Matrix Metalloproteinases Inhibition by Doxycycline Rescues Extracellular Matrix Organization and Partly Reverts Myofibroblast Differentiation in Hypermobile Ehlers-Danlos Syndrome Dermal Fibroblasts: A Potential Therapeutic Target?
Abstract
:1. Introduction
2. Materials and Methods
2.1. Ethical Compliance
2.2. Cell Cultures and Preparation of Conditioned Media (CM)
2.3. Immunofluorescence Microscopy (IF)
2.4. Western Blotting (WB)
2.5. Label-Free Liquid Chromatography with Tandem Mass Spectrometry (LC–MS/MS)
2.6. Functional Enrichment Analyses
2.7. Doxycycline Treatment
3. Results
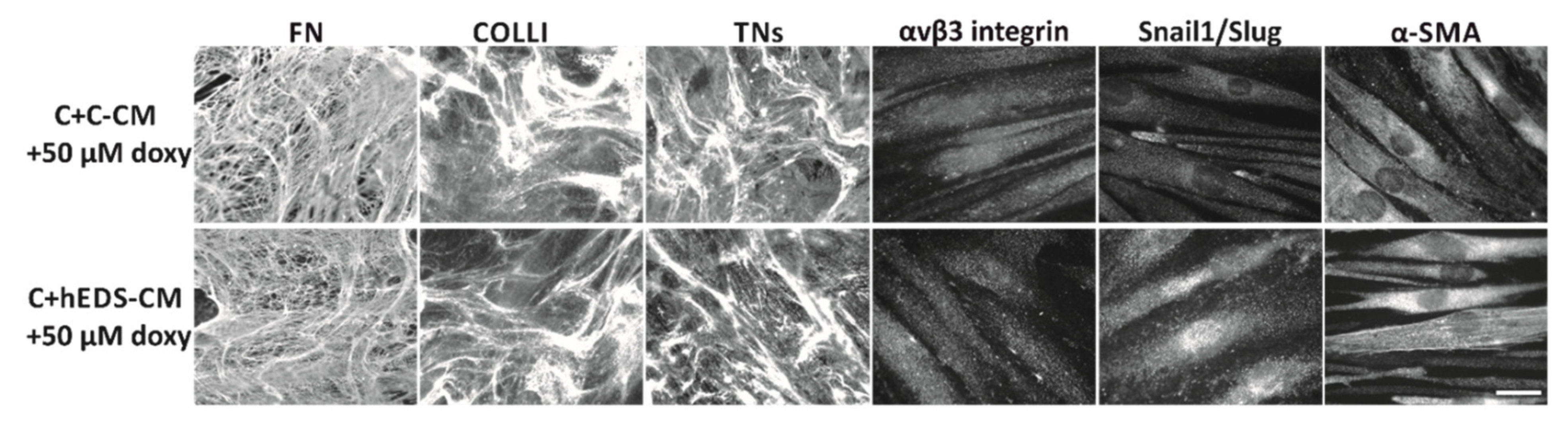
3.1. hEDS-CM Affects ECM Organization of Control Dermal Fibroblasts and Induces Myofibroblast Differentiation
3.2. Secretome Analysis of hEDS-CM
3.3. Doxycycline Treatment Rescues ECM Organization and Partly Reverts the Myofibroblast-like Phenotype of hEDS Cells
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AA | Ascorbic acid |
| α-SMA | Alpha smooth muscle actin |
| COLLs | Collagens |
| COLLI-fs | COLLI-fragments |
| CM | Conditioned media |
| C-CM | Control cells-derived conditioned media |
| DAMPs | Damage-associated molecular patterns |
| DEPs | Differentially expressed proteins |
| doxy | Doxycycline |
| ER | Endoplasmic reticulum |
| MEM | Earle’s modified Eagle medium |
| ECM | Extracellular matrix |
| EVs | Extracellular vesicles |
| FDR | False discovery rate |
| FN | Fibronectin |
| FN-fs | FN-fragments |
| GO | Gene Ontology |
| HCTD | Heritable connective tissue disorder |
| hEDS | Hypermobile Ehlers-Danlos syndrome |
| hEDS-CM | hEDS cells-derived conditioned media |
| IF | Immunofluorescence microscopy |
| JHM | Joint hypermobility |
| LC-MS/MS | Liquid chromatography with tandem mass spectrometry |
| MMPs | Matrix metalloproteinases |
| OA | Osteoarthritis |
| RA | Rheumatoid arthritis |
| TNs | Tenascins |
| TNs-fs | TNs-fragments |
| TIMPs | Tissue inhibitor of metalloproteinases |
| WB | Western blotting |
References
- Malfait, F.; Francomano, C.; Byers, P.; Belmont, J.; Berglund, B.; Black, J.; Bloom, L.; Bowen, J.M.; Brady, A.F.; Burrows, N.P.; et al. The 2017 international classification of the Ehlers–Danlos syndromes. Am. J. Med. Genet. Part C Semin. Med. Genet. 2017, 175, 8–26. [Google Scholar] [CrossRef] [Green Version]
- Tinkle, B.; Castori, M.; Berglund, B.; Cohen, H.; Grahame, R.; Kazkaz, H.; Levy, H. Hypermobile Ehlers–Danlos syndrome (a.k.a. Ehlers–Danlos syndrome Type III and Ehlers–Danlos syndrome hypermobility type): Clinical description and natural history. Am. J. Med. Genet. Part C Semin. Med. Genet. 2017, 175, 48–69. [Google Scholar] [CrossRef] [Green Version]
- Chiarelli, N.; Carini, G.; Zoppi, N.; Dordoni, C.; Ritelli, M.; Venturini, M.; Castori, M.; Colombi, M. Transcriptome-wide expression profiling in skin fibroblasts of patients with joint hypermobility syndrome/ehlers-danlos syndrome hypermobility type. PLoS ONE 2016, 11, e161347. [Google Scholar] [CrossRef] [Green Version]
- Zoppi, N.; Chiarelli, N.; Binetti, S.; Ritelli, M.; Colombi, M. Dermal fibroblast-to-myofibroblast transition sustained by αvß3 integrin-ILK-Snail1/Slug signaling is a common feature for hypermobile Ehlers-Danlos syndrome and hypermobility spectrum disorders. Biochim. Biophys. Acta-Mol. Basis Dis. 2018, 1864, 1010–1023. [Google Scholar] [CrossRef]
- Chiarelli, N.; Zoppi, N.; Ritelli, M.; Venturini, M.; Capitanio, D.; Gelfi, C.; Colombi, M. Biological insights in the pathogenesis of hypermobile Ehlers-Danlos syndrome from proteome profiling of patients’ dermal myofibroblasts. Biochim. Biophys. Acta-Mol. Basis Dis. 2021, 1867, 166051. [Google Scholar] [CrossRef] [PubMed]
- Holmberg, C.; Ghesquiè, B.; Impens, F.; Gevaert, K.; Dinesh Kumar, J.; Cash, N.; Kandola, S.; Hegyi, P.; Wang, T.C.; Dockray, G.J.; et al. Mapping Proteolytic Processing in the Secretome of Gastric Cancer-Associated Myofibroblasts Reveals Activation of MMP-1, MMP-2, and MMP-3. J. Proteome Res. 2013, 12, 3413–3422. [Google Scholar] [CrossRef]
- Klingberg, F.; Hinz, B.; White, E.S. The myofibroblast matrix: Implications for tissue repair andfibrosis. J. Pathol. 2013, 229, 298–309. [Google Scholar] [CrossRef] [Green Version]
- D’Urso, M.; Kurniawan, N.A. Mechanical and Physical Regulation of Fibroblast–Myofibroblast Transition: From Cellular Mechanoresponse to Tissue Pathology. Front. Bioeng. Biotechnol. 2020, 8, 1459. [Google Scholar] [CrossRef]
- Rogers, N.K.; Clements, D.; Dongre, A.; Harrison, T.W.; Shaw, D.; Johnson, S.R. Extra-cellular matrix proteins induce matrix metalloproteinase-1 (MMP-1) activity and increase airway smooth muscle contraction in asthma. PLoS ONE 2014, 9, e90565. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ciechomska, M.; O’Reilly, S.; Suwara, M.; Bogunia-Kubik, K.; Van Laar, J.M. MiR-29a reduces TIMP-1 production by dermal fibroblasts via targeting TGF-β activated kinase 1 binding protein 1, implications for systemic sclerosis. PLoS ONE 2014, 9, e115596. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maarof, M.; Lokanathan, Y.; Ruszymah, H.I.; Saim, A.; Chowdhury, S.R. Proteomic Analysis of Human Dermal Fibroblast Conditioned Medium (DFCM). Protein J. 2018, 37, 589–607. [Google Scholar] [CrossRef] [PubMed]
- Brown, K.J.; Seol, H.; Pillai, D.K.; Sankoorikal, B.J.; Formolo, C.A.; Mac, J.; Edwards, N.J.; Rose, M.C.; Hathout, Y. The human secretome atlas initiative: Implications in health and disease conditions. Biochim. Biophys. Acta-Proteins Proteom. 2013, 1834, 2454–2461. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mathivanan, S.; Ji, H.; Simpson, R.J. Exosomes: Extracellular organelles important in intercellular communication. J. Proteom. 2010, 73, 1907–1920. [Google Scholar] [CrossRef]
- Kendall, R.T.; Renaud, L.; Baatz, J.E.; Malaab, M.; Nguyen, X.-X.; Feghali-Bostwick, C.A. Systemic sclerosis biomarkers detection in the secretome of TGFβ1-activated primary human lung fibroblasts. J. Proteom. 2021, 242, 104243. [Google Scholar] [CrossRef] [PubMed]
- Lipphardt, M.; Song, J.W.; Matsumoto, K.; Dadafarin, S.; Dihazi, H.; Müller, G.; Goligorsky, M.S. The third path of tubulointerstitial fibrosis: Aberrant endothelial secretome. Kidney Int. 2017, 92, 558–568. [Google Scholar] [CrossRef] [PubMed]
- Del Galdo, F.; Shaw, M.A.; Jimenez, S.A. Proteomic Analysis Identification of a Pattern of Shared Alterations in the Secretome of Dermal Fibroblasts from Systemic Sclerosis and Nephrogenic Systemic Fibrosis. Am. J. Pathol. 2010, 177, 1638–1646. [Google Scholar] [CrossRef]
- Thio, C.L.-P.; Yusof, R.; Ashrafzadeh, A.; Bahari, S.; Abdul-Rahman, P.S.; Karsani, S.A. Differential Analysis of the Secretome of WRL68 Cells Infected with the Chikungunya Virus. PLoS ONE 2015, 10, e0129033. [Google Scholar] [CrossRef] [Green Version]
- Huang, D.W.; Sherman, B.T.; Tan, Q.; Kir, J.; Liu, D.; Bryant, D.; Guo, Y.; Stephens, R.; Baseler, M.W.; Lane, H.C.; et al. DAVID Bioinformatics Resources: Expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 2007, 35. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Muruganujan, A.; Ebert, D.; Huang, X.; Thomas, P.D. PANTHER version 14: More genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 2019, 47, D419–D426. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Platt, B.N.; Jacobs, C.A.; Conley, C.E.W.; Stone, A.V. Tetracycline use in treating osteoarthritis: A systematic review. Inflamm. Res. 2021, 70, 249–259. [Google Scholar] [CrossRef] [PubMed]
- Theocharis, A.D.; Manou, D.; Karamanos, N.K. The extracellular matrix as a multitasking player in disease. FEBS J. 2019, 286, 2830–2869. [Google Scholar] [CrossRef] [Green Version]
- Lamandé, S.R.; Bateman, J.F. Genetic Disorders of the Extracellular Matrix. Anat. Rec. 2020, 303, 1527–1542. [Google Scholar] [CrossRef]
- Bateman, J.F.; Boot-Handford, R.P.; Lamandé, S.R. Genetic diseases of connective tissues: Cellular and extracellular effects of ECM mutations. Nat. Rev. Genet. 2009, 10, 173–183. [Google Scholar] [CrossRef] [PubMed]
- Lu, P.; Takai, K.; Weaver, V.M.; Werb, Z. Extracellular Matrix degradation and remodeling in development and disease. Cold Spring Harb. Perspect. Biol. 2011, 3. [Google Scholar] [CrossRef] [PubMed]
- Iozzo, R.V.; Gubbiotti, M.A. Extracellular matrix: The driving force of mammalian diseases. Matrix Biol. 2018, 71–72, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Karamanos, N.K.; Theocharis, A.D.; Neill, T.; Iozzo, R.V. Matrix modeling and remodeling: A biological interplay regulating tissue homeostasis and diseases. Matrix Biol. 2019, 75–76, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Martin, A. An acquired or heritable connective tissue disorder? A review of hypermobile Ehlers Danlos Syndrome. Eur. J. Med. Genet. 2019, 62, 103672. [Google Scholar] [CrossRef]
- Gensemer, C.; Burks, R.; Kautz, S.; Judge, D.P.; Lavallee, M.; Norris, R.A. Hypermobile Ehlers-Danlos syndromes: Complex phenotypes, challenging diagnoses, and poorly understood causes. Dev. Dyn. 2021, 250, 318–344. [Google Scholar] [CrossRef] [PubMed]
- Scicluna, K.; Formosa, M.M.; Farrugia, R.; Borg, I. Hypermobile Ehlers–Danlos syndrome: A review and a critical appraisal of published genetic research to date. Clin. Genet. 2021. [Google Scholar] [CrossRef]
- Chiarelli, N.; Ritelli, M.; Zoppi, N.; Colombi, M. Cellular and molecular mechanisms in the pathogenesis of classical, vascular, and hypermobile ehlers‒danlos syndromes. Genes 2019, 10, 609. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zoppi, N.; Chiarelli, N.; Ritelli, M.; Colombi, M. Multifaced Roles of the αvβ3 Integrin in Ehlers–Danlos and Arterial Tortuosity Syndromes’ Dermal Fibroblasts. Int. J. Mol. Sci. 2018, 19, 982. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roh, J.S.; Sohn, D.H. Damage-associated molecular patterns in inflammatory diseases. Immune Netw. 2018, 18, e27. [Google Scholar] [CrossRef] [PubMed]
- Midwood, K.; Sacre, S.; Piccinini, A.M.; Inglis, J.; Trebaul, A.; Chan, E.; Drexler, S.; Sofat, N.; Kashiwagi, M.; Orend, G.; et al. Tenascin-C is an endogenous activator of Toll-like receptor 4 that is essential for maintaining inflammation in arthritic joint disease. Nat. Med. 2009, 15, 774–780. [Google Scholar] [CrossRef] [PubMed]
- Zheng, M.; Ambesi, A.; McKeown-Longo, P.J. Role of TLR4 Receptor Complex in the Regulation of the Innate Immune Response by Fibronectin. Cells 2020, 9, 216. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anders, H.J.; Schaefer, L. Beyond tissue injury—Damage-associated molecular patterns, toll-like receptors, and inflammasomes also drive regeneration and fibrosis. J. Am. Soc. Nephrol. 2014, 25, 1387–1400. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, M.; Martel-Pelletier, J.; Lajeunesse, D.; Pelletier, J.P.; Fahmi, H. Role of proinflammatory cytokines in the pathophysiology of osteoarthritis. Nat. Rev. Rheumatol. 2011, 7, 33–42. [Google Scholar] [CrossRef]
- Hwang, H.S.; Park, S.J.; Cheon, E.J.; Lee, M.H.; Kim, H.A. Fibronectin fragment-induced expression of matrix metalloproteinases is mediated by MyD88-dependent TLR-2 signaling pathway in human chondrocytes. Arthritis Res. Ther. 2015, 17. [Google Scholar] [CrossRef] [Green Version]
- Pérez-García, S.; Carrión, M.; Gutiérrez-Cañas, I.; Villanueva-Romero, R.; Castro, D.; Martínez, C.; González-Álvaro, I.; Blanco, F.J.; Juarranz, Y.; Gomariz, R.P. Profile of Matrix-Remodeling Proteinases in Osteoarthritis: Impact of Fibronectin. Cells 2019, 9, 40. [Google Scholar] [CrossRef] [Green Version]
- Zack, M.D.; Arner, E.C.; Anglin, C.P.; Alston, J.T.; Malfait, A.M.; Tortorella, M.D. Identification of fibronectin neoepitopes present in human osteoarthritic cartilage. Arthritis Rheum. 2006, 54, 2912–2922. [Google Scholar] [CrossRef]
- Pulai, J.I.; Chen, H.; Im, H.-J.; Kumar, S.; Hanning, C.; Hegde, P.S.; Loeser, R.F. NF-κB Mediates the Stimulation of Cytokine and Chemokine Expression by Human Articular Chondrocytes in Response to Fibronectin Fragments. J. Immunol. 2005, 174, 5781–5788. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Homandberg, G.A.; Wen, C.; Hui, F. Cartilage damaging activities of fibronectin fragments derived from cartilage and synovial fluid. Osteoarthr. Cartil. 1998, 6, 231–244. [Google Scholar] [CrossRef] [Green Version]
- Reed, K.S.M.; Ulici, V.; Kim, C.; Chubinskaya, S.; Loeser, R.F.; Phanstiel, D.H. Transcriptional response of human articular chondrocytes treated with fibronectin fragments: An in vitro model of the osteoarthritis phenotype. Osteoarthr. Cartil. 2021, 29, 235–247. [Google Scholar] [CrossRef]
- Pérez-García, S.; Calamia, V.; Hermida-Gómez, T.; Gutiérrez-Cañas, I.; Carrión, M.; Villanueva-Romero, R.; Castro, D.; Martínez, C.; Juarranz, Y.; Blanco, F.J.; et al. Proteomic analysis of synovial fibroblasts and articular chondrocytes co-cultures reveals valuable vip-modulated inflammatory and degradative proteins in osteoarthritis. Int. J. Mol. Sci. 2021, 22, 6441. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, M.; Yoshida, T.; Sudo, A. Tenascin-C in Osteoarthritis and Rheumatoid Arthritis. Front. Immunol. 2020, 11, 2585. [Google Scholar] [CrossRef] [PubMed]
- Patel, L.; Sun, W.; Glasson, S.S.; Morris, E.A.; Flannery, C.R.; Chockalingam, P.S. Tenascin-C induces inflammatory mediators and matrix degradation in osteoarthritic cartilage. BMC Musculoskelet. Disord. 2011, 12, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Fichter, M.; Körner, U.; Schömburg, J.; Jennings, L.; Cole, A.A.; Mollenhauer, J. Collagen degradation products modulate matrix metalloproteinase expression in cultured articular chondrocytes. J. Orthop. Res. 2006, 24, 63–70. [Google Scholar] [CrossRef]
- Klatt, A.R.; Paul-Klausch, B.; Klinger, C.; Kühn, C.; Renno, J.H.; Banerjee, M.; Malchau, G.; Wielckens, K. A critical role for collagen II in cartilage matrix degradation: Collagen II induces pro-inflammatory cytokines and MMPS in primary human chondrocytes. J. Orthop. Res. 2009, 27, 65–70. [Google Scholar] [CrossRef] [PubMed]
- Sofat, N. Analysing the role of endogenous matrix molecules in the development of osteoarthritis. Int. J. Exp. Pathol. 2009, 90, 463–479. [Google Scholar] [CrossRef]
- Capello, M.; Ferri-Borgogno, S.; Cappello, P.; Novelli, F. α-enolase: A promising therapeutic and diagnostic tumor target. FEBS J. 2011, 278, 1064–1074. [Google Scholar] [CrossRef] [Green Version]
- Sharma, A.; Khan, R.; Gupta, N.; Sharma, A.; Zaheer, M.S.; Abbas, M.; Khan, S.A. Acute phase reactant, Pentraxin 3, as a novel marker for the diagnosis of rheumatoid arthritis. Clin. Chim. Acta 2018, 480, 65–70. [Google Scholar] [CrossRef]
- Pilewski, J.M.; Liu, L.; Henry, A.C.; Knauer, A.V.; Feghali-Bostwick, C.A. Insulin-like growth factor binding proteins 3 and 5 are overexpressed in idiopathic pulmonary fibrosis and contribute to extracellular matrix deposition. Am. J. Pathol. 2005, 166, 399–407. [Google Scholar] [CrossRef] [Green Version]
- Tumbarello, D.A.; Temple, J.; Brenton, J.D. ß3 integrin modulates transforming growth factor beta induced (TGFBI) function and paclitaxel response in ovarian cancer cells. Mol. Cancer 2012, 11, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Albacete-Albacete, L. Extracellular Vesicles: An Emerging Mechanism Governing the Secretion and Biological Roles of Tenascin-C. Front. Immunol. 2021, 12, 671485. [Google Scholar] [CrossRef]
- Zanotti, S.; Gibertini, S.; Blasevich, F.; Bragato, C.; Ruggieri, A.; Saredi, S.; Fabbri, M.; Bernasconi, P.; Maggi, L.; Mantegazza, R.; et al. Exosomes and exosomal miRNAs from muscle-derived fibroblasts promote skeletal muscle fibrosis. Matrix Biol. 2018, 74, 77–100. [Google Scholar] [CrossRef] [PubMed]
- Asghar, S.; Litherland, G.J.; Lockhart, J.C.; Goodyear, C.S.; Crilly, A. Exosomes in intercellular communication and implications for osteoarthritis. Rheumatology 2020, 59, 57–68. [Google Scholar] [CrossRef]
- Wermuth, P.J.; Piera-Velazquez, S.; Jimenez, S.A. Exosomes isolated from serum of systemic sclerosis patients display alterations in their content of profibrotic and antifibrotic microRNA and induce a profibrotic phenotype in cultured normal dermal fibroblasts. Clin. Exp. Rheumatol. 2017, 35, S21–S30. [Google Scholar]
- Siller, S.S.; Broadie, K. Matrix metalloproteinases and minocycline: Therapeutic avenues for fragile X syndrome. Neural Plast. 2012, 2012, 124548. [Google Scholar] [CrossRef] [Green Version]
- Dziembowska, M.; Pretto, D.I.; Janusz, A.; Kaczmarek, L.; Leigh, M.J.; Gabriel, N.; Durbin-Johnson, B.; Hagerman, R.J.; Tassone, F. High MMP-9 activity levels in fragile X syndrome are lowered by minocycline. Am. J. Med. Genet. Part A 2013, 161, 1897–1903. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Tay, F.R.; Yiu, C.K.Y. The past, present and future perspectives of matrix metalloproteinase inhibitors. Pharmacol. Ther. 2020, 207, 107465. [Google Scholar] [CrossRef] [PubMed]

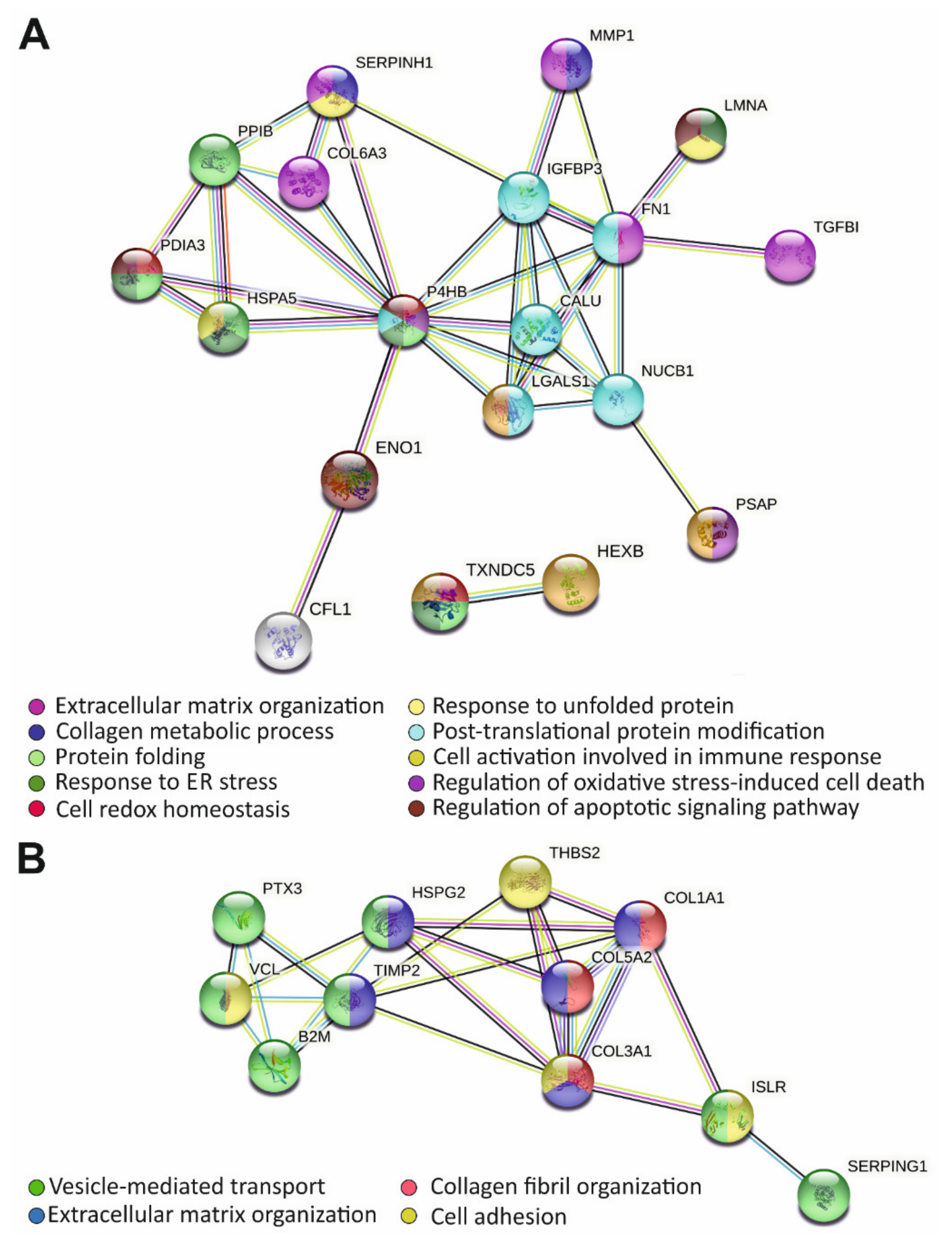

| Protein Accession N° a | Protein Name | Protein Description | Fold-Change | FDR Corrected p-Value |
|---|---|---|---|---|
| Up-regulated proteins | ||||
| P03956 | MMP1 | Matrix metalloproteinase 1 | 53.02 | 0.0016 |
| P50454 | SERPINH1 | Serpin family H member 1 | 6.19 | 0.002 |
| Q15582 | TGFBI | Transforming growth factor beta induced | 3.31 | 0.002 |
| A6XND1 | IGFBP3 | Insulin-like growth factor binding protein 3 | 3.12 | 0.024 |
| Q3BDU5 | LMNA | Lamin A/C | 2.60 | 0.003 |
| P02751 | FN1 | Fibronectin 1 | 2.44 | 0.019 |
| P09382 | LGALS1 | Galectin 1 | 2.31 | 0.001 |
| O43852 | CALU | Calumenin | 2.20 | 0.012 |
| P06733 | ENO1 | Enolase 1 | 2.14 | 0.042 |
| P11021 | HSPA5 | Heat shock protein family A (Hsp70) member 5 | 2.08 | 0.006 |
| P30101 | PDIA3 | Protein disulfide isomerase family A member 3 | 2.08 | 0.036 |
| Q07954 | LRP1 | LDL receptor related protein 1 | 2.07 | 0.002 |
| F6SYF8 | DKK3 | Dickkopf WNT signaling pathway inhibitor 3 | 2.04 | 0.041 |
| P07686 | HEXB | Hexosaminidase subunit beta | 1.90 | <0.001 |
| P12111 | COL6A3 | Collagen type VI alpha 3 chain | 1.61 | 0.029 |
| C9JIZ6 | PSAP | Prosaposin | 1.60 | 0.002 |
| Q8NBS9 | TXNDC5 | Thioredoxin domain containing 5 | 1.54 | 0.003 |
| P07237 | P4HB | Prolyl 4-hydroxylase subunit beta | 1.53 | 0.016 |
| Q02818 | NUCB1 | Nucleobindin 1 | 1.52 | 0.022 |
| P23528 | CFL1 | Cofilin 1 | 1.44 | 0.023 |
| P23284 | PPIB | Peptidylprolyl isomerase B | 1.40 | 0.040 |
| Down-regulated proteins | ||||
| P35442 | THBS2 | Thrombospondin 2 | 0.76 | 0.008 |
| P02452 | COL1A1 | Collagen type I alpha 1 chain | 0.71 | 0.010 |
| P98160 | HSPG2 | Heparan sulfate proteoglycan 2 | 0.67 | 0.031 |
| P18206 | VCL | Vinculin | 0.65 | 0.014 |
| O14498 | ISLR | Immunoglobulin superfamily containing leucine rich repeat | 0.64 | 0.012 |
| P61769 | B2M | Beta-2-microglobulin | 0.63 | 0.021 |
| P16035 | TIMP2 | TIMP metallopeptidase inhibitor 2 | 0.63 | 0.015 |
| P26022 | PTX3 | Pentraxin 3 | 0.59 | 0.004 |
| P02461 | COL3A1 | Collagen type III alpha 1 chain | 0.56 | 0.003 |
| P05155 | SERPING1 | Serpin family G member 1 | 0.56 | 0.012 |
| P05997 | COL5A2 | Collagen type V alpha 2 chain | 0.42 | 0.001 |
| Biological Process | ||
|---|---|---|
| GO term | Proteins | FDR |
| GO:0030198~extracellular matrix organization | COL1A1, COL3A1, COL5A2, FN1, COL6A3, TGFBI, HSPG2 | 0.0003 |
| GO:0030574~collagen catabolic process | COL1A1, COL3A1, MMP1, COL5A2, COL6A3 | 0.0005 |
| GO:0002576~platelet degranulation | ISLR, PSAP, FN1, SERPING1, VCL | 0.0026 |
| GO:0030199~collagen fibril organization | COL1A1, COL3A1, COL5A2, SERPINH1 | 0.0028 |
| GO:0007155~cell adhesion | COL1A1, ISLR, FN1, COL6A3, TGFBI, THBS2, VCL | 0.0073 |
| GO:0022617~extracellular matrix disassembly | MMP1, TIMP2, FN1, HSPG2 | 0.015 |
| Molecular Function | ||
| GO term | Proteins | FDR |
| GO:0005178~integrin binding | COL3A1, TIMP2, FN1, TGFBI, P4HB | 0.0021 |
| GO:0005518~collagen binding | SERPINH1, FN1, TGFBI, PPIB | 0.0053 |
| GO:0003756~protein disulfide isomerase activity | PDIA3, P4HB, TXNDC5 | 0.0094 |
| GO:0005509~calcium ion binding | LRP1, HSPA5, MMP1, CALU, NUCB1, THBS2, HSPG2 | 0.0143 |
| Cellular Component | ||
| GO term | Proteins | FDR |
| GO:0031012~extracellular matrix | HSPA5, MMP1, FN1, THBS2, HSPG2, COL1A1, COL3A1, LGALS1, LMNA, COL5A2, CFL1, TIMP2, COL6A3, TGFBI, P4HB | 1.32 × 10−15 |
| GO:0005576~extracellular region | MMP1, IGFBP3, FN1, THBS2, HSPG2, DKK3, COL1A1, COL3A1, ISLR, COL5A2, PSAP, TIMP2, CALU, SERPING1, COL6A3, PTX3, TGFBI, P4HB, B2M, VCL | 1.21 × 10−11 |
| GO:0005788~endoplasmic reticulum lumen | PDIA3, COL1A1, COL3A1, HSPA5, COL5A2, SERPINH1, COL6A3, P4HB, PPIB, B2M, TXNDC5 | 1.21 × 10−11 |
| GO:0070062~extracellular exosome | PDIA3, HSPA5, HEXB, IGFBP3, FN1, ENO1, HSPG2, ISLR, LGALS1, CFL1, PSAP, TIMP2, SERPINH1, SERPING1, COL6A3, NUCB1, TGFBI, P4HB, PPIB, B2M, VCL, TXNDC5 | 1.52 × 10−9 |
| GO:0005925~focal adhesion | PDIA3, LRP1, HSPA5, CFL1, P4HB, PPIB, B2M, HSPG2, VCL | 3.53 × 10−6 |
| GO:0005581~collagen trimer | COL1A1, COL3A1, MMP1, COL5A2, SERPINH1, COL6A3 | 6.36 × 10−6 |
| GO:0005793~ER-Golgi intermediate compartment | HSPA5, SERPINH1, FN1, NUCB1, P4HB | 5.69 × 10−5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chiarelli, N.; Zoppi, N.; Venturini, M.; Capitanio, D.; Gelfi, C.; Ritelli, M.; Colombi, M. Matrix Metalloproteinases Inhibition by Doxycycline Rescues Extracellular Matrix Organization and Partly Reverts Myofibroblast Differentiation in Hypermobile Ehlers-Danlos Syndrome Dermal Fibroblasts: A Potential Therapeutic Target? Cells 2021, 10, 3236. https://doi.org/10.3390/cells10113236
Chiarelli N, Zoppi N, Venturini M, Capitanio D, Gelfi C, Ritelli M, Colombi M. Matrix Metalloproteinases Inhibition by Doxycycline Rescues Extracellular Matrix Organization and Partly Reverts Myofibroblast Differentiation in Hypermobile Ehlers-Danlos Syndrome Dermal Fibroblasts: A Potential Therapeutic Target? Cells. 2021; 10(11):3236. https://doi.org/10.3390/cells10113236
Chicago/Turabian StyleChiarelli, Nicola, Nicoletta Zoppi, Marina Venturini, Daniele Capitanio, Cecilia Gelfi, Marco Ritelli, and Marina Colombi. 2021. "Matrix Metalloproteinases Inhibition by Doxycycline Rescues Extracellular Matrix Organization and Partly Reverts Myofibroblast Differentiation in Hypermobile Ehlers-Danlos Syndrome Dermal Fibroblasts: A Potential Therapeutic Target?" Cells 10, no. 11: 3236. https://doi.org/10.3390/cells10113236
APA StyleChiarelli, N., Zoppi, N., Venturini, M., Capitanio, D., Gelfi, C., Ritelli, M., & Colombi, M. (2021). Matrix Metalloproteinases Inhibition by Doxycycline Rescues Extracellular Matrix Organization and Partly Reverts Myofibroblast Differentiation in Hypermobile Ehlers-Danlos Syndrome Dermal Fibroblasts: A Potential Therapeutic Target? Cells, 10(11), 3236. https://doi.org/10.3390/cells10113236











