Understanding the Impact of Drought in Coffea Genotypes: Transcriptomic Analysis Supports a Common High Resilience to Moderate Water Deficit but a Genotype Dependent Sensitivity to Severe Water Deficit
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material and Growth Conditions
2.2. Water Stress Imposition and Leaf Water Status
2.3. RNA Extraction and Illumina Sequencing
2.4. Quality Analysis of Sequencing Data
2.5. Reference-Based Mapping and Assembly
2.6. Identification of Differentially Expressed Genes (DEGs)
2.7. Regulation Patterns of Transcription Factors
2.8. Enrichment Analysis of Gene Ontology
3. Results
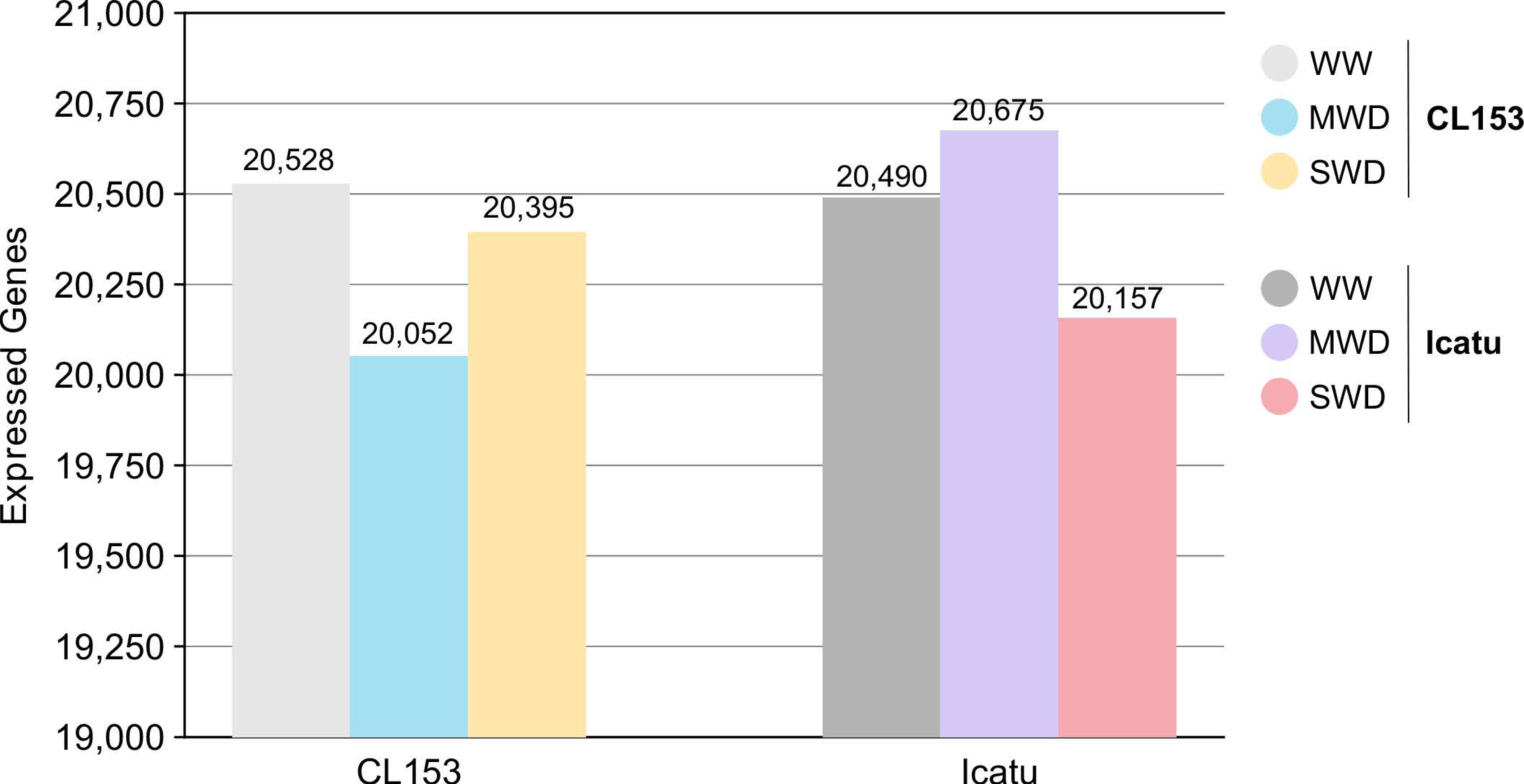
3.1. Overall Transcriptome Profiling and Mapping Statistics
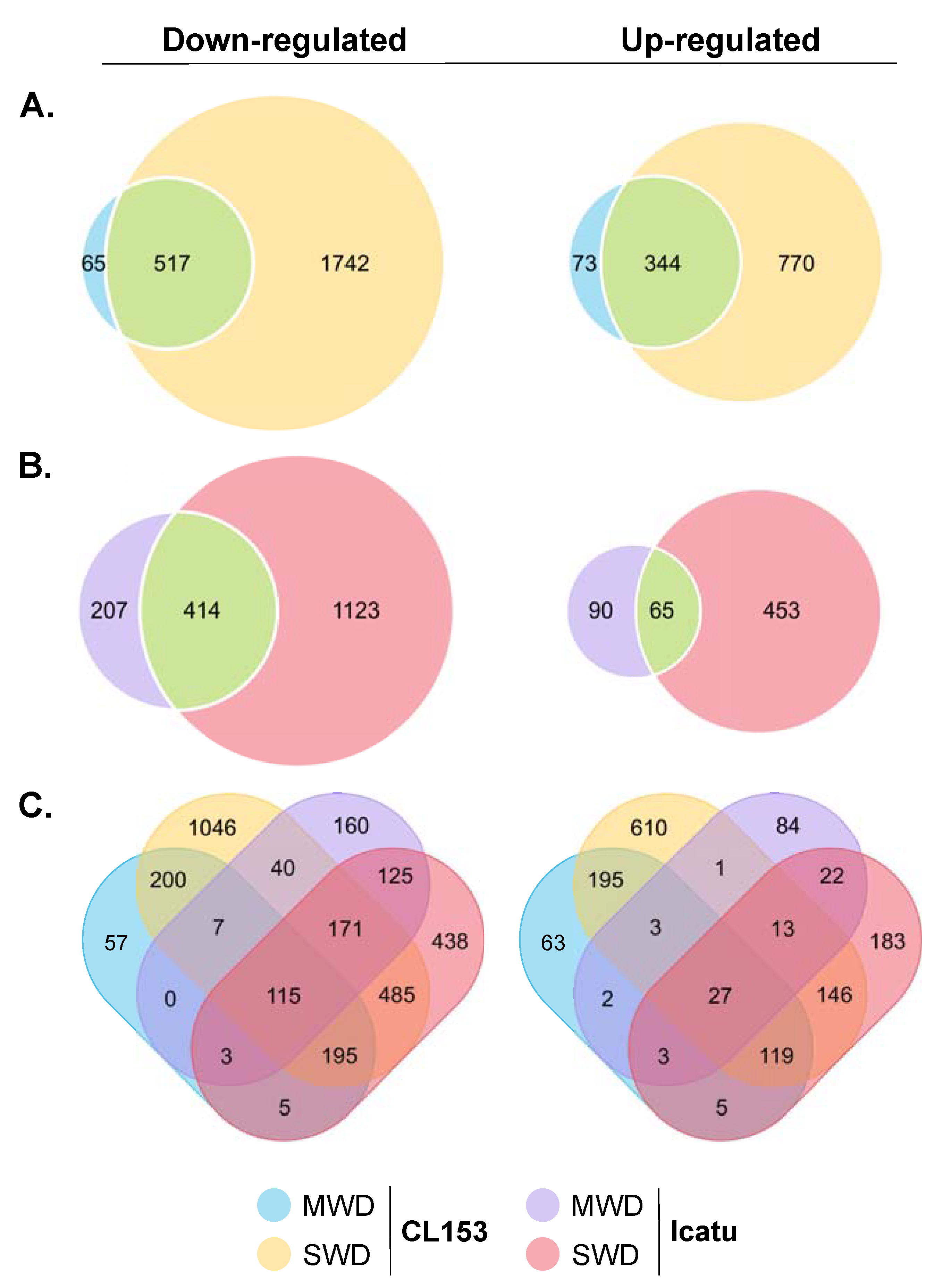
3.2. Differential Gene Changes of CL153 and Icatu in Response to Drought
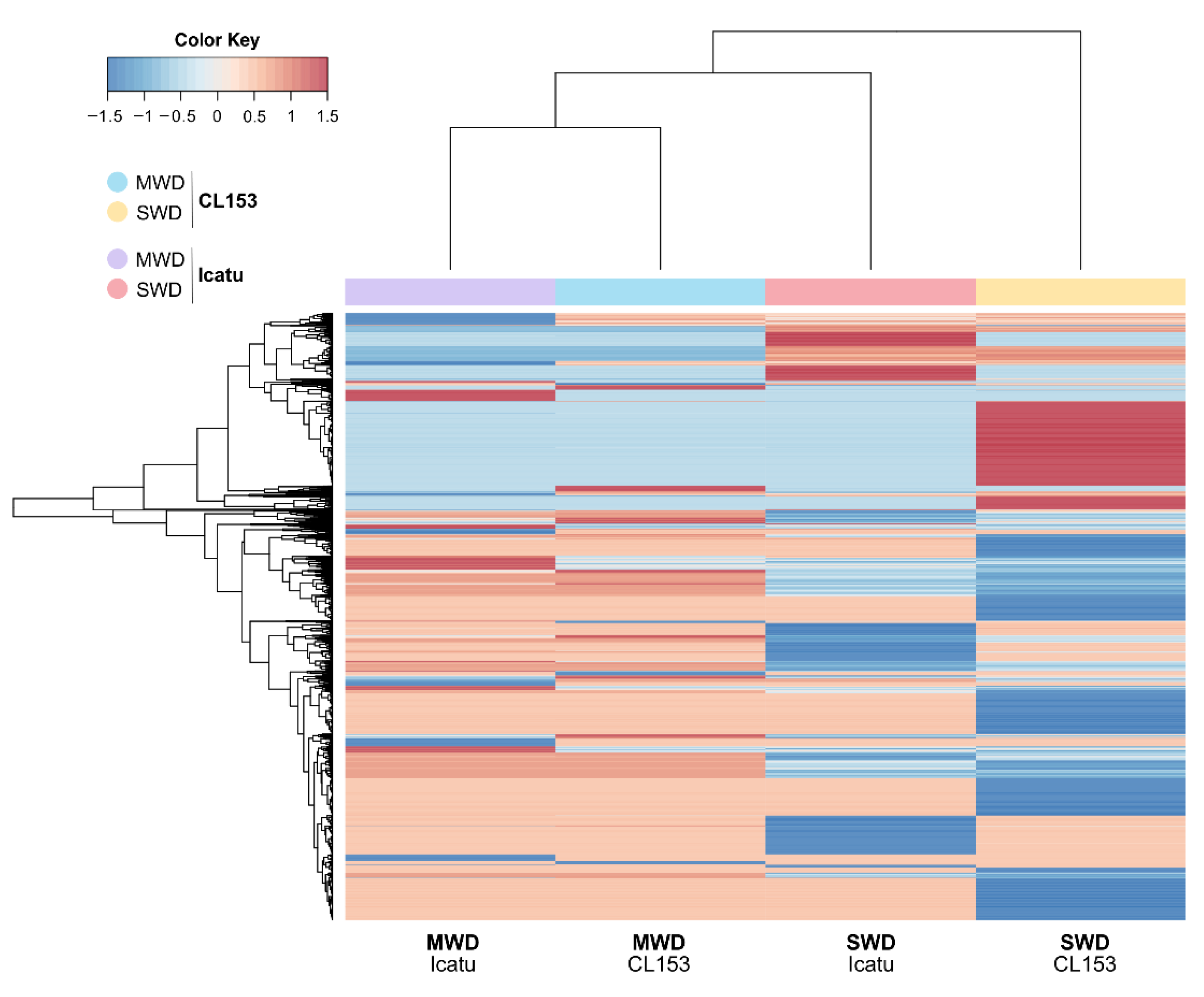
3.3. Identification and Classification of DEGs
3.4. Regulation Patterns of DEGs Directly Linked to Water Deprivation and Desiccation
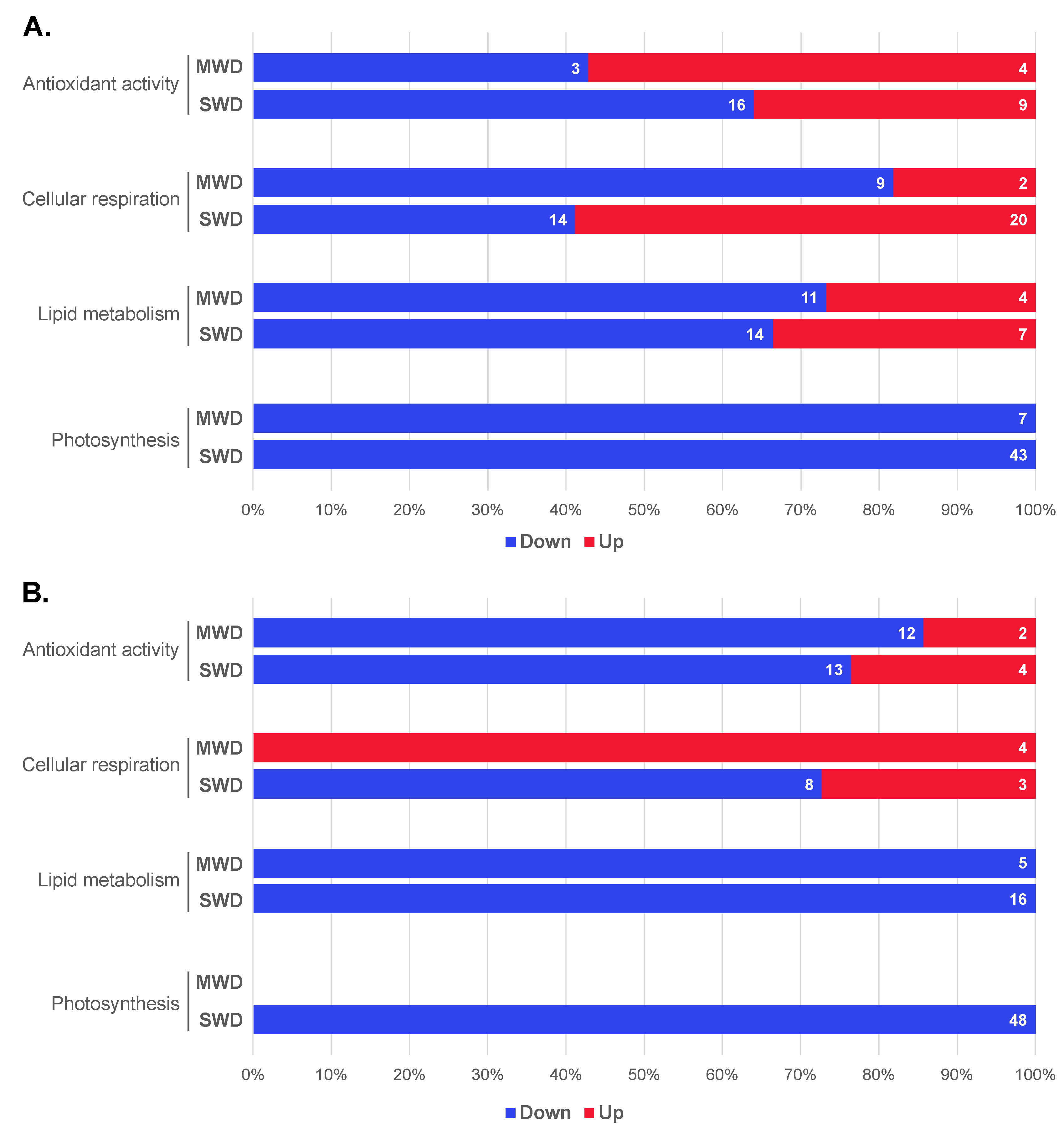
3.5. Regulation Patterns of Photosynthetic and Other Biochemical Coffee Related DEGs
3.6. Regulation Patterns of Transcription Factors among Responsiveness DEGs
3.7. Regulation Patterns of Phosphatases and Protein Kinases DEGs
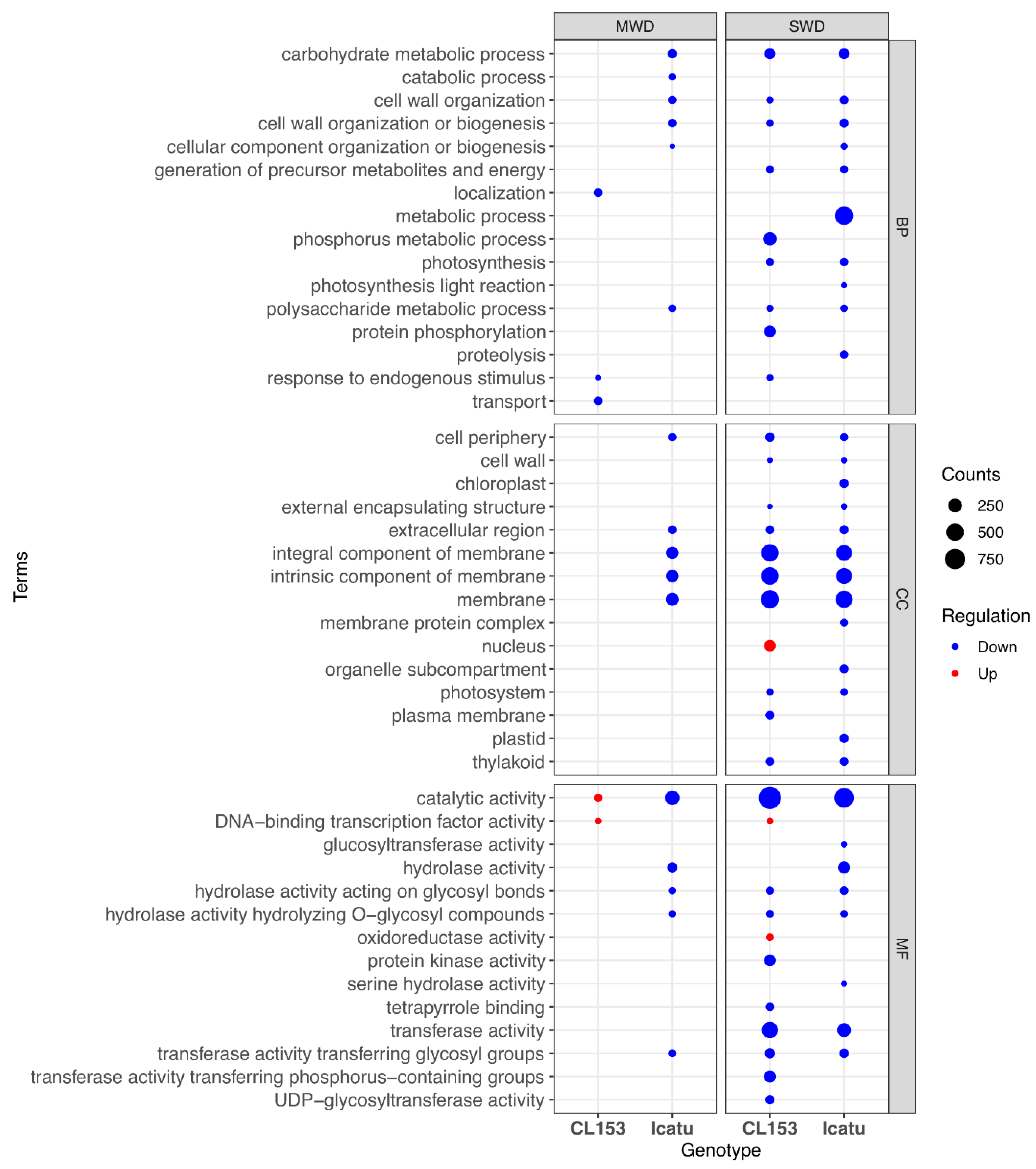
3.8. Enriched GO Terms of Drought-Related DEGs
4. Discussion
4.1. Impacts of Water Deficit on the Transcriptomic Profile of Coffea canephora (CL153) and C. arabica (Icatu)
4.2. Role of Aspartic Proteases and Protectant Proteins in Water Deprivation and Desiccation in Coffee
4.3. Drought-Responsiveness Transcription Factors
4.4. DEGs Involved in Phosphatases and Protein Kinases Affected by Drought
4.5. Coping with Drought: Lessons from Crossed Transcriptomic, Physiological, and Biochemical Studies
5. Conclusions
- Even though drought had an impact on the leaf transcriptome of both coffee genotypes, our results revealed that both genotypes are more drought-resistant than other coffee cultivars.
- Drought triggered a specific response associated with the magnitude of water deficit, which was also genotype-dependent since few DEGs and pathways were common to treatments and both genotypes. By comparison, MWD only had a minor effect on the transcripts of both genotypes.
- There was a predominance of protective genes (more in CL153) associated with antioxidant activities, including genes involved in water deprivation and desiccation, such as Lea and aspartic proteases.
- A significant number of TFs, including ERF, DREB, and the leucine zipper, were found to be significantly up-regulated under drought. Together with the large number of phosphatases and protein kinases we found, these results suggest the involvement of ABA signaling in the drought tolerance of these genotypes.
- Coupled with the previous physiological and metabolic results, our study provides novel and timely information showing several layers of response and suggesting the existence of post-transcriptional regulations in the two coffee genotypes, which should be further investigated.
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fahad, S.; Bajwa, A.A.; Nazir, U.; Anjum, S.A.; Farooq, A.; Zohaib, A.; Sadia, S.; Nasim, W.; Adkins, S.; Saud, S.; et al. Crop production under drought and heat stress: Plant responses and management options. Front. Plant Sci. 2017, 8, 1147. [Google Scholar] [CrossRef] [Green Version]
- Lamaoui, M.; Jemo, M.; Datla, R.; Bekkaoui, F. Heat and drought stresses in crops and approaches for their mitigation. Front. Chem. 2018, 6, 26. [Google Scholar] [CrossRef]
- Lang, Y.; Wang, M.; Xia, J.; Zhao, Q. Effects of soil drought stress on photosynthetic gas exchange traits and chlorophyll fluorescence in Forsythia suspensa. J. For. Res. 2018, 29, 45–53. [Google Scholar] [CrossRef]
- Chaves, M.M.; Maroco, J.P.; Pereira, J.S. Understanding plant responses to drought—From genes to the whole plant. Funct. Plant Biol. 2003, 30, 239–264. [Google Scholar] [CrossRef]
- Khan, A.; Pan, X.; Najeeb, U.; Tan, D.K.Y.; Fahad, S.; Zahoor, R.; Luo, H. Coping with drought: Stress and adaptive mechanisms, and management through cultural and molecular alternatives in cotton as vital constituents for plant stress resilience and fitness. Biol. Res. 2018, 51, 47. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Ghany, S.E.; Ullah, F.; Ben-Hur, A.; Reddy, A.S.N. Transcriptome analysis of drought-resistant and drought-sensitive sorghum (Sorghum bicolor) genotypes in response to peg-induced drought stress. Int. J. Mol. Sci. 2020, 21, 772. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iquebal, M.A.; Sharma, P.; Jasrotia, R.S.; Jaiswal, S.; Kaur, A.; Saroha, M.; Angadi, U.B.; Sheoran, S.; Singh, R.; Singh, G.P.; et al. RNAseq analysis reveals drought-responsive molecular pathways with candidate genes and putative molecular markers in root tissue of wheat. Sci. Rep. 2019, 9, 1–18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mashaki, K.M.; Garg, V.; Nasrollahnezhad Ghomi, A.A.; Kudapa, H.; Chitikineni, A.; Nezhad, K.Z.; Yamchi, A.; Soltanloo, H.; Varshney, R.K.; Thudi, M. RNA-Seq analysis revealed genes associated with drought stress response in kabuli chickpea (Cicer arietinum L.). PLoS ONE 2018, 13, e0199774. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jarzyniak, K.M.; Jasiński, M. Membrane transporters and drought resistance—A complex issue. Front. Plant Sci. 2014, 5, 687. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arisha, M.H.; Ahmad, M.Q.; Tang, W.; Liu, Y.; Yan, H.; Kou, M.; Wang, X.; Zhang, Y.; Li, Q. RNA-sequencing analysis revealed genes associated drought stress responses of different durations in hexaploid sweet potato. Sci. Rep. 2020, 10, 12573. [Google Scholar] [CrossRef]
- Cochetel, N.; Ghan, R.; Toups, H.S.; Degu, A.; Tillett, R.L.; Schlauch, K.A.; Cramer, G.R. Drought tolerance of the grapevine, Vitis champinii cv. Ramsey, is associated with higher photosynthesis and greater transcriptomic responsiveness of abscisic acid biosynthesis and signaling. BMC Plant Biol. 2020, 20, 1–25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meng, H.L.; Zhang, W.; Zhang, G.H.; Wang, J.J.; Meng, Z.G.; Long, G.Q.; Yang, S.C. Unigene-based RNA-seq provides insights on drought stress responses in Marsdenia tenacissima. PLoS ONE 2018, 13, e0202848. [Google Scholar] [CrossRef] [Green Version]
- Agarwal, P.K.; Gupta, K.; Lopato, S.; Agarwal, P. Dehydration responsive element binding transcription factors and their applications for the engineering of stress tolerance. J. Exp. Bot. 2017, 68, 2135–2148. [Google Scholar] [CrossRef] [PubMed]
- Chai, M.; Cheng, H.; Yan, M.; Priyadarshani, S.V.G.N.; Zhang, M.; He, Q.; Huang, Y.; Chen, F.; Liu, L.; Huang, X.; et al. Identification and expression analysis of the DREB transcription factor family in pineapple (Ananas comosus (L.) Merr.). Peer J 2020, 8, e9006. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.L.; Zhang, M.; Yan, X.; Bian, Y.W.; Zhen, S.M.; Yan, Y.M. Dynamic phosphoproteome analysis of seedling leaves in Brachypodium distachyon L. Reveals central phosphorylated proteins involved in the drought stress response. Sci. Rep. 2016, 6, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Hoang, X.L.T.; Nhi, D.N.H.; Thu, N.B.A.; Thao, N.P.; Tran, L.-S.P. Transcription factors and their roles in signal transduction in plants under abiotic stresses. Curr. Genom. 2017, 18, 483. [Google Scholar] [CrossRef] [PubMed]
- Hrmova, M.; Hussain, S.S. Plant transcription factors involved in drought and associated stresses. Int. J. Mol. Sci. 2021, 22, 5662. [Google Scholar] [CrossRef] [PubMed]
- DaMatta, F.M.; Rahn, E.; Läderach, P.; Ghini, R.; Ramalho, J.C. Why could the coffee crop endure climate change and global warming to a greater extent than previously estimated? Clim. Chang. 2019, 152, 167–178. [Google Scholar] [CrossRef] [Green Version]
- Semedo, J.N.; Rodrigues, W.P.; Dubberstein, D.; Martins, M.Q.; Martins, L.D.; Pais, I.P.; Rodrigues, A.P.; Leitão, A.E.; Partelli, F.L.; Campostrini, E.; et al. Coffee responses to drought, warming and high [CO2] in a context of future climate change scenarios. In Climate Change Management; Springer: Berlin/Heidelberg, Germany, 2018; pp. 465–477. [Google Scholar] [CrossRef]
- Davis, A.P.; Chadburn, H.; Moat, J.; O’Sullivan, R.; Hargreaves, S.; Nic Lughadha, E. High extinction risk for wild coffee species and implications for coffee sector sustainability. Sci. Adv. 2019, 5, eaav3473. [Google Scholar] [CrossRef] [Green Version]
- Van der Vossen, H.; Bertrand, B.; Charrier, A. Next generation variety development for sustainable production of arabica coffee (Coffea arabica L.): A review. Euphytica 2015, 204, 243–256. [Google Scholar] [CrossRef]
- Ramalho, J.C.; DaMatta, F.M.; Rodrigues, A.P.; Scotti-Campos, P.; Pais, I.; Batista-Santos, P.; Partelli, F.L.; Ribeiro, A.; Lidon, F.C.; Leitão, A.E. Cold impact and acclimation response of Coffea spp. plants. Theor. Exp. Plant Physiol. 2014, 26, 5–18. [Google Scholar] [CrossRef]
- Rodrigues, W.P.; Martins, M.Q.; Fortunato, A.S.; Rodrigues, A.P.; Semedo, J.N.; Simoes-Costa, M.C.; Pais, I.P.; Leitão, A.E.; Colwell, F.; Goulão, L.; et al. Long-term elevated air [CO2] strengthens photosynthetic functioning and mitigates the impact of supra-optimal temperatures in tropical Coffea arabica and Coffea canephora species. Glob. Chang. Biol. 2016, 22, 415–431. [Google Scholar] [CrossRef]
- Dubberstein, D.; Lidon, F.C.; Rodrigues, A.P.; Semedo, J.N.; Marques, I.; Rodrigues, W.P.; Gouveia, D.; Armengaud, J.; Semedo, M.C.; Martins, S.; et al. Resilient and sensitive key points of the photosynthetic machinery of Coffea spp. to the single and superimposed exposure to severe drought and heat stresses. Front. Plant Sci. 2020, 11, 1049. [Google Scholar] [CrossRef] [PubMed]
- Semedo, J.N.; Rodrigues, A.P.; Lidon, F.C.; Pais, I.P.; Marques, I.; Gouveia, D.; Armengaud, J.; Silva, M.J.; Martins, S.; Semedo, M.C.; et al. Intrinsic non-stomatal resilience to drought of the photosynthetic apparatus in Coffea spp. is strengthened by elevated air [CO2]. Tree Physiol. 2021, 41, 708–727. [Google Scholar] [CrossRef] [PubMed]
- DaMatta, F.M.; Ramalho, J.D.C. Impacts of drought and temperature stress on coffee physiology and production: A review. Braz. J. Plant Physiol. 2006, 18, 55–81. [Google Scholar] [CrossRef]
- Ramalho, J.C.; Pais, I.P.; Leitão, A.E.; Guerra, M.; Reboredo, F.H.; Máguas, C.M.; Carvalho, M.L.; Scotti-Campos, P.; Ribeiro-Barros, A.I.; Lidon, F.J.C.; et al. Can elevated air [CO2] conditions mitigate the predicted warming impact on the quality of coffee bean? Front. Plant Sci. 2018, 9, 287. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haggar, J.; Schepp, K. NRI Working Paper Series: Climate Change, Agriculture and Natural Resources Natural Resources Institute Coffee and Climate Change Impacts and options for adaption in Brazil, Guatemala, Tanzania and Vietnam Coffee and Climate Change Impacts and options for adaption in Brazil, 2012, Guatemala, Tanzania and Vietnam. Available online: www.nri.org (accessed on 20 September 2021).
- De Freitas Guedes, F.A.; Nobres, P.; Rodrigues Ferreira, D.C.; Menezes-Silva, P.E.; Ribeiro-Alves, M.; Correa, R.L.; DaMatta, F.M.; Alves-Ferreira, M. Transcriptional memory contributes to drought tolerance in coffee (Coffea canephora) plants. Environ. Exp. Bot. 2018, 147, 220–233. [Google Scholar] [CrossRef]
- Kath, J.; Byrareddy, V.M.; Craparo, A.; Nguyen-Huy, T.; Mushtaq, S.; Cao, L.; Bossolasco, L. Not so robust: Robusta coffee production is highly sensitive to temperature. Glob. Chang. Biol. 2020, 26, 3677–3688. [Google Scholar] [CrossRef]
- Venancio, L.P.; Filgueiras, R.; Mantovani, E.C.; do Amaral, C.H.; da Cunha, F.F.; dos Santos Silva, F.C.; Althoff, D.; dos Santos, R.A.; Cavatte, P.C. Impact of drought associated with high temperatures on Coffea canephora plantations: A case study in Espírito Santo State, Brazil. Sci. Rep. 2020, 10, 19719. [Google Scholar] [CrossRef]
- Ramalho, J.C.; Fortunato, A.S.; Goulao, L.F.; Lidon, F.C. Cold-induced changes in mineral content in leaves of Coffea spp. Identification of descriptors for tolerance assessment. Biol. Plant. 2013, 57, 495–506. [Google Scholar] [CrossRef]
- Ramalho, J.C.; Rodrigues, A.P.; Semedo, J.N.; Pais, I.P.; Martins, L.D.; Simões-Costa, M.C.; Leitão, A.E.; Fortunato, A.S.; Batista-Santos, P.; Palos, I.M.; et al. Sustained photosynthetic performance of Coffea spp. under long-term enhanced [CO2]. PLoS ONE 2013, 8, 82712. [Google Scholar] [CrossRef] [Green Version]
- Marques, I.; Fernandes, I.; Paulo, O.S.; Lidon, F.C.; DaMatta, F.M.; Ramalho, J.C.; Ribeiro-Barros, A.I. A transcriptomic approach to understanding the combined impacts of supra-optimal temperatures and CO2 revealed different responses in the polyploid Coffea arabica and its diploid progenitor C. canephora. Int. J. Mol. Sci. 2021, 22, 3125. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 20 May 2020).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Wingett, S.W.; Andrews, S. FastQ Screen: A tool for multi-genome mapping and quality control. F1000Research 2018, 7, 1338. [Google Scholar] [CrossRef]
- Denoeud, F.; Carretero-Paulet, L.; Dereeper, A.; Droc, G.; Guyot, R.; Pietrella, M.; Zheng, C.; Alberti, A.; Anthony, F.; Aprea, G.; et al. The coffee genome provides insight into the convergent evolution of caffeine biosynthesis. Science 2014, 345, 1181–1184. [Google Scholar] [CrossRef] [Green Version]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq—A Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The sequence alignment/map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pertea, G.G. gffread: GFF/GTF Utility Providing Format Conversions, Region Filtering, FASTA Sequence Extraction and More. 2015. Available online: https://github.com/gpertea/gffread (accessed on 15 January 2021).
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016; ISBN 978-3-319-24277-4. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Benjamini, Y.; Hochberg, Y. On the Adaptive Control of the False Discovery Rate in Multiple Testing with Independent Statistics. J. Edu. Behav. Stat. 2016, 25, 60–83. [Google Scholar] [CrossRef] [Green Version]
- Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- Martins, M.Q.; Rodrigues, W.P.; Fortunato, A.S.; Leitão, A.E.; Rodrigues, A.P.; Pais, I.P.; Martins, L.D.; Silva, M.J.; Reboredo, F.H.; Partelli, F.L.; et al. Protective response mechanisms to heat stress in interaction with high [CO2] conditions in Coffea spp. Front. Plant Sci. 2016, 7, 947. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.; Huang, H.; Wu, C.H. Protein bioinformatics databases and resources. Meth. Mol. Biol. 2017, 1558, 3–39. [Google Scholar] [CrossRef] [Green Version]
- Raudvere, U.; Kolberg, L.; Kuzmin, I.; Arak, T.; Adler, P.; Peterson, H.; Vilo, J. G:Profiler: A web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 2019, 47, W191–W198. [Google Scholar] [CrossRef] [Green Version]
- Supek, F.; Bošnjak, M.; Škunca, N.; Šmuc, T. Revigo summarizes and visualizes long lists of gene ontology terms. PLoS ONE 2011, 6, 21800. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mofatto, L.S.; de Araújo Carneiro, F.; Vieira, N.G.; Duarte, K.E.; Vidal, R.O.; Alekcevetch, J.C.; Cotta, M.G.; Verdeil, J.-L.; Lapeyre-Montes, F.; Lartaud, M.; et al. Identification of candidate genes for drought tolerance in coffee by high-throughput sequencing in the shoot apex of different Coffea arabica cultivars. BMC Plant Biol. 2016, 16, 1–18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thioune, E.-H.; Strickler, S.; Gallagher, T.; Charpagne, A.; Decombes, P.; Osborne, B.; McCarthy, J. Temperature Impacts the response of Coffea canephora to decreasing soil water availability. Trop. Plant Biol. 2020, 13, 236–250. [Google Scholar] [CrossRef] [Green Version]
- Estavillo, G.M.; Crisp, P.A.; Pornsiriwong, W.; Wirtz, M.; Collinge, D.; Carrie, C.; Giraud, E.; Whelan, J.; David, P.; Javot, H.; et al. Evidence for a SAL1-PAP chloroplast retrograde pathway that functions in drought and high light signaling in Arabidopsis. Plant Cell 2011, 23, 3992–4012. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marques, I.; Fernandes, I.; David, P.H.C.; Paulo, O.S.; Goulao, L.F.; Fortunato, A.S.; Lidon, F.C.; DaMatta, F.M.; Ramalho, J.C.; Ribeiro-Barros, A.I. Transcriptomic leaf profiling reveals differential responses of the two most traded coffee species to elevated [CO2]. Int. J. Mol. Sci. 2020, 21, 9211. [Google Scholar] [CrossRef]
- Giuliano, G.; Al-Babili, S.; Von Lintig, J. Carotenoid oxygenases: Cleave it or leave it. Trends Plant Sci. 2003, 8, 145–149. [Google Scholar] [CrossRef]
- Hou, X.; Rivers, J.; León, P.; McQuinn, R.P.; Pogson, B.J. Synthesis and function of apocarotenoid signals in plants. Trends Plant Sci. 2016, 21, 792–803. [Google Scholar] [CrossRef]
- Choi, Y.; Lee, Y.; Kim, S.Y.; Lee, Y.; Hwang, J.-U. Arabidopsis ROP-interactive CRIB motif-containing protein 1 (RIC1) positively regulates auxin signalling and negatively regulates abscisic acid (ABA) signalling during root development. Plant Cell Environ. 2013, 36, 945–955. [Google Scholar] [CrossRef]
- Cao, Y.; Luo, Q.; Tian, Y.; Meng, F. Physiological and proteomic analyses of the drought stress response in Amygdalus mira (Koehne) Yü et Lu roots. BMC Plant Biol. 2017, 17, 1–16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, N.-Q.; Sun, Y.; Guo, T.; Shi, C.-L.; Zhang, Y.-M.; Kan, Y.; Xiang, Y.-H.; Zhang, H.; Yang, Y.-B.; Li, Y.-C.; et al. UDP-glucosyltransferase regulates grain size and abiotic stress tolerance associated with metabolic flux redirection in rice. Nat. Commun. 2020, 11, 1–16. [Google Scholar] [CrossRef]
- Fortunato, A.; Lidon, F.C.; Batista-Santos, P.; Leitão, A.E.; Pais, I.P.; Ribeiro, A.I.; Ramalho, J.C. Biochemical and molecular characterization of the antioxidative system of Coffea spp. under cold conditions in genotypes with contrasting tolerance. J. Plant Physiol. 2010, 167, 333–342. [Google Scholar] [CrossRef] [PubMed]
- Ramalho, J.C.; Rodrigues, A.P.; Lidon, F.C.; Marques, L.M.C.; Leitão, A.E.; Fortunato, A.S.; Pais, I.P.; Silva, M.J.; Scotti-Campos, P.; Lopes, A.; et al. Stress cross-response of the antioxidative system promoted by superimposed drought and cold conditions in Coffea spp. PLoS ONE 2018, 13, e0198694. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, K.; Takeda, M.; Kidokoro, S.; Yamada, K.; Sakuma, Y.; Urano, K.; Fujita, M.; Yoshiwara, K.; Matsukura, S.; Morishita, Y.; et al. Metabolic Pathways involved in cold acclimation identified by integrated analysis of metabolites and transcripts regulated by DREB1A and DREB2A. Plant Physiol. 2009, 150, 1972. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xia, Y.; Suzuki, H.; Borevitz, J.; Blount, J.; Guo, Z.; Patel, K.; Dixon, R.A.; Lamb, C. An extracellular aspartic protease functions in Arabidopsis disease resistance signaling. EMBO J. 2004, 23, 980. [Google Scholar] [CrossRef]
- Gong, L.; Zhang, H.; Gan, X.; Zhang, L.; Chen, Y.; Nie, F.; Shi, L.; Li, M.; Guo, Z.; Zhang, G.; et al. Transcriptome profiling of the potato (Solanum tuberosum L.) plant under drought stress and water-stimulus conditions. PLoS ONE 2015, 10, e0128041. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, L.; Santos, R.B.; Figueiredo, A. Defense and Offense Strategies: The Role of Aspartic Proteases in Plant–Pathogen Interactions. Biology 2021, 10, 75. [Google Scholar] [CrossRef] [PubMed]
- Sebastián, D.I.; Fernando, F.D.; Raúl, D.G.; Gabriela, G.M. Overexpression of Arabidopsis aspartic protease APA1 gene confers drought tolerance. Plant Sci. 2020, 292, 110406. [Google Scholar] [CrossRef] [PubMed]
- Yao, X.; Xiong, W.; Ye, T.; Wu, Y. Overexpression of the aspartic protease ASPG1 gene confers drought avoidance in Arabidopsis. J. Exp. Bot. 2012, 63, 2579. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, S.; Guo, M.; Wang, C.; Xu, W.; Shi, T.; Tong, G.; Zhen, C.; Cheng, H.; Yang, C.; Elsheery, N.I.; et al. Genome-wide characterization of aspartic protease (AP) gene family in Populus trichocarpa and identification of the potential PtAPs involved in wood formation. BMC Plant Biol. 2019, 19, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Devnarain, N.; Crampton, B.G.; Olivier, N.; van der Westhuyzen, C.; Becker, J.V.W.; O’Kennedy, M.M. Transcriptomic analysis of a Sorghum bicolor landrace identifies a role for beta-alanine betaine biosynthesis in drought tolerance. S. Afr. J. Bot. 2019, 127, 244–255. [Google Scholar] [CrossRef]
- Magwanga, R.O.; Lu, P.; Kirungu, J.N.; Lu, H.; Wang, X.; Cai, X.; Zhou, Z.; Zhang, Z.; Salih, H.; Wang, K.; et al. Characterization of the late embryogenesis abundant (LEA) proteins family and their role in drought stress tolerance in upland cotton. BMC Genet. 2018, 19, 1–31. [Google Scholar] [CrossRef] [Green Version]
- Yu, Z.; Wang, X.; Tian, Y.; Zhang, D.; Zhang, L. The functional analysis of a wheat group 3 late embryogenesis abundant protein in Escherichia coli and Arabidopsis under abiotic stresses. Plant Signal. Behav. 2019, 14, 1667207. [Google Scholar] [CrossRef] [PubMed]
- Wei, S.; Xia, R.; Chen, C.; Shang, X.; Ge, F.; Wei, H.; Chen, H.; Wu, Y.; Xie, Q. ZmbHLH124 identified in maize recombinant inbred lines contributes to drought tolerance in crops. Plant Biotechnol. J. 2021. [Google Scholar] [CrossRef]
- Alves, G.S.C.; Torres, L.F.; Déchamp, E.; Breitler, J.C.; Joët, T.; Gatineau, F.; Andrade, A.C.; Bertrand, B.; Marraccini, P.; Etienne, H. Differential fine-tuning of gene expression regulation in coffee leaves by CcDREB1D promoter haplotypes under water deficit. J. Exp. Bot. 2017, 68, 3017–3031. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alves, G.S.C.; Torres, L.F.; de Aquino, S.O.; Reichel, T.; Freire, L.P.; Vieira, N.G.; Vinecky, F.; This, D.; Pot, D.; Etienne, H.; et al. Nucleotide diversity of the coding and promoter regions of DREB1D, a candidate gene for drought tolerance in Coffea species. Trop. Plant Biol. 2018, 11, 31–48. [Google Scholar] [CrossRef]
- Torres, L.F.; Reichel, T.; Déchamp, E.; de Aquino, S.O.; Duarte, K.E.; Alves, G.S.C.; Silva, A.T.; Cotta, M.G.; Costa, T.S.; Diniz, L.E.C.; et al. Expression of DREB-like genes in Coffea canephora and C. arabica subjected to various types of abiotic stress. Trop. Plant Biol. 2019, 12, 98–116. [Google Scholar] [CrossRef] [Green Version]
- Lata, C.; Prasad, M. Role of DREBs in regulation of abiotic stress responses in plants. J. Exp. Bot. 2011, 62, 4731–4748. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Phukan, U.J.; Jeena, G.S.; Tripathi, V.; Shukla, R.K. Regulation of Apetala2/Ethylene response factors in plants. Front. Plant Sci. 2017, 8, 150. [Google Scholar] [CrossRef] [Green Version]
- Xie, Z.; Nolan, T.M.; Jiang, H.; Yin, Y. AP2/ERF Transcription Factor regulatory networks in hormone and abiotic stress responses in Arabidopsis. Front Plant Sci. 2019, 10, 228. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singh, D.; Laxmi, A. Transcriptional regulation of drought response: A tortuous network of transcriptional factors. Front. Plant Sci. 2015, 6, 895. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Müller, M.; Munné-Bosch, S. Ethylene Response Factors: A Key Regulatory Hub in Hormone and Stress Signaling. Plant Physiol. 2015, 169, 32–41. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bian, Z.; Gao, H.; Wang, C. NAC Transcription factors as positive or negative regulators during ongoing battle between pathogens and our food crops. Int. J. Mol. Sci. 2020, 22, 81. [Google Scholar] [CrossRef]
- Furihata, T.; Maruyama, K.; Fujita, Y.; Umezawa, T.; Yoshida, R.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Abscisic acid-dependent multisite phosphorylation regulates the activity of a transcription activator AREB1. Pro. Natl. Acad. Sci. USA 2006, 103, 1988. [Google Scholar] [CrossRef] [Green Version]
- Hussain, R.M.; Ali, M.; Feng, X.; Li, X. The essence of NAC gene family to the cultivation of drought-resistant soybean (Glycine max L. Merr.) cultivars. BMC Plant Biol. 2017, 17, 55. [Google Scholar] [CrossRef] [Green Version]
- Wu, L.; Zhang, Z.; Zhang, H.; Wang, X.C.; Huang, R. Transcriptional modulation of ethylene response factor protein JERF3 in the oxidative stress response enhances tolerance of tobacco seedlings to salt, drought, and freezing. Plant Physiol. 2008, 148, 1953–1963. [Google Scholar] [CrossRef] [Green Version]
- Mitula, F.; Tajdel, M.; Cieśla, A.; Kasprowicz-Maluśki, A.; Kulik, A.; Babula-Skowrońska, D.; Michalak, M.; Dobrowolska, G.; Sadowski, J.; Ludwików, A. Arabidopsis ABA-activated kinase MAPKKK18 is regulated by protein Phosphatase 2C ABI1 and the Ubiquitin–Proteasome pathway. Plant Cell Physiol. 2015, 56, 2351–2367. [Google Scholar] [CrossRef]
- Jeong, S.; Lim, C.W.; Lee, S.C. The pepper MAP kinase CaAIMK1 positively regulates ABA and drought stress responses. Front. Plant Sci. 2020, 11, 720. [Google Scholar] [CrossRef]
- Yang, Q.; Liu, K.; Niu, X.; Wang, Q.; Wan, Y.; Yang, F.; Li, G.; Wang, Y.; Wang, R. Genome-wide identification of PP2C genes and their expression profiling in response to drought and cold stresses in Medicago truncatula. Sci. Rep. 2018, 8, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; Zhu, T.; Song, Y.; Feng, L.; Farag, E.A.H.; Ren, M. ABSCISIC ACID INSENSITIVE5 interacts with RIBOSOMAL S6 KINASE2 to mediate ABA responses during seedling growth in Arabidopsis. Front. Plant Sci. 2021, 11, 598654. [Google Scholar] [CrossRef]
- Fernández-Rivas, M.; Bolhaar, S.; González-Mancebo, E.; Asero, R.; van Leeuwen, A.; Bohle, B.; Ma, Y.; Ebner, C.; Rigby, N.; Sancho, A.I.; et al. Apple allergy across Europe: How allergen sensitization profiles determine the clinical expression of allergies to plant foods. J. Allergy Clinic. Immunol. 2006, 118, 481–488. [Google Scholar] [CrossRef] [PubMed]
- Van Loon, L.C.; Rep, M.; Pieterse, C.M.J. Significance of Inducible defense-related proteins in infected plants. Annu. Rev. Phytopathol. 2006, 44, 135–162. [Google Scholar] [CrossRef] [Green Version]
- Pagliarani, G.; Paris, R.; Arens, P.; Tartarini, S.; Ricci, G.; Smulders, M.J.; van de Weg, W.E. A qRT-PCR assay for the expression of all Mal d 1 isoallergen genes. BMC Plant Biol. 2013, 13, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Teixeira, M.A.; Rajewski, A.; He, J.; Castaneda, O.G.; Litt, A.; Kaloshian, I. Classification and phylogenetic analyses of the Arabidopsis and tomato G-type lectin receptor kinases. BMC Genom. 2018, 19, 1–20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, X.L.; Yu, Q.Y.; Tang, L.L.; Ji, W.; Bai, X.; Cai, H.; Liu, X.F.; Ding, X.D.; Zhu, Y.M. GsSRK, a G-type lectin S-receptor-like serine/threonine protein kinase, is a positive regulator of plant tolerance to salt stress. J. Plant Physiol. 2013, 170, 505–515. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.-F.; Zhao, W.-Y.; Fu, J.-D.; Liu, Y.-W.; Chen, M.; Zhou, Y.-B.; Ma, Y.-Z.; Xu, Z.-S.; Xi, Y.-J. Genome-wide analysis of CDPK family in foxtail millet and determination of SiCDPK24 functions in drought stress. Front. Plant Sci. 2018, 9, 651. [Google Scholar] [CrossRef]
- Geiger, D.; Scherzer, S.; Mumm, P.; Stange, A.; Marten, I.; Bauer, H.; Ache, P.; Matschi, S.; Liese, A.; Al-Rasheid, K.A.S.; et al. Activity of guard cell anion channel SLAC1 is controlled by drought-stress signaling kinase-phosphatase pair. Proc. Natl. Acad. Sci. USA 2009, 106, 21425–21430. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.C.; Lan, W.; Buchanan, B.B.; Luan, S. A protein kinase-phosphatase pair interacts with an ion channel to regulate ABA signaling in plant guard cells. Proc. Natl. Acad. Sci. USA 2009, 106, 21419–21424. [Google Scholar] [CrossRef] [Green Version]
- Lim, J.; Lim, C.W.; Lee, S.C. Pepper novel serine-threonine kinase CaDIK1 Regulates drought tolerance via modulating ABA sensitivity. Front. Plant Sci. 2020, 11, 1133. [Google Scholar] [CrossRef]
- Marraccini, P.; Freire, L.P.; Alves, G.S.; Vieira, N.G.; Vinecky, F.; Elbelt, S.; Ramos, H.J.; Montagnon, C.; Vieira, L.G.; Leroy, T.; et al. RBCS1 expression in coffee: Coffea orthologs, Coffea arabica homeologs, and expression variability between genotypes and under drought stress. BMC Plant Biol. 2011, 11, 1–24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marraccini, P.; Vinecky, F.; Alves, G.S.C.; Ramos, H.J.O.; Elbelt, S.; Vieira, N.G.; Carneiro, F.A.; Sujii, P.S.; Alekcevetch, J.C.; Silva, V.A.; et al. Differentially expressed genes and proteins upon drought acclimation in tolerant and sensitive genotypes of Coffea canephora. J. Exp. Bot. 2012, 63, 4191. [Google Scholar] [CrossRef]
- Vieira, N.G.; Carneiro, F.A.; Sujii, P.S.; Alekcevetch, J.C.; Freire, L.P.; Vinecky, F.; Elbelt, S.; Silva, V.A.; DaMatta, F.M.; Ferrão, M.A.G.; et al. Different molecular mechanisms account for drought tolerance in Coffea canephora var. Conilon. Trop. Plant Biol. 2013, 6, 181–190. [Google Scholar] [CrossRef] [Green Version]
- De Oliveira Santos, M.; Coelho, L.S.; Carvalho, G.R.; Botelho, C.E.; Torres, L.F.; Vilela, D.J.M.; Andrade, A.C.; Silva, V.A. Photochemical efficiency correlated with candidate gene expression promote coffee drought tolerance. Sci. Rep. 2021, 11, 1–14. [Google Scholar] [CrossRef]
- Bertrand, B.; Bardil, A.; Baraille, H.; Dussert, S.; Doulbeau, S.; Dubois, E.; Severac, D.; Dereeper, A.; Etienne, H. The greater phenotypic homeostasis of the allopolyploid Coffea arabica improved the transcriptional homeostasis over that of both diploid parents. Plant Cell Physiol. 2015, 56, 2035. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dussert, S.; Serret, J.; Bastos-Siqueira, A.; Morcillo, F.; Déchamp, E.; Rofidal, V.; Lashermes, P.; Etienne, H.; JOët, T. Integrative analysis of the late maturation programme and desiccation tolerance mechanisms in intermediate coffee seeds. J. Exp. Bot. 2018, 69, 1583–1597. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martins, M.Q.; Fortunato, A.S.; Rodrigues, W.P.; Partelli, F.L.; Campostrini, E.; Lidon, F.C.; DaMatta, F.M.; Ramalho, J.C.; Ribeiro-Barros, A.I. Selection and validation of reference genes for accurate RT-qPCR Data normalization in Coffea spp. under a climate changes context of interacting elevated [CO2] and temperature. Front. Plant Sci. 2017, 8, 307. [Google Scholar] [CrossRef] [Green Version]
- Toniutti, L.; Breitler, J.-C.; Guittin, C.; Doulbeau, S.; Etienne, H.; Campa, C.; Lambot, C.; Pinilla, J.-C.H.; Bertrand, B. An altered circadian clock coupled with a higher photosynthesis efficiency could explain the better agronomic performance of a new coffee clone when compared with a standard variety. Int. J. Mol. Sci. 2019, 20, 736. [Google Scholar] [CrossRef] [Green Version]
- Rodrigues, A.M.; Jorge, T.; Osorio, S.; Pott, D.M.; Lidon, F.C.; DaMatta, F.M.; Marques, I.; Ribeiro-Barros, A.I.; Ramalho, J.C.; António, C. Primary metabolite profile changes in Coffea spp. promoted by single and combined exposure to drought and elevated CO2 concentration. Metabolites 2021, 11, 427. [Google Scholar] [CrossRef] [PubMed]
- Zapalska-Sozoniuk, M.; Chrobak, L.; Kowalczyk, K.; Kankofer, M. Is it useful to use several “omics” for obtaining valuable results? Mol. Biol. Rep. 2019, 46, 3597–3606. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Gene ID | Homolog | Protein Name | Molecular Function | FC |
|---|---|---|---|---|
| Up-regulated | ||||
| Cc09_g07380 | AT1G30760 | Putative Reticuline oxidase-like protein | oxidoreductase activity; FAD binding | 11.17 |
| Cc10_g00760 | Putative Arabidopsis protein of unknown function | 11.00 | ||
| Cc09_g07390 | AT4G20820 | Putative Reticuline oxidase-like protein | oxidoreductase activity; FAD binding | 10.61 |
| Cc02_g03420 | ERF027 | Putative Ethylene-responsive transcription factor | DNA-binding; DNA-binding transcription factor | 10.11 |
| Cc04_g10590 | CYP82C2 | Putative Cytochrome P450 82A1 | oxidoreductase activity; heme binding; iron ion binding | 9.81 |
| Cc06_g15980 | LEA46 | 18 kDa seed maturation protein | 9.67 | |
| Cc02_g29220 | CYP75B1 | Putative Cytochrome P450 750A1 | oxidoreductase activity; heme binding; iron ion binding | 9.24 |
| Cc01_g19640 | Putative Basic 7S globulin | aspartic-type endopeptidase activity | 9.12 | |
| Cc05_g00080 | EM6 | Late embryogenesis abundant protein EMB564 | 8.96 | |
| Cc06_g06630 | PUB22 | E3 ubiquitin-protein ligase PUB22 | ubiquitin-protein transferase activity | 8.94 |
| Down-regulated | ||||
| Cc07_g09010 | Putative unknown protein | −9.32 | ||
| Cc02_g16790 | SAUR50 | Auxin-induced protein 15A | −9.59 | |
| Cc04_g01940 | HTH | Hothead | oxidoreductase activity; FAD binding; mandelonitrile lyase activity * | −9.61 |
| Cc11_g01760 | Protein of unknown function, DUF642 | −9.77 | ||
| Cc02_g09710 | TBL36 | Trichome Birefringence-like36 | O-acetyltransferase activity * | −10.07 |
| Cc00_g29810 | CCR1 | NAD(P)-binding Rossmann-fold superfamily protein | oxidoreductase activity * | −10.22 |
| Cc07_g15190 | DOT3 | Putative BTB/POZ domain-containing protein DOT3 | −10.51 | |
| Cc06_g01530 | GPAT6 | Glycerol-3-phosphate acyltransferase 6 | transferase activity; phosphatase activity | −10.58 |
| Cc05_g09310 | PDF1 | Putative uncharacterized protein | −11.24 | |
| Cc06_g08040 | AZI1 | Putative uncharacterized protein | protein self-association * | −12.36 |
| Gene ID | Homolog | Protein Name | Molecular Function | FC |
|---|---|---|---|---|
| Up-regulated | ||||
| Cc06_g05620 | PAP20 | Probable purple acid phosphatase 20 | hydrolase activity; metal ion binding | 13.03 |
| Cc09_g07380 | AT1G30760 | Putative Reticuline oxidase-like protein | oxidoreductase activity; FAD binding | 12.44 |
| Cc09_g07390 | AT4G20820 | Putative Reticuline oxidase-like protein | oxidoreductase activity; FAD binding | 11.53 |
| Cc04_g10590 | CYP82C2 | Putative Cytochrome P450 82A1 | oxidoreductase activity; heme binding; iron ion binding | 10.95 |
| Cc10_g00760 | Putative Arabidopsis protein of unknown function (DUF241) | 10.92 | ||
| Cc08_g11420 | EXLB1 | Expansin-like B1 | 10.92 | |
| Cc00_g15520 | CWINV2 | Beta-fructofuranosidase, insoluble isoenzyme 1 | hydrolase activity | 10.54 |
| Cc07_g04930 | TBL19 | Putative Trichome Birefringence-like 19 | transferase activity * | 10.20 |
| Cc02_g24750 | Carotenoid cleavage dioxygenase 7 | oxidoreductase activity; metal ion binding | 9.82 | |
| Cc03_g15270 | AT4G20820 | Putative Reticuline oxidase-like protein | oxidoreductase activity; FAD binding | 9.78 |
| Down-regulated | ||||
| Cc00_g18380 | SDR2a | Momilactone A synthase | oxidoreductase activity * | −9.93 |
| Cc10_g02800 | RL1 | RAD-like 1 | DNA-binding transcription factor activity * | −10.15 |
| Cc01_g12960 | EXPA4 | Expansin-A4 | −10.31 | |
| Cc01_g05920 | GASA6 | Protein GAST1 | −10.48 | |
| Cc04_g12550 | RGI3 | Probable LRR receptor-like serine/threonine-protein kinase At4g26540 | ATP binding; protein kinase activity; peptide receptor activity *; peptide-binding * | −10.56 |
| Cc01_g11300 | FLA11 | Fasciclin-like arabinogalactan protein 11 | −10.58 | |
| Cc11_g07530 | RLP55 | Putative LRR receptor-like serine/threonine-protein kinase GSO1 | −10.61 | |
| Cc11_g08360 | Putative MLP-like protein 28 | −10.72 | ||
| Cc04_g15520 | Putative Mitochondrial outer membrane protein porin of 36 kDa | transmembrane transporter activity | −10.93 | |
| Cc03_g10850 | RIC5 | Putative ROP-interactive CRIB motif-containing protein 4 | −21.55 |
| Gene ID | Homolog | Protein Name | Molecular Function | FC |
|---|---|---|---|---|
| Up-regulated | ||||
| Cc05_g08230 | AT5G34940 | Putative Glycoside hydrolase family 79, N-terminal | 21.23 | |
| Cc03_g06560 | MYB59 | Putative Transcription factor MYB48 | transcription regulator activity; DNA binding | 8.02 |
| Cc07_g18370 | Predicted protein | 7.06 | ||
| Cc11_g04480 | HSP18.1 | 18.5 kDa class I heat shock protein | protein self-association *; unfolded protein binding * | 6.68 |
| Cc00_g26020 | ribonuclease Ps | 6.31 | ||
| Cc07_g13110 | ERF1B | Putative Ethylene-responsive transcription factor 15 | transcription regulator activity; DNA binding | 6.28 |
| Cc11_g04470 | HSP18.1 | 18.5 kDa class I heat shock protein | protein self-association *; unfolded protein binding * | 6.13 |
| Cc02_g23670 | HSP22.0 | 22.7 kDa class IV heat shock protein | protein self-association *; unfolded protein binding * | 6.11 |
| Cc04_g05250 | HSP70-4 | Heat shock 70 kDa protein | ATP binding; ATPase activity; protein binding *; protein folding chaperone * | 6.05 |
| Cc00_g04790 | Hypothetical protein | 5.99 | ||
| Down-regulated | ||||
| Cc01_g10980 | UXT1 | Uncharacterized membrane protein At1g06890 | transmembrane transporter activity | −9.06 |
| Cc06_g02440 | Laccase-4 | copper ion binding; hydroquinone:oxygen oxidoreductase activity | −9.16 | |
| Cc06_g02050 | RHO guanyl-nucleotide exchange factor 7 | guanyl-nucleotide exchange factor activity | −9.17 | |
| Cc09_g03130 | AT5G33370 | GDSL esterase/lipase At5g33370 | hydrolase activity, acting on ester bonds | −9.33 |
| Cc07_g11210 | PER64 | Peroxidase 64 | heme binding; metal ion binding; peroxidase activity | −9.85 |
| Cc06_g01530 | GPAT6 | Glycerol-3-phosphate acyltransferase 6 | acyltransferase activity; phosphatase activity * | −10.32 |
| Cc04_g07330 | ASPG1 | Putative Protein Aspartic Protease in Guard Cell 1 | aspartic-type peptidase activity; DNA binding | −10.55 |
| Cc00_g31960 | COBL4 | Cobra-like protein 4 | −10.96 | |
| Cc02_g05960 | LAC5 | Laccase-5 | copper ion binding; hydroquinone:oxygen oxidoreductase activity | −11.3 |
| Cc06_g19110 | ORG2 | Putative Transcription factor ORG2 | transcription regulator activity; protein dimerization activity | −25.66 |
| Gene ID | Homolog | Protein Name | Molecular Function | FC |
|---|---|---|---|---|
| Up-regulated | ||||
| Cc09_g07390 | AT4G20820 | Putative Reticuline oxidase-like protein | oxidoreductase activity; FAD binding | 20.55 |
| Cc01_g08410 | Putative Lysosomal beta glucosidase | hydrolase activity, hydrolyzing O-glycosyl compounds | 20.47 | |
| Cc10_g04010 | Hypothetical protein | 20.47 | ||
| Cc03_g09220 | UGT75D1 | Putative UDP-glycosyltransferase 75D1 | transferase activity | 20.01 |
| Cc08_g11420 | EXLB1 | Expansin-like B1 | 10.97 | |
| Cc02_g27970 | Uncharacterized protein | 9.08 | ||
| Cc03_g06560 | MYB59 | Putative Transcription factor MYB48 | transcription regulator activity; DNA binding | 9.04 |
| Cc07_g18370 | Predicted protein | 8.24 | ||
| Cc01_g04050 | AT2G23950 | Putative G-type lectin S-receptor-like serine/threonine-protein kinase RLK1 | ATP binding; protein serine/threonine kinase activity; coreceptor activity * | 8.05 |
| Cc07_g09470 | Hypothetical protein | 8.04 | ||
| Down-regulated | ||||
| Cc00_g31960 | COBL4 | Cobra-like protein 4 | −11.58 | |
| Cc04_g02380 | AT3G16370 | GDSL esterase/lipase APG | hydrolase activity, acting on ester bonds | −11.64 |
| Cc03_g07230 | AT4G13710 | Probable pectate lyase 15 | metal ion binding; pectate lyase activity | −12.10 |
| Cc02_g16500 | PRP4 | Hypothetical protein | −12.45 | |
| Cc07_g07560 | XTH6 | Probable xyloglucan endotransglucosylase/hydrolase protein 6 | hydrolase activity, hydrolyzing O-glycosyl compounds; xyloglucan:xyloglucosyl transferase activity | −12.86 |
| Cc03_g09190 | XTH6 | Probable xyloglucan endotransglucosylase/hydrolase protein 16 | hydrolase activity, hydrolyzing O-glycosyl compounds; xyloglucan:xyloglucosyl transferase activity | −12.87 |
| Cc01_g11300 | FLA11 | Fasciclin-like arabinogalactan protein 11 | −13.20 | |
| Cc10_g04590 | SBT1.8 | Putative Subtilisin-like protease | serine-type endopeptidase activity | −13.34 |
| Cc08_g04660 | SOT15 | Putative Cytosolic sulfotransferase 15 | sulfotransferase activity | −13.39 |
| Cc02_g17500 | RL1 | Hypothetical protein | transcription regulator activity | −13.91 |
| Gene ID | Homolog | Protein Name | CL153 | Icatu | ||
|---|---|---|---|---|---|---|
| MWD | SWD | MWD | SWD | |||
| Response to Water Deprivation | ||||||
| Cc06_g15980 | AT5G06760 | 18 kDa seed maturation protein | 9.67 | 9.19 | ||
| Cc04_g07360 | AT3G18490 | Putative Protein Aspartic Protease in Guard Cell 1 | 5.24 | −4.43 | ||
| Cc04_g07380 | AT3G18490 | Putative Protein Aspartic Protease in Guard Cell 1 | 4.35 | |||
| Cc11_g05800 | Annexin D5 | 1.48 | 1.49 | |||
| Cc04_g09640 | AT3G18490 | Protein Aspartic Protease in Guard Cell 1 | −5.00 | −6.97 | ||
| Cc07_g07560 | AT5G65730 | Probable xyloglucan endotransglucosylase/hydrolase protein 6 | −6.94 | −12.86 | ||
| Cc04_g07330 | AT3G18490 | Putative Protein Aspartic Protease in Guard Cell 1 | −9.13 | −10.19 | ||
| Cc04_g07360 | AT3G18490 | Putative Protein Aspartic Protease in Guard Cell 1 | 4.42 | −3.48 | ||
| Cc04_g07380 | AT3G18490 | Putative Protein Aspartic Protease in Guard Cell 1 | 3.30 | −3.23 | ||
| Cc11_g05800 | Annexin D5 | 1.49 | ||||
| Cc07_g15660 | AT2G22125 | C2 domain-containing protein | −1.71 | -2.17 | ||
| Cc04_g08280 | AT5G08120 | Putative movement protein-binding protein 2C | −2.83 | |||
| Cc04_g09640 | AT3G18490 | Protein Aspartic Protease in Guard Cell 1 | −4.84 | −4.24 | ||
| Cc04_g07350 | AT3G18490 | Putative Protein Aspartic Protease in Guard Cell 1 | −5.66 | −3.86 | −6.28 | |
| Cc07_g07560 | AT5G65730 | Probable xyloglucan endotransglucosylase/hydrolase protein 6 | −5.71 | −4.75 | ||
| Cc02_g15480 | AT4G18780 | Cellulose synthase A catalytic subunit 8 (UDP-forming) | −6.4 | −7.5 | −8.73 | |
| Cc04_g07330 | AT3G18490 | Putative Protein Aspartic Protease in Guard Cell 1 | −9.27 | −10.55 | ||
| Cc04_g07370 | AT3G18490 | Putative Protein Aspartic Protease in Guard Cell 1 | −1.93 | −3.31 | ||
| Cc01_g21050 | Sucrose synthase 2 | 2.64 | ||||
| Response to desiccation | ||||||
| Cc02_g38620 | Desiccation protectant protein Lea14 homolog | 1.76 | 1.20 | 1.50 | ||
| Cc01_g08980 | Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family | −3.16 | ||||
| Cc04_g03400 | Putative Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family | 1.60 | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernandes, I.; Marques, I.; Paulo, O.S.; Batista, D.; Partelli, F.L.; Lidon, F.C.; DaMatta, F.M.; Ramalho, J.C.; Ribeiro-Barros, A.I. Understanding the Impact of Drought in Coffea Genotypes: Transcriptomic Analysis Supports a Common High Resilience to Moderate Water Deficit but a Genotype Dependent Sensitivity to Severe Water Deficit. Agronomy 2021, 11, 2255. https://doi.org/10.3390/agronomy11112255
Fernandes I, Marques I, Paulo OS, Batista D, Partelli FL, Lidon FC, DaMatta FM, Ramalho JC, Ribeiro-Barros AI. Understanding the Impact of Drought in Coffea Genotypes: Transcriptomic Analysis Supports a Common High Resilience to Moderate Water Deficit but a Genotype Dependent Sensitivity to Severe Water Deficit. Agronomy. 2021; 11(11):2255. https://doi.org/10.3390/agronomy11112255
Chicago/Turabian StyleFernandes, Isabel, Isabel Marques, Octávio S. Paulo, Dora Batista, Fábio L. Partelli, Fernando C. Lidon, Fábio M. DaMatta, José C. Ramalho, and Ana I. Ribeiro-Barros. 2021. "Understanding the Impact of Drought in Coffea Genotypes: Transcriptomic Analysis Supports a Common High Resilience to Moderate Water Deficit but a Genotype Dependent Sensitivity to Severe Water Deficit" Agronomy 11, no. 11: 2255. https://doi.org/10.3390/agronomy11112255
APA StyleFernandes, I., Marques, I., Paulo, O. S., Batista, D., Partelli, F. L., Lidon, F. C., DaMatta, F. M., Ramalho, J. C., & Ribeiro-Barros, A. I. (2021). Understanding the Impact of Drought in Coffea Genotypes: Transcriptomic Analysis Supports a Common High Resilience to Moderate Water Deficit but a Genotype Dependent Sensitivity to Severe Water Deficit. Agronomy, 11(11), 2255. https://doi.org/10.3390/agronomy11112255













