Abstract
Due to the widespread distribution of F. oxysporum, the search for mechanisms of tolerance to this disease in Solanum lycopersicum L. is an ongoing endeavor. This research aimed to identify F. oxysporum-tolerant genotypes at the germination and seedling stages in order to use them as sources of resistance. Ninety-six tomato lines were inoculated with the F. oxysporum strain with NCBI accession key PQ187438. The germination test was carried out in a germination chamber at a constant temperature of 28 ± 2 °C with 70 ± 5% relative humidity in darkness for the first 3 days and then 7 days with light. Clustering and discriminant analysis identified 14 genotypes with tolerance, showing great seed vigor and lower disease severity. Seedling evaluation was conducted in a floating raft system for 10 days after inoculation. Nine genotypes showed greater tolerance to the pathogen by developing a larger leaf area and accumulating more dry matter (p ≤ 0.05). No genotypes with tolerance were identified at both phenological stages (germination and seedling), indicating that tolerance mechanisms are independent at both phenological stages, so genotype selection should be carried out independently.
1. Introduction
Tomato (Solanum lycopersicum L.) is an important part of international culinary culture, consumed raw, cooked or processed, and is a significant source of nutrients, including lycopene, β-carotene, flavonoids and vitamin C [1]. World production of this crop in 2023 was 192.32 million tons harvested on an area of 5.4 million ha [2]. Due to its widespread global distribution, this vegetable is frequently exposed to increasingly severe biotic and abiotic stressors that diminish its yield (diseases, pests, extreme temperatures, drought, salinity, nutritional deficiencies, etc.) [3].
This vegetable, native to South America, possesses great genetic diversity thanks to the presence of 16 related wild species, among them: Solanum pimpinellifolium, Solanum pennellii, Solanum habrochaites, Solanum peruvianum and Solanum chilense [3]. However, domestication and selection have caused gene loss, which hampers crop resilience [4] and makes the task of breeding for tolerance to adverse factors arduous.
Tomatoes are attacked by more than 20 fungal diseases [5], of which Fusarium oxysporum (Schlechtendahl emend. Snyder and Hansen) is one of the most devastating. Its presence can occur from germination [6,7] to fully developed crops, which can decrease yields by up to 80%, and even to devastating levels [8,9].
F. oxysporum produces three types of asexual infectious spores: macroconidia, microconidia, and chlamydospores; a sexual stage for F. oxysporum has not been described [10]. Infection begins in exudates near the root by emission of spore germ tubes that invade and colonize the root xylem [11,12]. By transporting macro- and microconidia through the vascular tissue, similarly to nutrients, it enables infection throughout the plant [11,12]. The degree of damage depends on the biotrophic or necrotrophic state of the fungus [11].
Initially, the disease causes a decrease in plant development, e.g., plant height, leaf area, and shoot and root dry weight [13,14], and the visible symptoms start on lower leaves, which show wilting, vein discoloration and leaf blade yellowing, which later develop throughout the plant, resulting in a brown color [8,12]; this causes stunted growth and premature fruit ripening, and, ultimately, premature death of the plant [7].
Plants have the innate ability to defend themselves. At the genetic-molecular level, this is due to the interaction between endogenous receptors and derived molecules that determine the patterns and speed of expression of proteins related to plant defense. The difference between resistance and susceptibility depends on the activation rate and intensity of expression of the genes involved [15,16].
In plants, three possible signaling pathways activated by the presence of a pathogen have been identified. The most common is the nuclear-targeted signal, which triggers gene reprogramming. The second is organelle-targeted, where regulation occurs at the level of a particular metabolic pathway. Finally, the third pathway is manifested by changes in the cytoplasm [17,18]. These mechanisms are related to the signaling pathways of salicylic acid (SA), jasmonic acid (JA) and reactive oxygen species (ROS), substances that are produced in the initial presence of pathogens [19,20,21] and that allow for cell restructuring by reorganizing the cytoskeleton in order to mobilize defense resources.
In crop breeding, developing disease-tolerant varieties is a primary objective to minimize significant impacts on yield [5]. This strategy, together with integrated pest and disease management, promotes more sustainable agriculture by reducing the use of pesticides that can harm the environment and human health [5,22].
Having a wide genetic diversity in germplasm characterized morphologically, molecularly and agronomically [3,23] facilitates the identification of genes that encode resistance proteins, transcription factors and enzymes related to plant defense and adaptation to disease infection [16]. These genes can modify root proliferation, membrane permeability, and exudation of phenolic compounds such as caffeic acid, chlorogenic acid and antimicrobial proteins in the host [24].
The sources of resistance to Fusarium are varied. In tomato, major genes I-1, I-2, I-3 and I-7, originating from the wild tomato relatives S. pimpinellifolium and S. pennellii [22], confer resistance and protect the crop against different strains with widespread global distribution; however, the great genetic variability of the fungus requires a constant search for new genes and tolerance mechanisms given the genetic uniformity of the crop [22]. On the other hand, plant-pathogen interactions change according to plant development [25], a phenomenon known as ontogenetic resistance, which involves physiological and structural changes that depend on the phenological phase in which a disease infection occurs [26,27]. This requires the design of economical, short-term strategies that allow for the efficient evaluation of a large number of genotypes at different phenological stages to identify those resistant to adverse factors, whether biotic or abiotic, as proposed by Deanda-Tovar et al. [28].
This research aimed to identify tomato genotypes tolerant to F. oxysporum during germination and initial seedling development, with the purpose of using them as sources of resistance in breeding and generating short-term methods that enable the efficient evaluation of a large number of genotypes in the presence of F. oxysporum at both phenological stages.
2. Materials and Methods
Ninety-four homozygous lines (F12 to F15) from the Tomato Genetic Breeding Project operated by Chapingo Autonomous University (UACh), derived from 25 crosses between commercial hybrids and their respective segregating populations by individual selection, were evaluated. Table S1 presents some characteristics of the evaluated lines. Additionally, the Imperial® (Enza Zaden, Salinas, CA, USA) and Topanga® (Rogers Seeds, Boise, ID, USA) hybrids, both resistant to Fusarium, races 1 and 2, were evaluated.
The F. oxysporum strain used was obtained from the Phytopathogenic Fungi Bank of UACH’s Department of Agricultural Parasitology, isolated from tomato roots grown in the state of Sinaloa, Mexico. Its morphological characterization was performed according to Issac et al. [29] and using the identification guides of Leslie and Summerell [30]. Molecular characterization, DNA extraction, PCR amplification and sequencing of the strain were performed using the methodology described by Robles-Yerena et al. [31] with the primers ITS5/ITS4 [32] and RBP2-5F/RPB2-7R [33]. Fragment sequencing was performed by the Macrogen® company (Seoul, Republic of Korea), and this was compared and registered in the NCBI database using the BLASTn tool (PQ187438).
The strain was reactivated on PDA (200 g of potato, 15 g of dextrose, 20 g of agar-agar in 1 L of distilled water) at 25 °C for 10 days. A spore solution was then prepared with sterile water by liquefying the mycelium from 10 Petri dishes per liter of water and adding 5 drops of Tween 20® as an emulsifier. The inoculum was prepared from fresh, sporulating cultures, with a concentration of 2.2 ± 0.75 ×106 spores/mL of F. oxysporum, measured with a hematocytometer. This concentration ensured infection of the genotypes, since the minimum recommended concentration to achieve this in a hydroponic system is at least 1 ×106 spores/mL [34].
2.1. Response of Tomato Genotypes to F. oxysporum During Germination
The experiment was conducted in October 2023 in a germination chamber at 28 ± 2 °C and 70 ± 5% relative humidity (Lab-Tech Inc.®, model D-7140, Hopkinton, MA, USA), in darkness for the first 3 days and then under artificial light (linear fluorescent lamp, Philips brand, model F32T8/TL841, 32 W 800 Series, Amsterdam, The Netherlands) for 7 more days.
The experimental unit (EU) was a 5 cm diameter Petri dish with filter paper and 25 seeds. Two mL of the inoculum solution was added to the inoculated EUs, while 2 mL of distilled water was added to the control EUs. The dishes were irrigated with 0.5 mL of sterile distilled water on days 3, 6 and 9 after sowing (DAS). A randomized complete block experimental design with 4 replicates was used. The spatial arrangement of the EUs within the chamber defined the corresponding blocks.
The number of germinated seeds () and the number of seeds damaged by F. oxysporum () were recorded daily for 10 days.
The Maguire [35] germination rate index () was calculated:
and the damage rate index ():
where i = 1, 2 … n; 0, number of counts performed; number of germinated seeds in counts and , respectively; = number of damaged seeds in counts and , respectively.
The Fusarium incidence () in the seed was obtained with the formula:
The area under the disease progress curve () was calculated with:
where number of damaged seeds in counts i and , respectively; number of daily counts performed after inoculation, n = 10.
At 10 DAS, a sample of 5 seedlings per EU was obtained, and the average radicle length (RAL, in cm), stem length (SL, in cm), and total length (TL, in cm; TL = RAL + SL) were recorded. Total dry weight (DW, in mg) was obtained by drying the EU-generated material in an oven (Riossa®, model H-48, series 301007, Mexico city, Mexico) at 55 °C to constant, then using an OHAUS Discovery analytical balance (310 g ± 0.1 mg; OHAUS Corp., Parsippany, NJ, USA).
2.2. Response of Tomato Genotypes to F. oxysporum During Seedling Stage
The test was conducted in May 2023 in a medium-tech full-vent greenhouse with a 600-gauge plastic cover and 80% light transmission, relative humidity of 70 ± 10%, minimum temperature of 8.3 °C, and maximum temperature of 32.1 °C, with vents covered with anti-aphid mesh, located at the UACh Experimental Agricultural Field (19°29′23.5″ N 98°52′26.6″ W and 2267 masl).
The genotypes were sown in 2 × 2 × 2 cm agricultural sponge (Oasis®, Columbus, OH, USA). To facilitate uniform seedling infection, twenty DAS, the roots of the seedlings were cut to a length of two centimeters and immersed in the solution containing the F. oxysporum inoculum for 24; the control was immersed in water for the same period. They were subsequently transplanted onto a floating raft system, using polystyrene plates in containers measuring 120 × 240 × 20 cm and with a capacity of 450 L. A density of 166 plants·m2 was used. The nutrient solution used was Cadahia [36] for tomato. Due to the short duration of the trial (11 days), the nutrient solution, both for the control and for the Fusarium treatments, was maintained throughout the experiment. The nutrient solution was oxygenated using air pumps (Elite, 802) that maintained an O2 concentration of 5 ± 1 ppm.
The experimental unit (EU) consisted of five seedlings. A randomized complete block experimental design was used. Four replicates were used for inoculated seedlings, and two for controls.
The traits evaluated were:
Final plant height (PH, in cm). Average of three EU plants measured 9 days after inoculation (DAI).
At 10 DAI, digital photographs with known scales were obtained. They were subsequently processed using the ImageJ package version 1.54h [37]; a spatial calibration was performed to obtain root length (RL, in cm) and leaf area (LA in cm2).
The accumulated dry matter of roots (RDW, in g) and of the aerial part (ADW, in g) was determined. To do this, the structures were separated and placed in a forced-air dryer at 55 °C for 5 days, and weights were obtained using an OHAUS analytical balance (310 g ± 0.1 mg; OHAUS Corp., Parsippany, NJ, USA).
The seedling height growth rate (GR, in cm·d−1) was obtained from four measurements (0, 3, 6, and 9 DAI) and regressions using the exponential model in each EU.
2.3. Statistical Analysis
The same analyses were performed at both the germination and seedling stages. To eliminate the effects of genotypes due to differences in their development, indices were obtained by dividing the response obtained for the variable recorded against F. oxysporum in each EU by the average response obtained in the absence of the disease (control) of the corresponding genotype.
With the indices of all the variables evaluated, a cluster analysis was performed using Gower’s distances [38] and Ward’s minimum variance algorithm. The cutoff height was determined using Hotelling’s [39] pseudo-t2 and the pseudo-F [40]. To corroborate these groupings and verify the importance of the indices in forming groups, a discriminant analysis was applied along with resubstitution tests [40].
Analyses of variance and Tukey’s mean comparisons of the indices were performed, considering the identified groups and the genotypes nested within groups as sources of variance. Finally, Pearson correlations were calculated within each test and overall.
The analyses were performed using the SAS® Studio version 9.4 statistical package with the CLUSTER, DISCRIM, GLM and CORR procedures.
3. Results and Discussion
3.1. Morphological and Molecular Characterization of F. oxysporum
The isolate showed white, cottony and purple-violet mycelial growth on PDA medium and in germination tests and a growth rate of 1.2 cm per day (Figure 1A,B). Macroconidia were falcate, of short to medium length (29.2 ± 3 μm long by 3.7 ± 0.64 μm wide), thin-walled, triceptate, and with a short, slightly hooked apical cell and a pointed foot (Figure 1C). Microconidia were abundant in false heads on short monophialides, with dimensions of 7.3 ± 1.75 μm long and 2.95 ± 0.5 μm wide, without septa, generally elliptical, although oval ones were present (Figure 1D). Chlamydospores were formed singly or in pairs, with a terminal or intercalary shape and either smooth or rough walls (Figure 1E). All characteristics matched those reported by Leslie and Summerell [30] for F. oxysporum Schlechtendahl emend. Snyder and Hansen.
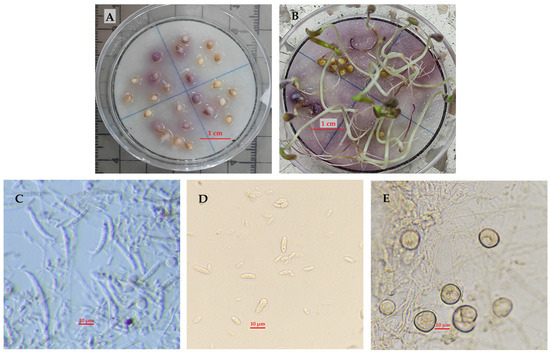
Figure 1.
Colony morphology and asexual reproduction structures of Fusarium oxysporum (Schlechtendahl emend. Snyder and Hansen) of GenBank accession strain PQ187438. (A) Seeds inoculated with F. oxysporum at 9 days after inoculation (DAI); (B) Seedlings damaged by F. oxysporum at 10 DAI; (C) Macroconidia and microconidia; (D) Microconidia; (E) Chlamydospores.
Molecular identification of the strain by sequencing the internal transcribed spacer (ITS5/ITS4) showed 100% similarity with the F. oxysporum (although it did not allow the special shape to be identified); sequence deposited at NCBI, which was consistent with the results of the morphological characterization. The obtained sequence was registered in the NCBI database under accession key PQ187438. This classification coincided with the results of the morphological characterization.
3.2. Response of Tomato Genotypes to F. oxysporum During Germination
At germination, three days after inoculation, lesions caused by F. oxysporum appeared on the seed, root and stem, with white and purple mycelial growth. New Fusarium colonies were obtained from 60 inoculated specimens selected at random, all of which had the same morphological characteristics as those originally inoculated, thus corroborating Koch’s postulates.
Cluster analysis using Gower’s [38] distances and Ward’s minimum variance algorithm (Figure 2) formed 4 groups at a semipartial r2 cutoff height of 0.055, determined using Hotelling’s [39] pseudo-t2 and pseudo-F [40]. The groups consisted of 52, 14, 10 and 20 genotypes.
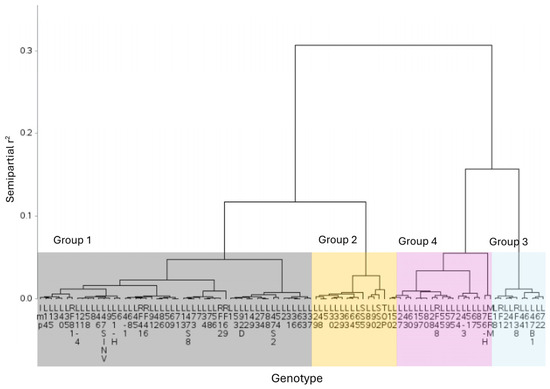
Figure 2.
Hierarchical dendrogram generated using Gower distances and Ward’s minimum variance algorithm based on tolerance indices for 96 tomato (Solanum lycopersicum L.) genotypes inoculated at germination with Fusarium oxysporum. A cutoff height of 0.055, determined using Hotelling’s pseudo-t2 and pseudo-F, formed 4 groups consisting of 52, 14, 10 and 20 genotypes.
Linear discriminant analysis verified the classification obtained by cluster analysis using two discriminant variables, DV1 and DV2, with eigenvalues of 5.1 and 1.28, respectively, and explained variance of 77% and 19%, respectively, both significant (p < 0.0001). Together, DV1 and DV2 accounted for 96% of the variance in the data.
The eigenvectors (Table 1) indicated that high DV1 values are associated with high expression of DRI, INC and AUC; that is, they identify susceptible genotypes, and vice versa. DV2 was positively associated with DW, RAL, SL and TL. Thus, high DV2 values identify genotypes with greater vigor, and vice versa. The resubstitution test showed that the assignments were correct for all genotypes.

Table 1.
Eigenvectors of the discriminant variables (DV1 and DV2) generated from indices of 96 tomato (Solanum lycopersicum L.) genotypes inoculated during the germination stage.
Figure 3 represents the genotypes based on DV1 and DV2. Group 2, with the greatest tolerance to F. oxysporum at germination, is located in quadrant 2 with the lowest disease damage (lower INC, DRI and AUC) and greatest vigor (higher GRI, DW, RAL, SL and TL). Group 1 had intermediate tolerance, while groups 3 and 4 were susceptible to F. oxysporum.
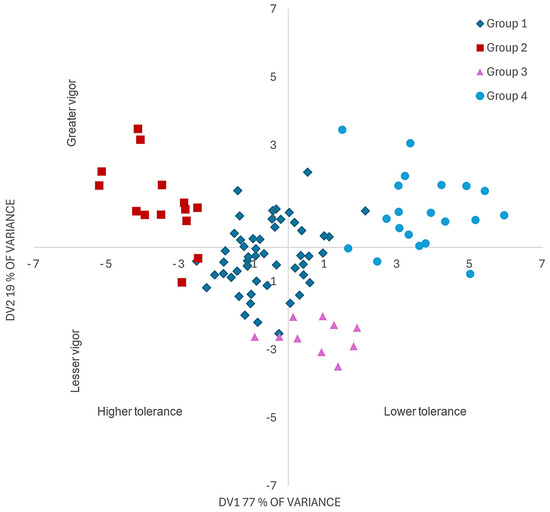
Figure 3.
Representation of discriminant variables for 96 tomato (Solanum lycopersicum L.) genotypes inoculated at germination with Fusarium oxysporum. DV1: discriminant variable 1, inversely associated with disease progress. DV2, discriminant variable 2, directly associated with seed vigor. Group 2 included 14 tolerant genotypes; group 1, included 52 genotypes with intermediate tolerance; and groups 3 and 4, with 10 and 20 genotypes, respectively, were susceptible.
Once the groupings were determined, analyses of variance were performed on the tolerance indices, considering the identified groups and the genotypes nested within groups as factors of variation (Table S2). These analyses detected differences (p ≤ 0.01) in the sources of variation in interest for all indices evaluated. The coefficients of variation were acceptable, with the exception of DW, RAL and TL, which were greater than 30%, although it should be noted that this is a test in which the genotypes are subjected to high stress, which increases random variation.
The comparison of Group means (Table 2) was consistent with the results of the discriminant analysis, as they identified the same pattern of behavior as in Figure 2; that is, Group 2 had the best performance against F. oxysporum, showing the greatest vigor, with higher expression of GER, GRI, DW, SL, RAL and TL (p ≤ 0.05), and less impairment of DRI, INC and AUC (p ≤ 0.05).

Table 2.
Comparisons of means of phenotypic traits for the Group factor in germination tests against Fusarium oxysporum.
Groups 3 and 4 suffered the greatest effects of F. oxysporum, with decreased vigor (GER, GRI, SL, RAL and TL). Group 3 decreased, relative to the control, by 22% in GER and 38% in GRI (p ≤ 0.05). In other crops, germination reductions (p ≤ 0.05) have been observed in the presence of Fusarium sp.; for example, in onion (Allium cepa L.) they ranged from 31 to 60% [41] and in sesame (Sesamum indicum L.) from 16.5 to 40% [42].
In the case of variables associated with vigor, the decreases observed in tomato in SL, RAL and DW (p ≤ 0.05) coincide with those of oats (Avena sativa) against F. sibiricum [43] and in corn (Zea mays L.) against F. verticillioides, which decreased germination percentage, plant height and number of leaves [44]. The attack of F. oxysporum in tomato seeds caused symptoms characteristic of phytopathogenic fungi and common to those observed in various crops, diminishing seed germination and vigor.
Since research such as that by Wang et al. [45] and Ghosal & Datta [46] indicates that germination damage caused by Fusarium is due to the alteration of protein, fiber and starch contents in the cotyledons, caused by toxic enzymes that degrade the extracellular wall (cellulase, polygalacturonase, and pectin lyase, among others), it is recommended that the activity of these enzymes be verified in Group 2. In addition, it will be necessary to verify whether the tolerance of genotypes in this group could be associated with the high production of enzymatic antioxidants such as phenylalanine ammonium lyase and lipoxygenase, which are involved in physiological, biochemical and defensive processes, as in the case of wheat [47].
Root growth of F. oxysporum-tolerant genotypes at germination (Table 2) was more than double that of the control (p ≤ 0.05); this behavior was detected as a tolerance mechanism in rice (Oryza sativa L.) against several Fusarium spp. species [48] and in tomato against Ralstonia [49]. This is why Meline et al. [49] proposed root length as a biomarker of fungal tolerance, which is consistent with the results obtained in this research.
Group 2 reacted to Fusarium with high root growth and proliferation levels, generating new xylem vessels that are the primary components of coordinated multiple resistance (CMR) to pathogen attack [50]. Other essential components of CMR include the production of phytoalexins, which are toxic to fungi [51], and the abundant production of tyloses, which clog xylem vessels and restrict the spread of the pathogen within the plant by trapping conidia and mycelium [12]. This set of traits should also be assessed in the case of tomato to verify their functionality as tolerance mechanisms.
Comparisons of genotype means within each group (Table S3) exhibited small variations; that is, the degree of tolerance or susceptibility was similar within each group, because the clustering and discriminant analyses achieved their purpose of classifying individuals. Therefore, the genotypes in Group 2 can be considered to have the same degree of tolerance, although it is necessary to verify which of the tolerance mechanisms are at work in each of the genotypes according to what was previously discussed.
3.3. Response of Tomato Genotypes to F. oxysporum During Seedling Stage
A quick way to assess the resistance of genotypes to disease-causing microorganisms is by exposing seedlings to high inoculum concentrations [52]. The floating raft system used allowed us to evaluate the behavior of a large number of genotypes in the presence of F. oxysporum and assess root development without causing damage to this structure. In contrast, evaluations of Fusarium spp. tolerance in soils or solid substrates impede the study of roots with high precision due to the loss of plant matter during root washing in tomato [34]. Although F. oxysporum is not an organism that subsists efficiently in an aqueous medium, such as a nutrient solution, inoculating the plant for 24 h with an exaggerated concentration of inoculum, a situation that does not occur in any production system, ensured that the pathogen invaded the plant through wounded roots, allowing the fungus to continue the infection inside the plant. For this reason, this method does not require the presence of the pathogen in the nutrient solution, a situation that significantly reduces the amount of inoculum needed.
In our evaluation, no lesions caused by F. oxysporum were observed in any of the genotypes, although seedling vigor was reduced (p ≤ 0.05), a situation similar to that observed in cape gooseberry (Physalis peruviana L.) [53]. This was due to the short 10 DAI test period compared to the symptom onset periods for F. oxysporum at 16 DAI in tomato [34], 14 DAI in Passiflora maliformis L. [54], and for Fusarium spp. at 15 and 50 DAI in garlic [55]. It is important to consider that F. oxysporum is a hemibiotrophic fungus that invades the vascular system without killing the plant until its infection cycles or environmental conditions change [9,56]; therefore, if disease symptoms are to be observed, the test should be prolonged.
The cluster analysis performed with Gower’s [38] distances and Ward’s minimum variance algorithm with a semipartial R2 cutoff height of 0.0621, determined using Hotelling’s [39] pseudo-t2 and pseudo-F [40], defined 4 groups consisting of 9, 14, 28 and 45 genotypes (Figure 4). Using these tools, the definition of genotype groups based on tolerance to F. oxysporum has been possible in species such as passionflower [57].
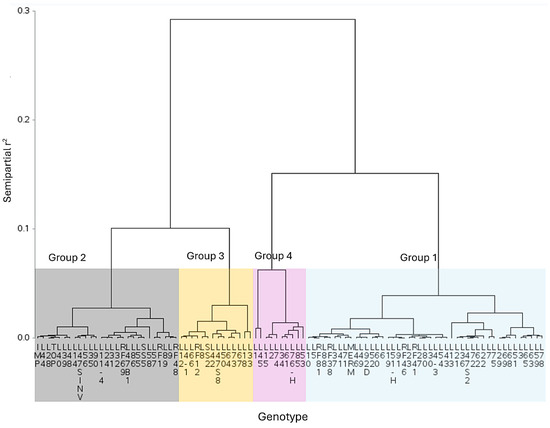
Figure 4.
Hierarchical dendrogram generated using Gower distances and Ward’s minimum variance algorithm based on tolerance indices to Fusarium oxysporum in 96 experimental tomato lines. Seedling stage test. A cutoff height of 0.0621, determined using Hotelling’s pseudo-t2 and pseudo-F, formed 4 groups consisting of 9, 14, 28 and 45 genotypes.
The classification of the cluster analysis was verified using linear discriminant analysis, where two discriminant variables (DV1 and DV2) had eigenvalues of 3.1 and 1.08, respectively, and explained variance of 68% and 23%, respectively; together, they explained 91% of the variance (p < 0.0001). The eigenvectors (Table 3) indicated that DV1 was positively associated with ADW, RDW, RL and LA. On the other hand, DV2 was positively associated with PH; that is, high values in DV1 and DV2 identify genotypes with greater tolerance, and vice versa.

Table 3.
Eigenvectors of the first two discriminant functions (DV1) and (DV2), generated from indices of 96 tomato (Solanum lycopersicum L.) genotypes during the seedling stage, inoculated with Fusarium oxysporum.
The resubstitution test [27] reassigned only two genotypes to the groups. The representation of genotypes based on DV1 and DV2 (Figure 5) places the genotypes of Group 4 with the greatest tolerance at seedling stage to F. oxysporum in quadrant 1, obtaining greater PH, ADW, RDM, RL and LA; that is, greater seedling development in the presence of the disease. Group 1 had intermediate tolerance, while Groups 2 and 3 were considered susceptible.
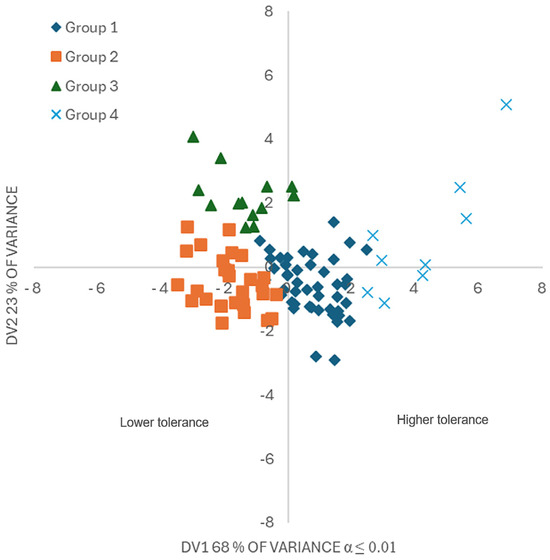
Figure 5.
Graphical representation of discriminant variables of 96 tomato (Solanum lycopersicum L.) genotypes inoculated in seedlings with Fusarium oxysporum. DV1: discriminant variable 1; high values indicate greater plant vigor and less disease damage, and vice versa; DV2, discriminant variable 2; high values indicate genotypes with greater seedling height and aerial dry weight. Group 4 included 9 tolerant genotypes; group 1 included 45 genotypes with intermediate tolerance; while groups 2 and 3, with 28 and 14 genotypes, respectively, were susceptible.
Analyses of variance (ANOVAs), which considered groups and genotypes nested within groups as sources of variation (Table S4), detected differences (p ≤ 0.01) in all sources of variation and traits evaluated; that is, the identified groups had different tolerances (p ≤ 0.01). The coefficients of variation were acceptable, with values less than 20%, except for RDW at 35%.
Comparisons of group means (Table 4) indicate that Group 4 had the greatest tolerance to F. oxysporum, showing higher ADW, RDW, RL and LA. Group 2 was the most susceptible, with the lowest development of PH, ADW, RDW, RL and LA, while Groups 1 and 3 were moderately affected by F. oxysporum. Research on wheat exposed to four isolates of F. graminearum showed a 39% decrease in seedling height [58], whereas celery exposed to F. oxysporum resulted in a 58% decrease in both seedling height and dry weight [59].

Table 4.
Comparisons of means of genotype groups against Fusarium oxysporum at seedling stage.
Genotypes belonging to a specific group exhibited similar behavior (p ≤ 0.05), as verified by comparisons of means (Table S5).
At the seedling stage, correlations between variables indicated that significant associations (p ≤ 0.01) occurred with ADW, RDW and LA, with values ranging from 0.6 to 0.8. In the germination test, associations were found between GER and GRI, DRI and AUC, and between SL and RL with ADW and RDW, with values ranging from 0.7 to 0.9. The rest of the values did not represent significant associations. Correlations between variables in the two tests did not detect significant associations (Table S6).
The classifications corroborated by canonical discriminant analyses (Table 5) show that tolerance to F. oxysporum at both phenological stages is independent; that is, the tolerance groups do not coincide with respect to the genotypes they contain. This result is consistent with the correlation analyses mentioned above (Table S6). Leyva-Mir et al. [60] evaluated in the field the tolerance in adult plants of 15 genotypes included in the present study, although in generations F6–F7, to Fusarium oxysporum; the tolerance classifications differed from those observed in this study. This situation is similar to that observed in cape gooseberry inoculated with Fusarium spp. during germination and seedling development [53], and to that observed in onion (Allium cepa) seedlings and bulbs infected with Fusarium under soil conditions, where the expression of resistance genes, identified with quantitative PCR, is not the same in the two phenological stages [61].

Table 5.
Composition of tomato genotypes within the groups resulting from the dendrogram and discriminant analysis in germination and seedling inoculated with Fusarium oxysporum.
Specifically in tomato, Zhang et al. [62] observed that tolerance to yellow leaf curl virus is differentially expressed throughout ontogenetic development; thus, young plants are more susceptible than adult plants, although there is a genotypic interaction, since the degree of tolerance among varieties varies.
Based on the above, the fact that in this research no genotypes with tolerance to F. oxysporum were identified at both germination and seedling stages (Table 5) suggests the presence of different genetic tolerance mechanisms, whose expression process is dynamic and affected by various factors such as the phenological stage, the environment, and the pathogen [53,61].
This implies that the fungus exhibits different levels of severity depending on the phenological stage of the host [63], since, for example, the thickness of root cell walls depends on the phenological stage, as lignification increases the capacity to act as a structural defense in plants with greater development [9]. Therefore, it is advisable to verify the presence of these physiological mechanisms in tolerant genotypes. In the present study, F. oxysporum damage was more severe during germination than in the seedling stage, where no visual symptoms were observed, although RL, SL and TL were reduced.
When identifying disease-tolerant genotypes, it is necessary to consider the plant-pathogen-environment interaction [6]. The aggressiveness and severity of the pathogen, the exposure time and the inoculum concentration are determining factors in the behavior of the pathogen, which can modify its morphology, physiology, metabolism and spectrum of proteins and metabolites secreted to achieve infection in plants, since they have different tolerance levels [64] according to their phenological development [65].
It is a fact that all the genotypes studied have the ability to defend themselves; however, there are differences in the ability to develop these defenses [15]. As a result, 14 tomato genotypes with underlying mechanisms of tolerance to F. oxysporum were identified during germination and 9 during seedling development. This may be associated with the response of the two commercial hybrids evaluated with vertical tolerance to races 1 and 2; both were susceptible to the disease at the seedling stage, and at germination, Imperial was moderately tolerant, and Topanga was tolerant. A challenge for the breeding of this species is to combine both complexes in a single variety in order to achieve long-term resistance to the disease; however, prolonged resistance has been repeatedly established as complex, demanding and challenging, because it involves traits governed by a high number of genes [65].
The traits studied in this research are quantitative, so they are associated with polygenic inheritance. The selection of this type of inheritance in breeding is a challenging and under-addressed problem in tomato [65], which is why the introgression of major genes that confer resistance in already improved cultivars is preferred [66]. Despite this, the methodology used allows the identification of Fusarium-tolerant genotypes associated with extensive root development as a tolerance mechanism by providing new vascular vessels as an adaptation strategy that reduces the impact of the disease.
Accurate phenotypic evaluations of quantitative traits in the presence of diseases, such as those carried out in this study, are highly useful tools for associating genetic and phenotypic information, which is necessary for genomic selection [67]. Additionally, these evaluations are suitable for short periods and with a large number of genotypes. Currently, GBS and GWAS analyses are being performed to associate SNPs with the quantitative traits evaluated in this research and to determine possible relationships with the expression of genes identified in the tomato genome.
4. Conclusions
Tomato tolerance to F. oxysporum at the germination stage and during seedling development is controlled by different sets of genes, as no tolerant genotypes were identified at both phenological stages, thereby establishing the need to study, in greater detail, the ontogenetic resistance of this crop.
Tests conducted on 96 genotypes allowed for an efficient, thorough and short-term assessment of the variation in tolerance to F. oxysporum.
Thus, 14 tolerant genotypes were identified during germination and 9 at the seedling stage, which will serve as germplasm for tomato breeding. However, it is necessary to corroborate the mechanisms that confer such resistance at different phenological stages.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/agronomy15092089/s1, Table S1: Phenotypic traits of tomato breeding lines developed under greenhouse conditions at Universidad Autónoma Chapingo; Table S2: Analysis of variance mean squares for phenotypic traits in 96 tomato lines inoculated with Fusarium oxysporum at the seed stage; Table S3: Mean comparisons among 96 tomato (Solanum lycopersicum L.) genotypes from all groups under Fusarium oxysporum challenge during germination. Table S4: Analysis of variance mean squares for phenotypic traits in 96 tomato lines inoculated with Fusarium oxysporum at the seedling stage; Table S5: Analysis of variance mean squares for phenotypic traits in 96 tomato lines inoculated with Fusarium oxysporum at the seedling stage; Table S6: Pearson correlation coefficients of phenotypic traits in germination and seedling stages.
Author Contributions
Conceptualization, A.E.P.-C., J.E.R.-P., J.S.-C. and S.G.L.-M.; methodology, A.E.P.-C., J.E.R.-P., J.S.-C., S.G.L.-M. and A.A.D.-T.; validation, A.E.P.-C., J.E.R.-P., M.H.-R. and A.A.D.-T.; formal analysis, A.E.P.-C., J.E.R.-P. and A.A.D.-T.; investigation, A.E.P.-C., J.E.R.-P. and A.A.D.-T.; resources, J.E.R.-P. and J.S.-C.; writing—original draft preparation, A.E.P.-C. and J.E.R.-P.; writing—review and editing, J.S.-C. and M.H.-R.; visualization, A.E.P.-C.; supervision, J.E.R.-P., J.S.-C., S.G.L.-M. and M.H.-R.; project administration, J.E.R.-P. and J.S.-C.; funding acquisition, J.E.R.-P. and J.S.-C. All authors have read and agreed to the published version of the manuscript.
Funding
This research received funding from Universidad Autónoma Chapingo through project D.G.I.P. 25001-EI.
Data Availability Statement
The original contributions presented in this study are included in the article and Supplementary Materials. Further inquiries can be directed to the corresponding author.
Acknowledgments
For their assistance in the conduction of the experiments, thanks to Jorge Luis Sánchez Galicia, Mariana Romualda Calzada Millán and Ricardo Gaspar Hernández.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Gerszberg, A.; Hnatuszko-Konka, K.; Kowalczyk, T.; Kononowicz, A.K. Tomato (Solanum lycopersicum L.) in the service of biotechnology. Plant Cell Tissue Organ Cult. 2015, 120, 881–902. [Google Scholar] [CrossRef]
- FAOSTAT. Crops and Livestock Products—Tomatoes: Production (Tonnes). Available online: https://www.fao.org/faostat/en/#data/QCL (accessed on 10 August 2025).
- Ramírez-Ojeda, G.; Rodríguez-Pérez, J.E.; Rodríguez-Guzmán, E.; Sahagún-Castellanos, J.; Chávez-Servia, J.L.; Peralta, I.E.; Barrera-Guzmán, L.Á. Distribution and climatic adaptation of wild tomato (Solanum lycopersicum L.) populations in Mexico. Plants 2022, 11, 2007. [Google Scholar] [CrossRef]
- Tamburino, R.; Sannino, L.; Cafasso, D.; Cantarella, C.; Orrù, L.; Cardi, T.; Cozzolino, S.; D’Agostino, N.; Scotti, N. Cultivated tomato (Solanum lycopersicum L.) suffered a severe cytoplasmic bottleneck during domestication: Implications from chloroplast genomes. Plants 2020, 9, 1443. [Google Scholar] [CrossRef]
- Rezk, A.; Abhary, M.; Akhkha, A. Tomato (Solanum lycopersicum L.) breeding strategies for biotic and abiotic stresses. In Advances in Plant Breeding Strategies: Vegetable Crops; Al-Khayri, J.M., Jain, S.M., Johnson, D.V., Eds.; Springer: Cham, Switzerland, 2021; pp. 363–405. [Google Scholar]
- Lamichhane, J.R.; Dürr, C.; Schwanck, A.A.; Robin, M.-H.; Sarthou, J.-P.; Cellier, V.; Messéan, A.; Aubertot, J.-N. Integrated management of damping-off diseases: A review. Agron. Sustain. Dev. 2017, 37, 10. [Google Scholar] [CrossRef]
- Kamil, D.; Toppo, R.S.; Devi, T.P.; Kumari, A. Diversity of seed-borne fungal phytopathogens. In Seed-Borne Diseases of Agricultural Crops: Detection, Diagnosis & Management; Springer: Singapore, 2020; pp. 293–306. [Google Scholar]
- Bawa, I. Management strategies of Fusarium wilt disease of tomato incited by Fusarium oxysporum f. sp. lycopersici (Sacc.): A review. Int. J. Adv. Acad. Res. 2016, 2, 31–42. [Google Scholar]
- Husaini, A.M.; Sakina, A.; Cambay, S.R. Host–pathogen interaction in Fusarium oxysporum infections: Where do we stand? Mol. Plant Microbe Interact. 2018, 31, 889–898. [Google Scholar] [CrossRef]
- McGovern, R.J. Management of tomato diseases caused by Fusarium oxysporum. Crop Prot. 2015, 73, 78–92. [Google Scholar] [CrossRef]
- Gordon, T.R. Fusarium oxysporum and the Fusarium wilt syndrome. Annu. Rev. Phytopathol. 2017, 55, 23–39. [Google Scholar] [CrossRef] [PubMed]
- Srinivas, C.; Nirmala Devi, D.; Narasimha Murthy, K.; Mohan, C.D.; Lakshmeesha, T.R.; Singh, B.; Kalagatur, N.K.; Niranjana, S.R.; Hashem, A.; Alqarawi, A.A.; et al. Fusarium oxysporum f. sp. lycopersici causal agent of vascular wilt disease of tomato: Biology to diversity–A review. Saudi J. Biol. Sci. 2019, 26, 1315–1324. [Google Scholar] [CrossRef]
- Razak, N.J.; Abass, M.H.; Awad, K.M. Multifactorial stressors: Linking Fusarium infection, heavy metals, and salinity to physiological dysfunction in tomato (Solanum lycopersicum L.). Thi-Qar J. Agric. Res. 2025, 14, 248–259. [Google Scholar]
- Yahaya, N.; Hayatu, M. Effect of Trichoderma harzianum on the incidence of Fusarium wilt disease, growth, and yield of selected tomato (Lycopersicon esculentum) varieties. Dutse J. Pure Appl. Sci. 2019, 5, 74–83. [Google Scholar]
- Tabassum, N.; Blilou, I. Cell-to-cell communication during plant-pathogen interaction. Mol. Plant Microbe Interact. 2022, 35, 98–108. [Google Scholar] [CrossRef]
- Ghosh, S.; Malukani, K.K.; Chandan, R.K.; Sonti, R.V.; Jha, G. How plants respond to pathogen attack: Interaction and communication. In Sensory Biology of Plants; Springer: Singapore, 2019; pp. 537–568. [Google Scholar]
- López-Zapata, S.P.; García-Jaramillo, D.J.; López, W.R.; Ceballos-Aguirre, N. Tomato (Solanum lycopersicum L.) and Fusarium oxysporum f. sp. lycopersici interaction: A review. Rev. UDCA Act. Divulg. Cient. 2021, 24. [Google Scholar] [CrossRef]
- Chitwood-Brown, J.; Vallad, G.E.; Lee, T.G.; Hutton, S.F. Breeding for resistance to Fusarium wilt of tomato: A review. Genes 2021, 12, 1673. [Google Scholar] [CrossRef]
- Cruz, J.L. Análisis de la Señalización Mediada por la Pared Celular y el Estrés Oxidativo en la Interacción Planta–Patógeno. Ph.D. Thesis, Universitat de València, Valencia, Spain, 2017. [Google Scholar]
- Naz, M.; Zhang, D.; Liao, K.; Chen, X.; Ahmed, N.; Wang, D.; Zhou, J.; Chen, Z. The past, present, and future of plant activators targeting the salicylic acid signaling pathway. Genes 2024, 15, 1237. [Google Scholar] [CrossRef] [PubMed]
- Zavala, J.A. Respuestas inmunológicas de las plantas frente al ataque de insectos. Cienc. Hoy 2010, 20, 52–59. [Google Scholar]
- Marín-Montes, I.M.; Rodríguez-Pérez, J.E.; Sahagún-Castellanos, J.; Hernández-Ibáñez, L.; Velasco-García, Á.M. Morphological and molecular variation in 55 native tomato collections from Mexico. Rev. Chapingo Ser. Hortic. 2016, 22, 117–132. [Google Scholar] [CrossRef]
- Gallé, Á.; Pelsőczi, A.; Benyó, D.; Podmaniczki, A.; Szabó-Hevér, Á.; Poór, P.; Tóth, B.; Horváth, E.; Erdei, L.; Csiszár, J. Systemic response to Fusarium graminearum and culmorum inoculations: Changes in detoxification of flag leaves in wheat. Cereal Res. Commun. 2022, 50, 1055–1063. [Google Scholar] [CrossRef]
- Driouich, A.; Follet-Gueye, M.L.; Vicré-Gibouin, M.; Hawes, M. Root border cells and secretions as critical elements in plant host defense. Curr. Opin. Plant Biol. 2013, 16, 489–495. [Google Scholar] [CrossRef]
- Ficke, A.; Gadoury, D.M.; Seem, R.C. Ontogenic resistance and plant disease management: A case study of grape powdery mildew. Phytopathology 2002, 92, 671–675. [Google Scholar] [CrossRef]
- Calonnec, A.; Jolivet, J.; Vivin, P.; Schnee, S. Pathogenicity traits correlate with the susceptible Vitis vinifera leaf physiology transition in the biotroph fungus Erysiphe necator: An adaptation to plant ontogenic resistance. Front. Plant Sci. 2018, 9, 1808. [Google Scholar] [CrossRef]
- Kus, J.V.; Zaton, K.; Sarkar, R.; Cameron, R.K. Age-related resistance in Arabidopsis is a developmentally regulated defense response to Pseudomonas syringae. Plant Cell 2002, 14, 479–490. [Google Scholar] [CrossRef] [PubMed]
- Deanda-Tovar, A.A.; Rodríguez-Pérez, J.E.; Sahagún-Castellanos, J.; Colinas-y-León, M.T.B.; Pérez-Rodríguez, P.; Paredes-Cervantes, A.E. Tomato lines tolerant to sodium chloride at early growth stages. Horticulturae 2025, 11, 532. [Google Scholar] [CrossRef]
- Isaac, M.R.; Leyva-Mir, S.G.; Sahagun-Castellanos, J.; Camara-Correia, K.; Tovar-Pedraza, J.M.; Rodriguez-Perez, J.E. Occurrence, identification, and pathogenicity of Fusarium spp. associated with tomato wilt in Mexico. Not. Bot. Horti Agrobot. Cluj-Napoca 2018, 46, 484–493. [Google Scholar] [CrossRef]
- Leslie, J.F.; Summerell, B.A. The Fusarium Laboratory Manual; John Wiley & Sons: Hoboken, NJ, USA, 2008. [Google Scholar] [CrossRef]
- Robles-Yerena, L.; Leyva Mir, S.G.; Cruz Gómez, A.; Camacho Tapia, M.; Nieto Ángel, D.; Tovar Pedraza, J.M. Fusarium oxysporum Schltdl. y Fusarium solani (Mart.) Sacc. causantes de la marchitez de plántulas de Pinus spp. en vivero. Rev. Mex. Cienc. For. 2016, 7, 25–36. [Google Scholar]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J.W. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerase II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef]
- Gayosso-Barragán, O.; López-Benítez, A.; Marroquín-Morales, J.Á.; Lopéz-Aguilar, K.; Hidalgo-Ramos, D.M.; Chávez-Aguilar, G. Evaluación de la respuesta de diferentes genotipos de tomate a Fusarium oxysporum raza 3. Rev. Mex. Cienc. Agríc. 2021, 12, 409–420. [Google Scholar]
- Maguire, J.D. Speed of germination-aid in selection and evaluation for seedling emergence and vigor. Crop Sci. 1962, 2, 176–177. [Google Scholar] [CrossRef]
- Cadahia López, C. Fertirrigación: Cultivos Hortícolas y Ornamentales, 1st ed.; Ediciones Mundi-Prensa: Madrid, España, 2000. [Google Scholar]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Gower, J.C. A comparison of some methods of cluster analysis. Biometrics 1967, 23, 623. [Google Scholar] [CrossRef]
- Hotelling, H. A generalized T test and measure of multivariate dispersion. In Proceedings of the Second Berkeley Symposium on Mathematical Statistics and Probability, Berkeley, CA, USA, 31 July–12 August 1950; University of California Press: Berkeley, CA, USA, 1951; Volume 1, pp. 23–41. [Google Scholar]
- Johnson, D.E. Métodos Multivariados Aplicados al Análisis de Datos, 1st ed.; Thompson Editores: Ciudad de México, México, 2000. [Google Scholar]
- Dabire, T.G.; Neya, B.F.; Somda, I.; Legrève, A. Pathogenicity study of some seed-borne fungi of onion (Allium cepa L.) from Burkina Faso. Int. J. Biol. Chem. Sci. 2021, 15, 1062–1072. [Google Scholar] [CrossRef]
- Nayyar, B.G.; Woodward, S.; Mur, L.A.; Akram, A.; Arshad, M.; Naqvi, S.S.; Akhund, S. Identification and pathogenicity of Fusarium species associated with sesame (Sesamum indicum L.) seeds from the Punjab, Pakistan. Physiol. Mol. Plant Pathol. 2018, 102, 128–135. [Google Scholar] [CrossRef]
- Chen, H.; White, J.F.; Malik, K.; Li, C. Molecular assessment of oat head blight fungus, including a new genus and species in a family of Nectriaceae. Int. J. Food Microbiol. 2024, 417, 110715. [Google Scholar] [CrossRef]
- Yasir, A.M.; Nasruddin, A. The evaluation effect of Fusarium verticillioides on seed quality in new high-yielding maize varieties. IOP Conf. Ser. Earth Environ. Sci. 2023, 1192, 012028. [Google Scholar] [CrossRef]
- Wang, J.; Wei, X.; Kamran, M.; Chen, T.; White, J.F.; Li, C. Quality and nutrition of oat seed as influenced by seed-borne fungal pathogens during storage. J. Plant Dis. Prot. 2022, 129, 243–252. [Google Scholar] [CrossRef]
- Ghosal, D.; Datta, B. Characterization, phylogenetic analysis and toxigenic potential of Fusarium incarnatum–equiseti species complex isolates associated with root rot disease in vegetables. J. Phytopathol. 2024, 172, e13362. [Google Scholar] [CrossRef]
- Khaledi, N.; Zare, L.; Hassani, F.; Moslemkhani, C. Current status of seed-borne Fusarium of wheat in Iran and its effect involved in seedling resistance and biochemical indicators. Indian Phytopathol. 2023, 76, 689–700. [Google Scholar] [CrossRef]
- Chhabra, R.; Kaur, N.; Bala, A. Differential biochemical response of basmati and non-basmati rice seeds upon bakanae (Fusarium fujikuroi) infection. Vegetos 2022, 36, 516–525. [Google Scholar] [CrossRef]
- Meline, V.; Hendrich, C.G.; Truchon, A.N.; Caldwell, D.; Hiles, R.; Leuschen-Kohl, R.; Tran, T.; Mitra, R.M.; Allen, C.; Iyer-Pascuzzi, A.S. Tomato deploys defence and growth simultaneously to resist bacterial wilt disease. Plant Cell Environ. 2023, 46, 3040–3058. [Google Scholar] [CrossRef]
- Gorshkov, V.; Tsers, I. Plant susceptible responses: The underestimated side of plant–pathogen interactions. Biol. Rev. 2022, 97, 45–66. [Google Scholar] [CrossRef]
- Liu, Y.; Esposto, D.; Mahdi, L.K.; Porzel, A.; Stark, P.; Hussain, H.; Scherr-Henning, A.; Isfort, S.; Bathe, U.; Acosta, I.F.; et al. Hordedane diterpenoid phytoalexins restrict Fusarium graminearum infection but enhance Bipolaris sorokiniana colonization of barley roots. Mol. Plant 2024, 17, 1307–1327. [Google Scholar] [CrossRef] [PubMed]
- Arunakumar, G.S.; Gnanesh, B.N. Evaluation of artificial inoculation methods to determine resistance reaction to dry root rot and black root rot disease in mulberry (Morus spp.). Arch. Phytopathol. Plant Prot. 2023, 56, 49–65. [Google Scholar] [CrossRef]
- López, M.M.; Aristizabal, A.M.O.; Echeverri, V.M.O. Control biológico contra Fusarium sp. en plantas de uchuva (Physalis peruviana L.) a nivel de invernadero mediante aplicación combinada de Trichoderma sp. y Bacillus sp. Rev. Investig. Agrar. Ambient. 2024, 15, 35–67. [Google Scholar] [CrossRef]
- Forero, R.; Ortiz, E.; De León, W.; Gómez, J.C.; Hoyos-Carvajal, L. Análisis de la resistencia a Fusarium oxysporum en plantas de Passiflora maliformis L. Rev. Colomb. Cienc. Hortic. 2015, 9, 197–208. [Google Scholar] [CrossRef]
- Delgado-Ortiz, J.C.; Ochoa-Fuentes, Y.M.; Cerna-Chávez, E.; Beltrán-Beache, M.; Rodríguez-Guerra, R.; Aguirre-Uribe, L.A.; Vázquez-Martínez, O. Patogenicidad de especies de Fusarium asociadas a la pudrición basal del ajo en el centro norte de México. Rev. Argent. Microbiol. 2016, 48, 222–228. [Google Scholar] [CrossRef]
- Ma, L.-J.; Geiser, D.M.; Proctor, R.H.; Rooney, A.P.; O’Donnell, K.; Trail, F.; Gardiner, D.M.; Manners, J.M.; Kazan, K. Fusarium pathogenomics. Annu. Rev. Microbiol. 2013, 67, 399–416. [Google Scholar] [CrossRef]
- Patiño-Pacheco, M.J.; Pérez-Cardona, O.Y. Evaluación de la Resistencia de Genotipos de Passifloras a Fusarium solani f. sp. passiflorae en Granadilla. Entramado 2021, 17, 256–267. [Google Scholar] [CrossRef]
- Semagn, K.; Henriquez, M.A.; Iqbal, M.; Brûlé-Babel, A.L.; Strenzke, K.; Ciechanowska, I.; Navabi, A.; N’Diaye, A.; Pozniak, C.; Spaner, D. Identification of Fusarium head blight sources of resistance and associated QTLs in historical and modern Canadian spring wheat. Front. Plant Sci. 2023, 14, 1190358. [Google Scholar] [CrossRef] [PubMed]
- Lori, G.A.; Malbran, I.; Mourelos, C.A.; Wolcan, S.M. First report of Fusarium oxysporum f. sp. apii race 2 causing Fusarium yellows on celery in Argentina. Plant Dis. 2016, 100, 1020. [Google Scholar] [CrossRef]
- Leyva-Mir, S.G.; González-Solano, C.M.; Rodríguez-Pérez, J.E.; Montalvo-Hernández, D. Comportamiento de líneas avanzadas de tomate (Solanum lycopersicum L.) a fitopatógenos en Chapingo, México. Rev. Chapingo Ser. Hortic. 2013, 19, 301–313. [Google Scholar] [CrossRef]
- Le, D.; Ameye, M.; Landschoot, S.; Audenaert, K.; Haesaert, G. Phenology-regulated defence mechanisms as drivers for Fusarium basal rot in onion (Allium cepa). Plant Pathol. 2022, 71, 1440–1453. [Google Scholar] [CrossRef]
- Zhang, J.R.; Liu, S.S.; Pan, L.L. Enhanced Age-Related Resistance to Tomato Yellow Leaf Curl Virus in Tomato Is Associated with Higher Basal Resistance. Front. Plant Sci. 2021, 12, 685382. [Google Scholar] [CrossRef] [PubMed]
- Chang, X.; Li, H.; Naeem, M.; Wu, X.; Yong, T.; Song, C.; Liu, T.; Chen, W.; Yang, W. Diversity of the seedborne fungi and pathogenicity of Fusarium species associated with intercropped soybean. Pathogens 2020, 9, 531. [Google Scholar] [CrossRef] [PubMed]
- Michielse, C.B.; Rep, M. Pathogen profile update: Fusarium oxysporum. Mol. Plant Pathol. 2009, 10, 311. [Google Scholar] [CrossRef]
- Robinson, R.A. Return to Resistance: Breeding Crops to Reduce Pesticide Dependence; AgAccess: Davis, CA, USA, 1996. [Google Scholar]
- Facundo-Angel, P.; Sahagún-Castellanos, J.; Rodríguez-Pérez, J.E.; Leyva-Mir, S.G. Molecular markers of pathogen resistance for assisted breeding of tomato (Solanum lycopersicum L.). Rev. Chapingo Ser. Hortic. 2024, 30, 21–33. [Google Scholar] [CrossRef]
- Tassone, M.R.; Bagnaresi, P.; Desiderio, F.; Bassolino, L.; Barchi, L.; Florio, F.E.; Sunseri, F.; Sirangelo, T.M.; Rotino, G.L.; Toppino, L. A Genomic BSAseq Approach for the Characterization of QTLs Underlying Resistance to Fusarium oxysporum in Eggplant. Cells 2022, 11, 2548. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).