Identification and Characterization of Diaporthe spp. Associated with Twig Cankers and Shoot Blight of Almonds in Spain
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling and Isolation
2.2. DNA Extraction, PCR Amplification and Sequencing
2.3. Phylogenetic Analyses
2.4. Taxonomy
2.5. Pathogenicity Tests
3. Results
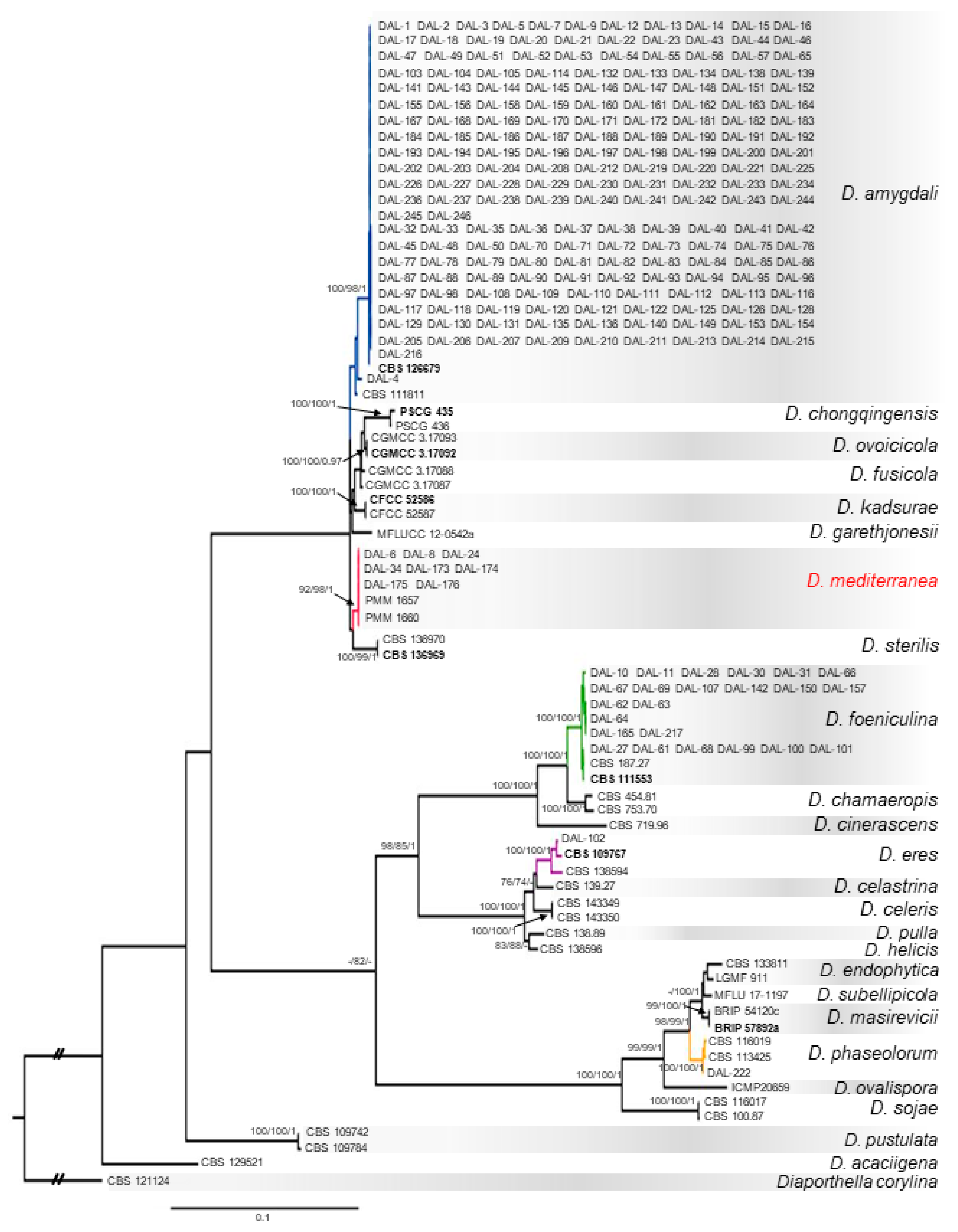
3.1. Phylogenetic Analyses
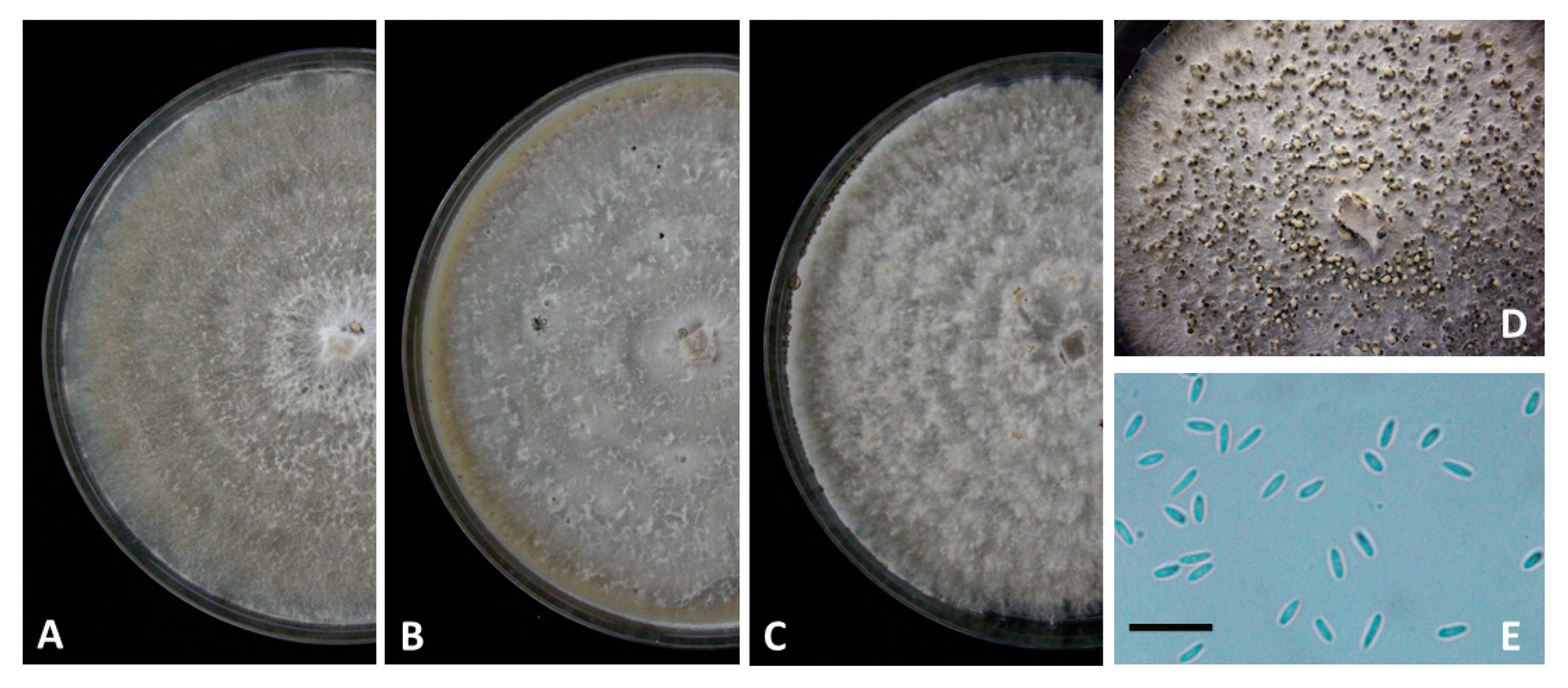
3.2. Taxonomy
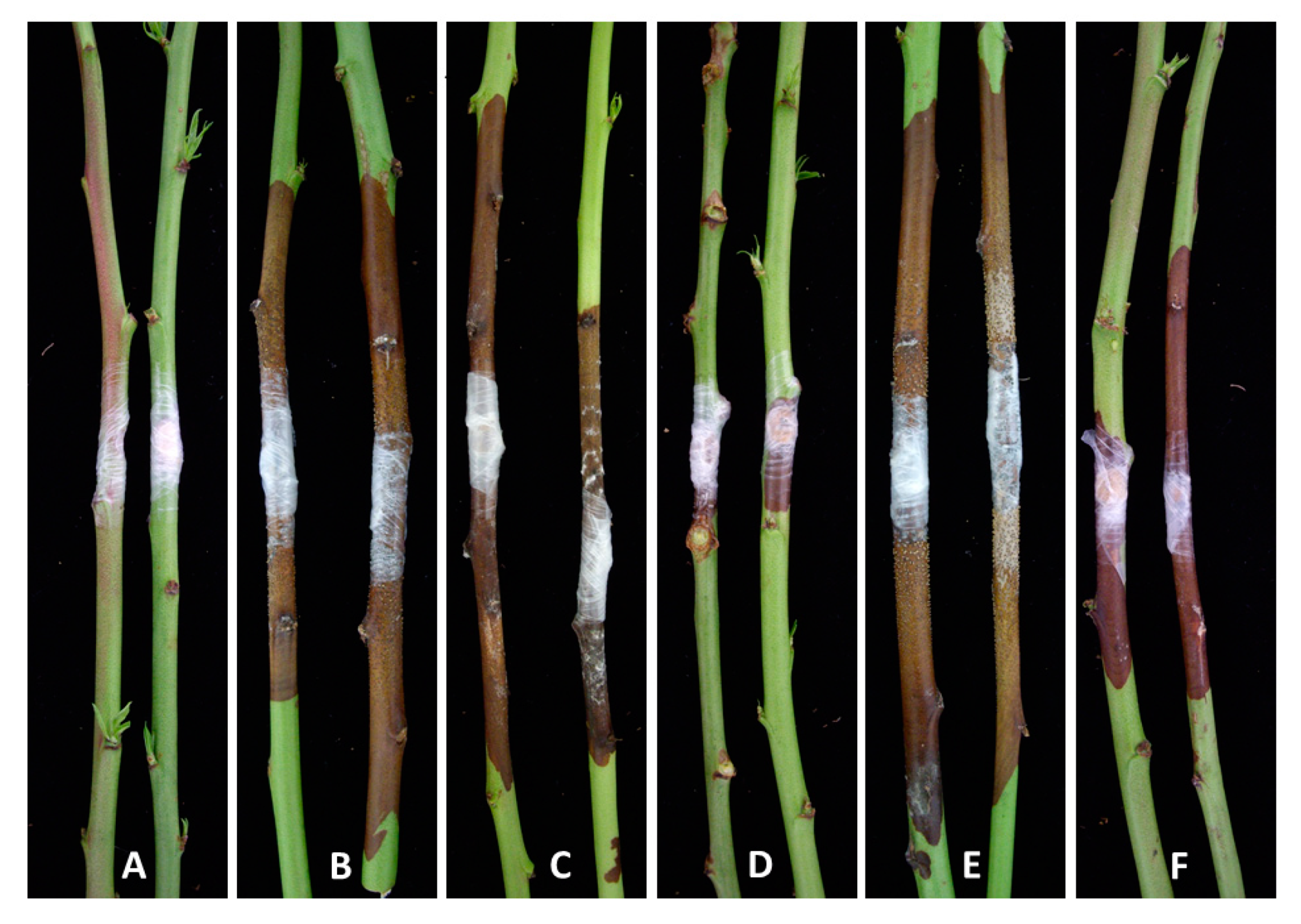
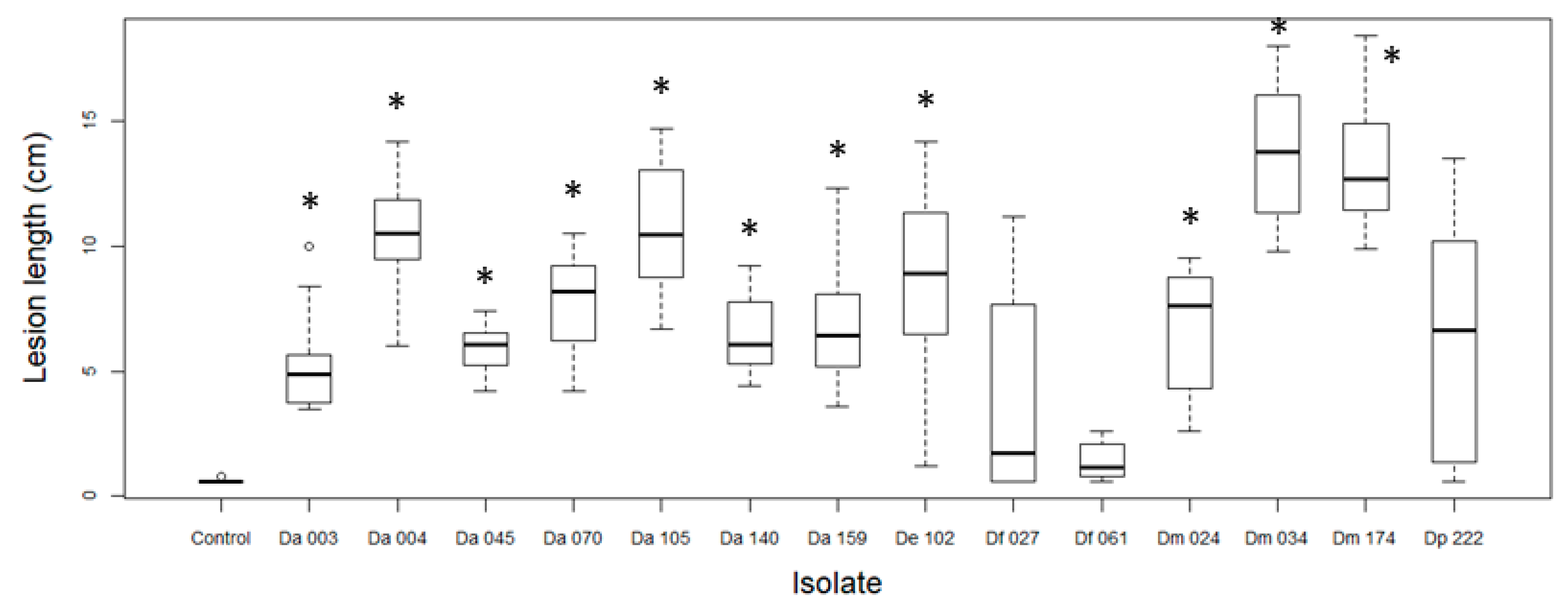
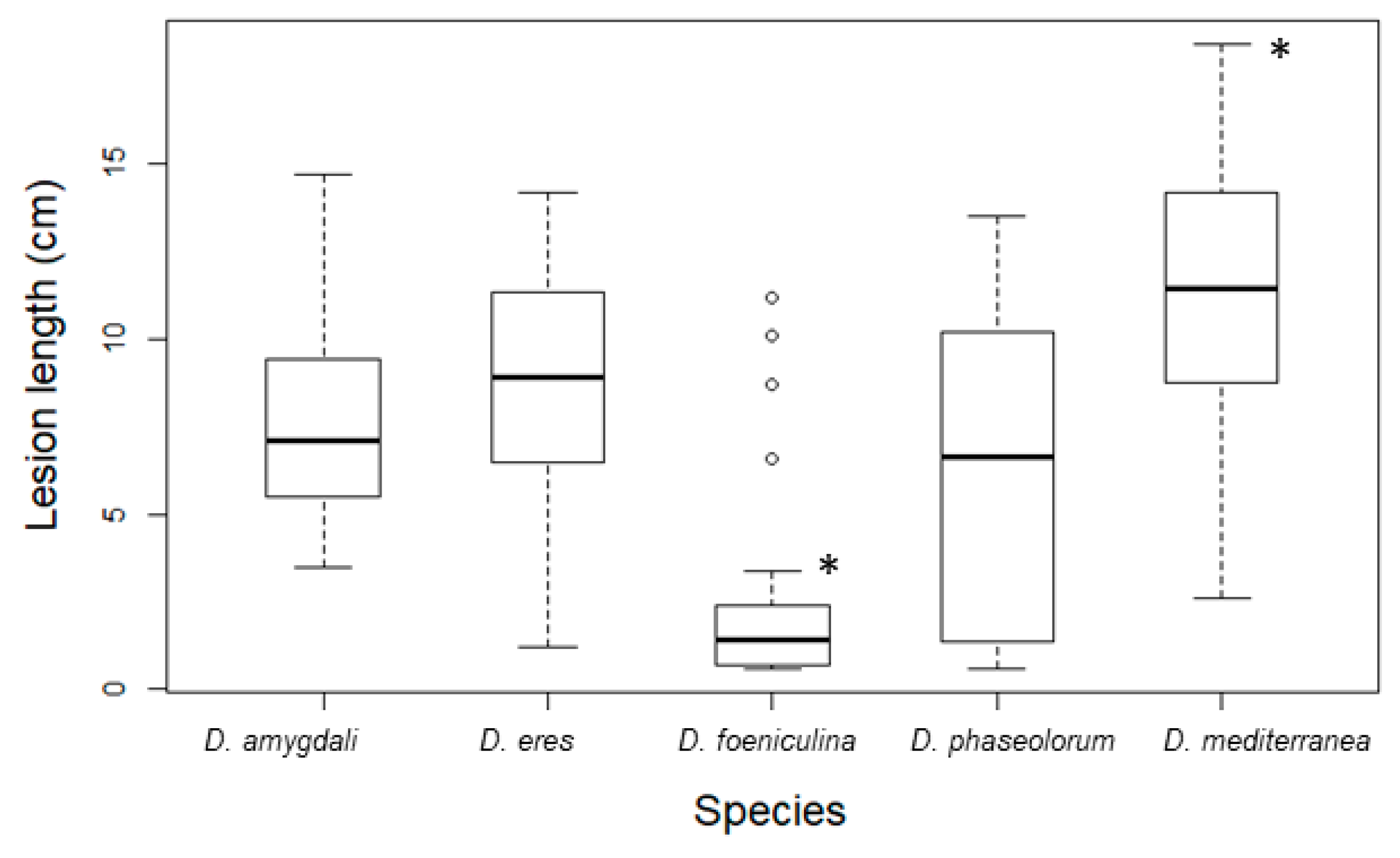
3.3. Pathogenicity Tests
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- FAOSTAT. Food and Agriculture Organization of the United Nations. 2018. Available online: http://www.fao.org/faostat/es/#dat (accessed on 1 June 2020).
- MAPA Anuario de Estadística Agraria. 2019. Available online: https://www.mapa.gob.es/es/estadistica/ (accessed on 1 June 2020).
- Egea, J.; Dicenta, F. Algunas consideraciones sobre el cultivo del almendro en secano. Fruticultura 2016, 49, 102–111. [Google Scholar]
- Miarnau, X.; Torguet, L.; Batlle, I.; Alegre, S. El cultivo del almendro en alta densidad. Fruticultura 2016, 49, 68–87. [Google Scholar]
- Barrios-Sanromà, G.; Aymamí-Besora, A. El futuro de la sanidad vegetal del almendro. Fruticultura 2016, 49, 128–151. [Google Scholar]
- Ollero-Lara, A.; López-Moral, A.; Lovera, M.; Raya, M.C.; Roca, L.F.; Arquero, O.; Trapero, A. Las enfermedades del almendro en Andalucía. Fruticultura 2016, 49, 166–183. [Google Scholar]
- Adaskaveg, J.E. Phomopsis canker and fruit rot. In Compendium of Nut Crop Diseases in Temperate Zones; Teviotdale, B.L., Michailides, T.J., Pscheidt, J.W., Eds.; APS Press: St. Paul, MN, USA, 2002; pp. 27–28. [Google Scholar]
- Diogo, E.L.F.; Santos, J.M.; Phillips, A.J.L. Phylogeny, morphology and pathogenicity of Diaporthe and Phomopsis species on almond in Portugal. Fungal Divers. 2010, 44, 107–115. [Google Scholar] [CrossRef]
- Tuset, J.J.; Portilla, M.T. Taxonomic status of Fusicoccum amygdali and Phomopsis amygdalina. Can. J. Bot. 1989, 67, 1275–1280. [Google Scholar] [CrossRef]
- Tuset, J.J.; Hinarejos, C.; Portilla, M.T. Incidence of Phomopsis amygdali, Botryosphaeria berengeriana and Valsa cincta diseases in almond under different control strategies. EPPO Bull. 1997, 27, 449–454. [Google Scholar] [CrossRef]
- Delacroix, G. Sur une maladie des amandiers en Provence. Bull. Trimest. Soc. Mycol. Fr. 1905, 21, 180–185. [Google Scholar]
- Udayanga, D.; Liu, X.; Crous, P.W.; McKenzie, E.H.C.; Chukeatirote, E.; Hyde, K.D. A multi-locus phylogenetic evaluation of Diaporthe (Phomopsis). Fungal Divers. 2012, 56, 157–171. [Google Scholar] [CrossRef]
- Rossman, A.Y.; Adams, G.C.; Cannon, P.F.; Castlebury, L.A.; Crous, P.W.; Gryzenhout, M.; Jaklitsch, W.M.; Mejía, L.C.; Stoykov, D.; Udayanga, D.; et al. Recommendations of generic names in Diaporthales competing for protection or use. IMA Fungus 2015, 6, 145–154. [Google Scholar] [CrossRef]
- Gomes, R.R.; Glienke, C.; Videira, S.I.R.; Lombard, L.; Groenewald, J.Z.; Crous, P.W. Diaporthe: A genus of endophytic, saprobic and plant pathogenic fungi. Persoonia 2013, 31, 1–41. [Google Scholar] [CrossRef]
- Gao, Y.; Lui, F.; Duan, W.; Crous, P.W.; Cai, L. Diaporthe is paraphyletic. IMA Fungus 2017, 8, 153–187. [Google Scholar] [CrossRef]
- Dissanayake, A.J.; Phillips, A.J.L.; Hyde, K.D.; Yan, J.Y.; Li, X.H. The current status of species in Diaporthe. Mycosphere 2017, 8, 1106–1156. [Google Scholar] [CrossRef]
- Santos, L.; Alves, A.; Alves, R. Evaluating multi-locus phylogenies for species boundaries determination in the genus Diaporthe. PeerJ 2017, 5, e3120. [Google Scholar] [CrossRef]
- Lawrence, D.P.; Travadon, R.; Baumgartner, K. Diversity of Diaporthe species associated with wood cankers of fruit and nut crops in northern California. Mycologia 2015, 107, 926–940. [Google Scholar] [CrossRef]
- Gramaje, D.; Agustí-Brisach, C.; Pérez-Sierra, A.; Moralejo, E.; Olmo, D.; Mostert, L.; Damm, U.; Armengol, J. Fungal trunk pathogens associated with Wood decay of almond trees on Mallorca (Spain). Persoonia 2012, 28, 1–13. [Google Scholar] [CrossRef]
- Gardes, M.; Bruns, T.D. ITS primers with enhanced specificity for basiodiomycetes-applications to the identification of mycorrhizae and rusts. Mol. Ecol. 1993, 2, 113–118. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.D.; Lee, S.B.; Taylor, J.W. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols—A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Alves, A.; Crous, P.W.; Correia, A.; Phillips, A.J.L. Morphological and molecular data reveal cryptic speciation in Lasiodiplodia theobromae. Fungal Divers. 2008, 28, 1–13. [Google Scholar]
- Travadon, R.; Lawrence, D.P.; Rooney-Latham, S.; Gubler, W.D.; Wilcox, W.F.; Rolshausen, P.E.; Baumgartner, K. Cadophora species associated with wood-decay of grapevine in North America. Fungal Biol. 2015, 119, 53–66. [Google Scholar] [CrossRef]
- O’Donnell, K.; Cigelnik, E. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol. Phylogenet. Evol. 1997, 7, 103–116. [Google Scholar]
- Glass, N.L.; Donaldson, G. Development of primer sets designed for use with PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microb. 1995, 61, 1323–1330. [Google Scholar] [CrossRef]
- Crous, P.W.; Groenewald, J.Z.; Risède, J.M.; Simoneau, P.; Hywel-Jones, N.L. Calonectria species and their Cylindrocladium anamorphs: Species with sphaeropedunculate vesicles. Stud. Mycol. 2004, 50, 415–430. [Google Scholar]
- Weir, B.S.; Johnston, P.R.; Damm, U. The Colletotrichum gloeosporioides species complex. Stud. Mycol. 2012, 73, 115–180. [Google Scholar] [CrossRef] [PubMed]
- Udayanga, D.; Castlebury, L.A.; Rossman, A.Y.; Hyde, K.D. Species limits in Diaporthe: Molecular re-assessment of D. citri, D. cytosporella, D. foeniculina and D. rudis. Persoonia 2014, 32, 83–101. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Vaidya, G.; Lohman, D.J.; Meier, R. SequenceMatrix: Concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 2011, 27, 171–180. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayers, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large modelspace. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In Proceedings of the Gateway Computing Environments Workshop (GCE), New Orleans, LA, USA, 14 November 2010; pp. 1–8. [Google Scholar]
- Nylander, J.A.A. MrModeltest v2. (Program Distributed by the Author.) Evolutionary Biology Centre; Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Rayner, R.W. A Mycological Colour Chart; Commonwealth Mycological Institute: Kew, UK, 1970. [Google Scholar]
- Duthie, J.A. Models of the response of foliar parasites to the combined effects of temperature and duration of wetness. Phytopathology 1997, 87, 1088–1095. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R. A Language and Environment for Statistical Computing; R Foundation for Statistical Computing; R Core Team. R: Vienna, Austria, 2020; Available online: https://www.R-project.org/ (accessed on 1 June 2020).
- Mendiburu, F. Agricolae: Statistical Procedures for Agricultural Research. R Package Version 1.2-3. 2015. Available online: http://CRAN.R-project.org/package=agricolae (accessed on 1 June 2020).
- Van Niekerk, J.M.; Groenewald, J.Z.; Farr, D.F.; Fourie, P.H.; Halleen, F.; Crous, P.W. Reassessment of Phomopsis species on grapevines. Australas. Plant Path. 2005, 34, 27–39. [Google Scholar] [CrossRef]
- Lesuthu, P.; Mostert, L.; Spies, C.F.J.; Moyo, P.; Regnier, T.; Halleen, F. Diaporthe nebulae sp. nov. and first feport of D. cynaroidis, D. novem, and D. serafiniae on Grapevines in South Africa. Plant Dis. 2019, 103, 808–817. [Google Scholar] [CrossRef] [PubMed]
- Lombard, L.; Van Leeuwen, G.C.; Guarnaccia, V.; Polizzi, G.; Van Rijswick, P.C.; Rosendahl, K.C.; Gabler, J.; Crous, P.W. Diaporthe species associated with Vaccinium, with specific reference to Europe. Phytopathol. Mediterr. 2014, 53, 287–299. [Google Scholar]
- Guarnaccia, V.; Groenewald, J.Z.; Woodhall, J.; Armengol, J.; Cinelli, T.; Eichmeier, A.; Ezra, D.; Fontaine, F.; Gramaje, D.; Gutiérrez-Aguirregabiria, A.; et al. Diaporthe diversity and pathogenicity revealed from a broad survey of grapevine diseases in Europe. Persoonia 2018, 40, 135–153. [Google Scholar] [CrossRef]
- Pantidou, M.E. Fungus-host index for Greece; Benaki Phytopathological Institute: Kiphissia, Athens, 1973; p. 382. [Google Scholar]
- Varjas, V.; Vajna, L.; Izsépi, F.; Nagy, G.; Pájtli, É. First report of Phomopsis amygdali causing twig canker on almond in Hungary. Plant Dis. 2017, 101, 1674. [Google Scholar] [CrossRef]
- Canonaco, A. Il seccume dei rameti di mandorlo in relazione ad alcuni micromiceti. Riv. Patol. Veget. 1936, 26, 145–164. [Google Scholar]
- Dias, M.R.S.; Lucas, M.T.; Lopes, M.C. Fungi Lusitaniae XXIX. Agron. Lusit. 1982, 41, 175–192. [Google Scholar]
- Trigui, A. Sur la présence en Tunisie de Fusicoccum amygdali Delacroix sur Amandier. Bull. ENSAT 1968, 18, 65–68. [Google Scholar]
- Garofalo, F. L’Albicocco “Tonda di Costigliole”, nuovo ospite di Fusicoccum amygdali Del. Inf. Fitopatol. 1973, 23, 13–15. [Google Scholar]
- Michailides, T.J.; Thomidis, T. First Report of Phomopsis amygdali Causing Fruit Rot on Peaches in Greece. Plant Dis. 2006, 90, 1551. [Google Scholar] [CrossRef] [PubMed]
- López-Moral, A.; Lovera, M.; Raya, M.C.; Cortés-Cosano, N.; Arquero, O.; Trapero, A.; Agustí-Brisach, C. Etiology of branch dieback and shoot blight of English walnut caused by Botryosphaeriaceae and Diaporthe Species in Southern Spain. Plant Dis. 2020, 104, 533–550. [Google Scholar] [CrossRef] [PubMed]
- Adaskaveg, J.E.; Forster, H.; Connell, J.H. First report of fruit rot and associated branch dieback of almond in California caused by a Phomopsis species tentatively identified as P. amygdali. Plant Dis. 1999, 83, 1073. [Google Scholar] [CrossRef] [PubMed]
- Farr, D.F.; Castlebury, L.A.; Pardo-Schultheiss, R. Phomopsis amygdali causes peach shoot blight of cultivated peach trees in the southeastern United States. Mycologia 1999, 91, 1008–1015. [Google Scholar] [CrossRef]
- Mostert, L.; Crous, P.W.; Kang, J.C.; Phillips, A.J.L. Species of Phomopsis and a Libertella sp. occurring on grapevines with specific reference to South Africa: Morphological, cultural, molecular and pathological characterization. Mycologia 2001, 93, 146–167. [Google Scholar] [CrossRef]
- Kanematsu, S.; Yokoyama, Y.; Kobayashi, T. Taxonomic reassessment of the causal fungus of peach Fusicoccum canker in Japan. Ann. Phytopathol. Soc. Jpn. 1999, 65, 531–536. [Google Scholar] [CrossRef]
- Álvarez, M.I.; Perdomo, E.; Martínez, E.S.; Mondino, P.; Alaniz, S. Phomopsis amygdali principal agente causal de la viruela de la púa en durazneros y nectarinos en Uruguay. In Abstracts of the 13th National Congress of Hortifruticulture; INIA—Sociedad Uruguaya de Horti-Fruticultura: Montevideo, Uruguay, 2014; p. 91. [Google Scholar]
- Sessa, L.; Abreo, E.; Bettucci, L.; Lupo, S. Diversity and virulence of Diaporthe species associated with wood disease symptoms in deciduous fruit trees in Uruguay. Phytopathol. Mediterr. 2017, 56, 431–444. [Google Scholar]
- Dai, F.M.; Zeng, R.; Lu, J.P. First report of twig canker on peach caused by Phomopsis amygdali in China. Plant Dis. 2012, 96, 288. [Google Scholar] [CrossRef]
- Bai, Q.; Zhai, L.; Chen, X.; Hong, N.; Xu, W.; Wang, G. Biological and molecular characterization of five Phomopsis species associated with pear shoot canker in China. Plant Dis. 2015, 99, 1704–1712. [Google Scholar] [CrossRef]
- Meng, L.; Yu, C.; Wang, C.; Li, G. First report of Diaporthe amygdali causing walnut twig canker in Shandong province of China. Plant Dis. 2018, 102, 1859. [Google Scholar] [CrossRef]
- Santos, L.; Phillips, A.J.L.; Crous, P.W.; Alves, A. Diaporthe species on Rosaceae with descriptions of D. pyracanthae sp. nov. and D. malorum sp. nov. Mycosphere 2017, 8, 485–511. [Google Scholar] [CrossRef]

| Species | Strain Number | Year | Location | Province/Region | GenBank Accession Numbers | ||||
|---|---|---|---|---|---|---|---|---|---|
| ITS | tef-1α | tub | his | cal | |||||
| D. amygdali | DAL-1 | 2014 | Sant Joan | Mallorca/Islas Baleares | MT007292 | MT006769 | MT006466 | - | - |
| DAL-2 | 2014 | Sant Joan | Mallorca/Islas Baleares | MT007293 | MT006770 | MT006467 | - | - | |
| DAL-3 | 2014 | Santa Margalida | Mallorca/Islas Baleares | MT007294 | MT006771 | MT006468 | MT006997 | MT006694 | |
| DAL-4 | 2014 | Santa Margalida | Mallorca/Islas Baleares | MT007295 | MT006772 | MT006469 | MT006998 | MT006695 | |
| DAL-5 | 2014 | Calvià | Mallorca/Islas Baleares | MT007296 | MT006773 | MT006470 | - | - | |
| DAL-7 | 2014 | Calvià | Mallorca/Islas Baleares | MT007297 | MT006774 | MT006471 | MT006999 | - | |
| DAL-9 | 2014 | Calvià | Mallorca/Islas Baleares | MT007298 | MT006775 | MT006472 | MT007000 | MT006696 | |
| DAL-12 | 2014 | Binissalem | Mallorca/Islas Baleares | MT007299 | MT006776 | MT006473 | MT007001 | - | |
| DAL-13 | 2014 | Llucmajor | Mallorca/Islas Baleares | MT007300 | MT006777 | MT006474 | - | - | |
| DAL-14 | 2014 | Llucmajor | Mallorca/Islas Baleares | MT007301 | MT006778 | MT006475 | - | - | |
| DAL-15 | 2014 | Marratxí | Mallorca/Islas Baleares | MT007302 | MT006779 | MT006476 | MT007002 | - | |
| DAL-16 | 2014 | Sa Pobla | Mallorca/Islas Baleares | MT007303 | MT006780 | MT006477 | MT007003 | MT006697 | |
| DAL-17 | 2014 | Sa Pobla | Mallorca/Islas Baleares | MT007304 | MT006781 | MT006478 | - | - | |
| DAL-18 | 2014 | Inca | Mallorca/Islas Baleares | MT007305 | MT006782 | MT006479 | MT007004 | - | |
| DAL-19 | 2014 | Binissalem | Mallorca/Islas Baleares | MT007306 | MT006783 | MT006480 | MT007005 | - | |
| DAL-20 | 2014 | Palma | Mallorca/Islas Baleares | MT007307 | MT006784 | MT006481 | - | - | |
| DAL-21 | 2014 | Binissalem | Mallorca/Islas Baleares | MT007308 | MT006785 | MT006482 | - | - | |
| DAL-22 | 2014 | Llucmajor | Mallorca/Islas Baleares | MT007309 | MT006786 | MT006483 | MT007006 | - | |
| DAL-23 | 2014 | Inca | Mallorca/Islas Baleares | MT007310 | MT006787 | MT006484 | - | - | |
| DAL-32 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007313 | MT006790 | MT006487 | - | - | |
| DAL-33 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007314 | MT006791 | MT006488 | - | - | |
| DAL-35 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007315 | MT006792 | MT006489 | - | - | |
| DAL-36 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007316 | MT006793 | MT006490 | - | - | |
| DAL-37 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007317 | MT006794 | MT006491 | - | - | |
| DAL-38 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007318 | MT006795 | MT006492 | - | - | |
| DAL-39 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007319 | MT006796 | MT006493 | - | - | |
| DAL-40 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007320 | MT006797 | MT006494 | - | - | |
| DAL-41 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007321 | MT006798 | MT006495 | - | - | |
| DAL-42 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007322 | MT006799 | MT006496 | MT007008 | MT006699 | |
| DAL-43 | 2017 | Bunyola | Mallorca/Islas Baleares | MT007323 | MT006800 | MT006497 | MT007009 | MT006700 | |
| DAL-44 | 2017 | Bunyola | Mallorca/Islas Baleares | MT007324 | MT006801 | MT006498 | - | - | |
| DAL-45 | 2017 | Bunyola | Mallorca/Islas Baleares | MT007325 | MT006802 | MT006499 | MT007010 | MT006701 | |
| DAL-46 | 2017 | Bunyola | Mallorca/Islas Baleares | MT007326 | MT006803 | MT006500 | - | - | |
| DAL-47 | 2017 | Bunyola | Mallorca/Islas Baleares | MT007327 | MT006804 | MT006501 | - | - | |
| DAL-48 | 2017 | Bunyola | Mallorca/Islas Baleares | MT007328 | MT006805 | MT006502 | MT007011 | MT006702 | |
| D. amygdali (cont.) | DAL-49 | 2017 | Bunyola | Mallorca/Islas Baleares | MT007329 | MT006806 | MT006503 | - | - |
| DAL-50 | 2017 | Bunyola | Mallorca/Islas Baleares | MT007330 | MT006807 | MT006504 | MT007012 | - | |
| DAL-51 | 2017 | Bunyola | Mallorca/Islas Baleares | MT007331 | MT006808 | MT006505 | - | - | |
| DAL-52 | 2017 | Palma | Mallorca/Islas Baleares | MT007332 | MT006809 | MT006506 | - | - | |
| DAL-53 | 2017 | Palma | Mallorca/Islas Baleares | MT007333 | MT006810 | MT006507 | - | - | |
| DAL-54 | 2017 | Palma | Mallorca/Islas Baleares | MT007334 | MT006811 | MT006508 | - | - | |
| DAL-55 | 2017 | Palma | Mallorca/Islas Baleares | MT007335 | MT006812 | MT006509 | - | - | |
| DAL-56 | 2017 | Palma | Mallorca/Islas Baleares | MT007336 | MT006813 | MT006510 | - | - | |
| DAL-57 | 2017 | Palma | Mallorca/Islas Baleares | MT007337 | MT006814 | MT006511 | MT007013 | - | |
| DAL-65 | 2017 | La Rinconada | Sevilla/Andalucía | MT007338 | MT006815 | MT006512 | MT007014 | - | |
| DAL-70 | 2018 | Godelleta | Valencia/Comunidad Valenciana | MT007339 | MT006816 | MT006513 | MT007015 | MT006703 | |
| DAL-71 | 2018 | Godelleta | Valencia/Comunidad Valenciana | MT007340 | MT006817 | MT006514 | - | - | |
| DAL-72 | 2018 | Godelleta | Valencia/Comunidad Valenciana | MT007341 | MT006818 | MT006515 | - | - | |
| DAL-73 | 2018 | Godelleta | Valencia/Comunidad Valenciana | MT007342 | MT006819 | MT006516 | - | - | |
| DAL-74 | 2018 | Godelleta | Valencia/Comunidad Valenciana | MT007343 | MT006820 | MT006517 | - | - | |
| DAL-75 | 2018 | Godelleta | Valencia/Comunidad Valenciana | MT007344 | MT006821 | MT006518 | - | - | |
| DAL-76 | 2018 | Montserrat | Valencia/Comunidad Valenciana | MT007345 | MT006822 | MT006519 | MT007016 | MT006704 | |
| DAL-77 | 2018 | Montserrat | Valencia/Comunidad Valenciana | MT007346 | MT006823 | MT006520 | - | - | |
| DAL-78 | 2018 | Montserrat | Valencia/Comunidad Valenciana | MT007347 | MT006824 | MT006521 | - | - | |
| DAL-79 | 2018 | Montserrat | Valencia/Comunidad Valenciana | MT007348 | MT006825 | MT006522 | - | - | |
| DAL-80 | 2018 | Montserrat | Valencia/Comunidad Valenciana | MT007349 | MT006826 | MT006523 | - | - | |
| DAL-81 | 2018 | Montserrat | Valencia/Comunidad Valenciana | MT007350 | MT006827 | MT006524 | - | - | |
| DAL-82 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007351 | MT006828 | MT006525 | MT007017 | MT006705 | |
| DAL-83 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007352 | MT006829 | MT006526 | - | - | |
| DAL-84 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007353 | MT006830 | MT006527 | - | - | |
| DAL-85 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007354 | MT006831 | MT006528 | MT007018 | MT006706 | |
| DAL-86 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007355 | MT006832 | MT006529 | - | - | |
| DAL-87 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007356 | MT006833 | MT006530 | - | - | |
| DAL-88 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007357 | MT006834 | MT006531 | - | - | |
| DAL-89 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007358 | MT006835 | MT006532 | - | - | |
| DAL-90 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007359 | MT006836 | MT006533 | - | - | |
| DAL-91 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007360 | MT006837 | MT006534 | - | - | |
| DAL-92 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007361 | MT006838 | MT006535 | - | - | |
| DAL-93 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007362 | MT006839 | MT006536 | - | - | |
| DAL-94 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007363 | MT006840 | MT006537 | MT007019 | - | |
| D. amygdali (cont.) | DAL-95 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007364 | MT006841 | MT006538 | MT007020 | - |
| DAL-96 | 2018 | Viver | Castellón/Comunidad Valenciana | MT007365 | MT006842 | MT006539 | - | - | |
| DAL-97 | 2018 | Fuente la Higuera | Valencia/Comunidad Valenciana | MT007366 | MT006843 | MT006540 | - | - | |
| DAL-98 | 2018 | Fuente la Higuera | Valencia/Comunidad Valenciana | MT007367 | MT006844 | MT006541 | - | - | |
| DAL-103 | 2017 | Gibraleón | Huelva/Andalucía | MT007368 | MT006845 | MT006542 | MT007021 | MT006707 | |
| DAL-104 | 2016 | El Contador | Almería/Andalucía | MT007369 | MT006848 | MT006543 | MT007022 | MT006708 | |
| DAL-105 | 2017 | Alcalá del Río | Sevilla/Andalucía | MT007370 | MT006846 | MT006544 | MT007023 | MT006709 | |
| DAL-108 | 2018 | Biar | Alicante/Comunidad Valenciana | MT007371 | MT006847 | MT006545 | MT007024 | MT006710 | |
| DAL-109 | 2018 | Biar | Alicante/Comunidad Valenciana | MT007372 | MT006849 | MT006546 | - | - | |
| DAL-110 | 2018 | Fuente la Higuera | Valencia/Comunidad Valenciana | MT007373 | MT006850 | MT006547 | - | - | |
| DAL-111 | 2018 | Fuente la Higuera | Valencia/Comunidad Valenciana | MT007374 | MT006851 | MT006548 | - | - | |
| DAL-112 | 2018 | Fuente la Higuera | Valencia/Comunidad Valenciana | MT007375 | MT006852 | MT006549 | - | - | |
| DAL-113 | 2018 | Fontanars dels Alforins | Valencia/Comunidad Valenciana | MT007376 | MT006853 | MT006550 | MT007025 | MT006711 | |
| DAL-114 | 2018 | Fontanars dels Alforins | Valencia/Comunidad Valenciana | MT007377 | MT006854 | MT006551 | MT007026 | MT006712 | |
| DAL-116 | 2018 | Alcublas | Valencia/Comunidad Valenciana | MT007378 | MT006855 | MT006552 | MT007027 | - | |
| DAL-117 | 2018 | Alcublas | Valencia/Comunidad Valenciana | MT007379 | MT006856 | MT006553 | - | - | |
| DAL-118 | 2018 | Casinos | Valencia/Comunidad Valenciana | MT007380 | MT006857 | MT006554 | - | - | |
| DAL-119 | 2018 | Casinos | Valencia/Comunidad Valenciana | MT007381 | MT006858 | MT006555 | - | - | |
| DAL-120 | 2018 | Casinos | Valencia/Comunidad Valenciana | MT007382 | MT006859 | MT006556 | - | - | |
| DAL-121 | 2018 | Vall d’Alba | Castellón/Comunidad Valenciana | MT007383 | MT006860 | MT006557 | MT007028 | - | |
| DAL-122 | 2018 | Vall d’Alba | Castellón/Comunidad Valenciana | MT007384 | MT006861 | MT006558 | - | - | |
| DAL-125 | 2018 | Vall d’Alba | Castellón/Comunidad Valenciana | MT007385 | MT006862 | MT006559 | - | - | |
| DAL-126 | 2018 | Vall d’Alba | Castellón/Comunidad Valenciana | MT007386 | MT006863 | MT006560 | - | - | |
| DAL-128 | 2018 | Godelleta | Valencia/Comunidad Valenciana | MT007387 | MT006864 | MT006561 | - | - | |
| DAL-129 | 2018 | Godelleta | Valencia/Comunidad Valenciana | MT007388 | MT006865 | MT006562 | - | - | |
| DAL-130 | 2018 | Torremendo | Alicante/Comunidad Valenciana | MT007389 | MT006866 | MT006563 | MT007029 | - | |
| DAL-131 | 2018 | Torremendo | Alicante/Comunidad Valencian | MT007390 | MT006867 | MT006564 | - | - | |
| DAL-132 | 2018 | Requena | Valencia/Comunidad Valenciana | MT007391 | MT006868 | MT006565 | MT007030 | MT006713 | |
| DAL-133 | 2018 | Requena | Valencia/Comunidad Valenciana | MT007392 | MT006869 | MT006566 | MT007031 | - | |
| DAL-134 | 2018 | Requena | Valencia/Comunidad Valenciana | MT007393 | MT006870 | MT006567 | MT007032 | - | |
| DAL-135 | 2018 | L’Eliana | Valencia/Comunidad Valenciana | MT007394 | MT006871 | MT006568 | MT007033 | - | |
| DAL-136 | 2018 | L’Eliana | Valencia/Comunidad Valenciana | MT007395 | MT006872 | MT006569 | - | - | |
| DAL-138 | 2005 | Constantí | Tarragona/Cataluña | MT007396 | MT006873 | MT006570 | - | - | |
| DAL-139 | 2005 | Constantí | Tarragona/Cataluña | MT007397 | MT006874 | MT006571 | MT007034 | - | |
| DAL-140 | 2012 | Ulldecona | Tarragona/Cataluña | MT007398 | MT006875 | MT006572 | MT007035 | MT006714 | |
| D. amygdali (cont.) | DAL-141 | 2016 | Gandesa | Tarragona/Cataluña | MT007399 | MT006876 | MT006573 | - | - |
| DAL-143 | 2018 | Gandesa | Tarragona/Cataluña | MT007400 | MT006877 | MT006574 | - | - | |
| DAL-144 | 2018 | Gandesa | Tarragona/Cataluña | MT007401 | MT006878 | MT006575 | - | - | |
| DAL-145 | 2018 | Gandesa | Tarragona/Cataluña | MT007402 | MT006879 | MT006576 | - | - | |
| DAL-146 | 2018 | Gandesa | Tarragona/Cataluña | MT007403 | MT006880 | MT006577 | MT007036 | - | |
| DAL-147 | 2018 | Constantí | Tarragona/Cataluña | MT007404 | MT006881 | MT006578 | MT007037 | - | |
| DAL-148 | 2018 | Constantí | Tarragona/Cataluña | MT007405 | MT006882 | MT006579 | MT007038 | - | |
| DAL-149 | 2018 | Constantí | Tarragona/Cataluña | MT007406 | MT006883 | MT006580 | MT007039 | MT006715 | |
| DAL-151 | 2018 | Constantí | Tarragona/Cataluña | MT007407 | MT006884 | MT006581 | - | - | |
| DAL-152 | 2018 | Constantí | Tarragona/Cataluña | MT007408 | MT006885 | MT006582 | MT007040 | MT006716 | |
| DAL-153 | 2018 | Constantí | Tarragona/Cataluña | MT007409 | MT006886 | MT006583 | - | - | |
| DAL-154 | 2018 | La Selva del Camp | Tarragona/Cataluña | MT007410 | MT006887 | MT006584 | MT007041 | MT006717 | |
| DAL-155 | 2018 | La Selva del Camp | Tarragona/Cataluña | MT007411 | MT006888 | MT006585 | MT007042 | MT006718 | |
| DAL-156 | 2018 | La Selva del Camp | Tarragona/Cataluña | MT007412 | MT006889 | MT006586 | - | - | |
| DAL-158 | 2018 | La Selva del Camp | Tarragona/Cataluña | MT007413 | MT006890 | MT006587 | - | - | |
| DAL-159 | 2018 | La Selva del Camp | Tarragona/Cataluña | MT007414 | MT006891 | MT006588 | - | - | |
| DAL-160 | 2018 | La Selva del Camp | Tarragona/Cataluña | MT007415 | MT006892 | MT006589 | - | - | |
| DAL-161 | 2018 | Constantí | Tarragona/Cataluña | MT007416 | MT006893 | MT006590 | - | - | |
| DAL-162 | 2018 | Constantí | Tarragona/Cataluña | MT007417 | MT006894 | MT006591 | - | - | |
| DAL-163 | 2018 | Estepa | Sevilla/Andalucía | MT007418 | MT006895 | MT006592 | - | - | |
| DAL-164 | 2018 | Estepa | Sevilla/Andalucía | MT007419 | MT006896 | MT006593 | MT007043 | MT006719 | |
| DAL-167 | 2018 | Los Palacios | Sevilla/Andalucía | MT007420 | MT006897 | MT006594 | MT007044 | MT006720 | |
| DAL-168 | 2018 | Los Palacios | Sevilla/Andalucía | MT007421 | MT006898 | MT006595 | - | - | |
| DAL-169 | 2018 | Los Palacios | Sevilla/Andalucía | MT007422 | MT006899 | MT006596 | - | - | |
| DAL-170 | 2018 | Los Palacios | Sevilla/Andalucía | MT007423 | MT006900 | MT006597 | - | - | |
| DAL-171 | 2018 | Los Palacios | Sevilla/Andalucía | MT007424 | MT006901 | MT006598 | - | - | |
| DAL-172 | 2018 | Los Palacios | Sevilla/Andalucía | MT007425 | MT006902 | MT006599 | MT007045 | - | |
| DAL-181 | 2018 | Córdoba | Córdoba/Andalucía | MT007426 | MT006903 | MT006600 | MT007046 | MT006721 | |
| DAL-182 | 2018 | Córdoba | Córdoba/Andalucía | MT007427 | MT006904 | MT006601 | - | - | |
| DAL-183 | 2018 | Córdoba | Córdoba/Andalucía | MT007428 | MT006905 | MT006602 | - | - | |
| DAL-184 | 2018 | Mairena del Alcor | Sevilla/Andalucía | MT007429 | MT006906 | MT006603 | MT007047 | - | |
| DAL-185 | 2018 | Mairena del Alcor | Sevilla/Andalucía | MT007430 | MT006907 | MT006604 | - | - | |
| DAL-186 | 2018 | Mairena del Alcor | Sevilla/Andalucía | MT007431 | MT006908 | MT006605 | - | - | |
| DAL-187 | 2018 | Mairena del Alcor | Sevilla/Andalucía | MT007432 | MT006909 | MT006606 | - | - | |
| D. amygdali (cont.) | DAL-188 | 2018 | Mairena del Alcor | Sevilla/Andalucía | MT007433 | MT006910 | MT006607 | - | - |
| DAL-189 | 2018 | Mairena del Alcor | Sevilla/Andalucía | MT007434 | MT006911 | MT006608 | MT007048 | - | |
| DAL-190 | 2018 | Mairena del Alcor | Sevilla/Andalucía | MT007435 | MT006912 | MT006609 | MT007049 | MT006722 | |
| DAL-191 | 2018 | Mairena del Alcor | Sevilla/Andalucía | MT007436 | MT006913 | MT006610 | - | - | |
| DAL-192 | 2018 | Mairena del Alcor | Sevilla/Andalucía | MT007437 | MT006914 | MT006611 | - | - | |
| DAL-193 | 2018 | Ronda | Málaga/Andalucía | MT007438 | MT006915 | MT006612 | MT007050 | MT006723 | |
| DAL-194 | 2018 | Ronda | Málaga/Andalucía | MT007439 | MT006916 | MT006613 | - | - | |
| DAL-195 | 2018 | Ronda | Málaga/Andalucía | MT007440 | MT006917 | MT006614 | - | - | |
| DAL-196 | 2018 | Ronda | Málaga/Andalucía | MT007441 | MT006918 | MT006615 | - | - | |
| DAL-197 | 2018 | Ronda | Málaga/Andalucía | MT007442 | MT006919 | MT006616 | MT007051 | MT006724 | |
| DAL-198 | 2018 | Ronda | Málaga/Andalucía | MT007443 | MT006920 | MT006617 | - | - | |
| DAL-199 | 2018 | Ronda | Málaga/Andalucía | MT007444 | MT006921 | MT006618 | - | - | |
| DAL-200 | 2018 | Ronda | Málaga/Andalucía | MT007445 | MT006922 | MT006619 | - | - | |
| DAL-201 | 2018 | Ronda | Málaga/Andalucía | MT007446 | MT006923 | MT006620 | - | - | |
| DAL-202 | 2018 | Ronda | Málaga/Andalucía | MT007447 | MT006924 | MT006621 | MT007052 | - | |
| DAL-203 | 2018 | Reus | Tarragona/Cataluña | MT007448 | MT006925 | MT006622 | - | - | |
| DAL-204 | 2018 | Reus | Tarragona/Cataluña | MT007449 | MT006926 | MT006623 | MT007053 | MT006725 | |
| DAL-205 | 2018 | Reus | Tarragona/Cataluña | MT007450 | MT006927 | MT006624 | MT007054 | MT006726 | |
| DAL-206 | 2018 | Riudoms | Tarragona/Cataluña | MT007451 | MT006928 | MT006625 | - | - | |
| DAL-207 | 2018 | Riudoms | Tarragona/Cataluña | MT007452 | MT006929 | MT006626 | - | - | |
| DAL-208 | 2018 | Riudoms | Tarragona/Cataluña | MT007453 | MT006930 | MT006627 | MT007055 | - | |
| DAL-209 | 2018 | Riudoms | Tarragona/Cataluña | MT007454 | MT006931 | MT006628 | MT007056 | - | |
| DAL-210 | 2018 | Riudoms | Tarragona/Cataluña | MT007455 | MT006932 | MT006629 | MT007057 | - | |
| DAL-211 | 2018 | Riudoms | Tarragona/Cataluña | MT007456 | MT006933 | MT006630 | - | - | |
| DAL-212 | 2018 | Riudoms | Tarragona/Cataluña | MT007457 | MT006934 | MT006631 | - | - | |
| DAL-213 | 2018 | Riudoms | Tarragona/Cataluña | MT007458 | MT006935 | MT006632 | - | - | |
| DAL-214 | 2018 | Botarell | Tarragona/Cataluña | MT007459 | MT006936 | MT006633 | - | - | |
| DAL-215 | 2018 | Botarell | Tarragona/Cataluña | MT007460 | MT006937 | MT006634 | MT007058 | - | |
| DAL-216 | 2018 | Botarell | Tarragona/Cataluña | MT007461 | MT006938 | MT006635 | MT007059 | - | |
| DAL-219 | 2018 | Les Borges Blanques | Lérida/Cataluña | MT007462 | MT006939 | MT006636 | MT007060 | MT006727 | |
| DAL-220 | 2018 | Isona i Conca Dellà | Lérida/Cataluña | MT007463 | MT006940 | MT006637 | MT007061 | MT006728 | |
| DAL-221 | 2018 | Isona i Conca Dellà | Lérida/Cataluña | MT007464 | MT006941 | MT006638 | MT007062 | - | |
| DAL-225 | 2019 | Murillo | Logroño/La Rioja | MT007465 | MT006942 | MT006639 | MT007063 | MT006729 | |
| DAL-226 | 2019 | Murillo | Logroño/La Rioja | MT007466 | MT006943 | MT006640 | - | - | |
| D. amygdali (cont.) | DAL-227 | 2019 | Santa Engracia de Jubera | Logroño/La Rioja | MT007467 | MT006944 | MT006641 | MT007064 | MT006730 |
| DAL-228 | 2019 | Santa Engracia de Jubera | Logroño/La Rioja | MT007468 | MT006945 | MT006642 | - | - | |
| DAL-229 | 2019 | Santa Engracia de Jubera | Logroño/La Rioja | MT007469 | MT006946 | MT006643 | - | - | |
| DAL-230 | 2019 | Santa Engracia de Jubera | Logroño/La Rioja | MT007470 | MT006947 | MT006644 | - | - | |
| DAL-231 | 2019 | Santa Engracia de Jubera | Logroño/La Rioja | MT007471 | MT006948 | MT006645 | - | - | |
| DAL-232 | 2019 | Santa Engracia de Jubera | Logroño/La Rioja | MT007472 | MT006949 | MT006646 | - | - | |
| DAL-233 | 2019 | Lagunilla | Logroño/La Rioja | MT007473 | MT006950 | MT006647 | MT007065 | MT006731 | |
| DAL-234 | 2019 | Santa Engracia de Jubera | Logroño/La Rioja | MT007474 | MT006951 | MT006648 | - | - | |
| DAL-236 | 2019 | Alcalá del Río | Sevilla/Andalucía | MT007475 | MT006952 | MT006649 | MT007066 | - | |
| DAL-237 | 2019 | Alcalá del Río | Sevilla/Andalucía | MT007476 | MT006953 | MT006650 | - | - | |
| DAL-238 | 2019 | Alcalá del Río | Sevilla/Andalucía | MT007477 | MT006954 | MT006651 | - | - | |
| DAL-239 | 2019 | Córdoba | Córdoba/Andalucía | MT007478 | MT006955 | MT006652 | - | - | |
| DAL-240 | 2019 | Córdoba | Córdoba/Andalucía | MT007479 | MT006956 | MT006653 | MT007067 | MT006732 | |
| DAL-241 | 2019 | Córdoba | Córdoba/Andalucía | MT007480 | MT006957 | MT006654 | - | - | |
| DAL-242 | 2019 | Santa Cruz | Córdoba/Andalucía | MT007481 | MT006958 | MT006655 | - | - | |
| DAL-243 | 2019 | Córdoba | Córdoba/Andalucía | MT007482 | MT006959 | MT006656 | - | - | |
| DAL-244 | 2019 | Villamanrique de la Condesa | Sevilla/Andalucía | MT007483 | MT006960 | MT006657 | MT007068 | MT006733 | |
| DAL-245 | 2019 | Villamanrique de la Condesa | Sevilla/Andalucía | MT007484 | MT006961 | MT006658 | - | - | |
| DAL-246 | 2019 | Santa Engracia de Jubera | Logroño/La Rioja | MT007485 | MT006962 | MT006659 | - | - | |
| D. eres | DAL-102 | 2016 | Córdoba | Córdoba/Andalucía | MN997106 | MT007104 | MT006462 | MT007106 | MT006465 |
| D. foeniculina | DAL-10 | 2014 | Santa Margalida i Calvià | Mallorca/Islas Baleares | MT007497 | MT006963 | MT006660 | MT007069 | MT006734 |
| DAL-11 | 2014 | Mallorca/Islas Baleares | MT007498 | MT006964 | MT006661 | MT007070 | MT006735 | ||
| DAL-27 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007499 | MT006965 | MT006662 | MT007071 | MT006736 | |
| DAL-28 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007500 | MT006966 | MT006663 | MT007072 | MT006737 | |
| DAL-30 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007501 | MT006967 | MT006664 | MT007073 | MT006738 | |
| DAL-31 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007502 | MT006968 | MT006665 | MT007074 | MT006739 | |
| DAL-61 | 2016 | Alcalá del Río | Sevilla/Andalucía | MT007503 | MT006969 | MT006666 | MT007075 | MT006740 | |
| DAL-62 | 2016 | Alcalá del Río | Sevilla/Andalucía | MT007504 | MT006970 | MT006667 | MT007076 | MT006741 | |
| DAL-63 | 2016 | Alcalá del Río | Sevilla/Andalucía | MT007505 | MT006971 | MT006668 | MT007077 | MT006742 | |
| DAL-64 | 2016 | Alcalá del Río | Sevilla/Andalucía | MT007506 | MT006972 | MT006669 | MT007078 | MT006743 | |
| DAL-66 | 2017 | La Rinconada | Sevilla/Andalucía | MT007507 | MT006973 | MT006670 | MT007079 | MT006744 | |
| DAL-67 | 2017 | La Rinconada | Sevilla/Andalucía | MT007508 | MT006974 | MT006671 | MT007080 | MT006745 | |
| DAL-68 | 2017 | La Rinconada | Sevilla/Andalucía | MT007509 | MT006975 | MT006672 | MT007081 | MT006746 | |
| DAL-69 | 2017 | La Rinconada | Sevilla/Andalucía | MT007510 | MT006976 | MT006673 | MT007082 | MT006747 | |
| DAL-99 | 2018 | Fuente la Higuera | Valencia/Comunidad Valenciana | MT007511 | MT006977 | MT006674 | MT007083 | MT006748 | |
| DAL-100 | 2018 | Fuente la Higuera | Valencia/Comunidad Valenciana | MT007512 | MT006978 | MT006675 | MT007084 | MT006749 | |
| D. foeniculina (cont.) | DAL-101 | 2018 | Fuente la Higuera | Valencia/Comunidad Valenciana | MT007513 | MT006979 | MT006676 | MT007085 | MT006750 |
| DAL-107 | 2018 | Marchena | Sevilla/Andalucía | MT007514 | MT006980 | MT006677 | MT007086 | MT006751 | |
| DAL-142 | 2018 | Cabrils | Barcelona/Cataluña | MT007515 | MT006981 | MT006678 | MT007087 | MT006752 | |
| DAL-150 | 2018 | Constantí | Tarragona/Cataluña | MT007516 | MT006982 | MT006679 | MT007088 | MT006753 | |
| DAL-157 | 2018 | La Selva del Camp | Tarragona/Cataluña | MT007517 | MT006983 | MT006680 | MT007089 | MT006754 | |
| DAL-165 | 2018 | Estepa | Sevilla/Andalucía | MT007518 | MT006984 | MT006681 | MT007090 | MT006755 | |
| DAL-217 | 2018 | Les Borges Blanques | Lérida/Cataluña | MT007519 | MT006985 | MT006682 | MT007091 | MT006756 | |
| D. mediterranea | DAL-6 | 2014 | Calvià | Mallorca/Islas Baleares | MT007486 | MT006986 | MT006683 | MT007092 | MT006758 |
| DAL-8 | 2014 | Consell | Mallorca/Islas Baleares | MT007487 | MT006987 | MT006684 | MT007093 | MT006759 | |
| DAL-24 | 2014 | Sant Llorenç d’Escardassar | Mallorca/Islas Baleares | MT007488 | MT006988 | MT006685 | MT007094 | MT006760 | |
| DAL-34 | 2017 | Alcalalí | Alicante/Comunidad Valenciana | MT007489 | MT006989 | MT006686 | MT007095 | MT006761 | |
| DAL-173 | 2018 | Altea la Vella | Alicante/Comunidad Valenciana | MT007493 | MT006993 | MT006691 | MT007099 | MT006765 | |
| DAL-174 | 2018 | Altea la Vella | Alicante/Comunidad Valenciana | MT007494 | MT006994 | MT006690 | MT007100 | MT006766 | |
| DAL-175 | 2018 | Altea la Vella | Alicante/Comunidad Valenciana | MT007495 | MT006995 | MT006692 | MT007101 | MT006767 | |
| DAL-176 | 2018 | Altea la Vella | Alicante/Comunidad Valenciana | MT007496 | MT006996 | MT006693 | MT007102 | MT006768 | |
| D. phaseolorum | DAL-222 | 2016 | Alcalá del Río | Sevilla/Andalucía | MN997107 | MT007103 | MT006463 | MT007105 | MT006464 |
| Species | Strain | Host | Country | GenBank Accession Numbers | ||||
|---|---|---|---|---|---|---|---|---|
| ITS | tef-1α | tub | his | cal | ||||
| D. acaciigena | CBS 129521 | Acacia retinodes | Australia | KC343005 | KC343731 | KC343973 | KC343489 | KC343247 |
| D. amygdali | CBS 126679 | Prunus dulcis | Portugal | KC343022 | KC343748 | KC343990 | KC343506 | KC343264 |
| CBS 111811 | Vitis vinifera | South Africa | KC343019 | KC343745 | KC343987 | KC343503 | KC343261 | |
| D. celastrina | CBS 139.27 | Celastrus scandens | USA | KC343047 | KC343773 | KC344015 | KC343531 | KC343289 |
| D. celeris | CBS 143349 | Vitis vinifera | UK | MG281017 | MG281538 | MG281190 | MG281363 | MG281712 |
| CBS 143350 | Vitis vinifera | UK | MG281018 | MG281539 | MG281191 | MG281364 | MG281713 | |
| D. chamaeropis | CBS 454.81 | Chamaerops humilis | Greece | KC343048 | KC343774 | KC344016 | KC343532 | KC343290 |
| CBS 753.70 | Spartium junceum | Croatia | KC343049 | KC343775 | KC344017 | KC343533 | KC343291 | |
| D. chongqingensis | PSCG 435 | Pyrus pyrifolia | China | MK626916 | MK654866 | MK691321 | MK726257 | MK691209 |
| PSCG 436 | Pyrus pyrifolia | China | MK626917 | MK654867 | MK691322 | MK726256 | MK691208 | |
| D. cinerascens | CBS 719.96 | Ficus carica | Bulgaria | KC343050 | KC343776 | KC344018 | KC343534 | KC343292 |
| D. endophytica | CBS 133811 | Schinus terebinthifolius | Brazil | KC343065 | KC343791 | KC344033 | KC343549 | KC343307 |
| LGMF911 | Schinus terebinthifolius | Brazil | KC343066 | KC343792 | KC344034 | KC343550 | KC343308 | |
| D. eres | CBS 138594 | Ulmus laevis | Germany | KJ210529 | KJ210550 | KJ420799 | KJ420850 | KJ434999 |
| CBS 109767 | Acer campestre | Austria | KC343075 | KC343801 | KC344043 | KC343559 | KC343317 | |
| D. foeniculina | CBS 111553 | Foeniculum vulgare | Spain | KC343101 | KC343827 | KC344069 | KC343585 | KC343343 |
| CBS 187.27 | Camellia sinensis | Italy | KC343107 | KC343833 | KC344075 | KC343591 | KC343349 | |
| D. fusicola | CGMCC 3.17087 | Lithocarpus glabra | China | KF576281 | KF576256 | KF576305 | - | KF576233 |
| CGMCC 3.17088 | Lithocarpus glabra | China | KF576263 | KF576238 | KF576287 | - | KF576221 | |
| D. garethjonesii | MFLUCC 12-0542A | Unknown dead leaf | Thailand | KT459423 | KT459457 | KT459441 | - | KT459470 |
| D. helicis | CBS 138596 | Hederahelix | Germany | KJ210538 | KJ210559 | KJ420828 | KJ420875 | KJ435043 |
| D. kadsurae | CFCC 52586 | Kadsura longipedunculata | China | MH121521 | MH121563 | MH121600 | MH121479 | MH121439 |
| CFCC 52587 | Kadsura longipedunculata | China | MH121522 | MH121564 | MH121601 | MH121480 | MH121440 | |
| D. masirevicii | BRIP 54120c | Zea mays | Australia | KJ197278 | KJ197240 | KJ197258 | - | - |
| BRIP 57892a | Helianthus annuus | Australia | KJ197276 | KJ197239 | KJ197257 | - | - | |
| D. ovalispora | ICMP20659 | Citrus limon | China | KJ490628 | KJ490507 | KJ490449 | KJ490570 | - |
| D. ovoicicola | CGMCC 3.17092 | Lithocarpus glabra | China | KF576264 | KF576239 | KF576288 | - | KF576222 |
| CGMCC 3.17093 | Citrus sp. | China | KF576265 | KF576240 | KF576289 | - | KF576223 | |
| D. phaseolorum | CBS 113425 | Olearia cf. rani | New Zealand | KC343174 | KC343900 | KC344142 | KC343658 | KC343416 |
| CBS 116019 | Caperonia palustris | USA | KC343175 | KC343901 | KC344143 | KC343659 | KC343417 | |
| D. pulla | CBS 338.89 | Hedera helix | Croatia | KC343152 | KC343878 | KC344120 | KC343636 | KC343394 |
| D. pustulata | CBS 109742 | Acer pseudoplatanus | Austria | KC343185 | KC343911 | KC344153 | KC343669 | KC343427 |
| CBS 109784 | Prunus padus | Austria | KC343187 | KC343913 | KC344155 | KC343671 | KC343429 | |
| D. sojae | CBS 100.87 | Glycine soja | Italy | KC343196 | KC343922 | KC344164 | KC343680 | KC343438 |
| CBS 116017 | Euphorbia nutans | USA | KC343197 | KC343923 | KC344165 | KC343681 | KC343439 | |
| D. sterilis | CBS 136969 | Vaccinium corymbosum | Italy | KJ160579 | KJ160611 | KJ160528 | MF418350 | KJ160548 |
| CBS 136970 | Vaccinium corymbosum | Italy | KJ160580 | KJ160612 | KJ160529 | - | KJ160549 | |
| D. subellipicola | MFLUCC 17-1197 | On dead wood | China | MG746632 | MG746633 | MG746634 | - | - |
| Diaporthella corylina | CBS 121124 | Corylus sp. | China | KC343004 | KC343730 | KC343972 | KC343488 | KC343246 |
| Phomopsis sp. 5 | PMM1657 | Vitis vinifera | South Africa | KY511331 | - | KY511363 | - | - |
| PMM1660 | Vitis vinifera | South Africa | KY511333 | - | KY511365 | - | - | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
León, M.; Berbegal, M.; Rodríguez-Reina, J.M.; Elena, G.; Abad-Campos, P.; Ramón-Albalat, A.; Olmo, D.; Vicent, A.; Luque, J.; Miarnau, X.; et al. Identification and Characterization of Diaporthe spp. Associated with Twig Cankers and Shoot Blight of Almonds in Spain. Agronomy 2020, 10, 1062. https://doi.org/10.3390/agronomy10081062
León M, Berbegal M, Rodríguez-Reina JM, Elena G, Abad-Campos P, Ramón-Albalat A, Olmo D, Vicent A, Luque J, Miarnau X, et al. Identification and Characterization of Diaporthe spp. Associated with Twig Cankers and Shoot Blight of Almonds in Spain. Agronomy. 2020; 10(8):1062. https://doi.org/10.3390/agronomy10081062
Chicago/Turabian StyleLeón, Maela, Mónica Berbegal, José M. Rodríguez-Reina, Georgina Elena, Paloma Abad-Campos, Antonio Ramón-Albalat, Diego Olmo, Antonio Vicent, Jordi Luque, Xavier Miarnau, and et al. 2020. "Identification and Characterization of Diaporthe spp. Associated with Twig Cankers and Shoot Blight of Almonds in Spain" Agronomy 10, no. 8: 1062. https://doi.org/10.3390/agronomy10081062
APA StyleLeón, M., Berbegal, M., Rodríguez-Reina, J. M., Elena, G., Abad-Campos, P., Ramón-Albalat, A., Olmo, D., Vicent, A., Luque, J., Miarnau, X., Agustí-Brisach, C., Trapero, A., Capote, N., Arroyo, F. T., Avilés, M., Gramaje, D., Andrés-Sodupe, M., & Armengol, J. (2020). Identification and Characterization of Diaporthe spp. Associated with Twig Cankers and Shoot Blight of Almonds in Spain. Agronomy, 10(8), 1062. https://doi.org/10.3390/agronomy10081062










