Potato Leaf Chlorophyll Content Estimation through Radiative Transfer Modeling and Active Learning
Abstract
:1. Introduction
2. Materials and Methods
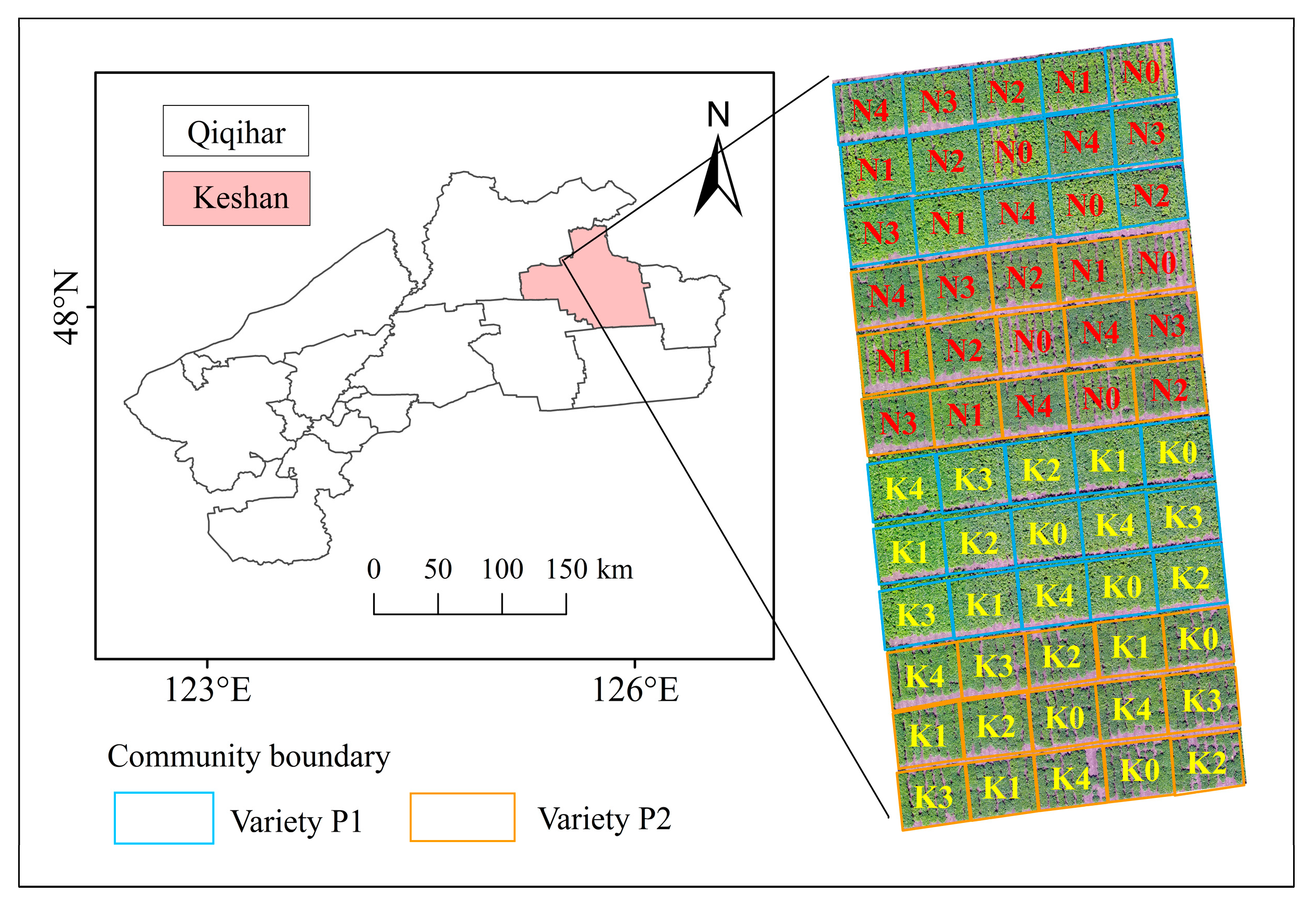
2.1. Experimental Design
2.2. Data Acquisition
2.2.1. Spectral Measurement of Potato Leaves
2.2.2. Determination of Leaf Chlorophyll Content
2.3. Data Analysis Flow and Methods
2.3.1. PROSPECT-4 Model and LUT Generation
2.3.2. Active Learning
2.3.3. Gaussian Process Regression
2.4. LCC Modelling and Accuracy Assessment
2.4.1. LCC Inversion Based on LUT and Cost Function
2.4.2. Hybrid Modeling Approach
2.4.3. Model Accuracy Assessment
3. Results and Analysis
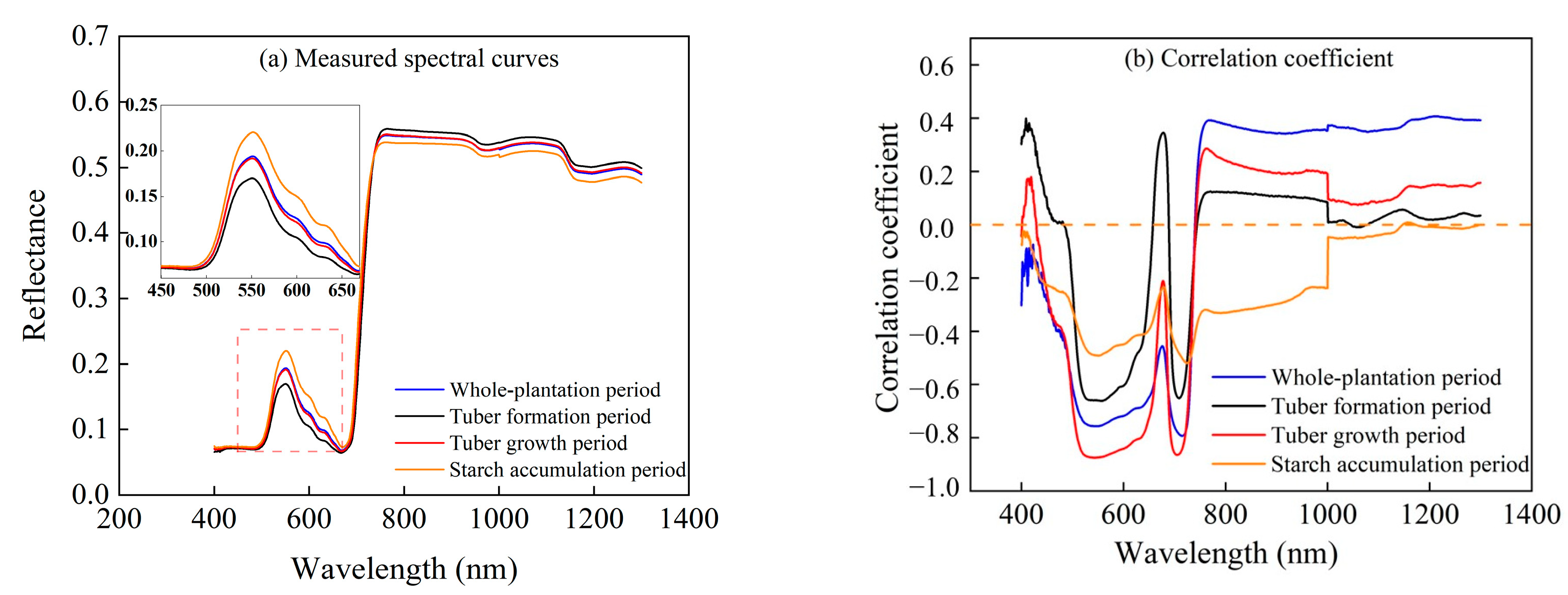
3.1. Relationship between Measured LCC and Spectrum
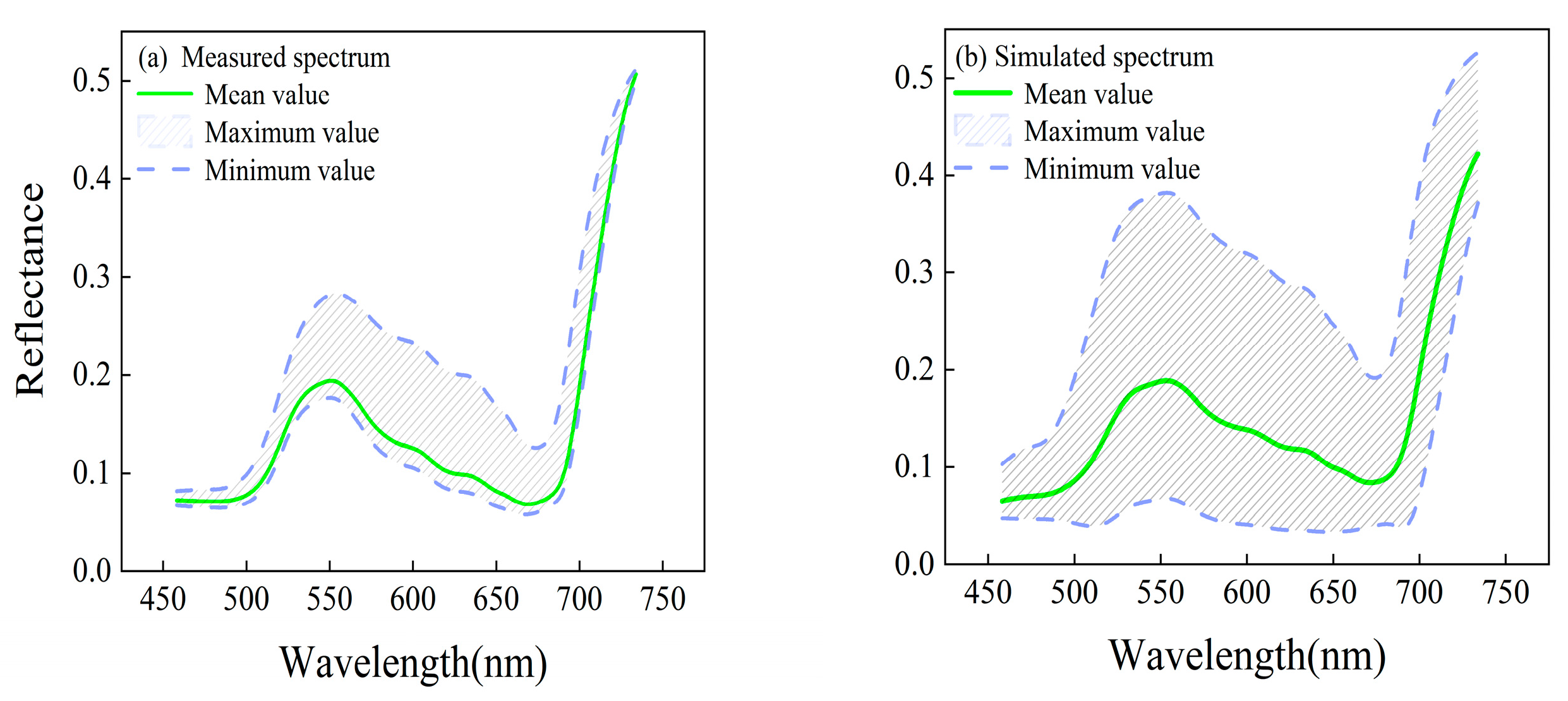
3.2. LUT Generation with PROSPECT-4 Model
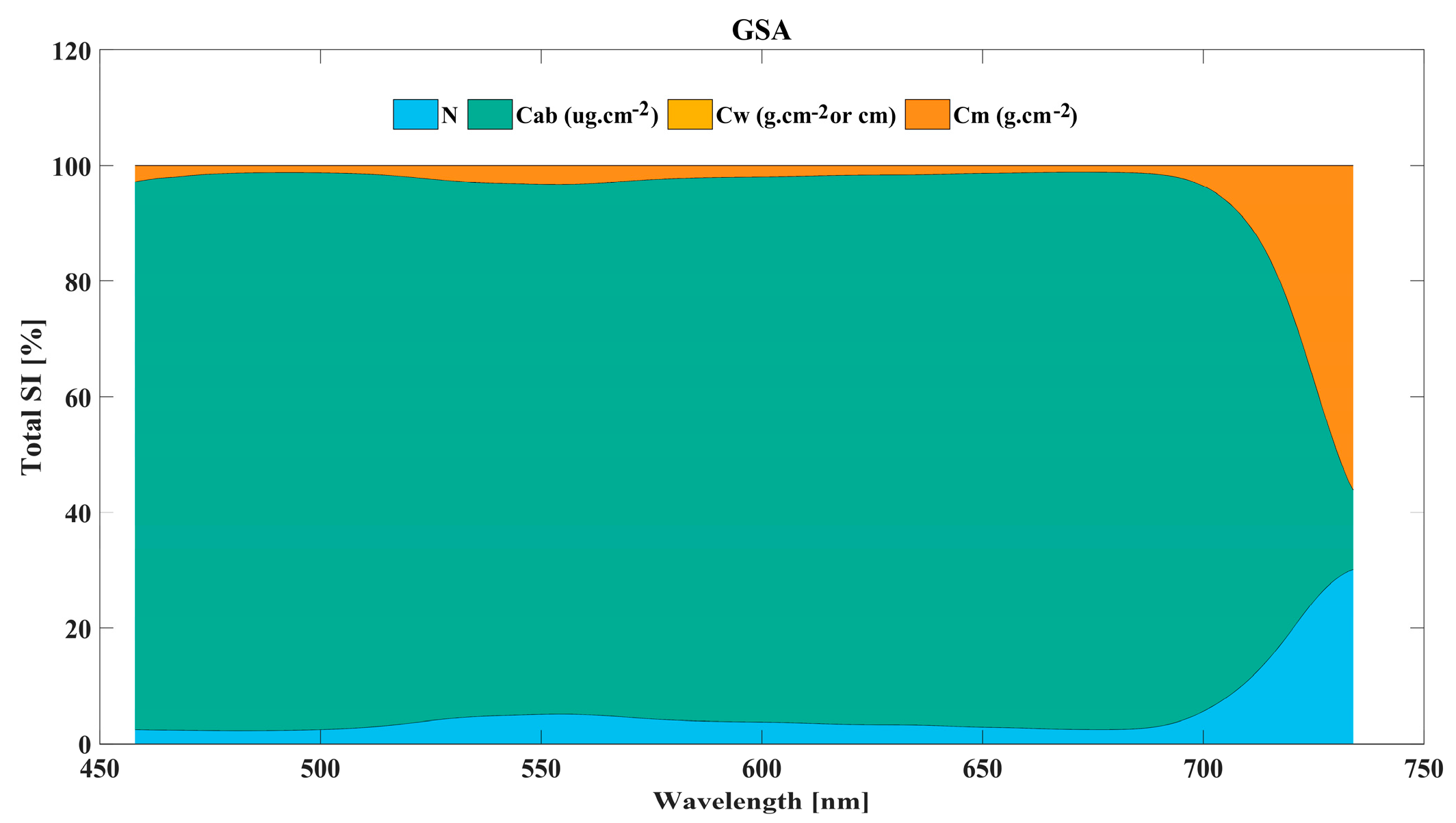
3.2.1. Global Sensitivity Analysis of PROSPECT-4 Model Input Parameters
3.2.2. PROSPECT-4 Input Parameters and LUT Generation
3.3. Potato LCC Inversion
3.3.1. GPR_PROSPECT Combined with AL Modeling Inversion
3.3.2. Chlorophyll Inversion Based on LUT CF Method
3.4. Validation of Inversion Model for Potato Chlorophyll Content
4. Discussion
4.1. AL for Hybrid Retrieval Methods
4.2. Analysis of Potato LCC Hybrid Inversion Model Construction
5. Conclusions
- (1)
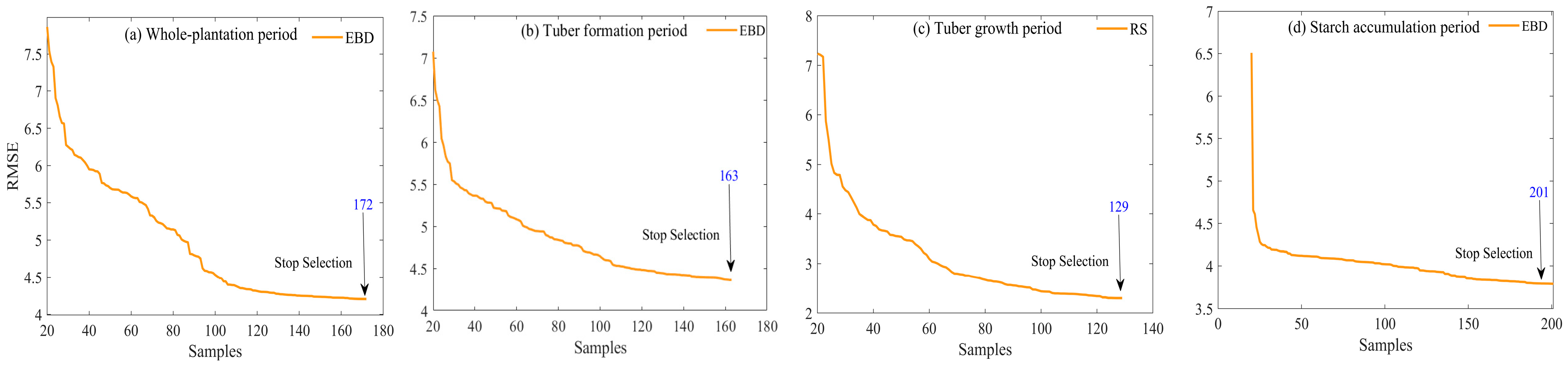
- This study demonstrated that the AL algorithm was able to screen the modeling samples efficiently. Based on the measured labeled dataset of potatoes at different growth periods, this study constructed effective modeling samples of potato LCC at different growth periods from the simulated data pool. The training samples were 172, 163, 129, and 201 for the whole plantation, tuber formation, tuber growth, and starch accumulation periods, respectively. The EBD algorithm in the AL algorithm was more efficient in the sample screening process. Based on whole- and single-fertility validation data, six different AL methods were used in this study to screen the training set from the simulated dataset. Each AL method converged faster to the lower error bound than a random sampling strategy. Diversity criteria (EBD, ABD, and CBD) generally performed the best both in terms of reaching high accuracies as well processing time.
- (2)
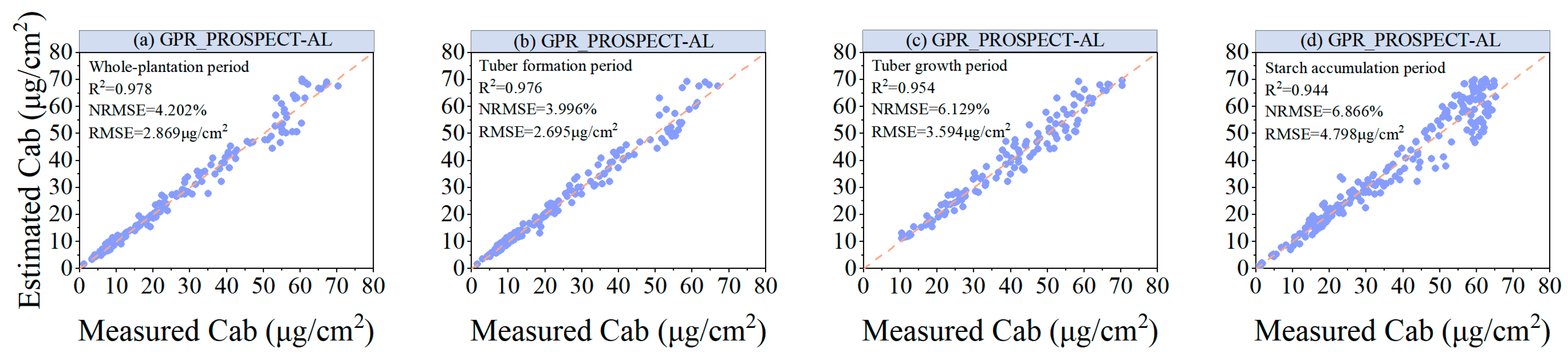
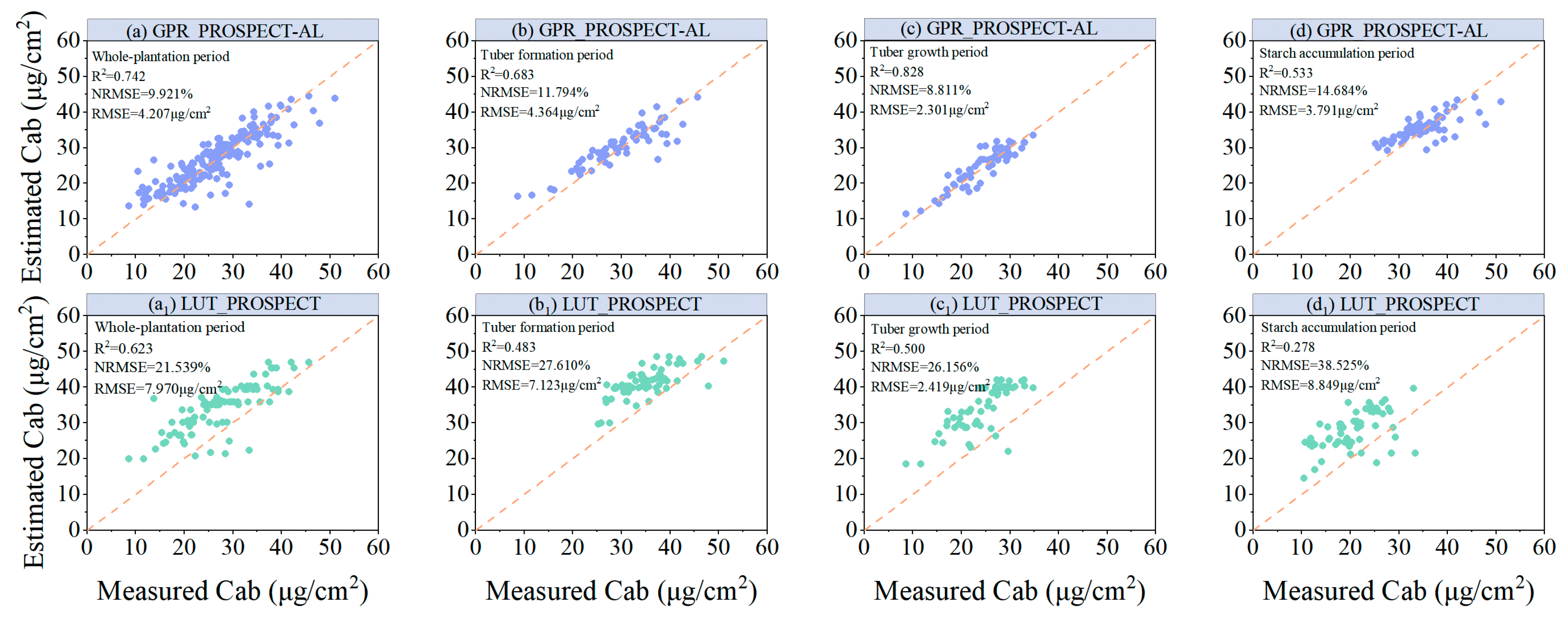
- Compared with the LUT CF method, the hybrid model constructed using GPR_PROSPECT-AL has higher modeling accuracy. This indicates that the RTM-based simulation can generate a sufficiently large training dataset and can be used for inverse LCC model training, while the AL approach helps to optimize the training samples for the RTM simulation and improves the accuracy of the model.
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Guan, K.; Wu, J.; Kimball, J.S.; Anderson, M.C.; Frolking, S.; Li, B.; Hain, C.R.; Lobell, D.B. The shared and unique values of optical, fluorescence, thermal and microwave satellite data for estimating large-scale crop yields. Remote Sens. Environ. 2017, 199, 333–349. [Google Scholar] [CrossRef]
- Houborg, R.; Soegaard, H.; Boegh, E. Combining vegetation index and model inversion methods for the extraction of key vegetation biophysical parameters using Terra and Aqua MODIS reflectance data. Remote Sens. Environ. 2007, 106, 39–58. [Google Scholar] [CrossRef]
- Guo, Y.; Yin, G.; Sun, H.; Wang, H.; Chen, S.; Senthilnath, J.; Wang, J.; Fu, Y. Scaling effects on chlorophyll content estimations with RGB camera mounted on a UAV platform using machine-learning methods. Sensors 2020, 20, 5130. [Google Scholar] [CrossRef]
- Zhang, H.; Ge, Y.; Xie, X.; Atefi, A.; Wijewardane, N.K.; Thapa, S. High throughput analysis of leaf chlorophyll content in sorghum using RGB, hyperspectral, and fluorescence imaging and sensor fusion. Plant Methods 2022, 18, 60. [Google Scholar] [CrossRef]
- Mondal, P.; Basu, M. Adoption of precision agriculture technologies in India and in some developing countries: Scope, present status and strategies. Prog. Nat. Sci. 2009, 19, 659–666. [Google Scholar] [CrossRef]
- Tang, J.-F.; Lin, C.-Y.; Yang, F.-C.; Chang, C.-L. Influence of nitrogen content and bias voltage on residual stress and the tribological and mechanical properties of CrAlN films. Coatings 2020, 10, 546. [Google Scholar] [CrossRef]
- Zhang, J.; Song, X.; Jing, X.; Yang, G.; Yang, C.; Feng, H.; Wang, J.; Ming, S. Remote Sensing Monitoring of Rice Grain Protein Content Based on a Multidimensional Euclidean Distance Method. Remote Sens. 2022, 14, 3989. [Google Scholar] [CrossRef]
- Erb, K.-H.; Kastner, T.; Plutzar, C.; Bais, A.L.S.; Carvalhais, N.; Fetzel, T.; Gingrich, S.; Haberl, H.; Lauk, C.; Niedertscheider, M. Unexpectedly large impact of forest management and grazing on global vegetation biomass. Nature 2018, 553, 73–76. [Google Scholar] [CrossRef]
- Cao, C.; Wang, T.; Gao, M.; Li, Y.; Li, D.; Zhang, H. Hyperspectral inversion of nitrogen content in maize leaves based on different dimensionality reduction algorithms. Comput. Electron. Agric. 2021, 190, 106461. [Google Scholar] [CrossRef]
- Verrelst, J.; Halabuk, A.; Atzberger, C.; Hank, T.; Steinhauser, S.; Berger, K. A comprehensive survey on quantifying non-photosynthetic vegetation cover and biomass from imaging spectroscopy. Ecol. Indic. 2023, 155, 110911. [Google Scholar] [CrossRef]
- Verrelst, J.; Rivera, J.P.; Moreno, J.; Camps-Valls, G. Gaussian processes uncertainty estimates in experimental Sentinel-2 LAI and leaf chlorophyll content retrieval. ISPRS J. Photogramm. Remote Sens. 2013, 86, 157–167. [Google Scholar] [CrossRef]
- Delegido, J.; Alonso, L.; Gonzalez, G.; Moreno, J. Estimating chlorophyll content of crops from hyperspectral data using a normalized area over reflectance curve (NAOC). Int. J. Appl. Earth Obs. Geoinf. 2010, 12, 165–174. [Google Scholar] [CrossRef]
- Wang, X.; Xiao, Z. Hyperspectral Inversion of Chlorophyll Content in Maize Leaves Based on Image and Spectrum Fusion. In Proceedings of the 2022 China Automation Congress (CAC), Xiamen, China, 25–27 November 2022; pp. 6445–6449. [Google Scholar]
- Ban, S.; Liu, W.; Tian, M.; Wang, Q.; Yuan, T.; Chang, Q.; Li, L. Rice leaf chlorophyll content estimation using UAV-based spectral images in different regions. Agronomy 2022, 12, 2832. [Google Scholar] [CrossRef]
- Zhu, X.; Yang, Q.; Chen, X.; Ding, Z. An Approach for Joint Estimation of Grassland Leaf Area Index and Leaf Chlorophyll Content from UAV Hyperspectral Data. Remote Sens. 2023, 15, 2525. [Google Scholar] [CrossRef]
- Kayad, A.; Rodrigues, F.A., Jr.; Naranjo, S.; Sozzi, M.; Pirotti, F.; Marinello, F.; Schulthess, U.; Defourny, P.; Gerard, B.; Weiss, M. Radiative transfer model inversion using high-resolution hyperspectral airborne imagery–Retrieving maize LAI to access biomass and grain yield. Field Crops Res. 2022, 282, 108449. [Google Scholar] [CrossRef] [PubMed]
- Verrelst, J.; Rivera, J.P.; Veroustraete, F.; Muñoz-Marí, J.; Clevers, J.G.; Camps-Valls, G.; Moreno, J. Experimental Sentinel-2 LAI estimation using parametric, non-parametric and physical retrieval methods—A comparison. ISPRS J. Photogramm. Remote Sens. 2015, 108, 260–272. [Google Scholar] [CrossRef]
- Rivera, J.P.; Verrelst, J.; Leonenko, G.; Moreno, J. Multiple cost functions and regularization options for improved retrieval of leaf chlorophyll content and LAI through inversion of the PROSAIL model. Remote Sens. 2013, 5, 3280–3304. [Google Scholar] [CrossRef]
- Binh, N.A.; Hauser, L.T.; Viet Hoa, P.; Thi Phuong Thao, G.; An, N.N.; Nhut, H.S.; Phuong, T.A.; Verrelst, J. Quantifying mangrove leaf area index from Sentinel-2 imagery using hybrid models and active learning. Int. J. Remote Sens. 2022, 43, 5636–5657. [Google Scholar] [CrossRef]
- Berger, K.; Verrelst, J.; Féret, J.-B.; Hank, T.; Wocher, M.; Mauser, W.; Camps-Valls, G. Retrieval of aboveground crop nitrogen content with a hybrid machine learning method. Int. J. Appl. Earth Obs. Geoinf. 2020, 92, 102174. [Google Scholar] [CrossRef]
- Xu, X.; Lu, J.; Zhang, N.; Yang, T.; He, J.; Yao, X.; Cheng, T.; Zhu, Y.; Cao, W.; Tian, Y. Inversion of rice canopy chlorophyll content and leaf area index based on coupling of radiative transfer and Bayesian network models. ISPRS J. Photogramm. Remote Sens. 2019, 150, 185–196. [Google Scholar] [CrossRef]
- Pascual-Venteo, A.B.; Portalés, E.; Berger, K.; Tagliabue, G.; Garcia, J.L.; Pérez-Suay, A.; Rivera-Caicedo, J.P.; Verrelst, J. Prototyping crop traits retrieval models for CHIME: Dimensionality reduction strategies applied to PRISMA data. Remote Sens. 2022, 14, 2448. [Google Scholar] [CrossRef]
- Zhu, J.; Lu, J.; Li, W.; Wang, Y.; Jiang, J.; Cheng, T.; Zhu, Y.; Cao, W.; Yao, X. Estimation of canopy water content for wheat through combining radiative transfer model and machine learning. Field Crops Res. 2023, 302, 109077. [Google Scholar] [CrossRef]
- Antonucci, G.; Impollonia, G.; Croci, M.; Potenza, E.; Marcone, A.; Amaducci, S. Evaluating biostimulants via high-throughput field phenotyping: Biophysical traits retrieval through PROSAIL inversion. Smart Agric. Technol. 2023, 3, 100067. [Google Scholar] [CrossRef]
- ElRafey, A.; Wojtusiak, J. A hybrid active learning and progressive sampling algorithm. Int. J. Mach. Learn. Comput. 2018, 8, 423–427. [Google Scholar]
- Meek, C.; Thiesson, B.; Heckerman, D. The learning-curve sampling method applied to model-based clustering. J. Mach. Learn. Res. 2002, 2, 397–418. [Google Scholar]
- Amitrano, D.; Giacco, G.; Marrone, S.; Pascarella, A.E.; Rigiroli, M.; Sansone, C. Forest Aboveground Biomass Estimation Using Machine Learning Ensembles: Active Learning Strategies for Model Transfer and Field Sampling Reduction. Remote Sens. 2023, 15, 5138. [Google Scholar] [CrossRef]
- Liu, P.; Wang, L.; Ranjan, R.; He, G.; Zhao, L. A survey on active deep learning: From model driven to data driven. ACM Comput. Surv. (CSUR) 2022, 54, 1–34. [Google Scholar] [CrossRef]
- Gulzar, Y.; Ünal, Z.; Aktaş, H.; Mir, M.S. Harnessing the power of transfer learning in sunflower disease detection: A comparative study. Agriculture 2023, 13, 1479. [Google Scholar] [CrossRef]
- Gulzar, Y. Fruit image classification model based on MobileNetV2 with deep transfer learning technique. Sustainability 2023, 15, 1906. [Google Scholar] [CrossRef]
- Dhiman, P.; Kaur, A.; Balasaraswathi, V.; Gulzar, Y.; Alwan, A.A.; Hamid, Y. Image Acquisition, Preprocessing and Classification of Citrus Fruit Diseases: A Systematic Literature Review. Sustainability 2023, 15, 9643. [Google Scholar] [CrossRef]
- Verrelst, J.; Dethier, S.; Rivera, J.P.; Munoz-Mari, J.; Camps-Valls, G.; Moreno, J. Active learning methods for efficient hybrid biophysical variable retrieval. IEEE Geosci. Remote Sens. Lett. 2016, 13, 1012–1016. [Google Scholar] [CrossRef]
- Wan, L.; Liu, Y.; He, Y.; Cen, H. Prior knowledge and active learning enable hybrid method for estimating leaf chlorophyll content from multi-scale canopy reflectance. Comput. Electron. Agric. 2023, 214, 108308. [Google Scholar] [CrossRef]
- Guo, A.; Ye, H.; Li, G.; Zhang, B.; Huang, W.; Jiao, Q.; Qian, B.; Luo, P. Evaluation of Hybrid Models for Maize Chlorophyll Retrieval Using Medium-and High-Spatial-Resolution Satellite Images. Remote Sens. 2023, 15, 1784. [Google Scholar] [CrossRef]
- Wocher, M.; Berger, K.; Verrelst, J.; Hank, T. Retrieval of carbon content and biomass from hyperspectral imagery over cultivated areas. ISPRS J. Photogramm. 2022, 193, 104–114. [Google Scholar] [CrossRef] [PubMed]
- Isharnani, C.E.; Nurcahyani, E.; Lande, M.L. Kandungan Klorofil Daun Planlet Anggrek Tanah (Spathoglottis plicata Blume.) Hasil Pengimbasan Ketahanan terhadap Asam Fusarat secara In Vitro. In Proceedings of the Prosiding Seminar Nasional Swasembada Pangan Polinela Bandar Lampung, Kota Bandar Lampung, Indonesia, 29 April 2015; pp. 86–92. [Google Scholar]
- Yáñez-Rausell, L.; Malenovsky, Z.; Clevers, J.; Schaepman, M.E. Performance of the PROSPECT leaf radiative transfer model version 4 for Norway spruce needles. In Proceedings of the Hyperspectral Workshop 2010, Frascati, Italy, 17–19 March 2010. [Google Scholar]
- Jacquemoud, S.; Baret, F. PROSPECT: A model of leaf optical properties spectra. Remote Sens. Environ. 1990, 34, 75–91. [Google Scholar] [CrossRef]
- Féret, J.-B.; Berger, K.; De Boissieu, F.; Malenovský, Z. PROSPECT-PRO for estimating content of nitrogen-containing leaf proteins and other carbon-based constituents. Remote Sens. Environ. 2021, 252, 112173. [Google Scholar] [CrossRef]
- Spafford, L.; Le Maire, G.; MacDougall, A.; De Boissieu, F.; Féret, J.-B. Spectral subdomains and prior estimation of leaf structure improves PROSPECT inversion on reflectance or transmittance alone. Remote Sens. Environ. 2021, 252, 112176. [Google Scholar] [CrossRef]
- Saltelli, A.; Tarantola, S.; Chan, K.-S. A quantitative model-independent method for global sensitivity analysis of model output. Technometrics 1999, 41, 39–56. [Google Scholar] [CrossRef]
- Berger, K.; Rivera Caicedo, J.P.; Martino, L.; Wocher, M.; Hank, T.; Verrelst, J. A survey of active learning for quantifying vegetation traits from terrestrial earth observation data. Remote Sens. 2021, 13, 287. [Google Scholar] [CrossRef] [PubMed]
- Douak, F.; Melgani, F.; Benoudjit, N. Kernel ridge regression with active learning for wind speed prediction. Appl. Energy 2013, 103, 328–340. [Google Scholar] [CrossRef]
- Tuia, D.; Volpi, M.; Copa, L.; Kanevski, M.; Munoz-Mari, J. A survey of active learning algorithms for supervised remote sensing image classification. IEEE J. Sel. Top. Signal Process. 2011, 5, 606–617. [Google Scholar] [CrossRef]
- Douak, F.; Benoudjit, N.; Melgani, F. A two-stage regression approach for spectroscopic quantitative analysis. Chemom. Intell. Lab. Syst. 2011, 109, 34–41. [Google Scholar] [CrossRef]
- Demir, B.; Persello, C.; Bruzzone, L. Batch-mode active-learning methods for the interactive classification of remote sensing images. IEEE Trans. Geosci. Remote Sens. 2010, 49, 1014–1031. [Google Scholar] [CrossRef]
- Patra, S.; Bruzzone, L. A cluster-assumption based batch mode active learning technique. Pattern Recognit. Lett. 2012, 33, 1042–1048. [Google Scholar] [CrossRef]
- Verrelst, J.; Berger, K.; Rivera-Caicedo, J.P. Intelligent sampling for vegetation nitrogen mapping based on hybrid machine learning algorithms. IEEE Geosci. Remote Sens. Lett. 2020, 18, 2038–2042. [Google Scholar] [CrossRef]
- Bacour, C.; Baret, F.; Béal, D.; Weiss, M.; Pavageau, K. Neural network estimation of LAI, fAPAR, fCover and LAI× Cab, from top of canopy MERIS reflectance data: Principles and validation. Remote Sens. Environ. 2006, 105, 313–325. [Google Scholar] [CrossRef]
- MacKay, D.J. Information-based objective functions for active data selection. Neural Comput. 1992, 4, 590–604. [Google Scholar] [CrossRef]
- Ki, W.C.; Rasmussen, C.E. Gaussian processes for machine learning. Int. J. Neural Syst. 2006, 14, 69–106. [Google Scholar]
- Solla, M.; Pérez-Gracia, V.; Fontul, S. A review of GPR application on transport infrastructures: Troubleshooting and best practices. Remote Sens. 2021, 13, 672. [Google Scholar] [CrossRef]
- Cerquera, M.R.P.; Montaño, J.D.C.; Mondragón, I.; Canbolat, H. UAV for landmine detection using SDR-based GPR technology. In Robots Operating in Hazardous Environments; IntechOpen: Rijeka, Croatia, 2017; pp. 26–55. [Google Scholar]
- Sinha, S.K.; Padalia, H.; Dasgupta, A.; Verrelst, J.; Rivera, J.P. Estimation of leaf area index using PROSAIL based LUT inversion, MLRA-GPR and empirical models: Case study of tropical deciduous forest plantation, North India. Int. J. Appl. Earth Obs. Geoinf. 2020, 86, 102027. [Google Scholar] [CrossRef]
- Locherer, M.; Hank, T.; Danner, M.; Mauser, W. Retrieval of seasonal leaf area index from simulated EnMAP data through optimized LUT-based inversion of the PROSAIL model. Remote Sens. 2015, 7, 10321–10346. [Google Scholar] [CrossRef]
- Darvishzadeh, R.; Skidmore, A.; Schlerf, M.; Atzberger, C. Inversion of a radiative transfer model for estimating vegetation LAI and chlorophyll in a heterogeneous grassland. Remote Sens. Environ. 2008, 112, 2592–2604. [Google Scholar] [CrossRef]
- Impollonia, G.; Croci, M.; Blandinières, H.; Marcone, A.; Amaducci, S. Comparison of PROSAIL Model Inversion Methods for Estimating Leaf Chlorophyll Content and LAI Using UAV Imagery for Hemp Phenotyping. Remote Sens. 2022, 14, 5801. [Google Scholar] [CrossRef]
- Willmott, C.J.; Matsuura, K. Advantages of the mean absolute error (MAE) over the root mean square error (RMSE) in assessing average model performance. Clim. Res. 2005, 30, 79–82. [Google Scholar] [CrossRef]
- Verrelst, J.; Rivera, J.P.; Leonenko, G.; Alonso, L.; Moreno, J. Optimizing LUT-based RTM inversion for semiautomatic mapping of crop biophysical parameters from Sentinel-2 and-3 data: Role of cost functions. IEEE Trans. Geosci. Remote Sens. 2013, 52, 257–269. [Google Scholar] [CrossRef]
- Sun, J.; Shi, S.; Wang, L.; Li, H.; Wang, S.; Gong, W.; Tagesson, T. Optimizing LUT-based inversion of leaf chlorophyll from hyperspectral lidar data: Role of cost functions and regulation strategies. Int. J. Appl. Earth Obs. Geoinf. 2021, 105, 102602. [Google Scholar] [CrossRef]
- Chakhvashvili, E.; Siegmann, B.; Muller, O.; Verrelst, J.; Bendig, J.; Kraska, T.; Rascher, U. Retrieval of crop variables from proximal multispectral UAV image data using PROSAIL in maize canopy. Remote Sens. 2022, 14, 1247. [Google Scholar] [CrossRef]
- Berger, K.; Hank, T.; Halabuk, A.; Rivera-Caicedo, J.P.; Wocher, M.; Mojses, M.; Gerhátová, K.; Tagliabue, G.; Dolz, M.M.; Venteo, A.B.P. Assessing non-photosynthetic cropland biomass from spaceborne hyperspectral imagery. Remote Sens. 2021, 13, 4711. [Google Scholar] [CrossRef]
- Williams, C.K.; Rasmussen, C.E. Gaussian Processes for Machine Learning; MIT Press: Cambridge, MA, USA, 2006; Volume 2. [Google Scholar]
- Verrelst, J.; Alonso, L.; Camps-Valls, G.; Delegido, J.; Moreno, J. Retrieval of vegetation biophysical parameters using Gaussian process techniques. IEEE Trans. Geosci. Remote Sens. 2011, 50, 1832–1843. [Google Scholar] [CrossRef]
- Verrelst, J.; Muñoz, J.; Alonso, L.; Delegido, J.; Rivera, J.P.; Camps-Valls, G.; Moreno, J. Machine learning regression algorithms for biophysical parameter retrieval: Opportunities for Sentinel-2 and-3. Remote Sens. Environ. 2012, 118, 127–139. [Google Scholar] [CrossRef]

| Parameter | Unit | Min | Max | Samples |
|---|---|---|---|---|
| Chlorophyll content (Cab) | µg/cm2 | 0 | 70 | 2 |
| Equivalent water thickness (Cw) | g/cm2 | 0.0001 | 0.08 | 2 |
| Leaf structure (N) | — | 1.5 | 2.5 | 2 |
| Dry matter content (Cm) | g/cm2 | 0.0001 | 0.05 | 2 |
| AL Selected Criterions | AL Algorithms | Equation | Literatures |
|---|---|---|---|
| Diversity Criteria Methods | Euclidean distance-based diversity (EBD) | [43] | |
| Angle-based diversity (ABD) | [46] | ||
| Cluster-based diversity (CBD) | clustering algorithm | [47] | |
| Uncertainty Criteria Methods | Pool of regressors (PAL) | [43] | |
| Residual regression AL (RSAL) | [45] | ||
| Entropy query by bagging (EQB) | [44] |
| Measured Datasets | Tuber Formation Period | Tuber Growth Period | Starch Accumulation Period | Whole Plantation Period | |
|---|---|---|---|---|---|
| Data collection Date | 21 July 2022 | 8 August 2022 | 28 August 2022 | — | |
| LCC (µg/cm2) | Sample size | 60 | 60 | 60 | 180 |
| Min | 25.565 | 8.797 | 10.526 | 8.797 | |
| Max | 51.123 | 34.828 | 33.846 | 51.123 | |
| Mean | 38.344 | 21.813 | 22.186 | 29.960 | |
| CV (%) | 0.471 | 0.844 | 0.743 | 0.998 | |
| LCC for N treatment (µg/cm2) | Sample size | 30 | 30 | 30 | 90 |
| Min | 25.565 | 8.797 | 13.945 | 8.797 | |
| Max | 46.192 | 33.473 | 33.846 | 46.192 | |
| Mean | 35.879 | 21.135 | 23.896 | 27.4945 | |
| CV (%) | 0.407 | 0.826 | 0.589 | 0.962 | |
| LCC for K treatment (µg/cm2) | Sample size | 30 | 30 | 30 | 90 |
| Min | 25.854 | 14.689 | 10.526 | 10.526 | |
| Max | 51.123 | 34.828 | 28.357 | 51.123 | |
| Mean | 38.489 | 24.759 | 19.442 | 30.825 | |
| CV (%) | 0.464 | 0.575 | 0.649 | 0.931 | |
| AL | Whole Plantation Period | Tuber Formation Period | Tuber Growth Period | Starch Accumulation Period | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GPR_PROSPECT-AL | GPR_PROSPECT-AL | GPR_PROSPECT-AL | GPR_PROSPECT-AL | |||||||||
| R2 | NRMSE | Time | R2 | NRMSE | Time | R2 | NRMSE | Time | R2 | NRMSE | Time | |
| RAL | 0.701 | 0.107 | 0.022 | 0.481 | 0.154 | 0.022 | 0.726 | 0.116 | 0.016 | 0.312 | 0.212 | 0.035 |
| RS | 0.732 | 0.012 | 0.022 | 0.517 | 0.148 | 0.02 | 0.828 * | 0.088 | 0.026 | 0.256 | 0.195 | 0.033 |
| PAL | 0.743 | 0.099 | 0.031 | 0.481 | 0.154 | 0.029 | 0.792 | 0.097 | 0.031 | 0.266 | 0.214 | 0.032 |
| ABD | 0.729 | 0.104 | 0.031 | 0.51 | 0.151 | 0.031 | 0.632 | 0.129 | 0.038 | 0.214 | 0.218 | 0.033 |
| CBD | 0.725 | 0.103 | 0.032 | 0.518 | 0.148 | 0.028 | 0.815 | 0.09 | 0.028 | 0.232 | 0.218 | 0.033 |
| EBD | 0.742 * | 0.099 | 0.026 | 0.683 * | 0.118 | 0.03 | 0.804 | 0.092 | 0.031 | 0.533 * | 0.147 | 0.003 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, Y.; Qiu, C.; Zhang, J.; Pan, D.; Zheng, C.; Sun, H.; Feng, H.; Song, X. Potato Leaf Chlorophyll Content Estimation through Radiative Transfer Modeling and Active Learning. Agronomy 2023, 13, 3071. https://doi.org/10.3390/agronomy13123071
Ma Y, Qiu C, Zhang J, Pan D, Zheng C, Sun H, Feng H, Song X. Potato Leaf Chlorophyll Content Estimation through Radiative Transfer Modeling and Active Learning. Agronomy. 2023; 13(12):3071. https://doi.org/10.3390/agronomy13123071
Chicago/Turabian StyleMa, Yuanyuan, Chunxia Qiu, Jie Zhang, Di Pan, Chunkai Zheng, Heguang Sun, Haikuan Feng, and Xiaoyu Song. 2023. "Potato Leaf Chlorophyll Content Estimation through Radiative Transfer Modeling and Active Learning" Agronomy 13, no. 12: 3071. https://doi.org/10.3390/agronomy13123071
APA StyleMa, Y., Qiu, C., Zhang, J., Pan, D., Zheng, C., Sun, H., Feng, H., & Song, X. (2023). Potato Leaf Chlorophyll Content Estimation through Radiative Transfer Modeling and Active Learning. Agronomy, 13(12), 3071. https://doi.org/10.3390/agronomy13123071







