Use of Cytokine Mix-, Imiquimod-, and Serum-Induced Monoculture and Lipopolysaccharide- and Interferon Gamma-Treated Co-Culture to Establish In Vitro Psoriasis-like Inflammation Models
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Lines and Maintenance of Mono- and Co-Cultures
2.2. Stimulation to Psoriasis (Ps)-Like Inflammation Status
2.2.1. Stimulation with a Pro-Inflammatory Mix of Cytokines
2.2.2. Stimulation with Imiquimod (IMQ)
2.2.3. Stimulation with Serum
2.2.4. Stimulation with Phorbol 12-Myristate 13-Acetate (PMA), Lipopolysaccharide (LPS), and Interferon Gamma (IFN-γ)
2.2.5. Cell Viability Assay
2.2.6. RNA Handling and Reverse Transcription
2.2.7. Real-Time Quantitative Reverse Transcriptase-Polymerase Chain Reaction (Real-Time qRT-PCR)
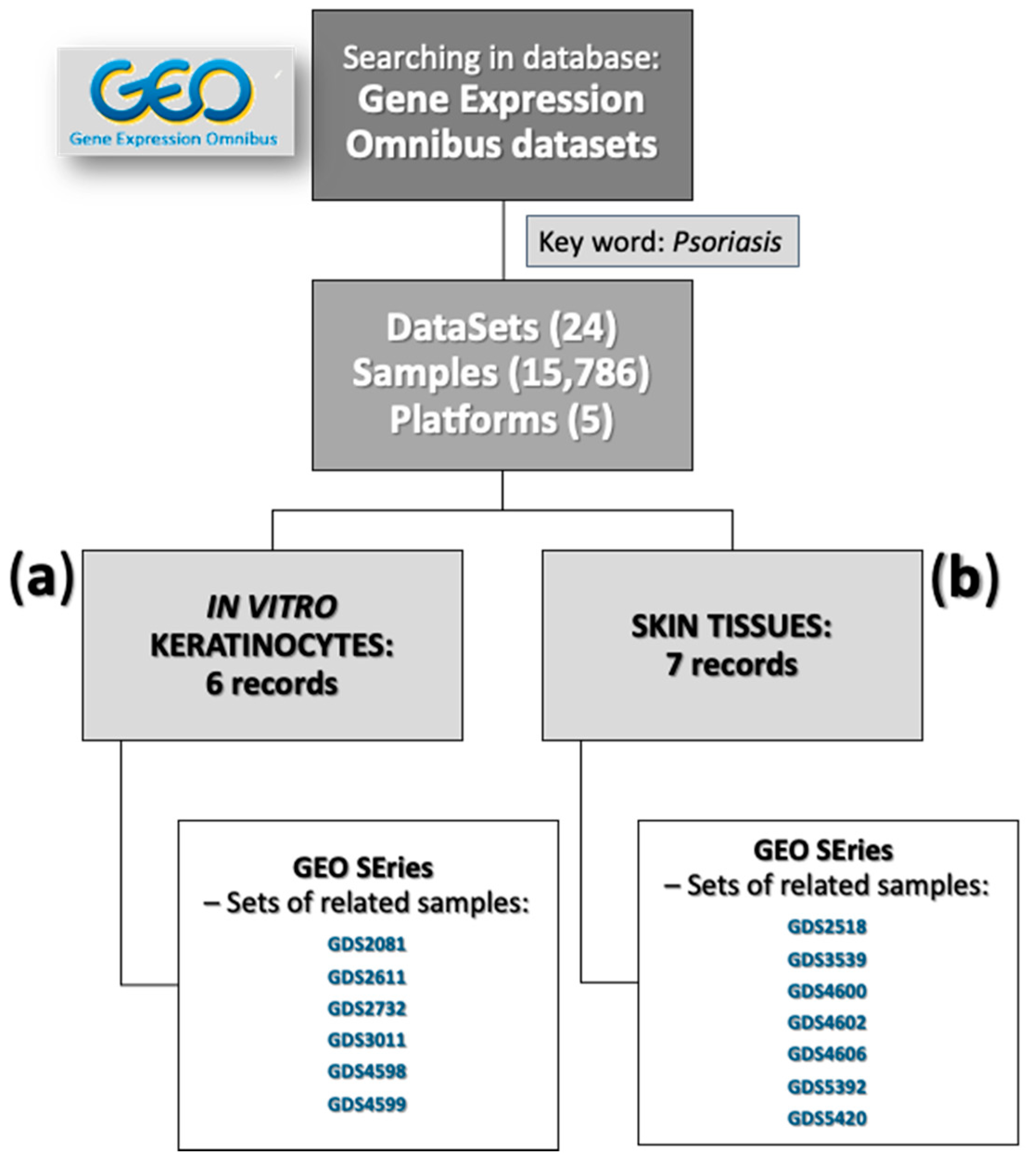
2.2.8. Gene Expression Omnibus (GEO) Analyses
3. Results
3.1. Designation of Gene Markers
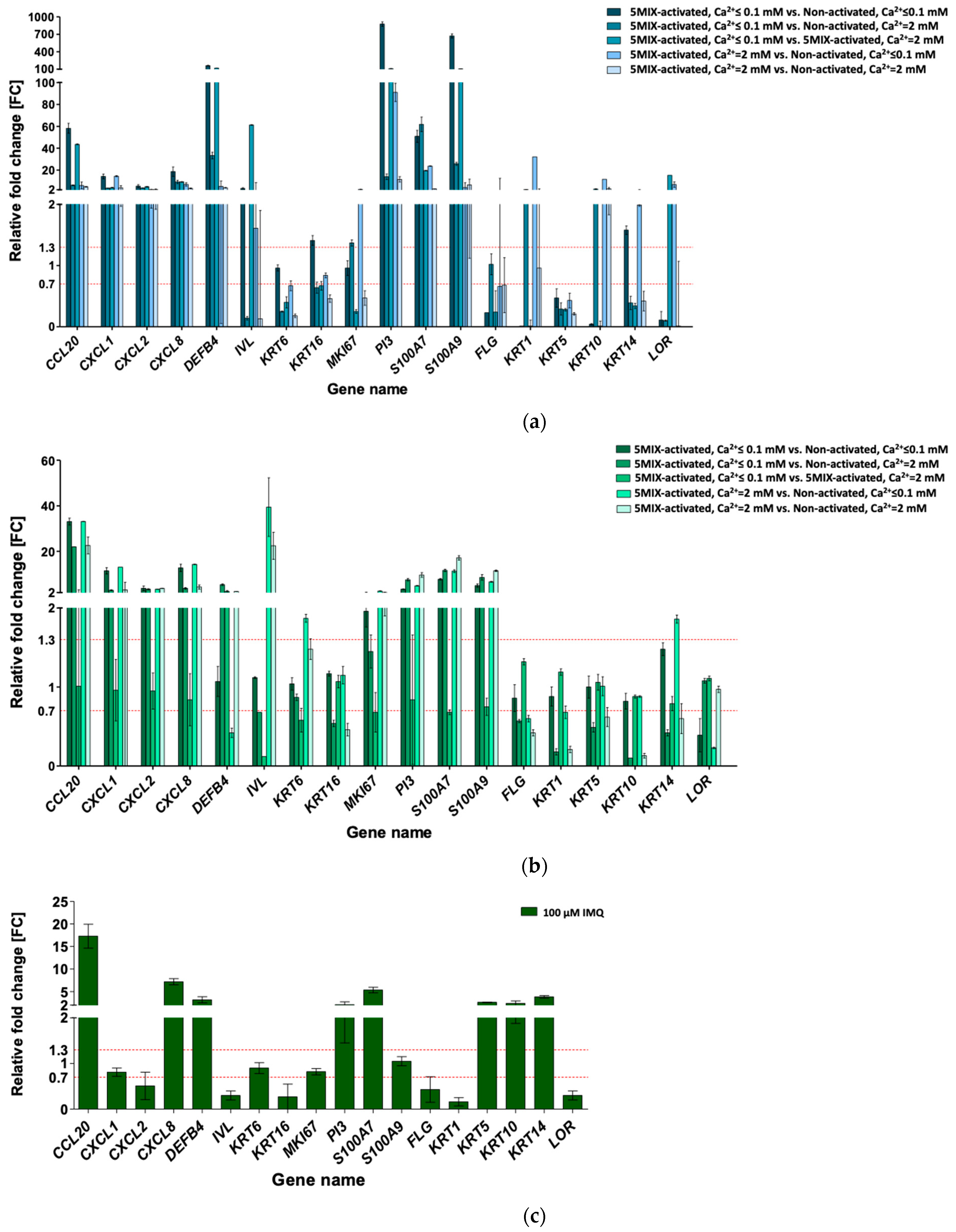
3.2. Cytokine Mix-Stimulated Psoriasis (Ps)-Like Inflammation Response in HaCaT and pKC
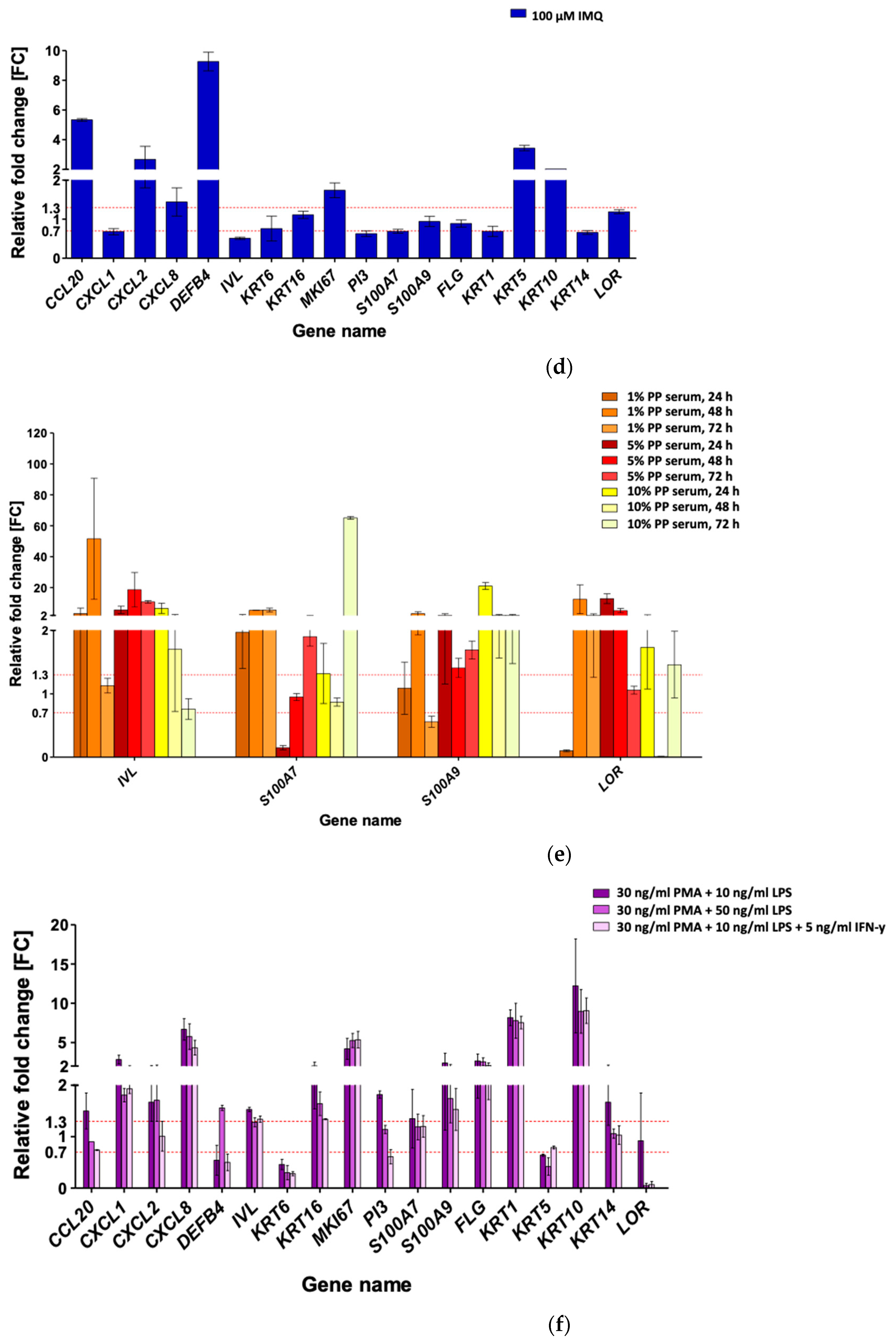
3.3. Imiquimod (IMQ) as Ps-Like Inflammation Factor in HaCaT and pKC
3.4. Serum-Mediated Ps-Like Inflammation Modulation in HaCaT
3.5. Phorbol 12-Myristate 13-Acetate (PMA), Lipopolysaccharide (LPS), and Interferon-Gamma (IFN-γ) for Ps-Like Inflammation Stimulation in HaCaT with THP-1
3.6. Statement of In Vitro Culture and Skin Biopsies Data from GEO in Relation to Our Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Benam, K.H.; Dauth, S.; Hassell, B.; Herland, A.; Jain, A.; Jang, K.J.; Karalis, K.; Kim, H.J.; MacQueen, L.; Mahmoodian, R.; et al. Engineered In Vitro Disease Models. Annu. Rev. Pathol. 2015, 10, 195–262. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sántha, M. Biologia Futura: Animal Testing in Drug Development-the Past, the Present and the Future. Biol. Futur. 2020, 71, 443–452. [Google Scholar] [CrossRef] [PubMed]
- Marzano, A.V.; Damiani, G.; Genovese, G.; Gattorno, M.A. Dermatologic Perspective on Autoinflammatory Diseases. Clin. Exp. Rheumatol. 2018, 110, 32–38. [Google Scholar]
- Raharja, A.; Mahil, S.K.; Barker, J.N. Psoriasis: A Brief Overview. Clin. Med. 2021, 21, 170–173. [Google Scholar] [CrossRef] [PubMed]
- Meglio, P.D.; Villanova, F.; Nestle, F.O. Psoriasis. Cold Spring Harb. Perspect. Med. 2014, 4, a015354. [Google Scholar] [CrossRef] [Green Version]
- Ramezani, M.; Shamshiri, A.; Zavattaro, E.; Khazaei, S.; Rezaei, M.; Mahmoodi, R.; Sadeghi, M. Immunohistochemical Expression of P53, Ki-67, and CD34 in Psoriasis and Psoriasiform Dermatitis. Biomedicine 2019, 9, 26. [Google Scholar] [CrossRef] [Green Version]
- Ganguly, D.; Chamilos, G.; Lande, R.; Gregorio, J.; Meller, S.; Facchinetti, V.; Homey, B.; Barrat, F.J.; Zal, T.; Gilliet, M. Self-RNA–Antimicrobial Peptide Complexes Activate Human Dendritic Cells through TLR7 and TLR8. J. Exp. Med. 2009, 206, 1983–1994. [Google Scholar] [CrossRef]
- Terhune, J.; Berk, E.; Czerniecki, B.J. Dendritic Cell-Induced Th1 and Th17 Cell Differentiation for Cancer Therapy. Vaccines 2013, 1, 527–549. [Google Scholar] [CrossRef] [Green Version]
- Palomino, D.C.T.; Marti, L.C. Chemokines and Immunity. Einstein 2015, 13, 469–473. [Google Scholar] [CrossRef] [Green Version]
- Abdallah, H.K.; Johansen, C.; Iversen, L. Key Signaling Pathways in Psoriasis: Recent Insights from Antipsoriatic Therapeutics. Psoriasis 2021, 11, 83–97. [Google Scholar] [CrossRef]
- Mylonas, A.; Conrad, C. Psoriasis: Classical vs. Paradoxical. The Yin-Yang of TNF and Type I Interferon. Front. Immunol. 2018, 9, 2746. [Google Scholar] [CrossRef]
- Lowes, M.A.; Suárez-Fariñas, M.; Krueger, J.G. Immunology of Psoriasis. Annu. Rev. Immunol. 2014, 32, 227–255. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Albanesi, C.; Madonna, S.; Gisondi, P.; Girolomoni, G. The Interplay Between Keratinocytes and Immune Cells in the Pathogenesis of Psoriasis. Front. Immunol. 2018, 9, 1549. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, T.; Lin, X.; Meng, X.; Lin, M. Phosphoinositide-3 Kinase/Protein Kinase-B/Mammalian Target of Rapamycin Pathway in Psoriasis Pathogenesis. A Potential Therapeutic Target? Acta. Derm. Venereol. 2014, 94, 371–379. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.; Yin, M.; Zhang, L.J. Keratin 6, 16 and 17—Critical Barrier Alarmin Molecules in Skin Wounds and Psoriasis. Cells 2019, 8, 807. [Google Scholar] [CrossRef] [Green Version]
- Kim, B.E.; Howell, M.D.; Guttman-Yassky, E.; Gilleaudeau, P.M.; Cardinale, I.R.; Boguniewicz, M.; Krueger, J.G.; Leung, D.Y.M. TNF-α Downregulates Filaggrin and Loricrin through c-Jun N-terminal Kinase: Role for TNF-α Antagonists to Improve Skin Barrier. J. Invest. Dermatol. 2011, 131, 1272–1279. [Google Scholar] [CrossRef] [Green Version]
- Bocheńska, K.; Smolińska, E.; Moskot, M.; Jakóbkiewicz-Banecka, J.; Gabig-Cimińska, M. Models in the Research Process of Psoriasis. Int. J. Mol. Sci. 2017, 4, 2514. [Google Scholar] [CrossRef] [Green Version]
- Giot, J.P.; Paris, I.; Levillain, P.; Huguier, V.; Charreau, S.; Delwail, A.; Garcia, M.; Garnier, J.; Bernard, F.X.; Dagregorio, G.; et al. Involvement of IL-1 and Oncostatin M in Acanthosis Associated With Hypertensive Leg Ulcer. Am. J. Pathol. 2013, 182, 806–818. [Google Scholar] [CrossRef]
- Guilloteau, K.; Paris, I.; Pedretti, N.; Boniface, K.; Juchaux, F.; Huguier, V.; Guillet, G.; Bernard, F.X.; Lecron, J.C.; Morel, F. Skin Inflammation Induced by the Synergistic Action of IL-17A, IL-22, Oncostatin M, IL-1{alpha}, and TNF-{alpha} Recapitulates Some Features of Psoriasis. J. Immunol. 2010, 184, 5263–5270. [Google Scholar] [CrossRef] [Green Version]
- Desmet, E.; Ramadhas, A.; Lambert, J.; Van Gele, M. In Vitro Psoriasis Models with Focus on Reconstructed Skin Models as Promising Tools in Psoriasis Research. Exp. Biol. Med. 2017, 242, 1158–1169. [Google Scholar] [CrossRef] [Green Version]
- Bocheńska, K.; Moskot, M.; Malinowska, M.; Jakóbkiewicz-Banecka, J.; Szczerkowska-Dobosz, A.; Purzycka-Bohdan, D.; Pleńkowska, J.; Słomiński, B.; Gabig-Cimińska, M. Lysosome Alterations in the Human Epithelial Cell Line HaCaT and Skin Specimens: Relevance to Psoriasis. Int. J. Mol. Sci. 2019, 20, 2255. [Google Scholar] [CrossRef] [Green Version]
- Ishida-Yamamoto, A.; Iizuka, H. Differences in Involucrin Immunolabeling within Cornified Cell Envelopes in Normal and Psoriatic Epidermis. J. Invest. Dermatol. 1995, 104, 391–395. [Google Scholar] [CrossRef] [Green Version]
- Bowcock, A.; Shannon, W.; Du, F.; Duncan, J.; Cao, K.; Aftergut, K.; Catier, J.; Fernandez-Vina, M.A.; Menter, A. Insights into Psoriasis and Other Inflammatory Diseases from Large-Scale Gene Expression Studies. Hum. Mol. Genet. 2001, 10, 1793–1805. [Google Scholar] [CrossRef] [Green Version]
- Elango, T.; Sun, J.; Zhu, C.; Zhou, F.; Zhang, Y.; Sun, L.; Yang, S.; Zhan, X. Mutational Analysis of Epidermal and Hyperproliferative Type I Keratins in Mild and Moderate Psoriasis Vulgaris Patients: A Possible Role in the Pathogenesis of Psoriasis along with Disease Severity. Hum. Genom. 2018, 12, 27. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, J.B.; Sedgewick, A.J.; Finnegan, A.I.; Harirchian, P.; Lee, J.; Kwon, S.; Fassett, M.S.; Golovato, J.; Gray, M.; Ghadially, R.; et al. Transcriptional Programming of Normal and Inflamed Human Epidermis at Single-Cell Resolution. Cell. Rep. 2018, 25, 871–883. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thewes, M.; Stadler, R.; Korge, B.; Mischke, D. Normal Psoriatic Epidermis Expression of Hyperproliferation-Associated Keratins. Arch. Dermatol. Res. 1991, 283, 465–471. [Google Scholar] [CrossRef]
- Giardina, E.; Capon, F.; De Rosa, M.; Mango, R.; Zambruno, G.; Orecchia, A.; Chimenti, S.; Giardina, B.; Novelli, G. Characterization of the Loricrin (LOR) Gene as a Positional Candidate for the PSORS4 Psoriasis Susceptibility Locus. Ann. Hum. Genet. 2004, 68, 639–645. [Google Scholar] [CrossRef] [PubMed]
- Hollox, E.J.; Huffmeier, U.; Zeeuwen, P.L.J.M.; Palla, R.; Lascorz, J.; Rodijk-Olthuis, D.; van de Kerkhof, P.C.M.; Traupe, H.; de Jongh, G.; den Heijer, M.; et al. Psoriasis is Associated with Increased Beta-Defensin Genomic Copy Number. Nat. Genet. 2008, 40, 23–25. [Google Scholar] [CrossRef] [Green Version]
- Schalkwijk, J.; de Roo, C.; de Jongh, G.J. Skin-Derived Antileukoproteinase (SKALP) an Elastase Inhibitor from Human Keratinocytes: Purification and Biochemical Properties. Biochim. Biophys. Acta. 1991, 1096, 148–154. [Google Scholar] [CrossRef]
- Madsen, P.; Rasmussen, H.H.; Leffers, H.; Honoré, B.; Dejgaard, K.; Olsen, E.; Kiil, J.; Walbum, E.; Andersen, A.H.; Basse, B.; et al. Molecular Cloning, Occurrence, and Expression of a Novel Partially Secreted Protein “Psoriasin” that is Highly Up-Regulated in Psoriatic Skin. J. Investig. Dermatol. 1991, 97, 701–712. [Google Scholar] [CrossRef] [Green Version]
- Harper, E.G.; Guo, C.; Rizzo, H.; Lillis, J.V.; Kurtz, S.E.; Skorcheva, I.; Purdy, D.; Fitch, E.; Iordanov, M.; Blauvelt, A. Th17 Cytokines Stimulate CCL20 Expression in Keratinocytes In Vitro and In Vivo: Implications for Psoriasis Pathogenesis. J. Invest. Dermatol. 2009, 129, 2175–2183. [Google Scholar] [CrossRef] [Green Version]
- Suárez-Fariñas, M.; Li, K.; Fuentes-Duculan, J.; Hayden, K.; Brodmerkel, C.; Krueger, J.G. Expanding the Psoriasis Disease Profile: Interrogation of the Skin and Serum of Patients with Moderate-to-Severe Psoriasis. J. Invest. Dermatol. 2012, 132, 2552–2564. [Google Scholar] [CrossRef] [Green Version]
- Kennedy-Crispin, M.; Billick, E.; Mitsui, H.; Gulati, N.; Fujita, H.; Gilleaudeau, P.; Sullivan-Whalen, M.; Johnson-Huang, L.M.; Suárez-Fariñas, M.; Krueger, J.G. Human Keratinocytes’ Response to Injury Upregulates CCL20 and Other Genes Linking Innate and Adaptive Immunity. J. Invest. Dermatol. 2012, 132, 105–113. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Mare, S.; de Jong, E.; Van Erp, P.E.; Van de Kerkhof, P.C. Markers for Proliferation and Keratinization in the Margin of the Active Psoriatic Lesion. Br. J. Dermatol. 1990, 122, 469–475. [Google Scholar] [CrossRef]
- Varma, S.R. Imiquim Od-Induced Psoriasis-Like Inflammation in Differentiated Human Keratinocytes: Its Evaluation Using Curcumin. Eur. J. Pharmacol. 2017, 813, 33–41. [Google Scholar] [CrossRef]
- Lund, M.E.; To, J.; O’Brien, B.A.; Donnelly, S. The Choice of Phorbol 12-Myristate 13-Acetate Differentiation Protocol Influences the Response of THP-1 Macrophages to a Pro-Inflammatory Stimulus. J. Immunol. Methods 2016, 430, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Kämpfer, A.A.M.; Urbán, P.; Gioria, S.; Kanase, N.; Stone, V.; Kinsner-Ovaskainen, A. Development of an In Vitro Co-Culture Model to Mimic the Human Intestine in Healthy and diseased state. Toxicologist 2017, 45, 31–43. [Google Scholar] [CrossRef] [PubMed]
- Smolińska, E.; Moskot, M.; Jakóbkiewicz-Banecka, J.; Węgrzyn, G.; Banecki, B.; Szczerkowska-Dobosz, A.; Purzycka-Bohdan, D.; Gabig-Cimińska, M. Molecular Action of Isoflavone Genistein in the Human Epithelial Cell Line HaCaT. PLoS ONE 2018, 13, e0192297. [Google Scholar] [CrossRef]
- Heard, C.M. An Ex Vivo Skin Model to Probe Modulation of Local Cutaneous Arachidonic Acid Inflammation Pathway. J. Biol. Methods 2020, 7, e138. [Google Scholar] [CrossRef]
- Doke, S.K.; Dhawale, S.C. Alternatives to Animal Testing: A Review. Saudi. Pharm. J. 2015, 23, 223–229. [Google Scholar] [CrossRef] [Green Version]
- Sivamani, R.K.; Correa, G.; Ono, Y.; Bowen, M.P.; Raychaudhuri, S.P.; Maverakis, E. Biological Therapy of Psoriasis. Indianj. Dermatol. 2010, 55, 161–170. [Google Scholar]
- Astashkina, A.; Mann, B.; Grainger, D.W. A Critical Evaluation of In Vitro Cell Culture Models for High-Throughput Drug Screening and Toxicity. Pharm. Ther. 2012, 134, 82–106. [Google Scholar] [CrossRef] [PubMed]
- Kanda, N.; Hoashi, T.; Saeki, H. The Defect in Regulatory T Cells in Psoriasis and Therapeutic Approaches. J. Clin. Med. 2021, 10, 3880. [Google Scholar] [CrossRef]
- Amigó, M.; Payá, M.; Braza-Boïls, A.; De Rosa, S.; Carmen Terencio, M. Avarol Inhibits TNF-Alpha Generation and NF-KappaB Activation in Human Cells and in Animal Models. Life Sci. 2008, 82, 256–264. [Google Scholar] [CrossRef] [PubMed]
- Weng, Z.; Patel, A.B.; Vasiadi, M.; Therianou, A.; Theoharides, T.C. Luteolin Inhibits Human Keratinocyte Activation and Decreases NF-κB Induction that is Increased in Psoriatic Skin. PLoS ONE 2014, 9, e90739. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olaru, F.; Jensen, L.E. Chemokine Expression by Human Keratinocyte Cell Lines after Activation of Toll-Like Receptors. Exp. Dermatol. 2010, 19, e314–e316. [Google Scholar] [CrossRef] [Green Version]
- Boukamp, P.; Petrussevska, R.T.; Breitkreutz, D.; Hornung, J.; Markham, A.; Fusenig, N.E. Normal Keratinization in a Spontaneously Immortalized Aneuploid Human Keratinocyte Cell Line. J. Cell. Biol. 1988, 106, 761–771. [Google Scholar] [CrossRef] [Green Version]
- Seo, M.D.; Kang, T.J.; Lee, C.H.; Lee, A.Y.; Noh, M. HaCaT Keratinocytes and Primary Epidermal Keratinocytes Have Different Transcriptional Profiles of Cornified Envelope-Associated Genes to T Helper Cell Cytokines. Biomol. Ther. 2012, 20, 171–176. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dallaglio, K.; Marconi, A.; Truzzi, F.; Lotti, R.; Palazzo, E.; Petrachi, T.; Saltari, A.; Coppini, M.; Pincelli, C. E-FABP Induces Differentiation in Normal Human Keratinocytes and Modulates the Differentiation Process in Psoriatic Keratinocytes In Vitro. Exp. Dermatol. 2013, 22, 255–261. [Google Scholar] [CrossRef] [PubMed]
- Leigh, I.M.; Navsaria, H.; Purkis, P.; Mckay, I.; Bowden, P.; Riddle, P. Keratins (K16 and K17) as Markers of Keratinocyte Hyperproliferation in Psoriasis In Vivo and In Vitro. Br. J. Dermatol. 1995, 133, 501–511. [Google Scholar] [CrossRef] [PubMed]
- Holzmann, H.; Bernd, A.; Hohlmaier, K. Proliferation of Non-Psoriatic Human Fibroblasts In Vitro by Serum from Patients with Psoriasis. Lancet 1988, 29, 1031. [Google Scholar] [CrossRef]
- Laubscher, T.A.; Harris, N.; Chalton, D.O. The Mitogenic Effects of Sera from Psoriatic Subjects on Normal Dermal Fibroblasts: An Absence of Correlation with the Clinical Activity of Psoriasis. Br. J. Dermatol. 1991, 124, 163–167. [Google Scholar] [CrossRef]
- Priestley, G.C.; Adams, L.W. Mitogenic Effects of Sera from Normal and Psoriatic Subjects on Human Skin Fibroblasts. Arch. Dermatol. Res. 1985, 277, 13–15. [Google Scholar] [CrossRef] [PubMed]
- Baxter, E.W.; Graham, A.E.; Re, N.A.; Carr, I.M.; Robinson, J.I.; Mackie, S.L.; Morgan, A.W. Standardized Protocols for Differentiation of THP-1 Cells to Macrophages with Distinct M(IFNγ+LPS), M(IL-4) and M(IL-10) Phenotypes. J. Immunol. Methods 2020, 478, 112721. [Google Scholar] [CrossRef] [PubMed]
- Caras, I.; Tucureanu, C.; Lerescu, L.; Pitica, R.; Melinceanu, L.; Neagu, S.; Salageanu, A. Influence of Tumor Cell Culture Supernatants on Macrophage Functional Polarization: In Vitro Models of Macrophage-Tumor Environment Interaction. Tumori 2011, 97, 647–654. [Google Scholar] [CrossRef] [PubMed]
- Rabeony, H.; Petit-Paris, I.; Garnier, J.; Barrault, C.; Pedretti, N.; Guilloteau, K.; Jegou, J.F.; Guillet, G.; Huguier, V.; Lecron, J.C.; et al. Inhibition of Keratinocyte Differentiation by the Synergistic Effect of IL-17A, IL-22, IL-1α, TNFα and Oncostatin, M. PLoS ONE 2014, 9, e101937. [Google Scholar] [CrossRef] [Green Version]
- Giardina, E.; Sinibaldi, C.; Novelli, G. The Psoriasis Genetics as a Model of Complex Disease. Curr. Drug Targets Inflamm. Allergy. 2004, 3, 129–136. [Google Scholar] [CrossRef]
- Hu, Z.; Xiong, Z.; Xu, X.; Li, F.; Lu, L.; Li, W.; Su, J.; Liu, Y.; Liu, D.; Xie, Z.; et al. Loss-of-Function Mutations in Filaggrin Gene Associate with Psoriasis Vulgaris in Chinese Population. Hum. Genet. 2012, 131, 1269–1274. [Google Scholar] [CrossRef]
- Suga, H. Skin Barrier Dysfunction and Low Antimicrobial Peptide Expression in Cutaneous T-Cell Lymphoma. Clin. Cancer. Res. 2014, 20, 4339–4348. [Google Scholar] [CrossRef] [Green Version]
- Swindell, W.R.; Sarkar, M.K.; Liang, Y.; Xing, X.; Baliwag, J.; Elder, J.T.; Johnston, A.; Ward, N.L.; Gudjonsson, J.E. RNA-Seq Identifies a Diminished Differentiation Gene Signature in Primary Monolayer Keratinocytes Grown from Lesional and Uninvolved Psoriatic Skin. Sci. Rep. 2017, 7, 18045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reichelt, J.; Furstenberger, G.; Magin, T.M. Loss of Keratin 10 Leads to Mitogen-Activated Protein Kinase (MAPK) Activation, Increased Keratinocyte Turnover, and Decreased Tumor Formation in Mice. J. Invest. Dermatol. 2004, 123, 973–981. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gudjonsson, J.E.; Ding, J.; Li, X.; Nair, R.P.; Tejasvi, T.; Qin, Z.S.; Ghosh, D.; Aphale, A.; Gumucio, D.L.; Voorhees, J.; et al. Global Gene Expression Analysis Reveals Evidence for Decreased Lipid Biosynthesis and Increased Innate Immunity in Uninvolved Psoriatic Skin. J. Invest. Dermatol. 2009, 129, 2795–2804. [Google Scholar] [CrossRef] [Green Version]
- Morizane, S.; Gallo, R.L. Antimicrobial peptides in the pathogenesis of psoriasis. J. Dermatol. 2012, 39, 225–230. [Google Scholar] [CrossRef] [PubMed]
- Gudjonsson, J.E.; Ding, J.; Johnston, A.; Tejasvi, T.; Guzman, A.M.; Nair, R.P.; Voorhees, J.J.; Abecasis, G.R.; Elder, J.T. Assessment of the Psoriatic Transcriptome in a Large Sample: Additional Regulated Genes and Comparisons with In Vitro Models. J. Invest. Dermatol. 2010, 130, 1829–1840. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Functionality Clusters | Gene Symbol | Effector Protein | Gene Expression in Ps Phenotype Referring to the Literature Data ↑ (Upregulation) or ↓ (Downregulation) |
|---|---|---|---|
| Keratinocyte differentiation markers | IVL | Involucrin | ↑ Ishida-Yamamoto et al., 1995 [22] |
| FLG | Profilaggrin | ↓ Bowcock et al., 2001 [23] | |
| KRT1 | Cytokeratin 1 | ↓ Elango et al., 2018 [24] | |
| KRT5 | Cytokeratin 5 | ↓ Cheng et al., 2018 [25] | |
| KRT6 | Cytokeratin 6 | ↑ Thewes et al., 1991 [26] | |
| KRT10 | Cytokeratin 10 | ↓ Thewes et al., 1991 [26] | |
| KRT14 | Cytokeratin 14 | ↓ Thewes et al., 1991 [26] | |
| KRT16 | Cytokeratin 16 | ↑ Thewes et al., 1991 [26] | |
| LOR | Loricrin | ↓ Giardina et al., 2004 [27] | |
| Antimicrobial peptides | DEFB4 | β-defensin | ↑ Hollox et al., 2008 [28] |
| PI3 | Peptidase Inhibitor 3 (SKALP) | ↑ Schalkwijk et al., 1993 [29] | |
| S100A7 | S100 Calcium Binding Protein A7 | ↑ Madsen et al., 1991 [30] | |
| S100A9 | S100 Calcium Binding Protein A9 | ↑ Madsen et al., 1991 [30] | |
| Chemokines | CCL20 | C-C Motif Chemokine Ligand 20 | ↑ Harper et al., 2009 [31] |
| CXCL1 | C-X-C Motif Chemokine Ligand 1 | ↑ Suárez-Fariñas et al., 2012 [32] | |
| CXCL2 | C-X-C Motif Chemokine Ligand 2 | ↑ Kennedy-Crispin et al., 2012 [33] | |
| CXCL8 | C-X-C Motif Chemokine Ligand 8 | ↑ Suárez-Fariñas at al., 2012 [32] | |
| Proliferation marker | MKI67 | Marker Of Proliferation Ki-67 | ↑ De Mare et al., 1990 [34] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bocheńska, K.; Moskot, M.; Gabig-Cimińska, M. Use of Cytokine Mix-, Imiquimod-, and Serum-Induced Monoculture and Lipopolysaccharide- and Interferon Gamma-Treated Co-Culture to Establish In Vitro Psoriasis-like Inflammation Models. Cells 2021, 10, 2985. https://doi.org/10.3390/cells10112985
Bocheńska K, Moskot M, Gabig-Cimińska M. Use of Cytokine Mix-, Imiquimod-, and Serum-Induced Monoculture and Lipopolysaccharide- and Interferon Gamma-Treated Co-Culture to Establish In Vitro Psoriasis-like Inflammation Models. Cells. 2021; 10(11):2985. https://doi.org/10.3390/cells10112985
Chicago/Turabian StyleBocheńska, Katarzyna, Marta Moskot, and Magdalena Gabig-Cimińska. 2021. "Use of Cytokine Mix-, Imiquimod-, and Serum-Induced Monoculture and Lipopolysaccharide- and Interferon Gamma-Treated Co-Culture to Establish In Vitro Psoriasis-like Inflammation Models" Cells 10, no. 11: 2985. https://doi.org/10.3390/cells10112985
APA StyleBocheńska, K., Moskot, M., & Gabig-Cimińska, M. (2021). Use of Cytokine Mix-, Imiquimod-, and Serum-Induced Monoculture and Lipopolysaccharide- and Interferon Gamma-Treated Co-Culture to Establish In Vitro Psoriasis-like Inflammation Models. Cells, 10(11), 2985. https://doi.org/10.3390/cells10112985







