Straw-Based Biopurification Systems to Remove Ibuprofen, Diclofenac and Triclosan from Wastewaters: Dominant Microbial Communities
Abstract
:1. Introduction
2. Material and Methods
2.1. Soil, Organic Material and Preparation of Biomixtures
2.2. Chemicals
2.3. Adsorption of PPCPs in the Biomixtures
2.4. Dissipation Study in Microcosm Biopurification System
2.5. Analysis of the PPCPs and Their Metabolites
2.6. DNA Extraction and Microbial Identification
2.7. Data Analysis
3. Results and Discussion
3.1. Comparison of Chemical Properties of Uncontaminated Biomixtures
3.2. PPCP Adsorption in the Biomixtures
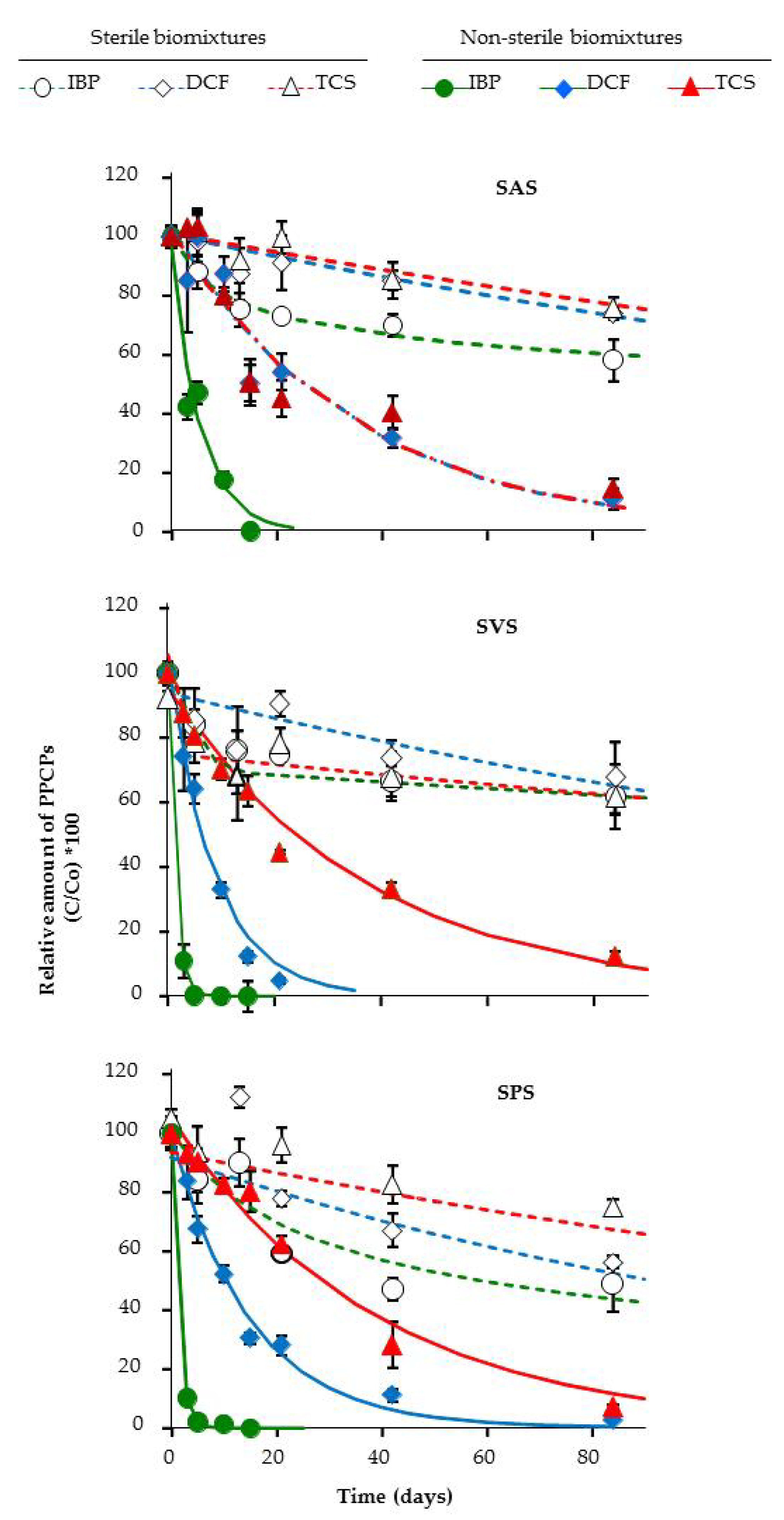
3.3. Dissipation of PPCPs in the Microcosm Biopurification Systems
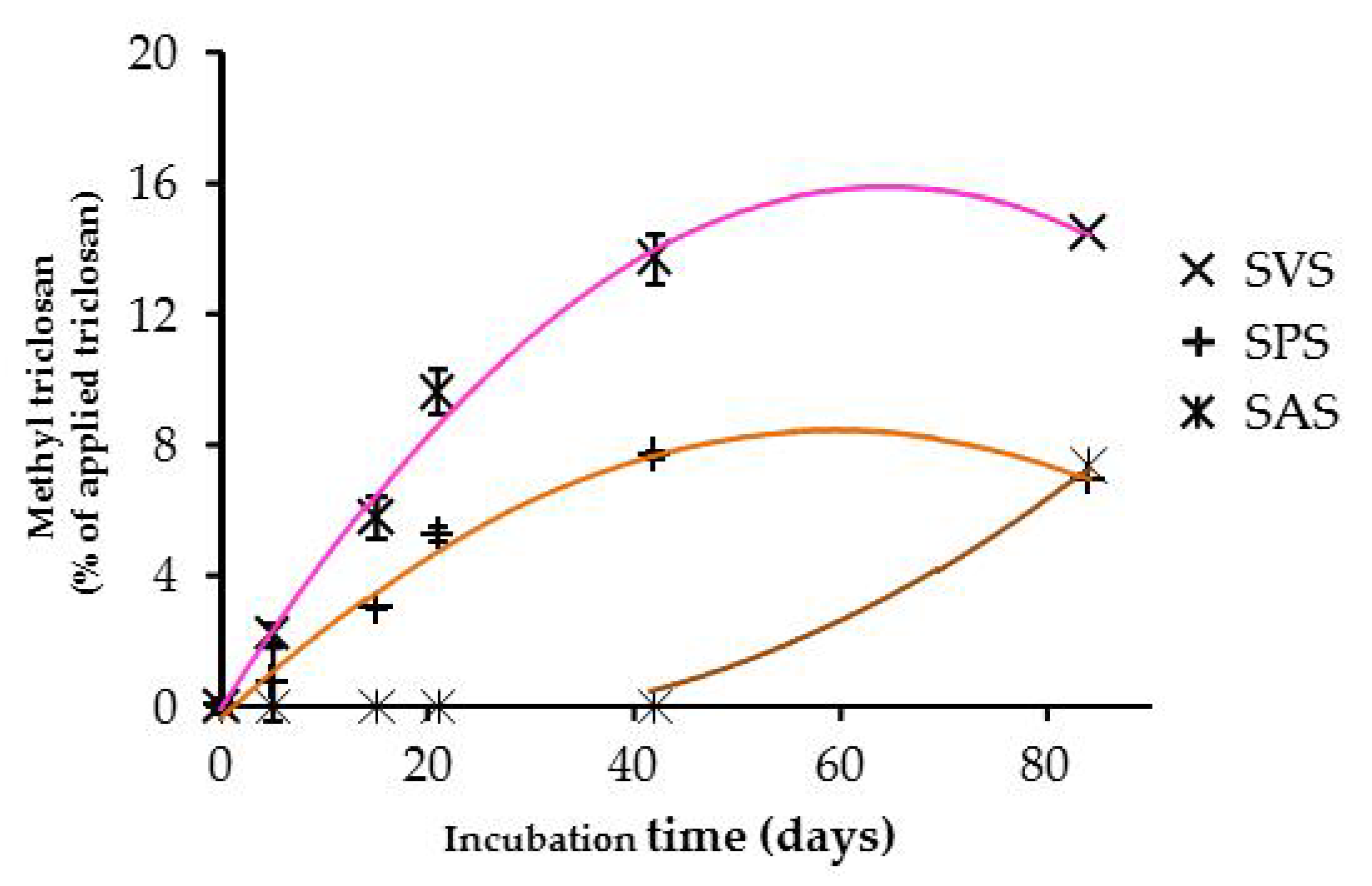
3.4. Degradation Products
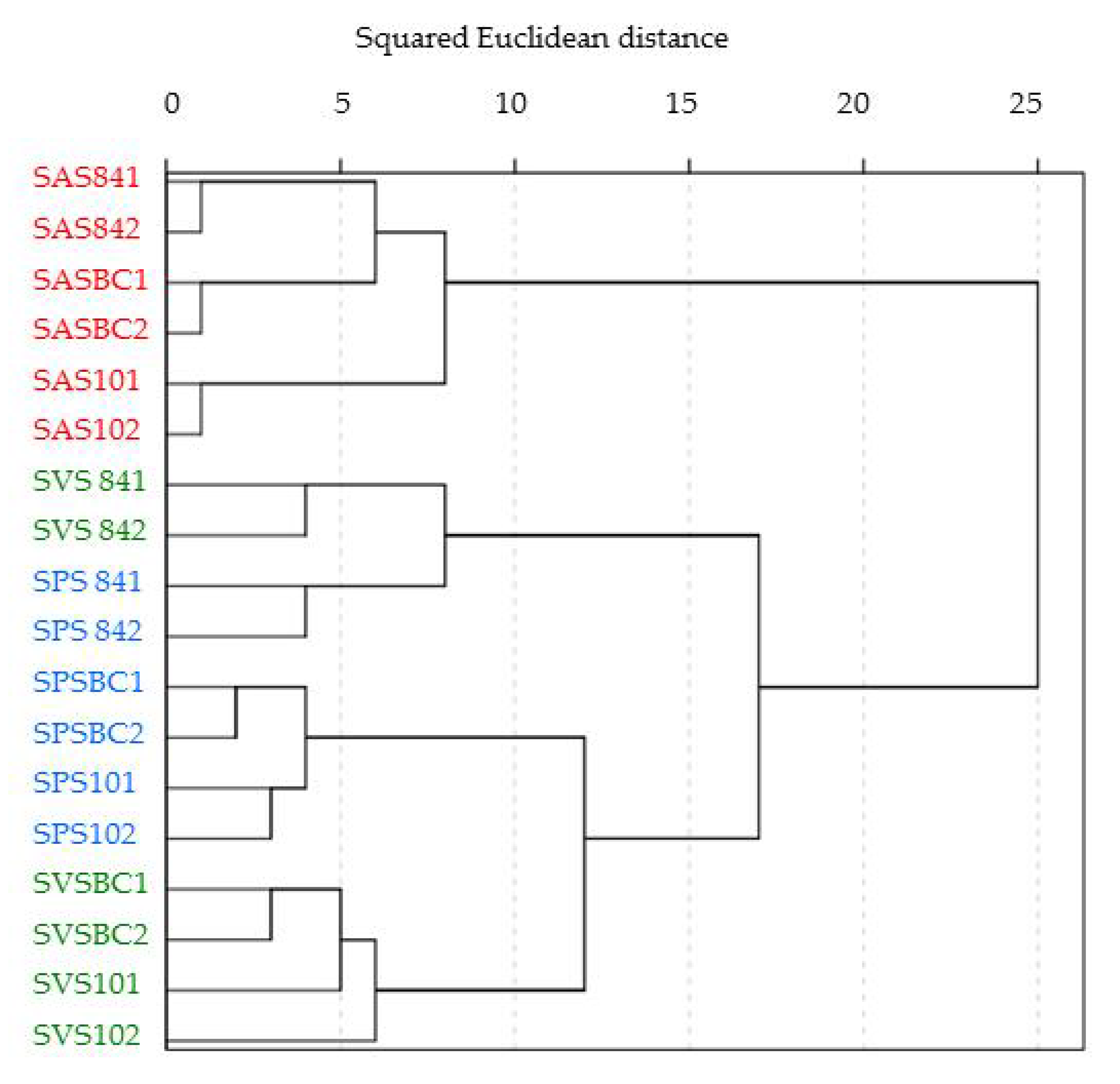
3.5. Bacterial Community Structure and Composition in Microcosm Biopurification Systems
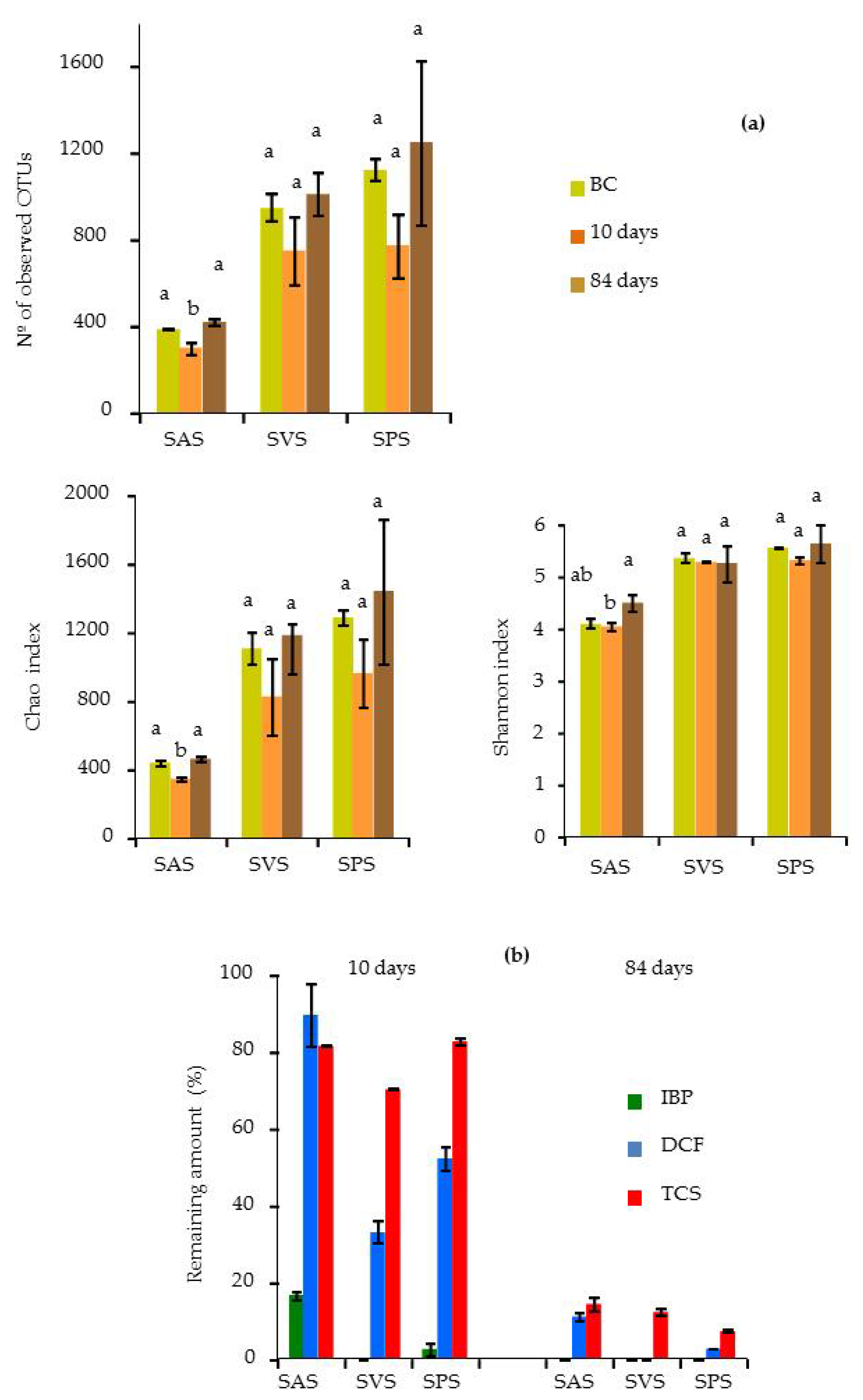
3.5.1. Bacterial Diversity in the Microcosm Biopurification Systems
3.5.2. Distribution of Bacterial Phyla in Biomixtures from Microcosm Biopurification Systems
3.5.3. Impact of PPCP Application on Bacterial Phyla Composition of Biomixtures from Microcosm Biopurification Systems
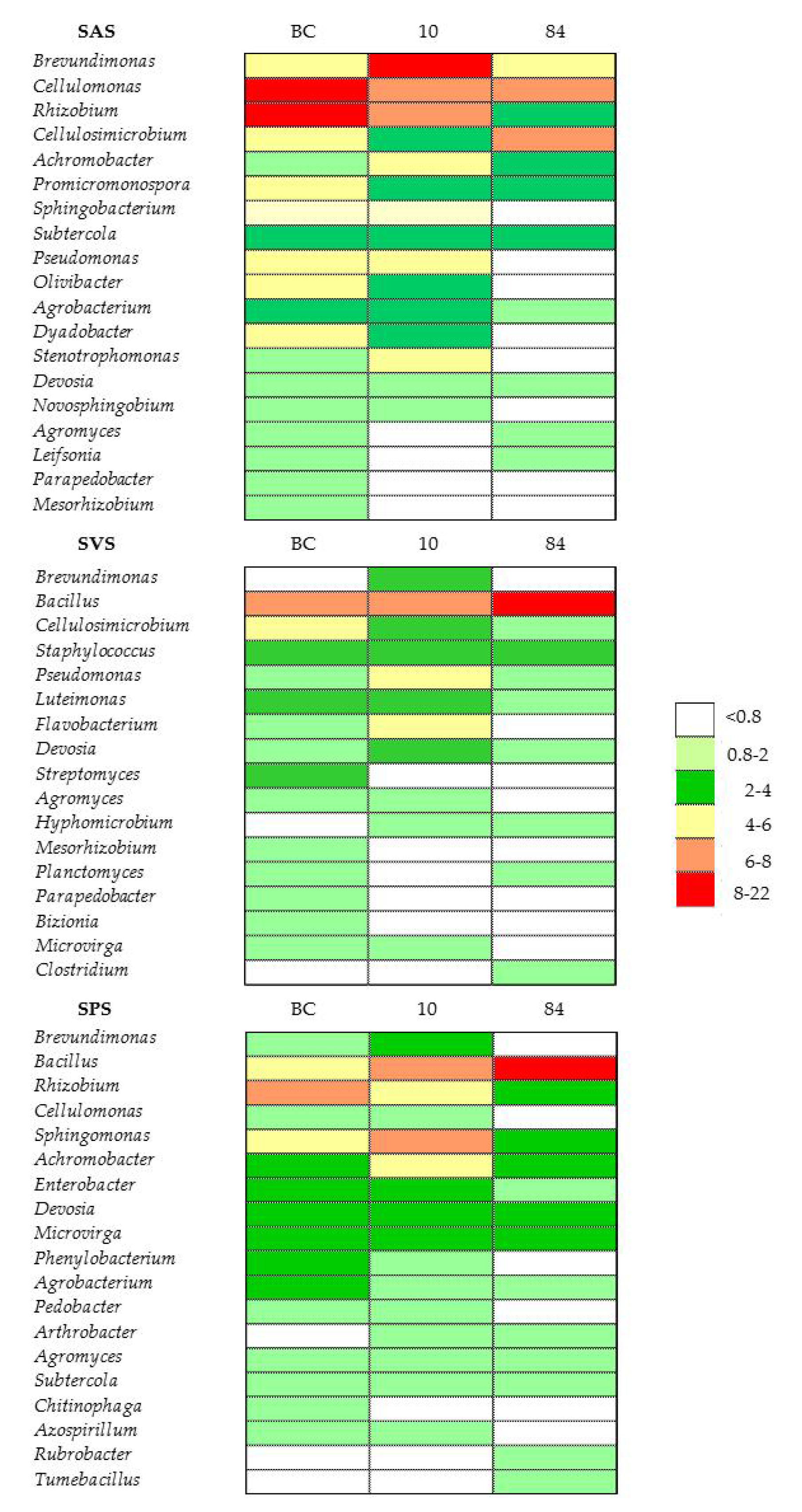
3.5.4. Bacterial Classes and Genera in Microcosm Biopurification Systems
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Aus der Beek, T.; Weber, F.A.; Bergmann, A.; Hickmann, S.; Ebert, I.; Hein, A.; Küster, A. Pharmaceuticals in the environment-Global occurrences and perspectives. Environ. Toxicol. Chem. 2016, 35, 823–835. [Google Scholar] [CrossRef]
- Fekadu, S.; Alemayehu, E.; Dewil, R.; Van der Bruggen, B. Pharmaceuticals in freshwater aquatic environments: A comparison of the African and European challenge. Sci. Total Environ. 2019, 654, 324–337. [Google Scholar] [CrossRef]
- Rivera-Utrilla, J.; Sánchez-Polo, M.; Ferro-García, M.Á.; Prados-Joya, G.; Ocampo-Pérez, R. Pharmaceuticals as emerging contaminants and their removal from water. A review. Chemosphere 2013, 93, 1268–1287. [Google Scholar] [CrossRef]
- Grenni, P.; Ancona, V.; Barra Caracciolo, A. Ecological effects of antibiotics on natural ecosystems: A review. Microchem. J. 2018, 136, 25–39. [Google Scholar] [CrossRef]
- Santos, L.H.M.L.M.; Araújo, A.N.; Fachini, A.; Pena, A.; Delerue-Matos, C.; Montenegro, M.C.B.S.M. Ecotoxicological aspects related to the presence of pharmaceuticals in the aquatic environment. J. Hazard. Mater. 2010, 175, 45–95. [Google Scholar] [CrossRef] [Green Version]
- Oaks, J.L.; Gilbert, M.; Virani, M.Z.; Watson, R.T.; Meteyer, C.U.; Rideout, B.A.; Shivaprasad, H.L.; Ahmed, S.; Chaudhry, M.J.I.; Arshad, M.; et al. Diclofenac residues as the cause of vulture population decline in Pakistan. Nature 2004, 427, 630–633. [Google Scholar] [CrossRef]
- Lonappan, L.; Brar, S.K.; Das, R.K.; Verma, M.; Surampalli, R.Y. Diclofenac and its transformation products: Environmental occurrence and toxicity—A review. Environ. Int. 2016, 96, 127–138. [Google Scholar] [CrossRef] [Green Version]
- Almeida, B.; Oehmen, A.; Marques, R.; Brito, D.; Carvalho, G.; Barreto Crespo, M.T. Modelling the biodegradation of non-steroidal anti-inflammatory drugs (NSAIDs) by activated sludge and a pure culture. Bioresour. Technol. 2013, 133, 31–37. [Google Scholar] [CrossRef] [PubMed]
- Gros, M.; Petrovic, M.; Barceló, D. Wastewater treatment plants as a pathway for aquatic contamination by pharmaceuticals in the Ebro river basin (northeast Spain). Environ. Toxicol. Chem. 2007, 26, 1553–1562. [Google Scholar] [CrossRef] [PubMed]
- Thomaidi, V.S.; Matsoukas, C.; Stasinakis, A.S. Risk assessment of triclosan released from sewage treatment plants in European rivers using a combination of risk quotient methodology and Monte Carlo simulation. Sci. Total Environ. 2017, 603–604, 487–494. [Google Scholar] [CrossRef] [PubMed]
- Quintana, J.B.; Weiss, S.; Reemtsma, T. Pathways and metabolites of microbial degradation of selected acidic pharmaceutical and their occurrence in municipal wastewater treated by a membrane bioreactor. Water Res. 2005, 39, 2654–2664. [Google Scholar] [CrossRef] [PubMed]
- Macherius, A.; Lapen, D.R.; Reemtsma, T.; Römbke, J.; Topp, E.; Coors, A. Triclocarban, triclosan and its transformation product methyl triclosan in native earthworm species four years after a commercial-scale biosolids application. Sci. Total Environ. 2014, 472, 235–238. [Google Scholar] [CrossRef]
- Ding, T.; Lin, K.; Bao, L.; Yang, M.; Li, J.; Yang, B.; Gan, J. Biouptake, toxicity and biotransformation of triclosan in diatom Cymbella sp. and the influence of humic acid. Environ. Pollut. 2018, 234, 231–242. [Google Scholar] [CrossRef] [PubMed]
- Butler, E.; Whelan, M.J.; Sakrabani, R.; Van Egmond, R. Fate of triclosan in field soils receiving sewage sludge. Environ. Pollut. 2012, 167, 101–109. [Google Scholar] [CrossRef]
- Macedo, S.; Torres, T.; Santos, M.M. Methyl-triclosan and triclosan impact embryonic development of Danio rerio and Paracentrotus lividus. Ecotoxicology 2017, 26, 482–489. [Google Scholar] [CrossRef] [PubMed]
- Delgado-Moreno, L.; Bazhari, S.; Nogales, R.; Romero, E. Innovative application of biobed bioremediation systems to remove emerging contaminants: Adsorption, degradation and bioaccesibility. Sci. Total Environ. 2019, 651. [Google Scholar] [CrossRef] [PubMed]
- Aguilar-Romero, I.; Romero, E.; Wittich, R.M.; van Dillewijn, P. Bacterial ecotoxicity and shifts in bacterial communities associated with the removal of ibuprofen, diclofenac and triclosan in biopurification systems. Sci. Total Environ. 2020, 741. [Google Scholar] [CrossRef]
- Castillo, M.D.P.; Torstensson, L.; Stenström, J. Biobeds for environmental protection from pesticide use—A review. J. Agric. Food Chem. 2008, 56, 6206–6219. [Google Scholar] [CrossRef]
- Larcher, S.; Yargeau, V. Biodegradation of 17α-ethinylestradiol by heterotrophic bacteria. Environ. Pollut. 2013, 173, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Aissaoui, S.; Ouled-Haddar, H.; Sifour, M.; Harrouche, K.; Sghaier, H. Metabolic and co-metabolic transformation of diclofenac by enterobacter hormaechei D15 isolated from activated sludge. Curr. Microbiol. 2017, 74, 381–388. [Google Scholar] [CrossRef]
- Holmsgaard, P.N.; Dealtry, S.; Dunon, V.; Heuer, H.; Hansen, L.H.; Springael, D.; Smalla, K.; Riber, L.; Sørensen, S.J. Response of the bacterial community in an on-farm biopurification system, to which diverse pesticides are introduced over an agricultural season. Environ. Pollut. 2017, 229, 854–862. [Google Scholar] [CrossRef] [PubMed]
- Góngora-Echeverría, V.R.; Quintal-Franco, C.; Arena-Ortiz, M.L.; Giácoman-Vallejos, G.; Ponce-Caballero, C. Identification of microbial species present in a pesticide dissipation process in biobed systems using typical substrates from southeastern Mexico as a biomixture at a laboratory scale. Sci. Total Environ. 2018, 628–629, 528–538. [Google Scholar] [CrossRef]
- Castro-Gutiérrez, V.; Masís-Mora, M.; Carazo-Rojas, E.; Mora-López, M.; Rodríguez-Rodríguez, C.E. Impact of oxytetracycline and bacterial bioaugmentation on the efficiency and microbial community structure of a pesticide-degrading biomixture. Environ. Sci. Pollut. Res. 2018, 25, 11787–11799. [Google Scholar] [CrossRef]
- Castillo Diaz, J.M.; Delgado-Moreno, L.; Núñez, R.; Nogales, R.; Romero, E. Enhancing pesticide degradation using indigenous microorganisms isolated under high pesticide load in bioremediation systems with vermicomposts. Bioresour. Technol. 2016, 214. [Google Scholar] [CrossRef] [PubMed]
- Delgado-Moreno, L.; Nogales, R.; Romero, E. Biodegradation of high doses of commercial pesticide products in pilot-scale biobeds using olive-oil agroindustry wastes. J. Environ. Manag. 2017, 204. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef]
- Delgado-Moreno, L.; Bazhari, S.; Gasco, G.; Méndez, A.; El Azzouzi, M.; Romero, E. New insights into the efficient removal of emerging contaminants by biochars and hydrochars derived from olive oil wastes. Sci. Total Environ. 2021, 752, 141838. [Google Scholar] [CrossRef] [PubMed]
- Karnjanapiboonwong, A.; Morse, A.N.; Maul, J.D.; Anderson, T.A. Sorption of estrogens, triclosan, and caffeine in a sandy loam and a silt loam soil. J. Soils Sediments 2010, 10, 1300–1307. [Google Scholar] [CrossRef]
- Behera, S.K.; Oh, S.Y.; Park, H.S. Sorption of triclosan onto activated carbon, kaolinite and montmorillonite: Effects of pH, ionic strength, and humic acid. J. Hazard. Mater. 2010, 179, 684–691. [Google Scholar] [CrossRef]
- Xu, J.; Wu, L.; Chang, A.C. Degradation and adsorption of selected pharmaceuticals and personal care products (PPCPs) in agricultural soils. Chemosphere 2009, 77, 1299–1305. [Google Scholar] [CrossRef]
- Castillo, J.M.; Romero, E.; Nogales, R. Dynamics of microbial communities related to biochemical parameters during vermicomposting and maturation of agroindustrial lignocellulose wastes. Bioresour. Technol. 2013, 146. [Google Scholar] [CrossRef]
- Yu, Y.; Han, P.; Zhou, L.J.; Li, Z.; Wagner, M.; Men, Y. Ammonia monooxygenase-mediated cometabolic biotransformation and hydroxylamine-mediated abiotic transformation of Micropollutants in an AOB/NOB Coculture. Environ. Sci. Technol. 2018, 52, 9196–9205. [Google Scholar] [CrossRef]
- Wu, G.; Geng, J.; Xu, K.; Ren, H. Removal of pharmaceuticals by ammonia oxidizers during nitrification. Appl. Microbiol. Biotechnol. 2021, 105, 909–921. [Google Scholar] [CrossRef] [PubMed]
- Iranzo, M.; Gamón, M.; Boluda, R.; Mormeneo, S. Analysis of pharmaceutical biodegradation of WWTP sludge using composting and identification of certain microorganisms involved in the process. Sci. Total Environ. 2018, 640–641, 840–848. [Google Scholar] [CrossRef]
- Romero, E.; Salido, A.; Cifuentes, C.; Fernández, J.-D.; Nogales, R. Effect of vermicomposting process on pesticide sorption capability using agro-industrial wastes. Int. J. Environ. Anal. Chem. 2006, 86. [Google Scholar] [CrossRef]
- Wu, C.; Spongberg, A.L.; Witter, J.D. Adsorption and degradation of triclosan and triclocarban in soils and biosolids-amended soils. J. Agric. Food Chem. 2009, 57, 4900–4905. [Google Scholar] [CrossRef]
- Fu, Q.; Sanganyado, E.; Ye, Q.; Gan, J. Meta-analysis of biosolid effects on persistence of triclosan and triclocarban in soil. Environ. Pollut. 2016, 210, 137–144. [Google Scholar] [CrossRef] [Green Version]
- Clayborn, A.B.; Toofan, S.N.; Champlin, F.R. Influence of methylation on the antibacterial properties of triclosan in Pasteurella multocida and Pseudomonas aeruginosa variant strains. J. Hosp. Infect. 2011, 77, 129–133. [Google Scholar] [CrossRef] [Green Version]
- Oh, S.; Choi, D.; Cha, C.J. Ecological processes underpinning microbial community structure during exposure to subinhibitory level of triclosan. Sci. Rep. 2019, 9, 1–12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saavedra, M.; Benitez, E.; Cifuentes, C.; Nogales, R. Enzyme activities and chemical changes in wet olive cake after treatment with Pleurotus ostreatus or Eisenia fetida. Biodegradation 2006, 17, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Vivas, A.; Moreno, B.; Garcia-Rodriguez, S.; Benitez, E. Assessing the impact of composting and vermicomposting on bacterial community size and structure, and microbial functional diversity of an olive-mill waste. Bioresour. Technol. 2009, 100, 1319–1326. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Geng, J.; Hu, H.; Ma, H.; Gao, X.; Ren, H. Impact of selected non-steroidal anti-inflammatory pharmaceuticals on microbial community assembly and activity in sequencing batch reactors. PLoS ONE 2017, 12, e0179236. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rutere, C.; Knoop, K.; Posselt, M.; Ho, A.; Horn, M.A. Ibuprofen degradation and associated bacterial communities in hyporheic zone sediments. Microorganisms 2020, 8, 1245. [Google Scholar] [CrossRef]
- Thelusmond, J.R.; Kawka, E.; Strathmann, T.J.; Cupples, A.M. Diclofenac, carbamazepine and triclocarban biodegradation in agricultural soils and the microorganisms and metabolic pathways affected. Sci. Total Environ. 2018, 640–641, 1393–1410. [Google Scholar] [CrossRef]
- Lawrence, J.R.; Swerhone, G.D.W.; Wassenaar, L.I.; Neu, T.R. Effects of selected pharmaceuticals on riverine biofilm communities. Can. J. Microbiol. 2005, 51, 655–669. [Google Scholar] [CrossRef]
- Xia, Z.; Xiao-chun, W.; Zhong-lin, C.; Hao, X.; Qing-fang, Z. Microbial community structure and pharmaceuticals and personal care products removal in a membrane bioreactor seeded with aerobic granular sludge. Appl. Microbiol. Biotechnol. 2015, 99, 425–433. [Google Scholar] [CrossRef]
- Fierer, N.; Lauber, C.L.; Ramirez, K.S.; Zaneveld, J.; Bradford, M.A.; Knight, R. Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J. 2012, 6, 1007–1017. [Google Scholar] [CrossRef] [Green Version]
- Nega, M.; Braun, B.; Künzel, S.; Szewzyk, U. Evaluating the impact ofwastewater effluent on microbial communities in the Panke, an Urban River. Water 2019, 11, 888. [Google Scholar] [CrossRef] [Green Version]
- Zhang, D.; Luo, J.; Lee, Z.M.P.; Maspolim, Y.; Gersberg, R.M.; Liu, Y.; Tan, S.K.; Ng, W.J. Characterization of bacterial communities in wetland mesocosms receiving pharmaceutical-enriched wastewater. Ecol. Eng. 2016, 90, 215–224. [Google Scholar] [CrossRef]
- Krzmarzick, M.J.; Crary, B.B.; Harding, J.J.; Oyerinde, O.O.; Leri, A.C.; Myneni, S.C.B.; Novak, P.J. Natural niche for organohalide-respiring chloroflexi. Appl. Environ. Microbiol. 2012, 78, 393–401. [Google Scholar] [CrossRef] [Green Version]
- Krzmarzick, M.J.; Taylor, D.K.; Fu, X.; McCutchan, A.L. Diversity and niche of archaea in bioremediation. Archaea 2018, 2018. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mcnamara, P.J.; Krzmarzick, M.J. Triclosan enriches for Dehalococcoides-like Chloroflexi in anaerobic soil at environmentally relevant concentrations. FEMS Microbiol. Lett. 2013, 344, 48–52. [Google Scholar] [CrossRef]
- Tortella, G.R.; Mella-Herrera, R.A.; Sousa, D.Z.; Rubilar, O.; Acuña, J.J.; Briceño, G.; Diez, M.C. Atrazine dissipation and its impact on the microbial communities and community level physiological profiles in a microcosm simulating the biomixture of on-farm biopurification system. J. Hazard. Mater. 2013, 260, 459–467. [Google Scholar] [CrossRef] [Green Version]
- Barra Caracciolo, A.; Topp, E.; Grenni, P. Pharmaceuticals in the environment: Biodegradation and effects on natural microbial communities. A review. J. Pharm. Biomed. Anal. 2015, 106, 25–36. [Google Scholar] [CrossRef]
- Butler, E.; Whelan, M.J.; Ritz, K.; Sakrabani, R.; Van Egmond, R. Effects of triclosan on soil microbial respiration. Environ. Toxicol. Chem. 2011, 30, 360–366. [Google Scholar] [CrossRef]
- Zhao, C.; Xie, H.J.; Xu, J.; Xu, X.; Zhang, J.; Hu, Z.; Liu, C.; Liang, S.; Wang, Q.; Wang, J. Bacterial community variation and microbial mechanism of triclosan (TCS) removal by constructed wetlands with different types of plants. Sci. Total Environ. 2015, 505, 633–639. [Google Scholar] [CrossRef]
- Guo, Q.; Yan, J.; Wen, J.; Hu, Y.; Chen, Y.; Wu, W. Rhamnolipid-enhanced aerobic biodegradation of triclosan (TCS) by indigenous microorganisms in water-sediment systems. Sci. Total Environ. 2016, 571, 1304–1311. [Google Scholar] [CrossRef]
- Martin, E.K.; Kemal, S.; Henrik, N.R.; Alexander, E.; Natalia, C.; Henrik, J.C.; Thomas, B.; Hans, B.; Erik, K. Triclosan changes community composition and selects for specific bacterial taxa in marine periphyton biofilms in low nanomolar concentrations. Ecotoxicology 2020, 29, 1083–1094. [Google Scholar] [CrossRef]
- Lear, J.C.; Maillard, J.Y.; Dettmar, P.W.; Goddard, P.A.; Russell, A.D. Chloroxylenol- and triclosan-tolerant bacteria from industrial sources. J. Ind. Microbiol. Biotechnol. 2002, 29, 238–242. [Google Scholar] [CrossRef] [PubMed]
- Murdoch, R.W.; Hay, A.G. Formation of catechols via removal of acid side chains from ibuprofen and related aromatic acids. Appl. Environ. Microbiol. 2005, 71, 6121–6125. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aulestia, M.; Flores, A.; Mangas, E.L.; Pérez-Pulido, A.J.; Santero, E.; Camacho, E.M. Isolation and genomic characterization of the ibuprofen-degrading bacterium Sphingomonas strain MPO218. Environ. Microbiol. 2021, 23, 267–280. [Google Scholar] [CrossRef]
- Arora, P.K. Bacilli-Mediated Degradation of Xenobiotic Compounds and Heavy Metals. Front. Bioeng. Biotechnol. 2020, 8, 1–28. [Google Scholar] [CrossRef] [PubMed]
- Marchlewicz, A.; Guzik, U.; Smułek, W.; Wojcieszyńska, D. Exploring the Degradation of Ibuprofen by Bacillus thuringiensis B1(2015b): The New Pathway and Factors Affecting Degradation. Molecules 2017, 22, 1676. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.W.; Hsiao, W.C.; Fan, C.H.; Chang, B.V. Bacterial communities associated with sulfonamide antibiotics degradation in sludge-amended soil. Environ. Sci. Pollut. Res. 2016, 23, 19754–19763. [Google Scholar] [CrossRef] [PubMed]
- Kraigher, B.; Mandic-Mulec, I. Influence of diclofenac on activated sludge bacterial communities in fed-batch reactors. Food Technol. Biotechnol. 2020, 58, 402–410. [Google Scholar] [CrossRef]
| Properties | Unit | SAS | SVS | SPS |
|---|---|---|---|---|
| pH | 8.30 ± 0.01 c | 8.17 ± 0.02 b | 7.04 ± 0.02 a | |
| EC | (dS m−1) | 2.23 ± 0.02 b | 2.06 ± 0.04 a | 3.00 ± 0.03 c |
| TC | (g kg−1) | 219.7 ± 7.60 b | 180.0 ± 6.2 a | 178.0 ± 1.00 a |
| TKN | (g kg−1) | 7.76 ± 0.19 a | 10.46 ± 0.45 b | 7.31 ± 0.18 a |
| TOC | (g kg−1) | 183.7 ± 5.50 b | 154.0 ± 8.70 a | 149.0 ± 10.40 a |
| C/N | 28.33 ± 1.65 c | 17.24 ± 1.32 a | 24.36 ± 0.54 b | |
| TEC | (g kg−1) | 3.18 ± 0.02 b | 2.32 ± 0.03 a | 5.87 ± 0.02 c |
| HA | (g kg−1) | 1.15 ± 0.06 b | 0.81 ± 0.06 a | 3.92 ± 0.02 c |
| FA | (g kg−1) | 2.03 ± 0.05 b | 1.51 ± 0.08 a | 1.96 ± 0.02 b |
| WSC | (g kg−1) | 14.68 ± 0.57 c | 6.50 ± 0.20 b | 1.66 ± 0.27 a |
| Lignin | (g kg−1) | 57.41 ± 1.09 b | 69.99 ± 3.72 c | 24.06 ± 1.10 a |
| Hemicellulose | (g kg−1) | 95.23 ± 10.94 b | 82.49 ± 12.35 b | 28.88 ± 9.40 a |
| Cellulose | (g kg−1) | 71.98 ± 3.62 c | 42.96 ± 1.89 b | 17.32 ± 0.25 a |
| SAS | SVS | SPS | |
|---|---|---|---|
| Ibuprofen | |||
| Kd ± SE (L kg−1) | 3.7 ± 0.2 b | 2.1 ± 0.5 a | 3.8 ± 2.3 b |
| R2 | 0.95 | 0.99 | 0.98 |
| Koc± SE | 20 ± 1 a | 18 ± 1 a | 25 ± 1 b |
| Adsorption (%) | 6 ± 1 | 5 ± 1 | 7 ± 1 |
| Diclofenac | |||
| Kd ± SE (L kg−1) | 12.3 ± 0.5 a | 13.2 ± 0.5 a | 14.9 ± 8.3 a |
| R2 | 1.00 | 0.97 | 1.00 |
| Koc ± SE | 67 ± 3 a | 86 ± 3 b | 100 ± 3 b |
| Adsorption (%) | 20 ± 2 | 20 ± 3 | 25 ± 3 |
| Triclosan | |||
| Kd ± SE (L kg−1) | 1371 ± 81 b | 1049 ± 11 a | 1278 ± 759 a,b |
| R2 | 0.94 | 0.97 | 0.99 |
| Koc ± SE | 7465 ± 436 a,b | 6809 ± 71 a | 8579 ± 104 b |
| Adsorption (%) | 96 ± 1 | 95 ± 1 | 97 ± 1 |
| SAS | SVS | SPS | ||
|---|---|---|---|---|
| Ibuprofen | ||||
| C0 ± sd | (%) | 98 ± 3 | 100 ± 1 | 100 ± 1 |
| k × 102 ± sd | (d−1) | 18.7 ± 1.7 a | 75.9 ± 4.5 b | 75.9 ± 2.4 b |
| R2 | 0.94 | 0.99 | 0.99 | |
| DT50 | (d) | 4 | 1 | 1 |
| DT90 | (d) | 12 | 3 | 3 |
| D ± sd | (% at 10 d) | 82 ± 3 | 100 ± 0 | 97 ± 1 |
| Diclofenac | ||||
| C0 ± sd | (%) | 104 ± 3 | 102 ± 2 | 99 ± 2 |
| k × 102 ± sd | (d−1) | 3.0 ± 0.3 a | 11.5 ± 0.6 c | 6.7 ± 0.3 b |
| R2 | 0.91 | 0.97 | 0.98 | |
| DT50 | (d) | 23 | 6 | 10 |
| DT90 | (d) | 77 | 20 | 34 |
| D ± sd | (% at 84 d) | 89 ± 4 | 95 ± 1 | 97 ± 0 |
| Triclosan | ||||
| C0 ± sd | (%) | 102 ± 3 | 95 ± 2 | 106 ± 2 |
| k × 102 ± sd | (d−1) | 2.9 ± 0.3 b | 2.9 ± 0.2 b | 2.6 ± 0.2 a |
| R2 | 0.89 | 0.96 | 0.95 | |
| DT50 | (d) | 24 | 24 | 26 |
| DT90 | (d) | 79 | 79 | 89 |
| D ± sd | (% at 84 d) | 96 ± 1 | 91 ± 1 | 94 ± 1 |
| SAS | SVS | SPS | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Time (days) | BC | 10 | 84 | BC | 10 | 84 | BC | 10 | 84 |
| Archaea | |||||||||
| Crenarchaeota | 0.07 | 0.01 | 0.07 | 0.55 a | 0.76 a | 1.99 b | 1.37 a | 0.64 a | 2.83 b |
| Bacteria Phylum | |||||||||
| Proteobacteria | 39.2 a | 54.1 b | 31.9 a | 23.4 | 27.5 | 22.1 | 44.5 b | 44.5 b | 33.5 a |
| Actinobacteria | 32.6 b | 16.7 a | 29.4 b | 19.4 | 11.1 | 11.2 | 9.99 | 8.25 | 8.54 |
| Bacteroidetes | 19.8 | 20.5 | 19.0 | 18.7 a,b | 21.5 b | 8.5 a | 10.5 b | 9.3 a,b | 5.2 a |
| Chloroflexi | 0.25 a | 0.11 a | 2.49 b | 4.8 a | 4.0 a | 9.3 b | 3.2 a | 1.3 a | 9.4 b |
| Firmicutes | 0.99 b | 0.50 a | 0.66 a,b | 11.4 | 12.8 | 27.0 | 6.8 | 10.5 | 15.9 |
| Planctomycetes | 0.49 | 0.46 | 1.36 | 4.9 a,b | 2.4 a | 6.6 b | 3.8 | 2.5 | 5.8 |
| Verrucomicrobia | 1.63 a | 3.2 b | 3.99 b | 1.34 a | 2.11 b | 0.79 a | 3.3 b | 3.6 b | 1.8 a |
| Acidobacteria | 0.01 | 0.00 | 0.04 | 1.17 | 0.63 | 1.79 | 1.2 | 1.9 | 1.4 |
| Bacteria Classes | |||||||||
| Alphaproteobact | 28.8 b | 36.4 c | 21.2 a | 11.8 | 14.0 | 12.3 | 30.1 b | 29.8 b | 20.3 a |
| Betaproteobact | 2.65 a | 8.01 b | 3.45 a | 0.61 | 0.95 | 0.67 | 4.91 | 6.51 | 4.83 |
| Sphingobacteriia | 14.2 b | 15.2 b | 12.3 a | 5.2 b | 6.5 b | 2.8 a | 7.0 b | 6.5 b | 2.8 a |
| Anaerolineae | 0.0 | 0.0 | 1.1 | 1.9 | 1.2 | 2.0 | 1.5 a | 0.4 a | 4.4 b |
| Caldilineae | 0.0 | 0.0 | 0.4 | 1.9 a | 1.5 a | 6.0 b | 0.7 a | 0.3 a | 3.0 b |
| Bacilli | 0.8 | 0.5 | 0.5 | 10.9 | 11.9 | 25.0 | 6.3 | 8.0 | 14.7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Delgado-Moreno, L.; van Dillewijn, P.; Nogales, R.; Romero, E. Straw-Based Biopurification Systems to Remove Ibuprofen, Diclofenac and Triclosan from Wastewaters: Dominant Microbial Communities. Agronomy 2021, 11, 1507. https://doi.org/10.3390/agronomy11081507
Delgado-Moreno L, van Dillewijn P, Nogales R, Romero E. Straw-Based Biopurification Systems to Remove Ibuprofen, Diclofenac and Triclosan from Wastewaters: Dominant Microbial Communities. Agronomy. 2021; 11(8):1507. https://doi.org/10.3390/agronomy11081507
Chicago/Turabian StyleDelgado-Moreno, Laura, Pieter van Dillewijn, Rogelio Nogales, and Esperanza Romero. 2021. "Straw-Based Biopurification Systems to Remove Ibuprofen, Diclofenac and Triclosan from Wastewaters: Dominant Microbial Communities" Agronomy 11, no. 8: 1507. https://doi.org/10.3390/agronomy11081507
APA StyleDelgado-Moreno, L., van Dillewijn, P., Nogales, R., & Romero, E. (2021). Straw-Based Biopurification Systems to Remove Ibuprofen, Diclofenac and Triclosan from Wastewaters: Dominant Microbial Communities. Agronomy, 11(8), 1507. https://doi.org/10.3390/agronomy11081507






