Voluntary Wheel Running Did Not Alter Gene Expression in 5xfad Mice, but in Wild-Type Animals Exclusively after One-Day of Physical Activity
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Tissue Preparation
2.3. Lactate Measurement
2.4. RNA-Seq Library Preparation and Sequencing
2.5. RNA-Seq Analysis
3. Results
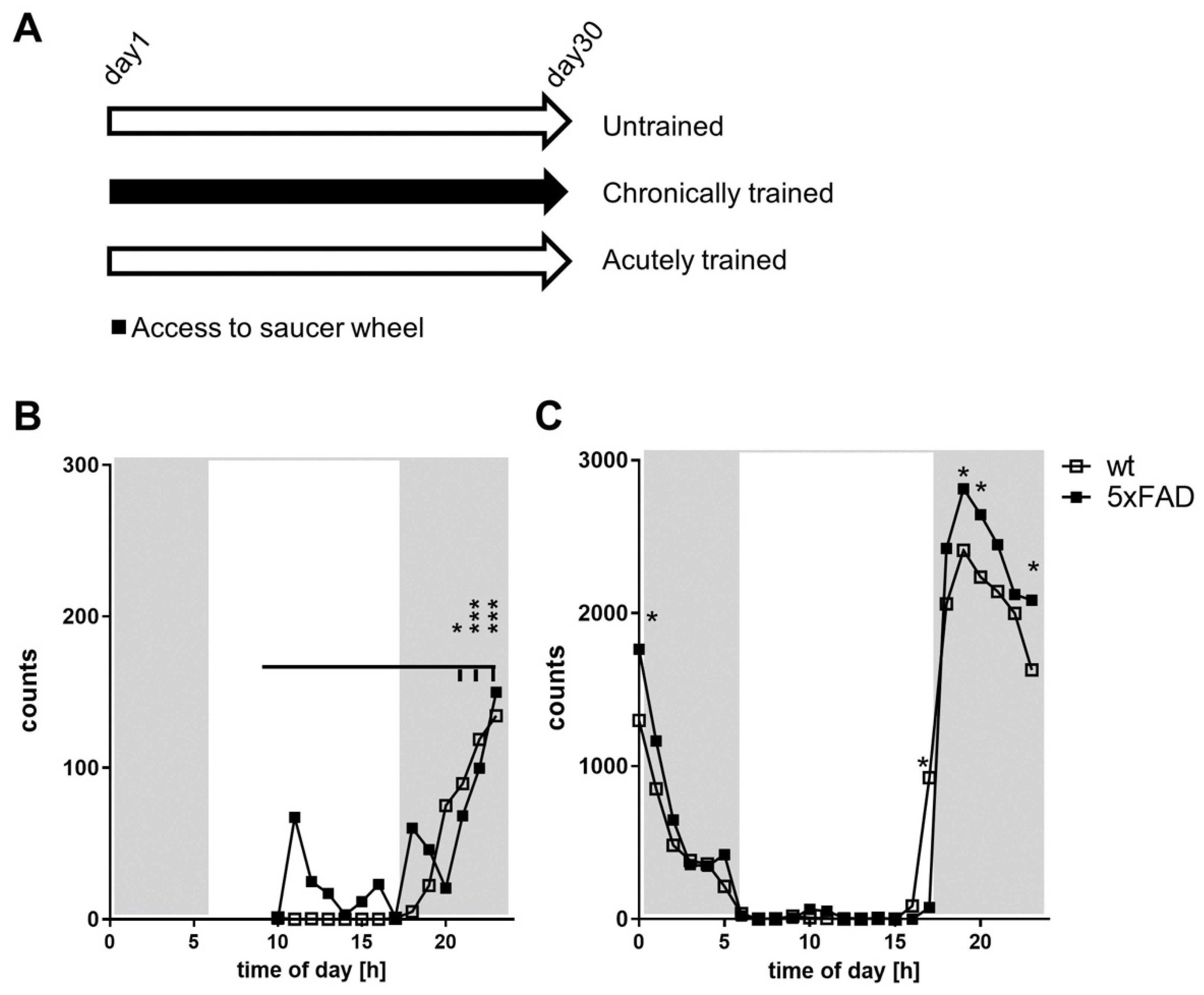
3.1. Voluntary Running Wheel Usage in 5xFAD Mice
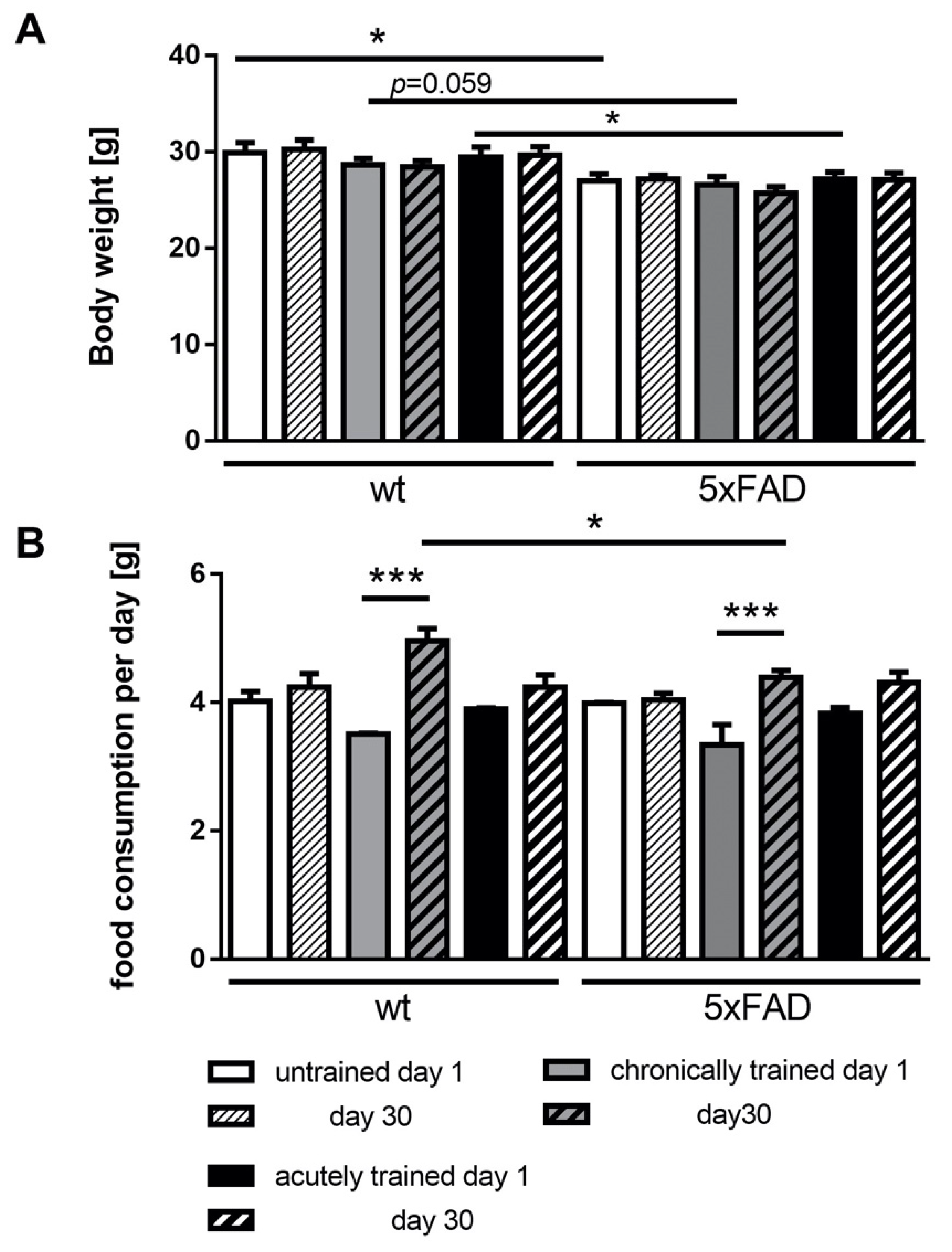
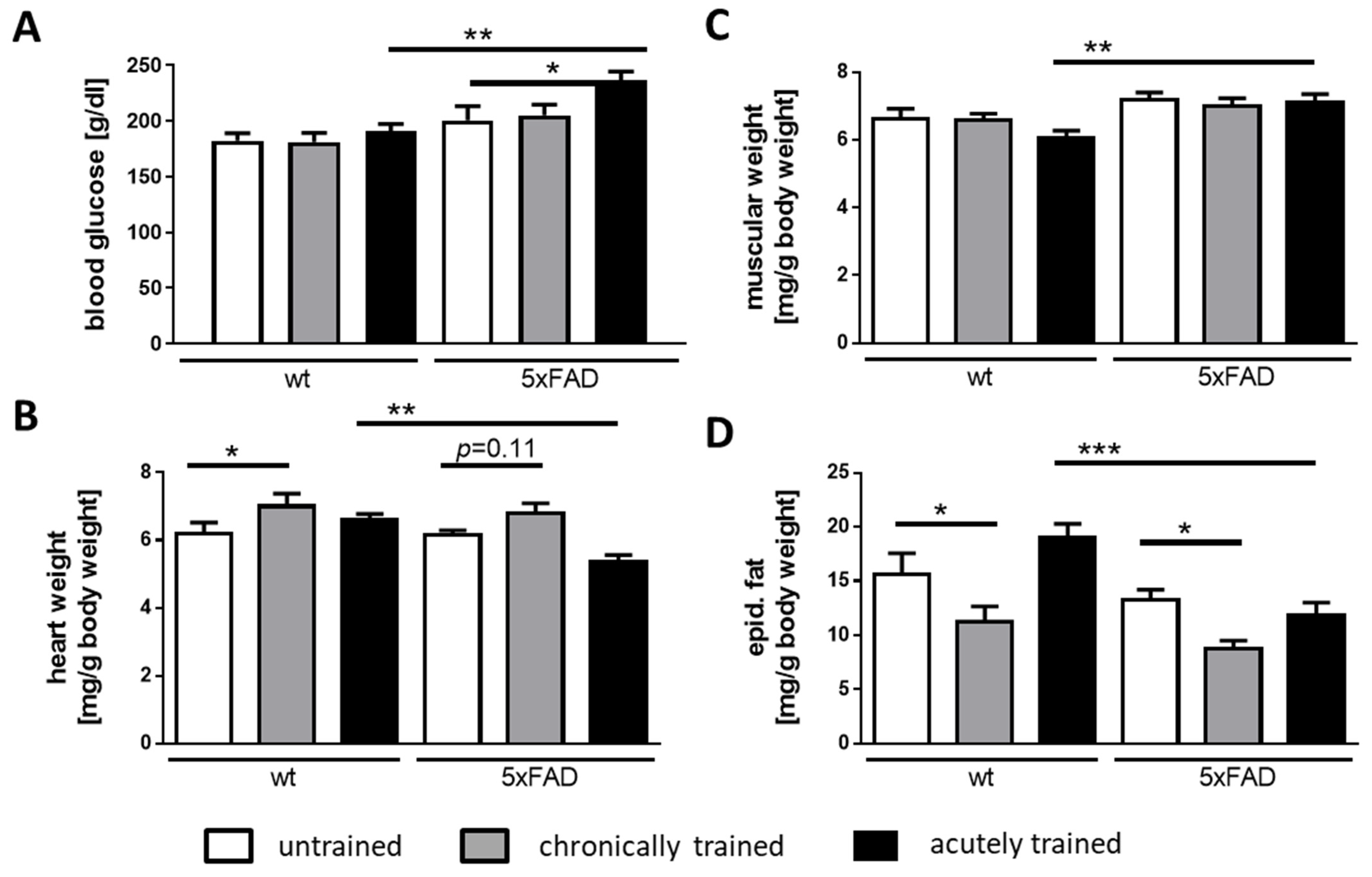
3.2. Physical Parameters of 5xFAD Mice under Acute and Chronic Saucer Wheel Usage
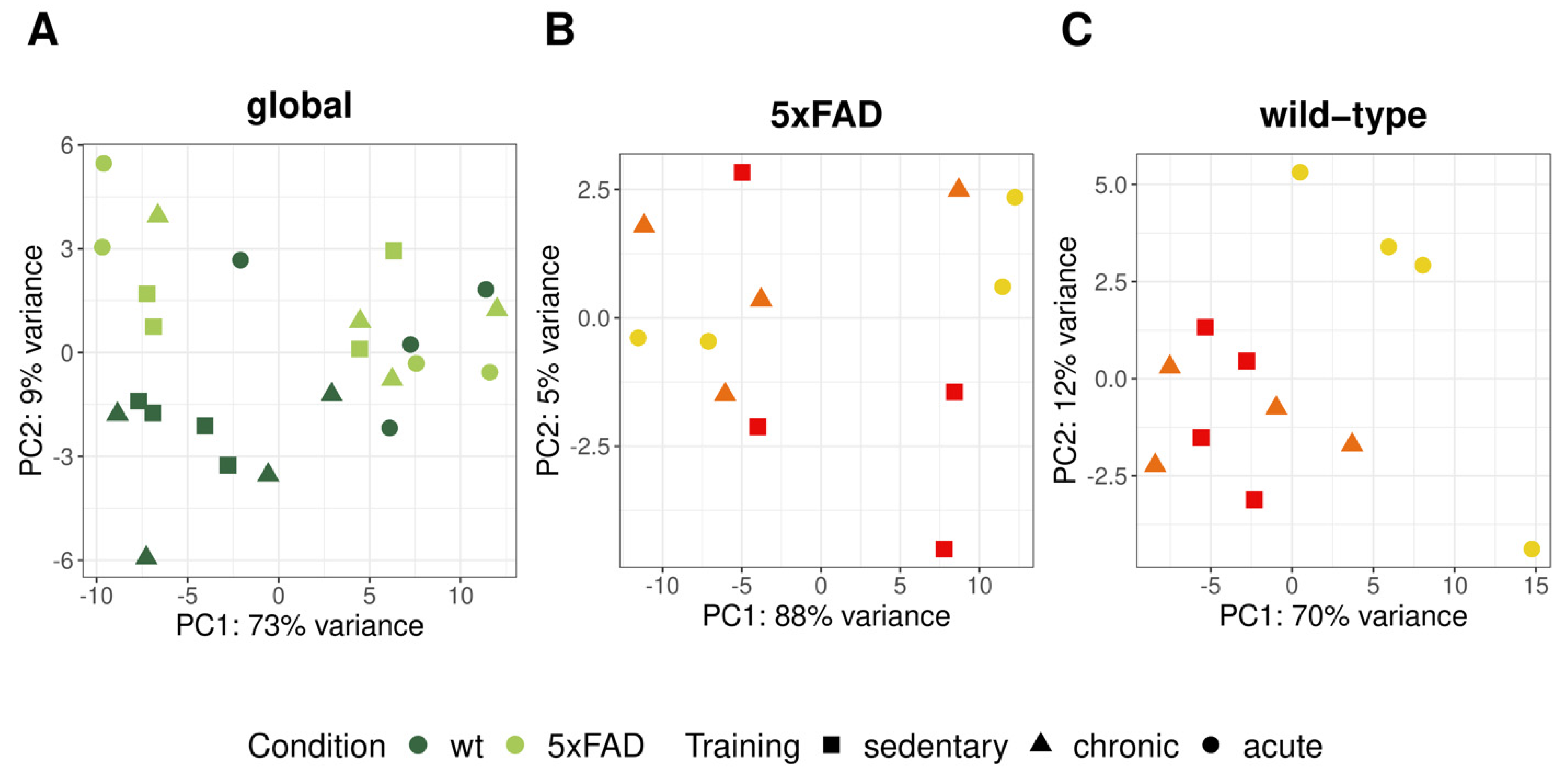
3.3. RNA-Seq Analysis of 5xFAD and Wild-Type Mice with Chronic and Acute Physical Activity
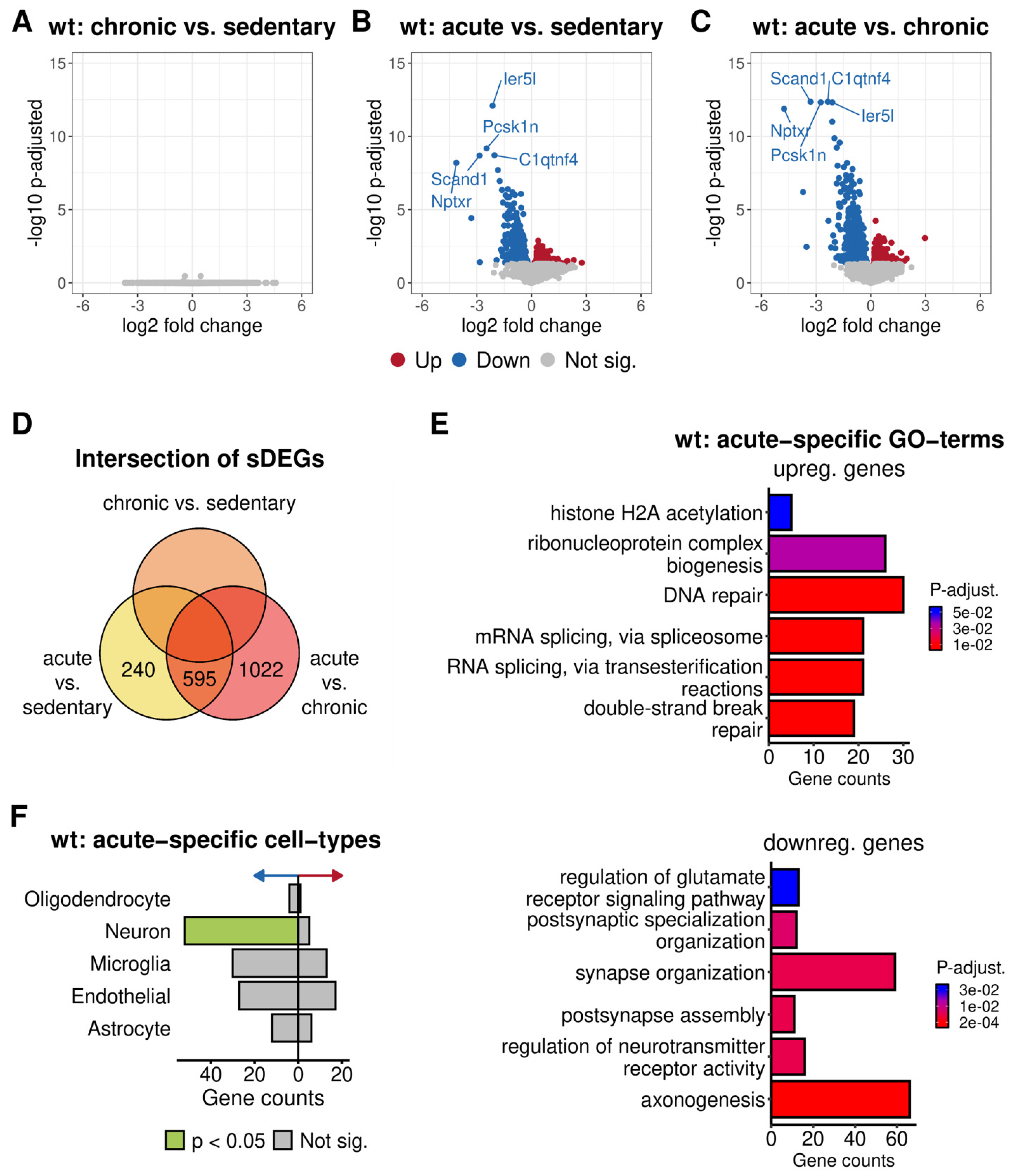
3.3.1. Differential Expression Analysis between Different Physical Activity Levels in 5xFAD and Wild-Type Mice
3.3.2. Functional Annotation of the Altered Gene Expression of Acutely Trained Wild-Type Mice
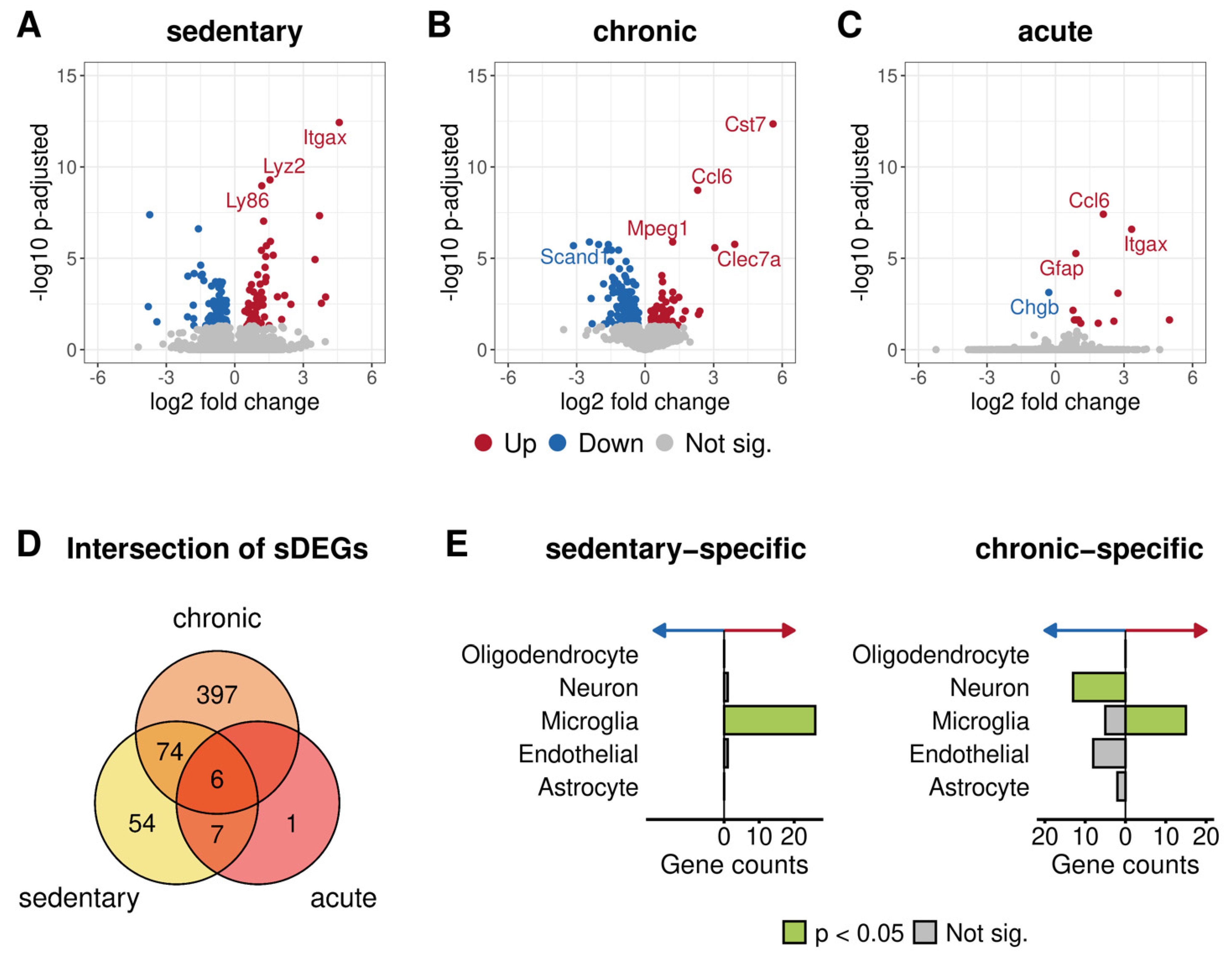
3.3.3. Differential Expression Analysis between 5xFAD and Wild-Type Mice after Different Physical Activity Levels
3.3.4. Functional Annotation of Genotype-Specific Genes Found after Chronic and No Training
4. Discussion
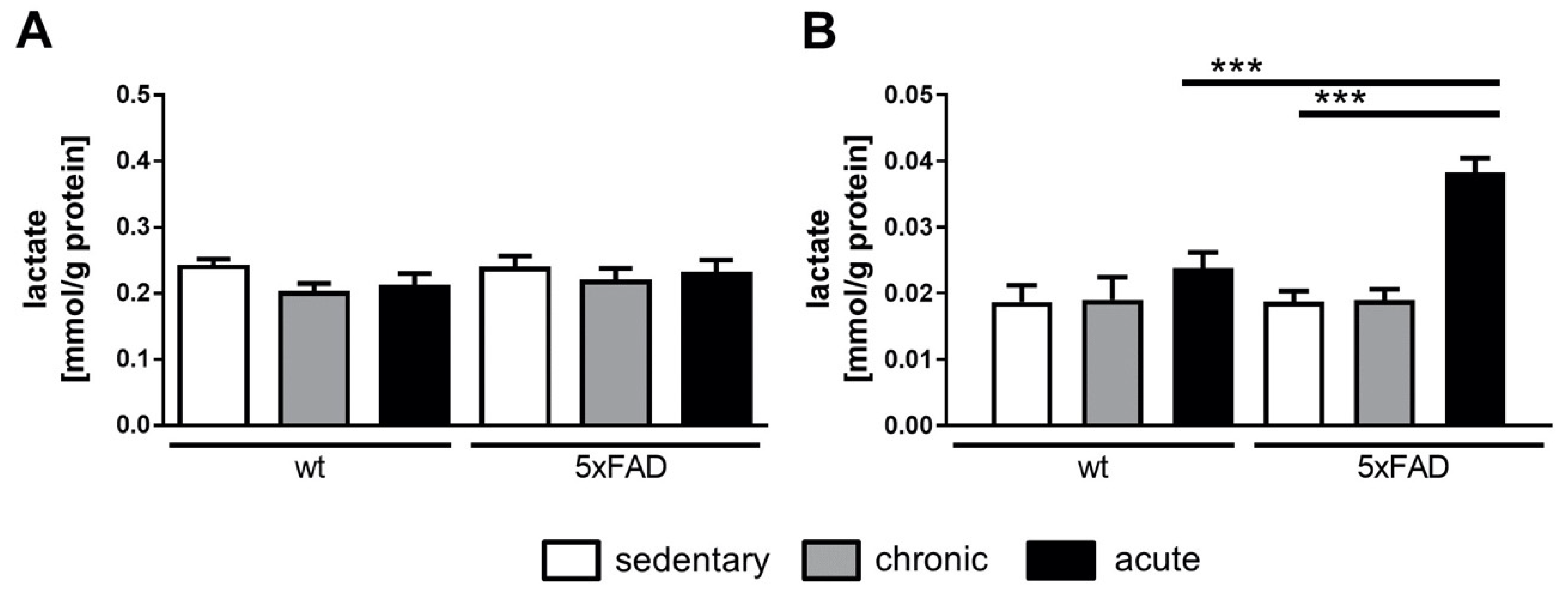
4.1. Brain Lactate Was Drastically Increased in 5xFAD Mice after Acute Physical Activity
4.2. One Day of Voluntary Wheel Running Altered Transcription Only in Wild-Type but Not in 5xFAD Mice
4.3. Immune-Related Genes Were Upregulated in Untrained and Chronically Trained 5xFAD Mice Compared to Their Wild-Type Counterparts
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Roth, G.A.; Abate, D.; Abate, K.H.; Abay, S.M.; Abbafati, C.; Abbasi, N.; Abbastabar, H.; Abd-Allah, F.; Abdela, J.; Abdelalim, A.; et al. Global, Regional, and National Age-Sex-Specific Mortality for 282 Causes of Death in 195 Countries and Territories, 1980–2017: A Systematic Analysis for the Global Burden of Disease Study 2017. Lancet 2018, 392, 1736–1788. [Google Scholar] [CrossRef]
- Conde-Sala, J.L.; Turró-Garriga, O.; Piñán-Hernández, S.; Portellano-Ortiz, C.; Viñas-Diez, V.; Gascón-Bayarri, J.; Reñé-Ramírez, R. Effects of Anosognosia and Neuropsychiatric Symptoms on the Quality of Life of Patients with Alzheimer’s Disease: A 24-Month Follow-up Study. Int. J. Geriatr. Psychiatry 2016, 31, 109–119. [Google Scholar] [CrossRef] [PubMed]
- McKhann, G.; Drachman, D.; Folstein, M.; Katzman, R.; Price, D.; Stadlan, E.M. Clinical Diagnosis of Alzheimer’s Disease: Report of the NINCDS-ADRDA Work Group under the Auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology 1984, 34, 939–944. [Google Scholar] [CrossRef]
- Luque-Contreras, D.; Carvajal, K.; Toral-Rios, D.; Franco-Bocanegra, D.; Campos-Peña, V. Oxidative Stress and Metabolic Syndrome: Cause or Consequence of Alzheimer’s Disease? Oxidative Med. Cell. Longev. 2014, 2014, 1–11. [Google Scholar] [CrossRef]
- Calsolaro, V.; Edison, P. Neuroinflammation in Alzheimer’s Disease: Current Evidence and Future Directions. Alzheimer’s Dement. 2016, 12, 719–732. [Google Scholar] [CrossRef]
- Hardy, J.A.; Higgins, G.A. Alzheimer’s Disease: The Amyloid Cascade Hypothesis. Science 1992, 256, 184–185. [Google Scholar] [CrossRef] [PubMed]
- Arnold, S.E.; Hyman, B.T.; Flory, J.; Damasio, A.R.; Van Hoesen, G.W. The Topographical and Neuroanatomical Distribution of Neurofibrillary Tangles and Neuritic Plaques in the Cerebral Cortex of Patients with Alzheimer’s Disease. Cereb. Cortex 1991, 1, 103–116. [Google Scholar] [CrossRef]
- de la Monte, S.M. Insulin Resistance and Neurodegeneration: Progress Towards the Development of New Therapeutics for Alzheimer’s Disease. Drugs 2017, 77, 47–65. [Google Scholar] [CrossRef] [PubMed]
- Hinz, F.I.; Geschwind, D.H. Molecular Genetics of Neurodegenerative Dementias. Cold Spring Harb. Perspect. Biol. 2017, 9, a023705. [Google Scholar] [CrossRef]
- Bellenguez, C.; Grenier-Boley, B.; Lambert, J.C. Genetics of Alzheimer’s Disease: Where We Are, and Where We Are Going. Curr. Opin. Neurobiol. 2020, 61, 40–48. [Google Scholar] [CrossRef]
- Ruffini, N.; Klingenberg, S.; Schweiger, S.; Gerber, S. Common Factors in Neurodegeneration: A Meta-Study Revealing Shared Patterns on a Multi-Omics Scale. Cells 2020, 9, 2642. [Google Scholar] [CrossRef]
- Hewel, C.; Kaiser, J.; Wierczeiko, A.; Linke, J.; Reinhardt, C.; Endres, K.; Gerber, S. Common MiRNA Patterns of Alzheimer’s Disease and Parkinson’s Disease and Their Putative Impact on Commensal Gut Microbiota. Front. Neurosci. 2019, 13, 113. [Google Scholar] [CrossRef]
- Zhang, L.; Tang, W.; Chao, F.; Zhou, C.; Jiang, L.; Zhang, Y.; Liang, X.; Tang, J.; Qi, Y.; Yang, H.; et al. Four-Month Treadmill Exercise Prevents the Decline in Spatial Learning and Memory Abilities and the Loss of Spinophilin-Immunoreactive Puncta in the Hippocampus of APP/PS1 Transgenic Mice. Neurobiol. Dis. 2020, 136, 104723. [Google Scholar] [CrossRef] [PubMed]
- Fagan, A.M.; Head, D.; Shah, A.R.; Marcus, D.; Mintun, M.; Morris, J.C.; Holtzman, D.M. Decreased Cerebrospinal Fluid Aβ42 Correlates with Brain Atrophy in Cognitively Normal Elderly. Ann. Neurol. 2009, 65, 176–183. [Google Scholar] [CrossRef]
- Deming, Y.; Li, Z.; Kapoor, M.; Harari, O.; Del-Aguila, J.L.; Black, K.; Carrell, D.; Cai, Y.; Fernandez, M.V.; Budde, J.; et al. Genome-Wide Association Study Identifies Four Novel Loci Associated with Alzheimer’s Endophenotypes and Disease Modifiers. Acta Neuropathol. 2017, 133, 839–856. [Google Scholar] [CrossRef] [PubMed]
- Morris, J.C.; Price, J.L. Pathologic Correlates of Nondemented Aging, Mild Cognitive Impairment, and Early-Stage Alzheimer’s Disease. J. Mol. Neurosci. 2001, 17, 101–118. [Google Scholar] [CrossRef]
- Jia, R.; Liang, J.; Xu, Y.; Wang, Y. Effects of Physical Activity and Exercise on the Cognitive Function of Patients with Alzheimer Disease: A Meta-Analysis. BMC Geriatr. 2019, 19, 181. [Google Scholar] [CrossRef]
- De la Rosa, A.; Olaso-Gonzalez, G.; Arc-Chagnaud, C.; Millan, F.; Salvador-Pascual, A.; García-Lucerga, C.; Blasco-Lafarga, C.; Garcia-Dominguez, E.; Carretero, A.; Correas, A.G.; et al. Physical Exercise in the Prevention and Treatment of Alzheimer’s Disease. J. Sport Health Sci. 2020, 9, 394–404. [Google Scholar] [CrossRef] [PubMed]
- Orgeta, V.; Mukadam, N.; Sommerlad, A.; Livingston, G. The Lancet Commission on Dementia Prevention, Intervention, and Care: A Call for Action. Ir. J. Psychol. Med. 2019, 36, 85–88. [Google Scholar] [CrossRef] [PubMed]
- Rashid, M.H.; Zahid, M.F.; Zain, S.; Kabir, A.; Hassan, S.U. The Neuroprotective Effects of Exercise on Cognitive Decline: A Preventive Approach to Alzheimer Disease. Cureus 2020, 12, e6958. [Google Scholar] [CrossRef]
- Roig, M.; Nordbrandt, S.; Geertsen, S.S.; Nielsen, J.B. The Effects of Cardiovascular Exercise on Human Memory: A Review with Meta-Analysis. Neurosci. Biobehav. Rev. 2013, 37, 1645–1666. [Google Scholar] [CrossRef]
- Erickson, K.I.; Hillman, C.; Stillman, C.M.; Ballard, R.M.; Bloodgood, B.; Conroy, D.E.; Macko, R.; Marquez, D.X.; Petruzzello, S.J.; Powell, K.E. Physical Activity, Cognition, and Brain Outcomes: A Review of the 2018 Physical Activity Guidelines. Med. Sci. Sports Exerc. 2019, 51, 1242–1251. [Google Scholar] [CrossRef]
- Panza, G.A.; Taylor, B.A.; MacDonald, H.V.; Johnson, B.T.; Zaleski, A.L.; Livingston, J.; Thompson, P.D.; Pescatello, L.S. Can Exercise Improve Cognitive Symptoms of Alzheimer’s Disease? J. Am. Geriatr. Soc. 2018, 66, 487–495. [Google Scholar] [CrossRef]
- Brown, B.M.; Peiffer, J.J.; Martins, R.N. Multiple Effects of Physical Activity on Molecular and Cognitive Signs of Brain Aging: Can Exercise Slow Neurodegeneration and Delay Alzheimer’s Disease? Mol. Psychiatry 2013, 18, 864–874. [Google Scholar] [CrossRef]
- Sellami, M.; Gasmi, M.; Denham, J.; Hayes, L.D.; Stratton, D.; Padulo, J.; Bragazzi, N. Effects of Acute and Chronic Exercise on Immunological Parameters in the Elderly Aged: Can Physical Activity Counteract the Effects of Aging? Front. Immunol. 2018, 9, 2187. [Google Scholar] [CrossRef]
- Segal, S.K.; Cotman, C.W.; Cahill, L.F. Exercise-Induced Noradrenergic Activation Enhances Memory Consolidation in Both Normal Aging and Patients with Amnestic Mild Cognitive Impairment. J. Alzheimer’s Dis. 2012, 32, 1011–1018. [Google Scholar] [CrossRef]
- Von Ah Morano, A.E.; Dorneles, G.P.; Peres, A.; Lira, F.S. The Role of Glucose Homeostasis on Immune Function in Response to Exercise: The Impact of Low or Higher Energetic Conditions. J. Cell. Physiol. 2020, 235, 3169–3188. [Google Scholar] [CrossRef]
- Adlard, P.A.; Perreau, V.M.; Pop, V.; Cotman, C.W. Voluntary Exercise Decreases Amyloid Load in a Transgenic Model of Alzheimer’s Disease. J. Neurosci. 2005, 25, 4217–4221. [Google Scholar] [CrossRef]
- Trigiani, L.J.; Lacalle-Aurioles, M.; Bourourou, M.; Li, L.; Greenhalgh, A.D.; Zarruk, J.G.; David, S.; Fehlings, M.G.; Hamel, E. Benefits of Physical Exercise on Cognition and Glial White Matter Pathology in a Mouse Model of Vascular Cognitive Impairment and Dementia. Glia 2020, 68, 1925–1940. [Google Scholar] [CrossRef]
- Liu, Y.; Chu, J.M.T.; Yan, T.; Zhang, Y.; Chen, Y.; Chang, R.C.C.; Wong, G.T.C. Short-Term Resistance Exercise Inhibits Neuroinflammation and Attenuates Neuropathological Changes in 3xTg Alzheimer’s Disease Mice. J. Neuroinflammation 2020, 17, 4. [Google Scholar] [CrossRef]
- Belarbi, K.; Burnouf, S.; Fernandez-Gomez, F.J.; Laurent, C.; Lestavel, S.; Figeac, M.; Sultan, A.; Troquier, L.; Leboucher, A.; Caillierez, R.; et al. Beneficial Effects of Exercise in a Transgenic Mouse Model of Alzheimer’s Disease-like Tau Pathology. Neurobiol. Dis. 2011, 43, 486–494. [Google Scholar] [CrossRef]
- Ryan, S.M.; Kelly, Á.M. Exercise as a Pro-Cognitive, pro-Neurogenic and Anti-Inflammatory Intervention in Transgenic Mouse Models of Alzheimer’s Disease. Ageing Res. Rev. 2016, 27, 77–92. [Google Scholar] [CrossRef]
- Lu, X.; Moeini, M.; Li, B.; de Montgolfier, O.; Lu, Y.; Bélanger, S.; Thorin, É.; Lesage, F. Voluntary Exercise Increases Brain Tissue Oxygenation and Spatially Homogenizes Oxygen Delivery in a Mouse Model of Alzheimer’s Disease. Neurobiol. Aging 2020, 88, 11–23. [Google Scholar] [CrossRef]
- Basso, J.C.; Suzuki, W.A. The Effects of Acute Exercise on Mood, Cognition, Neurophysiology, and Neurochemical Pathways: A Review. Brain Plast. 2017, 2, 127–152. [Google Scholar] [CrossRef]
- Madden, D.J.; Blumenthal, J.A.; Allen, P.A.; Emery, C.F. Improving Aerobic Capacity in Healthy Older Adults Does Not Necessarily Lead to Improved Cognitive Performance. Psychol. Aging 1989, 4, 307–320. [Google Scholar] [CrossRef]
- Toots, A.; Littbrand, H.; Boström, G.; Hornsten, C.; Holmberg, H.; Lillemor, L.O.; Lindelöf, N.; Nordstrom, P.; Gustafson, Y.; Rosendahl, E. Effects of Exercise on Cognitive Function in Older People with Dementia: A Randomized Controlled Trial. J. Alzheimer’s Dis. 2017, 60, 323–332. [Google Scholar] [CrossRef]
- Hansson, O.; Svensson, M.; Gustavsson, A.M.; Andersson, E.; Yang, Y.; Nägga, K.; Hållmarker, U.; James, S.; Deierborg, T. Midlife Physical Activity Is Associated with Lower Incidence of Vascular Dementia but Not Alzheimer’s Disease. Alzheimer’s Res. Ther. 2019, 11, 87. [Google Scholar] [CrossRef]
- Svensson, M.; Andersson, E.; Manouchehrian, O.; Yang, Y.; Deierborg, T. Voluntary Running Does Not Reduce Neuroinflammation or Improve Non-Cognitive Behavior in the 5xFAD Mouse Model of Alzheimer’s Disease. Sci. Rep. 2020, 10, 1–10. [Google Scholar] [CrossRef]
- Frederiksen, K.S.; Gjerum, L.; Waldemar, G.; Hasselbalch, S.G. Effects of Physical Exercise on Alzheimer’s Disease Biomarkers: A Systematic Review of Intervention Studies. J. Alzheimer’s Dis. 2018, 61, 359–372. [Google Scholar] [CrossRef]
- Valenzuela, P.L.; Castillo-García, A.; Morales, J.S.; de la Villa, P.; Hampel, H.; Emanuele, E.; Lista, S.; Lucia, A. Exercise Benefits on Alzheimer’s Disease: State-of-the-Science. Ageing Res. Rev. 2020, 62, 101108. [Google Scholar] [CrossRef]
- Lista, I.; Sorrentino, G. Biological Mechanisms of Physical Activity in Preventing Cognitive Decline. Cell. Mol. Neurobiol. 2010, 30, 493–503. [Google Scholar] [CrossRef]
- Archer, T. Physical Exercise Alleviates Debilities of Normal Aging and Alzheimer’s Disease. Acta Neurol. Scand. 2011, 123, 221–238. [Google Scholar] [CrossRef]
- Lin, C.-Y.; Chen, S.-H.; Kou, G.-H.; Kuo, C.-M. An Enzymatic Microassay for Lactate Concentration in Blood and Hemolymph. Acta Zool. Taiwanica 1999, 10, 91–101. [Google Scholar]
- Zerbino, D.R.; Achuthan, P.; Akanni, W.; Amode, M.R.; Barrell, D.; Bhai, J.; Billis, K.; Cummins, C.; Gall, A.; Girón, C.G.; et al. Ensembl 2018. Nucleic Acids Res. 2018, 46, D754–D761. [Google Scholar] [CrossRef] [PubMed]
- Illumina, I. IGenomes. Available online: https://support.illumina.com/sequencing/sequencing_software/igenome.html (accessed on 21 October 2020).
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast Universal RNA-Seq Aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Smyth, G.K.; Shi, W. FeatureCounts: An Efficient General Purpose Program for Assigning Sequence Reads to Genomic Features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- R Studio Team. R Studio: Integrated Development Environment for R; R Studio, Inc.: Boston, MA, USA, 2020. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Core Team: Vienna, Austria, 2020. [Google Scholar]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Oakley, H.; Cole, S.L.; Logan, S.; Maus, E.; Shao, P.; Craft, J.; Guillozet-Bongaarts, A.; Ohno, M.; Disterhoft, J.; Van Eldik, L.; et al. Intraneuronal β-Amyloid Aggregates, Neurodegeneration, and Neuron Loss in Transgenic Mice with Five Familial Alzheimer’s Disease Mutations: Potential Factors in Amyloid Plaque Formation. J. Neurosci. 2006, 26, 10129–10140. [Google Scholar] [CrossRef]
- Todorov, H.; Fournier, D.; Gerber, S. Principal Components Analysis: Theory and Application to Gene Expression Data Analysis. Genom. Comput. Biol. 2018, 4, e100041. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B (Methodol.) 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.-G.; Han, Y.; He, Q.-Y. ClusterProfiler: An R Package for Comparing Biological Themes among Gene Clusters. Omics: A J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Fisher, R.A. Statistical Methods for Research Workers; Springer: New York, NY, USA, 1992; pp. 66–70. [Google Scholar]
- Bagot, R.C.C.; Cates, H.M.M.; Purushothaman, I.; Lorsch, Z.S.S.; Walker, D.M.M.; Wang, J.; Huang, X.; Schlüter, O.M.M.; Maze, I.; Peña, C.J.J.; et al. Circuit-Wide Transcriptional Profiling Reveals Brain Region-Specific Gene Networks Regulating Depression Susceptibility. Neuron 2016, 90, 969–983. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, K.; Sloan, S.A.; Bennett, M.L.; Scholze, A.R.; O’Keeffe, S.; Phatnani, H.P.; Guarnieri, P.; Caneda, C.; Ruderisch, N.; et al. An RNA-Sequencing Transcriptome and Splicing Database of Glia, Neurons, and Vascular Cells of the Cerebral Cortex. J. Neurosci. 2014, 34, 11929–11947. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]
- Dusa, A. Venn: Draw Venn Diagrams. 2018. Available online: https://CRAN.R-project.org/package=venn (accessed on 9 February 2021).
- Brandscheid, C.; Schuck, F.; Reinhardt, S.; Schäfer, K.-H.; Pietrzik, C.U.; Grimm, M.; Hartmann, T.; Schwiertz, A.; Endres, K. Altered Gut Microbiome Composition and Tryptic Activity of the 5xFAD Alzheimer’s Mouse Model. J. Alzheimer’s Dis. 2017, 56, 775–788. [Google Scholar] [CrossRef] [PubMed]
- Landel, V.; Baranger, K.; Virard, I.; Loriod, B.; Khrestchatisky, M.; Rivera, S.; Benech, P.; Féron, F. Temporal Gene Profiling of the 5XFAD Transgenic Mouse Model Highlights the Importance of Microglial Activation in Alzheimer’s Disease. Mol. Neurodegener. 2014, 9, 33. [Google Scholar] [CrossRef] [PubMed]
- Proia, P.; di Liegro, C.M.; Schiera, G.; Fricano, A.; Di Liegro, I. Lactate as a Metabolite and a Regulator in the Central Nervous System. Int. J. Mol. Sci. 2016, 17, 1450. [Google Scholar] [CrossRef]
- Valvona, C.J.; Fillmore, H.L.; Nunn, P.B.; Pilkington, G.J. The Regulation and Function of Lactate Dehydrogenase A: Therapeutic Potential in Brain Tumor. Brain Pathol. 2016, 26, 3–17. [Google Scholar] [CrossRef] [PubMed]
- Dienel, G.A.; Cruz, N.F. Aerobic Glycolysis during Brain Activation: Adrenergic Regulation and Influence of Norepinephrine on Astrocytic Metabolism. J. Neurochem. 2016, 138, 14–52. [Google Scholar] [CrossRef]
- Marti, H.H.; Jung, H.H.; Pfeilschifter, J.; Bauer, C. Hypoxia and Cobalt Stimulate Lactate Dehydrogenase (LDH) Activity in Vascular Smooth Muscle Cells. Pflügers Arch. Eur. J. Physiol. 1994, 429, 216–222. [Google Scholar] [CrossRef]
- Lev-Vachnish, Y.; Cadury, S.; Rotter-Maskowitz, A.; Feldman, N.; Roichman, A.; Illouz, T.; Varvak, A.; Nicola, R.; Madar, R.; Okun, E. L-Lactate Promotes Adult Hippocampal Neurogenesis. Front. Neurosci. 2019, 13, 403. [Google Scholar] [CrossRef] [PubMed]
- van Hall, G. Lactate Kinetics in Human Tissues at Rest and during Exercise. Acta Physiol. 2010, 199, 499–508. [Google Scholar] [CrossRef]
- Hering, G.O.; Hennig, E.M.; Riehle, H.J.; Stepan, J. A Lactate Kinetics Method for Assessing the Maximal Lactate Steady State Workload. Front. Physiol. 2018, 9, 310. [Google Scholar] [CrossRef]
- Larrabee, M.G. Lactate Metabolism and Its Effects on Glucose Metabolism in an Excised Neural Tissue. J. Neurochem. 1995, 64, 1734–1741. [Google Scholar] [CrossRef]
- Smith, D.; Pernet, A.; Hallett, W.A.; Bingham, E.; Marsden, P.K.; Amiel, S.A. Lactate: A Preferred Fuel for Human Brain Metabolism in Vivo. J. Cereb. Blood Flow Metab. 2003, 23, 658–664. [Google Scholar] [CrossRef] [PubMed]
- Dienel, G.A. The Metabolic Trinity, Glucose–Glycogen–Lactate, Links Astrocytes and Neurons in Brain Energetics, Signaling, Memory, and Gene Expression. Neurosci. Lett. 2017, 637, 18–25. [Google Scholar] [CrossRef]
- Suzuki, A.; Stern, S.A.; Bozdagi, O.; Huntley, G.W.; Walker, R.H.; Magistretti, P.J.; Alberini, C.M. Astrocyte-Neuron Lactate Transport Is Required for Long-Term Memory Formation. Cell 2011, 144, 810–823. [Google Scholar] [CrossRef]
- El Hayek, L.; Khalifeh, M.; Zibara, V.; Abi Assaad, R.; Emmanuel, N.; Karnib, N.; El-Ghandour, R.; Nasrallah, P.; Bilen, M.; Ibrahim, P.; et al. Lactate Mediates the Effects of Exercise on Learning and Memory through Sirt1-Dependent Activation of Hippocampal Brain-Derived Neurotrophic Factor (BDNF). J. Neurosci. 2019, 39, 2369–2382. [Google Scholar] [CrossRef] [PubMed]
- Cunnane, S.; Nugent, S.; Roy, M.; Courchesne-Loyer, A.; Croteau, E.; Tremblay, S.; Castellano, A.; Pifferi, F.; Bocti, C.; Paquet, N.; et al. Brain Fuel Metabolism, Aging, and Alzheimer’s Disease. Nutrition 2011, 27, 3–20. [Google Scholar] [CrossRef]
- Zuend, M.; Saab, A.S.; Wyss, M.T.; Ferrari, K.D.; Hösli, L.; Looser, Z.J.; Stobart, J.L.; Duran, J.; Guinovart, J.J.; Barros, L.F.; et al. Arousal-Induced Cortical Activity Triggers Lactate Release from Astrocytes. Nat. Metab. 2020, 2, 179–191. [Google Scholar] [CrossRef]
- Magistretti, P.J.; Allaman, I. Lactate in the Brain: From Metabolic End-Product to Signalling Molecule. Nat. Rev. Neurosci. 2018, 19, 235–249. [Google Scholar] [CrossRef]
- Díaz-García, C.M.; Mongeon, R.; Lahmann, C.; Koveal, D.; Zucker, H.; Yellen, G. Neuronal Stimulation Triggers Neuronal Glycolysis and Not Lactate Uptake. Cell Metab. 2017, 26, 361–374.e4. [Google Scholar] [CrossRef] [PubMed]
- Kuo, C.-C.; Chiang, A.W.; Lewis, N.E. Dysregulation of the Secretory Pathway Connects Alzheimer’s Disease Genetics to Aggregate Formation. bioRxiv 2020. [Google Scholar] [CrossRef]
- Hendrix, J.; Nijs, J.; Ickmans, K.; Godderis, L.; Ghosh, M.; Polli, A. The Interplay between Oxidative Stress, Exercise, and Pain in Health and Disease: Potential Role of Autonomic Regulation and Epigenetic Mechanisms. Antioxidants 2020, 9, 1166. [Google Scholar] [CrossRef]
- Kakanis, M.W.; Peake, J.; Brenu, E.W.; Simmonds, M.; Gray, B.; Hooper, S.L.; Marshall-Gradisnik, S.M. The Open Window of Susceptibility to Infection after Acute Exercise in Healthy Young Male Elite Athletes. Exerc. Immunol. Rev. 2010, 16, 119–137. [Google Scholar] [CrossRef]
- Guerriero, F.; Sgarlata, C.; Francis, M.; Maurizi, N.; Faragli, A.; Perna, S.; Rondanelli, M.; Rollone, M.; Ricevuti, G. Neuroinflammation, Immune System and Alzheimer Disease: Searching for the Missing Link. Aging Clin. Exp. Res 2017, 29, 821–831. [Google Scholar] [CrossRef]
- Boza-Serrano, A.; Yang, Y.; Paulus, A.; Deierborg, T. Innate Immune Alterations Are Elicited in Microglial Cells before Plaque Deposition in the Alzheimer’s Disease Mouse Model 5xFAD. Sci. Rep. 2018, 8, 1550. [Google Scholar] [CrossRef] [PubMed]
- Passamonti, X.L.; Tsvetanov, X.K.; Jones, P.; Bevan-Jones, W.; Arnold, R.; Borchert, R.; Mak, E.; Su, L.; Rowe, X.J. Neurobiology of Disease Neuroinflammation and Functional Connectivity in Alzheimer’s Disease: Interactive Influences on Cognitive Performance. J. Neurosci. 2019, 39, 7218–7226. [Google Scholar] [CrossRef]
- Heppner, F.L.; Ransohoff, R.M.; Becher, B. Immune Attack: The Role of Inflammation in Alzheimer Disease. Nat. Rev. Neurosci. 2015, 16, 358–372. [Google Scholar] [CrossRef]
- Chi, L.M.; Wang, X.; Nan, G.X. In Silico Analyses for Molecular Genetic Mechanism and Candidate Genes in Patients with Alzheimer’s Disease. Acta Neurol. Belg. 2016, 116, 543–547. [Google Scholar] [CrossRef]
- Edgar, R.; Domrachev, M.; Lash, A.E. Gene Expression Omnibus: NCBI Gene Expression and Hybridization Array Data Repository. Nucleic Acids Res. 2002, 30, 207–210. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wierczeiko, A.; Gammel, L.; Radyushkin, K.; Nguyen, V.T.T.; Todorov, H.; Gerber, S.; Endres, K. Voluntary Wheel Running Did Not Alter Gene Expression in 5xfad Mice, but in Wild-Type Animals Exclusively after One-Day of Physical Activity. Cells 2021, 10, 693. https://doi.org/10.3390/cells10030693
Wierczeiko A, Gammel L, Radyushkin K, Nguyen VTT, Todorov H, Gerber S, Endres K. Voluntary Wheel Running Did Not Alter Gene Expression in 5xfad Mice, but in Wild-Type Animals Exclusively after One-Day of Physical Activity. Cells. 2021; 10(3):693. https://doi.org/10.3390/cells10030693
Chicago/Turabian StyleWierczeiko, Anna, Lena Gammel, Konstantin Radyushkin, Vu Thu Thuy Nguyen, Hristo Todorov, Susanne Gerber, and Kristina Endres. 2021. "Voluntary Wheel Running Did Not Alter Gene Expression in 5xfad Mice, but in Wild-Type Animals Exclusively after One-Day of Physical Activity" Cells 10, no. 3: 693. https://doi.org/10.3390/cells10030693
APA StyleWierczeiko, A., Gammel, L., Radyushkin, K., Nguyen, V. T. T., Todorov, H., Gerber, S., & Endres, K. (2021). Voluntary Wheel Running Did Not Alter Gene Expression in 5xfad Mice, but in Wild-Type Animals Exclusively after One-Day of Physical Activity. Cells, 10(3), 693. https://doi.org/10.3390/cells10030693






