Glycosylation of Immune Receptors in Cancer
Abstract
1. Introduction
2. Background of N-Linked Glycosylation
3. N-Glycosylation in Cancer Immunity
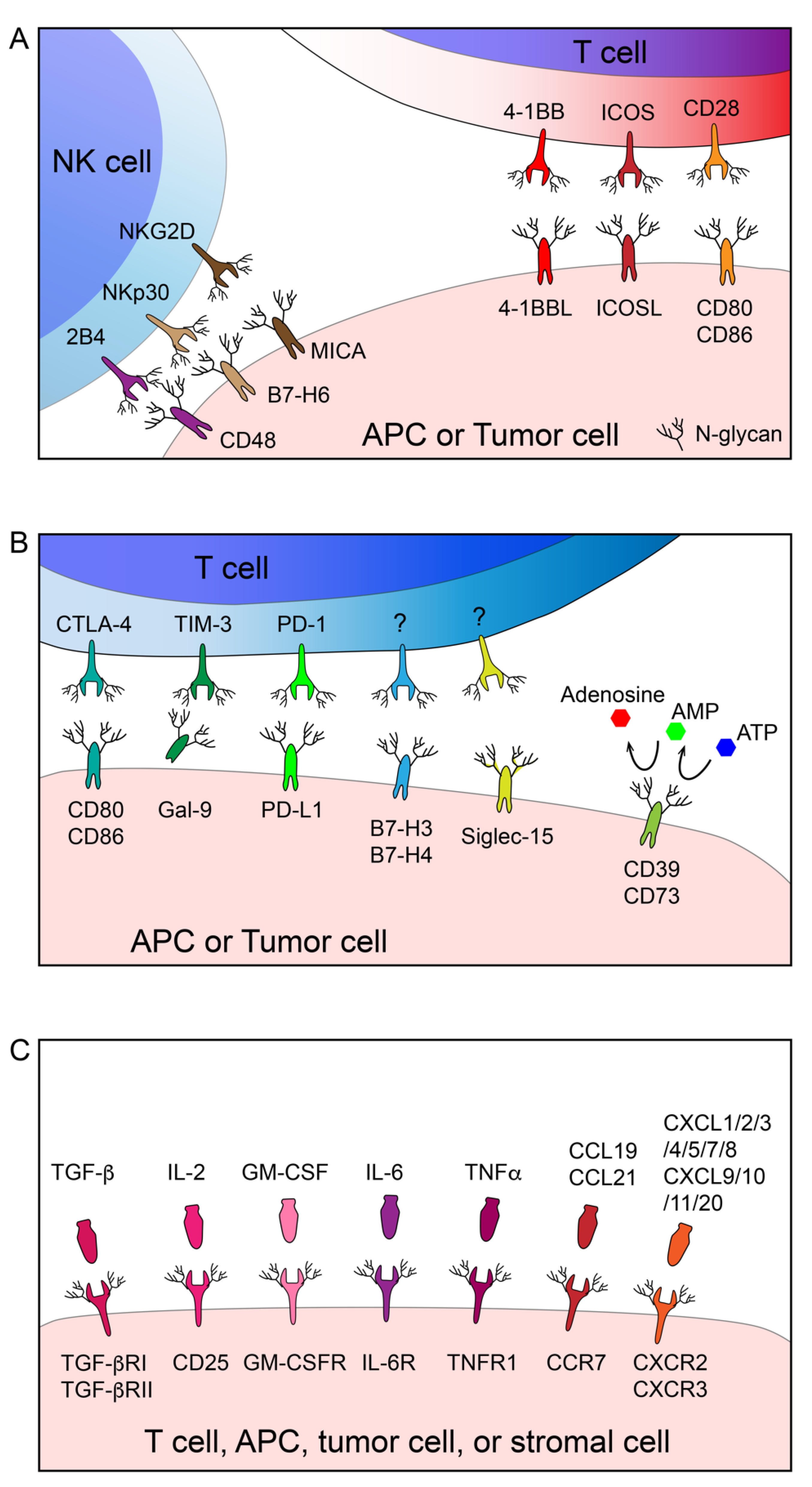
3.1. N-Glycosylation of Stimulatory Immune Receptors
3.2. N-Glycosylation of Inhibitory Immune Receptors
3.3. N-Glycosylation of Cytokine/Chemokine Receptors
4. N-Glycosylation as a Target for Cancer Immunotherapy
5. Concluding Remarks
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chen, D.S.; Mellman, I. Oncology meets immunology: The cancer-immunity cycle. Immunity 2013, 39, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, T.N.; Schreiber, R.D. Neoantigens in cancer immunotherapy. Science 2015, 348, 69–74. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, J.S.; Teng, M.W.L.; Smyth, M.J. Cancer immunoediting and resistance to T cell-based immunotherapy. Nat. Rev. Clin. Oncol. 2019, 16, 151–167. [Google Scholar] [CrossRef]
- An, H.J.; Froehlich, J.W.; Lebrilla, C.B. Determination of glycosylation sites and site-specific heterogeneity in glycoproteins. Curr. Opin. Chem. Biol. 2009, 13, 421–426. [Google Scholar] [CrossRef]
- Helenius, A.; Aebi, M. Intracellular functions of N-linked glycans. Science 2001, 291, 2364–2369. [Google Scholar] [CrossRef]
- Aebi, M. N-linked protein glycosylation in the ER. Biochim. Biophys. Acta 2013, 1833, 2430–2437. [Google Scholar] [CrossRef]
- Stanley, P.; Taniguchi, N.; Aebi, M. N-Glycans. In Essentials of Glycobiology, 3rd ed.; Varki, A., Cummings, R.D., Esko, J.D., Stanley, P., Hart, G.W., Aebi, M., Darvill, A.G., Kinoshita, T., Packer, N.H., Eds.; Cold Spring: Harbor, NY, USA, 2015; pp. 99–111. [Google Scholar]
- Varki, A.; Kannagi, R.; Toole, B.P. Glycosylation Changes in Cancer. In Essentials of Glycobiology, 2nd ed.; Varki, A., Cummings, R.D., Esko, J.D., Freeze, H.H., Stanley, P., Bertozzi, C.R., Hart, G.W., Etzler, M.E., Eds.; Cold Spring: Harbor, NY, USA, 2009. [Google Scholar]
- Beatson, R.; Tajadura-Ortega, V.; Achkova, D.; Picco, G.; Tsourouktsoglou, T.D.; Klausing, S.; Hillier, M.; Maher, J.; Noll, T.; Crocker, P.R.; et al. The mucin MUC1 modulates the tumor immunological microenvironment through engagement of the lectin Siglec-9. Nat. Immunol. 2016, 17, 1273–1281. [Google Scholar] [CrossRef]
- Cabral, J.; Hanley, S.A.; Gerlach, J.Q.; O’Leary, N.; Cunningham, S.; Ritter, T.; Ceredig, R.; Joshi, L.; Griffin, M.D. Distinctive Surface Glycosylation Patterns Associated With Mouse and Human CD4(+) Regulatory T Cells and Their Suppressive Function. Front. Immunol. 2017, 8, 987. [Google Scholar] [CrossRef]
- Chester, C.; Sanmamed, M.F.; Wang, J.; Melero, I. Immunotherapy targeting 4-1BB: Mechanistic rationale, clinical results, and future strategies. Blood 2018, 131, 49–57. [Google Scholar] [CrossRef]
- Song, X.; Zhou, Z.; Li, H.; Xue, Y.; Lu, X.; Bahar, I.; Kepp, O.; Hung, M.C.; Kroemer, G.; Wan, Y. Pharmacologic Suppression of B7-H4 Glycosylation Restores Antitumor Immunity in Immune-Cold Breast Cancers. Cancer Discov. 2020, 10, 1872–1893. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.; Sun, R.; Oh, W.; Kim, A.M.J.; Schwarz, J.R.; Lim, S.O. Saccharide analog, 2-deoxy-d-glucose enhances 4-1BB-mediated antitumor immunity via PD-L1 deglycosylation. Mol. Carcinog. 2020, 59, 691–700. [Google Scholar] [CrossRef]
- Apweiler, R.; Hermjakob, H.; Sharon, N. On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim. Biophys. Acta 1999, 1473, 4–8. [Google Scholar] [CrossRef]
- Ruiz-Canada, C.; Kelleher, D.J.; Gilmore, R. Cotranslational and posttranslational N-glycosylation of polypeptides by distinct mammalian OST isoforms. Cell 2009, 136, 272–283. [Google Scholar] [CrossRef]
- Shrimal, S.; Cherepanova, N.A.; Gilmore, R. Cotranslational and posttranslocational N-glycosylation of proteins in the endoplasmic reticulum. Semin. Cell Dev. Biol. 2015, 41, 71–78. [Google Scholar] [CrossRef]
- Olzmann, J.A.; Kopito, R.R.; Christianson, J.C. The mammalian endoplasmic reticulum-associated degradation system. Cold Spring Harb. Perspect. Biol. 2013, 5, a013185. [Google Scholar] [CrossRef]
- Ruggiano, A.; Foresti, O.; Carvalho, P. Quality control: ER-associated degradation: Protein quality control and beyond. J. Cell Biol. 2014, 204, 869–879. [Google Scholar] [CrossRef] [PubMed]
- Breitling, J.; Aebi, M. N-linked protein glycosylation in the endoplasmic reticulum. Cold Spring Harb. Perspect. Biol. 2013, 5, a013359. [Google Scholar] [CrossRef] [PubMed]
- Costa, A.F.; Campos, D.; Reis, C.A.; Gomes, C. Targeting Glycosylation: A New Road for Cancer Drug Discovery. Trends Cancer 2020, 6, 757–766. [Google Scholar] [CrossRef]
- Pinho, S.S.; Reis, C.A. Glycosylation in cancer: Mechanisms and clinical implications. Nat. Rev. Cancer 2015, 15, 540–555. [Google Scholar] [CrossRef] [PubMed]
- Sanmamed, M.F.; Chen, L. A Paradigm Shift in Cancer Immunotherapy: From Enhancement to Normalization. Cell 2018, 175, 313–326. [Google Scholar] [CrossRef]
- Khalil, D.N.; Smith, E.L.; Brentjens, R.J.; Wolchok, J.D. The future of cancer treatment: Immunomodulation, CARs and combination immunotherapy. Nat. Rev. Clin. Oncol. 2016, 13, 273–290. [Google Scholar] [CrossRef]
- Kuball, J.; Hauptrock, B.; Malina, V.; Antunes, E.; Voss, R.H.; Wolfl, M.; Strong, R.; Theobald, M.; Greenberg, P.D. Increasing functional avidity of TCR-redirected T cells by removing defined N-glycosylation sites in the TCR constant domain. J. Exp. Med. 2009, 206, 463–475. [Google Scholar] [CrossRef]
- Demetriou, M.; Granovsky, M.; Quaggin, S.; Dennis, J.W. Negative regulation of T-cell activation and autoimmunity by Mgat5 N-glycosylation. Nature 2001, 409, 733–739. [Google Scholar] [CrossRef]
- Dias, A.M.; Dourado, J.; Lago, P.; Cabral, J.; Marcos-Pinto, R.; Salgueiro, P.; Almeida, C.R.; Carvalho, S.; Fonseca, S.; Lima, M.; et al. Dysregulation of T cell receptor N-glycosylation: A molecular mechanism involved in ulcerative colitis. Hum. Mol. Genet. 2014, 23, 2416–2427. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Flies, D.B. Molecular mechanisms of T cell co-stimulation and co-inhibition. Nat. Rev. Immunol. 2013, 13, 227–242. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.Y.; Mikolajczak, S.A.; Yoshida, T.; Yoshida, R.; Kelvin, D.J.; Ochi, A. CD28 T cell costimulatory receptor function is negatively regulated by N-linked carbohydrates. Biochem. Biophys. Res. Commun. 2004, 317, 60–67. [Google Scholar] [CrossRef] [PubMed]
- Vinay, D.S.; Kwon, B.S. 4-1BB signaling beyond T cells. Cell Mol. Immunol. 2011, 8, 281–284. [Google Scholar] [CrossRef] [PubMed]
- Zapata, J.M.; Perez-Chacon, G.; Carr-Baena, P.; Martinez-Forero, I.; Azpilikueta, A.; Otano, I.; Melero, I. CD137 (4-1BB) Signalosome: Complexity Is a Matter of TRAFs. Front. Immunol. 2018, 9, 2618. [Google Scholar] [CrossRef]
- Bitra, A.; Doukov, T.; Wang, J.; Picarda, G.; Benedict, C.A.; Croft, M.; Zajonc, D.M. Crystal structure of murine 4-1BB and its interaction with 4-1BBL support a role for galectin-9 in 4-1BB signaling. J. Biol. Chem. 2018, 293, 1317–1329. [Google Scholar] [CrossRef]
- Li, Y.; Tan, S.; Zhang, C.; Chai, Y.; He, M.; Zhang, C.W.; Wang, Q.; Tong, Z.; Liu, K.; Lei, Y.; et al. Limited Cross-Linking of 4-1BB by 4-1BB Ligand and the Agonist Monoclonal Antibody Utomilumab. Cell Rep. 2018, 25, 909–920.e4. [Google Scholar] [CrossRef]
- Madireddi, S.; Eun, S.Y.; Lee, S.W.; Nemcovicova, I.; Mehta, A.K.; Zajonc, D.M.; Nishi, N.; Niki, T.; Hirashima, M.; Croft, M. Galectin-9 controls the therapeutic activity of 4-1BB-targeting antibodies. J. Exp. Med. 2014, 211, 1433–1448. [Google Scholar] [CrossRef] [PubMed]
- Agresta, L.; Hoebe, K.H.N.; Janssen, E.M. The Emerging Role of CD244 Signaling in Immune Cells of the Tumor Microenvironment. Front. Immunol. 2018, 9, 2809. [Google Scholar] [CrossRef]
- Chlewicki, L.K.; Velikovsky, C.A.; Balakrishnan, V.; Mariuzza, R.A.; Kumar, V. Molecular basis of the dual functions of 2B4 (CD244). J. Immunol. 2008, 180, 8159–8167. [Google Scholar] [CrossRef] [PubMed]
- Brown, M.H.; Boles, K.; van der Merwe, P.A.; Kumar, V.; Mathew, P.A.; Barclay, A.N. 2B4, the natural killer and T cell immunoglobulin superfamily surface protein, is a ligand for CD48. J. Exp. Med. 1998, 188, 2083–2090. [Google Scholar] [CrossRef]
- Margraf-Schonfeld, S.; Bohm, C.; Watzl, C. Glycosylation affects ligand binding and function of the activating natural killer cell receptor 2B4 (CD244) protein. J. Biol. Chem. 2011, 286, 24142–24149. [Google Scholar] [CrossRef] [PubMed]
- Amatore, F.; Gorvel, L.; Olive, D. Role of Inducible Co-Stimulator (ICOS) in cancer immunotherapy. Expert Opin. Biol. Ther. 2020, 20, 141–150. [Google Scholar] [CrossRef] [PubMed]
- Hutloff, A.; Dittrich, A.M.; Beier, K.C.; Eljaschewitsch, B.; Kraft, R.; Anagnostopoulos, I.; Kroczek, R.A. ICOS is an inducible T-cell co-stimulator structurally and functionally related to CD28. Nature 1999, 397, 263–266. [Google Scholar] [CrossRef]
- Kamei, N.; Fukui, R.; Suzuki, Y.; Kajihara, Y.; Kinoshita, M.; Kakehi, K.; Hojo, H.; Tezuka, K.; Tsuji, T. Definitive evidence that a single N-glycan among three glycans on inducible costimulator is required for proper protein trafficking and ligand binding. Biochem. Biophys. Res. Commun. 2010, 391, 557–563. [Google Scholar] [CrossRef]
- Rujas, E.; Cui, H.; Sicard, T.; Semesi, A.; Julien, J.P. Structural characterization of the ICOS/ICOS-L immune complex reveals high molecular mimicry by therapeutic antibodies. Nat. Commun. 2020, 11, 5066. [Google Scholar] [CrossRef]
- Hartmann, J.; Tran, T.V.; Kaudeer, J.; Oberle, K.; Herrmann, J.; Quagliano, I.; Abel, T.; Cohnen, A.; Gatterdam, V.; Jacobs, A.; et al. The stalk domain and the glycosylation status of the activating natural killer cell receptor NKp30 are important for ligand binding. J. Biol. Chem. 2012, 287, 31527–31539. [Google Scholar] [CrossRef]
- Skorepa, O.; Pazicky, S.; Kalouskova, B.; Blaha, J.; Abreu, C.; Jecmen, T.; Rosulek, M.; Fish, A.; Sedivy, A.; Harlos, K.; et al. Natural Killer Cell Activation Receptor NKp30 Oligomerization Depends on Its N-Glycosylation. Cancers 2020, 12, 1998. [Google Scholar] [CrossRef]
- Ferrari de Andrade, L.; Tay, R.E.; Pan, D.; Luoma, A.M.; Ito, Y.; Badrinath, S.; Tsoucas, D.; Franz, B.; May, K.F., Jr.; Harvey, C.J.; et al. Antibody-mediated inhibition of MICA and MICB shedding promotes NK cell-driven tumor immunity. Science 2018, 359, 1537–1542. [Google Scholar] [CrossRef] [PubMed]
- Datema, R.; Schwarz, R.T. Formation of 2-deoxyglucose-containing lipid-linked oligosaccharides. Interference with glycosylation of glycoproteins. Eur. J. Biochem. 1978, 90, 505–516. [Google Scholar] [CrossRef] [PubMed]
- Mellergaard, M.; Skovbakke, S.L.; Schneider, C.L.; Lauridsen, F.; Andresen, L.; Jensen, H.; Skov, S. N-glycosylation of asparagine 8 regulates surface expression of major histocompatibility complex class I chain-related protein A (MICA) alleles dependent on threonine 24. J. Biol. Chem. 2014, 289, 20078–20091. [Google Scholar] [CrossRef] [PubMed]
- Yamada, N.; Kato-Kogoe, N.; Yamanegi, K.; Nishiura, H.; Fujihara, Y.; Fukunishi, S.; Okamura, H.; Terada, N.; Nakasho, K. Inhibition of Asparagine-linked Glycosylation Participates in Hypoxia-induced Down-regulation of Cell-surface MICA Expression. Anticancer Res. 2018, 38, 1353–1359. [Google Scholar] [CrossRef]
- Andresen, L.; Skovbakke, S.L.; Persson, G.; Hagemann-Jensen, M.; Hansen, K.A.; Jensen, H.; Skov, S. 2-deoxy D-glucose prevents cell surface expression of NKG2D ligands through inhibition of N-linked glycosylation. J. Immunol. 2012, 188, 1847–1855. [Google Scholar] [CrossRef]
- Hui, E.; Cheung, J.; Zhu, J.; Su, X.; Taylor, M.J.; Wallweber, H.A.; Sasmal, D.K.; Huang, J.; Kim, J.M.; Mellman, I.; et al. T cell costimulatory receptor CD28 is a primary target for PD-1-mediated inhibition. Science 2017, 355, 1428–1433. [Google Scholar] [CrossRef]
- Yokosuka, T.; Takamatsu, M.; Kobayashi-Imanishi, W.; Hashimoto-Tane, A.; Azuma, M.; Saito, T. Programmed cell death 1 forms negative costimulatory microclusters that directly inhibit T cell receptor signaling by recruiting phosphatase SHP2. J. Exp. Med. 2012, 209, 1201–1217. [Google Scholar] [CrossRef]
- Sun, L.; Li, C.W.; Chung, E.M.; Yang, R.; Kim, Y.S.; Park, A.H.; Lai, Y.J.; Yang, Y.; Wang, Y.H.; Liu, J.; et al. Targeting Glycosylated PD-1 Induces Potent Antitumor Immunity. Cancer Res. 2020, 80, 2298–2310. [Google Scholar] [CrossRef]
- Zhou, X.A.; Zhou, J.; Zhao, L.; Yu, G.; Zhan, J.; Shi, C.; Yuan, R.; Wang, Y.; Chen, C.; Zhang, W.; et al. KLHL22 maintains PD-1 homeostasis and prevents excessive T cell suppression. Proc. Natl. Acad. Sci. USA 2020, 117, 28239–28250. [Google Scholar] [CrossRef]
- Liu, K.; Tan, S.; Jin, W.; Guan, J.; Wang, Q.; Sun, H.; Qi, J.; Yan, J.; Chai, Y.; Wang, Z.; et al. N-glycosylation of PD-1 promotes binding of camrelizumab. EMBO Rep. 2020, 21, e51444. [Google Scholar] [CrossRef]
- Chen, L.; Han, X. Anti-PD-1/PD-L1 therapy of human cancer: Past, present, and future. J. Clin. Investig. 2015, 125, 3384–3391. [Google Scholar] [CrossRef]
- Hsu, J.M.; Xia, W.; Hsu, Y.H.; Chan, L.C.; Yu, W.H.; Cha, J.H.; Chen, C.T.; Liao, H.W.; Kuo, C.W.; Khoo, K.H.; et al. STT3-dependent PD-L1 accumulation on cancer stem cells promotes immune evasion. Nat. Commun. 2018, 9, 1908. [Google Scholar] [CrossRef]
- Li, C.W.; Lim, S.O.; Xia, W.; Lee, H.H.; Chan, L.C.; Kuo, C.W.; Khoo, K.H.; Chang, S.S.; Cha, J.H.; Kim, T.; et al. Glycosylation and stabilization of programmed death ligand-1 suppresses T-cell activity. Nat. Commun. 2016, 7, 12632. [Google Scholar] [CrossRef]
- Cha, J.H.; Yang, W.H.; Xia, W.; Wei, Y.; Chan, L.C.; Lim, S.O.; Li, C.W.; Kim, T.; Chang, S.S.; Lee, H.H.; et al. Metformin Promotes Antitumor Immunity via Endoplasmic-Reticulum-Associated Degradation of PD-L1. Mol. Cell 2018, 71, 606–620.e7. [Google Scholar] [CrossRef] [PubMed]
- D’Arrigo, P.; Russo, M.; Rea, A.; Tufano, M.; Guadagno, E.; Del Basso De Caro, M.L.; Pacelli, R.; Hausch, F.; Staibano, S.; Ilardi, G.; et al. A regulatory role for the co-chaperone FKBP51s in PD-L1 expression in glioma. Oncotarget 2017, 8, 68291–68304. [Google Scholar] [CrossRef] [PubMed]
- Li, C.W.; Lim, S.O.; Chung, E.M.; Kim, Y.S.; Park, A.H.; Yao, J.; Cha, J.H.; Xia, W.; Chan, L.C.; Kim, T.; et al. Eradication of Triple-Negative Breast Cancer Cells by Targeting Glycosylated PD-L1. Cancer Cell 2018, 33, 187–201.e10. [Google Scholar] [CrossRef]
- Benicky, J.; Sanda, M.; Brnakova Kennedy, Z.; Grant, O.C.; Woods, R.J.; Zwart, A.; Goldman, R. PD-L1 Glycosylation and Its Impact on Binding to Clinical Antibodies. J. Proteome Res. 2021, 20, 485–497. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.H.; Wang, Y.N.; Xia, W.; Chen, C.H.; Rau, K.M.; Ye, L.; Wei, Y.; Chou, C.K.; Wang, S.C.; Yan, M.; et al. Removal of N-Linked Glycosylation Enhances PD-L1 Detection and Predicts Anti-PD-1/PD-L1 Therapeutic Efficacy. Cancer Cell 2019, 36, 168–178.e4. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Lee, Y.; Li, Y.; Jiang, Y.; Lu, H.; Zang, W.; Zhao, X.; Liu, L.; Chen, Y.; Tan, H.; et al. Co-inhibitory Molecule B7 Superfamily Member 1 Expressed by Tumor-Infiltrating Myeloid Cells Induces Dysfunction of Anti-tumor CD8(+) T Cells. Immunity 2018, 48, 773–786.e5. [Google Scholar] [CrossRef] [PubMed]
- MacGregor, H.L.; Ohashi, P.S. Molecular Pathways: Evaluating the Potential for B7-H4 as an Immunoregulatory Target. Clin. Cancer Res. 2017, 23, 2934–2941. [Google Scholar] [CrossRef]
- Chen, D.; Li, G.; Ji, C.; Lu, Q.; Qi, Y.; Tang, C.; Xiong, J.; Hu, J.; Yasar, F.B.A.; Zhang, Y.; et al. Enhanced B7-H4 expression in gliomas with low PD-L1 expression identifies super-cold tumors. J. Immunother. Cancer 2020, 8, e000154. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.H.; Martin-Orozco, N.; Zheng, P.; Li, J.; Zhang, P.; Tan, H.; Park, H.J.; Jeong, M.; Chang, S.H.; Kim, B.S.; et al. Inhibition of the B7-H3 immune checkpoint limits tumor growth by enhancing cytotoxic lymphocyte function. Cell Res. 2017, 27, 1034–1045. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.T.; Chen, C.H.; Ku, K.L.; Hsiao, M.; Chiang, C.P.; Hsu, T.L.; Chen, M.H.; Wong, C.H. Glycoprotein B7-H3 overexpression and aberrant glycosylation in oral cancer and immune response. Proc. Natl. Acad. Sci. USA 2015, 112, 13057–13062. [Google Scholar] [CrossRef] [PubMed]
- Alegre, M.L.; Frauwirth, K.A.; Thompson, C.B. T-cell regulation by CD28 and CTLA-4. Nat. Rev. Immunol. 2001, 1, 220–228. [Google Scholar] [CrossRef]
- Chambers, C.A.; Kuhns, M.S.; Egen, J.G.; Allison, J.P. CTLA-4-mediated inhibition in regulation of T cell responses: Mechanisms and manipulation in tumor immunotherapy. Annu Rev. Immunol. 2001, 19, 565–594. [Google Scholar] [CrossRef]
- Lau, K.S.; Partridge, E.A.; Grigorian, A.; Silvescu, C.I.; Reinhold, V.N.; Demetriou, M.; Dennis, J.W. Complex N-glycan number and degree of branching cooperate to regulate cell proliferation and differentiation. Cell 2007, 129, 123–134. [Google Scholar] [CrossRef]
- Zhu, C.; Anderson, A.C.; Schubart, A.; Xiong, H.; Imitola, J.; Khoury, S.J.; Zheng, X.X.; Strom, T.B.; Kuchroo, V.K. The Tim-3 ligand galectin-9 negatively regulates T helper type 1 immunity. Nat. Immunol. 2005, 6, 1245–1252. [Google Scholar] [CrossRef]
- Yang, R.; Sun, L.; Li, C.F.; Wang, Y.H.; Yao, J.; Li, H.; Yan, M.; Chang, W.C.; Hsu, J.M.; Cha, J.H.; et al. Galectin-9 interacts with PD-1 and TIM-3 to regulate T cell death and is a target for cancer immunotherapy. Nat. Commun. 2021, 12, 832. [Google Scholar] [CrossRef]
- Ohta, A. A Metabolic Immune Checkpoint: Adenosine in Tumor Microenvironment. Front. Immunol. 2016, 7, 109. [Google Scholar] [CrossRef]
- Vijayan, D.; Young, A.; Teng, M.W.L.; Smyth, M.J. Targeting immunosuppressive adenosine in cancer. Nat. Rev. Cancer 2017, 17, 709–724. [Google Scholar] [CrossRef]
- Wu, J.J.; Choi, L.E.; Guidotti, G. N-linked oligosaccharides affect the enzymatic activity of CD39: Diverse interactions between seven N-linked glycosylation sites. Mol. Biol. Cell 2005, 16, 1661–1672. [Google Scholar] [CrossRef] [PubMed]
- Alcedo, K.P.; Guerrero, A.; Basrur, V.; Fu, D.; Richardson, M.L.; McLane, J.S.; Tsou, C.C.; Nesvizhskii, A.I.; Welling, T.H.; Lebrilla, C.B.; et al. Tumor-Selective Altered Glycosylation and Functional Attenuation of CD73 in Human Hepatocellular Carcinoma. Hepatol. Commun. 2019, 3, 1400–1414. [Google Scholar] [CrossRef]
- Adams, O.J.; Stanczak, M.A.; von Gunten, S.; Laubli, H. Targeting sialic acid-Siglec interactions to reverse immune suppression in cancer. Glycobiology 2018, 28, 640–647. [Google Scholar] [CrossRef]
- van de Wall, S.; Santegoets, K.C.M.; van Houtum, E.J.H.; Bull, C.; Adema, G.J. Sialoglycans and Siglecs Can Shape the Tumor Immune Microenvironment. Trends Immunol. 2020, 41, 274–285. [Google Scholar] [CrossRef]
- Wang, J.; Sun, J.; Liu, L.N.; Flies, D.B.; Nie, X.; Toki, M.; Zhang, J.; Song, C.; Zarr, M.; Zhou, X.; et al. Siglec-15 as an immune suppressor and potential target for normalization cancer immunotherapy. Nat. Med. 2019, 25, 656–666. [Google Scholar] [CrossRef]
- Chen, X.; Dang, X.; Song, J.; Wang, G.; Liu, C.; Cui, L.; Huang, Z. N-glycosylation of Siglec-15 decreases its lysosome-dependent degradation and promotes its transportation to the cell membrane. Biochem. Biophys. Res. Commun. 2020, 533, 77–82. [Google Scholar] [CrossRef]
- Barkal, A.A.; Brewer, R.E.; Markovic, M.; Kowarsky, M.; Barkal, S.A.; Zaro, B.W.; Krishnan, V.; Hatakeyama, J.; Dorigo, O.; Barkal, L.J.; et al. CD24 signalling through macrophage Siglec-10 is a target for cancer immunotherapy. Nature 2019, 572, 392–396. [Google Scholar] [CrossRef] [PubMed]
- Forgione, R.E.; Di Carluccio, C.; Guzman-Caldentey, J.; Gaglione, R.; Battista, F.; Chiodo, F.; Manabe, Y.; Arciello, A.; Del Vecchio, P.; Fukase, K.; et al. Unveiling Molecular Recognition of Sialoglycans by Human Siglec-10. iScience 2020, 23, 101231. [Google Scholar] [CrossRef] [PubMed]
- Mariathasan, S.; Turley, S.J.; Nickles, D.; Castiglioni, A.; Yuen, K.; Wang, Y.; Kadel, E.E., III; Koeppen, H.; Astarita, J.L.; Cubas, R.; et al. TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 2018, 554, 544–548. [Google Scholar] [CrossRef]
- Tauriello, D.V.F.; Palomo-Ponce, S.; Stork, D.; Berenguer-Llergo, A.; Badia-Ramentol, J.; Iglesias, M.; Sevillano, M.; Ibiza, S.; Canellas, A.; Hernando-Momblona, X.; et al. TGFbeta drives immune evasion in genetically reconstituted colon cancer metastasis. Nature 2018, 554, 538–543. [Google Scholar] [CrossRef] [PubMed]
- Batlle, E.; Massague, J. Transforming Growth Factor-beta Signaling in Immunity and Cancer. Immunity 2019, 50, 924–940. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Kuo, F.; Capistrano, K.J.; Kang, D.; Nixon, B.G.; Shi, W.; Chou, C.; Do, M.H.; Stamatiades, E.G.; Gao, S.; et al. TGF-beta suppresses type 2 immunity to cancer. Nature 2020, 587, 115–120. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.W.; Park, J.; Lee, H.J.; Lee, S.Y.; Kim, S.J. TGF-beta sensitivity is determined by N-linked glycosylation of the type II TGF-beta receptor. Biochem. J. 2012, 445, 403–411. [Google Scholar] [CrossRef] [PubMed]
- Niu, L.; Heaney, M.L.; Vera, J.C.; Golde, D.W. High-affinity binding to the GM-CSF receptor requires intact N-glycosylation sites in the extracellular domain of the beta subunit. Blood 2000, 95, 3357–3362. [Google Scholar] [CrossRef]
- Lin, H.; Wang, D.; Wu, T.; Dong, C.; Shen, N.; Sun, Y.; Sun, Y.; Xie, H.; Wang, N.; Shan, L. Blocking core fucosylation of TGF-beta1 receptors downregulates their functions and attenuates the epithelial-mesenchymal transition of renal tubular cells. Am. J. Physiol. Renal. Physiol. 2011, 300, F1017–F1025. [Google Scholar] [CrossRef]
- Shen, N.; Lin, H.; Wu, T.; Wang, D.; Wang, W.; Xie, H.; Zhang, J.; Feng, Z. Inhibition of TGF-beta1-receptor posttranslational core fucosylation attenuates rat renal interstitial fibrosis. Kidney Int. 2013, 84, 64–77. [Google Scholar] [CrossRef]
- Arce Vargas, F.; Furness, A.J.S.; Solomon, I.; Joshi, K.; Mekkaoui, L.; Lesko, M.H.; Miranda Rota, E.; Dahan, R.; Georgiou, A.; Sledzinska, A.; et al. Fc-Optimized Anti-CD25 Depletes Tumor-Infiltrating Regulatory T Cells and Synergizes with PD-1 Blockade to Eradicate Established Tumors. Immunity 2017, 46, 577–586. [Google Scholar] [CrossRef]
- Solomon, I.; Amann, M.; Goubier, A.; Vargas, F.A.; Zervas, D.; Qing, C.; Henry, J.Y.; Ghorani, E.; Akarca, A.U.; Marafioti, T.; et al. CD25-Treg-depleting antibodies preserving IL-2 signaling on effector T cells enhance effector activation and antitumor immunity. Nat. Cancer 2020, 1, 1153–1166. [Google Scholar] [CrossRef]
- Chien, M.W.; Lin, M.H.; Huang, S.H.; Fu, S.H.; Hsu, C.Y.; Yen, B.L.; Chen, J.T.; Chang, D.M.; Sytwu, H.K. Glucosamine Modulates T Cell Differentiation through Down-regulating N-Linked Glycosylation of CD25. J. Biol. Chem. 2015, 290, 29329–29344. [Google Scholar] [CrossRef]
- Montfort, A.; Colacios, C.; Levade, T.; Andrieu-Abadie, N.; Meyer, N.; Segui, B. The TNF Paradox in Cancer Progression and Immunotherapy. Front. Immunol. 2019, 10, 1818. [Google Scholar] [CrossRef]
- Kearney, C.J.; Vervoort, S.J.; Hogg, S.J.; Ramsbottom, K.M.; Freeman, A.J.; Lalaoui, N.; Pijpers, L.; Michie, J.; Brown, K.K.; Knight, D.A.; et al. Tumor immune evasion arises through loss of TNF sensitivity. Sci. Immunol. 2018, 3, eaar3451. [Google Scholar] [CrossRef]
- Bertrand, F.; Montfort, A.; Marcheteau, E.; Imbert, C.; Gilhodes, J.; Filleron, T.; Rochaix, P.; Andrieu-Abadie, N.; Levade, T.; Meyer, N.; et al. TNFalpha blockade overcomes resistance to anti-PD-1 in experimental melanoma. Nat. Commun. 2017, 8, 2256. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Zhang, D.; Tao, T.; Sun, X.; Liu, X.; Zhu, G.; Xu, Z.; Zhu, L.; Zhang, Y.; Liu, W.; et al. The role of N-glycan modification of TNFR1 in inflammatory microglia activation. Glycoconj. J. 2015, 32, 685–693. [Google Scholar] [CrossRef]
- de Vreede, G.; Morrison, H.A.; Houser, A.M.; Boileau, R.M.; Andersen, D.; Colombani, J.; Bilder, D. A Drosophila Tumor Suppressor Gene Prevents Tonic TNF Signaling through Receptor N-Glycosylation. Dev. Cell 2018, 45, 595–605.e4. [Google Scholar] [CrossRef]
- Maude, S.L.; Barrett, D.; Teachey, D.T.; Grupp, S.A. Managing cytokine release syndrome associated with novel T cell-engaging therapies. Cancer J. 2014, 20, 119–122. [Google Scholar] [CrossRef]
- Le, R.Q.; Li, L.; Yuan, W.; Shord, S.S.; Nie, L.; Habtemariam, B.A.; Przepiorka, D.; Farrell, A.T.; Pazdur, R. FDA Approval Summary: Tocilizumab for Treatment of Chimeric Antigen Receptor T Cell-Induced Severe or Life-Threatening Cytokine Release Syndrome. Oncologist 2018, 23, 943–947. [Google Scholar] [CrossRef]
- Waetzig, G.H.; Chalaris, A.; Rosenstiel, P.; Suthaus, J.; Holland, C.; Karl, N.; Valles Uriarte, L.; Till, A.; Scheller, J.; Grotzinger, J.; et al. N-linked glycosylation is essential for the stability but not the signaling function of the interleukin-6 signal transducer glycoprotein 130. J. Biol. Chem. 2010, 285, 1781–1789. [Google Scholar] [CrossRef]
- Riethmueller, S.; Somasundaram, P.; Ehlers, J.C.; Hung, C.W.; Flynn, C.M.; Lokau, J.; Agthe, M.; Dusterhoft, S.; Zhu, Y.; Grotzinger, J.; et al. Proteolytic Origin of the Soluble Human IL-6R In Vivo and a Decisive Role of N-Glycosylation. PLoS Biol. 2017, 15, e2000080. [Google Scholar] [CrossRef]
- Nagarsheth, N.; Wicha, M.S.; Zou, W. Chemokines in the cancer microenvironment and their relevance in cancer immunotherapy. Nat. Rev. Immunol. 2017, 17, 559–572. [Google Scholar] [CrossRef]
- Hauser, M.A.; Kindinger, I.; Laufer, J.M.; Spate, A.K.; Bucher, D.; Vanes, S.L.; Krueger, W.A.; Wittmann, V.; Legler, D.F. Distinct CCR7 glycosylation pattern shapes receptor signaling and endocytosis to modulate chemotactic responses. J. Leukoc. Biol. 2016, 99, 993–1007. [Google Scholar] [CrossRef]
- Steele, C.W.; Karim, S.A.; Leach, J.D.G.; Bailey, P.; Upstill-Goddard, R.; Rishi, L.; Foth, M.; Bryson, S.; McDaid, K.; Wilson, Z.; et al. CXCR2 Inhibition Profoundly Suppresses Metastases and Augments Immunotherapy in Pancreatic Ductal Adenocarcinoma. Cancer Cell 2016, 29, 832–845. [Google Scholar] [CrossRef]
- Ludwig, A.; Ehlert, J.E.; Flad, H.D.; Brandt, E. Identification of distinct surface-expressed and intracellular CXC-chemokine receptor 2 glycoforms in neutrophils: N-glycosylation is essential for maintenance of receptor surface expression. J. Immunol. 2000, 165, 1044–1052. [Google Scholar] [CrossRef]
- Sun, C.; Zhu, X.; Tao, T.; Zhang, D.; Wang, Y.; Xu, H.; Ren, Y.; Wang, Y. The beta4GalT1 affects the fibroblast-like synoviocytes invasion in rheumatoid arthritis by modifying N-linked glycosylation of CXCR3. Eur. J. Cell Biol. 2017, 96, 172–181. [Google Scholar] [CrossRef]
- Mereiter, S.; Balmana, M.; Campos, D.; Gomes, J.; Reis, C.A. Glycosylation in the Era of Cancer-Targeted Therapy: Where Are We Heading? Cancer Cell 2019, 36, 6–16. [Google Scholar] [CrossRef]
- Okada, M.; Chikuma, S.; Kondo, T.; Hibino, S.; Machiyama, H.; Yokosuka, T.; Nakano, M.; Yoshimura, A. Blockage of Core Fucosylation Reduces Cell-Surface Expression of PD-1 and Promotes Anti-tumor Immune Responses of T Cells. Cell Rep. 2017, 20, 1017–1028. [Google Scholar] [CrossRef]
- Zhang, N.; Li, M.; Xu, X.; Zhang, Y.; Liu, Y.; Zhao, M.; Li, P.; Chen, J.; Fukuda, T.; Gu, J.; et al. Loss of core fucosylation enhances the anticancer activity of cytotoxic T lymphocytes by increasing PD-1 degradation. Eur J. Immunol. 2020, 50, 1820–1833. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Jiang, L.; Li, S.C.; He, Q.J.; Yang, B.; Cao, J. Small molecule inhibitors targeting the PD-1/PD-L1 signaling pathway. Acta Pharmacol. Sin. 2021, 42, 1–9. [Google Scholar] [CrossRef]
- Guzik, K.; Zak, K.M.; Grudnik, P.; Magiera, K.; Musielak, B.; Torner, R.; Skalniak, L.; Domling, A.; Dubin, G.; Holak, T.A. Small-Molecule Inhibitors of the Programmed Cell Death-1/Programmed Death-Ligand 1 (PD-1/PD-L1) Interaction via Transiently Induced Protein States and Dimerization of PD-L1. J. Med. Chem. 2017, 60, 5857–5867. [Google Scholar] [CrossRef] [PubMed]
- Skalniak, L.; Zak, K.M.; Guzik, K.; Magiera, K.; Musielak, B.; Pachota, M.; Szelazek, B.; Kocik, J.; Grudnik, P.; Tomala, M.; et al. Small-molecule inhibitors of PD-1/PD-L1 immune checkpoint alleviate the PD-L1-induced exhaustion of T-cells. Oncotarget 2017, 8, 72167–72181. [Google Scholar] [CrossRef]
- Chen, F.F.; Li, Z.; Ma, D.; Yu, Q. Small-molecule PD-L1 inhibitor BMS1166 abrogates the function of PD-L1 by blocking its ER export. Oncoimmunology 2020, 9, 1831153. [Google Scholar] [CrossRef]
- Shao, B.; Li, C.W.; Lim, S.O.; Sun, L.; Lai, Y.J.; Hou, J.; Liu, C.; Chang, C.W.; Qiu, Y.; Hsu, J.M.; et al. Deglycosylation of PD-L1 by 2-deoxyglucose reverses PARP inhibitor-induced immunosuppression in triple-negative breast cancer. Am. J. Cancer Res. 2018, 8, 1837–1846. [Google Scholar] [PubMed]
- Bull, C.; Boltje, T.J.; Balneger, N.; Weischer, S.M.; Wassink, M.; van Gemst, J.J.; Bloemendal, V.R.; Boon, L.; van der Vlag, J.; Heise, T.; et al. Sialic Acid Blockade Suppresses Tumor Growth by Enhancing T-cell-Mediated Tumor Immunity. Cancer Res. 2018, 78, 3574–3588. [Google Scholar] [CrossRef] [PubMed]
- Stanczak, M.A.; Siddiqui, S.S.; Trefny, M.P.; Thommen, D.S.; Boligan, K.F.; von Gunten, S.; Tzankov, A.; Tietze, L.; Lardinois, D.; Heinzelmann-Schwarz, V.; et al. Self-associated molecular patterns mediate cancer immune evasion by engaging Siglecs on T cells. J. Clin. Investig. 2018, 128, 4912–4923. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.; Gu, S.; Han, T.; Zhang, W.; Huang, L.; Li, Z.; Pan, D.; Fu, J.; Ge, J.; Brown, M.; et al. Inhibition of MAN2A1 Enhances the Immune Response to Anti-PD-L1 in Human Tumors. Clin. Cancer Res. 2020, 26, 5990–6002. [Google Scholar] [CrossRef] [PubMed]
- Xiao, H.; Woods, E.C.; Vukojicic, P.; Bertozzi, C.R. Precision glycocalyx editing as a strategy for cancer immunotherapy. Proc. Natl. Acad. Sci. USA 2016, 113, 10304–10309. [Google Scholar] [CrossRef]
- Shi, X.; Zhang, D.; Li, F.; Zhang, Z.; Wang, S.; Xuan, Y.; Ping, Y.; Zhang, Y. Targeting glycosylation of PD-1 to enhance CAR-T cell cytotoxicity. J. Hematol. Oncol. 2019, 12, 127. [Google Scholar] [CrossRef] [PubMed]
- Rillahan, C.D.; Antonopoulos, A.; Lefort, C.T.; Sonon, R.; Azadi, P.; Ley, K.; Dell, A.; Haslam, S.M.; Paulson, J.C. Global metabolic inhibitors of sialyl- and fucosyltransferases remodel the glycome. Nat. Chem. Biol. 2012, 8, 661–668. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, S.A.; Restifo, N.P. Adoptive cell transfer as personalized immunotherapy for human cancer. Science 2015, 348, 62–68. [Google Scholar] [CrossRef]
- McGowan, E.; Lin, Q.; Ma, G.; Yin, H.; Chen, S.; Lin, Y. PD-1 disrupted CAR-T cells in the treatment of solid tumors: Promises and challenges. Biomed. Pharmacother. 2020, 121, 109625. [Google Scholar] [CrossRef]
- John, L.B.; Devaud, C.; Duong, C.P.; Yong, C.S.; Beavis, P.A.; Haynes, N.M.; Chow, M.T.; Smyth, M.J.; Kershaw, M.H.; Darcy, P.K. Anti-PD-1 antibody therapy potently enhances the eradication of established tumors by gene-modified T cells. Clin. Cancer Res. 2013, 19, 5636–5646. [Google Scholar] [CrossRef] [PubMed]
- Cherkassky, L.; Morello, A.; Villena-Vargas, J.; Feng, Y.; Dimitrov, D.S.; Jones, D.R.; Sadelain, M.; Adusumilli, P.S. Human CAR T cells with cell-intrinsic PD-1 checkpoint blockade resist tumor-mediated inhibition. J. Clin. Investig. 2016, 126, 3130–3144. [Google Scholar] [CrossRef]
- Rafiq, S.; Yeku, O.O.; Jackson, H.J.; Purdon, T.J.; van Leeuwen, D.G.; Drakes, D.J.; Song, M.; Miele, M.M.; Li, Z.; Wang, P.; et al. Targeted delivery of a PD-1-blocking scFv by CAR-T cells enhances anti-tumor efficacy in vivo. Nat. Biotechnol. 2018, 36, 847–856. [Google Scholar] [CrossRef] [PubMed]
- Singh, A. Glycoproteomics. Nat. Methods 2021, 18, 28. [Google Scholar] [CrossRef]
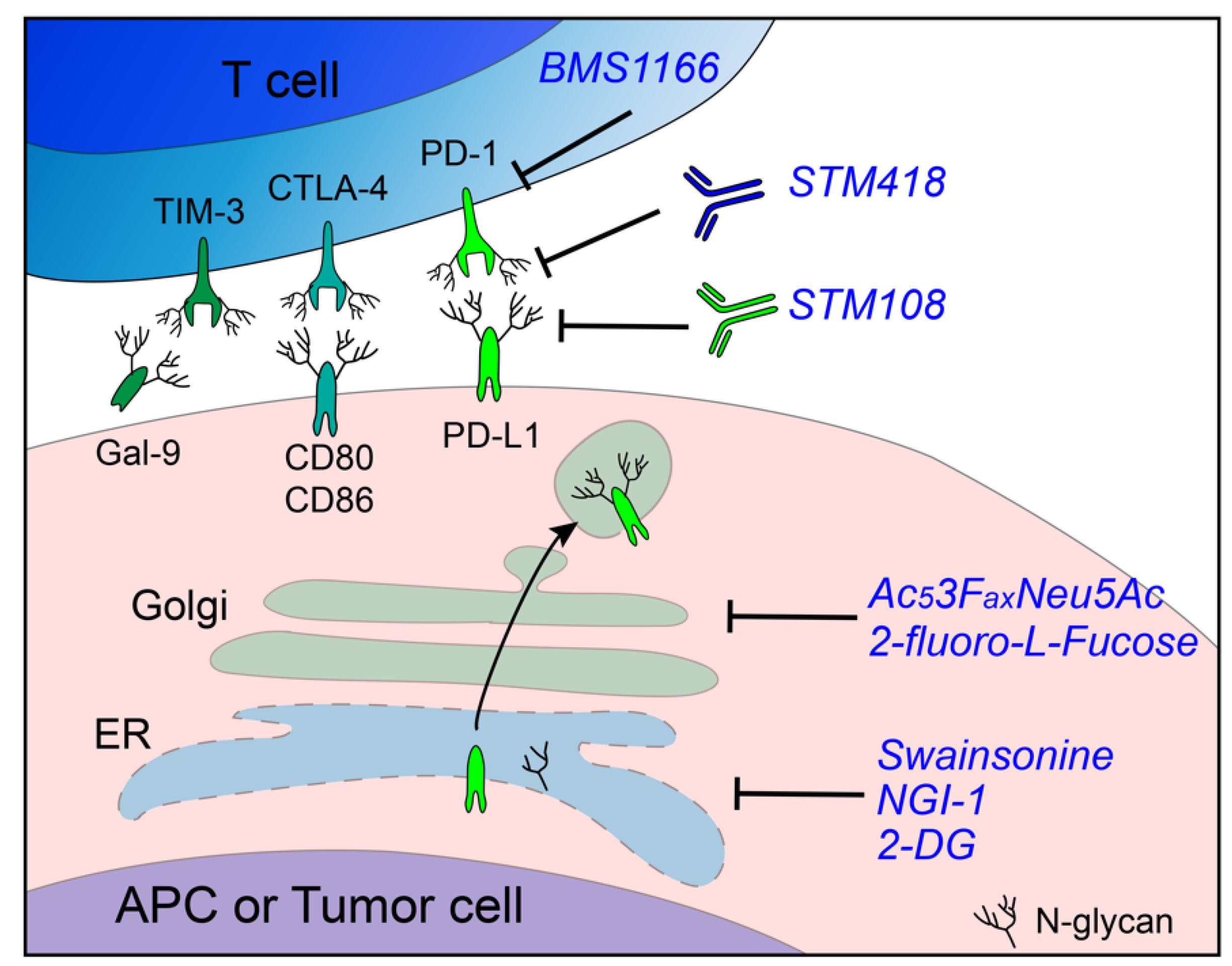
| Drug name | Type | Target | Reference |
|---|---|---|---|
| 2-DG | Sugar analog | Synthesis of lipid-linked oligosaccharides | [13,114] |
| 2-fluoro-l-Fucose | Sugar analog | α-(1,6)-Fucosyltransferase | [108] |
| Ac53FaxNeu5Ac | Sugar analog | Sialyltransferase | [115,116] |
| Swainsonine | Small molecule | α-Mannosidase II | [117] |
| NGI-1 | Small molecule | STT3A/B | [12] |
| BMS1166 | Small molecule | PD-L1 | [113] |
| STM108 | Monoclonal antibody | (N-glycosylated) PD-L1 | [59] |
| STM418 | Monoclonal antibody | (N-glycosylated) PD-1 | [51] |
| T-Sia | Enzyme–antibody conjugate | Tumor surface sialic acid | [118] |
| Base-edited CAR-T | CAR-T | PD-1 (N74) | [119] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, R.; Kim, A.M.J.; Lim, S.-O. Glycosylation of Immune Receptors in Cancer. Cells 2021, 10, 1100. https://doi.org/10.3390/cells10051100
Sun R, Kim AMJ, Lim S-O. Glycosylation of Immune Receptors in Cancer. Cells. 2021; 10(5):1100. https://doi.org/10.3390/cells10051100
Chicago/Turabian StyleSun, Ruoxuan, Alyssa Min Jung Kim, and Seung-Oe Lim. 2021. "Glycosylation of Immune Receptors in Cancer" Cells 10, no. 5: 1100. https://doi.org/10.3390/cells10051100
APA StyleSun, R., Kim, A. M. J., & Lim, S.-O. (2021). Glycosylation of Immune Receptors in Cancer. Cells, 10(5), 1100. https://doi.org/10.3390/cells10051100






