Effects of Exogenous Spermidine on Root Metabolism of Cucumber Seedlings under Salt Stress by GC-MS
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Stress Treatments
2.2. Determination of Plant Growth
2.3. Metabolite Extraction
2.4. GC-MS Analysis and QC
2.5. Statistical Analysis
3. Results
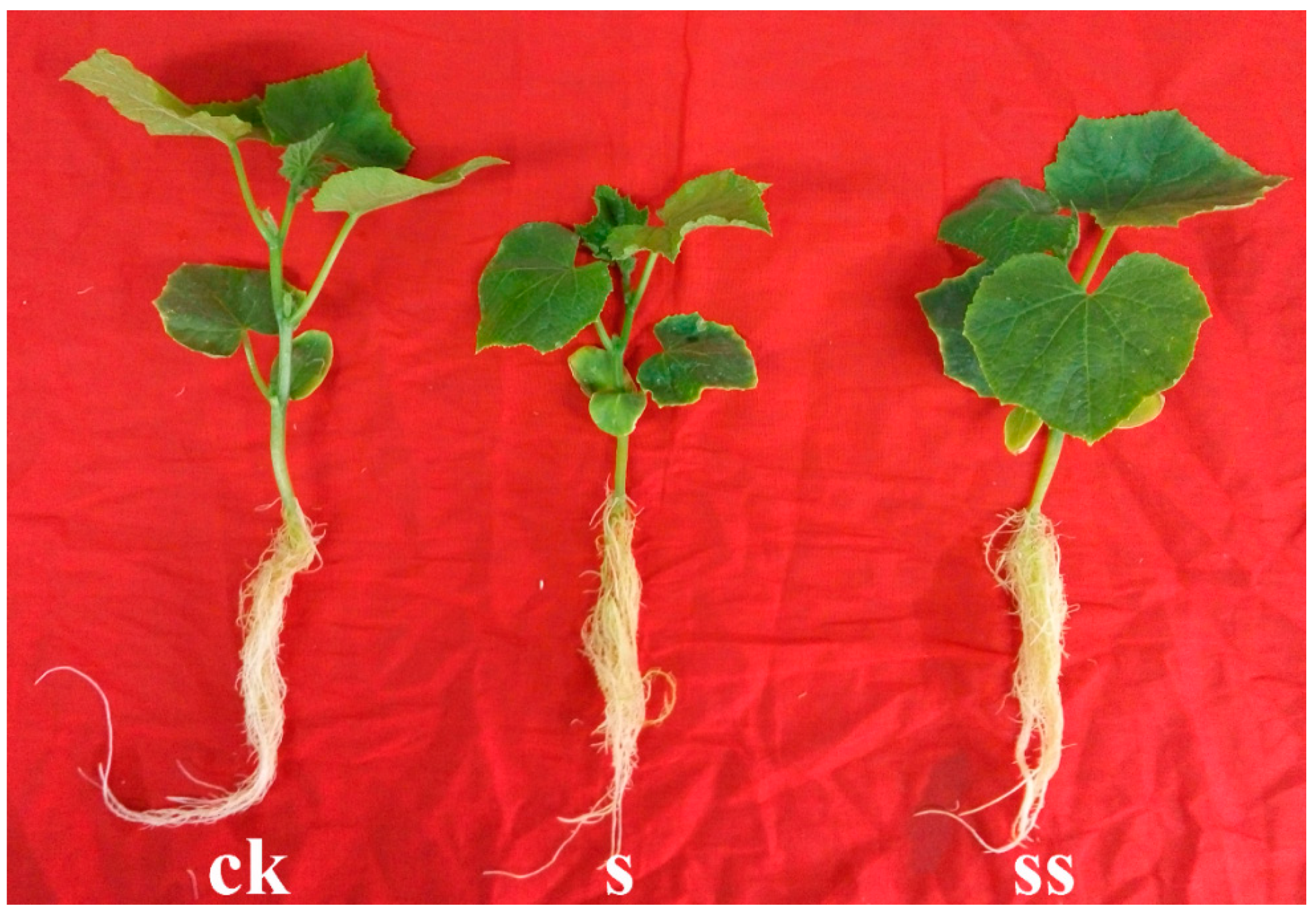
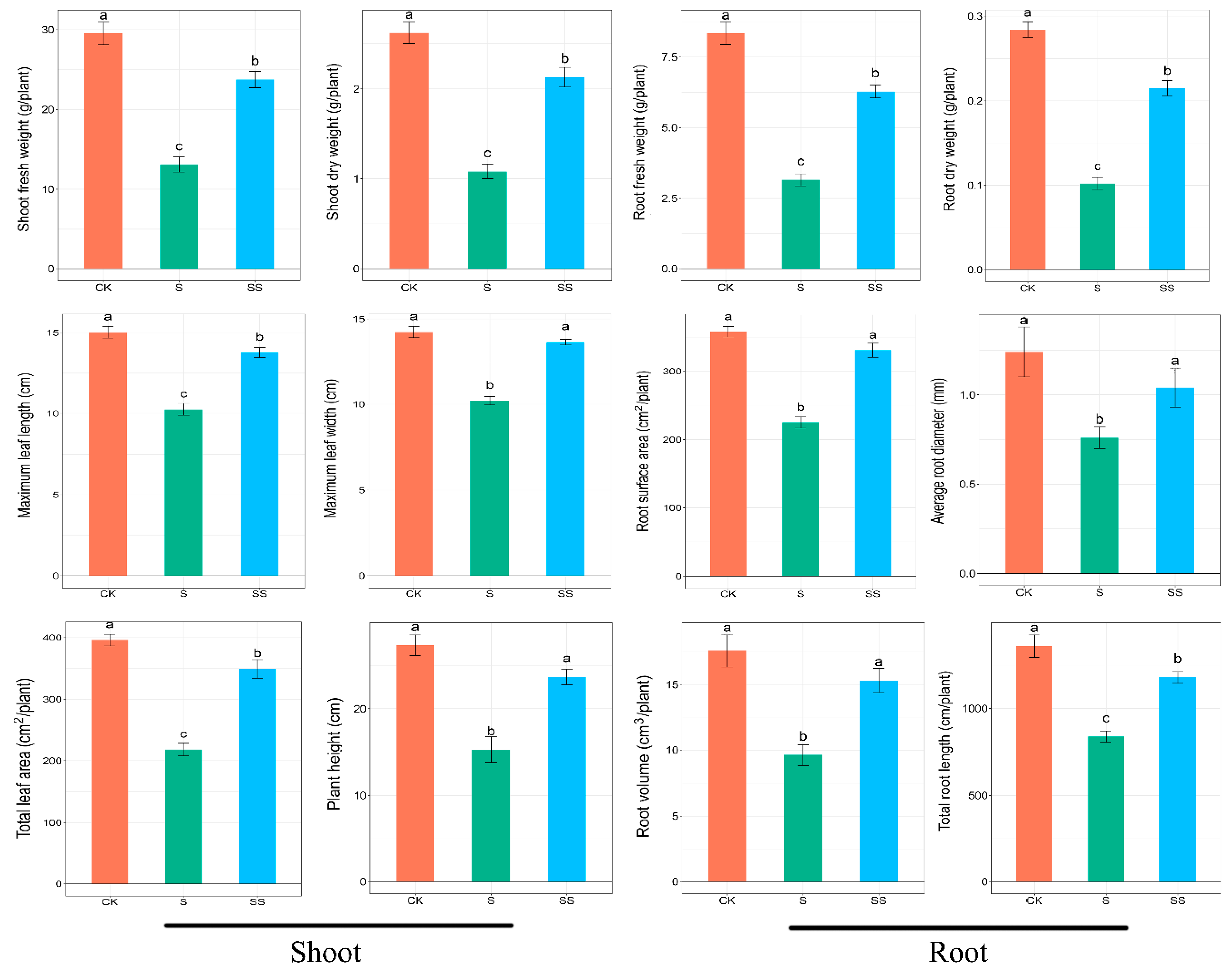
3.1. Growth Parameters of Cucumber Seedlings
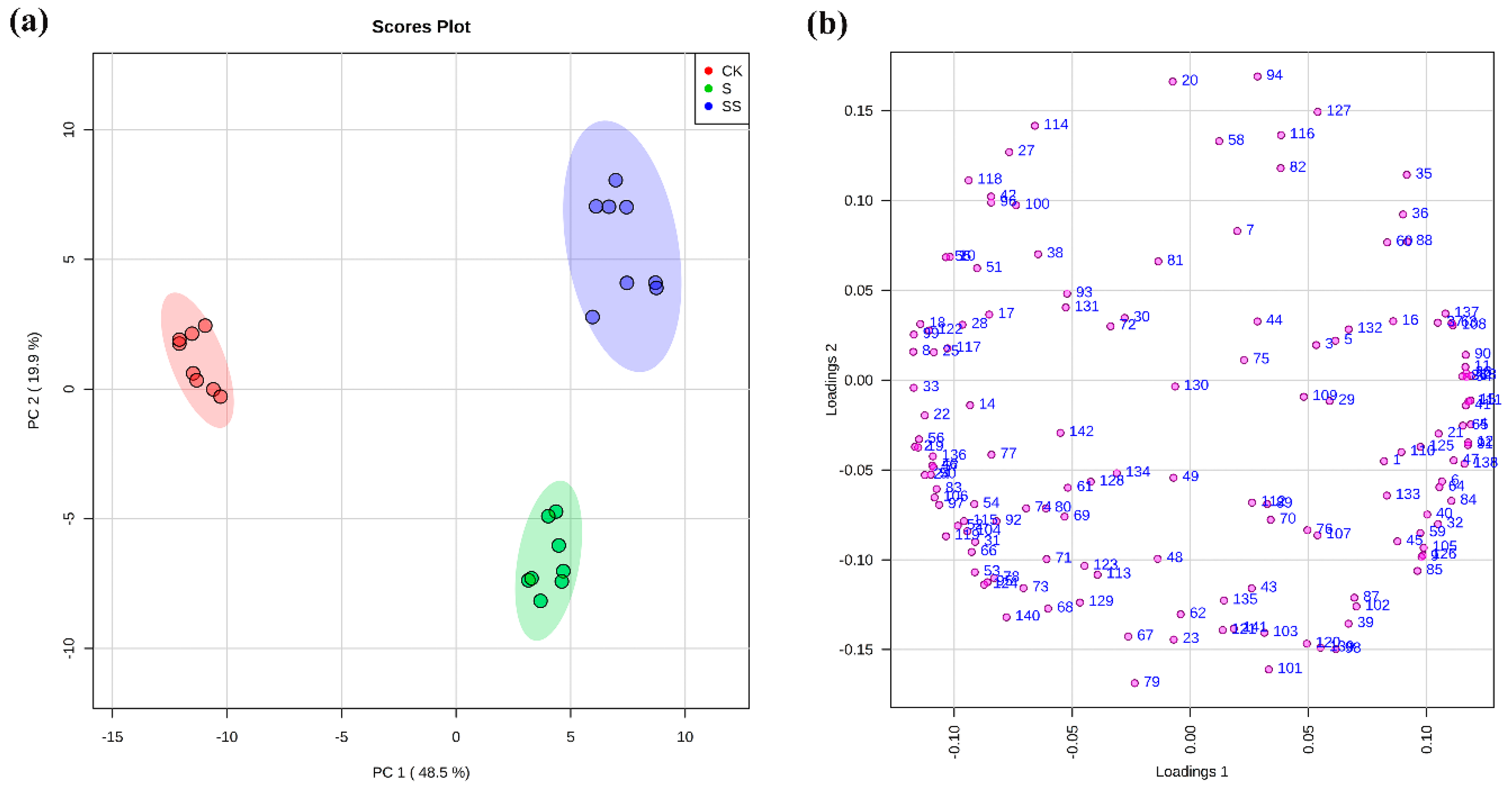
3.2. Metabolic Changes in Cucumber Seedlings
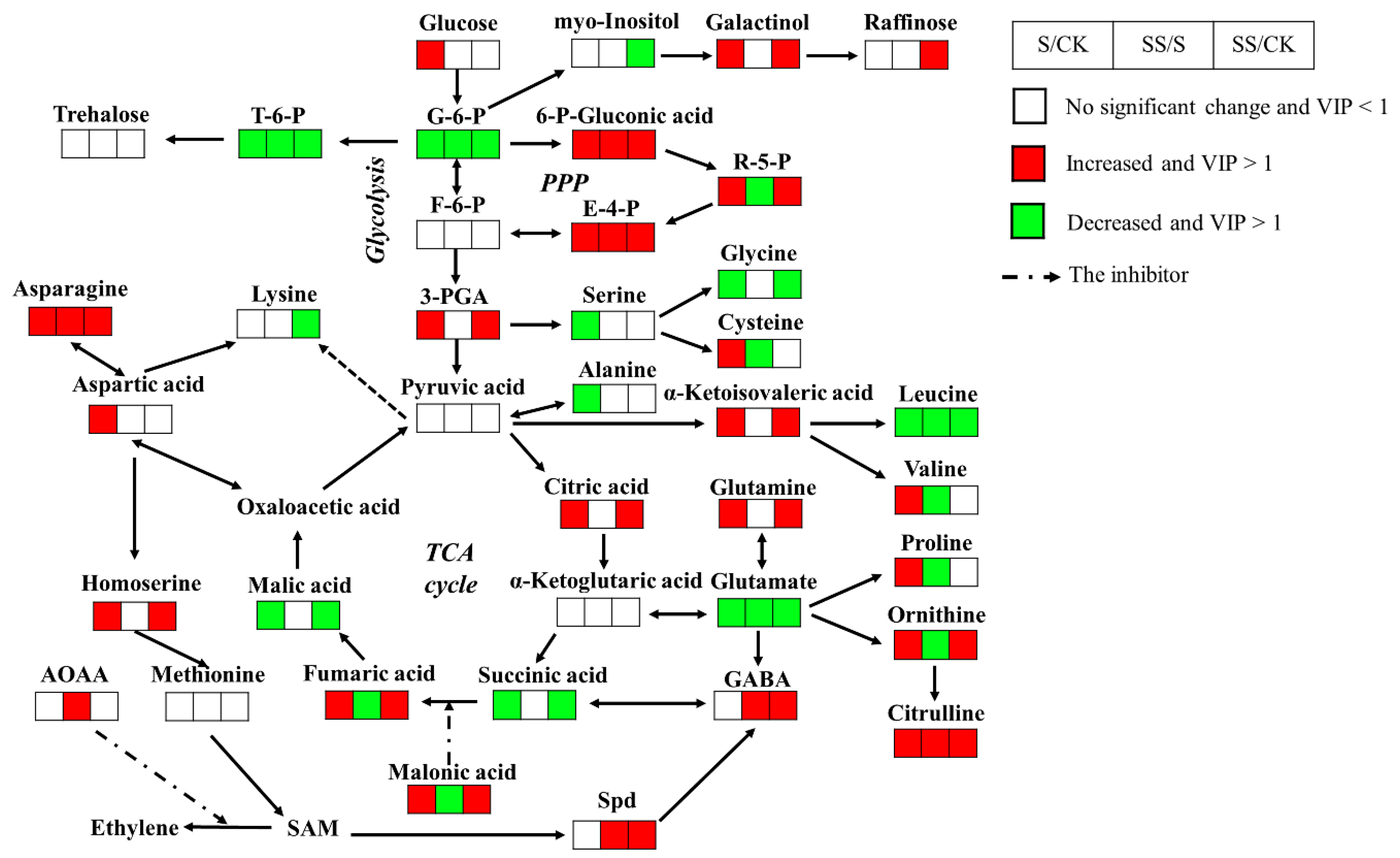
3.3. Differentially Metabolites under Salt Stress with or without Spd
3.3.1. Sugar Metabolic Changes
3.3.2. Organic Acids Changes
3.3.3. Amino Acids Changes
3.3.4. Other Metabolites Changes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Machado, R.M.A.; Serralheiro, R.P. Soil salinity: Effect on vegetable crop growth. Management practices to prevent and mitigate soil salinization. Horticulturae 2017, 3, 30. [Google Scholar] [CrossRef]
- D’Amelia, L.; Dell’Aversana, E.; Woodrow, P.; Ciarmiello, L.F.; Carillo, P. Metabolomics for crop improvement against salinity stress. In Salinity Responses and Tolerance in Plants; Kumar, V., Wani, S.H., Suprasanna, P., Tran, L.S., Eds.; Springer: Cham, Switzerland, 2018; Volume 2, pp. 267–287. [Google Scholar] [CrossRef]
- Mustafa, G.; Akhtar, M.S.; Abdullah, R. Global Concern for Salinity on Various Agro-Ecosystems. In Salt Stress, Microbes, and Plant Interactions: Causes and Solution; Akhtar, M., Ed.; Springer: Singapore, 2019; pp. 1–19. [Google Scholar] [CrossRef]
- Türkan, I.; Demiral, T. Recent developments in understanding salinity tolerance. Environ. Exp. Bot. 2009, 67, 2–9. [Google Scholar] [CrossRef]
- Wu, D.; Cai, S.; Chen, M.; Ye, L.; Chen, Z.; Zhang, H.; Zhang, G. Tissue metabolic responses to salt stress in wild and cultivated barley. PLoS ONE 2013, 8, e55431. [Google Scholar] [CrossRef] [PubMed]
- Munns, R.; Day, D.A.; Fricke, W.; Watt, M.; Arsova, B.; Barkla, B.J.; Gilliham, M. Energy costs of salt tolerance in crop plants. New Phytol. 2020, 225, 1072–1090. [Google Scholar] [CrossRef]
- Arbona, V.; Manzi, M.; Ollas, C.D.; Gómez-Cadenas, A. Metabolomics as a tool to investigate abiotic stress tolerance in plants. Int. J. Mol. Sci. 2013, 14, 4885–4911. [Google Scholar] [CrossRef]
- Baniasadi, F.; Saffari, V.R.; Moud, A.A.M. Physiological and growth responses of Calendula officinalis L. plants to the interaction effects of polyamines and salt stress. Sci. Hortic. 2018, 234, 312–317. [Google Scholar] [CrossRef]
- Jia, X.; Li, H.; Guo, H.; Wang, T.; Zhang, Z.; Qiao, Y.; Guo, Z. Metabolomics advances in horticultural crops. Chin. Agr. Sci. Bul. 2014, 30, 75–83. [Google Scholar] [CrossRef]
- Kim, H.K.; Verpoorte, R. Sample preparation for plant metabolomics. Phytochem. Anal. 2010, 21, 4–13. [Google Scholar] [CrossRef] [PubMed]
- Fiehn, O. Metabolomics—The link between genotypes and phenotypes. Plant Mol. Biol. 2002, 48, 155–171. [Google Scholar] [CrossRef]
- Jiménez-Bremont, J.F.; Ruiz, O.A.; Rodríguez-Kessler, M. Modulation of spermidine and spermine levels in maize seedlings subjected to long-term salt stress. Plant Physiol. Biochem. 2007, 45, 812–821. [Google Scholar] [CrossRef]
- López-Gómez, M.; Hidalgo-Castellanos, J.; Muñoz-Sánchez, J.R.; Marín-Peña, A.J.; Lluch, C.; Herrera-Cervera, J.A. Polyamines contribute to salinity tolerance in the symbiosis Medicago truncatula-Sinorhizobium meliloti by preventing oxidative damage. Plant Physiol. Biochem. 2017, 116, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Alcázar, R.; Altabella, T.; Marco, F.; Bortolotti, C.; Reymond, M.; Koncz, C.; Tiburcio, A.F. Polyamines: Molecules with regulatory functions in plant abiotic stress tolerance. Planta 2010, 231, 1237–1249. [Google Scholar] [CrossRef] [PubMed]
- Pál, M.; Tajti, J.; Szalai, G.; Peeva, V.; Végh, B.; Janda, T. Interaction of polyamines, abscisic acid and proline under osmotic stress in the leaves of wheat plants. Sci. Rep. 2018, 8, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Sang, T.; Shan, X.; Li, B.; Shu, S.; Sun, J.; Guo, S. Comparative proteomic analysis reveals the positive effect of exogenous spermidine on photosynthesis and salinity tolerance in cucumber seedlings. Plant Cell Rep. 2016, 35, 1769–1782. [Google Scholar] [CrossRef] [PubMed]
- Zhong, M.; Yuan, Y.; Shu, S.; Sun, J.; Guo, S.; Yuan, R.; Tang, Y. Effects of exogenous putrescine on glycolysis and Krebs cycle metabolism in cucumber leaves subjected to salt stress. Plant Growth Regul. 2016, 79, 319–330. [Google Scholar] [CrossRef]
- Zhou, H.; Guo, S.; Shao, H.; Chen, X.; Wei, B.; Sun, J. Effects of iso-smotic Ca(NO3)2 and NaCl stress on growth and physiological characteristics of cucumber seedlings. Acta Ecol. Sin. 2014, 34, 1880–1890. [Google Scholar] [CrossRef]
- Wu, J.; Shu, S.; Li, C.; Sun, J.; Guo, S. Spermidine-mediated hydrogen peroxide signaling enhances the antioxidant capacity of salt-stressed cucumber roots. Plant Physiol. Biochem. 2018, 128, 152–162. [Google Scholar] [CrossRef]
- Han, B.; Xu, G.; Guo, S.; He, C.; Sun, Y.; Gao, W.; Shi, L. Growth and physiological metabolism of cucumber seedlings in response to different concentrations of salt stress. Jiangsu J. Agr. Sci. 2014, 30, 172–177. [Google Scholar] [CrossRef]
- Haling, R.E.; Richardson, A.E.; Culvenor, R.A.; Lambers, H.; Simpson, R.J. Root morphology, root-hair development and rhizosheath formation on perennial grass seedlings is influenced by soil acidity. Plant Soil 2010, 335, 457–468. [Google Scholar] [CrossRef]
- Li, B.; He, L.; Guo, S.; Li, J.; Yang, Y.; Yan, B.; Li, J. Proteomics reveal cucumber Spd-responses under normal condition and salt stress. Plant Physiol. Biochem. 2013, 67, 7–14. [Google Scholar] [CrossRef]
- Guo, R.; Yang, Z.; Li, F.; Yan, C.; Zhong, X.; Liu, Q.; Zhao, L. Comparative metabolic responses and adaptive strategies of wheat (Triticum aestivum) to salt and alkali stress. BMC Plant Biol. 2015, 15, 170. [Google Scholar] [CrossRef] [PubMed]
- Liang, W.; Ma, X.; Wan, P.; Liu, L. Plant salt-tolerance mechanism: A review. Biochem. Biophys. Res. Commun. 2018, 495, 286–291. [Google Scholar] [CrossRef] [PubMed]
- Patterson, J.H.; Newbigin, E.D.; Tester, M.; Bacic, A.; Roessner, U. Metabolic responses to salt stress of barley (Hordeum vulgare L.) cultivars, Sahara and Clipper, which differ in salinity tolerance. J. Exp. Bot. 2009, 60, 4089–4103. [Google Scholar] [CrossRef]
- Hare, P.D.; Cress, W.A.; Van Staden, J. Dissecting the roles of osmolyte accumulation during stress. Plant Cell Environ. 1998, 21, 535–553. [Google Scholar] [CrossRef]
- Williams, T.C.R.; Poolman, M.G.; Howden, A.J.M.; Schwarzlander, M.; Fell, D.A.; Ratcliffe, R.G. A genome-scale metabolic model accurately predicts fluxes in central carbon metabolism under stress conditions. Plant Physiol. 2010, 154, 311–323. [Google Scholar] [CrossRef]
- Turner, J.F.; Tomlinson, J.D.; Caldwell, R.A. Effect of salts on the activity of carrot phosphofructokinase. Plant Physiol. 1980, 66, 973–977. [Google Scholar] [CrossRef]
- Shao, Q.; Shu, S.; Du, J.; Yuan, Y.; Xing, W.; Guo, S.; Sun, J. Proteome analysis of roots in cucumber seedlings under iso-osmotic NaCl and Ca (NO3)2 stresses. Plant Mol. Biol. Rep. 2016, 34, 303–317. [Google Scholar] [CrossRef]
- Krasensky, J.; Jonak, C. Drought, salt, and temperature stress-induced metabolic rearrangements and regulatory networks. J. Exp. Bot. 2012, 63, 1593–1608. [Google Scholar] [CrossRef]
- Li, M.; Guo, R.; Jiao, Y.; Jin, X.; Zhang, H.; Shi, L. Comparison of salt tolerance in Soja based on metabolomics of seedling roots. Front. Plant Sci. 2017, 8, 1101. [Google Scholar] [CrossRef]
- Shen, Q.; Yu, J.; Fu, L.; Wu, L.; Dai, F.; Jiang, L.; Zhang, G. Ionomic, metabolomic and proteomic analyses reveal molecular mechanisms of root adaption to salt stress in Tibetan wild barley. Plant Physiol. Biochem. 2018, 123, 319–330. [Google Scholar] [CrossRef]
- Chandler, A.M. Amino acid metabolism. In Biochemistry, 2nd ed.; Briggs, T., Chandler, A.M., Eds.; Springer: New York, NY, USA, 1995; pp. 80–103. [Google Scholar] [CrossRef]
- Yu, J.; Si, Y. Metabolic carbon fluxes and biosynthesis of polyhydroxyalkanoates in Ralstonia eutropha on short chain fatty acids. Biotechnol. Prog. 2004, 20, 1015–1024. [Google Scholar] [CrossRef] [PubMed]
- Feng, R.; Lei, L.; Liu, B.; Chen, W.; Zhang, R.; Wang, L.; Lin, Z. Effects of different inhibitors such as malonic acid, Na3PO4 and HgCl2 on uptake of different forms of antimony in rice plant. Plant Soil 2019, 445, 259–271. [Google Scholar] [CrossRef]
- Huan, L.; Xie, X.; Zheng, Z.; Sun, F.; Wu, S.; Li, M.; Wang, G. Positive correlation between PSI response and oxidative pentose phosphate pathway activity during salt stress in an intertidal macroalga. Plant Cell Physiol. 2014, 55, 1395–1403. [Google Scholar] [CrossRef] [PubMed]
- Orwat, J.; Sarkar, D.; Osorno, J.; Shetty, K. Improved salinity resilience in black bean by seed elicitation using organic compounds. Agron. J. 2017, 109, 1991–2003. [Google Scholar] [CrossRef]
- Valderrama, R.; Corpas, F.J.; Carreras, A.; Gomez-Rodriguez, M.V.; Chaki, M.; Pedrajas, J.R.; Barroso, J.B. The dehydrogenase-mediated recycling of NADPH is a key antioxidant system against salt-induced oxidative stress in olive plants. Plant Cell Environ. 2006, 29, 1449–1459. [Google Scholar] [CrossRef] [PubMed]
- Cho, E.S.; Cha, Y.H.; Kim, H.S.; Kim, N.H.; Yook, J.I. The pentose phosphate pathway as a potential target for cancer therapy. Biomol. Ther. 2018, 26, 29. [Google Scholar] [CrossRef] [PubMed]
- Goyal, R.K.; Fatima, T.; Topuz, M.; Bernadec, A.; Sicher, R.; Handa, A.K.; Mattoo, A.K. Pathogenesis-related protein 1b1 (PR1b1) is a major tomato fruit protein responsive to chilling temperature and upregulated in high polyamine transgenic genotypes. Front. Plant Sci. 2016, 7, 901. [Google Scholar] [CrossRef] [PubMed]
- Kruger, N.J.; Schaewen, A.V. The oxidative pentose phosphate pathway: Structure and organisation. Curr. Opin. Plant Biol. 2003, 6, 236–246. [Google Scholar] [CrossRef]
- Handley, L.W.; Pharr, D.M.; McFeeters, R.F. Carbohydrate changes during maturation of cucumber fruit: Implications for sugar metabolism and transport. Plant Physiol. 1983, 72, 498–502. [Google Scholar] [CrossRef]
- Irving, D.E.; Hurst, P.L.; Ragg, J.S. Changes in carbohydrates and carbohydrate metabolizing enzymes during the development, maturation, and ripening of buttercup squash (Cucurbita maxima D. Delica’). J. Am. Soc. Hortic. Sci. 1997, 122, 310–314. [Google Scholar] [CrossRef]
- Taji, T.; Ohsumi, C.; Iuchi, S.; Seki, M.; Kasuga, M.; Kobayashi, M.; Shinozaki, K. Important roles of drought- and cold- inducible genes for galactinol synthase in stress tolerance in Arabidopsis thaliana. Plant J. 2002, 29, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos, T.B.; Budzinski, I.G.; Marur, C.J.; Petkowicz, C.L.; Pereira, L.F.; Vieira, L.G. Expression of three galactinol synthase isoforms in Coffea arabica L. and accumulation of raffinose and stachyose in response to abiotic stresses. Plant Physiol. Biochem. 2011, 49, 441–448. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, A.; Chakraborty, M.; Saha, J.; Gupta, B.; Gupta, K. Polyamines: Osmoprotectants in plant abiotic stress adaptation. In Osmolytes and Plants Acclimation to Changing Environment: Emerging Omics Technologies; Iqbal, N., Nazar, R., Eds.; Springer: New Delhi, India, 2016; pp. 97–127. [Google Scholar] [CrossRef]
- Fernandez, O.; Béthencourt, L.; Quero, A.; Sangwan, R.S.; Clément, C. Trehalose and plant stress responses: Friend or foe? Trends Plant Sci. 2010, 15, 409–417. [Google Scholar] [CrossRef] [PubMed]
- O’Hara, L.E.; Paul, M.J.; Wingler, A. How do sugars regulate plant growth and development? New insight into the role of trehalose-6-phosphate. Mol. Plant. 2013, 6, 261–274. [Google Scholar] [CrossRef] [PubMed]
- Smeekens, S.; Ma, J.; Hanson, J.; Rolland, F. Sugar signals and molecular networks controlling plant growth. Curr. Opin. Plant Biol. 2010, 13, 273–278. [Google Scholar] [CrossRef] [PubMed]
- Gazzarrini, S.; Tsai, A.Y.L. Trehalose-6-phosphate and SnRK1 kinases in plant development and signaling: The emerging picture. Front. Plant Sci. 2014, 5, 119. [Google Scholar] [CrossRef]
- Hardie, D.G. AMPK and SNF1: Snuffing out stress. Cell Metab. 2007, 6, 339–340. [Google Scholar] [CrossRef]
- Baena-González, E.; Rolland, F.; Thevelein, J.M.; Sheen, J. A central integrator of transcription networks in plant stress and energy signalling. Nature 2007, 448, 938. [Google Scholar] [CrossRef]
- Emanuelle, S.; Doblin, M.S.; Stapleton, D.I.; Bacic, A.; Gooley, P.R. Molecular insights into the enigmatic metabolic regulator, SnRK. Trends Plant Sci. 2016, 21, 341–353. [Google Scholar] [CrossRef]
- Oszvald, M.; Primavesi, L.F.; Griffiths, C.A.; Cohn, J.; Basu, S.S.; Nuccio, M.L.; Paul, M.J. Trehalose 6-phosphate regulates photosynthesis and assimilate partitioning in reproductive tissue. Plant Physiol. 2018, 176, 2623–2638. [Google Scholar] [CrossRef]
- Shu, S.; Chen, L.; Lu, W.; Sun, J.; Guo, S.; Yuan, Y.; Li, J. Effects of exogenous spermidine on photosynthetic capacity and expression of Calvin cycle genes in salt-stressed cucumber seedlings. J. Plant Res. 2014, 127, 763–773. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Shu, S.; An, Y.; Zhou, H.; Guo, S.; Sun, J. Influence of exogenous spermidine on carbon–nitrogen metabolism under Ca(NO3)2 stress in cucumber root. Plant Growth Regul. 2017, 81, 103–115. [Google Scholar] [CrossRef]
- Joshi, V.; Fernie, A.R. Citrulline metabolism in plants. Amino Acids 2017, 49, 1543–1559. [Google Scholar] [CrossRef] [PubMed]
- Okumoto, S.; Funck, D.; Trovato, M.; Forlani, G. Amino acids of the glutamate family: Functions beyond primary metabolism. Front. Plant Sci. 2016, 7, 318. [Google Scholar] [CrossRef]
- Sanchez, D.H.; Pieckenstain, F.L.; Szymanski, J.; Erban, A.; Bromke, M.; Hannah, M.A.; Udvardi, M.K. Comparative functional genomics of salt stress in related model and cultivated plants identifies and overcomes limitations to translational genomics. PLoS ONE 2011, 6, e17094. [Google Scholar] [CrossRef]
- Sicher, R.C.; Timlin, D.; Bailey, B. Responses of growth and primary metabolism of water-stressed barley roots to rehydration. J. Plant Physiol. 2012, 169, 686–695. [Google Scholar] [CrossRef]
- Triplett, E.W.; Blevins, D.G.; Randall, D.D. Allantoic acid synthesis in soybean root nodule cytosol via xanthine dehydrogenase. Plant Physiol. 1980, 65, 1203–1206. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ray, R.M.; Bhattacharya, S.; Bavaria, M.N.; Viar, M.J.; Johnson, L.R. Spermidine, a sensor for antizyme 1 expression regulates intracellular polyamine homeostasis. Amino Acids 2014, 46, 2005–2013. [Google Scholar] [CrossRef][Green Version]
- Liu, D.; He, S.; Zhai, H.; Wang, L.; Zhao, Y.; Wang, B.; Liu, Q. Overexpression of IbP5CR enhances salt tolerance in transgenic sweet potato. Plant Cell Tissue Organ. Cult. 2014, 117, 1–16. [Google Scholar] [CrossRef]
- López-Gómez, M.; Hidalgo-Castellanos, J.; Iribarne, C.; Lluch, C. Proline accumulation has prevalence over polyamines in nodules of Medicago sativa in symbiosis with Sinorhizobium meliloti during the initial response to salinity. Plant Soil 2014, 374, 149–159. [Google Scholar] [CrossRef]
- Ludwig, R.A. Arabidopsis chloroplasts dissimilate L-arginine and L-citrulline for use as N source. Plant Physiol. 1993, 101, 429–434. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, D.E.; Madore, M.A. Patterns of assimilate production and translocation in muskmelon (Cucumis melo L.): II. Low temperature effects. Plant Physiol. 1992, 99, 966–971. [Google Scholar] [CrossRef] [PubMed]
- Akashi, K.; Miyake, C.; Yokota, A. Citrulline, a novel compatible solute in drought-tolerant wild watermelon leaves, is an efficient hydroxyl radical scavenger. FEBS Lett. 2001, 508, 438–442. [Google Scholar] [CrossRef]
- Massange-Sanchez, J.A.; Palmeros-Suarez, P.A.; Espitia-Rangel, E.; Rodriguez-Arevalo, I.; Sanchez-Segura, L.; Martinez-Gallardo, N.A.; Delano-Frier, J.P. Overexpression of grain amaranth (Amaranthus hypochondriacus) AhERF or AhDOF transcription factors in Arabidopsis thaliana increases water deficit-and salt-stress tolerance, respectively, via contrasting stress-amelioration mechanisms. PLoS ONE 2016, 11, e0164280. [Google Scholar] [CrossRef] [PubMed]
- Roosens, N.H.; Thu, T.T.; Iskandar, H.M.; Jacobs, M. Isolation of the ornithine-δ-aminotransferase cDNA and effect of salt stress on its expression in Arabidopsis thaliana. Plant Physiol. 1998, 117, 263–271. [Google Scholar] [CrossRef] [PubMed]
- De Campos, M.K.F.; de Carvalho, K.; de Souza, F.S.; Marur, C.J.; Pereira, L.F.P.; Filho, J.C.; Vieira, L.G.E. Drought tolerance and antioxidant enzymatic activity in transgenic ‘Swingle’ citrumelo plants over-accumulating proline. Environ. Exp. Bot. 2011, 72, 242–250. [Google Scholar] [CrossRef]
- Ghars, M.A.; Parre, E.; Debez, A.; Bordenave, M.; Richard, L.; Leport, L.; Abdelly, C. Comparative salt tolerance analysis between Arabidopsis thaliana and Thellungiella halophila, with special emphasis on K+/Na+ selectivity and proline accumulation. J. Plant Physiol. 2008, 165, 588–599. [Google Scholar] [CrossRef]
- Gessler, P.; Buchal, P.; Schwenk, H.U.; Wermuth, B. Favourable long-term outcome after immediate treatment of neonatal hyperammonemia due to N-acetylglutamate synthase deficiency. Eur. J. Pediatr. 2010, 169, 197–199. [Google Scholar] [CrossRef]
- Ramesh, S.A.; Tyerman, S.D.; Gilliham, M.; Xu, B. γ-Aminobutyric acid (GABA) signalling in plants. Cell. Mol. Life Sci. 2017, 74, 1577–1603. [Google Scholar] [CrossRef]
- Xu, J.; Liu, T.; Yang, S.; Jin, X.; Qu, F.; Huang, N.; Hu, X. Polyamines are involved in GABA-regulated salinity-alkalinity stress tolerance in muskmelon. Environ. Exp. Bot. 2019, 164, 181–189. [Google Scholar] [CrossRef]
- Jin, X.; Liu, T.; Xu, J.; Gao, Z.; Hu, X. Exogenous GABA enhances muskmelon tolerance to salinity-alkalinity stress by regulating redox balance and chlorophyll biosynthesis. BMC Plant Biol. 2019, 19, 48. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Xu, Z.; Xu, W.; Li, J.; Zhao, N.; Zhou, Y. Application of γ-aminobutyric acid demonstrates a protective role of polyamine and GABA metabolism in muskmelon seedlings under Ca(NO3)2 stress. Plant Physiol. Biochem. 2015, 92, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Radhakrishnan, R.; Lee, I.J. Regulation of salicylic acid, jasmonic acid and fatty acids in cucumber (Cucumis sativus L.) by spermidine promotes plant growth against salt stress. Acta Physiol. Plant. 2013, 35, 3315–3322. [Google Scholar] [CrossRef]
- Hao, J.H.; Wang, X.L.; Dong, C.J.; Zhang, Z.G.; Shang, Q.M. Salicylic acid induces stomatal closure by modulating endogenous hormone levels in cucumber cotyledons. Russ. J. Plant Physiol. 2011, 58, 906. [Google Scholar] [CrossRef]
- Feng, H.Y.; Wang, Z.M.; Kong, F.N.; Zhang, M.J.; Zhou, S.L. Roles of carbohydrate supply and ethylene, polyamines in maize kernel set. J. Integr. Plant Biol. 2011, 53, 388–398. [Google Scholar] [CrossRef]
- Hagège, D.; Kevers, C.; Geuns, J.; Gaspar, T. Ethylene production and polyamine content of fully habituated sugarbeet calli. J. Plant Physiol. 1994, 143, 722–725. [Google Scholar] [CrossRef]
- Kacperska, A.; Kubacka-Zębalska, M. Formation of stress ethylene depends both on ACC synthesis and on the activity of free radical-generating system. Physiol. Plant. 1989, 77, 231–237. [Google Scholar] [CrossRef]
- Siddikee, M.A.; Glick, B.R.; Chauhan, P.S.; Yim, W.; Sa, T. Enhancement of growth and salt tolerance of red pepper seedlings (Capsicum annuum L.) by regulating stress ethylene synthesis with halotolerant bacteria containing 1-aminocyclopropane-1-carboxylic acid deaminase activity. Plant Physiol. Biochem. 2011, 49, 427–434. [Google Scholar] [CrossRef]

| Metabolite Names | Relative Concentration | Fold Changes | VIP | ||||||
|---|---|---|---|---|---|---|---|---|---|
| CK | S | SS | Log2 (S/CK) | Log2 (SS/S) | Log2 (SS/CK) | S/CK | SS/S | SS/CK | |
| Simple sugars | |||||||||
| Fructose | 15.74 ± 0.61 | 11.86 ± 0.69 | 11.95 ± 0.76 | −0.41 ** | 0.01 | −0.40 ** | 0.95 | 0.02 | 0.9 |
| Mannose | 132.87 ± 6.19 | 56.64 ± 3.17 | 55.38 ± 3.63 | −1.23 *** | −0.03 | −1.26 *** | 1.21 | 0.13 | 1.21 |
| Xylose | 21.64 ± 0.67 | 8.44 ± 0.58 | 5.25 ± 0.3 | −1.36 *** | −0.68 *** | −2.04 *** | 1.22 | 1.2 | 1.26 |
| Erythrose | 5.66 ± 0.32 | 2.14 ± 0.22 | 8.61 ± 0.08 | −1.41 *** | 2.01 *** | 0.61 *** | 1.16 | 1.44 | 1.14 |
| Galactose | 31.89 ± 1.98 | 154.64 ± 17.37 | 125.09 ± 6.81 | 2.28 *** | −0.31 | 1.97 *** | 1.13 | 0.25 | 1.25 |
| Sedoheptulose | 101.88 ± 6.41 | 45.73 ± 2.47 | 34.28 ± 1.29 | −1.16 *** | −0.42 ** | −1.57 *** | 1.18 | 1.12 | 1.24 |
| Glucoheptose | 6.38 ± 0.34 | 8.03 ± 0.58 | 5.35 ± 0.49 | 0.33 * | −0.59 ** | −0.25 | 0.71 | 1.03 | 0.57 |
| Talose | 3.38 ± 0.12 | 2.09 ± 0.07 | 1.51 ± 0.03 | −0.69 *** | −0.47 *** | −1.16 *** | 1.19 | 1.35 | 1.26 |
| Allose | 0.82 ± 0.04 | 0.39 ± 0.02 | 0.38 ± 0.03 | −1.07 *** | −0.04 | −1.11 *** | 1.18 | 0.22 | 1.15 |
| Altrose | 3.16 ± 0.25 | 11.29 ± 0.6 | 14.74 ± 0.77 | 1.84 *** | 0.38 ** | 2.22 *** | 1.22 | 1.03 | 1.25 |
| Oligoses | |||||||||
| Maltotriose | 3.83 ± 0.27 | 1.37 ± 0.04 | 2.61 ± 0.2 | −1.48 *** | 0.93 *** | −0.55 ** | 1.22 | 1.34 | 0.89 |
| Cellobiose | 9.14 ± 0.53 | 1.99 ± 0.27 | 1.69 ± 0.09 | −2.20 *** | −0.23 | −2.43 *** | 1.01 | 0.06 | 1.26 |
| Leucrose | 0.03 ± 0.01 | 0.04 ± 0 | 0.05 ± 0.01 | 0.58 | 0.2 | 0.78 | 0.55 | 0.24 | 0.61 |
| Melibiose | 12.94 ± 0.63 | 9.48 ± 0.91 | 4.89 ± 0.44 | −0.45 ** | −0.96 *** | −1.40 *** | 0.83 | 1.19 | 1.18 |
| Polysaccharide | |||||||||
| Levoglucosan | 4.04 ± 0.27 | 12.13 ± 0.77 | 9.15 ± 0.59 | 1.59 *** | −0.41 * | 1.18 *** | 1.22 | 0.96 | 1.18 |
| Sugar phosphate esters | |||||||||
| Glucose-1-P | 69.52 ± 6.18 | 357.65 ± 20.12 | 529.59 ± 81.17 | 2.36 *** | 0.57 * | 2.93 *** | 1.24 | 0.82 | 1.23 |
| Sugar acids | |||||||||
| Saccharic acid | 0.56 ± 0.03 | 0.94 ± 0.03 | 8.95 ± 0.42 | 0.76 *** | 3.25 *** | 4.00 *** | 1.13 | 1.49 | 1.27 |
| Glucuronic acid | 8.78 ± 0.38 | 9.7 ± 0.14 | 13.95 ± 0.42 | 0.14 * | 0.53 *** | 0.67 *** | 0.68 | 1.41 | 1.18 |
| Gluconic acid | 20.39 ± 0.84 | 25.3 ± 0.65 | 30.2 ± 0.9 | 0.31 *** | 0.26 ** | 0.57 *** | 0.99 | 1.16 | 1.16 |
| Gluconic lactone | 2.08 ± 0.07 | 1.05 ± 0.1 | 1.64 ± 0.28 | −0.98 *** | 0.64 | −0.35 | 1.13 | 0.58 | 0.55 |
| Glucoheptonic acid | 0.66 ± 0.04 | 1.32 ± 0.06 | 0.83 ± 0.07 | 1.00 *** | −0.67 *** | 0.33 | 1.17 | 1.19 | 0.65 |
| Galactonic acid | 5.68 ± 0.26 | 10 ± 0.35 | 8.8 ± 0.56 | 0.82 *** | −0.18 | 0.63 *** | 1.18 | 0.67 | 1.06 |
| Threonic acid | 14.82 ± 0.61 | 35.47 ± 2.16 | 39.15 ± 1.01 | 1.26 *** | 0.14 | 1.40 *** | 1.22 | 0.63 | 1.26 |
| Lactobionic acid | 14.07 ± 1.03 | 5.69 ± 0.38 | 8.69 ± 0.7 | −1.31 *** | 0.61 ** | −0.69 *** | 1.18 | 1.08 | 0.98 |
| Glucosaminic acid | 2.54 ± 0.07 | 3 ± 0.17 | 2.6 ± 0.16 | 0.24 * | 0.21 | 0.03 | 0.73 | 0.66 | 0.07 |
| Amino sugar | |||||||||
| N-acetyl-β-D-mannosamine | 1.08 ± 0.1 | 1.46 ± 0.22 | 1.83 ± 0.07 | 0.43 | 0.33 | 0.76 *** | 0.03 | 0.47 | 1 |
| Deoxy suger | |||||||||
| 2-Deoxy-D-galactose | 0.22 ± 0.02 | 0.41 ± 0.01 | 0.32 ± 0.02 | 0.89 *** | −0.36 ** | 0.53 ** | 1.12 | 1.12 | 0.92 |
| Sugar alcohols | |||||||||
| Ribitol | 5.53 ± 0.18 | 3.01 ± 0.18 | 2.03 ± 0.14 | −0.88 *** | −0.57 ** | −1.44 *** | 1.18 | 1.14 | 1.23 |
| Threitol | 0.93 ± 0.01 | 2.23 ± 0.15 | 2.11 ± 0.12 | 1.25 *** | −0.08 | 1.18 *** | 1.21 | 0.19 | 1.24 |
| Allo-inositol | 0.31 ± 0.02 | 0.34 ± 0.02 | 0.28 ± 0.02 | 0.13 | −0.31 * | −0.18 | 0.36 | 0.83 | 0.49 |
| 1,5-Anhydroglucitol | 1.84 ± 0.42 | 1.92 ± 0.04 | 1.84 ± 0.1 | 0.06 | −0.06 | 0 | 0.33 | 0.41 | 0.21 |
| Carboxylic acids | |||||||||
| Maleic acid | 0.6 ± 0.02 | 0.29 ± 0.03 | 0.18 ± 0.02 | −1.06 *** | −0.70 ** | −1.77 *** | 1.15 | 1.11 | 1.24 |
| Glyceric acid | 14.17 ± 0.35 | 10.99 ± 0.33 | 11.57 ± 0.43 | −0.37 *** | 0.07 | −0.29 *** | 1.11 | 0.41 | 1 |
| Glycolic acid | 0.89 ± 0.07 | 0.55 ± 0.07 | 0.25 ± 0.02 | −0.7 ** | −1.14 *** | −1.83 *** | 0.9 | 1.24 | 1.22 |
| 2-Furoic acid | 2.17 ± 0.56 | 0.39 ± 0.09 | 0 | −2.46 *** | OS ** | OS ** | 0.99 | 1.37 | 1.23 |
| Lactic acid | 65.74 ± 4.89 | 48.74 ± 3.45 | 32.82 ± 2.03 | −0.43 * | −0.58 ** | −1.00 *** | 0.76 | 1.08 | 1.14 |
| 3-Hydroxypropionic acid | 0.74 ± 0.01 | 0.44 ± 0.01 | 0.51 ± 0.02 | −0.76 *** | 0.23 ** | −0.53 *** | 1.25 | 1.13 | 1.2 |
| 3-Hydroxybutyric acid | 2.21 ± 0.11 | 0.9 ± 0.05 | 0.62 ± 0.04 | −1.30 *** | −0.54 *** | −1.83 *** | 1.22 | 1.17 | 1.25 |
| 2-Hydroxybutanoic acid | 363.26 ± 17.41 | 240.05 ± 6.36 | 185.64 ± 5.56 | −0.60 *** | −0.37 *** | −0.97 *** | 1.13 | 1.3 | 1.22 |
| Benzoic acid | 0.72 ± 0.04 | 0.98 ± 0.13 | 1.64 ± 0.1 | 0.44 | 0.75 ** | 1.18 *** | 0.58 | 1.11 | 1.2 |
| Quinoline-4-carboxylic acid | 0.21 ± 0.01 | 0.19 ± 0.01 | 0.12 ± 0.01 | −0.19 | −0.61 | −0.79 | 0.42 | 0.78 | 0.66 |
| Pyrrole-2-carboxylic acid | 0.26 ± 0.04 | 0.34 ± 0.03 | 0.19 ± 0.02 | 0.39 | −0.81 *** | −0.42 | 0.59 | 1.25 | 0.41 |
| Pipecolinic acid | 15.85 ± 0.49 | 21.37 ± 0.74 | 25.23 ± 0.24 | 0.43 *** | 0.24 *** | 0.67 *** | 1.1 | 1.18 | 1.24 |
| 2-Ketobutyric acid | 1.67 ± 0.08 | 4.75 ± 0.38 | 4.76 ± 0.1 | 1.51 *** | 0 | 1.51 *** | 1.21 | 0.1 | 1.26 |
| Polyphenols | |||||||||
| Gallic acid | 7.11 ± 0.39 | 5.53 ± 0.35 | 3.6 ± 0.18 | −0.36 * | −0.62 *** | −0.98 *** | 0.81 | 1.22 | 1.19 |
| Pyrogallol | 0.13 ± 0 | 0.16 ± 0.01 | 0.11 ± 0.01 | 0.3 * | −0.54 ** | −0.24 * | 0.74 | 1.09 | 0.73 |
| 1,2,4-Benzenetriol | 0.17 ± 0.01 | 0.18 ± 0.01 | 0.1 ± 0.01 | 0.04 | −0.87 ** | −0.83 *** | 0.07 | 1.15 | 1 |
| Neohesperidin | 2.95 ± 0.17 | 2.08 ± 0.1 | 1.58 ± 0.17 | −0.51 *** | −0.4 | −0.9 * | 0.98 | 0.79 | 0.72 |
| Alcohols | |||||||||
| Acetol | 0.55 ± 0.08 | 0.82 ± 0.08 | 0.57 ± 0.04 | 0.57 | −0.52 * | 0.05 | 0.63 | 0.87 | 0.18 |
| Cuminic alcohol | 0.56 ± 0.1 | 0.36 ± 0.01 | 0.33 ± 0.09 | −0.64 | −0.14 ** | −0.78 ** | 0.63 | 1.03 | 0.92 |
| Cyclohexane-1,2-diol | 2.09 ± 0.06 | 0.97 ± 0.25 | 2.73 ± 0.17 | −1.10 * | 1.49 | 0.38 | 0.86 | 0.25 | 0.25 |
| Dodecanol | 0.56 ± 0.04 | 0.52 ± 0.04 | 0.34 ± 0.02 | −0.11 | −0.60 *** | −0.71 *** | 0.26 | 1.17 | 1.09 |
| Diglycerol | 4.29 ± 0.03 | 1.46 ± 0.06 | 0.84 ± 0.11 | −1.56 *** | −0.79 * | −2.34 ** | 1.26 | 0.86 | 0.87 |
| Glycerol | 522.34 ± 77.62 | 390.08 ± 12.75 | 475.12 ± 21.25 | −0.42 | 0.28 *** | −0.13 | 0.19 | 1.03 | 0.26 |
| Fatty acids | |||||||||
| Stearic acid | 0.25 ± 0.01 | 0.32 ± 0.02 | 0.28 ± 0.01 | 0.34 ** | −0.21 | 0.14 | 0.83 | 0.65 | 0.43 |
| Palmitic acid | 10.3 ± 0.86 | 5.81 ± 0.64 | 4.48 ± 0.86 | −0.83 *** | −0.37 | −1.20 ** | 0.97 | 0.56 | 0.96 |
| Heptadecanoic acid | 1.13 ± 0.25 | 0.43 ± 0.12 | 0.05 ± 0.02 | −1.38 ** | −3.08 *** | −4.46 *** | 0.89 | 1.3 | 1.13 |
| Azelaic acid | 0.22 ± 0.01 | 0.26 ± 0.01 | 0.17 ± 0.01 | 0.25 * | −0.60 *** | −0.34 ** | 0.8 | 1.37 | 0.96 |
| Simple lipids | |||||||||
| Monostearin | 0.64 ± 0 | 0.3 ± 0.03 | 0.56 ± 0.07 | −1.06 *** | 0.87 | −0.19 * | 1.13 | 0.68 | 0.76 |
| 1-Monopalmitin | 2.5 ± 0.33 | 1.47 ± 0.1 | 2.27 ± 0.16 | −0.77 | 0.63 ** | −0.14 | 0.47 | 1.11 | 0.04 |
| 2-Monopalmitin | 0.79 ± 0.09 | 0.7 ± 0.05 | 1.18 ± 0.11 | −0.18 | 0.75 ** | 0.57 * | 0.25 | 1.13 | 0.78 |
| Other aspartate family | |||||||||
| Threonine | 2.36 ± 0.04 | 1.77 ± 0.04 | 3.42 ± 0.03 | −0.41 *** | 0.95 *** | 0.54 *** | 1.21 | 1.48 | 1.26 |
| Isoleucine | 33.33 ± 0.54 | 30.55 ± 1.28 | 23.53 ± 0.61 | −0.13 | −0.38 *** | −0.50 *** | 0.61 | 1.21 | 1.22 |
| Aromatic amino acids | |||||||||
| Tryptophan | 8.75 ± 0.36 | 12.96 ± 0.49 | 10.2 ± 0.43 | 0.57 *** | −0.35 ** | 0.22 * | 1.13 | 1.13 | 0.74 |
| Phenylalanine | 0.83 ± 0.05 | 1.43 ± 0.08 | 0.86 ± 0.1 | 0.79 *** | −0.73 ** | 0.06 | 1.1 | 1.13 | 0.03 |
| Tyrosine | 161.38 ± 8 | 125.23 ± 6.4 | 92.65 ± 3.99 | −0.37 ** | −0.43 ** | −0.80 *** | 0.88 | 1.14 | 1.17 |
| Other amino acids | |||||||||
| Oxoproline | 224.1 ± 16.39 | 615.38 ± 17.81 | 428.64 ± 20.67 | 1.46 *** | −0.52 *** | 0.94 *** | 1.22 | 1.27 | 1.13 |
| α-Aminoadipic acid | 11.92 ± 0.44 | 8.1 ± 0.17 | 5.48 ± 0.15 | −0.56 *** | −0.56 *** | −1.12 *** | 1.17 | 1.42 | 1.25 |
| Creatine degr | 5.88 ± 0.36 | 7.06 ± 0.21 | 6.46 ± 0.29 | 0.27 * | −0.13 | 0.14 | 0.74 | 0.64 | 0.42 |
| N-Carbamylglutamate | 3.61 ± 0.31 | 6.87 ± 0.31 | 9.74 ± 0.14 | 0.93 *** | 0.50 *** | 1.43 *** | 1.1 | 1.33 | 1.21 |
| Cycloleucine | 0.33 ± 0.02 | 0.48 ± 0.06 | 0.53 ± 0.11 | 0.55 | 0.16 | 0.71 | 0.59 | 0.05 | 0.51 |
| Cysteinylglycine | 0 | 1.05 ± 0.27 | 0.51 ± 0.09 | OS ** | −1.04 | OS ** | 1.11 | 0.42 | 0.99 |
| Maleimide | 0.1 ± 0 | 0.23 ± 0.01 | 0.25 ± 0.01 | 1.20 *** | 0.15 | 1.35 *** | 1.26 | 0.62 | 1.25 |
| Oxamide | 0.68 ± 0.01 | 6.66 ± 0.36 | 6.14 ± 0.23 | 3.30 *** | −0.12 | 3.18 | 1.27 | 0.58 | 0.02 |
| Nucleotide metabolic pathway | |||||||||
| Uracil | 4.28 ± 0.47 | 4.36 ± 0.44 | 2.94 ± 0.15 | 0.03 | −0.57 ** | −0.54 * | 0.08 | 1.07 | 0.75 |
| Thymine | 0.24 ± 0.01 | 0.11 ± 0 | 0.19 ± 0 | −1.13 *** | 0.84 *** | −0.30 ** | 1.23 | 1.46 | 0.92 |
| Adenine | 3.22 ± 0.1 | 2.59 ± 0.08 | 2 ± 0.09 | −0.31 *** | −0.38 ** | −0.69 *** | 1.01 | 1.16 | 1.17 |
| Uridine | 4.03 ± 0.52 | 2.98 ± 0.34 | 7.76 ± 0.35 | −0.44 | 1.38 *** | 0.95 ** | 0.42 | 1.3 | 0.96 |
| Guanosine | 12.33 ± 1.66 | 4.06 ± 0.25 | 3.79 ± 0.38 | −1.60 *** | −0.1 | −1.70 *** | 1.11 | 0.3 | 1.1 |
| Adenosine | 97.24 ± 8 | 15.08 ± 0.51 | 35.96 ± 1.01 | −2.69 *** | 1.25 *** | −1.44 *** | 1.25 | 1.47 | 1.22 |
| Inosine | 1.37 ± 0.09 | 0.83 ± 0.05 | 0.42 ± 0.02 | −0.72 ** | −1.00 *** | −1.72 *** | 1.07 | 1.37 | 1.24 |
| Hypoxanthine | 0 | 0.1 ± 0.02 | 0.06 | OS *** | OS * | OS | 1.24 | 1.29 | 1.28 |
| Carbamoyl-aspartic acid | 2.46 ± 0.16 | 3.8 ± 0.22 | 2.26 ± 0.23 | 0.63 *** | −0.75 ** | −0.12 | 1.01 | 1 | 0.27 |
| β-Alanine | 5.64 ± 0.13 | 3.96 ± 0.09 | 4.05 ± 0.06 | −0.51 *** | 0.03 | −0.48 *** | 1.21 | 0.34 | 1.22 |
| 3-Aminoisobutyric acid | 45.2 ± 2.04 | 48.34 ± 1.87 | 27.57 ± 5.41 | 0.1 | −0.81 * | −0.71 * | 0.38 | 0.91 | 0.73 |
| 5,6-Dihydrouracil | 41.11 ± 2.4 | 33.48 ± 1.93 | 19.5 ± 1.2 | −0.30 * | −0.78 *** | −1.08 *** | 0.7 | 1.28 | 1.17 |
| Allantoic acid | 0 | 3.22 ± 0.12 | 6.3 ± 0.04 | OS *** | 0.97 | OS *** | 1.27 | 0.37 | 0.99 |
| Urea | 10.22 ± 0.32 | 20.86 ± 0.89 | 16.08 ± 0.67 | 1.03 *** | −0.38 ** | 0.65 *** | 1.23 | 1.15 | 1.18 |
| Polyamines | |||||||||
| Putrescine | 28.26 ± 2.26 | 25.54 ± 1.62 | 23.03 ± 1.74 | −0.15 | −0.15 | −0.31 | 0.29 | 0.41 | 0.55 |
| Hormones and precursors | |||||||||
| Salicin | 2.73 ± 1.62 | 4.45 ± 0.84 | 6.71 ± 0.93 | 0.71 | 0.59 * | 1.3 * | 0.51 | 1.2 | 0.97 |
| Melatonin | 0.79 ± 0.01 | 0.82 ± 0.02 | 1.49 ± 0.03 | 0.06 | 0.86 | 0.92 | 0.5 | 0.06 | 0.09 |
| α-Ecdysone | 20.49 ± 1.18 | 16.73 ± 0.61 | 17.87 ± 0.99 | −0.29 * | 0.1 | −0.2 | 0.74 | 0.33 | 0.5 |
| Sphingosine | |||||||||
| Phytosphingosine | 0.8 ± 0.09 | 1.61 ± 0.28 | 2.74 ± 0.56 | 1.01 | 0.77 | 1.78 ** | 0.51 | 0.55 | 0.96 |
| D-Erythro-sphingosine | 2.87 ± 0.33 | 7.94 ± 0.62 | 6.38 ± 0.85 | 1.47 *** | −0.32 | 1.15 *** | 1.15 | 0.51 | 0.84 |
| Dihydrosphingosine | 3.58 ± 0.17 | 3.63 ± 0.09 | 5.14 ± 0.15 | 0.02 | 0.5 | 0.52 | 0.12 | 0.46 | 0.41 |
| Catecholamine metabolism pathway | |||||||||
| L-dopa | 1.23 ± 0.07 | 1.75 ± 0.17 | 1.13 ± 0.07 | 0.51 * | −0.63 ** | −0.12 | 0.81 | 1.06 | 0.3 |
| Dehydroascorbic acid | 0.37 ± 0.02 | 0.22 ± 0.01 | 0.17 ± 0.01 | −0.75 *** | −0.41 *** | −1.16 *** | 1.11 | 1.21 | 1.21 |
| Noradrenaline | 9.82 ± 0.63 | 17.28 ± 1.15 | 26.68 ± 2.05 | 0.82 *** | 0.63 *** | 1.44 *** | 1.07 | 1.12 | 1.2 |
| Steroids | |||||||||
| 22-Ketocholesterol | 5.12 ± 0.28 | 132.18 ± 5.73 | 96.63 ± 3.18 | 4.69 *** | −0.45 *** | 4.24 *** | 1.27 | 1.24 | 1.28 |
| 21-Hydroxypregnenolone | 0.35 ± 0.01 | 0.5 ± 0.02 | 0.38 ± 0.01 | 0.51 *** | −0.40 *** | 0.11 | 1.12 | 1.2 | 0.63 |
| 7-Hydroxy-4-androstene-3,17-dione | 0.28 ± 0.02 | 0.19 ± 0.01 | 0 | −0.54 *** | OS *** | OS *** | 1.02 | 1.36 | 1.17 |
| Vitamin | |||||||||
| Nicotinic acid | 2.2 ± 0.04 | 3.03 ± 0.13 | 2.06 ± 0.15 | 0.46 *** | −0.56 ** | −0.1 | 1.12 | 1.14 | 0.35 |
| Nicotinamide | 0.95 ± 0.05 | 0.85 ± 0.11 | 0.72 ± 0.07 | −0.16 | −0.24 | −0.41 * | 0.4 | 0.44 | 0.8 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, B.; Peng, X.; Han, L.; Hou, L.; Li, B. Effects of Exogenous Spermidine on Root Metabolism of Cucumber Seedlings under Salt Stress by GC-MS. Agronomy 2020, 10, 459. https://doi.org/10.3390/agronomy10040459
Liu B, Peng X, Han L, Hou L, Li B. Effects of Exogenous Spermidine on Root Metabolism of Cucumber Seedlings under Salt Stress by GC-MS. Agronomy. 2020; 10(4):459. https://doi.org/10.3390/agronomy10040459
Chicago/Turabian StyleLiu, Bing, Xujian Peng, Lingjuan Han, Leiping Hou, and Bin Li. 2020. "Effects of Exogenous Spermidine on Root Metabolism of Cucumber Seedlings under Salt Stress by GC-MS" Agronomy 10, no. 4: 459. https://doi.org/10.3390/agronomy10040459
APA StyleLiu, B., Peng, X., Han, L., Hou, L., & Li, B. (2020). Effects of Exogenous Spermidine on Root Metabolism of Cucumber Seedlings under Salt Stress by GC-MS. Agronomy, 10(4), 459. https://doi.org/10.3390/agronomy10040459




