Molecular Characterizations of Kenyan Brachiaria Grass Ecotypes with Microsatellite (SSR) Markers
Abstract
:1. Introduction
2. Results
2.1. Descriptive Statistics for Simple Sequence Repeat (SSR) Markers
2.2. Population Diversity Indices
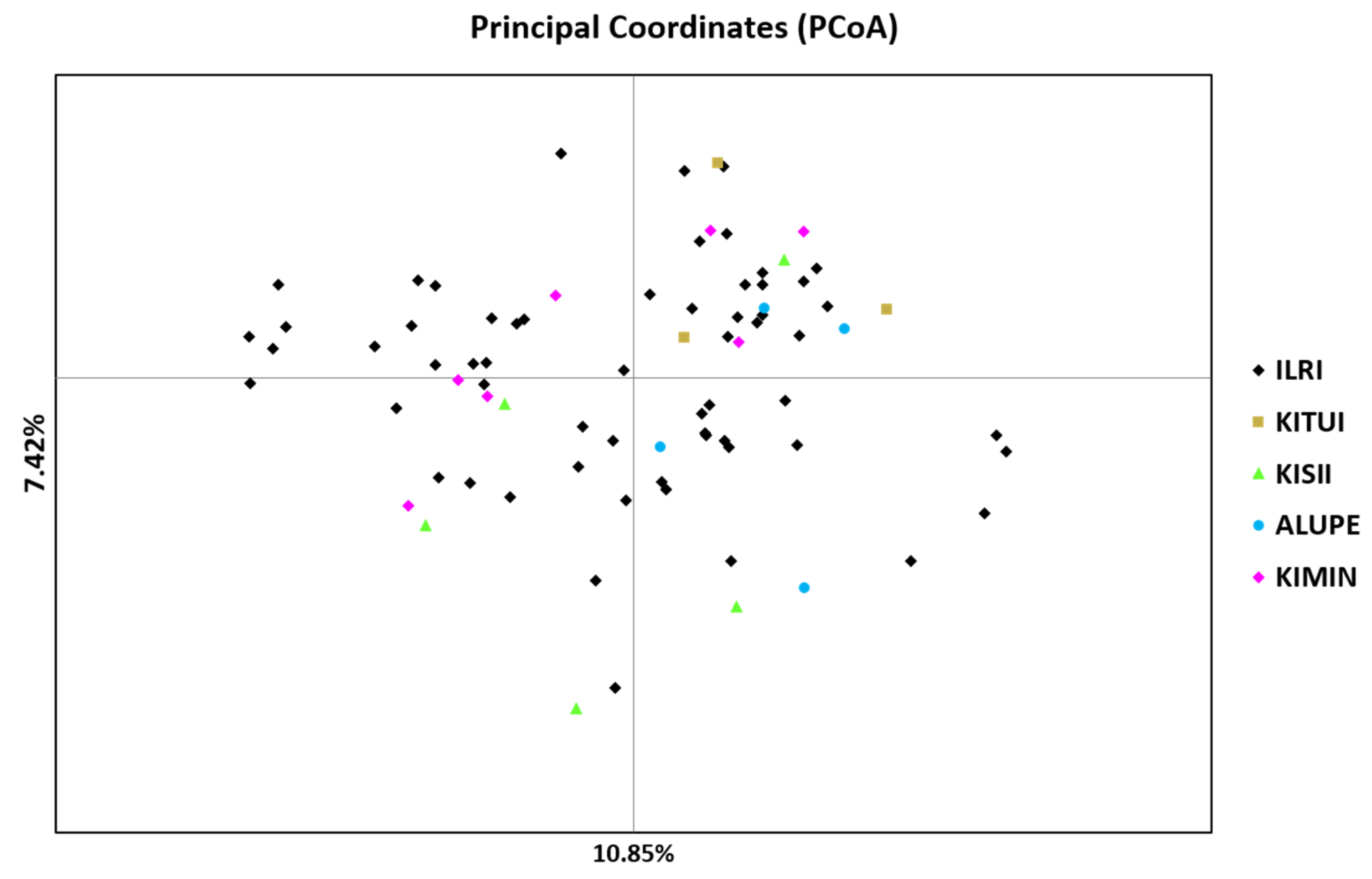
2.3. Genetic Diversity and Relationships
2.4. Analysis of Molecular Variance (AMOVA)
3. Discussion
4. Experimental Section
4.1. Source of Plant Materials
4.2. Genomic DNA Extraction
4.3. PCR Amplification and Genotyping
4.4. Data Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Renvoize, S.A.; Clayton, W.D.; Kabuye, C.H.S. Morphology, taxonomy and natural distribution of Brachiaria (Trin.) Griseb. In Brachiaria: Biology, Agronomy, and Improvement; Miles, J.W., Maass, B.L., do Valle, C.B., Eds.; International Centre for Tropical Agriculture: Cali, Colombia, 1996; pp. 1–15. [Google Scholar]
- Miles, J.W.; Maass, B.L.; do Valle, C.B. Brachiaria: Biology, Agronomy, and Improvement; International Centre for Tropical Agriculture: Cali, Colombia, 1996; p. 288. [Google Scholar]
- Holmann, F.; Rivas, L.; Argel, P.J.; Perez, E. Impact of the adoption of Brachiaria grasses: Central America and Mexico. Livest. Res. Rural Dev. 2004, 16. Available online: http://www.lrrd.org/lrrd16/12/holm16098.htm (accessed on 21 January 2017). [Google Scholar]
- Jotee, D. Evaluation of the potential of some common forage germplasms in Mauritius. In Proceedings of the Third PANESA Workshop, Arusha, Tanzania, 27–30 April 1987; Dzowela, B.H., Ed.; ILCA: Addis Ababa, Ethiopia, 1988; pp. 81–90. [Google Scholar]
- Subbarao, G.V.; Nakahara, K.; Hurtado, M.P.; Ono, H.; Moreta, D.E.; Salcedo, A.F.; Yoshihashi, A.T.; Ishikawa, T.; Ishitani, M.; Ohnishi-Kameyama, M.; et al. Evidence for biological nitrification inhibition in Brachiaria pastures. Proc. Natl. Acad. Sci. USA 2009, 106, 17302–17307. [Google Scholar] [CrossRef] [PubMed]
- Arango, J.; Moreta, D.; Núñez, J.; Hartmann, K.; Domínguez, M.; Ishitani, M.; Miles, J.; Subbarao, G.; Peters, M.; Rao, I. Developing methods to evaluate phenotypic variability in biological nitrification inhibition (BNI) capacity of Brachiaria grasses. Trop. Grassl.-Forrajes Trop. 2014, 2, 6–8. [Google Scholar] [CrossRef]
- Moreta, D.E.; Arango, J.; Sotelo, M.; Vergara, D.; Rincón, A.; Ishitani, N.; Castro, A.; Miles, J.; Peters, M.; Tohme, J.; et al. Biological nitrification inhibition (BNI) in Brachiaria pastures: A novel strategy to improve eco-efficiency of crop-livestock systems and to mitigate climate change. Trop. Grassl.-Forrajes Trop. 2014, 2, 88–91. [Google Scholar] [CrossRef]
- Rao, I.; Ishitani, M.; Miles, J.; Peters, M.; Tohme, J.; Arango, J.; Moreta, D.E.; Lopez, H.; Castro, A.; Hoek, R.V.D.; et al. Climate-smart crop-livestock systems for smallholders in the tropics: Integration of new forage hybrids to intensify agriculture and to mitigate climate change through regulation of nitrification in soil. Trop. Grassl.-Forrajes Trop. 2014, 2, 130–132. [Google Scholar] [CrossRef]
- Jank, L.; Barrios, S.C.; do Valle, C.B.; Simeão, R.M.; Alves, G.F. The value of improved pastures to Brazilian beef production. Crop Pasture Sci. 2014, 65, 1132–1137. [Google Scholar] [CrossRef]
- Ndikumana, J.; de Leeuw, P.N. Regional experiences with Brachiaria: Sub-Saharan Africa. In Brachiaria: Biology, Agronomy, and Improvement; Miles, J.W., Maass, B.L., do Valle, C.B., Eds.; International Centre for Tropical Agriculture: Cali, Colombia, 1996; pp. 247–257. [Google Scholar]
- Maass, B.L.; Midega, A.O.; Mutimura, M.; Rahetlah, V.B.; Salgado, P.; Kabirizi, J.M.; Khan, Z.R.; Ghimire, S.; Rao, I.M. Homecoming of Brachiaria: Improved hybrids prove useful for African animal agriculture. E. Afr. Agric. For. J. 2015, 81, 1–78. [Google Scholar]
- Ghimire, S.; Njarui, D.; Mutimura, M.; Cardoso, J.; Johnson, L.; Gichangi, E.; Teasdale, S.; Odokonyero, K.; Caradus, J.; Rao, I.; et al. Climate-smart Brachiaria for improving livestock production in East Africa: Emerging opportunities. In Sustainable Use of Grassland Resources for Forage Production, Biodiversity and Environmental Protection; Vijaya, D., Srivastava, M., Gupta, C., Malaviya, D., Roy, M., Mahanta, S., Singh, J., Maity, A., Ghos, P., Eds.; Range Management Society of India; ICAR-Indian Grassland and Fodder Research Institute: New Delhi, India, 2015; pp. 361–370. [Google Scholar]
- Keller-Grein, G.; Maass, B.L.; Hanson, J. Natural variation in Brachiaria and existing germplasm collection. In Brachiaria: Biology, Agronomy, and Improvement; Miles, J.W., Maass, B.L., do Valle, C.B., Eds.; International Centre for Tropical Agriculture: Cali, Colombia, 1996; pp. 16–42. [Google Scholar]
- Silva, P.I.; Martins, A.M.; Gouvea, E.G.; Pessoa-Filho, M.; Ferreira, M.E. Development and validation of microsatellite markers for Brachiaria ruziziensis obtained by partial genome assembly of Illumina single-end reads. BMC Genom. 2013, 14, 17. [Google Scholar] [CrossRef] [PubMed]
- Jungmann, L.; Vigna, B.B.Z.; Paiva, J.; Sousa, A.C.B.; do Valle, C.B.; Laborda, P.R. Development of microsatellite markers for Brachiaria humidicola (Rendle) Schweick. Conserv. Genet. Resour. 2009, 1, 475–479. [Google Scholar] [CrossRef]
- Vigna, B.B.Z.; Jungmann, L.; Francisco, P.M.; Zucchi, M.I.; do Valle, C.B.; Souza, A.P. Genetic diversity and population structure of the Brachiaria brizantha germplasm. Trop. Plant Biol. 2011, 4, 157–169. [Google Scholar] [CrossRef]
- Jungmann, L.; Sousa, A.C.B.; Paiva, J.; Francisco, P.M.; Vigna, B.B.Z.; do Valle, C.B. Isolation and characterization of microsatellite markers for Brachiaria brizantha (Hochst. ex A. Rich.) Stap. Conserv. Genet. 2009, 10, 1873–1876. [Google Scholar] [CrossRef]
- Pessoa-Filho, M.; Azevedo, A.L.S.; Sobrinho, F.S.; Gouvea, E.G.; Martins, A.M.; Ferreira, M.E. Genetic diversity and structure of Ruzigrass germplasm collected in Africa and Brazil. Crop Sci. 2015, 55, 2736–2745. [Google Scholar] [CrossRef]
- Szpiech, Z.A.; Jakobsson, M.; Rosenberg, N.A. ADZE: A rarefaction approach for counting alleles private to combinations of populations. Bioinformatics 2008, 24, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Kalinowski, S.T. Counting alleles with rarefaction: Private alleles and hierarchical sampling designs. Conserv. Genet. 2004, 5, 539–543. [Google Scholar] [CrossRef]
- Boonman, J.G. Experimental studies on seed production of tropical grasses in Kenya part 2 tillering and heading on in seed crops of eight grasses. Neth. J. Agric. Sci. 1971, 19, 237–249. [Google Scholar]
- Boonman, J.G. East Africa’s Grasses and Fodders: Their Ecology and Husbandry; Kluwer Academic Publishers: Dordrecht, The Netherlands, 1993; p. 343. [Google Scholar]
- Wandera, J.L. Forage research and production in western Kenya. In A Review of Agricultural Practices and Constraints in the North Rift Valley Province; Rees, D.J., Nkonge, C., Wandera, J.L., Eds.; Kenya Agricultural Research Institute: Nairobi, Kenya, 1997; pp. 169–187. [Google Scholar]
- Garcia, M.; Vigna, B.B.Z.; Sousa, A.C.B.; Jungmann, L.; Cidade, F.W.; Toledo-Silva, G.; Francisco, P.M.; Chiari, L.; Carvalho, M.A.; Karia, C.T.; et al. Molecular genetic variability, population structure and mating system in tropical forages. Trop. Grassl.-Forrajes Trop. 2013, 1, 25–30. [Google Scholar] [CrossRef]
- Azevedo, A.L.S.; Costa, P.P.; Machado, M.A.; de Paula, C.M.P.; Sobrinho, F.S. High degree of genetic diversity among genotypes of the forage grass Brachiaria ruziziensis (Poaceae) detected with ISSR markers. Genet. Mol. Res. 2011, 10, 3530–3538. [Google Scholar] [CrossRef] [PubMed]
- Jungmann, L.; Vigna, B.B.Z.; Boldrini, K.R.; Sousa, A.C.B.; do Valle, C.B.; Resende, R.M.S.; Pagliarini, M.S.; Zucchi, M.I.; de Souza, A.P. Genetic diversity and population structure analysis of the tropical pasture grass Brachiaria humidicola based on microsatellites, cytogenetics, morphological traits, and geographical origin. Genome 2010, 53, 698–709. [Google Scholar] [PubMed]
- Wright, S. The genetical structure of populations. Ann Eugen 1951, 15, 323–354. [Google Scholar] [CrossRef] [PubMed]
- Vucetich, J.A.; Waite, T.A. Is one migrant per generation sufficient for the genetic management of fluctuating populations? Anim. Conserv. 2000, 3, 261–266. [Google Scholar] [CrossRef]
- Do Valle, C.B.; Savidan, Y.H. Genetics, cytogenetic and reproductive biology of Brachiaria. In Brachiaria: Biology, Agronomy, and Improvement; Miles, J.W., Maass, B.L., do Valle, C.B., Eds.; International Centre for Tropical Agriculture: Cali, Colombia, 1996; pp. 147–163. [Google Scholar]
- Harrington, K.C.; Beskow, W.B.; Hodgson, J. Recovery and viability of seeds ingested by goats. N. Z. Plant Prot. 2011, 64, 75–80. [Google Scholar]
- Leitch, A.R.; Leitch, I.J. Genomic plasticity and the diversity of polyploid plants. Science 2008, 320, 481–483. [Google Scholar] [CrossRef] [PubMed]
- Malo, J.E.; Suárez, F. Herbivorous mammals as seed dispersers in a Mediterranean dehesa. Oecologia 1995, 104, 246–255. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. Isolation of plant DNA from fresh tissue. Focus 1990, 12, 13–15. [Google Scholar]
- Cordeiro, G.M.; Pan, Y.; Henry, R.J. Sugarcane microsatellites for the assessments of genetic diversity in sugarcane germplasm. Plant Sci. 2003, 165, 181–189. [Google Scholar] [CrossRef]
- Cidade, F.W.; Vigna, B.B.Z.; de Sauza, F.H.D.; Valls, J.F.M.; Dall’Agnol, M.; Zucchi, M.I.; de Sauza-Chies, T.T.; Sauza, A.P. Genetic variation in polyploid forage grass: Assessing the molecular genetic variability in the Paspalum genus. BMC Genet. 2013, 14, 50. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Zhou, H.; Pan, Y.B.; Chen, C.Y.; Zhu, J.R.; Chen, P.H.; Li, Y.R.; Cai, Q.; Chen, R.K. Seggregation analysis of microsatellite (SSR) markers in sugarcane polyploids. Genet. Mol. Res. 2015, 14, 18384–18395. [Google Scholar] [CrossRef] [PubMed]
- Schie, S.; Chaudhary, R.; Debener, T. Analysis of a complex polyploid plant genome using molecular markers: Strong evidence for segmental allooctoploidy in garden Dahalias. Plant Genome 2014, 7, 3. [Google Scholar] [CrossRef]
- Prasanth, V.P.; Chandra, S. ALS-Binary. A Program for Converting Allele Size in Microsatellite Markers into 0–1 (Binary) Data; ICRISAT: Hyderabad, India, 1997. [Google Scholar]
- Prasanth, V.P.; Chandra, S.; Jayashree, B.; Hoisington, D. AlleloBin. A Software for Allele Binning of Microsatellite Markers Based on the Algorithms of Idury and Cardon; ICRISAT: Hyderabad, India, 1997. [Google Scholar]
- Liu, K.; Muse, S.V. PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 2005, 21, 2128–2129. [Google Scholar] [CrossRef] [PubMed]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Perrier, X.; Jacquemoud-Collet, J.P. DARwin: Dissimilarity Analysis and Representation for Windows, Version 5.0.157. Computer Program. 2006. Available online: http://darwin.cirad.fr/darwin (accessed on 21 January 2017).
- Dice, LR. Measures of the amount of ecologic association between species. Ecology 1945, 26, 297–302. [Google Scholar] [CrossRef]

| Marker | MAF | NDA | I | PIC |
|---|---|---|---|---|
| Brz0012 | 0.4304 | 5 | 0.7101 | 0.6670 |
| Brz0028 | 0.4304 | 5 | 0.6521 | 0.5892 |
| Brz0029 | 0.6203 | 3 | 0.5124 | 0.4327 |
| Brz0067 | 0.4051 | 5 | 0.7419 | 0.7061 |
| Brz0076 | 0.8228 | 3 | 0.3169 | 0.3087 |
| Brz0087 | 0.481 | 8 | 0.6983 | 0.6649 |
| Brz0092 | 0.8101 | 5 | 0.3352 | 0.3240 |
| Brz0100 | 0.4684 | 4 | 0.6614 | 0.6052 |
| Brz0115 | 0.3671 | 7 | 0.8021 | 0.7829 |
| Brz0117 | 0.6076 | 6 | 0.5371 | 0.4676 |
| Brz0118 | 0.5063 | 4 | 0.5573 | 0.4613 |
| Brz0122 | 0.4557 | 6 | 0.6739 | 0.6225 |
| Brz0130 | 0.3418 | 10 | 0.7947 | 0.7706 |
| Brz0149 | 0.7722 | 5 | 0.3874 | 0.3679 |
| Brz0156 | 0.6456 | 4 | 0.5365 | 0.497 |
| Brz0203 | 0.3671 | 7 | 0.7685 | 0.7379 |
| Brz0212 | 0.5823 | 8 | 0.6195 | 0.5906 |
| Brz0213 | 0.7468 | 4 | 0.4192 | 0.3932 |
| Brz0214 | 0.4304 | 7 | 0.7432 | 0.7138 |
| Brz0235 | 0.4051 | 4 | 0.7438 | 0.709 |
| Brz3002 | 0.2405 | 5 | 0.854 | 0.8384 |
| Brz3009 | 0.4684 | 5 | 0.6313 | 0.5643 |
| Mean | 0.5184 | 5.45 | 0.6225 | 0.5825 |
| Population | N | Na | Np | Ae | I | Ho | He | PL (%) |
|---|---|---|---|---|---|---|---|---|
| ILRI | 60 | 3.633 | 0.833 | 2.21 | 0.887 | 0.76 | 0.499 | 86.67 |
| KITUI | 3 | 1.233 | 0.133 | 1.171 | 0.408 | 0.417 | 0.261 | 46.67 |
| KISII | 5 | 1.567 | 0.067 | 1.396 | 0.498 | 0.537 | 0.315 | 56.67 |
| ALUPE | 4 | 1.6 | 0.0133 | 1.486 | 0.524 | 0.544 | 0.333 | 60.00 |
| KIMIN | 7 | 2.133 | 0.1 | 1.833 | 0.678 | 0.647 | 0.41 | 70.00 |
| Mean | 15.8 | 2.033 | 0.22926 | 1.619 | 0.599 | 0.581 | 0.364 | 64.00 |
| Population | Alupe | ILRI | Kiminini | Kisii | Kitui |
|---|---|---|---|---|---|
| Alupe | - | 0.462 | 0.388 | 0.323 | 0.235 |
| ILRI | 0.393 | - | 0.636 | 0.440 | 0.327 |
| Kiminini | 0.448 | 0.307 | - | 0.399 | 0.299 |
| Kisii | 0.467 | 0.392 | 0.446 | - | 0.247 |
| Kitui | 0.510 | 0.441 | 0.413 | 0.503 | - |
| Source | Degree of Freedom | Sum of Squares | Mean Squares | Estimated Variance | Variation (%) | p Values |
|---|---|---|---|---|---|---|
| Among Populations | 4 | 43.440 | 10.860 | 0.155 | 2% | 0.023 |
| Among Individual | 74 | 619.649 | 8.374 | 1.215 | 17% | 0.001 |
| Within Individual | 79 | 469.500 | 5.943 | 5.943 | 81% | 0.001 |
| Total | 157 | 1132.589 | 7.313 | 100% | ||
| FST = 0.021 and Nm = 11.580 | ||||||
| Marker | Forward Primer | Reverse Primer | Ta (°C) | Allele Size (bp) | Repeat Motif |
|---|---|---|---|---|---|
| Brz0012 | ACTCAAACAATCTCCAACACG | CCCACAAATGGTGAATGTAAC | 59 | 160 | (AT)8 |
| Brz0028 | CATGGACAAGGAGAAGATTGA | TGGGAGTTAACATTAGTGTTTTT | 57 | 158 | (TA)8 |
| Brz0029 | TTTGTGCCAAAGTCCAAATAG | TATTCCAGCTTCTTCTGCCTA | 56 | 150 | (AG)14 |
| Brz0067 | TTAGATTCCTCAGGACATTGG | TCCTATATGCCGTCGTACTCA | 51 | 156 | (AT)9 |
| Brz0076 | CCTAGAATGCGGAAGTAGTGA | TTACGTGTTCCTCGACTCAAC | 58 | 151 | (AT)7 |
| Brz0087 | TTCCCCCACTACTCATCTCA | AACAGCACACCGTAGCAAGT | 60 | 243 | (GA)9 |
| Brz0092 | TTGATCAGTGGGAGGTAGGA | TGAAACTTGTCCCTTTTTCG | 54 | 251 | (AT)6 |
| Brz0100 | CCATCTGCAATTATTCAGGAAA | GTTCTTGGTGCTTGACCATT | 56 | 256 | (AT)11 |
| Brz0115 | AATTCATGATCGGAGCACAT | TGAACAATGGCTTTGAATGA | 59 | 252 | (AT)6 |
| Brz0117 | AGCTAAGGGGCTACTGTTGG | CGCGATCTCCAAAATGTAAT | 60 | 260 | (TA)5 |
| Brz0118 | AGGAGGTCCAAATCACCAAT | CGTCAGCAATTCGTACCAC | 57 | 252 | (CT)11 |
| Brz0122 | CATTGCTCCTCTCGCACTAT | CTGCAGTTAGCAGGTTGGTT | 57 | 253 | (CA)6 |
| Brz0130 | TCCTTTCATGAACCCCTGTA | CATCGCACGCTTATATGACA | 57 | 248 | (CT)14 |
| Brz0149 | GCAAGACCGCTGTTAGAGAA | CTAACATGGACACCGCTCTT | 57 | 245 | (AT)11 |
| Brz0156 | GCCATGATGTTTCATTGGTT | TTTTGCACCTTTCATTGCTT | 58 | 260 | (AC)7 |
| Brz0203 | CGCTTGAGAAGCTAGCAAGT | TAGCCTTTTGCATGGGTTAG | 57 | 301 | (GA)8 |
| Brz0212 | ACTCATTTTCACACGCACAA | CGAAGAATTGCAGCAGAAGT | 57 | 301 | (CA)5 |
| Brz0213 | TGAAGCCCTTTCTAAATGATG | GAACTAGGAAGCCATGGACA | 57 | 296 | (CA)7 |
| Brz0214 | TCTGGTGTCTCTTTGCTCCT | TCCATGGTACCTGAATGACA | 57 | 309 | (AT)8 |
| Brz0235 | CACACTCACACACGGAGAGA | CATCCAGAGCCTGATGAAGT | 57 | 298 | (TC)9 |
| Brz3002 | GCTGGAATCAGAATCGATGA | GAACTGCAGTGGCTGATCTT | 57 | 160 | (AAT)7 |
| Brz3009 | AGACTCTGTGCGGGAAATTA | ACTTCGCTTGTCCTACTTGG | 55 | 151 | (AAT)10 |
| Ecotype | Species | Status | Location | Alt. (m a.s.l.) | Lat. (S) | Lon. (E) | Collection Year |
|---|---|---|---|---|---|---|---|
| ke_1 | Brachiaria spp. | Wild | ILRI Farm | 1761 | 1.27085 | 36.72204 | 2013 |
| ke_2 | Brachiaria spp. | Wild | ILRI Farm | 1783 | 1.27091 | 36.72200 | 2013 |
| ke_3 | Brachiaria spp. | Wild | ILRI Farm | 1787 | 1.27117 | 36.72206 | 2013 |
| ke_4 | Brachiaria spp. | Wild | ILRI Farm | 1805 | 1.27152 | 36.72212 | 2013 |
| ke_5 | Brachiaria spp. | Wild | ILRI Farm | 1798 | 1.27306 | 36.72255 | 2013 |
| ke_6 | Brachiaria spp. | Wild | ILRI Farm | 1804 | 1.27307 | 36.72384 | 2013 |
| ke_7 | Brachiaria spp. | Wild | ILRI Farm | 1810 | 1.27292 | 36.72390 | 2013 |
| ke_8 | Brachiaria spp. | Wild | ILRI Farm | 1813 | 1.27281 | 36.72404 | 2013 |
| ke_9 | Brachiaria spp. | Wild | ILRI Farm | 1815 | 1.27269 | 36.72436 | 2013 |
| ke_10 | Brachiaria spp. | Wild | ILRI Farm | 1814 | 1.27262 | 36.72483 | 2013 |
| ke_11 | Brachiaria spp. | Wild | ILRI Farm | 1808 | 1.27275 | 36.72517 | 2013 |
| ke_12 | Brachiaria spp. | Wild | ILRI Farm | 1871 | 1.27077 | 36.72224 | 2013 |
| ke_13 | Brachiaria spp. | Wild | ILRI Farm | 1814 | 1.27076 | 36.72532 | 2013 |
| ke_14 | Brachiaria spp. | Wild | ILRI Farm | 1870 | 1.27073 | 36.72562 | 2013 |
| ke_15 | Brachiaria spp. | Wild | ILRI Farm | 1852 | 1.27088 | 36.72697 | 2013 |
| ke_16 | Brachiaria spp. | Wild | ILRI Farm | 1851 | 1.27091 | 36.72702 | 2013 |
| ke_17 | Brachiaria spp. | Wild | ILRI Farm | 1840 | 1.27135 | 36.72716 | 2013 |
| ke_18 | Brachiaria spp. | Wild | ILRI Farm | 1836 | 1.27152 | 36.72699 | 2013 |
| ke_19 | Brachiaria spp. | Wild | ILRI Farm | 1832 | 1.27214 | 36.72649 | 2013 |
| ke_20 | Brachiaria spp. | Wild | ILRI Farm | 1830 | 1.27236 | 36.72605 | 2013 |
| ke_21 | Brachiaria spp. | Wild | ILRI Farm | 1828 | 1.2725 | 36.72592 | 2013 |
| ke_22 | Brachiaria spp. | Wild | ILRI Farm | 1823 | 1.27268 | 36.72547 | 2013 |
| ke_23 | Brachiaria spp. | Wild | ILRI Farm | 1825 | 1.27263 | 36.72520 | 2013 |
| ke_24 | Brachiaria spp. | Wild | ILRI Farm | 1825 | 1.27273 | 36.72519 | 2013 |
| ke_25 | Brachiaria spp. | Wild | ILRI Farm | 1825 | 1.27261 | 36.72560 | 2013 |
| ke_26 | Brachiaria spp. | Wild | ILRI Farm | 1833 | 1.27213 | 36.72660 | 2013 |
| ke_27 | Brachiaria spp. | Wild | ILRI Farm | 1835 | 1.27196 | 36.72673 | 2013 |
| ke_28 | Brachiaria spp. | Wild | ILRI Farm | 1843 | 1.27144 | 36.72709 | 2013 |
| ke_29 | Brachiaria spp. | Wild | ILRI Farm | 1852 | 1.27109 | 36.72713 | 2013 |
| ke_30 | Brachiaria spp. | Wild | ILRI Farm | 1876 | 1.27067 | 36.72585 | 2013 |
| ke_31 | Brachiaria spp. | Wild | ILRI Farm | 1837 | 1.27086 | 36.72210 | 2014 |
| ke_32 | Brachiaria spp. | Wild | ILRI Farm | 1882 | 1.27084 | 36.72208 | 2014 |
| ke_33 | Brachiaria spp. | Wild | ILRI Farm | 1854 | 1.27252 | 36.72235 | 2014 |
| ke_34 | Brachiaria spp. | Wild | ILRI Farm | 1839 | 1.27264 | 36.72424 | 2014 |
| ke_35 | Brachiaria spp. | Wild | ILRI Farm | 1826 | 1.27274 | 36.72518 | 2014 |
| ke_36 | Brachiaria spp. | Wild | ILRI Farm | 1824 | 1.27233 | 36.72612 | 2014 |
| ke_37 | Brachiaria spp. | Wild | ILRI Farm | 1830 | 1.27257 | 36.72567 | 2014 |
| ke_38 | Brachiaria spp. | Wild | ILRI Farm | 1835 | 1.27165 | 36.72692 | 2014 |
| ke_39 | Brachiaria spp. | Wild | ILRI Farm | 1847 | 1.27101 | 36.72718 | 2014 |
| ke_40 | Brachiaria spp. | Wild | ILRI Farm | 1871 | 1.27077 | 36.72536 | 2014 |
| ke_41 | Brachiaria spp. | Wild | ILRI Farm | 1866 | 1.2708 | 36.72210 | 2014 |
| ke_42 | Brachiaria spp. | Wild | ILRI Farm | 1859 | 1.27134 | 36.72213 | 2014 |
| ke_43 | Brachiaria spp. | Wild | ILRI Farm | 1842 | 1.27285 | 36.72249 | 2014 |
| ke_44 | Brachiaria spp. | Wild | ILRI Farm | 1835 | 1.27242 | 36.72230 | 2014 |
| ke_45 | Brachiaria spp. | Wild | ILRI Farm | 1829 | 1.2734 | 36.72302 | 2014 |
| ke_46 | Brachiaria spp. | Wild | ILRI Farm | 1828 | 1.27315 | 36.72381 | 2014 |
| ke_47 | Brachiaria spp. | Wild | ILRI Farm | 1829 | 1.27271 | 36.72427 | 2014 |
| ke_48 | Brachiaria spp. | Wild | ILRI Farm | 1828 | 1.27269 | 36.72454 | 2014 |
| ke_49 | Brachiaria spp. | Wild | ILRI Farm | 1816 | 1.27261 | 36.72550 | 2014 |
| ke_50 | Brachiaria spp. | Wild | ILRI Farm | 1829 | 1.2717 | 36.72688 | 2014 |
| ke_51 | Brachiaria spp. | Wild | Kitui | 1163 | NA | NA | 2014 |
| ke_52 | Brachiaria spp. | Wild | Kitui | 1163 | NA | NA | 2014 |
| ke_53 | Brachiaria spp. | Wild | Kitui | 1163 | NA | NA | 2014 |
| ke_54 | Brachiaria spp. | Wild | ILRI Farm | 1754 | 1.27778 | 36.38821 | 2014 |
| ke_55 | Brachiaria spp. | Wild | ILRI Farm | 1857 | 1.2708 | 36.72206 | 2014 |
| ke_56 | Brachiaria spp. | Wild | ILRI Farm | 1856 | 1.27284 | 36.72204 | 2014 |
| ke_57 | Brachiaria spp. | Wild | ILRI Farm | 1844 | 1.27162 | 36.72208 | 2014 |
| ke_58 | Brachiaria spp. | Wild | ILRI Farm | 1840 | 1.27203 | 36.72217 | 2014 |
| ke_59 | Brachiaria spp. | Wild | ILRI Farm | 1822 | 1.2732 | 36.72357 | 2014 |
| ke_60 | Brachiaria spp. | Wild | ILRI Farm | 1822 | 1.27321 | 36.72358 | 2014 |
| ke_61 | Brachiaria spp. | Wild | ILRI Farm | 1810 | 1.27281 | 36.72506 | 2014 |
| ke_62 | Brachiaria spp. | Wild | ILRI Farm | 1821 | 1.27176 | 36.72678 | 2014 |
| ke_63 | Brachiaria spp. | Wild | ILRI Farm | 1824 | 1.27155 | 36.72697 | 2014 |
| ke_67 | Brachiaria spp. | Wild | Kisii | 1750 | 0.68575 | 34.78978 | 2014 |
| ke_68 | Brachiaria spp. | Wild | Kisii | 1750 | 0.68486 | 34.78914 | 2014 |
| ke_69 | Brachiaria spp. | Wild | Kisii | 1750 | 0.68484 | 34.78910 | 2014 |
| ke_70 | Brachiaria spp. | Wild | Kisii | 1750 | 0.68471 | 34.78896 | 2014 |
| ke_71 | Brachiaria spp. | Wild | Kisii | 1750 | 0.68473 | 34.78884 | 2014 |
| ke_72 | Brachiaria spp. | Wild | Alupe | 1200 | 0.49766 | 34.12480 | 2014 |
| ke_73 | Brachiaria spp. | Wild | Alupe | 1200 | 0.49781 | 34.12480 | 2014 |
| ke_74 | Brachiaria spp. | Wild | Alupe | 1200 | 0.49847 | 34.12319 | 2014 |
| ke_76 | Brachiaria spp. | Wild | Alupe | 1200 | 0.49855 | 34.12284 | 2014 |
| ke_82 | Brachiaria spp. | Wild | Kiminini | 1750 | 0.89104 | 34.91368 | 2014 |
| ke_83 | Brachiaria spp. | Wild | Kiminini | 1750 | 0.89102 | 34.91378 | 2014 |
| ke_84 | Brachiaria spp. | Wild | Kiminini | 1750 | 0.89126 | 34.91338 | 2014 |
| ke_85 | Brachiaria spp. | Wild | Kiminini | 1750 | 0.89144 | 34.91310 | 2014 |
| ke_86 | Brachiaria spp. | Wild | Kiminini | 1750 | 0.89139 | 34.91302 | 2014 |
| ke_87 | Brachiaria spp. | Wild | Kiminini | 1750 | 0.8913 | 34.91272 | 2014 |
| ke_88 | Brachiaria spp. | Wild | Kiminini | 1750 | 0.89131 | 34.91264 | 2014 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ondabu, N.; Maina, S.; Kimani, W.; Njarui, D.; Djikeng, A.; Ghimire, S. Molecular Characterizations of Kenyan Brachiaria Grass Ecotypes with Microsatellite (SSR) Markers. Agronomy 2017, 7, 8. https://doi.org/10.3390/agronomy7010008
Ondabu N, Maina S, Kimani W, Njarui D, Djikeng A, Ghimire S. Molecular Characterizations of Kenyan Brachiaria Grass Ecotypes with Microsatellite (SSR) Markers. Agronomy. 2017; 7(1):8. https://doi.org/10.3390/agronomy7010008
Chicago/Turabian StyleOndabu, Naftali, Solomon Maina, Wilson Kimani, Donald Njarui, Appolinaire Djikeng, and Sita Ghimire. 2017. "Molecular Characterizations of Kenyan Brachiaria Grass Ecotypes with Microsatellite (SSR) Markers" Agronomy 7, no. 1: 8. https://doi.org/10.3390/agronomy7010008
APA StyleOndabu, N., Maina, S., Kimani, W., Njarui, D., Djikeng, A., & Ghimire, S. (2017). Molecular Characterizations of Kenyan Brachiaria Grass Ecotypes with Microsatellite (SSR) Markers. Agronomy, 7(1), 8. https://doi.org/10.3390/agronomy7010008





