Expression of Mst89B and CG31287 is Needed for Effective Sperm Storage and Egg Fertilization in Drosophila
Abstract
1. Introduction
2. Materials and Methods
2.1. Gene Selection
2.2. Fly Stocks and Maintenance
2.3. Gene Knockdown
2.4. Fecundity and Sperm Competition
2.5. Sperm Production, Sperm Transfer, Sperm Storage, Egg Hatchability, and Larva Viability
2.6. Statistical Analysis
3. Results
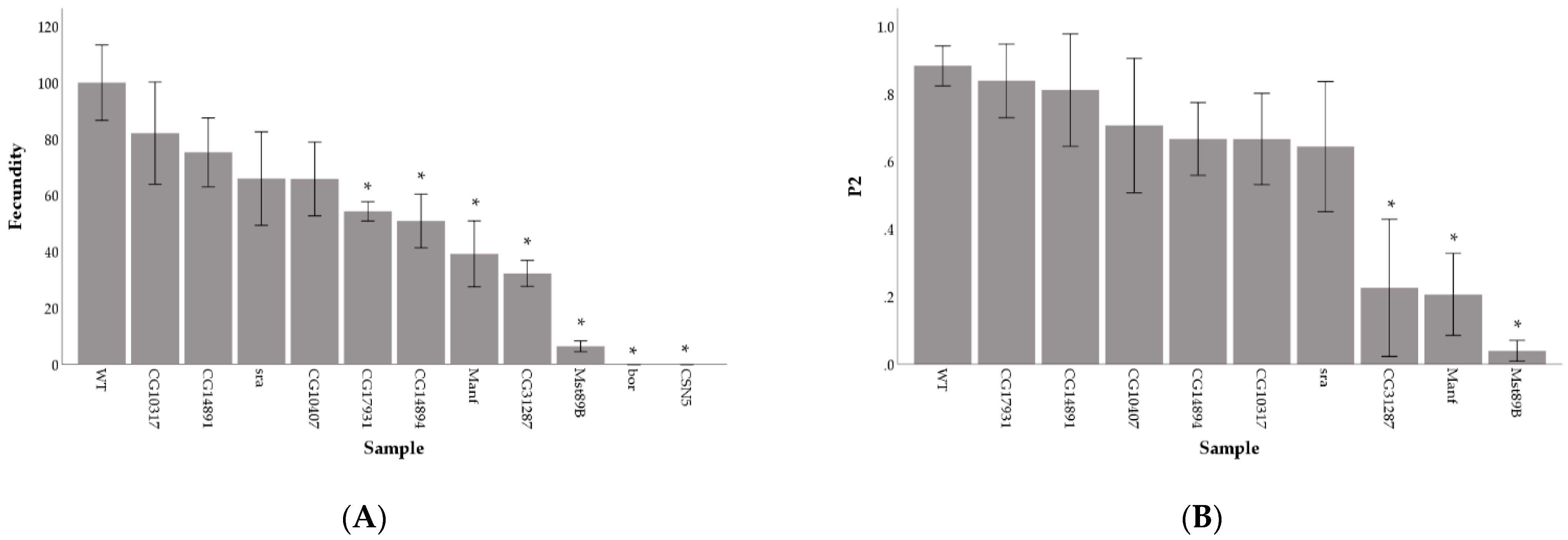
3.1. Reduced Germline Expression via RNAi Identifies Genes Needed for Male Fecundity
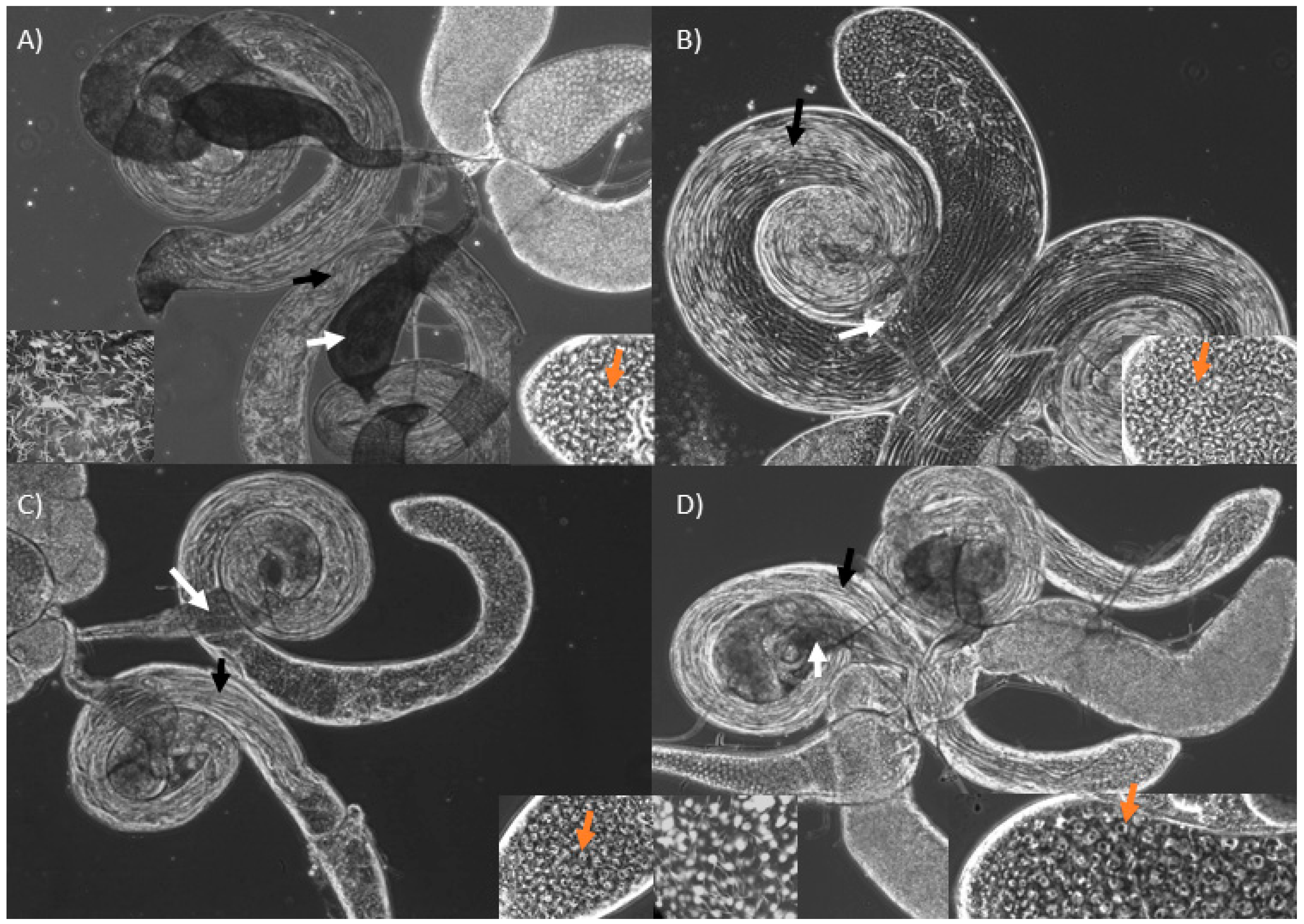
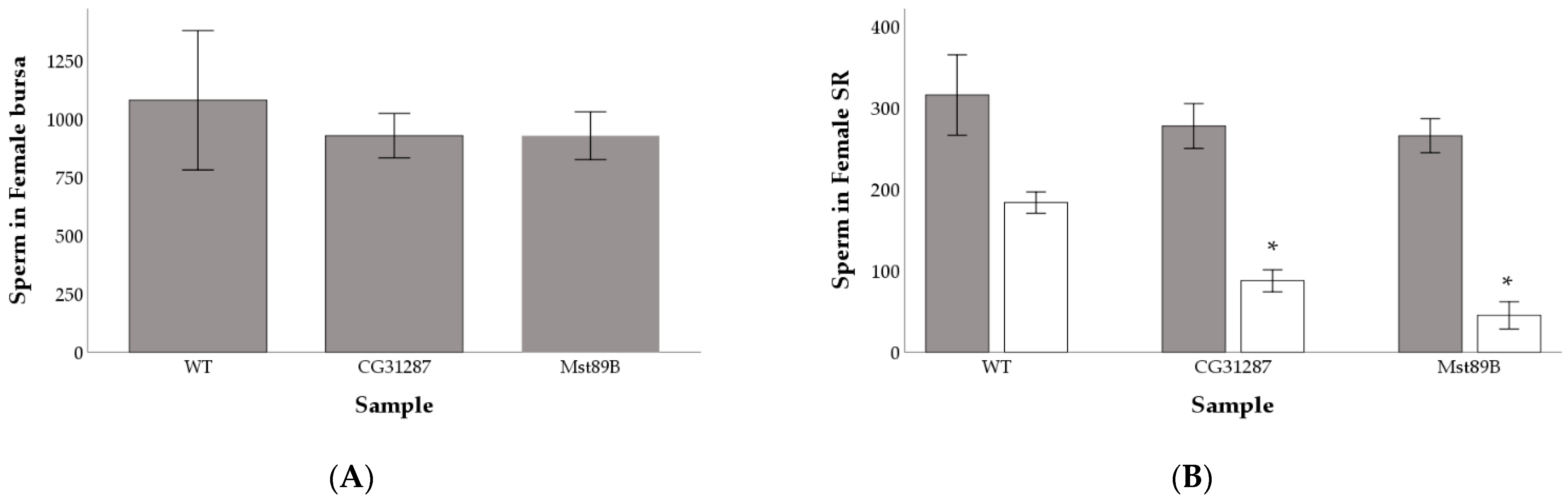
3.2. Knockdown of Mst89B and CG31287 Does Not Affect Sperm Transfer but Impairs Long-Term Sperm Storage
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lewontin, R.C. The Genetic Basis of Evolutionary Change; Columbia University Press: New York, NY, USA, 1974; Volume 560. [Google Scholar]
- Gubala, A.M.; Schmitz, J.F.; Kearns, M.J.; Vinh, T.T.; Bornberg-Bauer, E.; Wolfner, M.F.; Findlay, G.D. The goddard and saturn genes are essential for Drosophila male fertility and may have arisen de novo. Mol. Biol. Evol. 2017, 34, 1066–1082. [Google Scholar] [CrossRef]
- Neubaum, D.M.; Wolfner, M.F. Mated Drosophila melanogaster females require a seminal fluid protein, Acp36DE, to store sperm efficiently. Genetics 1999, 153, 845–857. [Google Scholar] [PubMed]
- Qazi, M.C.B.; Wolfner, M.F. An early role for the Drosophila melanogaster male seminal protein Acp36DE in female sperm storage. J. Exp. Biol. 2003, 206, 3521–3528. [Google Scholar] [CrossRef] [PubMed]
- Hansen, J.M.; Chavez, D.R.; Stanfield, G.M. COMP-1 promotes competitive advantage of nematode sperm. Elife 2015, 4, e05423. [Google Scholar] [CrossRef] [PubMed]
- Ram, K.R.; Wolfner, M.F. Sustained post-mating response in Drosophila melanogaster requires multiple seminal fluid proteins. PLoS Genet. 2007, 3, e238. [Google Scholar] [CrossRef]
- Ohsako, T.; Yamamoto, M.-T. Sperm of the wasted mutant are wasted when females utilize the stored sperm in Drosophila melanogaster. Genes Genet. Syst. 2011, 86, 97–108. [Google Scholar] [CrossRef]
- Yeh, S.-D.; Do, T.; Chan, C.; Cordova, A.; Carranza, F.; Yamamoto, E.A.; Abbassi, M.; Gandasetiawan, K.A.; Librado, P.; Damia, E. Functional evidence that a recently evolved Drosophila sperm-specific gene boosts sperm competition. Proc. Natl. Acad. Sci. USA 2012, 109, 2043–2048. [Google Scholar] [CrossRef]
- Kawano, N.; Araki, N.; Yoshida, K.; Hibino, T.; Ohnami, N.; Makino, M.; Kanai, S.; Hasuwa, H.; Yoshida, M.; Miyado, K. Seminal vesicle protein SVS2 is required for sperm survival in the uterus. Proc. Natl. Acad. Sci. USA 2014, 111, 4145–4150. [Google Scholar] [CrossRef]
- Dosselli, R.; Grassl, J.; den Boer, S.P.A.; Kratz, M.; Moran, J.M.; Boomsma, J.J.; Baer, B. Protein-level interactions as mediators of sexual conflict in ants. Mol. Cell. Proteom. 2019, 18, S34–S45. [Google Scholar] [CrossRef]
- Civetta, A.; Ranz, J.M. Genetic factors influencing sperm competition. Front. Genet. 2019, 10, 820. [Google Scholar] [CrossRef]
- Lefevre, G., Jr.; Jonsson, U.B. Sperm transfer, storage, displacement, and utilization in Drosophila melanogaster. Genetics 1962, 47, 1719. [Google Scholar] [CrossRef] [PubMed]
- Adams, E.M.; Wolfner, M.F. Seminal proteins but not sperm induce morphological changes in the Drosophila melanogaster female reproductive tract during sperm storage. J. Insect Physiol. 2007, 53, 319–331. [Google Scholar] [CrossRef] [PubMed]
- Avila, F.W.; Wolfner, M.F. Acp36DE is required for uterine conformational changes in mated Drosophila females. Proc. Natl. Acad. Sci. USA 2009, 106, 15796–15800. [Google Scholar] [CrossRef] [PubMed]
- Fowler, G.L. Some aspects of the reproductive biology of Drosophila: Sperm transfer, sperm storage, and sperm utilization. In Advances in Genetics; Elsevier: Amsterdam, The Netherlands, 1973; Volume 17, pp. 293–360. [Google Scholar]
- Manier, M.K.; Belote, J.M.; Berben, K.S.; Novikov, D.; Stuart, W.T.; Pitnick, S. Resolving mechanisms of competitive fertilization success in Drosophila melanogaster. Science 2010, 328, 354–357. [Google Scholar] [CrossRef]
- Lee, K.-M.; Daubnerová, I.; Isaac, R.E.; Zhang, C.; Choi, S.; Chung, J.; Kim, Y.-J. A neuronal pathway that controls sperm ejection and storage in female Drosophila. Curr. Biol. 2015, 25, 790–797. [Google Scholar] [CrossRef]
- Gilbert, D.G. Ejaculate esterase 6 and initial sperm use by female Drosophila melanogaster. J. Insect Physiol. 1981, 27, 641–650. [Google Scholar] [CrossRef]
- Bloch Qazi, M.C.; Wolfner, M.F. Emergence of sperm from female storage sites has egg-influenced and egg-independent phases in Drosophila melanogaster. Biol. Lett. 2006, 2, 128–130. [Google Scholar] [CrossRef][Green Version]
- Pitnick, S.; Wolfner, M.F.; Suarez, S.S. Ejaculate–female and sperm–female interactions. In Sperm Biology; Elsevier: Amsterdam, The Netherlands, 2009; pp. 247–304. [Google Scholar]
- Lüpold, S.; Pitnick, S. Sperm form and function: What do we know about the role of sexual selection? Reproduction 2018, 155, R229–R243. [Google Scholar] [CrossRef]
- Schnakenberg, S.L.; Matias, W.R.; Siegal, M.L. Sperm-storage defects and live birth in Drosophila females lacking spermathecal secretory cells. PLoS Biol. 2011, 9, e1001192. [Google Scholar] [CrossRef]
- Avila, F.W.; Qazi, M.C.B.; Rubinstein, C.D.; Wolfner, M.F. A requirement for the neuromodulators octopamine and tyramine in Drosophila melanogaster female sperm storage. Proc. Natl. Acad. Sci. USA 2012, 109, 4562–4567. [Google Scholar] [CrossRef]
- Chen, D.S.; Delbare, S.Y.N.; White, S.L.; Sitnik, J.; Chatterjee, M.; DoBell, E.; Weiss, O.; Clark, A.G.; Wolfner, M.F. Female genetic contributions to sperm competition in Drosophila melanogaster. Genetics 2019, 212, 789–800. [Google Scholar] [CrossRef] [PubMed]
- Levesque, L.; Brouwers, B.; Sundararajan, V.; Civetta, A. Third chromosome candidate genes for conspecific sperm precedence between D. simulans and D. mauritiana. BMC Genet. 2010, 11, 21. [Google Scholar] [CrossRef] [PubMed]
- Civetta, A.; Finn, S. Do candidate genes mediating conspecific sperm precedence affect sperm competitive ability within species? A test case in Drosophila. G3 Genes Genomes Genet. 2014, 4, 1701–1707. [Google Scholar] [CrossRef][Green Version]
- White-Cooper, H. Tissue, cell type and stage-specific ectopic gene expression and RNAi induction in the Drosophila testis. Spermatogenesis 2012, 2, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.-Q.; Markstein, M.; Binari, R.; Pfeiffer, B.; Liu, L.-P.; Villalta, C.; Booker, M.; Perkins, L.; Perrimon, N. Vector and parameters for targeted transgenic RNA interference in Drosophila melanogaster. Nat. Methods 2008, 5, 49–51. [Google Scholar] [CrossRef]
- Barron, A.B. Anaesthetising Drosophila for behavioural studies. J. Insect Physiol. 2000, 46, 439–442. [Google Scholar] [CrossRef]
- Manier, M.K.; Lüpold, S.; Pitnick, S.; Starmer, W.T. An analytical framework for estimating fertilization bias and the fertilization set from multiple sperm-storage organs. Am. Nat. 2013, 182, 552–561. [Google Scholar] [CrossRef]
- Pitnick, S.; Marrow, T.; Spicer, G.S. Evolution of multiple kinds of female sperm-storage organs in Drosophila. Evolution 1999, 53, 1804–1822. [Google Scholar] [CrossRef]
- Halligan, D.L.; Keightley, P.D. Ubiquitous selective constraints in the Drosophila genome revealed by a genome-wide interspecies comparison. Genome Res. 2006, 16, 875–884. [Google Scholar] [CrossRef]
- Karak, S.; Jacobs, J.S.; Kittelmann, M.; Spalthoff, C.; Katana, R.; Sivan-Loukianova, E.; Schon, M.A.; Kernan, M.J.; Eberl, D.F.; Göpfert, M.C. Diverse roles of axonemal dyneins in Drosophila auditory neuron function and mechanical amplification in hearing. Sci. Rep. 2015, 5, 17085. [Google Scholar] [CrossRef]
- Köttgen, M.; Hofherr, A.; Li, W.; Chu, K.; Cook, S.; Montell, C.; Watnick, T. Drosophila sperm swim backwards in the female reproductive tract and are activated via TRPP2 ion channels. PLoS ONE 2011, 6, e20031. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Lu, X. Drosophila sperm motility in the reproductive tract. Biol. Reprod. 2011, 84, 1005–1015. [Google Scholar] [CrossRef] [PubMed]
- Tomaru, M.; Ohsako, T.; Watanabe, M.; Juni, N.; Matsubayashi, H.; Sato, H.; Takahashi, A.; Yamamoto, M.-T. Severe fertility effects of sheepish sperm caused by failure to enter female sperm storage organs in Drosophila melanogaster. G3 Genes Genomes Genet. 2018, 8, 149–160. [Google Scholar] [CrossRef] [PubMed]
- Herndon, L.A.; Wolfner, M.F. A Drosophila seminal fluid protein, Acp26Aa, stimulates egg laying in females for 1 day after mating. Proc. Natl. Acad. Sci. USA 1995, 92, 10114–10118. [Google Scholar] [CrossRef] [PubMed]
- Heifetz, Y.; Lung, O.; Frongillo, E.A., Jr.; Wolfner, M.F. The Drosophila seminal fluid protein Acp26Aa stimulates release of oocytes by the ovary. Curr. Biol. 2000, 10, 99–102. [Google Scholar] [CrossRef]
- Ohsako, T.; Hirai, K.; Yamamoto, M.-T. The Drosophila misfire gene has an essential role in sperm activation during fertilization. Genes Genet. Syst. 2003, 78, 253–266. [Google Scholar] [CrossRef][Green Version]
- Heigwer, F.; Port, F.; Boutros, M. RNA interference (RNAi) screening in Drosophila. Genetics 2018, 208, 853–874. [Google Scholar] [CrossRef]
- Hu, Y.; Roesel, C.; Flockhart, I.; Perkins, L.; Perrimon, N.; Mohr, S.E. UP-TORR: Online tool for accurate and Up-to-Date annotation of RNAi Reagents. Genetics 2013, 195, 37–45. [Google Scholar] [CrossRef]
- Qian, Y.; Ng, C.L.; Schulz, C. CSN maintains the germline cellular microenvironment and controls the level of stem cell genes via distinct CRLs in testes of Drosophila melanogaster. Dev. Biol. 2015, 398, 68–79. [Google Scholar] [CrossRef]
- Hoffmann, M.; Bellance, N.; Rossignol, R.; Koopman, W.J.H.; Willems, P.H.G.M.; Mayatepek, E.; Bossinger, O.; Distelmaier, F. C. elegans ATAD-3 is essential for mitochondrial activity and development. PLoS ONE 2009, 4, e7644. [Google Scholar] [CrossRef]
- Harel, T.; Yoon, W.H.; Garone, C.; Gu, S.; Coban-Akdemir, Z.; Eldomery, M.K.; Posey, J.E.; Jhangiani, S.N.; Rosenfeld, J.A.; Cho, M.T. Recurrent de novo and biallelic variation of ATAD3A, encoding a mitochondrial membrane protein, results in distinct neurological syndromes. Am. J. Hum. Genet. 2016, 99, 831–845. [Google Scholar] [CrossRef] [PubMed]
- Palgi, M.; Lindström, R.; Peränen, J.; Piepponen, T.P.; Saarma, M.; Heino, T.I. Evidence that DmMANF is an invertebrate neurotrophic factor supporting dopaminergic neurons. Proc. Natl. Acad. Sci. USA 2009, 106, 2429–2434. [Google Scholar] [CrossRef] [PubMed]
- Lindström, R.; Lindholm, P.; Kallijärvi, J.; Yu, L.; Piepponen, T.P.; Arumäe, U.; Saarma, M.; Heino, T.I. Characterization of the structural and functional determinants of MANF/CDNF in Drosophila in vivo model. PLoS ONE 2013, 8, e73928. [Google Scholar] [CrossRef] [PubMed]
- Lindström, R.; Lindholm, P.; Palgi, M.; Saarma, M.; Heino, T.I. In vivo screening reveals interactions between Drosophila Manf and genes involved in the mitochondria and the ubiquinone synthesis pathway. BMC Genet. 2017, 18, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Tokuyasu, K.T. Dynamics of spermiogenesis in Drosophila melanogaster. VI. Significance of “onion” nebenkern formation. J. Ultrastruct. Res. 1975, 53, 93–112. [Google Scholar] [CrossRef]
- Noguchi, T.; Koizumi, M.; Hayashi, S. Sustained elongation of sperm tail promoted by local remodeling of giant mitochondria in Drosophila. Curr. Biol. 2011, 21, 805–814. [Google Scholar] [CrossRef] [PubMed]
- Insco, M.L.; Bailey, A.S.; Kim, J.; Olivares, G.H.; Wapinski, O.L.; Tam, C.H.; Fuller, M.T. A self-limiting switch based on translational control regulates the transition from proliferation to differentiation in an adult stem cell lineage. Cell Stem Cell 2012, 11, 689–700. [Google Scholar] [CrossRef]
- Stebbings, L.; Grimes, B.R.; Bownes, M. A testis-specifically expressed gene is embedded within a cluster of maternally expressed genes at 89B in Drosophila melanogaster. Dev. Genes Evol. 1998, 208, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Jayaswal, V.; Jimenez, J.; Magie, R.; Nguyen, K.; Clifton, B.; Yeh, S.; Ranz, J.M. A species-specific multigene family mediates differential sperm displacement in Drosophila melanogaster. Evolution 2018, 72, 399–403. [Google Scholar] [CrossRef]
- Gao, Z.; Ruden, D.M.; Lu, X. PKD2 cation channel is required for directional sperm movement and male fertility. Curr. Biol. 2003, 13, 2175–2178. [Google Scholar] [CrossRef]
- Avila, F.W.; Ram, K.R.; Qazi, M.C.B.; Wolfner, M.F. Sex peptide is required for the efficient release of stored sperm in mated Drosophila females. Genetics 2010, 186, 595–600. [Google Scholar] [CrossRef] [PubMed]
- Wong, A.; Albright, S.N.; Giebel, J.D.; Ram, K.R.; Ji, S.; Fiumera, A.C.; Wolfner, M.F. A role for Acp29AB, a predicted seminal fluid lectin, in female sperm storage in Drosophila melanogaster. Genetics 2008, 180, 921–931. [Google Scholar] [CrossRef][Green Version]
- LaFlamme, B.A.; Ram, K.R.; Wolfner, M.F. The Drosophila melanogaster seminal fluid protease “seminase” regulates proteolytic and post-mating reproductive processes. PLoS Genet. 2012, 8, e1002435. [Google Scholar] [CrossRef] [PubMed]
- Findlay, G.D.; Sitnik, J.L.; Wang, W.; Aquadro, C.F.; Clark, N.L.; Wolfner, M.F. Evolutionary rate covariation identifies new members of a protein network required for Drosophila melanogaster female post-mating responses. PLoS Genet. 2014, 10, e1004108. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Buehner, N.A.; Lin, H.; Baranowski, K.J.; Findlay, G.D.; Wolfner, M.F. Long-term interaction between Drosophila sperm and sex peptide is mediated by other seminal proteins that bind only transiently to sperm. Insect Biochem. Mol. Biol. 2018, 102, 43–51. [Google Scholar] [CrossRef] [PubMed]

| Treatment | N | P/L | A/P |
|---|---|---|---|
| Control | 96 | 0.89 | 0.93 |
| CG31287 | 86 | 0.84 | 0.99 |
| Mst89B | 44 | 0.94 | 0.97 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grewal, G.; Patlar, B.; Civetta, A. Expression of Mst89B and CG31287 is Needed for Effective Sperm Storage and Egg Fertilization in Drosophila. Cells 2021, 10, 289. https://doi.org/10.3390/cells10020289
Grewal G, Patlar B, Civetta A. Expression of Mst89B and CG31287 is Needed for Effective Sperm Storage and Egg Fertilization in Drosophila. Cells. 2021; 10(2):289. https://doi.org/10.3390/cells10020289
Chicago/Turabian StyleGrewal, Gurman, Bahar Patlar, and Alberto Civetta. 2021. "Expression of Mst89B and CG31287 is Needed for Effective Sperm Storage and Egg Fertilization in Drosophila" Cells 10, no. 2: 289. https://doi.org/10.3390/cells10020289
APA StyleGrewal, G., Patlar, B., & Civetta, A. (2021). Expression of Mst89B and CG31287 is Needed for Effective Sperm Storage and Egg Fertilization in Drosophila. Cells, 10(2), 289. https://doi.org/10.3390/cells10020289







