stebbings, L.A.; et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 2011, 144, 27–40. [Google Scholar] [CrossRef]
- Holland, A.J.; Cleveland, D.W. Chromoanagenesis and cancer: Mechanisms and consequences of localized, complex chromosomal rearrangements. Nat. Med. 2012, 18, 1630–1638. [Google Scholar] [CrossRef]
- Pellestor, F.; Gaillard, J.B.; Schneider, A.; Puechberty, J.; Gatinois, V. Chromoanagenesis, the mechanisms of a genomic chaos. Semin Cell Dev. Biol. 2021, S1084-9521. [Google Scholar] [CrossRef]
- Pellestor, F.; Gatinois, V. Chromoanasynthesis: Another way for the formation of complex chromosomal abnormalities in human reproduction. Hum. Reprod. 2018, 33, 1381–1387. [Google Scholar] [CrossRef]
- Shen, M.M. Chromoplexy: A New Category of Complex Rearrangements in the Cancer Genome. Cancer Cell 2013, 23, 567–569. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Wang, Y.; Collins, C.C. Chromoplexy: A new paradigm in genome remodeling and evolution. Asian J. 2013, 15, 711–712. [Google Scholar] [CrossRef] [PubMed]
- Baca, S.C.; Prandi, D.; Lawrence, M.S.; Mosquera, J.M.; Romanel, A.; Drier, Y.; Park, K.; Kitabayashi, N.; MacDonald, T.Y.; Ghandi, M.; et al. Punctuated evolution of prostate cancer genomes. Cell 2013, 153, 666–677. [Google Scholar] [CrossRef]
- Righolt, C.; Mai, S. Shattered and stitched chromosomes-chromothripsis and chromoanasynthesis-manifestations of a new chromosome crisis? Genes Chromosom. Cancer 2012, 51, 975–981. [Google Scholar] [CrossRef]
- Kinsella, M.; Patel, A.; Bafna, V. The elusive evidence for chromothripsis. Nucleic Acids Res. 2014, 42, 8231–8242. [Google Scholar] [CrossRef]
- Korbel, J.O.; Campbell, P.J. Criteria for inference of chromothripsis in cancer genomes. Cell 2013, 152, 1226–1236. [Google Scholar] [CrossRef] [PubMed]
- Simons, A.; Shaffer, L.G.; Hastings, R.J. Cytogenetic nomenclature: Changes in the ISCN 2013 compared to the 2009 edition. Cytogenet. Genome Res. 2013, 141, 1–6. [Google Scholar] [CrossRef]
- Stevens, J.B.; Abdallah, B.Y.; Liu, G.; Ye, C.J.; Horne, S.D.; Wang, G.; Savasan, S.; Shekhar, M.; Krawetz, S.A.; Hüttemann, M.; et al. Diverse system stresses: Common mechanisms of chromosome fragmentation. Cell Death Dis. 2011, 2, e178. [Google Scholar] [CrossRef] [PubMed]
- Morishita, M.; Muramatsu, T.; Suto, Y.; Hirai, M.; Konishi, T.; Hayashi, S.; Shigemizu, D.; Tsunoda, T.; Moriyama, K.; Inazawa, J. Chromothripsis-like chromosomal rearrangements induced by ionizing radiation using proton microbeam Irradiation system. Oncotarget 2016, 7, 10182–10192. [Google Scholar] [CrossRef]
- Leibowitz, M.L.; Zhang, C.Z.; Pellman, D. Chromothripsis: A New Mechanism for Rapid Karyotype Evolution. Annu. Rev. Genet. 2015, 49, 183–211. [Google Scholar] [CrossRef] [PubMed]
- Fenech, M.; Kirsch-Volders, M.; Natarajan, A.T.; Surralles, J.; Crott, J.W.; Parry, J.; Norppa, H.; Eastmond, D.A.; Tucker, J.D.; Thomas, P. Molecular mechanisms of micronucleus, nucleoplasmic bridge and nuclear bud formation in mammalian and human cells. Mutagenesis 2011, 26, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Thompson, S.L.; Compton, D.A. Chromosome missegregation in human cells arises through specific types of kinetochore-microtubule attachment errors. Proc. Natl. Acad. Sci. USA 2011, 108, 17974–17978. [Google Scholar] [CrossRef] [PubMed]
- Crasta, K.; Ganem, N.J.; Dagher, R.; Lantermann, A.B.; Ivanova, E.V.; Pan, Y.; Nezi, L.; Protopopov, A.; Chowdhury, D.; Pellman, D. DNA breaks and chromosome pulverization from errors in mitosis. Nature 2012, 482, 53–58. [Google Scholar] [CrossRef]
- Terzoudi, G.I.; Karakosta, M.; Pantelias, A.; Hatzi, V.I.; Karachristou, I.; Pantelias, G. Stress induced by premature chromatin condensation triggers chromosome shattering and chromothripsis at DNA sites still replicating in micronuclei or multinucleate cells when primary nuclei enter mitosis. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2015, 793, 185–198. [Google Scholar] [CrossRef]
- Terradas, M.; Martín, M.; Genescà, A. Impaired nuclear functions in micronuclei results in genome instability and chromothripsis. Arch. Toxicol. 2016, 90, 2657–2667. [Google Scholar] [CrossRef] [PubMed]
- Forment, J.V.; Kaidi, A.; Jackson, S.P. Chromothripsis and cancer: Causes and consequences of chromosome shattering. Nat. Rev. Cancer 2012, 12, 663–670. [Google Scholar] [CrossRef]
- Zhang, C.Z.; Spektor, A.; Cornils, H.; Francis, J.M.; Jackson, E.K.; Liu, S.; Meyerson, M.; Pellman, D. Chromothripsis from DNA damage in micronuclei. Nature 2015, 522, 179–184. [Google Scholar] [CrossRef] [PubMed]
- Luijten, M.N.H.; Lee, J.X.T.; Chen, S.; Crasta, K.C. Generation of micronuclei and detection of chromosome pulverization. Methods Mol. Biol. 2018, 1769, 183–195. [Google Scholar] [CrossRef] [PubMed]
- Pantelias, A.; Karachristou, I.; Georgakilas, A.G.; Terzoudi, G.I. Interphase cytogenetic analysis of micronucleated and multinucleated cells supports the premature chromosome condensation hypothesis as the mechanistic origin of chromothripsis. Cancers (Basel) 2019, 11, 1123. [Google Scholar] [CrossRef]
- Sorzano, C.O.S.; Pascual-Montano, A.; De Diego, A.S.; Martińez-A, C.; Van Wely, K.H.M. Chromothripsis: Breakage-fusion-bridge over and over again. Cell Cycle 2013, 12, 2016–2023. [Google Scholar] [CrossRef] [PubMed]
- Fukami, M.; Shima, H.; Suzuki, E.; Ogata, T.; Matsubara, K.; Kamimaki, T. Catastrophic cellular events leading to complex chromosomal rearrangements in the germline. Clin. Genet. 2017, 91, 653–660. [Google Scholar] [CrossRef]
- McClintock, B. The Behavior in Successive Nuclear Divisions of a Chromosome Broken at Meiosis. Proc. Natl. Acad. Sci. USA 1939, 25, 405–416. [Google Scholar] [CrossRef]
- Maciejowski, J.; Li, Y.; Bosco, N.; Campbell, P.J.; De Lange, T. Chromothripsis and Kataegis Induced by Telomere Crisis. Cell 2015, 163, 1641–1654. [Google Scholar] [CrossRef]
- Maciejowski, J.; De Lange, T. Telomeres in cancer: Tumour suppression and genome instability. Nat. Rev. Mol. Cell Biol. 2017, 18, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Schwab, C.; Ryan, S.L.; Papaemmanuil, E.; Robinson, H.M.; Jacobs, P.; Moorman, A.V.; Dyer, S.; Borrow, J.; Griffiths, M.; et al. Constitutional and somatic rearrangement of chromosome 21 in acute lymphoblastic leukaemia. Nature 2014, 508, 98–102. [Google Scholar] [CrossRef]
- Garsed, D.W.; Marshall, O.J.; Corbin, V.D.A.; Hsu, A.; DiStefano, L.; Schröder, J.; Li, J.; Feng, Z.P.; Kim, B.W.; Kowarsky, M.; et al. The Architecture and Evolution of Cancer Neochromosomes. Cancer Cell 2014, 26, 653–667. [Google Scholar] [CrossRef] [PubMed]
- Umbreit, N.T.; Zhang, C.Z.; Lynch, L.D.; Blaine, L.J.; Cheng, A.M.; Tourdot, R.; Sun, L.; Almubarak, H.F.; Judge, K.; Mitchell, T.J.; et al. Mechanisms generating cancer genome complexity from a single cell division error. Science 2020, 368, 0712. [Google Scholar] [CrossRef] [PubMed]
- Tubio, J.M.C.; Estivill, X. Cancer: When catastrophe strikes a cell. Nature 2011, 470, 476–477. [Google Scholar] [CrossRef]
- Tang, H.L.; Tang, H.M.; Mak, K.H.; Hu, S.; Wang, S.S.; Wong, K.M.; Wong, C.S.T.; Wu, H.Y.; Law, H.T.; Liu, K.; et al. Cell survival, DNA damage, and oncogenic transformation after a transient and reversible apoptotic response. Mol. Biol. Cell 2012, 23, 2240–2252. [Google Scholar] [CrossRef]
- Mardin, B.R.; Drainas, A.P.; Waszak, S.M.; Weischenfeldt, J.; Isokane, M.; Stütz, A.M.; Raeder, B.; Efthymiopoulos, T.; Buccitelli, C.; Segura-Wang, M.; et al. A cell-based model system links chromothripsis with hyperploidy. Mol. Syst. Biol. 2015, 11, 828. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.R.; Koren, S.; Sutton, G. Assembly algorithms for next-generation sequencing data. Genomics 2010, 95, 315–327. [Google Scholar] [CrossRef]
- Gao, G.; Smith, D.I. How can mate-pair sequencing be utilized for cancer patients? Expert Rev. Mol. Diagn. 2017, 17, 1–3. [Google Scholar] [CrossRef]
- Keren, B. The advantages of SNP arrays over CGH arrays. Mol. Cytogenet. 2014, 7, I31. [Google Scholar] [CrossRef]
- Balajee, A.S.; Hande, M.P. History and evolution of cytogenetic techniques: Current and future applications in basic and clinical research. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2018, 836, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Shilina, M.A.; Grinchuk, T.M.; Nikolsky, N.N. Genetic stability of human endometrial mesenchymal stem cells assessed with morphological and molecular karyotyping. Cell Tiss. Biol. 2017, 11, 35–41. [Google Scholar] [CrossRef]
- Cortés-Ciriano, I.; Lee, J.J.K.; Xi, R.; Jain, D.; Jung, Y.L.; Yang, L.; Gordenin, D.; Klimczak, L.J.; Zhang, C.Z.; Pellman, D.S.; et al. Comprehensive analysis of chromothripsis in 2,658 human cancers using whole-genome sequencing. Nat. Genet. 2020, 52, 331–341. [Google Scholar] [CrossRef]
- Voronina, N.; Wong, J.K.L.; Hübschmann, D.; Hlevnjak, M.; Uhrig, S.; Heilig, C.E.; Horak, P.; Kreutzfeldt, S.; Mock, A.; Stenzinger, A.; et al. The landscape of chromothripsis across adult cancer types. Nat. Commun. 2020, 11, 2320. [Google Scholar] [CrossRef] [PubMed]
- Rode, A.; Maass, K.K.; Willmund, K.V.; Lichter, P.; Ernst, A. Chromothripsis in cancer cells: An update. Int. J. Cancer 2016, 138, 2322–2333. [Google Scholar] [CrossRef]
- Mamaev, N.; Gindina, T.; Boichenko, E. Chromothripsis in Oncology: Literature Review and Case Report. Clin. Oncohematology 2017, 10, 191–205. [Google Scholar] [CrossRef]
- Malhotra, A.; Lindberg, M.; Faust, G.G.; Leibowitz, M.L.; Clark, R.A.; Layer, R.M.; Quinlan, A.R.; Hall, I.M. Breakpoint profiling of 64 cancer genomes reveals numerous complex rearrangements spawned by homology-independent mechanisms. Genome Res. 2013, 23, 762–776. [Google Scholar] [CrossRef]
- Kloosterman, W.P.; Koster, J.; Molenaar, J.J. Prevalence and clinical implications of chromothripsis in cancer genomes. Curr. Opin. Oncol. 2014, 26, 64–72. [Google Scholar] [CrossRef] [PubMed]
- Behjati, S.; Tarpey, P.S.; Haase, K.; Ye, H.; Young, M.D.; Alexandrov, L.B.; Farndon, S.J.; Collord, G.; Wedge, D.C.; Martincorena, I.; et al. Recurrent mutation of IGF signalling genes and distinct patterns of genomic rearrangement in osteosarcoma. Nat. Commun. 2017, 8, 15936. [Google Scholar] [CrossRef]
- Erson-Omay, E.Z.; Henegariu, O.; Omay, S.B.; Harmanci, A.S.; Youngblood, M.W.; Mishra-Gorur, K.; Li, J.; Özduman, K.; Carrión-Grant, G.; Clark, V.E.; et al. Longitudinal analysis of treatment-induced genomic alterations in gliomas. Genome Med. 2017, 9, 12. [Google Scholar] [CrossRef][Green Version]
- Landuzzi, L.; Manara, M.C.; Lollini, P.L.; Scotlandi, K. Patient Derived Xenografts for Genome-Driven Therapy of Osteosarcoma. Cells 2021, 10, 416. [Google Scholar] [CrossRef]
- Rausch, T.; Jones, D.T.W.; Zapatka, M.; Stütz, A.M.; Zichner, T.; Weischenfeldt, J.; Jäger, N.; Remke, M.; Shih, D.; Northcott, P.A.; et al. Genome sequencing of pediatric medulloblastoma links catastrophic DNA rearrangements with TP53 mutations. Cell 2012, 148, 59–71. [Google Scholar] [CrossRef]
- Ratnaparkhe, M.; Hlevnjak, M.; Kolb, T.; Jauch, A.; Maass, K.K.; Devens, F.; Rode, A.; Hovestadt, V.; Korshunov, A.; Pastorczak, A.; et al. Genomic profiling of Acute lymphoblastic leukemia in ataxia telangiectasia patients reveals tight link between ATM mutations and chromothripsis. Leukemia 2017, 31, 2048–2056. [Google Scholar] [CrossRef]
- Mehine, M.; Kaasinen, E.; Mäkinen, N.; Katainen, R.; Kämpjärvi, K.; Pitkänen, E.; Heinonen, H.-R.; Bützow, R.; Kilpivaara, O.; Kuosmanen, A.; et al. Characterization of Uterine Leiomyomas by Whole-Genome Sequencing. N. Engl. J. Med. 2013, 369, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Mehine, M.; Mäkinen, N.; Heinonen, H.R.; Aaltonen, L.A.; Vahteristo, P. Genomics of uterine leiomyomas: Insights from high-throughput sequencing. Fertil. Steril. 2014, 102, 621–629. [Google Scholar] [CrossRef]
- Holzmann, C.; Markowski, D.N.; Koczan, D.; Küpker, W.; Helmke, B.M.; Bullerdiek, J. Cytogenetically normal uterine leiomyomas without MED12-mutations -A source to identify unknown mechanisms of the development of uterine smooth muscle tumors. Mol. Cytogenet. 2014, 7, 88. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Baltus, C.; Toffoli, S.; London, F.; Delrée, P.; Gilliard, C.; Gustin, T. Chromothripsis in an Early Recurrent Chordoid Meningioma. World Neurosurg. 2019, 130, 380–385. [Google Scholar] [CrossRef]
- Koltsova, A.S.; Pendina, A.A.; Efimova, O.A.; Chiryaeva, O.G.; Kuznetzova, T.V.; Baranov, V.S. On the complexity of mechanisms and consequences of chromothripsis: An update. Front. Genet. 2019, 10, 393. [Google Scholar] [CrossRef]
- De Pagter, M.S.; Van Roosmalen, M.J.; Baas, A.F.; Renkens, I.; Duran, K.J.; Van Binsbergen, E.; Tavakoli-Yaraki, M.; Hochstenbach, R.; Van Der Veken, L.T.; Cuppen, E.; et al. Chromothripsis in healthy individuals affects multiple protein-coding genes and can result in severe congenital abnormalities in offspring. Am. J. Hum. Genet. 2015, 96, 651–656. [Google Scholar] [CrossRef] [PubMed]
- Kloosterman, W.P.; Cuppen, E. Chromothripsis in congenital disorders and cancer: Similarities and differences. Curr. Opin. Cell Biol. 2013, 25, 341–348. [Google Scholar] [CrossRef]
- Chiang, C.; Jacobsen, J.C.; Ernst, C.; Hanscom, C.; Heilbut, A.; Blumenthal, I.; Mills, R.E.; Kirby, A.; Lindgren, A.M.; Rudiger, S.R.; et al. Complex reorganization and predominant non-homologous repair following chromosomal breakage in karyotypically balanced germline rearrangements and transgenic integration. Nat. Genet. 2012, 44, 390–397. [Google Scholar] [CrossRef]
- Collins, R.L.; Brand, H.; Redin, C.E.; Hanscom, C.; Antolik, C.; Stone, M.R.; Glessner, J.T.; Mason, T.; Pregno, G.; Dorrani, N.; et al. Defining the diverse spectrum of inversions, complex structural variation, and chromothripsis in the morbid human genome. Genome Biol. 2017, 18, 36. [Google Scholar] [CrossRef] [PubMed]
- Itani, O.A.; Flibotte, S.; Dumas, K.J.; Moerman, D.G.; Hu, P.J. Chromoanasynthetic genomic rearrangement identified in a N-Ethyl-N-Nitrosourea (ENU) mutagenesis screen in caenorhabditis elegans. G3 Genes Genomes Genet. 2016, 6, 351–356. [Google Scholar] [CrossRef]
- Tan, E.H.; Henry, I.M.; Ravi, M.; Bradnam, K.R.; Mandakova, T.; Marimuthu, M.P.A.; Korf, I.; Lysak, M.A.; Comai, L.; Chan, S.W.L. Catastrophic chromosomal restructuring during genome elimination in plants. eLife 2015, 4, e06516. [Google Scholar] [CrossRef]
- Carbonell-Bejerano, P.; Royo, C.; Torres-Pérez, R.; Grimplet, J.; Fernandez, L.; Franco-Zorrilla, J.M.; Lijavetzky, D.; Baroja, E.; Martínez, J.; García-Escudero, E.; et al. Catastrophic unbalanced genome rearrangements cause somatic loss of berry color in grapevine. Plant. Physiol. 2017, 175, 786–801. [Google Scholar] [CrossRef]
- Liu, P.; Erez, A.; Nagamani, S.C.S.; Dhar, S.U.; Kołodziejska, K.E.; Dharmadhikari, A.V.; Cooper, M.L.; Wiszniewska, J.; Zhang, F.; Withers, M.A.; et al. Chromosome catastrophes involve replication mechanisms generating complex genomic rearrangements. Cell 2011, 146, 889–903. [Google Scholar] [CrossRef]
- Nazaryan, L.; Stefanou, E.G.; Hansen, C.; Kosyakova, N.; Bak, M.; Sharkey, F.H.; Mantziou, T.; Papanastasiou, A.D.; Velissariou, V.; Liehr, T.; et al. The strength of combined cytogenetic and mate-pair sequencing techniques illustrated by a germline chromothripsis rearrangement involving FOXP2. Eur. J. Hum. Genet. 2014, 22, 338–343. [Google Scholar] [CrossRef]
- Northcott, P.A.; Shih, D.J.H.; Peacock, J.; Garzia, L.; Sorana Morrissy, A.; Zichner, T.; Stútz, A.M.; Korshunov, A.; Reimand, J.; Schumacher, S.E.; et al. Subgroup-specific structural variation across 1,000 medulloblastoma genomes. Nature 2012, 487, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Waszak, S.M.; Northcott, P.A.; Buchhalter, I.; Robinson, G.W.; Sutter, C.; Groebner, S.; Grund, K.B.; Brugières, L.; Jones, D.T.W.; Pajtler, K.W.; et al. Spectrum and prevalence of genetic predisposition in medulloblastoma: A retrospective genetic study and prospective validation in a clinical trial cohort. Lancet Oncol. 2018, 19, 785–798. [Google Scholar] [CrossRef]
- Bochtler, T.; Granzow, M.; Stölzel, F.; Kunz, C.; Mohr, B.; Kartal-Kaess, M.; Hinderhofer, K.; Heilig, C.E.; Kramer, M.; Thiede, C.; et al. Marker chromosomes can arise from chromothripsis and predict adverse prognosis in acute myeloid leukemia. Blood 2017, 129, 1333–1342. [Google Scholar] [CrossRef]
- Cai, H.; Kumar, N.; Bagheri, H.C.; von Mering, C.; Robinson, M.D.; Baudis, M. Chromothripsis-like patterns are recurring but heterogeneously distributed features in a survey of 22,347 cancer genome screens. BMC Genom. 2014, 15, 82. [Google Scholar] [CrossRef] [PubMed]
- Rücker, F.G.; Dolnik, A.; Blätte, T.J.; Teleanu, V.; Ernst, A.; Thol, F.; Heuser, M.; Ganser, A.; Döhner, H.; Döhner, K.; et al. Chromothripsis is linked to TP53 alteration, cell cycle impairment, and dismal outcome in acute myeloid leukemia with complex karyotype. Haematologica 2018, 103, e17–e20. [Google Scholar] [CrossRef]
- Fontana, M.C.; Marconi, G.; Feenstra, J.D.M.; Fonzi, E.; Papayannidis, C.; Ghelli Luserna Di Rorá, A.; Padella, A.; Solli, V.; Franchini, E.; Ottaviani, E.; et al. Chromothripsis in acute myeloid leukemia: Biological features and impact on survival. Leukemia 2018, 32, 1609–1620. [Google Scholar] [CrossRef]
- Abáigar, M.; Robledo, C.; Benito, R.; Ramos, F.; Díez-Campelo, M.; Hermosín, L.; Sánchez-Del-Real, J.; Alonso, J.M.; Cuello, R.; Megido, M.; et al. Chromothripsis is a recurrent genomic abnormality in high-risk myelodysplastic syndromes. PLoS ONE 2016, 11, e0164370. [Google Scholar] [CrossRef] [PubMed]
- Cohen, A.; Sato, M.; Aldape, K.; Mason, C.C.; Alfaro-Munoz, K.; Heathcock, L.; South, S.T.; Abegglen, L.M.; Schiffman, J.D.; Colman, H. DNA copy number analysis of Grade II-III and Grade IV gliomas reveals differences in molecular ontogeny including chromothripsis associated with IDH mutation status. Acta Neuropathol. Commun. 2015, 3, 34. [Google Scholar] [CrossRef] [PubMed]
- Furgason, J.M.; Koncar, R.F.; Michelhaugh, S.K.; Sarkar, F.H.; Mittal, S.; Sloan, A.E.; Barnholtz-Sloan, J.S.; Bahassi, E.M. Whole genome sequence analysis links chromothripsis to EGFR, MDM2, MDM4, and CDK4 amplification in glioblastoma. Oncoscience 2015, 2, 618–628. [Google Scholar] [CrossRef]
- Fernandez-Banet, J.; Lee, N.P.; Chan, K.T.; Gao, H.; Liu, X.; Sung, W.K.; Tan, W.; Fan, S.T.; Poon, R.T.; Li, S.; et al. Decoding complex patterns of genomic rearrangement in hepatocellular carcinoma. Genomics 2014, 103, 189–203. [Google Scholar] [CrossRef] [PubMed]
- Morrison, C.D.; Liu, P.; Woloszynska-Read, A.; Zhang, J.; Luo, W.; Qin, M.; Bshara, W.; Conroy, J.M.; Sabatini, L.; Vedell, P.; et al. Whole-genome sequencing identifies genomic heterogeneity at a nucleotide and chromosomal level in bladder cancer. Proc. Natl. Acad. Sci. USA 2014, 111, E672–E681. [Google Scholar] [CrossRef]
- Simovic, M.; Ernst, A. Chromothripsis, DNA repair and checkpoints defects. Semin Cell Dev. Biol. 2021. [Google Scholar] [CrossRef]
- The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 2020, 578, 82. [Google Scholar] [CrossRef]
- Bolkestein, M.; Wong, J.K.L.; Thewes, V.; Körber, V.; Hlevnjak, M.; Elgaafary, S.; Schulze, M.; Kommoss, F.K.F.; Sinn, H.P.; Anzeneder, T.; et al. Chromothripsis in Human Breast Cancer. Cancer Res. 2020, 80, 4918–4931. [Google Scholar] [CrossRef] [PubMed]
- Gröbner, S.N.; Worst, B.C.; Weischenfeldt, J.; Buchhalter, I.; Kleinheinz, K.; Rudneva, V.A.; Johann, P.D.; Balasubramanian, G.P.; Segura-Wang, M.; Brabetz, S.; et al. The landscape of genomic alterations across childhood cancers. Nature 2018, 555, 321–327. [Google Scholar] [CrossRef]
- Habermann, N.; Mardin, B.R.; Yakneen, S.; Korbel, J.O. Using large-scale genome variation cohorts to decipher the molecular mechanism of cancer. Comptes Rendus Biol. 2016, 339, 308–313. [Google Scholar] [CrossRef]
- Przybytkowski, E.; Lenkiewicz, E.; Barrett, M.T.; Klein, K.; Nabavi, S.; Greenwood, C.M.T.; Basik, M. Chromosome-breakage genomic instability and chromothripsis in breast cancer. BMC Genom. 2014, 15, 579. [Google Scholar] [CrossRef]
- Chen, H.; Singh, R.R.; Lu, X.; Huo, L.; Yao, H.; Aldape, K.; Abraham, R.; Virani, S.; Mehrotra, M.; Mishra, B.M.; et al. Genome-wide copy number aberrations and HER2 and FGFR1 alterations in primary breast cancer by molecular inversion probe microarray. Oncotarget 2017, 8, 10845–10857. [Google Scholar] [CrossRef]
- Zack, T.I.; Schumacher, S.E.; Carter, S.L.; Cherniack, A.D.; Saksena, G.; Tabak, B.; Lawrence, M.S.; Zhang, C.Z.; Wala, J.; Mermel, C.H.; et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 2013, 45, 1134–1140. [Google Scholar] [CrossRef]
- Skuja, E.; Kalniete, D.; Nakazawa-Miklasevica, M.; Daneberga, Z.; Abolins, A.; Purkalne, G.; Miklasevics, E. Chromothripsis and progression-free survival in metastatic colorectal cancer. Mol. Clin. Oncol. 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Molenaar, J.J.; Koster, J.; Zwijnenburg, D.A.; van Sluis, P.; Valentijn, L.J.; van der Ploeg, I.; Hamdi, M.; van Nes, J.; Westerman, B.A.; van Arkel, J.; et al. Sequencing of neuroblastoma identifies chromothripsis and defects in neuritogenesis genes. Nature 2012, 483, 589–593. [Google Scholar] [CrossRef]
- Shoshani, O.; Brunner, S.F.; Yaeger, R.; Ly, P.; Nechemia-Arbely, Y.; Kim, D.H.; Fang, R.; Castillon, G.A.; Yu, M.; Li, J.S.Z.; et al. Chromothripsis drives the evolution of gene amplification in cancer. Nature 2021, 591, 137–141. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shorokhova, M.; Nikolsky, N.; Grinchuk, T. Chromothripsis—Explosion in Genetic Science. Cells 2021, 10, 1102. https://doi.org/10.3390/cells10051102
Shorokhova M, Nikolsky N, Grinchuk T. Chromothripsis—Explosion in Genetic Science. Cells. 2021; 10(5):1102. https://doi.org/10.3390/cells10051102
Chicago/Turabian StyleShorokhova, Mariia, Nikolay Nikolsky, and Tatiana Grinchuk. 2021. "Chromothripsis—Explosion in Genetic Science" Cells 10, no. 5: 1102. https://doi.org/10.3390/cells10051102
APA StyleShorokhova, M., Nikolsky, N., & Grinchuk, T. (2021). Chromothripsis—Explosion in Genetic Science. Cells, 10(5), 1102. https://doi.org/10.3390/cells10051102





