The Melon Zym Locus Conferring Resistance to ZYMV: High Resolution Mapping and Candidate Gene Identification
Abstract
:1. Introduction
2. Materials and Methods
2.1. Mapping Populations and Overall Experimental Design
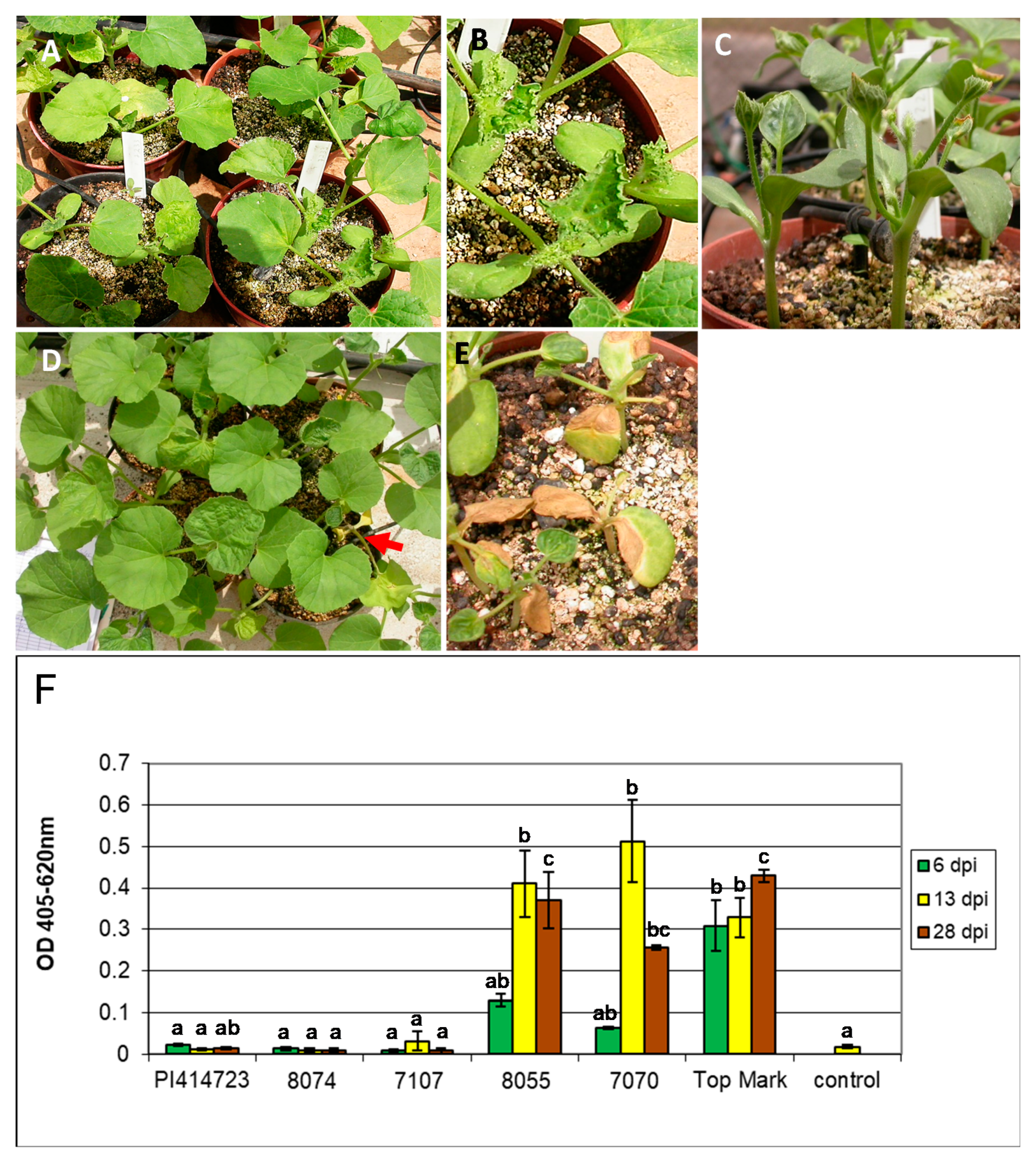
2.2. Scoring for ZYMV Resistance
2.3. BAC Library Screening and Chromosome Walking
2.4. BAC End-Sequencing and Marker Development
2.5. BAC Contig Assembly Using Clone Fingerprinting and PCR Analysis
2.6. Whole BAC Sequencing and Sequence Assembly
2.7. Sequence Annotation and Comparison
2.8. Amplification and Sequencing of Zym Alleles from a ZYMV Resistant Genotype
2.9. QTL Mapping of Zym with High Density Markers Derived from RNAseq
3. Results
3.1. ZYMV Resistance in PI 414723
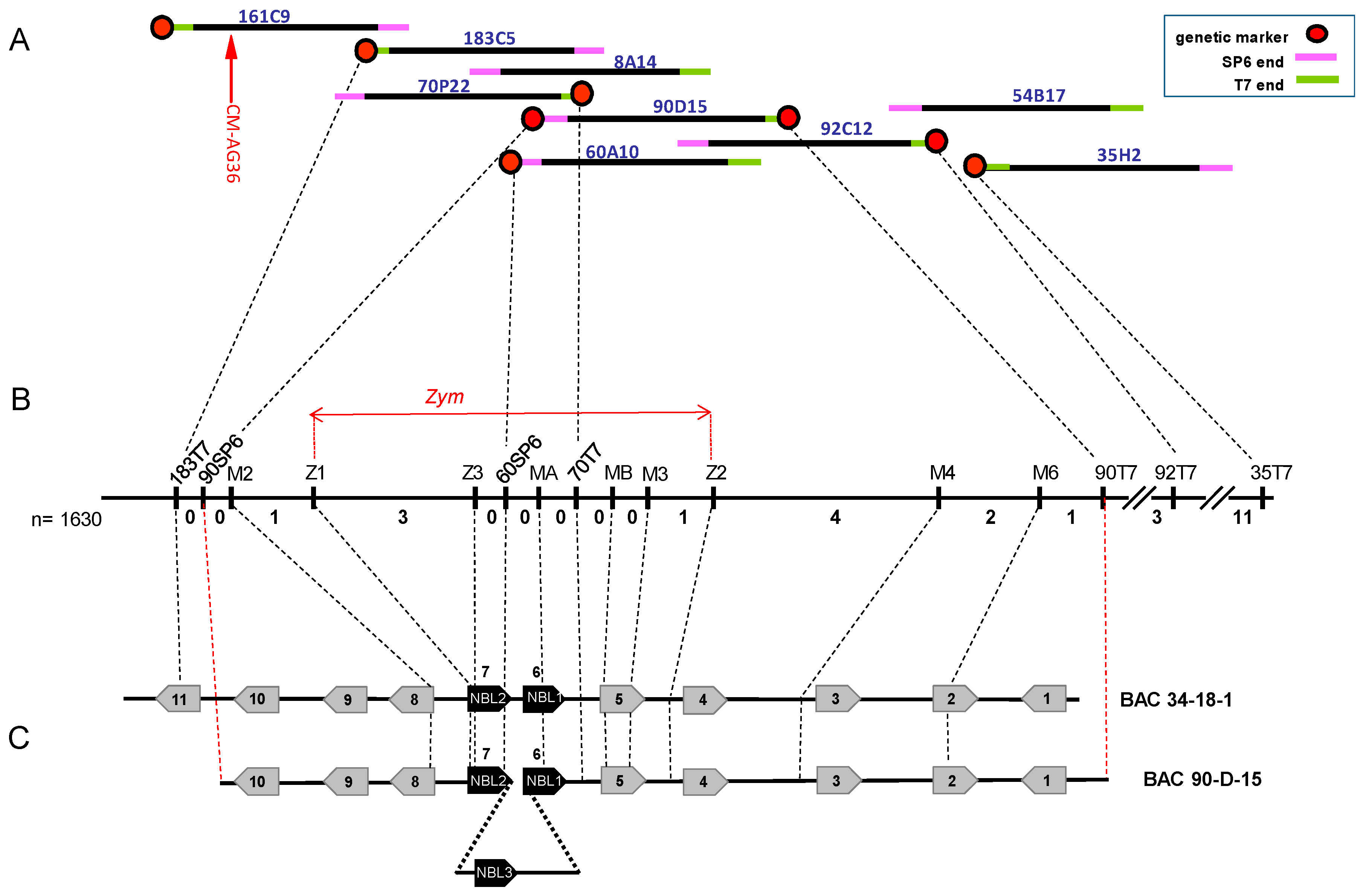
3.2. Chromosome Walking towards the Zym Locus
3.3. Fine Mapping of Zym
3.4. Clones 34-18-1 and 90D15 Contain a Small Cluster of Resistance Gene Homologs
3.5. Candidate Genes for the Zym Locus Determined by the Closest Flanking Markers
3.6. QTL Approach with Saturated SNP Detection Supports Genetic Mapping
3.7. Sequencing the Zym Locus from a ZYMV-Resistant Genotype
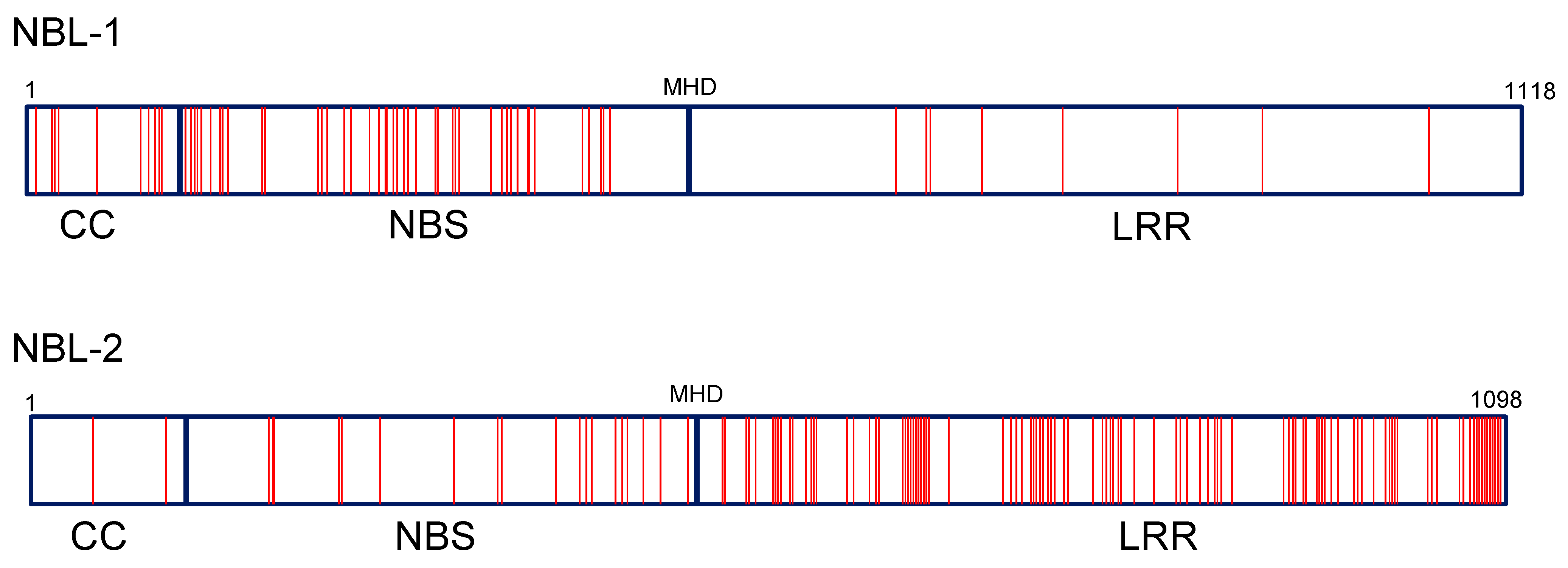
3.8. Protein Motifs and Polymorphism of Zym Candidates NBL-1, NBL-2 and NBL-3
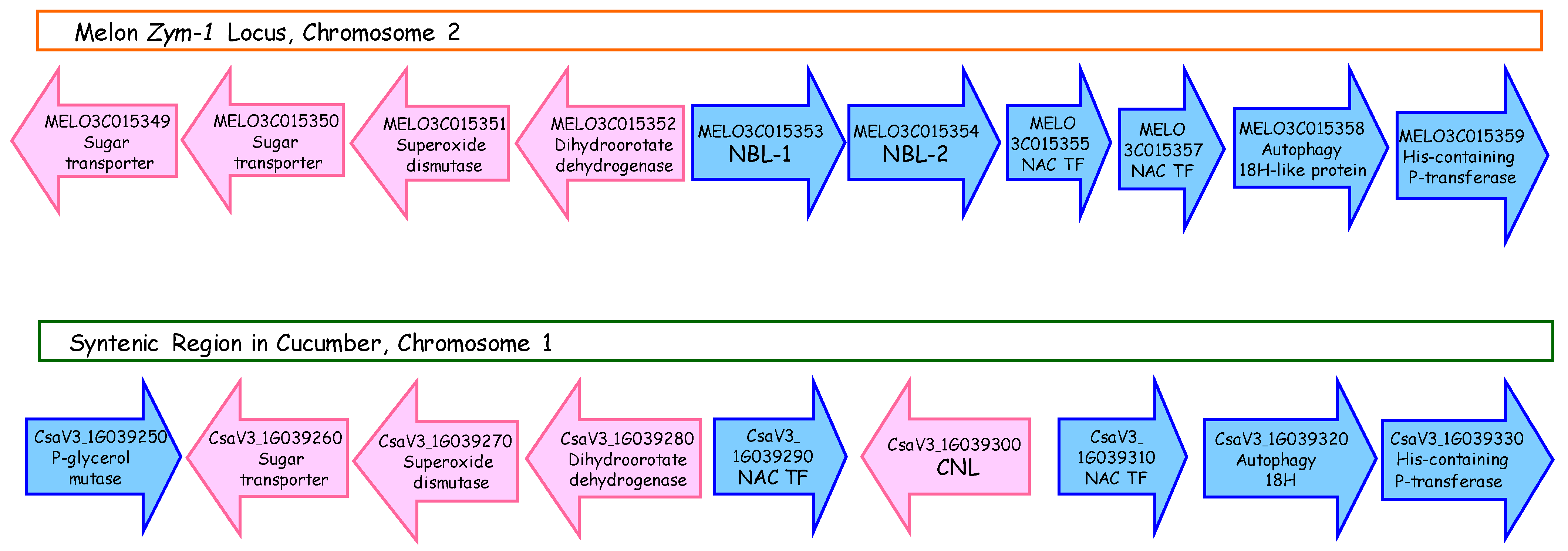
3.9. Synteny of the ZYMV Locus with a Homologous Genomic Region in Cucumber
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Arumuganathan, K.; Earle, E.D. Nuclear DNA content of some important plant species. Plant Mol. Biol. Rep. 1991, 9, 208–218. [Google Scholar] [CrossRef]
- Garcia-Mas, J.; Benjak, A.; Sanseverino, W.; Bourgeois, M.; Mir, G.; Gonźalez, V.M.; Heńaff, E.; Camara, F.; Cozzuto, L.; Lowy, E.; et al. The genome of melon (Cucumis melo L.). Proc. Natl. Acad. Sci. USA 2012, 109, 11872–11877. [Google Scholar] [CrossRef] [Green Version]
- Castanera, R.; Ruggieri, V.; Pujol, M.; Garcia-Mas, J.; Casacuberta, J.M. An improved melon reference genome with single-molecule sequencing uncovers a recent burst of transposable elements with potential impact on genes. Front. Plant Sci. 2020, 10, 1815. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yano, R.; Ariizumi, T.; Nonaka, S.; Kawazu, Y.; Zhong, S.; Mueller, L.; Giovannoni, J.J.; Rose, J.K.C.; Ezura, H. Comparative genomics of muskmelon reveals a potential role for retrotransposons in the modification of gene expression. Commun. Biol. 2020, 3, 432. [Google Scholar] [CrossRef] [PubMed]
- Yano, R.; Nonaka, S.; Ezura, H. Melonet-DB, a Grand RNA-Seq gene expression atlas in melon (Cucumis melo L.). Plant Cell Physiol. 2017, 59, E4. [Google Scholar] [CrossRef]
- Zhang, H.; Li, X.; Yu, H.; Zhang, Y.; Li, M.; Wang, H.; Wang, D.; Wang, H.; Fu, Q.; Liu, M.; et al. A high-quality melon genome assembly provides insights into genetic basis of fruit trait improvement. iScience 2019, 22, 16–27. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Deng, G.; Lian, J.; Garraway, J.; Niu, Y.; Hu, Z.; Yu, J.; Zhang, M. The chromosome-scale genome of melon dissects genetic architecture of important agronomic traits. iScience 2020, 23, 101422. [Google Scholar] [CrossRef]
- Périn, C.; Hagen, L.S.; de Conto, V.; Katzir, N.; Danin-Poleg, Y.; Portnoy, V.; Baudracco-Arnas, S.; Chadoeuf, J.; Dogimont, C.; Pitrat, M. A reference map of Cucumis melo based on two recombinant inbred line populations. Theor. Appl. Genet. 2002, 104, 1017–1034. [Google Scholar] [CrossRef]
- Fukino, N.; Ohara, T.; Monforte, A.J.; Sugiyama, M.; Sakata, Y.; Kunihisa, M.; Matsumoto, S. Identification of QTLs for resistance to powdery mildew and SSR markers diagnostic for powdery mildew resistance genes in melon (Cucumis melo L.). Theor. Appl. Genet. 2008, 118, 165–175. [Google Scholar] [CrossRef]
- Diaz, A.; Fergany, M.; Formisano, G.; Ziarsolo, P.; Blanca, J.; Fei, Z.; Staub, J.E.; Zalapa, J.E.; Cuevas, H.E.; Dace, G.; et al. A consensus linkage map for molecular markers and Quantitative Trait Loci associated with economically important traits in melon (Cucumis melo L.). BMC Plant Biol. 2011, 11, 111. [Google Scholar] [CrossRef] [Green Version]
- Brotman, Y.; Kovalski, I.; Dogimont, C.; Pitrat, M.; Portnoy, V.; Katzir, N.; Perl-Treves, R. Molecular markers linked to papaya ring spot virus resistance and Fusarium race 2 resistance in melon. Theor. Appl. Genet. 2005, 110, 337–345. [Google Scholar] [CrossRef] [PubMed]
- Brotman, Y.; Silberstein, L.; Kovalski, I.; Perin, C.; Dogimont, C.; Pitrat, M.; Klingler, J.; Thompson, G.A.; Perl-Treves, R. Resistance gene homologues in melon are linked to genetic loci conferring disease and pest resistance. Theor. Appl. Genet. 2002, 104, 1055–1063. [Google Scholar] [CrossRef] [PubMed]
- Argyris, J.M.; Pujol, M.; Martín-Hernández, A.M.; Garcia-Mas, J. Combined use of genetic and genomics resources to understand virus resistance and fruit quality traits in melon. Physiol. Plant. 2015, 155, 4–11. [Google Scholar] [CrossRef]
- Pereira, L.; Ruggieri, V.; Pérez, S.; Alexiou, K.G.; Fernández, M.; Jahrmann, T.; Pujol, M.; Garcia-Mas, J. QTL mapping of melon fruit quality traits using a high-density GBS-based genetic map. BMC Plant Biol. 2018, 18, 324. [Google Scholar] [CrossRef]
- Argyris, J.M.; Díaz, A.; Ruggieri, V.; Fernández, M.; Jahrmann, T.; Gibon, Y.; Picó, B.; Martín-Hernández, A.M.; Monforte, A.J.; Garcia-Mas, J. QTL analyses in multiple populations employed for the fine mapping and identification of candidate genes at a locus affecting sugar accumulation in melon (Cucumis melo L.). Front. Plant Sci. 2017, 8, 1679. [Google Scholar] [CrossRef] [Green Version]
- Wang, P.; Xu, X.; Zhao, G.; He, Y.; Hou, C.; Kong, W.; Zhang, J.; Liu, S.; Xu, Y.; Xu, Z. Genetic mapping and candidate gene analysis for melon resistance to Phytophthora capsici. Sci. Rep. 2020, 10, 20456. [Google Scholar] [CrossRef]
- Giner, A.; Pascual, L.; Bourgeois, M.; Gyetvai, G.; Rios, P.; Picó, B.; Troadec, C.; Bendahmane, A.; Garcia-Mas, J.; Martín-Hernández, A.M. A mutation in the melon Vacuolar Protein Sorting 41 prevents systemic infection of Cucumber mosaic virus. Sci. Rep. 2017, 7, 10471. [Google Scholar] [CrossRef] [Green Version]
- Brotman, Y.; Normantovich, M.; Goldenberg, Z.; Zvirin, Z.; Kovalski, I.; Stovbun, N.; Doniger, T.; Bolger, A.M.; Troadec, C.; Bendahmane, A.; et al. Dual resistance of melon to fusarium oxysporum races 0 and 2 and to papaya ring-spot virus is controlled by a pair of head-to-head-oriented nb-lrr genes of unusual architecture. Mol. Plant 2013, 6, 235–238. [Google Scholar] [CrossRef] [Green Version]
- Shukla, D.D.; Ward, C.W.; Brunt, A.A. The Potyviridae; CAB International: Wallingford, UK, 1994. [Google Scholar]
- Desbiez, C.; Lecoq, H. Zucchini yellow mosaic virus. Plant Pathol. 1997, 46, 809–829. [Google Scholar] [CrossRef]
- Simmons, H.E.; Holmes, E.C.; Stephenson, A.G. Rapid evolutionary dynamics of zucchini yellow mosaic virus. J. Gen. Virol. 2008, 89, 1081–1085. [Google Scholar] [CrossRef]
- Riechmann, J.L.; Lain, S.; Garcia, J.A. Highlights and prospects of potyvirus molecular biology. J. Gen. Virol. 1992, 73, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Urcuqui-Inchima, S.; Haenni, A.L.; Bernardi, F. Potyvirus proteins: A wealth of functions. Virus Res. 2001, 74, 157–175. [Google Scholar] [CrossRef]
- Wang, D.; Li, G. Biological and molecular characterization of Zucchini yellow mosaic virus isolates infecting melon in Xinjiang, China. Can. J. Plant Pathol. 2017, 39, 48–59. [Google Scholar] [CrossRef]
- Bubici, G.; Navarro, B.; Carluccio, A.V.; Ciuffo, M.; Di Serio, F.; Cillo, F. Genomic sequence variability of an Italian Zucchini yellow mosaic virus isolate. Eur. J. Plant Pathol. 2019, 156, 325–332. [Google Scholar] [CrossRef]
- Chinnadurai, C.; Kollam, M.; Ramsubhag, A.; Jayaraman, J. Genome characterization of zucchini yellow mosaic virus infecting cucurbits reveals the presence of a new genotype in Trinidad and Tobago in the Caribbean region. Arch. Virol. 2021, 166, 1661–1669. [Google Scholar] [CrossRef]
- Cordero, T.; Cerdan, L.; Carbonell, A.; Katsarou, K.; Kalantidis, K.; Daròs, J.A. Dicer-like 4 is involved in restricting the systemic movement of zucchini yellow mosaic virus in Nicotiana benthamiana. Mol. Plant-Microbe Interact. 2017, 30, 63–71. [Google Scholar] [CrossRef]
- Revers, F.; García, J.A. Molecular biology of potyviruses. Adv. Virus Res. 2015, 92, 101–199. [Google Scholar] [CrossRef]
- McCreight, J.D.; Bohn, G.W.; Kishaba, A.N. ‘Pedigree’ PI414723 melon. Cucurbit. Genet. Coop. Rep. 1992, 15, 51–52. [Google Scholar]
- Pitrat, M.; Lecoq, H. Inheritance of zucchini yellow mosaic virus resistance in Cucumis melo L. Euphytica 1984, 33, 57–61. [Google Scholar] [CrossRef]
- Fang, G.; Grumet, R. Genetic engineering of potyvirus resistance using constructs derived from the zucchini yellow mosaic virus coat protein gene. Mol. Plant Microbe Interact. 1993, 6, 358–367. [Google Scholar] [CrossRef]
- Fuchs, M.; Gonsalves, D. Resistance of transgenic hybrid squash zw-20 expressing the coat protein genes of zucchini yellow mosaic virus and watermelon mosaic virus 2 to mixed infections by both potyviruses. Bio/Tecnologye 1995, 13, 1466–1473. [Google Scholar] [CrossRef]
- Leibman, D.; Wolf, D.; Saharan, V.; Zelcer, A.; Arazi, T.; Yoel, S.; Gaba, V.; Gal-On, A. A high level of transgenic viral small RNA is associated with broad potyvirus resistance in cucurbits. Mol. Plant-Microbe Interact. 2011, 24, 1220–1238. [Google Scholar] [CrossRef] [PubMed]
- Danin-Poleg, Y.; Tadmor, Y.; Tzuri, G.; Reis, N.; Hirschberg, J.; Katzir, N. Construction of a genetic map of melon with molecular markers and horticultural traits, and localization of genes associated with ZYMV resistance. Euphytica 2002, 125, 373–384. [Google Scholar] [CrossRef]
- Harel-Beja, R.; Tzuri, G.; Portnoy, V.; Lotan-Pompan, M.; Lev, S.; Cohen, S.; Dai, N.; Yeselson, L.; Meir, A.; Libhaber, S.E.; et al. A genetic map of melon highly enriched with fruit quality QTLs and EST markers, including sugar and carotenoid metabolism genes. Theor. Appl. Genet. 2010, 121, 511–533. [Google Scholar] [CrossRef] [PubMed]
- Galpaz, N.; Gonda, I.; Shem-Tov, D.; Barad, O.; Tzuri, G.; Lev, S.; Fei, Z.; Xu, Y.; Mao, L.; Jiao, C.; et al. Deciphering genetic factors that determine melon fruit-quality traits using RNA-Seq-based high-resolution QTL and eQTL mapping. Plant J. 2018, 94, 169–191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Herman, R.; Perl-Treves, R. Characterization and inheritance of a new source of resistance to Fusarium oxysporum f. sp. melonis race 1.2 in Cucumis melo. Plant Dis. 2007, 91, 1180–1186. [Google Scholar] [CrossRef]
- Clark, M.F.; Adams, A.N. Characteristics of the microplate method of enzyme linked immunosorbent assay for the detection of plant viruses. J. Gen. Virol. 1977, 34, 475–483. [Google Scholar] [CrossRef]
- Luo, M.; Wang, Y.H.; Frisch, D.; Joobeur, T.; Wing, R.A.; Dean, R.A. Melon bacterial artificial chromosome (BAC) library construction using improved methods and identification of clones linked to the locus conferring resistance to melon Fusarium wilt (Fom-2). Genome 2001, 44, 154–162. [Google Scholar] [CrossRef]
- Bohn, G.W.; Kishaba, A.N.; McCreight, J.D. WMR-29 muskmelon breeding line. HortScience 1980, 15, 539–540. [Google Scholar]
- Budiman, M.A.; Mao, L.; Wood, T.C.; Wing, R.A. A deep-coverage tomato BAC library and prospects toward development of an STC framework for genome sequencing. Genome Res. 2000, 10, 129–136. [Google Scholar] [CrossRef]
- Neff, M.M.; Turk, E.; Kalishman, M. Web-based primer design for single nucleotide polymorphism analysis. Trends Genet. 2002, 18, 613–615. [Google Scholar] [CrossRef]
- Sonnhammer, E.L.L.; Eddy, S.R.; Birney, E.; Bateman, A.; Durbin, R. Pfam: Multiple sequence alignments and HMM-profiles of protein domains. Nucleic Acids Res. 1998, 26, 320–322. [Google Scholar] [CrossRef] [Green Version]
- Kurtz, S.; Phillippy, A.; Delcher, A.L.; Smoot, M.; Shumway, M.; Antonescu, C.; Salzberg, S.L. Versatile and open software for comparing large genomes. Genome Biol. 2004, 5, R12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Slater, G.S.C.; Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinform. 2005, 6, 31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Silberstein, L.; Kovalski, I.; Brotman, Y.; Perin, C.; Dogimont, C.; Pitrat, M.; Klingler, J.; Thompson, G.; Portnoy, V.; Katzir, N.; et al. Linkage map of Cucumis melo including phenotypic traits and sequence-characterized genes. Genome 2003, 46, 761–773. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative Genome Viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meyers, B.C.; Kozik, A.; Griego, A.; Kuang, H.; Michelmore, R.W. Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant Cell 2003, 15, 809–834. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, S.; Li, R.; Zhang, Z.; Li, L.; Gu, X.; Fan, W.; Lucas, W.J.; Wang, X.; Xie, B.; Ni, P.; et al. The genome of the cucumber, Cucumis sativus L. Nat. Genet. 2009, 41, 1275–1281. [Google Scholar] [CrossRef] [Green Version]
- Kapos, P.; Devendrakumar, K.T.; Li, X. Plant NLRs: From discovery to application. Plant Sci. 2019, 279, 3–18. [Google Scholar] [CrossRef]
- Dogimont, C.; Chovelon, V.; Pauquet, J.; Boualem, A.; Bendahmane, A. The Vat locus encodes for a CC-NBS-LRR protein that confers resistance to Aphis gossypii infestation and A. gossypii-mediated virus resistance. Plant J. 2014, 80, 993–1004. [Google Scholar] [CrossRef]
- Joobeur, T.; King, J.J.; Nolin, S.J.; Thomas, C.E.; Dean, R.A. The fusarium wilt resistance locus Fom-2 of melon contains a single resistance gene with complex features. Plant J. 2004, 39, 283–297. [Google Scholar] [CrossRef]
- Nieto, C.; Morales, M.; Orjeda, G.; Clepet, C.; Monfort, A.; Sturbois, B.; Puigdomènech, P.; Pitrat, M.; Caboche, M.; Dogimont, C.; et al. An eIF4E allele confers resistance to an uncapped and non-polyadenylated RNA virus in melon. Plant J. 2006, 48, 452–462. [Google Scholar] [CrossRef]
- Taler, D.; Galperin, M.; Benjamin, I.; Cohen, Y.; Kenigsbuch, D. Plant eR genes that encode photorespiratory enzymes confer resistance against disease. Plant Cell 2004, 16, 172–184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Danin-Poleg, Y.; Paris, H.S.; Cohen, S.; Rabinowitch, H.D.; Karchi, Z. Oligogenic inheritance of resistance to zucchini yellow mosaic virus in melons. Euphytica 1997, 93, 331–337. [Google Scholar] [CrossRef]
- Goldenberg, T. High Resolution Mapping and Physical Delimitation of the Fom-1/Prv and Zym Loci in Melon. Ph.D. Thesis, Bar Ilan University, Ramat Gan, Israel, 2009. [Google Scholar]
- Díaz, J.A.; Mallor, C.; Carmen Soria, C.; Rocío Camero, R.; Elisa Garzo, E.; Alberto Fereres, A.; Alvarez, J.M.; Gómez-Guillamón, M.L.; Luis-Arteaga, E.M. Potential sources of resistance for melon to nonpersistently aphid-borne viruses. Plant Dis. 2003, 87, 960–964. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Romay, G.; Pitrat, M.; Lecoq, H.; Wipf-Scheibel, C.; Millot, P.; Girardot, G.; Desbiez, C. Resistance against melon chlorotic mosaic virus and tomato leaf curl New Delhi virus in melon. Plant Dis. 2019, 103, 2913–2919. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- van Wersch, R.; Li, X.; Zhang, Y. Mighty dwarfs: Arabidopsis autoimmune mutants and their usages in genetic dissection of plant immunity. Front. Plant Sci. 2016, 7, 1717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Venkatesh, J.; Kang, B.C. Current views on temperature-modulated R gene-mediated plant defense responses and tradeoffs between plant growth and immunity. Curr. Opin. Plant Biol. 2019, 50, 9–17. [Google Scholar] [CrossRef]
- Olsen, A.N.; Ernst, H.A.; Leggio, L.L.; Skriver, K. Transcriptional networks in plants NAC transcription factors: Structurally distinct, functionally diverse. Trends Plant Sci. 2005, 10, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Koyama, H.; Bhati, K.K.; Alok, A. The biotechnological importance of the plant-specific NAC transcription factor family in crop improvement. J. Plant Res. 2021, 134, 475–495. [Google Scholar] [CrossRef]
- Wang, X.; Goregaoker, S.P.; Culver, J.N. Interaction of the tobacco mosaic virus replicase protein with a NAC domain transcription factor is associated with the suppression of systemic host defenses. J. Virol. 2009, 83, 9720–9730. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, Q.; Sanz-Burgos, A.P.; Guo, H.; García, J.A.; Gutiérrez, C. GRAB proteins, novel members of the NAC domain family, isolated by their interaction with a geminivirus protein. Plant Mol. Biol. 1999, 39, 647–656. [Google Scholar] [CrossRef] [PubMed]
- Selth, L.A.; Dogra, S.C.; Rasheed, M.S.; Healy, H.; Randles, J.W.; Rezaian, M.A. A NAC domain protein interacts with Tomato leaf curl virus replication accessory protein and enhances viral replication. Plant Cell 2005, 17, 311–325. [Google Scholar] [CrossRef] [Green Version]
- Bergelson, J.; Kreitman, M.; Stahl, E.A.; Tian, D. Evolutionary dynamics of plant R-genes. Science 2001, 292, 2281–2285. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Friedman, A.R.; Baker, B.J. The evolution of resistance genes in multi-protein plant resistance systems. Curr. Opin. Genet. Dev. 2007, 17, 493–499. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kabelka, E.; Ullah, Z.; Grumet, R. Multiple alleles for zucchini yellow mosaic virus resistance at the zym locus in cucumber. Theor. Appl. Genet. 1997, 95, 997–1004. [Google Scholar] [CrossRef]
- Amano, M.; Mochizuki, A.; Kawagoe, Y.; Iwahori, K.; Niwa, K.; Svoboda, J.; Maeda, T.; Imura, Y. High-resolution mapping of zym, a recessive gene for Zucchini yellow mosaic virus resistance in cucumber. Theor. Appl. Genet. 2013, 126, 2983–2993. [Google Scholar] [CrossRef] [PubMed]
- Pascual, L.; Yan, J.; Pujol, M.; Monforte, A.J.; Picó, B.; Martín-Hernández, A.M. CmVPS41 is a general gatekeeper for resistance to cucumber mosaic virus phloem entry in melon. Front. Plant Sci. 2019, 10, 1219. [Google Scholar] [CrossRef] [Green Version]
- García-Almodóvar, R.C.; Gosalvez, B.; Aranda, M.A.; Burgos, L. Production of transgenic diploid Cucumis melo plants. Plant Cell. Tissue Organ Cult. 2017, 130, 323–333. [Google Scholar] [CrossRef]
- Chovelon, V.; Restier, V.; Giovinazzo, N.; Dogimont, C.; Aarrouf, J. Histological study of organogenesis in Cucumis melo L. after genetic transformation: Why is it difficult to obtain transgenic plants? Plant Cell Rep. 2011, 30, 2001–2011. [Google Scholar] [CrossRef]


| Population Number | Type | ZYMV-Resistant Parent | ZYMV-Susceptible Parent | Reference | # Offspring |
|---|---|---|---|---|---|
| 1 | RIL | PI 414723 | Dulce | Harel-Beja et al., 2010 | 97 |
| 2 | RIL | PI 414723 | Top Mark | This study | 103 |
| 3 | RIL | PI 414723 | Védrantais | Périn et al., 2002 | 61 |
| 4 | F2 | PI 414723 | BIZ | Herman & Perl-Treves, 2007 [37] | 141 |
| 5 | BC1 | PI 414723 | Védrantais | Brotman et al., 2013 | 1228 |
| Marker Name | Primer Name | Primer Sequence 5′-3′ | Tm (°C) | Size (bp) | Diagnostic Enzyme |
|---|---|---|---|---|---|
| 183C5-T7 | 183C5F-T7 | CCAAAAACACAAAGGAAGTTTATA | 56 | 385 | Csp6I |
| 183C5R-T7 | AGAATCACTTGTAAACATAGCCTT | ||||
| 90D15-sp6 | 90D15-FSP6 | ACAAACGGATAAATGAAAAGGA | 56–60 | 320 | HindIII |
| 90D15-RSP6 | ACTTCCTGCTAGTGGCGTTT | ||||
| M2-new | M2Fnew | GAATCATTTAACAAAACAAAAGTA | 55–60 | 1306 | TaqI, MseI, TaiI |
| M2-R | GCATCTCTTATTAGTTTTCGAG | ||||
| Z1 | Z1-F | GTATCGAGTTTCTCCCTGATCCT | 53 | 1112 | TruI |
| Z1-R | TGTGTGTTGGACTTGTGATTTGT | ||||
| Z3 | Z3-F | ACAATCATTTTCTACAACGGAACA | 60 | 1160 | EcoRV, TasI |
| Z3-R | GGATCAGGGAGAAACTCGATACTA | ||||
| 60A10-sp6 | 60A10-F-SP6 | TTCCAACGATTTTTCATCACC | 52 | 412 | TaqI |
| 60A10-R-SP6 | TGCAAATCCTTTGGCAAATA | ||||
| 60A10-F4-sp6 | 60A10-F4-SP6 | TTCGAAAGTTGGAATTACATTGG | 55–57 | 385 | BfmI, HincII, ApoI, HincII |
| 60A10-R4-SP6 | GGTGATGAAAAATCGTTGGAA | ||||
| MA | MA-F | CACTTGGCAGT CCCATTAGTCTTTGAC | 60 | 1395 | MspI, XpaI |
| MA-R | GAAGGAGAAGTAATTGGTCGAGATGAT | ||||
| MB | MB-F | GTGAAG ATGATACTCG TGTATAACC | 60 | 1404 | TaqI, TasI |
| MB-R | GAACACGATATTATGGTTAAATCC | ||||
| 70P22-T7 | 70P22-F-T7 | TTTCTGTTGTTTTCTTTTCCTTGC | 56 | 345 | SalI |
| 70P22-R-T7 | CCATTGTATTGTTGTTGGTGGAGA | ||||
| M3 | M3-F | CTTCATGAGG GTGGACAGAG GAAAG | 55–56 | 1161 | SatI |
| M3-R | CTATGTTCTTTCATGATCGCTC | ||||
| Z2 | Z2-F | TTAAATTTGGGTTTTCCATTGACT | 55 | 1054 | FspBI, TruI, TaiI |
| Z2-R | ATTAAGGGTTTGAATTGCAAATGT | ||||
| M4 | M4-F | GATTTACAAAATTAGTTGAC AAG | 60 | 1653 | AluI |
| M4-R | GTTAAAATTAGGTTTCCACAAC | ||||
| M6 | M6-F | CTAACGTT TAGTTTGAAC AACC | 55–56 | 1378 | TaiI |
| M6-R | GAGGCATTGGCATCCAAAAACTC | ||||
| 90D15-T7 | 90D15-FT7 | GCCTAGCAGTGGATTTTTCG | 59 | 471 | TasI |
| 90D15-RT7 | TCGAAACTTTTAAAGGCAGCA | ||||
| 92C12-T7 | 92C12-FT7 | ACTTTCCATCCCACCAATGCT | 56–60 | 200 | PCR product presence |
| 92C12-RT7 | AATCCCCACTTTCTTGGTGGA | ||||
| 35H2FN-T7 | 35H2FN-T7 | TTCTTTGTCTCCTTTCTCCTC | 56 | 415 | DraI |
| 35H2R-T7 | ATATTGGTTTGGTCGGTTGG | ||||
| 35H2F4-T7 | 35H2F4-T7 | TGAATCGAGAGGGGTCATTG | 56 | 330 | HphI |
| 35H2Rx-T7 | ATATCTGAGGTACGAAGGAGAAAC |
| № | Gene ID | Sequence Homology | Gene Model (Exon Positions on BAC Clone 34-18-1) |
|---|---|---|---|
| 1 | MELO 3C015360 | TF IIIB 60 kD subunit | 128..243, 1025..1105, 1335..1472, 1557..1580, 2041..2145, 2354..2407, 2961..3176 |
| 2 | MELO 3C015359 | His-containing phospho-trasferase | 8369..8440, 8743..8815, 9706..9774, 9933..10,063, 10,571..10,633 |
| 3 | MELO 3C015358 | Autophagy 18H-like protein | 18,420..18,424, 19,353..19,877, 19,977..20,104, 24,688..24,863, 24,942..25,736, 25,813..26,058, 26,831..27,265, 27,612..28,070, 29,451..29,654 |
| 4 | MELO 3C015357 | NAC domain Transcrtiption Factor | 36,438..36,710, 36,919..37,188, 37,439..37,765, 37,859..37,984, 38,217..38,491, 38,571..38,748 |
| 5 | MELO 3C015355 | NAC domain Transcrtiption Factor | 404,93..40,840, 40,960..42,132, 42,340..42,654, 42,742..42,882, 43,402..43,664, 43,738..43,915 |
| 6 | MELO 3C015354 | CC-NBS-LRR (NBL1) | 45,722..49,075 |
| 7 | MELO 3C015353 | CC-NBS-LRR (NBL2) | 52,452..55,706 |
| 8 | MELO 3C015352 | Dihidroorotate Dehydrogenase | 58,807..58,941, 60,448..60,531, 61,677..62,000, 62,216..62,470, 63,432..63,512, 63,593..63,730, 63,940..64,077, 64,813..64,848, 64,967..65,068, 65,791..65,862 |
| 9 | MELO 3C015351 | Superoxide Dismutase | 68,656..68,712, 69,076..69,183, 70,036..70,230, 71,369..71,536, 71,942..72,016, 72,091..72,117, 72,209..72,232, 72,771..72,893 |
| 10 | MELO 3C015350 | sugar transporter ERD-like | 73,954..74,019, 75,011..75,073, 75,370..75,459, 75,558..75,626, 76,044..76,100, 76,193..76,258, 76,956..77,036, 77,122..77,181, 77,548..77,640, 77,736..77,825, 77,930..78,010, 78,098..78,133, 78,218..78,283, 78,366..78,419, 78,519..78,578, 78,901..78,960, 79,262..79,378, 79,468..79,593 |
| 11 | MELO 3C015349 | sugar transporter ERD-like | 81,079..81,225, 81,969..82,031, 82,758..82,844, 82,993..83,055, 83,542..83,604, 83,711..83,773, 84,162..84,248, 84,336..84,392, 84,701..84,793, 84,876..84,959, 85,289..85,381, 85,856..85,942, 86,026..86,085 |
| NBL Paralogue Identity, % | ||
|---|---|---|
| Gene | NBL-2 | NBL-3 |
| NBL-1 | 88 (80) | 88 (81) |
| NBL-2 | 92 (86) | |
| Sequence | Total Length | Polymorphic Sites No. | Unique to WMR 29 | Unique to MR-1 | Unique to PI 414723 | |
|---|---|---|---|---|---|---|
| NBL-1 | DNA | 3354 | 120 | 4 | 5 | 111 |
| Protein | 1118 | 63 | 2 | 4 | 57 | |
| NBL-2 | DNA | 3294 | 171 | 94 | 57 | 73 |
| Protein | 1098 | 126 | 43 | 39 | 60 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Adler-Berke, N.; Goldenberg, Y.; Brotman, Y.; Kovalski, I.; Gal-On, A.; Doniger, T.; Harel-Beja, R.; Troadec, C.; Bendahmane, A.; Pitrat, M.; et al. The Melon Zym Locus Conferring Resistance to ZYMV: High Resolution Mapping and Candidate Gene Identification. Agronomy 2021, 11, 2427. https://doi.org/10.3390/agronomy11122427
Adler-Berke N, Goldenberg Y, Brotman Y, Kovalski I, Gal-On A, Doniger T, Harel-Beja R, Troadec C, Bendahmane A, Pitrat M, et al. The Melon Zym Locus Conferring Resistance to ZYMV: High Resolution Mapping and Candidate Gene Identification. Agronomy. 2021; 11(12):2427. https://doi.org/10.3390/agronomy11122427
Chicago/Turabian StyleAdler-Berke, Nastacia, Yitzchak Goldenberg, Yariv Brotman, Irina Kovalski, Amit Gal-On, Tirza Doniger, Rotem Harel-Beja, Christelle Troadec, Abdelhafid Bendahmane, Michel Pitrat, and et al. 2021. "The Melon Zym Locus Conferring Resistance to ZYMV: High Resolution Mapping and Candidate Gene Identification" Agronomy 11, no. 12: 2427. https://doi.org/10.3390/agronomy11122427
APA StyleAdler-Berke, N., Goldenberg, Y., Brotman, Y., Kovalski, I., Gal-On, A., Doniger, T., Harel-Beja, R., Troadec, C., Bendahmane, A., Pitrat, M., Dogimont, C., Katzir, N., & Perl-Treves, R. (2021). The Melon Zym Locus Conferring Resistance to ZYMV: High Resolution Mapping and Candidate Gene Identification. Agronomy, 11(12), 2427. https://doi.org/10.3390/agronomy11122427







