Study of Variability in Root System Architecture of Spanish Triticum turgidum L. Subspecies and Analysis of the Presence of a MITE Element Inserted in the TtDro1B Gene: Evolutionary Implications
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. RSA Analysis
2.3. DNA Extraction
2.4. MITE Detection in Triticum turgidum Subspecies
2.5. MITE Detection in Aegilops Species
2.6. Statistical Analysis
3. Results and Discussion
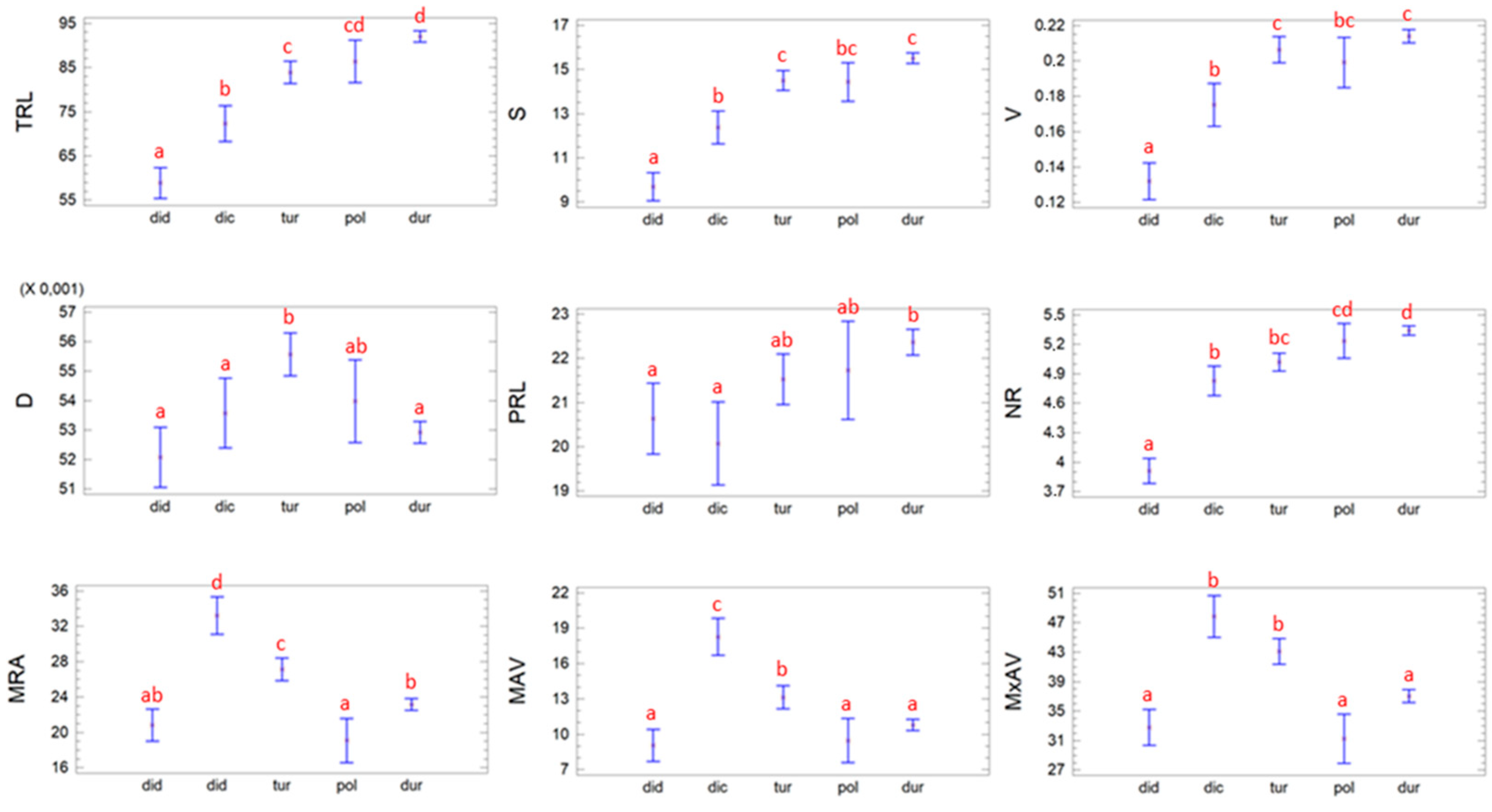
3.1. Study of the RSA in T. turgidum Subspecies
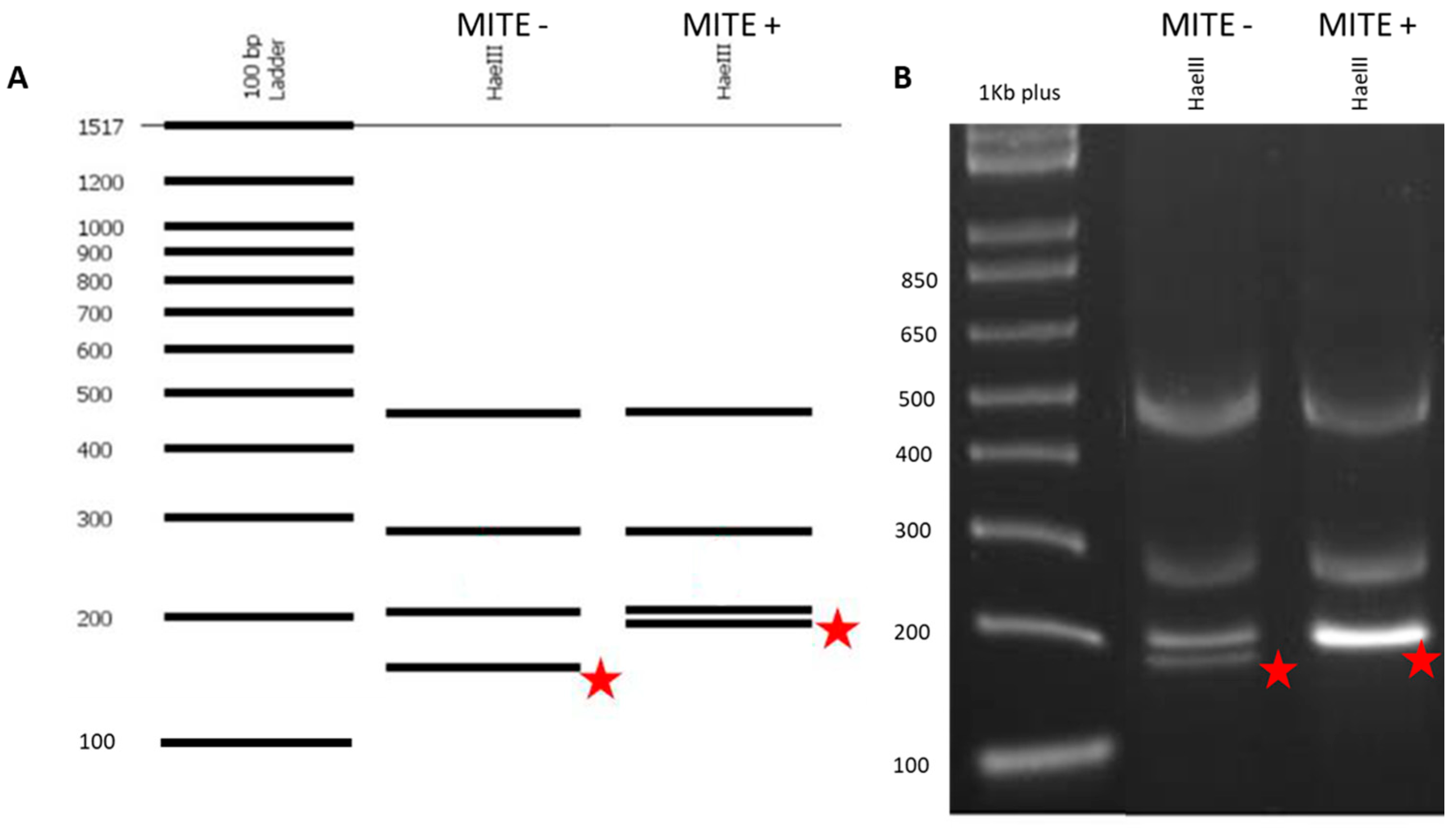
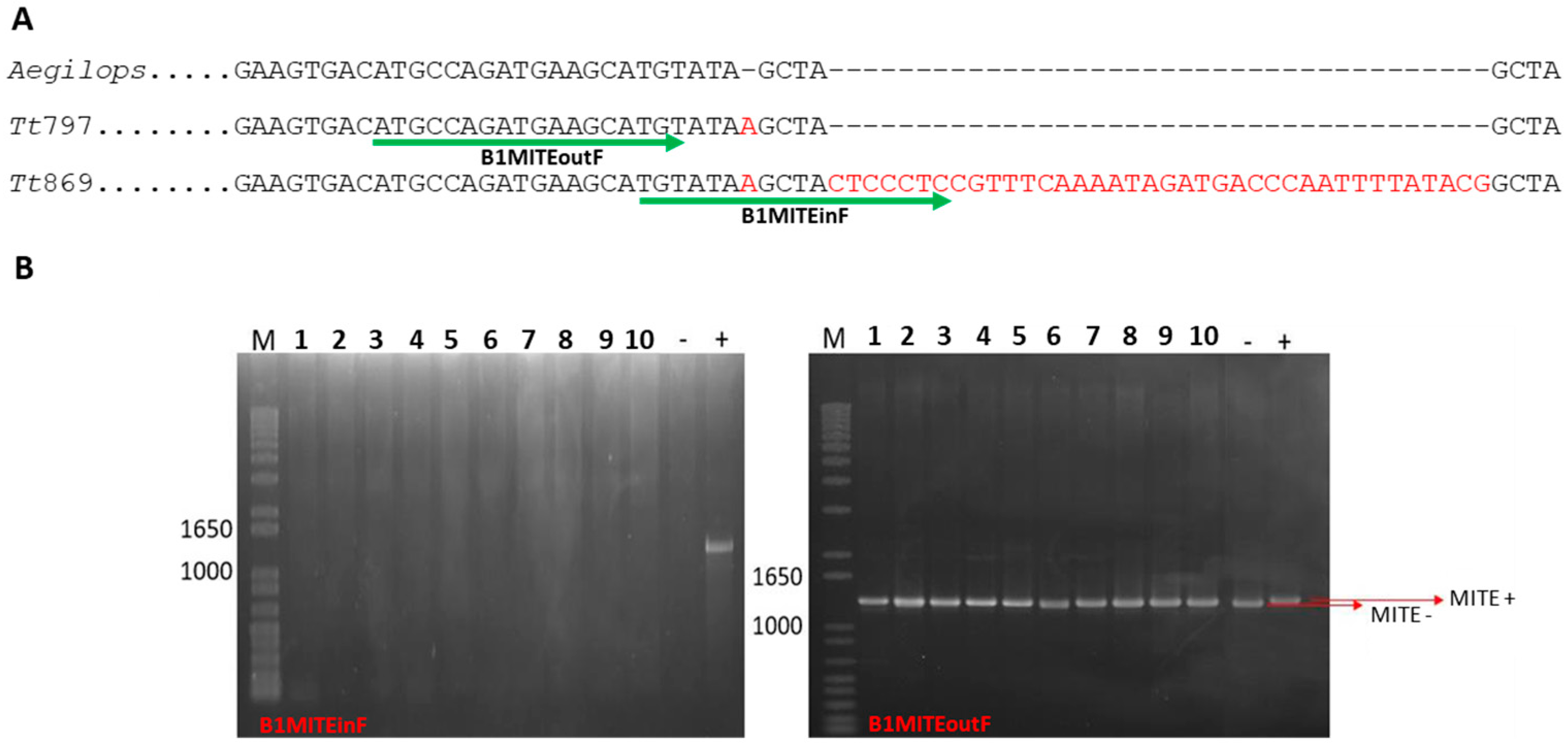
3.2. Analysis of a MITE Element
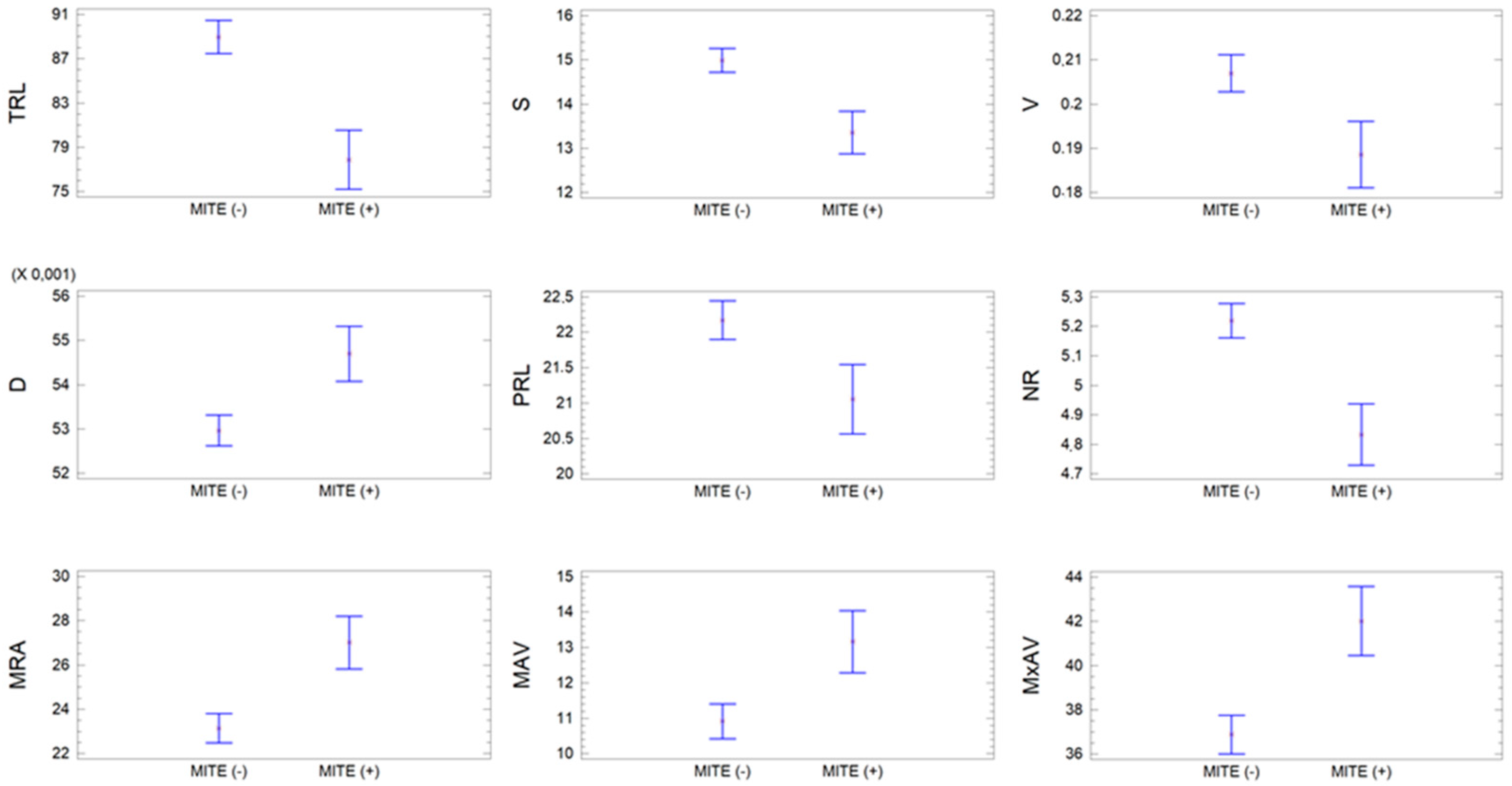
3.3. Relationship between the Presence of the MITE Element Insertion and the RSA
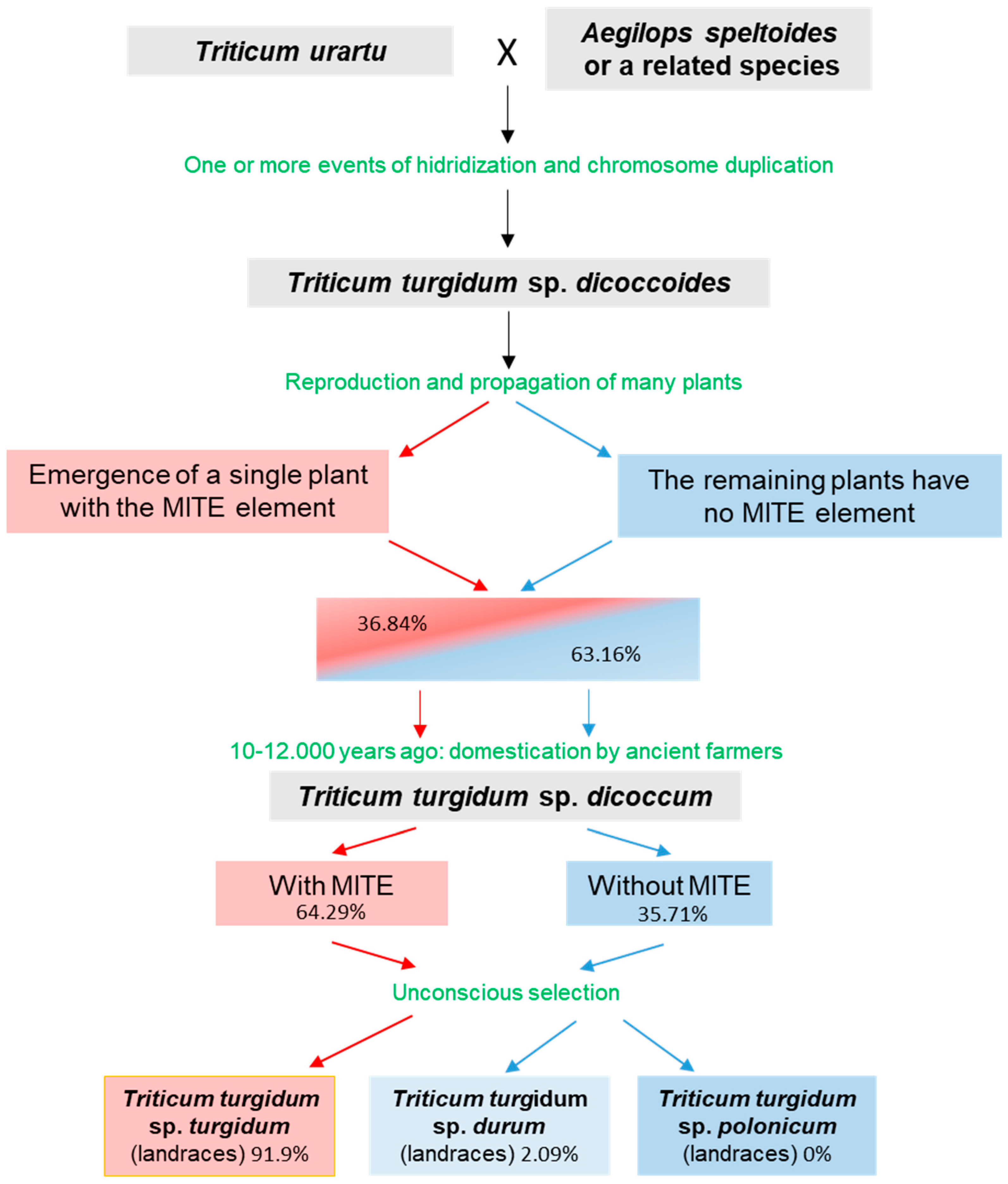
3.4. MITE as a Subspecies Marker
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Petersen, G.; Seberg, O.; Yde, M.; Berthelsen, K. Phylogenetic relationships of Triticum and Aegilops and evidence for the origin of the A, B, and D genomes of common wheat (Triticum aestivum). Mol. Phylogenet. Evol. 2006, 39, 70–82. [Google Scholar] [CrossRef]
- Dizkirici, A.; Kansu, C.; Onde, S.; Birsin, M.; Özgen, M.; Kaya, Z. Phylogenetic relationships among Triticum L. and Aegilops L. species as genome progenitors of bread wheat based on sequence diversity in trnT-F region of chloroplast DNA. Genet. Resour. Crop. Evol. 2013, 60, 2227–2240. [Google Scholar] [CrossRef]
- Igrejas, G.; Ikeda, T.M.; Guzmán, C. (Eds.) Wheat Quality for Improving Processing and Human Health; Springer: Cham, Switzerland, 2020. [Google Scholar]
- Tadesse, W.; Sanchez-Garcia, M.; Gizaw Assefa, S.; Amri, A.; Bishaw, Z.; Ogbonnaya, F.C.; Baum, M. Genetic Gains in Wheat Breeding and Its Role in Feeding the World. Crop Breed. Genet. Genom. 2019, 1, e190005. [Google Scholar] [CrossRef] [Green Version]
- FAO. Available online: http://www.fao.org/faostat/es/#home (accessed on 20 September 2021).
- Xynias, I.N.; Mylonas, I.; Korpetis, E.G.; Ninou, E.; Tsaballa, A.; Avdikos, I.D.; Mavromatis, A.G. Durum Wheat Breeding in the Mediterranean Region: Current Status and Future Prospects. Agronomy 2020, 21, 432. [Google Scholar] [CrossRef] [Green Version]
- Tang, Y.; Kang, H.Y.; Tang, L.; Diao, C.D.; Li, D.Y.; Zhu, W.; Fan, X.; Wang, Y.; Zeng, J.; Xu, L.L.; et al. Phylogenetic analysis of tetraploid wheat based on nuclear DMC1 gene. Biochem. Syst. Ecol. 2017, 70, 239–246. [Google Scholar] [CrossRef]
- Zhang, W.; Zhang, M.; Zhu, X.; Cao, Y.; Sun, Q.; Ma, G.; Chao, S.; Yan, C.; Xu, S.S.; Cai, X. Molecular cytogenetic and genomic analyses reveal new insights into the origin of the wheat B genome. Theor. Appl. Genet. 2018, 131, 365–375. [Google Scholar] [CrossRef]
- Miki, Y.; Yoshida, K.; Mizuno, N.; Nasuda, S.; Sato, K.; Takumi, S. Origin of wheat B-genome chromosomes inferred from RNA sequencing analysis of leaf transcripts from section Sitopsis species of Aegilops. DNA Res. 2019, 26, 171–182. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zohary, D.; Hopf, M.; Weiss, E. Domestication of Plants in the Old World: The Origin and Spread of Domesticated Plants in Southwest Asia, Europe, and the Mediterranean Basin, 4th ed.; Oxford University Press: Oxford, UK, 2012. [Google Scholar]
- Gioia, T.; Nagel, K.A.; Beleggia, R.; Fragasso, M.; Ficco, D.B.; Pieruschka, R.; De Vita, P.; Fiorani, F.; Papa, R. Impact of domestication on the phenotypic architecture of durum wheat under contrasting nitrogen fertilization. EXBOTJ 2015, 66, 5519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gupta, P.K.; Mir, R.R.; Mohan, A.; Kumar, J. Wheat Genomics: Present Status and Future Prospects. Int. J. Plant Genom. 2008, 2008, 896451. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Moreno, F.; Solís, I.; Noguero, D.; Blanco, A.; Özberk, İ.; Nsarellah, N.; Elias, E.; Mylonas, I.; Soriano, J.M. Durum wheat in the Mediterranean Rim: Historical evolution and genetic resources. Genet. Resour. Crop Evol. 2020, 67, 1415–1436. [Google Scholar] [CrossRef]
- Khoury, C.K.; Bjorkman, A.D.; Dempewolf, H.; Ramirez-Villegas, J.; Guarino, L.; Jarvis, A.; Rieseberg, L.H.; Struik, P.C. Increasing homogeneity in global food supplies and the implications for food security. Proc. Natl. Acad. Sci. USA 2014, 111, 4001–4006. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kabbaj, H.; Sall, A.T.; Al-Abdallat, A.; Geleta, M.; Amri, A.; Filali-Maltouf, A.; Belkadi, B.; Ortiz, R.; Bassi, F.M. Genetic Diversity within a Global Panel of Durum Wheat (Triticum durum) Landraces and Modern Germplasm Reveals the History of Alleles Exchange. Front. Plant Sci. 2017, 8, 1277. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Igartua, E.; Gracia, M.P.; Lasa, J.M.; Medina, B.; Molina-Cano, J.L.; Montoya, J.L.; Romagosa, I. The Spanish barley core collection. Genet. Resour. Crop Evol. 1998, 45, 475–481. [Google Scholar] [CrossRef] [Green Version]
- Ruiz, M.; Giraldo, P.; Royo, C.; Carrillo, J.M. Creation and Validation of the Spanish Durum Wheat Core Collection. Crop Sci. 2013, 53, 2530–2537. [Google Scholar] [CrossRef] [Green Version]
- Pascual, L.; Fernández, M.; Aparicio, N.; López-Fernández, M.; Fité, R.; Giraldo, P.; Ruiz, M. Development of a Multipurpose Core Collection of Bread Wheat Based on High-Throughput Genotyping Data. Agronomy 2020, 10, 534. [Google Scholar] [CrossRef] [Green Version]
- Lynch, J.P. Root architecture and plant productivity. Plant Physiol. (Bethesda) 1995, 109, 7–13. [Google Scholar] [CrossRef]
- Lynch, J.P.; Brown, K.B. Topsoil foraging—An architectural adaptation of plants to low phosphorus availability. Plant Soil 2001, 237, 225–237. [Google Scholar] [CrossRef]
- Lynch, J.P. Steep, cheap and deep: An ideotype to optimize water and N acquisition by maize root systems. Ann. Bot. 2013, 112, 347–357. [Google Scholar] [CrossRef] [Green Version]
- White, P.J.; George, T.S.; Dupuy, L.X.; Karley, A.J.; Valentine, T.A.; Wiesel, L.; Wishart, J. Root traits for infertile soils. Front. Plant Sci. 2013, 4, 193. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vives-Peris, V.; López-Climent, M.F.; Pérez-Clemente, R.M.; Gómez-Cadenas, A. Root Involvement in Plant Responses to Adverse Environmental Conditions. Agronomy 2020, 10, 942. [Google Scholar] [CrossRef]
- Paez-Garcia, A.; Motes, C.M.; Scheible, W.; Chen, R.; Blancaflor, E.B.; Monteros, M.J. Root Traits and Phenotyping Strategies for Plant Improvement. Plants 2015, 4, 334–355. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, M.; Giraldo, P.; González, J. Phenotypic variation in root architecture traits and their relationship with eco-geographical and agronomic features in a core collection of tetraploid wheat landraces (Triticum turgidum L.). Euphytica 2018, 214, 1–17. [Google Scholar] [CrossRef]
- Sanguineti, M.C.; Li, S.; Maccaferri, M.; Corneti, S.; Rotondo, Y.; Chiari, T.; Tuberosa, R. Genetic dissection of seminal root architecture in elite durum wheat germplasm. Ann. Appl. Biol. 2007, 151, 291–305. [Google Scholar] [CrossRef]
- Sinha, S.; Rani, M.; Kumar, A.; Kumar, S.; Venkatesh, K.; Mandal, P. Natural variation in root system architecture in diverse wheat genotypes grown under different nitrate conditions and root growth media. Theor. Exp. Plant Physiol. 2018, 30, 223–234. [Google Scholar] [CrossRef]
- Roselló, M.; Royo, C.; Sanchez-Garcia, M.; Soriano, J.M. Genetic Dissection of the Seminal Root System Architecture in Mediterranean Durum Wheat Landraces by Genome-Wide Association Study. Agronomy 2019, 9, 364. [Google Scholar] [CrossRef] [Green Version]
- Boudiar, R.; González, J.M.; Mekhlouf, A.; Casas, A.M.; Igartua, E. Durum Wheat Seminal Root Traits within Modern and Landrace Germplasm in Algeria. Agronomy 2020, 10, 713. [Google Scholar] [CrossRef]
- Uga, Y.; Sugimoto, K.; Ogawa, S.; Rane, J.; Ishitani, M.; Hará, N.; Kitomi, Y.; Inukai, Y.; Ono, K.; Kanno, N. Control of root system architecture by DEEPER ROOTING 1 increases rice yield under drought conditions. Nat. Genet. 2013, 45, 1097–1102. [Google Scholar] [CrossRef]
- Uga, Y.; Kitomi, Y.; Ishikawa, S.; Yano, M. Genetic improvement for root growth angle to enhance crop production. Breed. Sci. 2015, 65, 111–119. [Google Scholar] [CrossRef] [Green Version]
- Uga, Y.; Kitomi, Y.; Yamamoto, E.; Kanno, N.; Kawai, S.; Mizubayashi, T.; Fukuoka, S. A QTL for root growth angle on rice chromosome 7 is involved in the genetic pathway of DEEPER ROOTING 1. Rice 2015, 8, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Guseman, J.M.; Webb, K.; Srinivasan, C.; Dardick, C. DRO1 influences root system architecture in Arabidopsis and Prunus species. Plant J. Cell Mol. Biol. 2017, 89, 1093–1105. [Google Scholar] [CrossRef] [Green Version]
- Kitomi, Y.; Hanzawa, E.; Kuya, N.; Inoue, H.; Hará, N.; Kawai, S.; Kanno, N.; Endo, M.; Sugimoto, K.; Yamazaki, T.; et al. Root angle modifications by the DRO1 homolog improve rice yields in saline paddy fields. Proc. Natl. Acad. Sci. USA 2020, 117, 21242–21250. [Google Scholar] [CrossRef]
- Loarce, Y.; Cabeza, A.; Cañas, R.; González, J.M. Identification and analysis of TtDro1A and TtDro1B genes of Triticum turgidum subsp. durum and turgidum. Expression study to reveal their role in seedling root angle. BMC Plant Biol. 2021. under review. [Google Scholar]
- Yaakov, B.; Kashkush, K. Mobilization of Stowaway-like MITEs in newly formed allohexaploid wheat species. Plant Mol. Biol. 2012, 80, 419–427. [Google Scholar] [CrossRef] [PubMed]
- Yaakov, B.; Ceylan, E.; Domb, K.; Kashkush, K. Marker utility of miniature inverted-repeat transposable elements for wheat biodiversity and evolution. Theor. Appl. Genet. 2012, 124, 1365–1373. [Google Scholar] [CrossRef]
- Shen, J.; Liu, J.; Xie, K.; Xing, F.; Xiong, F.; Xiao, J.; Li, X.; Xiong, L. Translational repression by a miniature inverted-repeat transposable element in the 3′ untranslated region. Nat. Commun. 2017, 8, 14651. [Google Scholar] [CrossRef] [Green Version]
- Sahebi, M.; Hanafi, M.M.; van Wijnen, A.J.; Rice, D.; Rafii, M.Y.; Azizi, P.; Osman, M.; Taheri, S.; Abu Bakar, M.F.; Isa, M.N.M.; et al. Contribution of transposable elements in the plant’s genome. Gene 2018, 665, 155–166. [Google Scholar] [CrossRef] [PubMed]
- Feldman, M.; Levy, A.A. Genome evolution in allopolyploid wheat—A revolutionary reprogramming followed by gradual changes. J. Genet. Genom. 2009, 36, 511. [Google Scholar] [CrossRef]
- Muterko, A.; Salina, E. Divergence of VRN-B3 alleles during the evolution of domesticated wheat. Mol. Genet. Genom. 2019, 294, 263–275. [Google Scholar] [CrossRef]
- Sahri, A.; Chentoufi, L.; Arbaoui, M.; Ardisson, M.; Belqadi, L.; Birouk, A.; Roumet, P.; Muller, M.H. Towards a comprehensive characterization of durum wheat landraces in Moroccan traditional agrosystems: Analysing genetic diversity in the light of geography, farmers’ taxonomy and tetraploid wheat domestication history. BMC Evol. Biol. 2014, 14, 264. [Google Scholar] [CrossRef] [Green Version]
- De Oliveira, R.; Rimbert, H.; Balfourier, F.; Kitt, J.; Dynomant, E.; Vrána, J.; Doležel, J.; Cattonaro, F.; Paux, E.; Choulet, F. Structural Variations Affecting Genes and Transposable Elements of Chromosome 3B in Wheats. Front. Genet. 2020, 11, 891. [Google Scholar] [CrossRef]
- Özkan, H.; Willcox, G.; Graner, A.; Graner, A.; Salamini, F.; Kilian, B. Geographic distribution and domestication of wild emmer wheat (Triticum dicoccoides). Genet. Resour. Crop Evol. 2011, 58, 11–53. [Google Scholar] [CrossRef]
- Zaharieva, M.; Ayana, N.G.; Hakimi, A.A.; Misra, S.C.; Monneveux, P. Cultivated emmer wheat (Triticum dicoccon Schrank), an old crop with promising future: A review. Genet. Resour. Crop Evol. 2010, 57, 937. [Google Scholar] [CrossRef]
- Lucas, S.J.; Salantur, A.; Yazar, S.; Budak, H. High-throughput SNP genotyping of modern and wild emmer wheat for yield and root morphology using a combined association and linkage analysis. Funct. Integr. Genom. 2017, 17, 667–685. [Google Scholar] [CrossRef] [PubMed]
- Mohammadi, M.; Mirlohi, A.; Majidi, M.M.; Soleimani Kartalaei, E. Emmer wheat as a source for trait improvement in durum wheat: A study of general and specific combining ability. Euphytica 2021, 217, 64. [Google Scholar] [CrossRef]
- Van Hintum, T.J.L.; Brown, A.H.D.; Spillane, C.; Hodgkin, T. Core Collections of Plant Genetic Resources; IPGRI Technical Bulletin No. 3; International Plant Genetic Resources Institute: Rome, Italy, 2000. [Google Scholar]
- MacKey, J. Species relationship in Triticum. Hereditas 1966, 2, 237–276. [Google Scholar]
- Pascual, L.; Ruiz, M.; López-Fernández, M.; Pérez-Peña, H.; Benavente, E.; Vázquez, J.F.; Sansaloni, C.; Giraldo, P. Genomic analysis of Spanish wheat landraces reveals their variability and potential for breeding. BMC Genom. 2020, 21, 122. [Google Scholar] [CrossRef] [Green Version]
- Wiwart, M.; Suchowilska, E.; Kandler, W.; Sulyok, M.; Groenwald, P.; Krska, R. Can Polish wheat (Triticum polonicum L.) be an interesting gene source for breeding wheat cultivars with increased resistance to Fusarium head blight? Genet. Resour. Crop Evol. 2013, 60, 2359–2373. [Google Scholar] [CrossRef]
- Bieńkowska, T.; Suchowilska, E.; Kandler, W.; Krska, R.; Wiwart, M. Triticum polonicum L. as potential source material for the biofortification of wheat with essential micronutrients. Plant Genet. Resour. Charact. Util. 2019, 17, 213–220. [Google Scholar] [CrossRef]
- Wicker, T.; Gundlach, H.; Spannagl, M.; Uauy, C.; Borrill, P.; Ramírez-González, R.H.; De Oliveira, R.; Mayer, K.F.; Paux, E.; Choulet, F. Impact of transposable elements on genome structure and evolution in bread wheat. Genome Biol. 2018, 19, 103. [Google Scholar] [CrossRef]
- Matsuoka, Y.; Takumi, S.; Nasuda, S. Genetic Mechanisms of Allopolyploid Speciation Through Hybrid Genome Doubling. Int. Rev. Cell Mol. Biol. 2014, 309, 199–258. [Google Scholar]
- He, F.; Pasam, R.; Shi, F.; Kant, S.; Keeble-Gagnere, G.; Kay, P.; Forrest, K.; Fritz, A.; Hucl, P.; Wiebe, K.; et al. Exome sequencing highlights the role of wild-relative introgression in shaping the adaptive landscape of the wheat genome. Nat. Genet. 2019, 51, 896–904. [Google Scholar] [CrossRef] [PubMed]
- Walkowiak, S.; Gao, L.; Monat, C.; Haberer, G.; Kassa, M.T.; Brinton, J.; Ramirez-Gonzalez, R.H.; Kolodziej, M.C.; Delorean, E.; Thambugala, D.; et al. Multiple wheat genomes reveal global variation in modern breeding. Nature 2020, 588, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Hao, M.; Luo, J.; Zhang, L.; Yuan, Z.; Zheng, Y.; Zhang, H.; Liu, D. In situ hybridization analysis indicates that 4AL–5AL–7BS translocation preceded subspecies differentiation of Triticum turgidum. Genome 2013, 56, 303–305. [Google Scholar] [CrossRef] [PubMed]
- Devos, K.M.; Dubcovsky, J.; Dvorak, J.; Chinoy, C.N.; Gale, M.D. Structural evolution of wheat chromosomes 4A, 5A, and 7B and its impact on recombination. Theor. Appl. Genet. 1995, 91, 282–288. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feldman, M.; Levy, A.A. Allopolyploidy—A shaping force in the evolution of wheat genomes. Cytogenet. Genome Res. 2005, 109, 250–258. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.; Gao, Y.; Lynch, J.P. Large Crown Root Number Improves Topsoil Foraging and Phosphorus Acquisition. Plant Physiol. 2018, 177, 90. [Google Scholar] [CrossRef] [Green Version]
- Czajkowska, B.I.; Oliveira, H.R.; Brown, T.A. A discriminatory test for the wheat B and G genomes reveals misclassified accessions of Triticum timopheevii and Triticum turgidum. PLoS ONE 2019, 14, e0215175. [Google Scholar] [CrossRef] [PubMed]
| Subspecies | Variable | Mean | SD | CV | Min | Max |
|---|---|---|---|---|---|---|
| dicoccoides (n = 19) | TRL | 58.8 | 10.41 | 17.68 | 42.53 | 75.34 |
| S | 9.70 | 2.06 | 21.22 | 5.96 | 13.19 | |
| V | 0.13 | 0.03 | 25.09 | 0.07 | 0.19 | |
| D | 0.05 | 0.00 | 6.89 | 0.04 | 0.06 | |
| PRL | 20.6 | 4.03 | 19.51 | 12.64 | 27.84 | |
| NR | 3.91 | 0.67 | 17.18 | 3.00 | 5.33 | |
| MRA | 20.8 | 5.59 | 26.83 | 12.35 | 29.57 | |
| MAV | 9.07 | 4.00 | 44.10 | 2.30 | 17.53 | |
| MxAV | 32.7 | 8.72 | 26.60 | 20.75 | 51.50 | |
| dicoccum (n = 14) | TRL | 72.3 | 12.08 | 16.71 | 51.53 | 91.31 |
| S | 12.3 | 2.43 | 19.61 | 9.35 | 16.57 | |
| V | 0.18 | 0.04 | 23.50 | 0.13 | 0.25 | |
| D | 0.05 | 0.00 | 6.32 | 0.05 | 0.06 | |
| PRL | 20.0 | 2.17 | 10.80 | 16.99 | 23.22 | |
| NR | 4.83 | 0.31 | 6.37 | 4.08 | 5.42 | |
| MRA | 33.1 | 7.50 | 22.60 | 20.21 | 43.23 | |
| MAV | 18.2 | 6.12 | 33.49 | 11.22 | 32.83 | |
| MxAV | 47.8 | 8.93 | 18.68 | 31.83 | 58.75 | |
| turgidum (n = 37) | TRL | 83.8 | 10.46 | 12.47 | 53.55 | 101.63 |
| S | 14.5 | 1.87 | 12.88 | 10.08 | 18.20 | |
| V | 0.21 | 0.03 | 14.50 | 0.14 | 0.27 | |
| D | 0.06 | 0.00 | 7.55 | 0.05 | 0.07 | |
| PRL | 21.5 | 2.17 | 10.08 | 15.70 | 25.29 | |
| NR | 5.02 | 0.25 | 5.05 | 4.08 | 5.50 | |
| MRA | 27.1 | 6.44 | 23.76 | 13.78 | 42.76 | |
| MAV | 13.1 | 4.23 | 32.22 | 3.83 | 21.17 | |
| MxAV | 43.1 | 7.68 | 17.83 | 30.14 | 64.33 | |
| polonicum (n = 10) | TRL | 86.3 | 10.98 | 12.72 | 75.24 | 108.15 |
| S | 14.4 | 1.59 | 11.01 | 12.97 | 18.29 | |
| V | 0.20 | 0.02 | 11.55 | 0.17 | 0.26 | |
| D | 0.05 | 0.00 | 4.77 | 0.05 | 0.06 | |
| PRL | 21.7 | 3.10 | 14.29 | 16.15 | 26.36 | |
| NR | 5.23 | 0.36 | 6.95 | 4.75 | 6.00 | |
| MRA | 19.0 | 5.53 | 28.99 | 11.78 | 30.57 | |
| MAV | 9.48 | 6.21 | 65.53 | 1.75 | 20.40 | |
| MxAV | 31.2 | 7.46 | 23.85 | 20.00 | 46.63 | |
| durum (n = 143) | TRL | 91.9 | 10.98 | 11.94 | 47.35 | 115.89 |
| S | 15.5 | 1.99 | 12.85 | 8.69 | 20.43 | |
| V | 0.21 | 0.03 | 15.22 | 0.11 | 0.30 | |
| D | 0.05 | 0.00 | 5.32 | 0.04 | 0.06 | |
| PRL | 22.3 | 2.33 | 10.40 | 12.77 | 28.05 | |
| NR | 5.34 | 0.40 | 7.41 | 4.50 | 6.58 | |
| MRA | 23.1 | 5.25 | 22.69 | 11.78 | 37.27 | |
| MAV | 10.8 | 3.87 | 35.86 | 4.25 | 24.92 | |
| MxAV | 37.0 | 7.21 | 19.47 | 18.83 | 58.12 |
| MITE | Variable | Mean | SD | CV | Min | Max |
|---|---|---|---|---|---|---|
| Without (n = 170) | TRL | 88.95 | 13.66 | 15.35 | 42.53 | 115.8 |
| S | 14.98 | 2.47 | 16.51 | 5.96 | 20.43 | |
| V | 20.69 | 3.87 | 18.71 | 6.87 | 30.15 | |
| D | 52.97 | 2.97 | 5.60 | 42.14 | 60.49 | |
| PRL | 22.17 | 2.63 | 11.87 | 12.64 | 28.05 | |
| NR | 5.22 | 5.40 | 10.34 | 30.00 | 65.83 | |
| MRA | 23.14 | 5.63 | 24.33 | 11.78 | 39.99 | |
| MAV | 10.91 | 4.13 | 37.84 | 1.75 | 24.92 | |
| MxAV | 36.88 | 7.82 | 21.21 | 18.83 | 58.12 | |
| With (n = 53) | TRL | 77.89 | 14.78 | 18.97 | 43.03 | 112.3 |
| S | 13.35 | 2.65 | 19.84 | 6.99 | 18.20 | |
| V | 18.86 | 4.08 | 21.64 | 8.62 | 26.92 | |
| D | 54.70 | 4.01 | 7.33 | 48.23 | 72.34 | |
| PRL | 21.05 | 2.28 | 10.82 | 15.70 | 25.71 | |
| NR | 4.83 | 5.58 | 11.54 | 30.00 | 56.67 | |
| MRA | 27.01 | 7.72 | 28.58 | 12.35 | 43.23 | |
| MAV | 13.16 | 5.84 | 44.35 | 2.30 | 32.83 | |
| MxAV | 42.01 | 9.23 | 21.97 | 20.75 | 64.33 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
González, J.M.; Cañas, R.; Cabeza, A.; Ruiz, M.; Giraldo, P.; Loarce, Y. Study of Variability in Root System Architecture of Spanish Triticum turgidum L. Subspecies and Analysis of the Presence of a MITE Element Inserted in the TtDro1B Gene: Evolutionary Implications. Agronomy 2021, 11, 2294. https://doi.org/10.3390/agronomy11112294
González JM, Cañas R, Cabeza A, Ruiz M, Giraldo P, Loarce Y. Study of Variability in Root System Architecture of Spanish Triticum turgidum L. Subspecies and Analysis of the Presence of a MITE Element Inserted in the TtDro1B Gene: Evolutionary Implications. Agronomy. 2021; 11(11):2294. https://doi.org/10.3390/agronomy11112294
Chicago/Turabian StyleGonzález, Juan M., Rodrigo Cañas, Alejandra Cabeza, Magdalena Ruiz, Patricia Giraldo, and Yolanda Loarce. 2021. "Study of Variability in Root System Architecture of Spanish Triticum turgidum L. Subspecies and Analysis of the Presence of a MITE Element Inserted in the TtDro1B Gene: Evolutionary Implications" Agronomy 11, no. 11: 2294. https://doi.org/10.3390/agronomy11112294
APA StyleGonzález, J. M., Cañas, R., Cabeza, A., Ruiz, M., Giraldo, P., & Loarce, Y. (2021). Study of Variability in Root System Architecture of Spanish Triticum turgidum L. Subspecies and Analysis of the Presence of a MITE Element Inserted in the TtDro1B Gene: Evolutionary Implications. Agronomy, 11(11), 2294. https://doi.org/10.3390/agronomy11112294







