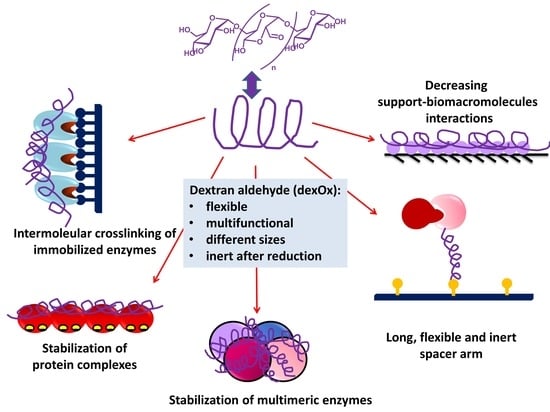
Dextran Aldehyde in Biocatalysis: More Than a Mere Immobilization System
Abstract
1. Introduction
2. Dextran and Dextran Aldehyde (DexOx)
2.1. Dextran
2.2. Dextran Aldehyde (DexOx)
2.3. Coupling of Enzymes to DexOx
3. Aplications of Dextran and DexOx in Biocatalysis
3.1. Glycosylation of Enzymes and Promotion of Intramolecular Crosslinkings
3.2. Dextran Aldehyde in the Preparation of Crosslinked Enzyme Aggregates (CLEAS)
3.3. Dextran Aldehyde for Controlling the Adsorption of Biomacromolecules on Support Surfaces
3.4. Dextran Aldehyde as Active Group and Spacer Arm in the Support
3.5. Generation of Hydrophilic Microenvironments in Immobilized Enzymes
3.6. Stabilization of Multimeric Enzymes
- The polymer size and the presence of multiple reactive groups make the one-point chemical modification less probable than the desired crosslinking.
- Similarly, also caused by the number and size of reactive groups, it is not necessary that the distance between the chemical groups involved in the crosslinking must fit one specific size.
- The flexibility of dextran allows one single molecule of dextran to react with a large percentage of the enzyme surface.
3.7. Enzyme Coimmobilization Using DexOx as Glue
3.8. -Use of dexOx for Intermolecular Crosslinking of Immobilized Enzyme Molecules
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hoyos, P.; Hernáiz, M.J.; Alcántara, A.R. Biocatalyzed production of fine chemicals. In Reference Module in Life Sciences; Elsevier: Amsterdam, The Netherlands, 2017. [Google Scholar] [CrossRef]
- Sheldon, R.A.; Pereira, P.C. Biocatalysis engineering: The big picture. Chem. Soc. Rev. 2017, 46, 2678–2691. [Google Scholar] [CrossRef]
- Alcántara, A.R. Biotransformations in drug synthesis: A green and powerful tool for medicinal chemistry. J. Med. Chem. Drug Des. 2018, 1, 1–7. [Google Scholar]
- Devine, P.N.; Howard, R.M.; Kumar, R.; Thompson, M.P.; Truppo, M.D.; Turner, N.J. Extending the application of biocatalysis to meet the challenges of drug development. Nat. Rev. Chem. 2018, 2, 409–421. [Google Scholar] [CrossRef]
- Hoyos, P.; Pace, V.; Alcántara, A.R. Chiral building blocks for drugs synthesis via biotransformations. In Asymmetric Synthesis of Drugs and Natural Products; Nag, A., Ed.; CRC Press: Boca Raton, FL, USA, 2018; pp. 346–448. [Google Scholar]
- Patel, R.N. Biocatalysis for synthesis of pharmaceuticals. Bioorg. Med. Chem. 2018, 26, 1252–1274. [Google Scholar] [CrossRef] [PubMed]
- Rosenthal, K.; Lütz, S. Recent developments and challenges of biocatalytic processes in the pharmaceutical industry. Curr. Opin. Green Sustain. Chem. 2018, 11, 58–64. [Google Scholar] [CrossRef]
- Sheldon, R.A.; Brady, D. The limits to biocatalysis: Pushing the envelope. Chem. Comm. 2018, 54, 6088–6104. [Google Scholar] [CrossRef] [PubMed]
- Sheldon, R.A.; Woodley, J.M. Role of Biocatalysis in Sustainable Chemistry. Chem. Rev. 2018, 118, 801–838. [Google Scholar] [CrossRef]
- Sun, H.; Zhang, H.; Ang, E.L.; Zhao, H. Biocatalysis for the synthesis of pharmaceuticals and pharmaceutical intermediates. Bioorg. Med. Chem. 2018, 26, 1275–1284. [Google Scholar] [CrossRef]
- Turner, N.J.; Kumar, R. Editorial overview: Biocatalysis and biotransformation: The golden age of biocatalysis. Curr. Opin. Chem. Biol. 2018, 43, A1–A3. [Google Scholar] [CrossRef]
- Adams, J.P.; Brown, M.J.B.; Diaz-Rodriguez, A.; Lloyd, R.C.; Roiban, G.D. Biocatalysis: A Pharma Perspective. Adv. Synth. Catal. 2019, in press. [Google Scholar] [CrossRef]
- Foley, A.M.; Maguire, A.R. The impact of recent developments in technologies which enable the increased use of biocatalysts. Eur. J. Org. Chem. 2019, in press. [Google Scholar] [CrossRef]
- Sheldon, R.A.; Brady, D. Broadening the scope of biocatalysis in sustainable organic synthesis. ChemSusChem 2019, in press. [Google Scholar] [CrossRef] [PubMed]
- Woodley, J.M. Accelerating the implementation of biocatalysis in industry. Appl. Microbiol. Biotechnol. 2019, in press. [Google Scholar] [CrossRef] [PubMed]
- Schoemaker, H.E.; Mink, D.; Wubbolts, M.G. Dispelling the myths—Biocatalysis in industrial synthesis. Science 2003, 299, 1694–1697. [Google Scholar] [CrossRef] [PubMed]
- Robles-Medina, A.; Gonzalez-Moreno, P.A.; Esteban-Cerdan, L.; Molina-Grima, E. Biocatalysis: Towards ever greener biodiesel production. Biotechnol. Adv. 2009, 27, 398–408. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, C.B.; Madeira, J.V.; Macedo, G.A. Biocatalysis combined with physical technologies for development of a green biodiesel process. Renew. Sust. Energ. Rev. 2014, 33, 333–343. [Google Scholar] [CrossRef]
- Budzaki, S.; Miljic, G.; Tisma, M.; Sundaram, S.; Hessel, V. Is there a future for enzymatic biodiesel industrial production in microreactors? Appl. Energy 2017, 201, 124–134. [Google Scholar] [CrossRef]
- Amini, Z.; Ilham, Z.; Ong, H.C.; Mazaheri, H.; Chen, W.H. State of the art and prospective of lipase-catalyzed transesterification reaction for biodiesel production. Energy Conv. Manag. 2017, 141, 339–353. [Google Scholar] [CrossRef]
- Tacias-Pascacio, V.G.; Torrestiana-Sanchez, B.; Dal Magro, L.; Virgen-Ortiz, J.J.; Suarez-Ruiz, F.J.; Rodrigues, R.C.; Fernandez-Lafuente, R. Comparison of acid, basic and enzymatic catalysis on the production of biodiesel after RSM optimization. Renew. Energy 2019, 135, 1–9. [Google Scholar] [CrossRef]
- Oort, M.V.; Whitehurst, R.J. Enzymes in Food Technology, 2th ed.; Wiley-Blackwell: West Sussex, UK, 2010. [Google Scholar]
- Pandey, A.; Du, G.; Sanromán, M.A.N.; Soccol, C.R.; Dussap, C.-G. Current Developments in Biotechnology and Bioengineering: Food and Beverages Industry; Elsevier Science: Amsterdam, The Netherlands, 2016. [Google Scholar]
- Kuddus, M.; Kahali; Kuddus. Enzymes in Food Technology, 1st ed.; Springer: Singapore, 2018. [Google Scholar]
- Raveendran, S.; Parameswaran, B.; Ummalyma, S.B.; Abraham, A.; Mathew, A.K.; Madhavan, A.; Rebello, S.; Pandey, A. Applications of microbial enzymes in food industry. Food Technol. Biotechnol. 2018, 56, 16–30. [Google Scholar] [CrossRef]
- Nguyen, H.H.; Lee, S.H.; Lee, U.J.; Fermin, C.D.; Kim, M. Immobilized enzymes in biosensor applications. Materials 2019, 12, 121. [Google Scholar] [CrossRef] [PubMed]
- Monteiro, T.; Almeida, M.G. Electrochemical enzyme biosensors revisited: Old solutions for new problems. Crit. Rev. Anal. Chem. 2019, 49, 44–66. [Google Scholar] [CrossRef] [PubMed]
- Gahlaut, A.; Hooda, V.; Gothwal, A.; Hooda, V. Enzyme-based ultrasensitive electrochemical biosensors for rapid assessment of nitrite toxicity: Recent advances and perspectives. Crit. Rev. Anal. Chem. 2019, 49, 32–43. [Google Scholar] [CrossRef] [PubMed]
- Campaña, A.L.; Florez, S.L.; Noguera, M.J.; Fuentes, O.P.; Puentes, P.R.; Cruz, J.C.; Osma, J.F. Enzyme-based electrochemical biosensors for microfluidic platforms to detect pharmaceutical residues in wastewater. Biosensors 2019, 9, 41. [Google Scholar] [CrossRef] [PubMed]
- Husain, Q. Carbon nanotubes mediated immobilized glucose oxidase biosensors as an effective and sensitive analytical tool. Biointerface Res. Appl. Chem. 2018, 8, 3060–3074. [Google Scholar]
- Luong, J.H.T.; Glennon, J.D.; Gedanken, A.; Vashist, S.K. Achievement and assessment of direct electron transfer of glucose oxidase in electrochemical biosensing using carbon nanotubes, graphene, and their nanocomposites. Microchim. Acta 2017, 184, 369–388. [Google Scholar] [CrossRef]
- Cormode, D.P.; Gao, L.Z.; Koo, H. Emerging biomedical applications of enzyme-like catalytic nanomaterials. Trends Biotechnol. 2018, 36, 15–29. [Google Scholar] [CrossRef] [PubMed]
- Bakunina, I.Y.; Balabanova, L.A.; Pennacchio, A.; Trincone, A. Hooked on alpha-D-galactosidases: From biomedicine to enzymatic synthesis. Crit. Rev. Biotechnol. 2016, 36, 233–245. [Google Scholar] [CrossRef] [PubMed]
- Iyer, P.V.; Ananthanarayan, L. Enzyme stability and stabilization—Aqueous and non-aqueous environment. Process Biochem. 2008, 43, 1019–1032. [Google Scholar] [CrossRef]
- Suplatov, D.; Voevodin, V.; Svedas, V. Robust enzyme design: Bioinformatic tools for improved protein stability. Biotechnol. J. 2015, 10, 344–355. [Google Scholar] [CrossRef] [PubMed]
- Reyes-De-Corcuera, J.I.; Olstad, H.E.; Garcia-Torres, R. Stability and stabilization of enzyme biosensors: The key to successful application and commercialization. In Annual Review of Food Science and Technology; Doyle, M.P., Klaenhammer, T.R., Eds.; Annual Reviews: Palo Alto, CA, USA, 2018; Volume 9, pp. 293–322. [Google Scholar]
- Hernandez, K.; Berenguer-Murcia, A.; Rodrigues, R.C.; Fernandez-Lafuente, R. Hydrogen peroxide in biocatalysis. A dangerous liaison. Curr. Org. Chem. 2012, 16, 2652–2672. [Google Scholar] [CrossRef]
- Rocchitta, G.; Spanu, A.; Babudieri, S.; Latte, G.; Madeddu, G.; Galleri, G.; Nuvoli, S.; Bagella, P.; Demartis, M.I.; Fiore, V.; et al. Enzyme biosensors for biomedical applications: Strategies for safeguarding analytical performances in biological fluids. Sensors 2016, 16, 780. [Google Scholar] [CrossRef] [PubMed]
- Deng, S.; Fu, A.L.; Junaid, M.; Wang, Y.; Yin, Q.; Fu, C.; Liu, L.; Su, D.S.; Bian, W.P.; Pei, D.S. Nitrogen-doped graphene quantum dots (N-GQDs) perturb redox-sensitive system via the selective inhibition of antioxidant enzyme activities in zebrafish. Biomaterials 2019, 206, 61–72. [Google Scholar] [CrossRef] [PubMed]
- Hdiouech, S.; Bruna, F.; Batisson, I.; Besse-Hoggan, P.; Prevot, V.; Mousty, C. Amperometric detection of the herbicide mesotrione based on competitive reactions at nitroreductase@layered double hydroxide bioelectrode. J. Electroanal. Chem. 2019, 835, 324–328. [Google Scholar] [CrossRef]
- Kuwahara, T.; Ogawa, K.; Sumita, D.; Kondo, M.; Shimomura, M. Amperometric glucose sensing with polyaniline/poly(acrylic acid) composite film bearing glucose oxidase and catalase based on competitive oxygen consumption reactions. J. Electroanal. Chem. 2018, 811, 62–67. [Google Scholar] [CrossRef]
- Daems, D.; Lu, J.; Delport, F.; Marien, N.; Orbie, L.; Aernouts, B.; Adriaens, I.; Huybrechts, T.; Saeys, W.; Spasic, D.; et al. Competitive inhibition assay for the detection of progesterone in dairy milk using a fiber optic SPR biosensor. Anal. Chim. Acta 2017, 950, 1–6. [Google Scholar] [CrossRef]
- Hopkins, S.P.; Pant, J.; Goudie, M.J.; Schmiedt, C.; Handa, H. Achieving long-term biocompatible silicone via covalently immobilized S-nitroso-N-acetylpenicillamine (SNAP) that exhibits 4 months of sustained nitric oxide release. ACS Appl. Mater. Interfaces 2018, 10, 27316–27325. [Google Scholar] [CrossRef] [PubMed]
- Grano, V.; Diano, N.; Portaccio, M.; Bencivenga, U.; De Maio, A.; De Santo, N.; Perna, A.; Salamino, F.; Mita, D.G. The alpha (1)-antitrypsin/elastase complex as an experimental model for hemodialysis in acute catabolic renal failure, extracorporeal blood circulation and cardiocirculatory bypass. Int. J. Artif. Organs 2002, 25, 297–305. [Google Scholar] [CrossRef] [PubMed]
- Grovender, E.A.; Cooney, C.L.; Langer, R.S.; Ameer, G.A. Modeling of the mixing behavior of the novel fluidized extracorporeal immunoadsorber. Chem. Eng. Sci. 2001, 56, 5437–5441. [Google Scholar] [CrossRef]
- Ameer, G.A.; Harmon, W.; Sasisekharan, R.; Langer, R. Investigation of a whole blood fluidized bed Taylor-Couette flow device for enzymatic heparin neutralization. Biotechnol. Bioeng. 1999, 62, 602–608. [Google Scholar] [CrossRef]
- Sheldon, R.A.; van Pelt, S. Enzyme immobilisation in biocatalysis: Why, what and how. Chem. Soc. Rev. 2013, 42, 6223–6235. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, R.C.; Ortiz, C.; Berenguer-Murcia, A.; Torres, R.; Fernandez-Lafuente, R. Modifying enzyme activity and selectivity by immobilization. Chem. Soc. Rev. 2013, 42, 6290–6307. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.K.; Tiwari, M.K.; Singh, R.; Lee, J.K. From protein engineering to immobilization: Promising strategies for the upgrade of industrial enzymes. Int. J. Mol. Sci. 2013, 14, 1232–1277. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, O.; Ortiz, C.; Berenguer-Murcia, A.; Torres, R.; Rodrigues, R.C.; Fernandez-Lafuente, R. Strategies for the one-step immobilization-purification of enzymes as industrial biocatalysts. Biotechnol. Adv. 2015, 33, 435–456. [Google Scholar] [CrossRef] [PubMed]
- Bezerra, C.S.; Lemos, C.; de Sousa, M.; Goncalves, L.R.B. Enzyme immobilization onto renewable polymeric matrixes: Past, present, and future trends. J. Appl. Polym. Sci. 2015, 132, 15. [Google Scholar] [CrossRef]
- Eş, I.; Vieira, J.D.G.; Amaral, A.C. Principles, techniques, and applications of biocatalyst immobilization for industrial application. Appl. Microbiol. Biotechnol. 2015, 99, 2065–2082. [Google Scholar] [CrossRef] [PubMed]
- Gutarra, M.L.E.; Miranda, L.S.M.; de Souza, R.O.M.A. Chapter 4—Enzyme Immobilization for Organic Synthesis A2—Goswami, Animesh. In Organic Synthesis Using Biocatalysis; Stewart, J.D., Ed.; Academic Press; Elsevier: Amsterdam, The Netherlands, 2016; pp. 99–126. [Google Scholar] [CrossRef]
- Schmidt-Dannert, S.; Zhang, G.Q.; Johnston, T.; Quin, M.B.; Schmidt-Dannert, C. Building a toolbox of protein scaffolds for future immobilization of biocatalysts. Appl. Microbiol. Biotechnol. 2018, 102, 8373–8388. [Google Scholar] [CrossRef] [PubMed]
- DiCosimo, R.; McAuliffe, J.; Poulose, A.J.; Bohlmann, G. Industrial use of immobilized enzymes. Chem. Soc. Rev. 2013, 42, 6437–6474. [Google Scholar] [CrossRef]
- Cantone, S.; Ferrario, V.; Corici, L.; Ebert, C.; Fattor, D.; Spizzo, P.; Gardossi, L. Efficient immobilisation of industrial biocatalysts: Criteria and constraints for the selection of organic polymeric carriers and immobilisation methods. Chem. Soc. Rev. 2013, 42, 6262–6276. [Google Scholar] [CrossRef]
- Almeida, O.G.G.; De Martinis, E.C.P. Bioinformatics tools to assess metagenomic data for applied microbiology. Appl. Microbiol. Biotechnol. 2019, 103, 69–82. [Google Scholar] [CrossRef]
- Ngara, T.R.; Zhang, H.J. Recent Advances in Function-based Metagenomic Screening. Genom. Proteom. Bioinform. 2019, 16, 405–415. [Google Scholar] [CrossRef] [PubMed]
- Bilal, T.; Malik, B.; Hakeem, K.R. Metagenomic analysis of uncultured microorganisms and their enzymatic attributes. J. Microbiol. Methods 2018, 155, 65–69. [Google Scholar] [CrossRef] [PubMed]
- Van der Helm, E.; Genee, H.J.; Sommer, M.O.A. The evolving interface between synthetic biology and functional metagenomics. Nat. Chem. Biol. 2018, 14, 752–759. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.H.; Alper, H.S. Strategies for directed and adapted evolution as part of microbial strain engineering. J. Chem. Technol. Biotechnol. 2019, 94, 366–376. [Google Scholar] [CrossRef]
- Zeymer, C.; Hilvert, D. Directed evolution of protein catalysts. In Annual Review of Biochemistry; Kornberg, R.D., Ed.; Annual Reviews: Palo Alto, CA USA, 2018; Volume 87, pp. 131–157. [Google Scholar]
- Bornscheuer, U.T.; Hauer, B.; Jaeger, K.E.; Schwaneberg, U. Directed evolution empowered redesign of natural proteins for the sustainable production of chemicals and pharmaceuticals. Angew. Chem. Int. Edit. 2019, 58, 36–40. [Google Scholar] [CrossRef] [PubMed]
- Li, G.Y.; Wang, J.B.; Reetz, M.T. Biocatalysts for the pharmaceutical industry created by structure-guided directed evolution of stereoselective enzymes. Bioorg. Med. Chem. 2018, 26, 1241–1251. [Google Scholar] [CrossRef]
- Arnold, F.H. Directed Evolution: Bringing New Chemistry to Life. Angew. Chem. Int. Edit. 2018, 57, 4143–4148. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, K.; Fernandez-Lafuente, R. Control of protein immobilization: Coupling immobilization and site-directed mutagenesis to improve biocatalyst or biosensor performance. Enzym. Microb. Technol. 2011, 48, 107–122. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, O.; Torres, R.; Ortiz, C.; Berenguer-Murcia, A.; Rodrigues, R.C.; Fernandez-Lafuente, R. Heterofunctional supports in enzyme immobilization: From traditional immobilization protocols to opportunities in tuning enzyme properties. Biomacromolecules 2013, 14, 2433–2462. [Google Scholar] [CrossRef]
- Cowan, D.A.; Fernandez-Lafuente, R. Enhancing the functional properties of thermophilic enzymes by chemical modification and immobilization. Enzym. Microb. Technol. 2011, 49, 326–346. [Google Scholar] [CrossRef]
- Rodrigues, R.C.; Berenguer-Murcia, A.; Fernandez-Lafuente, R. Coupling chemical modification and immobilization to improve the catalytic performance of enzymes. Adv. Synth. Catal. 2011, 353, 2216–2238. [Google Scholar] [CrossRef]
- Rueda, N.; dos Santos, J.C.S.; Ortiz, C.; Torres, R.; Barbosa, O.; Rodrigues, R.C.; Berenguer-Murcia, A.; Fernandez-Lafuente, R. Chemical modification in the design of immobilized enzyme biocatalysts: Drawbacks and opportunities. Chem. Rec. 2016, 16, 1436–1455. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Lafuente, R. Stabilization of multimeric enzymes: Strategies to prevent subunit dissociation. Enzym. Microb. Technol. 2009, 45, 405–418. [Google Scholar] [CrossRef]
- BeMiller, J.N. Dextran. In Encyclopedia of Food Sciences and Nutrition, 2nd ed.; Caballero, B., Ed.; Academic Press: Oxford, UK, 2003; pp. 1772–1773. [Google Scholar] [CrossRef]
- IUPAC. Dextrans. Available online: https://goldbook.iupac.org/html/D/D01655.html (accessed on 6 May 2019).
- Sidebotham, R.L. Dextrans. In Advances in Carbohydrate Chemistry and Biochemistry; Tipson, R.S., Horton, D., Eds.; Academic Press: Cambridge, MA, USA, 1974; Volume 30, pp. 371–444. [Google Scholar]
- Naessens, M.; Cerdobbel, A.; Soetaert, W.; Vandamme, E.J. Leuconostoc dextransucrase and dextran: Production, properties and applications. J. Chem. Technol. Biotechnol. 2005, 80, 845–860. [Google Scholar] [CrossRef]
- Bounaix, M.S.; Gabriel, V.; Robert, H.; Morel, S.; Rernaud-Simeon, M.; Gabriel, B.; Fontagne-Faucher, C. Characterization of glucan-producing Leuconostoc strains isolated from sourdough. Int. J. Food Microbiol. 2010, 144, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Fusco, V.; Quero, G.M.; Cho, G.S.; Kabisch, J.; Meske, D.; Neve, H.; Bockelmann, W.; Franz, C. The genus Weissella: Taxonomy, ecology and biotechnological potential. Front. Microbiol. 2015, 6, 22. [Google Scholar] [CrossRef] [PubMed]
- Bounaix, M.S.; Robert, H.; Gabriel, V.; Morel, S.; Remaud-Simeon, M.; Gabriel, B.; Fontagne-Faucher, C. Characterization of dextran-producing Weissella strains isolated from sourdoughs and evidence of constitutive dextransucrase expression. FEMS Microbiol. Lett. 2010, 311, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, Z.; Ahmad, A. Biopolymer Produced by the Lactic Acid Bacteria: Production and Practical Application; Academic Press Ltd-Elsevier Science Ltd.: London, UK, 2017; Volume 5, pp. 217–257. [Google Scholar]
- Van Hijum, S.; Kralj, S.; Ozimek, L.K.; Dijkhuizen, L.; van Geel-Schutten, I.G.H. Structure-function relationships of glucansucrase and fructansucrase enzymes from lactic acid bacteria. Microbiol. Mol. Biol. Rev. 2006, 70, 157–176. [Google Scholar] [CrossRef] [PubMed]
- Stepanov, N.A.; Senko, O.V.; Efremenko, E.N. Biocatalytic production of extracellular exopolysaccharide dextran synthesized by cells of Leuconostoc mesenteroides. Catal. Ind. 2017, 9, 339–343. [Google Scholar] [CrossRef]
- Siddiqui, N.N.; Aman, A.; Silipo, A.; Qader, S.A.U.; Molinaro, A. Structural analysis and characterization of dextran produced by wild and mutant strains of Leuconostoc mesenteroides. Carbohydr. Polym. 2014, 99, 331–338. [Google Scholar] [CrossRef]
- Leung, A.D.; Wong, K.H.K.; Tien, J. Plasma expanders stabilize human microvessels in microfluidic scaffolds. J. Biomed. Mater. Res. Part A 2012, 100A, 1815–1822. [Google Scholar] [CrossRef] [PubMed]
- McCahon, R.; Hardman, J. Pharmacology of plasma expanders. Anaesth. Intensive Care 2017, 18, 418–420. [Google Scholar] [CrossRef]
- Khalikova, E.; Susi, P.; Korpela, T. Microbial dextran-hydrolyzing enzymes: Fundamentals and applications. Microbiol. Mol. Biol. Rev. 2005, 69, 306–325. [Google Scholar] [CrossRef] [PubMed]
- Varshosaz, J. Dextran conjugates in drug delivery. Expert Opin. Drug Deliv. 2012, 9, 509–523. [Google Scholar] [CrossRef] [PubMed]
- Kirschning, A.; Dibbert, N.; Drager, G. Chemical Functionalization of Polysaccharides-Towards Biocompatible Hydrogels for Biomedical Applications. Chem. Eur. J. 2018, 24, 1231–1240. [Google Scholar] [CrossRef] [PubMed]
- Peesa, J.P.; Yalavarthi, P.R.; Rasheed, A.; Mandava, V.B.R. A perspective review on role of novel NSAID prodrugs in the management of acute inflammation. J. Acute Dis. 2016, 5, 364–381. [Google Scholar] [CrossRef]
- Larsen, C. Dextran prodrugs—Structure and stability in relation to therapeutic activity. Adv. Drug. Deliv. Rev. 1989, 3, 103–154. [Google Scholar] [CrossRef]
- Engelmann, G.; Jobmann, M.; Rafler, G. Dextran carbamates—Materials for microencapsulation. Ind. Crops Prod. 2004, 20, 37–48. [Google Scholar] [CrossRef]
- Van Tomme, S.R.; Hennink, W.E. Biodegradable dextran hydrogels for protein delivery applications. Expert Rev. Med. Devices 2007, 4, 147–164. [Google Scholar] [CrossRef]
- Mukwaya, V.; Wang, C.L.; Dou, H.J. Saccharide-based nanocarriers for targeted therapeutic and diagnostic applications. Polym. Int. 2019, 68, 306–319. [Google Scholar] [CrossRef]
- Huang, S.Y.; Huang, G.L. Preparation and drug delivery of dextran-drug complex. Drug Deliv. 2019, 26, 10. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.L.; Huang, H.L. Application of dextran as nanoscale drug carriers. Nanomedicine 2018, 13, 3149–3158. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A.; Bandopadhyay, R. Use of dextran nanoparticle: A paradigm shift in bacterial exopolysaccharide based biomedical applications. Int. J. Biol. Macromol. 2016, 87, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Wang, L.N.; Lin, X.; Shen, L.; Feng, Y. Drug delivery for bioactive polysaccharides to improve their drug-like properties and curative efficacy. Drug Deliv. 2017, 24, 70–80. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Gong, Y.S.; Gern, J.E.; Ikeda, S.; Lucey, J.A. Glycation of whey protein with dextrans of different molar mass: Effect on immunoglobulin E-binding capacity with blood sera obtained from patients with cow milk protein allergy. J. Dairy Sci. 2018, 101, 6823–6834. [Google Scholar] [CrossRef] [PubMed]
- Nodake, Y.; Fukumoto, S.; Fukasawa, M.; Sakakibara, R.; Yamasaki, N. Reduction of the Immunogenicity of beta-Lactoglobulin from Cow’s Milk by Conjugation with a Dextran Derivative. Biosci. Biotech. Bioch. 2010, 74, 721–726. [Google Scholar] [CrossRef] [PubMed]
- Maia, J.; Evangelista, M.; Gil, H.; Ferreira, L. Dextran-based materials for biomedical applications. In Carbohydrates Applications in Medicine; Gil, M.H., Ed.; Research Signpost: Trivandrum, India, 2014; pp. 31–53. [Google Scholar]
- Tiwari, S.; Patil, R.; Bahadur, P. Polysaccharide Based Scaffolds for Soft Tissue Engineering Applications. Polymers 2019, 11, 1. [Google Scholar] [CrossRef] [PubMed]
- Wolf, M.; Koch, T.A.; Bregman, D.B. Effects of iron deficiency anemia and its treatment on fibroblast growth factor 23 and phosphate homeostasis in women. J. Bone Miner. Res. 2013, 28, 1793–1803. [Google Scholar] [CrossRef] [PubMed]
- Auerbach, M.; Pappadakis, J.A.; Bahrain, H.; Auerbach, S.A.; Ballard, H.; Dahl, N.V. Safety and efficacy of rapidly administered (one hour) one gram of low molecular weight iron dextran (INFeD) for the treatment of iron deficient anemia. Am. J. Hematol. 2011, 86, 860–862. [Google Scholar] [CrossRef]
- Alhareth, K.; Vauthier, C.; Bourasset, F.; Gueutin, C.; Ponchel, G.; Moussa, F. Conformation of surface-decorating dextran chains affects the pharmacokinetics and biodistribution of doxorubicin-loaded nanoparticles. Eur. J. Pharm. Biopharm. 2012, 81, 453–457. [Google Scholar] [CrossRef]
- Merdan, T.; Kopecek, J.; Kissel, T. Prospects for cationic polymers in gene and oligonucleotide therapy against cancer. Adv. Drug Deliv. Rev. 2002, 54, 715–758. [Google Scholar] [CrossRef]
- Morille, M.; Passirani, C.; Vonarbourg, A.; Clavreul, A.; Benoit, J.P. Progress in developing cationic vectors for non-viral systemic gene therapy against cancer. Biomaterials 2008, 29, 3477–3496. [Google Scholar] [CrossRef] [PubMed]
- Lachelt, U.; Wagner, E. Nucleic acid therapeutics using polyplexes: A journey of 50 years (and beyond). Chem. Rev. 2015, 115, 11043–11078. [Google Scholar] [CrossRef] [PubMed]
- Togo, Y.; Takahashi, K.; Saito, K.; Kiso, H.; Huang, B.; Tsukamoto, H.; Hyon, S.-H.; Bessho, K. Aldehyded dextran and epsilon-poly (L-lysine) hydrogel as nonviral gene carrier. Stem Cells Int. 2013. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.J.; Rekha, M.R.; Sharma, C.P. Unraveling the intracellular efficacy of dextran-histidine polycation as an efficient nonviral gene delivery system. Mol. Pharm. 2012, 9, 121–134. [Google Scholar] [CrossRef] [PubMed]
- Chu, M.; Dong, C.Y.; Zhu, H.Y.; Cai, X.J.; Dong, H.Q.; Ren, T.B.; Su, J.S.; Li, Y.Y. Biocompatible polyethylenimine-graft-dextran catiomer for highly efficient gene delivery assisted by a nuclear targeting ligand. Polym. Chem. 2013, 4, 2528–2539. [Google Scholar] [CrossRef]
- Ochrimenko, S.; Vollrath, A.; Tauhardt, L.; Kempe, K.; Schubert, S.; Schubert, U.S.; Fischer, D. Dextran-graft-linear poly(ethylene imine)s for gene delivery: Importance of the linking strategy. Carbohydr. Polym. 2014, 113, 597–606. [Google Scholar] [CrossRef]
- Kloypan, C.; Prapan, A.; Suwannasom, N.; Chaiwaree, S.; Kaewprayoon, W.; Steffen, A.; Xiong, Y.; Baisaeng, N.; Georgieva, R.; Baumler, H. Improved oxygen storage capacity of haemoglobin submicron particles by one-pot formulation. Artif. Cells Nanomed. Biotechnol. 2018, 46, S964–S972. [Google Scholar] [CrossRef]
- Kloypan, C.; Suwannasom, N.; Chaiwaree, S.; Prapan, A.; Smuda, K.; Baisaeng, N.; Pruss, A.; Georgieva, R.; Baumler, H. In-Vitro haemocompatibility of dextran-protein submicron particles. Artif. Cells Nanomed. Biotechnol. 2019, 47, 241–249. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, Y.; Zhang, S.; You, G.X.; Li, P.L.; Wang, Q.; Yu, W.L.; Hu, T.; Zhou, H.; Zhao, L. In vitro and in vivo investigation of the novel Dex-bHb as oxygen carriers. Artif. Cells Nanomed. Biotechnol. 2018, 46, S133–S137. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, S.; Zhang, J.; Yu, W.L.; Gao, D.W.; Wang, Q.; You, G.X.; Hu, T.; Zhao, L.; Zhou, H. Structural, functional and physiochemical properties of dextran-bovine hemoglobin conjugate as a hemoglobin-based oxygen carrier. Process Biochem. 2017, 60, 67–73. [Google Scholar] [CrossRef]
- Dossabhoy, N.R.; Turley, S.; Gascoyne, R.; Tapolyai, M.; Sulaiman, K. Safety of total dose iron dextran infusion in geriatric patients with chronic kidney disease and iron deficiency anemia. Ren. Fail. 2014, 36, 1033–1037. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cooke, M.; Lamplugh, A.; Naudeer, S.; Edey, M.; Bhandari, S. Efficacy and tolerability of accelerated-dose low-molecular-weight iron dextran (cosmofer) in patients with chronic kidney disease. Am. J. Nephrol. 2012, 35, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Yessayan, L.; Sandhu, A.; Besarab, A.; Yessayan, A.; Frinak, S.; Zasuwa, G.; Yee, J. Intravenous iron dextran as a component of anemia management in chronic kidney disease: A report of safety and efficacy. Int. J. Nephrol. 2013, 2013, 9. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Wang, Y.; You, G.X.; Wang, Q.; Zhang, S.; Yu, W.L.; Hu, T.; Zhao, L.; Zhou, H. Conjugation with 20kDa dextran decreases the autoxidation rate of bovine hemoglobin. Artif. Cells Nanomed. Biotechnol. 2018, 46, 1436–1443. [Google Scholar] [CrossRef] [PubMed]
- Malaprade, L. The oxydation of certain polyhydric alcohol by the application of periodic acid. C. R. Hebd. Seances Acad. Sci. 1928, 186, 382–385. [Google Scholar]
- Wang, Z. Malaprade Reaction. In Comprehensive Organic Name Reactions and Reagents; Wang, Z., Ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2010. [Google Scholar] [CrossRef]
- Kent, P.W. Periodate oxidation in the study of the structure of dextrans. Science 1949, 110, 689–690. [Google Scholar] [CrossRef] [PubMed]
- Dimler, R.J.; Wolff, I.A.; Sloan, J.W.; Rist, C.E. Interpretation of periodate oxidation data on degraded dextran. J. Am. Chem. Soc. 1955, 77, 6568–6573. [Google Scholar] [CrossRef]
- Maia, J.; Carvalho, R.A.; Coelho, J.F.J.; Simoes, P.N.; Gil, M.H. Insight on the periodate oxidation of dextran and its structural vicissitudes. Polymer 2011, 52, 258–265. [Google Scholar] [CrossRef]
- Jeanes, A.; Wilham, C.A. Periodate oxidation of dextran. J. Am. Chem. Soc. 1950, 72, 2655–2657. [Google Scholar] [CrossRef]
- Ishak, M.F.; Painter, T.J. Kinetic evidence for hemiacetal formation during oxidation of dextran in aqueous periodate. Carbohydr. Res. 1978, 64, 189–197. [Google Scholar] [CrossRef]
- Drobchenko, S.N.; IsaevaIvanova, L.S.; Kleiner, A.R.; Eneyskaya, E.V. Aldo-enol transition in periodate-oxidized dextrans. Carbohydr. Res. 1996, 280, 171–176. [Google Scholar] [CrossRef]
- Yu, R.J.; Bishop, C.T. Novel oxidations of methyl glycopyranosides by periodic acid in dimethyl sulfoxide. Can. J. Chem. 1967, 45, 2195–2203. [Google Scholar] [CrossRef]
- Maia, J.; Ferreira, L.; Carvalho, R.; Ramos, M.A.; Gil, M.H. Synthesis and characterization of new injectable and degradable dextran-based hydrogels. Polymer 2005, 46, 9604–9614. [Google Scholar] [CrossRef]
- Bouhadir, K.H.; Hausman, D.S.; Mooney, D.J. Synthesis of cross-linked poly (aldehyde guluronate) hydrogels. Polymer 1999, 40, 3575–3584. [Google Scholar] [CrossRef]
- Lenders, J.P.; Crichton, R.R. Thermal stabilization of amylolytic enzymes by covalent coupling to soluble polysaccharides. Biotechnol. Bioeng. 1984, 26, 1343–1351. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Heindel, N.D. Determination of degree of substitution of formyl groups in polyaldehyde dextran by the hydroxylamine hydrochloride method. Pharm. Res. 1991, 8, 400–402. [Google Scholar] [CrossRef] [PubMed]
- Al-Noor, T.; El-Ajaily, M.M.; Maihub, A.A. A Review: On Chemistry Important of Schiff’s Bases Complexes, 1st ed.; Noor Publishing: Saarbrücken, Germany, 2017. [Google Scholar]
- Hermanson, G.T. Bioconjugate Techniques; Elsevier Science & Technology: San Diego, CA, USA, 2013. [Google Scholar]
- Lundblad, R.L.; Lundblad, R.L. Modification of Proteins with Reducing Agents; CRC Press-Taylor & Francis Group: Boca Raton, FL, USA, 2014; pp. 185–216. [Google Scholar]
- Tarasevich, V.A. Reductive amination of oxygen-containing organic compounds. Russ. Chem. Rev. 1999, 68, 55–72. [Google Scholar] [CrossRef]
- Baxter, E.W.; Reitz, A.B. Reductive aminations of carbonyl compounds with borohydride and borane reducing agents. In Organic Reactions; Paquette, L., Ed.; John Wiley & Sons: Hoboken, NJ, USA, 2002; Volume 59, pp. 1–57. [Google Scholar]
- Sato, S.; Sakamoto, T.; Miyazawa, E.; Kikugawa, Y. One-pot reductive amination of aldehydes and ketones with alpha-picoline-borane in methanol, in water, and in neat conditions. Tetrahedron 2004, 60, 7899–7906. [Google Scholar] [CrossRef]
- Ruhaak, L.R.; Steenvoorden, E.; Koeleman, C.A.M.; Deelder, A.M.; Wuhrer, M. 2-Picoline-borane: A non-toxic reducing agent for oligosaccharide labeling by reductive amination. Proteomics 2010, 10, 2330–2336. [Google Scholar] [CrossRef] [PubMed]
- Cosenza, V.A.; Navarro, D.A.; Stortz, C.A. Usage of alpha-picoline borane for the reductive amination of carbohydrates. Arkivoc 2011, 182–194. [Google Scholar]
- Kawase, Y.; Yamagishi, T.; Kato, J.Y.; Kutsuma, T.; Kataoka, T.; Iwakuma, T.; Yokomatsu, T. Reductive alkylation of hydrazine derivatives with alpha-picoline-borane and its applications to the syntheses of useful compounds related to active pharmaceutical ingredients. Synthesis 2014, 46, 455–464. [Google Scholar]
- Orrego, A.H.; Romero-Fernandez, M.; Millan-Linares, M.D.; Yust, M.D.; Guisan, J.M.; Rocha-Martin, J. Stabilization of enzymes by multipoint covalent attachment on aldehyde-supports: 2-picoline borane as an alternative reducing agent. Catalysts 2018, 8, 333. [Google Scholar] [CrossRef]
- Ruzicka, J.; Carroll, A.D.; Lahdesmaki, I. Immobilization of proteins on agarose beads, monitored in real time by bead injection spectroscopy. Analyst 2006, 131, 799–808. [Google Scholar] [CrossRef] [PubMed]
- Chimpibul, W.; Nagashima, T.; Hayashi, F.; Nakajima, N.; Hyon, S.H.; Matsumura, K. Dextran oxidized by a Malaprade reaction shows main chain scission through a Maillard reaction triggered by Schiff base formation between aldehydes and amines. J. Polym. Sci. Pol. Chem. 2016, 54, 2254–2260. [Google Scholar] [CrossRef]
- Arslan, A.; Kuzu, H.; Altikatoglu, M. Functional stabilization of cellulase from Aspergillus niger by conjugation with dextran-aldehyde. J. Carbohydr. Chem. 2010, 29, 222–235. [Google Scholar] [CrossRef]
- Altikatoglu, M.; Kuzu, H. Water soluble covalent conjugates of Aspergillus oryzae alpha amylase with dextran sulphate and their fluorescence characteristics. Fresenius Environ. Bull. 2009, 18, 2108–2113. [Google Scholar]
- Altikatoglu, M.; Arioz, C.; Basaran, Y.; Kuzu, H. Stabilization of horseradish peroxidase by covalent conjugation with dextran aldehyde against temperature and pH changes. Cent. Eur. J. Chem. 2009, 7, 423–428. [Google Scholar] [CrossRef]
- Maksimenko, A.V.; Schechilina, Y.V.; Tischenko, E.G. Resistance of dextran-modified hyaluronidase to inhibition by heparin. Biochem. Moscow 2001, 66, 456–463. [Google Scholar] [CrossRef]
- Maksimenko, A.V.; Petrova, M.L.; Tischenko, E.G.; Schechilina, Y.V. Chemical modification of hyaluronidase regulates its inhibition by heparin. Eur. J. Pharm. Biopharm. 2001, 51, 33–38. [Google Scholar] [CrossRef]
- Venkatesh, R.; Warrier, S.R.; Shubhada, S.; Sundaram, P.V. Monopolyaldehyde-mediated, dipolyaldehyde-mediated, and polyaldehyde-mediated chemical modification of urease. In Enzym. Engineering XI; Clark, D.S., Estell, D.A., Eds.; New York Acad Sciences: New York, NY, USA, 1992; Volume 672, pp. 580–582. [Google Scholar]
- Maksimenko, A.V.; Nadirashvili, L.A.; Romaschenko, A.D.; Erkomaishvili, G.S.; Abramova, V.V.; Torchilin, V.P. Complex of papaya proteinases modified with soluble polymer and its possible medical application. J. Control. Release 1989, 10, 131–143. [Google Scholar] [CrossRef]
- Lenders, J.P.; Crichton, R.R. Chemical stabilization of glucoamylase from Aspergillus niger against thermal inactivation. Biotechnol. Bioeng. 1988, 31, 267–277. [Google Scholar] [CrossRef] [PubMed]
- Maksimenko, A.V.; Grigor’eva, E.L.; Morozkin, A.D.; Tishchenko, E.G.; Minkovskiĭ, E.B.; Torchilin, V.P. Assessment of the composition and structure of covalent complexes of superoxide dismutase with aldehyde dextran by analytical ultracentrifugation. Biokhimiia 1991, 56, 1330–1336. [Google Scholar] [PubMed]
- Maksimenko, A.V.; Grigoreva, E.L.; Bezrukavnikova, L.M.; Petrov, A.D.; Tishchenko, E.G.; Arkhipova, O.G.; Torchilin, V.P. Antifibrous action of aldehyde-dextran modified superoxide-dismutase in experimental silicosis. Bull. Exp. Biol. Med. 1991, 112, 1268–1271. [Google Scholar] [CrossRef]
- Gutarra, M.L.E.; Romero, O.; Abian, O.; Torres, F.A.G.; Freire, D.M.G.; Castro, A.M.; Guisan, J.M.; Palomo, J.M. Enzyme surface glycosylation in the solid phase: Improved activity and selectivity of Candida antarctica lipase B. ChemCatChem 2011, 3, 1902–1910. [Google Scholar] [CrossRef]
- Orrego, A.H.; Ghobadi, R.; Moreno-Perez, S.; Mendoza, A.J.; Fernandez-Lorente, G.; Guisan, J.M.; Rocha-Martin, J. Stabilization of immobilized lipases by intense intramolecular cross-linking of their surfaces by using aldehyde-dextran polymers. Int. J. Mol. Sci. 2018, 19, 553. [Google Scholar] [CrossRef] [PubMed]
- Manoel, E.A.; dos Santos, J.C.S.; Freire, D.M.G.; Rueda, N.; Fernandez-Lafuente, R. Immobilization of lipases on hydrophobic supports involves the open form of the enzyme. Enzym. Microb. Technol. 2015, 71, 53–57. [Google Scholar] [CrossRef]
- Rodrigues, R.C.; Virgen-Ortiz, J.J.; Dos Santos, J.C.S.; Berenguer-Murcia, A.; Alcantara, A.R.; Barbosa, O.; Ortiz, C.; Fernandez-Lafuente, R. Immobilization of lipases on hydrophobic supports: Immobilization mechanism, advantages, problems, and solutions. Biotechnol. Adv. 2019, in press. [Google Scholar] [CrossRef]
- Carraway, K.L.; Koshland, D.E., Jr. Carbodiimide modification of proteins. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 1972; Volume 25, pp. 616–623. [Google Scholar]
- Rodrigues, R.C.; Barbosa, O.; Ortiz, C.; Berenguer-Murcia, A.; Torres, R.; Fernandez-Lafuente, R. Amination of enzymes to improve biocatalyst performance: Coupling genetic modification and physicochemical tools. RSC Adv. 2014, 4, 38350–38374. [Google Scholar] [CrossRef]
- Yano, Y.F. Kinetics of protein unfolding at interfaces. J. Phys. Condens. Matter. 2012, 24, 503101. [Google Scholar] [CrossRef]
- Findrik, Z.; Presecki, A.V.; Vasic-Racki, D. The influence of aeration on activity and operational stability of two snake venom amino acid oxidases. Biochem. Eng. J. 2012, 60, 91–98. [Google Scholar] [CrossRef]
- Findrik, Z.; Valentovic, I.; Vasic-Racki, D. A mathematical model of oxidative deamination of amino acid catalyzed by two D-amino acid oxidases and influence of aeration on enzyme stability. Appl. Biochem. Biotechnol. 2014, 172, 3092–3105. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Fan, Y.; Sheng, J.; Sui, S.F. Induction of changes in the secondary structure of globular proteins by a hydrophobic surface. Eur. Biophys. J. 1993, 22, 201–205. [Google Scholar] [CrossRef] [PubMed]
- Ross, A.C.; Bell, G.; Halling, P.J. Organic solvent functional group effect on enzyme inactivation by the interfacial mechanism. J. Mol. Catal. B Enzym. 2000, 8, 183–192. [Google Scholar] [CrossRef]
- Ghatorae, A.S.; Guerra, M.J.; Bell, G.; Halling, P.J. New technique for monitoring interfacial inactivation of enzymes by organic solvents. In Studies in Organic Chemistry; Elsevier: Amsterdam, The Netherlands, 1993; Volume 47, pp. 329–336. [Google Scholar]
- Halling, P.J.; Ross, A.C.; Bell, G. Inactivation of enzymes at the aqueous-organic interface. In Progress in Biotechnology; Elsevier: Amsterdam, The Netherlands, 1998; Volume 15, pp. 365–372. [Google Scholar]
- Garcia-Galan, C.; Berenguer-Murcia, A.; Fernandez-Lafuente, R.; Rodrigues, R.C. Potential of different enzyme immobilization strategies to improve enzyme performance. Adv. Synth. Catal. 2011, 353, 2885–2904. [Google Scholar] [CrossRef]
- Garcia-Galan, C.; Barbosa, O.; Fernandez-Lafuente, R. Stabilization of the hexameric glutamate dehydrogenase from Escherichia coli by cations and polyethyleneimine. Enzym. Microb. Technol. 2013, 52, 211–217. [Google Scholar] [CrossRef] [PubMed]
- Bolivar, J.M.; Rocha-Martin, J.; Mateo, C.; Cava, F.; Berenguer, J.; Fernandez-Lafuente, R.; Guisan, J.M. Coating of soluble and immobilized enzymes with ionic polymers: Full stabilization of the quaternary structure of multimeric enzymes. Biomacromolecules 2009, 10, 742–747. [Google Scholar] [CrossRef] [PubMed]
- Montes, T.; Grazu, V.; López-Gallego, F.; Hermoso, J.A.; Guisán, J.M.; Fernández-Lafuente, R. Chemical modification of protein surfaces to improve their reversible enzyme immobilization on ionic exchangers. Biomacromolecules 2006, 7, 3052–3058. [Google Scholar] [CrossRef] [PubMed]
- Betancor, L.; Lopez-Gallego, F.; Hidalgo, A.; Alonso-Morales, N.; Fuentes, M.; Fernandez-Lafuente, R.; Guisan, J.M. Prevention of interfacial inactivation of enzymes by coating the enzyme surface with dextran-aldehyde. J. Biotechnol. 2004, 110, 201–207. [Google Scholar] [CrossRef] [PubMed]
- Altikatoglu, M.; Celebi, M. Enhanced stability and decolorization of Coomassie Brilliant Blue R-250 by dextran aldehyde-modified horseradish peroxidase. Artif. Cells Blood Substit. Biotechnol. 2011, 39, 185–190. [Google Scholar] [CrossRef] [PubMed]
- Betancor, L.; Fuentes, M.; Dellamora-Ortiz, G.; Lopez-Gallego, F.; Hidalgo, A.; Alonso-Morales, N.; Mateo, C.; Guisan, J.M.; Fernandez-Lafuente, R. Dextran aldehyde coating of glucose oxidase immobilized on magnetic nanoparticles prevents its inactivation by gas bubbles. J. Mol. Catal. B Enzym. 2005, 32, 97–101. [Google Scholar] [CrossRef]
- Fujihara, M.; Takahashi, D.; Abe, H.; Sakai, H.; Horinouchi, H.; Kobayashi, K.; Ikeda, H.; Azuma, H. Primary and secondary immune responses to keyhole limpet hemocyanin in rats after infusion of hemoglobin vesicle, an artificial oxygen carrier. Artif. Organs 2014, 38, 234–238. [Google Scholar] [CrossRef] [PubMed]
- Chapman, P.B.; Morrissey, D.M.; Panageas, K.S.; Hamilton, W.B.; Zhan, C.; Destro, A.N.; Williams, L.; Israel, R.J.; Livingston, P.O. Induction of antibodies against GM2 ganglioside by immunizing melanoma patients using GM2-keyhole limpet hemocyanin + QS21 vaccine: A dose-response study. Clin. Canc. Res. 2000, 6, 874–879. [Google Scholar]
- Adluri, S.; Helling, F.; Ogata, S.; Zhang, S.; Itzkowitz, S.H.; Lloyd, K.O.; Livingston, P.O. Immunogenicity of synthetic TF-KLH (keyhole limpet hemocyanin) and sTn-KLH conjugates in colorectal carcinoma patients. Cancer Immunol. Immunother. 1995, 41, 185–192. [Google Scholar] [CrossRef] [PubMed]
- Helling, F.; Shang, A.; Calves, M.; Zhang, S.; Ren, S.; Yu, R.K.; Oettgen, H.F.; Livingston, P.O. GD3 vaccines for melanoma: Superior immunogenicity of keyhole limpet hemocyanin conjugate vaccines. Cancer Res. 1994, 54, 197–203. [Google Scholar] [PubMed]
- Fuentes, M.; Mateo, C.; Fernandez-Lafuente, R.; Guisan, J.M. Aldehyde-dextran-protein conjugates to immobilize amino-haptens: Avoiding cross-reactions in the immunodetection. Enzym. Microb. Technol. 2005, 36, 510–513. [Google Scholar] [CrossRef]
- Azam, M.G.; Shibata, T.; Kabashima, T.; Kai, M. Sensitive chemiluminescence detection of prion protein on a membrane by using a peroxidase-labeled dextran probe. Anal. Sci. 2011, 27, 715–720. [Google Scholar] [CrossRef]
- Shanazarova, I.M.; Vanchugova, L.V.; Valuev, L.I.; Platé, N.A. Interaction of ovomucoid from duck egg proteins with aldehydes-dextrans. Prikl. Biokhim. Mikrobiol. 1992, 28, 292–296. [Google Scholar]
- Cao, L.Q.; van Rantwijk, F.; Sheldon, R.A. Cross-linked enzyme aggregates: A simple and effective method for the immobilization of penicillin acylase. Org. Lett. 2000, 2, 1361–1364. [Google Scholar] [CrossRef]
- Sheldon, R.A. Cross-linked enzyme aggregates as industrial biocatalysts. Org. Process Res. Dev. 2011, 15, 213–223. [Google Scholar] [CrossRef]
- Schoevaart, R.; Wolbers, M.W.; Golubovic, M.; Ottens, M.; Kieboom, A.P.G.; van Rantwijk, F.; van der Wielen, L.A.M.; Sheldon, R.A. Preparation, optimization, and structures of cross-linked enzyme aggregates (CLEAs). Biotechnol. Bioeng. 2004, 87, 754–762. [Google Scholar] [CrossRef] [PubMed]
- Sheldon, R.A. Cross-linked enzyme aggregates (CLEA® s): Stable and recyclable biocatalysts. Biochem. Soc. Trans. 2007, 35, 1583–1587. [Google Scholar] [CrossRef] [PubMed]
- Sheldon, R.A. Enzyme immobilization: The quest for optimum performance. Adv. Synth. Catal. 2007, 349, 1289–1307. [Google Scholar] [CrossRef]
- Tukel, S.S.; Hurrem, F.; Yildirim, D.; Alptekin, O. Preparation of crosslinked enzyme aggregates (CLEA) of catalase and its characterization. J. Mol. Catal. B Enzym. 2013, 97, 252–257. [Google Scholar] [CrossRef]
- Araujo-Silva, R.; Mafra, A.C.O.; Rojas, M.J.; Kopp, W.; Giordano, R.D.; Fernandez-Lafuente, R.; Tardioli, P.W. Maltose production using starch from cassava bagasse catalyzed by cross-linked beta-amylase aggregates. Catalysts 2018, 8, 170. [Google Scholar] [CrossRef]
- Ramos, M.D.; Miranda, L.P.; Giordano, R.L.C.; Fernandez-Lafuente, R.; Kopp, W.; Tardioli, P.W. 1, 3-Regiospecific ethanolysis of soybean oil catalyzed by crosslinked porcine pancreas lipase aggregates. Biotechnol. Prog. 2018, 34, 910–920. [Google Scholar] [CrossRef] [PubMed]
- Amaral-Fonseca, M.; Kopp, W.; Giordano, R.D.C.; Fernandez-Lafuente, R.; Tardioli, P.W. Preparation of magnetic cross-linked amyloglucosidase aggregates: Solving some activity problems. Catalysts 2018, 8, 496. [Google Scholar] [CrossRef]
- Torres, M.D.G.; Foresti, M.L.; Ferreira, M.L. Cross-linked enzyme aggregates (CLEAs) of selected lipases: A procedure for the proper calculation of their recovered activity. AMB Express 2013, 3, 11. [Google Scholar]
- Rojas, M.J.; Amaral-Fonseca, M.; Zanin, G.M.; Fernandez-Lafuente, R.; Giordano, R.D.C.; Tardioli, P.W. Preparation of crosslinked enzyme aggregates of a thermostable cyclodextrin glucosyltransferase from Thermoanaerobacter sp. Critical effect of the crosslinking agent. Catalysts 2019, 9, 120. [Google Scholar] [CrossRef]
- Tirunagari, H.; Basetty, S.; Rode, H.B.; Fadnavis, N.W. Crosslinked enzyme aggregates (CLEA) of phytase with soymilk proteins. J. Biotechnol. 2018, 282, 67–69. [Google Scholar] [CrossRef] [PubMed]
- Cruz, J.; Barbosa, O.; Rodrigues, R.C.; Fernandez-Lafuente, R.; Torres, R.; Ortiz, C. Optimized preparation of CALB-CLEAs by response surface methodology: The necessity to employ a feeder to have an effective crosslinking. J. Mol. Catal. B Enzym. 2012, 80, 7–14. [Google Scholar] [CrossRef]
- Lopez-Gallego, F.; Betancor, L.; Hidalgo, A.; Alonso, N.; Fernandez-Lafuente, R.; Guisan, J.M. Co-aggregation of enzymes and polyethyleneimine: A simple method to prepare stable and immobilized derivatives of glutaryl acylase. Biomacromolecules 2005, 6, 1839–1842. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Gui, X.; Wang, G.; Yan, Y. Improving stability and activity of cross-linked enzyme aggregates based on polyethylenimine in hydrolysis of fish oil for enrichment of polyunsaturated fatty acids. Appl. Biochem. Biotechnol. 2012, 166, 925–932. [Google Scholar] [CrossRef] [PubMed]
- Galvis, M.; Barbosa, O.; Ruiz, M.; Cruz, J.; Ortiz, C.; Torres, R.; Fernandez-Lafuente, R. Chemical amination of lipase B from Candida antarctica is an efficient solution for the preparation of crosslinked enzyme aggregates. Process Biochem. 2012, 47, 2373–2378. [Google Scholar] [CrossRef]
- Vaidya, B.K.; Kuwar, S.S.; Golegaonkar, S.B.; Nene, S.N. Preparation of cross-linked enzyme aggregates of L-aminoacylase via co-aggregation with polyethyleneimine. J. Mol. Catal. B Enzym. 2012, 74, 184–191. [Google Scholar] [CrossRef]
- Pan, J.A.; Kong, X.D.; Li, C.X.; Ye, Q.; Xu, J.H.; Imanaka, T. Crosslinking of enzyme coaggregate with polyethyleneimine: A simple and promising method for preparing stable biocatalyst of Serratia marcescens lipase. J. Mol. Catal. B Enzym. 2011, 68, 256–261. [Google Scholar] [CrossRef]
- Velasco-Lozano, S.; Lopez-Gallego, F.; Vazquez-Duhalt, R.; Mateos-Diaz, J.C.; Guisan, J.M.; Favela-Torres, E. Carrier-free immobilization of lipase from Candida rugosa with polyethyleneimines by carboxyl-activated cross-linking. Biomacromolecules 2014, 15, 1896–1903. [Google Scholar] [CrossRef] [PubMed]
- Migneault, I.; Dartiguenave, C.; Bertrand, M.J.; Waldron, K.C. Glutaraldehyde: Behavior in aqueous solution, reaction with proteins, and application to enzyme crosslinking. BioTechniques 2004, 37, 790–802. [Google Scholar] [CrossRef]
- Barbosa, O.; Ortiz, C.; Berenguer-Murcia, A.; Torres, R.; Rodrigues, R.C.; Fernandez-Lafuente, R. Glutaraldehyde in bio-catalysts design: A useful crosslinker and a versatile tool in enzyme immobilization. RSC Adv. 2014, 4, 1583–1600. [Google Scholar] [CrossRef]
- Valdes, E.C.; Soto, L.W.; Arcaya, G.A. Influence of the pH of glutaraldehyde and the use of dextran aldehyde on the preparation of cross-linked enzyme aggregates (CLEAs) of lipase from Burkholderia cepacia. Electron. J. Biotechnol. 2011, 14, 7. [Google Scholar] [CrossRef]
- Mateo, C.; Palomo, J.M.; van Langen, L.M.; van Rantwijk, F.; Sheldon, R.A. A new, mild cross-linking methodology to prepare cross-linked enzyme aggregates. Biotechnol. Bioeng. 2004, 86, 273–276. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.H.; Park, S.; Kim, Y.H.; Won, K.; Lee, S.H. Immobilization of formate dehydrogenase from Candida boidinii through cross-linked enzyme aggregates. J. Mol. Catal. B Enzym. 2013, 97, 209–214. [Google Scholar] [CrossRef]
- Nadar, S.S.; Rathod, V.K. Magnetic macromolecular cross linked enzyme aggregates (CLEAs) of glucoamylase. Enzym. Microb. Technol. 2016, 83, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Ba, S.; Haroune, L.; Cruz-Morato, C.; Jacquet, C.; Touahar, I.E.; Bellenger, J.P.; Legault, C.Y.; Jones, J.P.; Cabana, H. Synthesis and characterization of combined cross-linked laccase and tyrosinase aggregates transforming acetaminophen as a model phenolic compound in wastewaters. Sci. Total Environ. 2014, 487, 748–755. [Google Scholar] [CrossRef] [PubMed]
- Ortiz-Soto, M.E.; Rudino-Pinera, E.; Rodriguez-Alegria, M.E.; Munguia, A.L. Evaluation of cross-linked aggregates from purified Bacillus subtilis levansucrase mutants for transfructosylation reactions. BMC Biotechnol. 2009, 9, 8. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.T.; Yang, K.L. Combined cross-linked enzyme aggregates of horseradish peroxidase and glucose oxidase for catalyzing cascade chemical reactions. Enzym. Microb. Technol. 2017, 100, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Zhen, Q.N.; Wang, M.F.; Qi, W.; Su, R.X.; He, Z.M. Preparation of beta-mannanase CLEAs using macromolecular cross-linkers. Catal. Sci. Technol. 2013, 3, 1937–1941. [Google Scholar] [CrossRef]
- Nadar, S.S.; Muley, A.B.; Ladole, M.R.; Joshi, P.U. Macromolecular cross-linked enzyme aggregates (M-CLEAs) of alpha-amylase. Int. J. Biol. Macromol. 2016, 84, 69–78. [Google Scholar] [CrossRef] [PubMed]
- Sahutoglu, A.S.; Akgul, C. Immobilisation of Aspergillus oryzae alpha-amylase and Aspergillus niger glucoamylase enzymes as cross-linked enzyme aggregates. Chem. Pap. 2015, 69, 433–439. [Google Scholar] [CrossRef]
- Qin, Y.Z.; Ye, M.; Li, N.; Zong, M.H. Preparation, characterization, and application of cross-linked enzyme aggregates of β-glycosidases from bovine liver. Mod. Food Sci. Technol. 2015, 31, 210–214. [Google Scholar]
- Do, M.Q.; Henry, E.; Kato, M.; Cheruzel, L. Cross-linked cytochrome P450 BM3 aggregates promoted by Ru (II)-diimine complexes bearing aldehyde groups. J. Inorg. Biochem. 2018, 186, 130–134. [Google Scholar] [CrossRef] [PubMed]
- Martwiset, S.; Koh, A.E.; Chen, W. Nonfouling characteristics of dextran-containing surfaces. Langmuir 2006, 22, 8192–8196. [Google Scholar] [CrossRef] [PubMed]
- Pessela, B.C.C.; Vian, A.; Mateo, U.; Fernandez-Lafuente, R.; Garcia, J.L.; Guisan, J.M.; Carrascosa, A.V. Overproduction of Thermus sp strain T2 beta-galactosidase in Escherichia coli and preparation by using tailor-made metal chelate supports. Appl. Environ. Microbiol. 2003, 69, 1967–1972. [Google Scholar] [CrossRef] [PubMed]
- Pessela, B.C.C.; Mateo, C.; Filho, M.; Carrascosa, A.; Fernandez-Lafuente, R.; Guisan, J.M. Selective adsorption of large proteins on highly activated IMAC supports in the presence of high imidazole concentrations: Purification, reversible immobilization and stabilization of thermophilic alpha-and beta-galactosidases. Enzym. Microb. Technol. 2007, 40, 242–248. [Google Scholar] [CrossRef]
- Murza, A.; Fernández-Lafuente, R.; Guisán, J.M. Essential role of the concentration of immobilized ligands in affinity chromatography: Purification of guanidinobenzoatase on an ionized ligand. J. Chromatogr. B Biomed. Sci. Appl. 2000, 740, 211–218. [Google Scholar] [CrossRef]
- Sulkowski, E. Purification of proteins by IMAC. Trends Biotechnol. 1985, 3, 1–7. [Google Scholar] [CrossRef]
- Franken, K.; Hiemstra, H.S.; van Meijgaarden, K.E.; Subronto, Y.; den Hartigh, J.; Ottenhoff, T.H.M.; Drijfhout, J.W. Purification of His-tagged proteins by immobilized chelate affinity chromatography: The benefits from the use of organic solvent. Protein Expr. Purif. 2000, 18, 95–99. [Google Scholar] [CrossRef]
- Hemdan, E.S.; Zhao, Y.J.; Sulkowski, E.; Porath, J. Surface-topography of histidine-residues—A facile probe by immobilized metal-ion affinity-chromatography. Proc. Natl. Acad. Sci. USA 1989, 86, 1811–1815. [Google Scholar] [CrossRef]
- Porath, J. Immobilized metal ion affinity chromatography. Protein Expr. Purif. 1992, 3, 263–281. [Google Scholar] [CrossRef]
- Porath, J.; Carlsson, J.; Olsson, I.; Belfrage, G. Metal chelate affinity chromatography, a new approach to protein fractionation. Nature 1975, 258, 598–599. [Google Scholar] [CrossRef]
- Wong, J.W.; Albright, R.L.; Wang, N.H.L. Immobilized metal-ion affinity-chromatography (imac) chemistry and bioseparation applications. Sep. Purif. Methods 1991, 20, 49–106. [Google Scholar] [CrossRef]
- Armisén, P.; Mateo, C.; Cortés, E.; Barredo, J.L.; Salto, F.; Diez, B.; Rodés, L.; García, J.L.; Fernández-Lafuente, R.; Guisán, J.M. Selective adsorption of poly-His tagged glutaryl acylase on tailor-made metal chelate supports. J. Chromatogr. A 1999, 848, 61–70. [Google Scholar] [CrossRef]
- Mateo, C.; Fernandez-Lorente, G.; Pessela, B.C.C.; Vian, A.; Carrascosa, A.V.; Garcia, J.L.; Fernandez-Lafuente, R.; Guisan, J.M. Affinity chromatography of polyhistidine tagged enzymes—New dextran-coated immobilized metal ion affinity chromatography matrices for prevention of undesired multipoint adsorptions. J. Chromatogr. A 2001, 915, 97–106. [Google Scholar] [CrossRef]
- Fuentes, M.; Mateo, C.; Pessela, B.C.C.; Batalla, P.; Fernandez-Lafuente, R.; Guisan, J.M. Solid phase proteomics: Dramatic reinforcement of very weak protein-protein interactions. J. Chromatogr. B 2007, 849, 243–250. [Google Scholar] [CrossRef] [PubMed]
- Bolivar, J.M.; Batalla, P.; Mateo, C.; Carrascosa, A.V.; Pessela, B.C.; Guisan, J.M. Selective adsorption of small proteins on large-pore anion exchangers coated with medium size proteins. Colloid Surf. B Biointerfaces 2010, 78, 140–145. [Google Scholar] [CrossRef] [PubMed]
- Mateo, C.; Palomo, J.M.; Fuentes, M.; Betancor, L.; Grazu, V.; Lopez-Gallego, F.; Pessela, B.C.C.; Hidalgo, A.; Fernandez-Lorente, G.; Fernandez-Lafuente, R.; et al. Glyoxyl agarose: A fully inert and hydrophilic support for immobilization and high stabilization of proteins. Enzym. Microb. Technol. 2006, 39, 274–280. [Google Scholar] [CrossRef]
- Bhalla, N.; Jolly, P.; Formisano, N.; Estrela, P. Introduction to biosensors. In Biosensor Technologies for Detection of Biomolecules; Estrela, P., Ed.; Portland Press Ltd.: London, UK, 2016; Volume 60, pp. 1–8. [Google Scholar]
- Batalla, P.; Fuentes, M.; Grazu, V.; Mateo, C.; Fernandez-Lafuente, R.; Guisan, J.M. Oriented covalent immobilization of antibodies on physically inert and hydrophilic support surfaces through their glycosidic chains. Biomacromolecules 2008, 9, 719–723. [Google Scholar] [CrossRef]
- Kang, J.H.; Choi, H.J.; Hwang, S.Y.; Han, S.H.; Jeon, J.Y.; Lee, E.K. Improving immunobinding using oriented immobilization of an oxidized antibody. J. Chromatogr. A 2007, 1161, 9–14. [Google Scholar] [CrossRef]
- Nisnevitch, M.; Firer, M.A. The solid phase in affinity chromatography: Strategies for antibody attachment. J. Biochem. Biophys. Methods 2001, 49, 467–480. [Google Scholar] [CrossRef]
- Shmanai, V.V.; Nikolayeva, T.A.; Vinokurova, L.G.; Litoshka, A.A. Oriented antibody immobilization to polystyrene macrocarriers for immunoassay modified with hydrazide derivatives of poly (meth) acrylic acid. BMC Biotechnol. 2001, 1, 4. [Google Scholar] [CrossRef]
- Weiping, Q.; Bin, X.; Lei, W.; Chunxiao, W.; Danfeng, Y.; Fang, Y.; Chunwei, Y.; Yu, W. Controlled site-directed assembly of antibodies by their oligosaccharide moieties onto APTES derivatized surfaces. J. Colloid Interface Sci. 1999, 214, 16–19. [Google Scholar] [CrossRef]
- Jesionowski, T.; Zdarta, J.; Krajewska, B. Enzyme immobilization by adsorption: A review. Adsorpt. J. Int. Adsorpt. Soc. 2014, 20, 801–821. [Google Scholar] [CrossRef]
- Fuentes, M.; Mateo, C.; Guisan, J.M.; Fernandez-Lafuente, R. Preparation of inert magnetic nano-particles for the directed immobilization of antibodies. Biosens. Bioelectron. 2005, 20, 1380–1387. [Google Scholar] [CrossRef] [PubMed]
- Fuentes, M.; Mateo, C.; Rodriguez, A.; Casqueiro, M.; Tercero, J.C.; Riese, H.H.; Fernandez-Lafuente, R.; Guisan, J.M. Detecting minimal traces of DNA using DNA covalently attached to superparamagnetic nanoparticles and direct PCR-ELISA. Biosens. Bioelectron. 2006, 21, 1574–1580. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.; Yang, X.; Liu, T.; Chen, Z.; Chen, L.L.; Li, H.D.; Deng, L. Preparing a highly specific inert immunomolecular-magnetic beads for rapid detection and separation of S. aureus and group G Streptococcus. Appl. Microbiol. Biotechnol. 2007, 75, 1209–1216. [Google Scholar] [CrossRef]
- Farrell, M.; Beaudoin, S. Surface forces and protein adsorption on dextran- and polyethylene glycol-modified polydimethylsiloxane. Colloid Surf. B Biointerfaces 2010, 81, 468–475. [Google Scholar] [CrossRef]
- Bi, Y.C.; Yu, M.; Zhou, H.Y.; Zhou, H.; Wei, P. Biosynthesis of oleyl oleate in solvent-free system by Candida rugosa Lipase (CRL) immobilized in macroporous resin with cross-linking of aldehyde-dextran. J. Mol. Catal. B Enzym. 2016, 133, 1–5. [Google Scholar] [CrossRef]
- Avseenko, N.V.; Morozova, T.Y.; Ataullakhanov, F.I.; Morozov, V.N. Immobilization of proteins in immunochemical microarrays fabricated by electrospray deposition. Anal. Chem. 2001, 73, 6047–6052. [Google Scholar] [CrossRef]
- Nouaimi, M.; Möschel, K.; Bisswanger, H. Immobilization of trypsin on polyester fleece via different spacers. Enzym. Microb. Technol. 2001, 29, 567–574. [Google Scholar] [CrossRef]
- Matthijs, G.; Schacht, E. Comparative study of methodologies for obtaining β-glucosidase immobilized on dextran-modified silica. Enzym. Microb. Technol. 1996, 19, 601–605. [Google Scholar] [CrossRef]
- Moser, I.; Dworsky, P.; Pittner, F. Degradation of bacterial-cell walls by immobilized lysozyme. Appl. Biochem. Biotechnol. 1988, 19, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Germain, P.; Crichton, R.R. Characterization of a chemically modified β-amylase immobilized on porous silica. J. Chem. Technol. Biotechnol. 1988, 41, 297–315. [Google Scholar] [CrossRef]
- Palubinskas, V.I.; Yankevich, N.B.; Yanulaitene, K.K.; Vesa, V.S.; Bendikene, V.G.; Maksimenko, A.V.; Torchilin, V.P.; Ilyina, E.V.; Smirnov, V.N.; Krestyanova, I.N.; et al. Trypsin-like-enzyme from Streptomyces-771—Purification and properties of native and immobilized enzyme. Appl. Biochem. Biotechnol. 1984, 9, 231–241. [Google Scholar] [CrossRef] [PubMed]
- Penzol, G.; Armisen, P.; Fernandez-Lafuente, R.; Rodes, L.; Guisan, J.M. Use of dextrans as long and hydrophilic spacer arms to improve the performance of immobilized proteins acting on macromolecules. Biotechnol. Bioeng. 1998, 60, 518–523. [Google Scholar] [CrossRef]
- Azam, M.G.; Shibata, T.; Kabashima, T.; Kai, M. Alkaline phosphatase-labeled macromolecular probe for sensitive chemiluminescence detection of proteins on a solid-phase membrane. Anal. Bioanal. Chem. 2011, 401, 1211–1217. [Google Scholar] [CrossRef] [PubMed]
- Alvaro, G.; Fernandez-Lafuente, R.; Blanco, R.M.; Guisán, J.M. Immobilization-stabilization of Penicillin G acylase from Escherichia coli. Appl. Biochem. Biotechnol. 1990, 26, 181–195. [Google Scholar] [CrossRef]
- Abian, O.; Grazú, V.; Hermoso, J.; González, R.; García, J.L.; Fernández-Lafuente, R.; Guisán, J.M. Stabilization of penicillin G acylase from Escherichia coli: Site-directed mutagenesis of the protein surface to increase multipoint covalent attachment. Appl. Environ. Microbiol. 2004, 70, 1249–1251. [Google Scholar] [CrossRef]
- Fernandez-Lafuente, R.; Rosell, C.M.; Caanan-Haden, L.; Rodes, L.; Guisan, J.M. Facile synthesis of artificial enzyme nano-environments via solid-phase chemistry of immobilized derivatives: Dramatic stabilization of penicillin acylase versus organic solvents. Enzym. Microb. Technol. 1999, 24, 96–103. [Google Scholar] [CrossRef]
- Virgen-Ortíz, J.J.; Dos Santos, J.C.S.; Berenguer-Murcia, Á.; Barbosa, O.; Rodrigues, R.C.; Fernandez-Lafuente, R. Polyethylenimine: A very useful ionic polymer in the design of immobilized enzyme biocatalysts. J. Mater. Chem. B 2017, 5, 7461–7490. [Google Scholar] [CrossRef]
- Abian, O.; Wilson, L.; Mateo, C.; Fernandez-Lorente, G.; Palomo, J.M.; Fernandez-Lafuente, R.; Guisan, J.M.; Re, D.; Tam, A.; Daminatti, M. Preparation of artificial hyper-hydrophilic micro-environments (polymeric salts) surrounding enzyme molecules—New enzyme derivatives to be used in any reaction medium. J. Mol. Catal. B Enzym. 2002, 19, 295–303. [Google Scholar] [CrossRef]
- Abian, O.; Mateo, C.; Fernandez-Lorente, G.; Palomo, J.M.; Fernandez-Lafuente, R.; Guisan, J.M. Stabilization of immobilized enzymes against water-soluble organic cosolvents and generation of hyper-hydrophilic micro-environments surrounding enzyme molecules. Biocatal. Biotransform. 2001, 19, 489–503. [Google Scholar] [CrossRef]
- Abian, O.; Mateo, C.; Fernandez-Lorente, G.; Guisan, J.M.; Fernandez-Lafuente, R. Improving the industrial production of 6-APA: Enzymatic hydrolysis of penicillin G in the presence of organic solvents. Biotechnol. Prog. 2003, 19, 1639–1642. [Google Scholar] [CrossRef] [PubMed]
- Abian, O.; Mateo, C.; Fernandez-Lorente, G.; Guisan, J.M.; Fernandez-Lafuente, R. Thermodynamically controlled synthesis of amide bonds catalyzed by highly organic solvent-resistant penicillin acylase derivatives. Biotechnol. Prog. 2004, 20, 117–121. [Google Scholar] [CrossRef] [PubMed]
- Abian, O.; Mateo, C.; Palomo, J.M.; Fernandez-Lorente, G.; Guisian, J.M.; Fernandez-Lafuente, R. Enantioselective synthesis of phenylacetamides in the presence of high organic cosolvent concentrations catalyzed by stabilized penicillin G acylase. Effect of the acyl donor. Biotechnol. Prog. 2004, 20, 984–988. [Google Scholar] [CrossRef] [PubMed]
- Irazoqui, G.; Giacomini, C.; Batista-Viera, F.; Brena, B.M. Hydrophilization of immobilized model enzymes suggests a widely applicable method for enhancing protein stability in polar organic co-solvents. J. Mol. Catal. B Enzym. 2007, 46, 43–51. [Google Scholar] [CrossRef]
- Fernández-Lafuente, R.; Rodríguez, V.; Mateo, C.; Fernández-Lorente, G.; Arminsen, P.; Sabuquillo, P.; Guisán, J.M. Stabilization of enzymes (D-amino acid oxidase) against hydrogen peroxide via immobilization and post-immobilization techniques. J. Mol. Catal. B Enzym. 1999, 7, 173–179. [Google Scholar] [CrossRef]
- Lencki, R.W.; Arul, J.; Neufeld, R.J. Effect of subunit dissociation, denaturation, aggregation, coagulation, and decomposition on enzyme inactivation kinetics: II. Biphasic and grace period behavior. Biotechnol. Bioeng. 1992, 40, 1427–1434. [Google Scholar] [CrossRef] [PubMed]
- Bechtold, M.; Panke, S. Model-based characterization of operational stability of multimeric enzymes with complex deactivation behavior: An in-silico investigation. Chem. Eng. Sci. 2012, 80, 435–450. [Google Scholar] [CrossRef]
- Vrábel, P.; Polakovič, M.; Báleš, V. Modelling of inactivation phenomena in oligomeric enzymes. Comput. Chem. Eng. 1994, 18, S681–S685. [Google Scholar] [CrossRef]
- Ahern, T.J.; Casal, J.I.; Petsko, G.A.; Klibanov, A.M. Control of oligomeric enzyme thermostability by protein engineering. Proc. Natl. Acad. Sci. USA 1987, 84, 675–679. [Google Scholar] [CrossRef] [PubMed]
- Peimbert, M.; Dominguez-Ramirez, L.; Fernandez-Velasco, D.A. Hydrophobic repacking of the dimer interface of triosephosphate isomerase by in silico design and directed evolution. Biochemistry 2008, 47, 5556–5564. [Google Scholar] [CrossRef] [PubMed]
- Kabashima, T.; Li, Y.; Kanada, N.; Ito, K.; Yoshimoto, T. Enhancement of the thermal stability of pyroglutamyl peptidase I by introduction of an intersubunit disulfide bond. Biochim. Biophys. Acta Protein Struct. Molec. Enzym. 2001, 1547, 214–220. [Google Scholar] [CrossRef]
- Velanker, S.S.; Gokhale, R.S.; Ray, S.S.; Gopal, B.; Parthasarathy, S.; Santi, D.V.; Balaram, P.; Murthy, M.R.N. Disulfide engineering at the dimer interface of Lactobacillus casei thymidylate synthase: Crystal structure of the T155C/E188C/C244T mutant. Protein Sci. 1999, 8, 930–933. [Google Scholar] [CrossRef] [PubMed]
- Wilson, L.; Betancor, L.; Fernandez-Lorente, G.; Fuentes, M.; Hidalgo, A.; Guisan, J.M.; Pessela, B.C.C.; Fernandez-Lafuente, R. Cross-linked aggregates of multimeric enzymes: A simple and efficient methodology to stabilize their quaternary structure. Biomacromolecules 2004, 5, 814–817. [Google Scholar] [CrossRef] [PubMed]
- Bolivar, J.M.; Wilson, L.; Ferrarotti, S.A.; Guisan, J.M.; Fernandez-Lafuente, R.; Mateo, C. Improvement of the stability of alcohol dehydrogenase by covalent immobilization on glyoxyl-agarose. J. Biotechnol. 2006, 125, 85–94. [Google Scholar] [CrossRef] [PubMed]
- Gouin, S.G.; Fernandez, J.M.G.; Vanquelef, E.; Dupradeau, F.Y.; Salomonsson, E.; Leffler, H.; Ortega-Munoz, M.; Nilsson, U.J.; Kovensky, J. Multimeric lactoside “click clusters” as tools to investigate the effect of linker length in specific interactions with peanut lectin, galectin-1, and-3. ChemBiochem 2010, 11, 1430–1442. [Google Scholar] [CrossRef] [PubMed]
- Aissaoui, N.; Landoulsi, J.; Bergaoui, L.; Boujday, S.; Lambert, J.F. Catalytic activity and thermostability of enzymes immobilized on silanized surface: Influence of the crosslinking agent. Enzym. Microb. Technol. 2013, 52, 336–343. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Lafuente, R.; Rodríguez, V.; Mateo, C.; Penzol, G.; Hernández-Justiz, O.; Irazoqui, G.; Villarino, A.; Ovsejevi, K.; Batista, F.; Guisán, J.M. Stabilization of multimeric enzymes via immobilization and post-immobilization techniques. J. Mol. Catal. B Enzym. 1999, 7, 181–189. [Google Scholar] [CrossRef]
- Balcao, V.M.; Mateo, C.; Fernandez-Lafuente, R.; Malcata, F.X.; Guisan, J.M. Structural and functional stabilization of L-asparaginase via multisubunit immobilization onto highly activated supports. Biotechnol. Prog. 2001, 17, 537–542. [Google Scholar] [CrossRef]
- Fernandez-Lafuente, R.; Hernandez-Justiz, O.; Mateo, C.; Terreni, M.; Fernandez-Lorente, G.; Moreno, M.A.; Alonso, J.; Garcia-Lopez, J.L.; Guisan, J.M. Biotransformations catalyzed by multimeric enzymes: Stabilization of tetrameric ampicillin acylase permits the optimization of ampicillin synthesis under dissociation conditions. Biomacromolecules 2001, 2, 95–104. [Google Scholar] [CrossRef]
- Hidalgo, A.; Betancor, L.; Lopez-Gallego, F.; Moreno, R.; Berenguer, J.; Fernández-Lafuente, R.; Guisán, J.M. Design of an immobilized preparation of catalase from Thermus thermophilus to be used in a wide range of conditions.: Structural stabilization of a multimeric enzyme. Enzym. Microb. Technol. 2003, 33, 278–285. [Google Scholar] [CrossRef]
- Betancor, L.; Hidalgo, A.; Fernandez-Lorente, G.; Mateo, C.; Fernandez-Lafuente, R.; Guisan, J.M. Preparation of a stable biocatalyst of bovine liver catalase using immobilization and postimmobilization techniques. Biotechnol. Prog. 2003, 19, 763–767. [Google Scholar] [CrossRef]
- Kaddour, S.; Lopez-Gallego, F.; Sadoun, T.; Fernandez-Lafuente, R.; Guisan, J.M. Preparation of an immobilized-stabilized catalase derivative from Aspergillus niger having its multimeric structure stabilized: The effect of Zn2+ on enzyme stability. J. Mol. Catal. B Enzym. 2008, 55, 142–145. [Google Scholar] [CrossRef]
- Terrasan, C.R.F.; Trobo-Maseda, L.; Moreno-Perez, S.; Carmona, E.C.; Pessela, B.C.; Guisan, J.M. Co-immobilization and stabilization of xylanase, beta-xylosidase and alpha-L-arabinofuranosidase from Penicillium janczewskii for arabinoxylan hydrolysis. Process Biochem. 2016, 51, 614–623. [Google Scholar] [CrossRef]
- Rocha-Martin, J.; Acosta, A.; Berenguer, J.; Guisan, J.M.; Lopez-Gallego, F. Selective oxidation of glycerol to 1,3-dihydroxyacetone by covalently immobilized glycerol dehydrogenases with higher stability and lower product inhibition. Bioresour. Technol. 2014, 170, 445–453. [Google Scholar] [CrossRef]
- Serra, I.; Conti, S.; Piskur, J.; Clausen, A.R.; Munch-Petersen, B.; Terreni, M.; Ubiali, D. Immobilized Drosophila melanogaster deoxyribonucleoside kinase (DmdNK) as a high performing biocatalyst for the synthesis of purine arabinonucleotides. Adv. Synth. Catal. 2014, 356, 563–570. [Google Scholar] [CrossRef]
- Lopez-Gallego, F.; Betancor, L.; Hidalgo, A.; Dellamora-Ortiz, G.; Mateo, C.; Fernandez-Lafuente, R.; Guisan, J.M. Stabilization of different alcohol oxidases via immobilization and post immobilization techniques. Enzym. Microb. Technol. 2007, 40, 278–284. [Google Scholar] [CrossRef]
- Li, D.; Zhao, Y.; Fei, T.; Wang, Y.; Lee, B.H.; Shim, J.H.; Xu, B.; Li, Z.; Li, X. Effects of Streptococcus thermophilus GtfB enzyme on dough rheology, bread quality and starch digestibility. Food Hydrocoll. 2019, 96, 134–139. [Google Scholar] [CrossRef]
- Bala, A.; Singh, B. Cellulolytic and xylanolytic enzymes of thermophiles for the production of renewable biofuels. Renew. Energy 2019, 136, 1231–1244. [Google Scholar] [CrossRef]
- Atalah, J.; Caceres-Moreno, P.; Espina, G.; Blamey, J.M. Thermophiles and the applications of their enzymes as new biocatalysts. Bioresour. Technol. 2019, 280, 478–488. [Google Scholar] [CrossRef]
- Han, H.W.; Ling, Z.M.; Khan, A.; Virk, A.K.; Kulshrestha, S.; Li, X.K. Improvements of thermophilic enzymes: From genetic modifications to applications. Bioresour. Technol. 2019, 279, 350–361. [Google Scholar] [CrossRef] [PubMed]
- Karnaouri, A.; Antonopoulou, I.; Zerva, A.; Dimarogona, M.; Topakas, E.; Rova, U.; Christakopoulos, P. Thermophilic enzyme systems for efficient conversion of lignocellulose to valuable products: Structural insights and future perspectives for esterases and oxidative catalysts. Bioresour. Technol. 2019, 279, 362–372. [Google Scholar] [CrossRef] [PubMed]
- Bhalla, A.; Bansal, N.; Kumar, S.; Bischoff, K.M.; Sani, R.K. Improved lignocellulose conversion to biofuels with thermophilic bacteria and thermostable enzymes. Bioresour. Technol. 2013, 128, 751–759. [Google Scholar] [CrossRef] [PubMed]
- Turner, P.; Mamo, G.; Karlsson, E.N. Potential and utilization of thermophiles and thermostable enzymes in biorefining. Microb. Cells Fact. 2007, 6, 23. [Google Scholar] [CrossRef] [PubMed]
- Wolf-Watz, M.; Thai, V.; Henzler-Wildman, K.; Hadjipavlou, G.; Eisenmesser, E.Z.; Kern, D. Linkage between dynamics and catalysis in a thermophilic-mesophilic enzyme pair. Nat. Struct. Mol. Biol. 2004, 11, 945–949. [Google Scholar] [CrossRef] [PubMed]
- Pessela, B.C.; Mateo, C.; Filho, M.; Carrascosa, A.V.; Fernandez-Lafuente, R.; Guisan, J.M. Stabilization of the quaternary structure of a hexameric alpha-galactosidase from Thermus sp T2 by immobilization and post-immobilization techniques. Process Biochem. 2008, 43, 193–198. [Google Scholar] [CrossRef]
- Pessela, B.C.C.; Mateo, C.; Fuentes, M.; Vian, A.; Garcia, J.L.; Carrascosa, A.V.; Guisan, J.M.; Fernandez-Lafuente, R. Stabilization of a multimeric beta-galactosidase from Thermus sp strain T2 by immobilization on novel heterofunctional epoxy supports plus aldehyde-dextran cross-linking. Biotechnol. Prog. 2004, 20, 388–392. [Google Scholar] [CrossRef]
- Terrasan, C.R.F.; Aragon, C.C.; Masui, D.C.; Pessela, B.C.; Fernandez-Lorente, G.; Carmona, E.C.; Guisan, J.M. beta-xylosidase from Selenomonas ruminantium: Immobilization, stabilization, and application for xylooligosaccharide hydrolysis. Biocatal. Biotransform. 2016, 34, 161–171. [Google Scholar] [CrossRef]
- Trobo-Maseda, L.; Orrego, A.H.; Moreno-Perez, S.; Fernandez-Lorente, G.; Guisan, J.M.; Rocha-Martin, J. Stabilization of multimeric sucrose synthase from Acidithiobacillus caldus via immobilization and post-immobilization techniques for synthesis of UDP-glucose. Appl. Microbiol. Biotechnol. 2018, 102, 773–787. [Google Scholar] [CrossRef]
- Rajdeo, K.; Harini, T.; Lavanya, K.; Fadnavis, N.W. Immobilization of pectinase on reusable polymer support for clarification of apple juice. Food Bioprod. Process. 2016, 99, 12–19. [Google Scholar] [CrossRef]
- Fresco-Taboada, A.; Serra, I.; Fernandez-Lucas, J.; Acebal, C.; Arroyo, M.; Terreni, M.; de la Mata, I. Nucleoside 2′-deoxyribosyltransferase from psychrophilic bacterium Bacillus psychrosaccharolyticus—Preparation of an immobilized biocatalyst for the enzymatic synthesis of therapeutic nucleosides. Molecules 2014, 19, 11231–11249. [Google Scholar] [CrossRef] [PubMed]
- Rocchietti, S.; Ubiali, D.; Terreni, M.; Albertini, A.M.; Fernandez-Lafuente, R.; Guisan, J.M.; Pregnolato, M. Immobilization and stabilization of recombinant multimeric uridine and purine nucleoside phosphorylases from Bacillus subtilis. Biomacromolecules 2004, 5, 2195–2200. [Google Scholar] [CrossRef] [PubMed]
- Ubiali, D.; Rocchietti, S.; Scaramozzino, F.; Terreni, M.; Albertini, A.M.; Fernandez-Lafuente, R.; Guisan, J.M.; Pregnolato, M. Synthesis of 2′-deoxynucleosides by transglycosylation with new immobilized and stabilized uridine phosphorylase and purine nucleoside phosphorylase. Adv. Synth. Catal. 2004, 346, 1361–1366. [Google Scholar] [CrossRef]
- Ricca, E.; Brucher, B.; Schrittwieser, J.H. Multi-enzymatic cascade reactions: Overview and perspectives. Adv. Synth. Catal. 2011, 353, 2239–2262. [Google Scholar] [CrossRef]
- Zhang, G.; Quin, M.B.; Schmidt-Dannert, C. Self-Assembling Protein Scaffold System for Easy in Vitro Coimmobilization of Biocatalytic Cascade Enzymes. ACS Catal. 2018, 8, 5611–5620. [Google Scholar] [CrossRef]
- Walsh, C.T.; Moore, B.S. Enzymatic cascade reactions in biosynthesis. Angew. Chem. Int. Ed. 2019, 58, 6846–6879. [Google Scholar] [CrossRef] [PubMed]
- Kuzmak, A.; Carmali, S.; von Lieres, E.; Russell, A.J.; Kondrat, S. Can enzyme proximity accelerate cascade reactions? Sci. Rep. 2019, 9, 455. [Google Scholar] [CrossRef] [PubMed]
- Arana-Peña, S.; Mendez-Sanchez, C.; Rios, N.S.; Ortiz, C.; Gonçalves, L.R.B.; Fernandez-Lafuente, R. New applications of glyoxyl-octyl agarose in lipases co-immobilization: Strategies to reuse the most stable lipase. Int. J. Biol. Macromol. 2019, 131, 989–997. [Google Scholar] [CrossRef]
- Rios, N.S.; Arana-Peña, S.; Mendez-Sanchez, C.; Ortiz, C.; Gonçalves, L.R.B.; Fernandez-Lafuente, R. Reuse of lipase from Pseudomonas fluorescens via its step-by-step coimmobilization on glyoxyl-octyl agarose beads with least stable lipases. Catalysts 2019, 9, 487. [Google Scholar] [CrossRef]
- Zaak, H.; Kornecki, J.F.; Siar, E.H.; Fernandez-Lopez, L.; Corberán, V.C.; Sassi, M.; Fernandez-Lafuente, R. Coimmobilization of enzymes in bilayers using PEI as a glue to reuse the most stable enzyme: Preventing PEI release during inactivated enzyme desorption. Process Biochem. 2017, 61, 95–101. [Google Scholar] [CrossRef]
- Peirce, S.; Virgen-Ortíz, J.J.; Tacias-Pascacio, V.G.; Rueda, N.; Bartolome-Cabrero, R.; Fernandez-Lopez, L.; Russo, M.E.; Marzocchella, A.; Fernandez-Lafuente, R. Development of simple protocols to solve the problems of enzyme coimmobilization. Application to coimmobilize a lipase and a β-galactosidase. RSC Adv. 2016, 6, 61707–61715. [Google Scholar] [CrossRef]
- Pereira-Rodriguez, A.; Fernandez-Leiro, R.; Gonzalez-Siso, M.I.; Cerdan, M.E.; Becerra, M.; Sanz-Aparicio, J. Structural basis of specificity in tetrameric Kluyveromyces lactis beta-galactosidase. J. Struct. Biol. 2012, 177, 392–401. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.S.; Ji, E.S.; Oh, D.K. Expression and characterization of Kluyveromyces lactis beta-galactosidase in Escherichia coli. Biotechnol. Lett. 2003, 25, 1769–1774. [Google Scholar] [CrossRef] [PubMed]
- Henriques, R.O.; Bork, J.A.; Fernandez-Lorente, G.; Guisan, J.M.; Furigo, A.; de Oliveira, D.; Pessela, B.C. Co-immobilization of lipases and beta-D-galactosidase onto magnetic nanoparticle supports: Biochemical characterization. Mol. Catal. 2018, 453, 12–21. [Google Scholar] [CrossRef]
- Santos, J.C.S.D.; Barbosa, O.; Ortiz, C.; Berenguer-Murcia, A.; Rodrigues, R.C.; Fernandez-Lafuente, R. Importance of the support properties for immobilization or purification of enzymes. ChemCatChem 2015, 7, 2413–2432. [Google Scholar] [CrossRef]
- Virgen-Ortíz, J.J.; Peirce, S.; Tacias-Pascacio, V.G.; Cortes-Corberan, V.; Marzocchella, A.; Russo, M.E.; Fernandez-Lafuente, R. Reuse of anion exchangers as supports for enzyme immobilization: Reinforcement of the enzyme-support multiinteraction after enzyme inactivation. Process Biochem. 2016, 51, 1391–1396. [Google Scholar] [CrossRef]
- Virgen-Ortíz, J.J.; Pedrero, S.G.; Fernandez-Lopez, L.; Lopez-Carrobles, N.; Gorines, B.C.; Otero, C.; Fernandez-Lafuente, R. Desorption of lipases immobilized on octyl-agarose beads and coated with ionic polymers after thermal inactivation. Stronger adsorption of polymers/unfolded protein composites. Molecules 2017, 22, 91. [Google Scholar] [CrossRef]
- Virgen-Ortíz, J.J.; Tacias-Pascacio, V.G.; Hirata, D.B.; Torrestiana-Sanchez, B.; Rosales-Quintero, A.; Fernandez-Lafuente, R. Relevance of substrates and products on the desorption of lipases physically adsorbed on hydrophobic supports. Enzym. Microb. Technol. 2017, 96, 30–35. [Google Scholar] [CrossRef]
- Hirata, D.B.; Albuquerque, T.L.; Rueda, N.; Virgen-Ortiz, J.J.; Tacias-Pascacio, V.G.; Fernandez-Lafuente, R. Evaluation of different immobilized lipases in transesterification reactions using tributyrin: Advantages of the heterofunctional octyl agarose beads. J. Mol. Catal. B Enzym. 2016, 133, 117–123. [Google Scholar] [CrossRef]
- Hirata, D.B.; Albuquerque, T.L.; Rueda, N.; Sanchez-Montero, J.M.; Garcia-Verdugo, E.; Porcar, R.; Fernandez-Lafuente, R. Advantages of heterofunctional octyl supports: Production of 1,2-dibutyrin by specific and selective hydrolysis of tributyrin catalyzed by immobilized lipases. ChemistrySelect 2016, 1, 3259–3270. [Google Scholar] [CrossRef]
- Rios, N.S.; Mendez-Sanchez, C.; Arana-Peña, S.; Rueda, N.; Ortiz, C.; Gonçalves, L.R.B.; Fernandez-Lafuente, R. Immobilization of lipase from Pseudomonas fluorescens on glyoxyl-octyl-agarose beads: Improved stability and reusability. Biochim. Biophys. Acta Proteins Proteom. 2019, in press. [Google Scholar] [CrossRef] [PubMed]
- Rueda, N.; dos Santos, J.C.S.; Torres, R.; Ortiz, C.; Barbosa, O.; Fernandez-Lafuente, R. Improved performance of lipases immobilized on heterofunctional octyl-glyoxyl agarose beads. RSC Adv. 2015, 5, 11212–11222. [Google Scholar] [CrossRef]
- Fernandez-Lopez, L.; Virgen-Ortiz, J.J.; Pedrero, S.G.; Lopez-Carrobles, N.; Gorines, B.C.; Otero, C.; Fernandez-Lafuente, R. Optimization of the coating of octyl-CALB with ionic polymers to improve stability and decrease enzyme leakage. Biocatal. Biotransform. 2018, 36, 47–56. [Google Scholar] [CrossRef]
- Zaak, H.; Fernandez-Lopez, L.; Otero, C.; Sassi, M.; Fernandez-Lafuente, R. Improved stability of immobilized lipases via modification with polyethylenimine and glutaraldehyde. Enzym. Microb. Technol. 2017, 106, 67–74. [Google Scholar] [CrossRef]
- Fernandez-Lopez, L.; Pedrero, S.G.; Lopez-Carrobles, N.; Virgen-Ortiz, J.J.; Gorines, B.C.; Otero, C.; Fernandez-Lafuente, R. Physical crosslinking of lipase from Rhizomucor miehei immobilized on octyl agarose via coating with ionic polymers avoiding enzyme release from the support. Process Biochem. 2017, 54, 81–88. [Google Scholar] [CrossRef]
- Fernandez-Lorente, G.; Filice, M.; Lopez-Vela, D.; Pizarro, C.; Wilson, L.; Betancor, L.; Avila, Y.; Guisan, J.M. Cross-linking of lipases adsorbed on hydrophobic supports: Highly selective hydrolysis of fish oil catalyzed by RML. J. Am. Oil Chem. Soc. 2011, 88, 801–807. [Google Scholar] [CrossRef]
- Pizarro, C.; Branes, M.C.; Markovits, A.; Fernandez-Lorente, G.; Guisan, J.M.; Chamy, R.; Wilson, L. Influence of different immobilization techniques for Candida cylindracea lipase on its stability and fish oil hydrolysis. J. Mol. Catal. B Enzym. 2012, 78, 111–118. [Google Scholar] [CrossRef]
- Fuentes, M.; Segura, R.L.; Abian, O.; Betancor, L.; Hidalgo, A.; Mateo, C.; Fernandez-Lafuente, R.; Guisan, J.M. Determination of protein-protein interactions through aldehyde-dextran intermolecular cross-linking. Proteomics 2004, 4, 2602–2607. [Google Scholar] [CrossRef]
- Pessela, B.C.C.; Munilla, R.; Betancor, L.; Fuentes, M.; Carrascosa, A.V.; Vian, A.; Fernandez-Lafuente, R.; Guisan, J.M. Ion exchange using poorly activated supports, an easy way for purification of large proteins. J. Chromatogr. A 2004, 1034, 155–159. [Google Scholar] [CrossRef]
- Fuentes, M.; Mateo, C.; Pessela, B.C.C.; Guisan, J.M.; Fernandez-Lafuente, R. Purification, stabilization, and concentration of very weak protein-protein complexes: Shifting the association equilibrium via complex selective adsorption on lowly activated supports. Proteomics 2005, 5, 4062–4069. [Google Scholar] [CrossRef]
- Kvasnov, B.A.; Shalaev, P.V.; Dolgushin, S.A. Study of the aggregation of bovine serum albumin monomers in aqueous dispersions with different acid-base value pH. In Proceedings of the 2018 IEEE Conference of Russian Young Researchers in Electrical and Electronic Engineering (ElConRus), Moscow, Russia, 29 January–1 February 2018; pp. 1907–1909. [Google Scholar]
- Polyanichko, A.M.; Mikhailov, N.V.; Romanov, N.M.; Baranova, Y.G.; Chikhirzhina, E.V. Intermolecular Interactions in Solutions of Serum Albumin. Cell Tissue Biol. 2017, 11, 9–15. [Google Scholar] [CrossRef]
- Militello, V.; Casarino, C.; Emanuele, A.; Giostra, A.; Pullara, F.; Leone, M. Aggregation kinetics of bovine serum albumin studied by FTIR spectroscopy and light scattering. Biophys. Chem. 2004, 107, 175–187. [Google Scholar] [CrossRef] [PubMed]
- Militello, V.; Vetri, V.; Leone, M. Conformational changes involved in thermal aggregation processes of bovine serum albumin. Biophys. Chem. 2003, 105, 133–141. [Google Scholar] [CrossRef]
- Fuentes, M.; Pessela, B.C.C.; Mateo, C.; Palomo, J.M.; Batalla, P.; Fernandez-Lafuente, R.; Guisan, J.M. Adsorption behavior of bovine serum albumin on lowly activated anionic exchangers suggests a new strategy for solid-phase proteomics. Biomacromolecules 2006, 7, 1357–1361. [Google Scholar] [CrossRef] [PubMed]
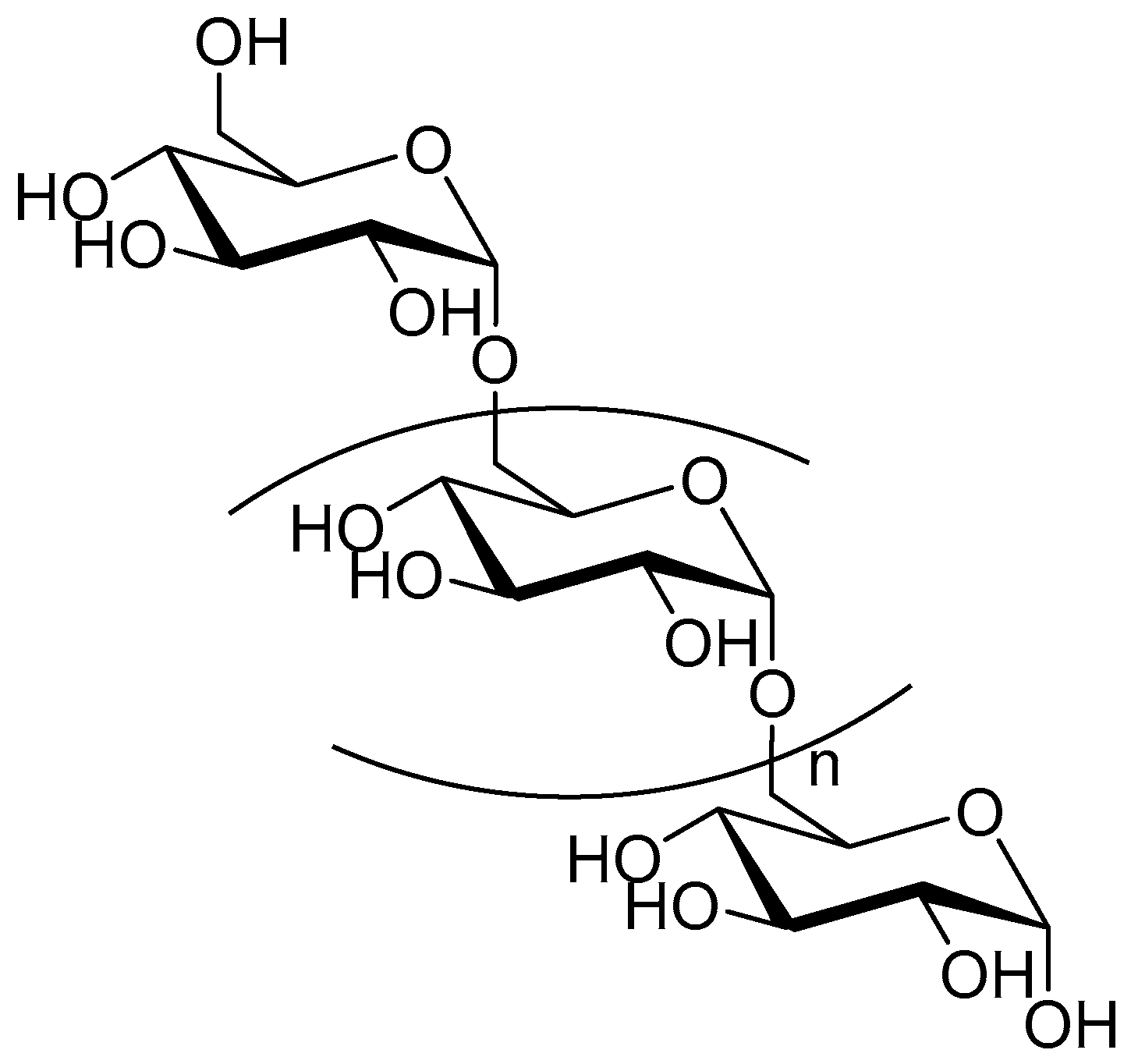


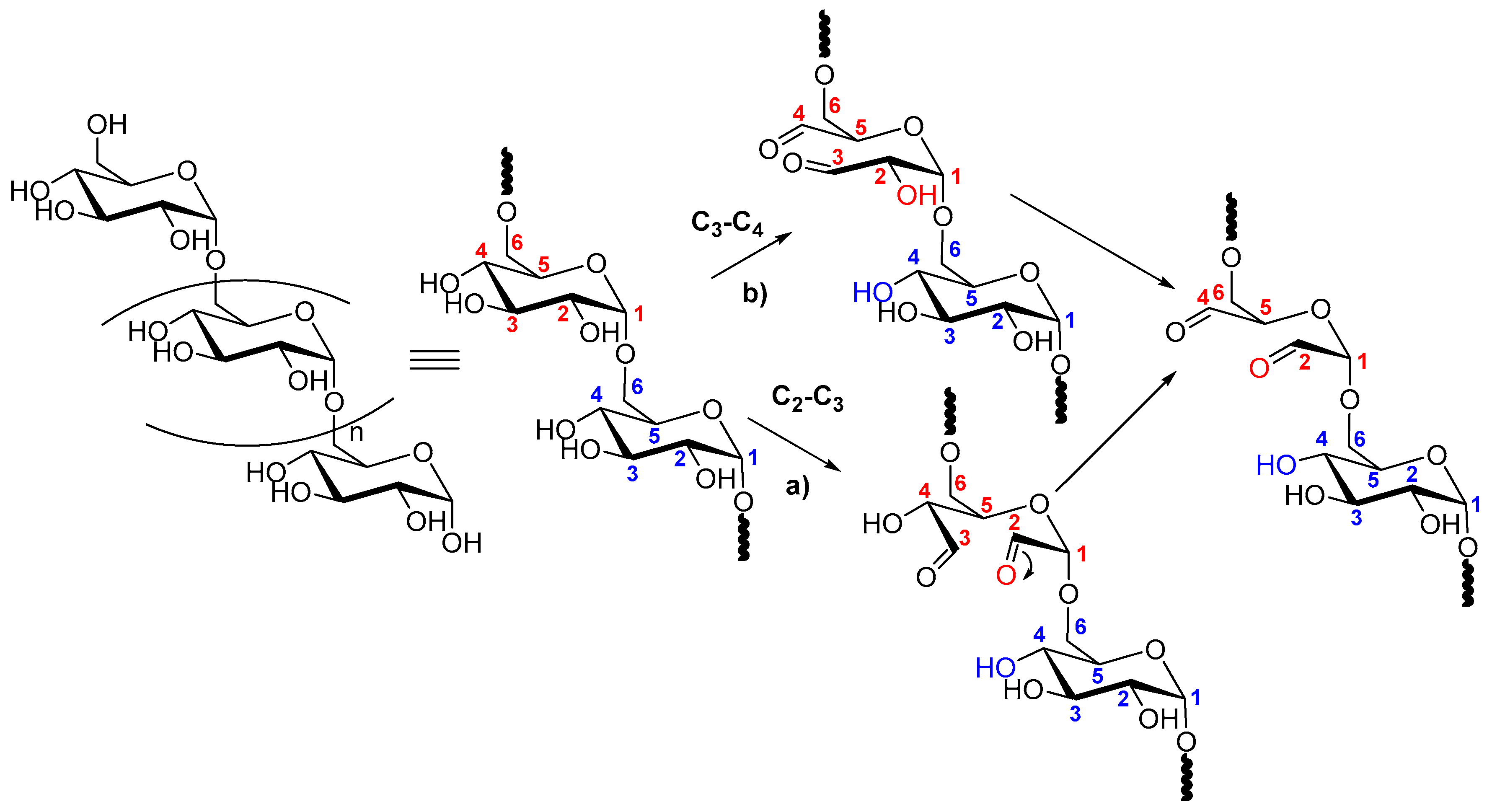
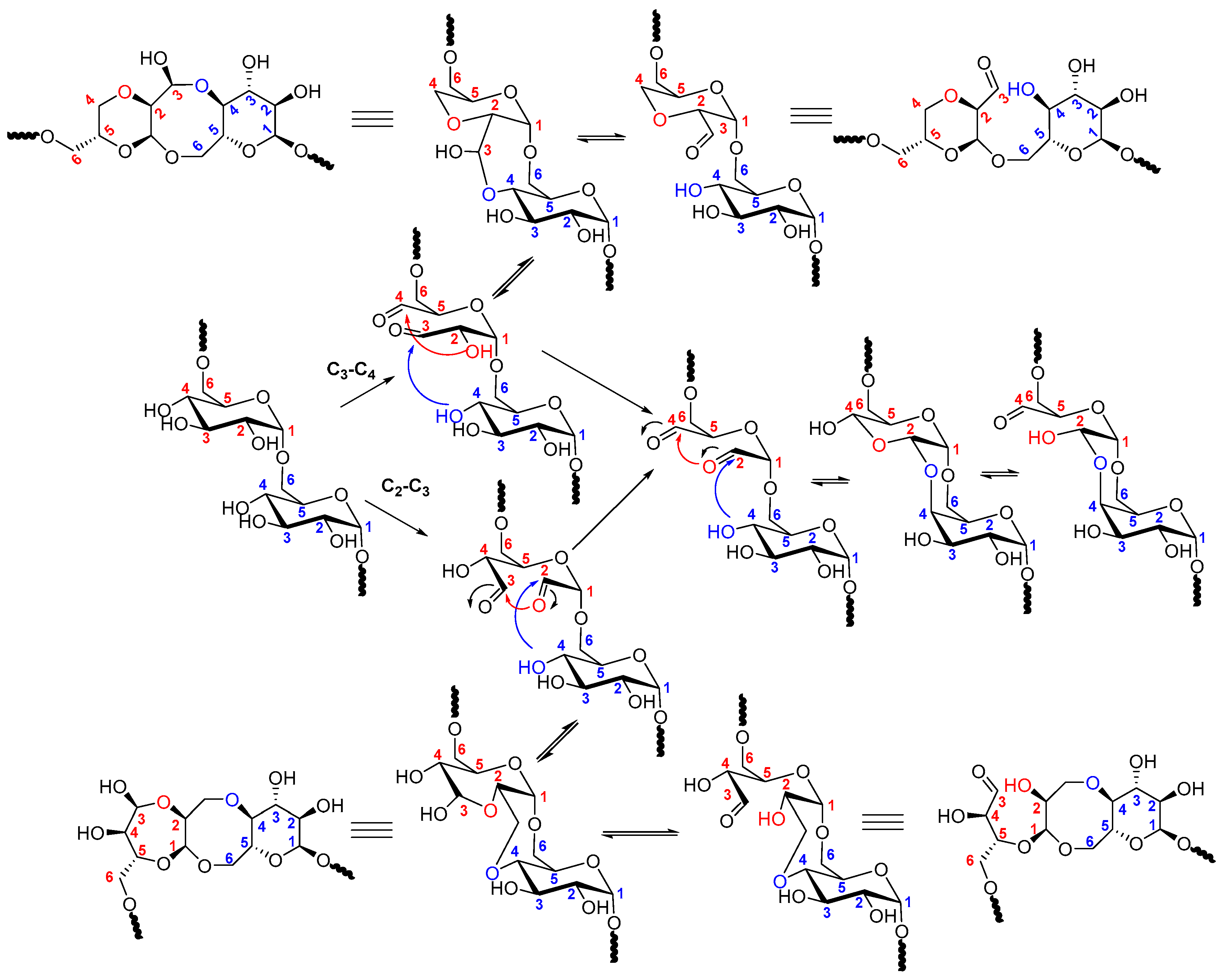
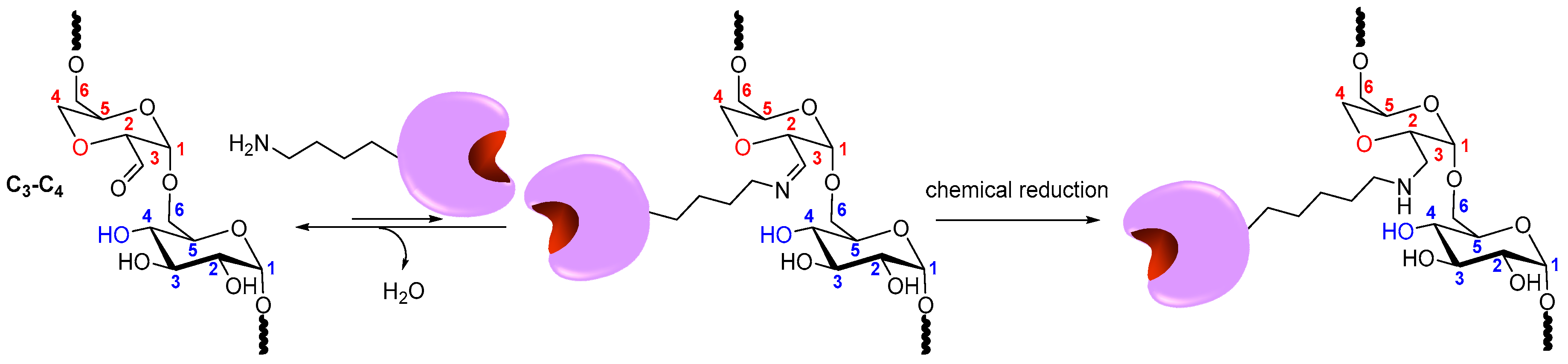
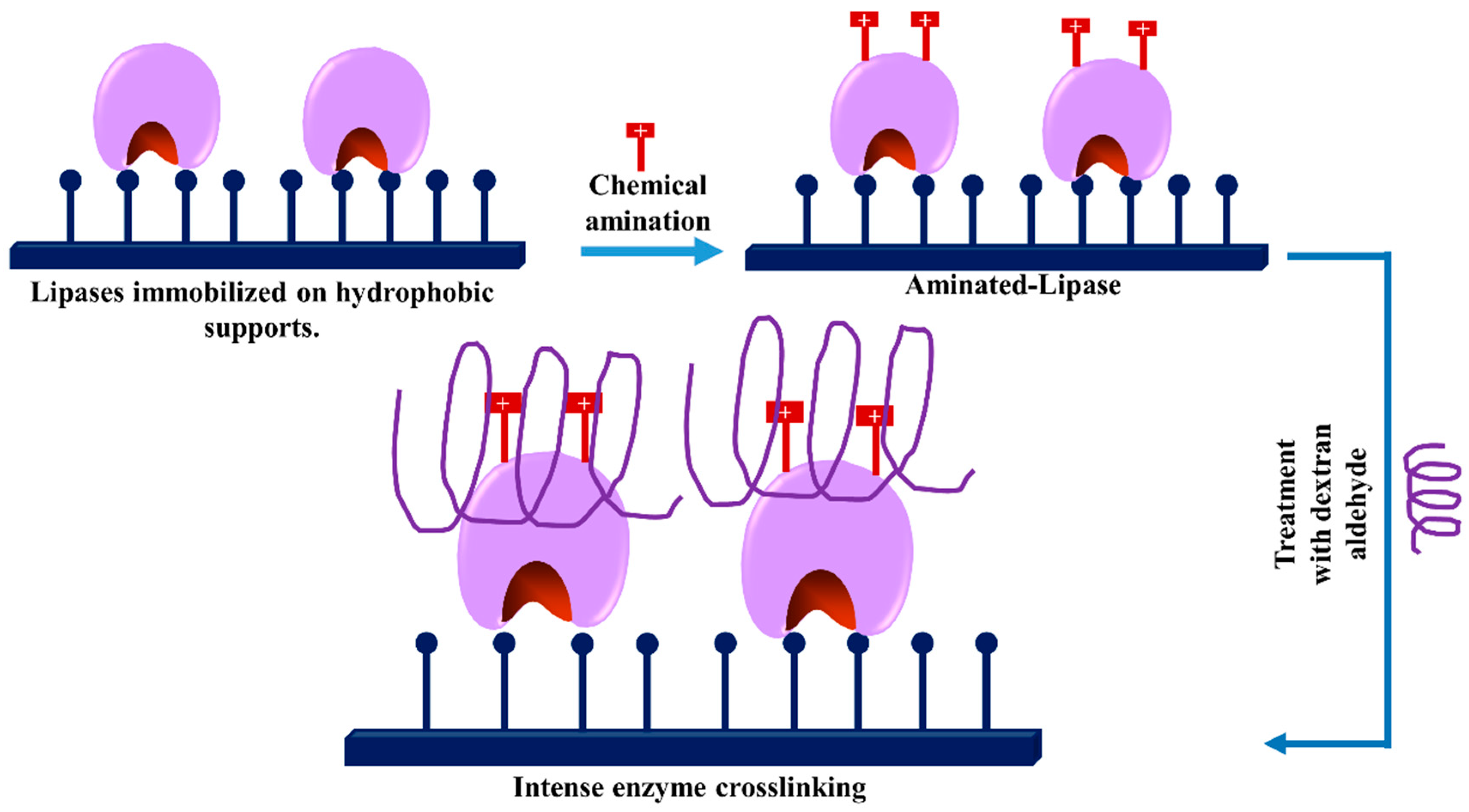

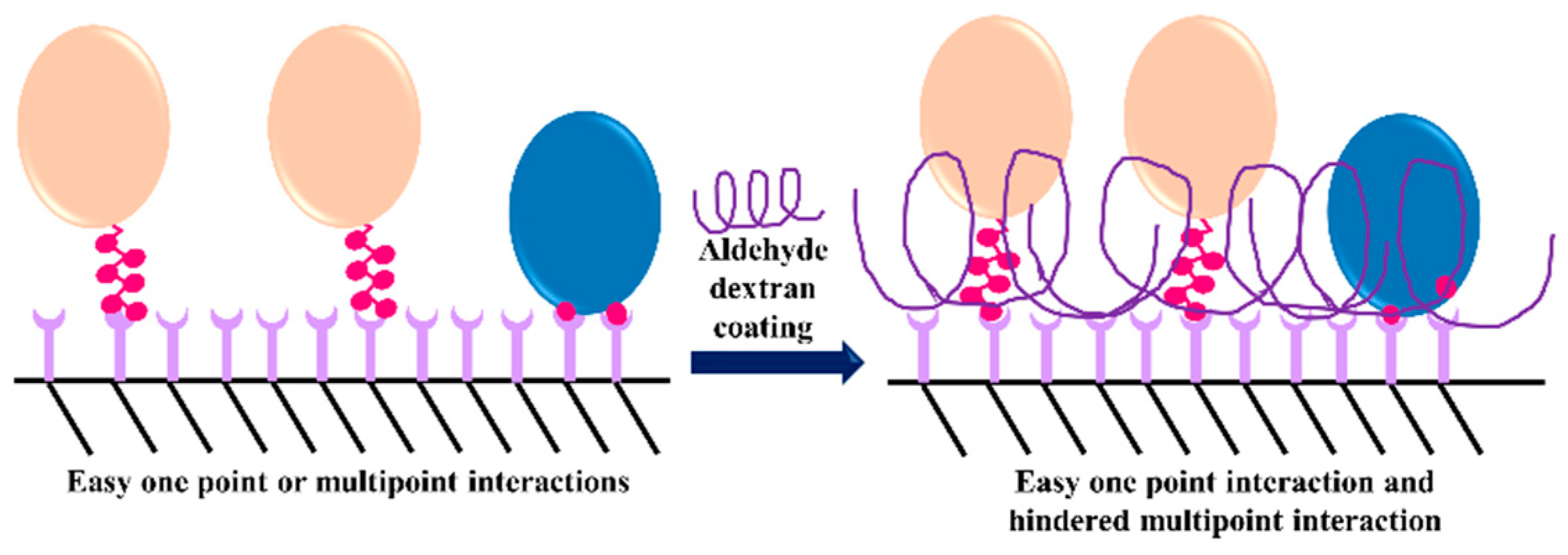

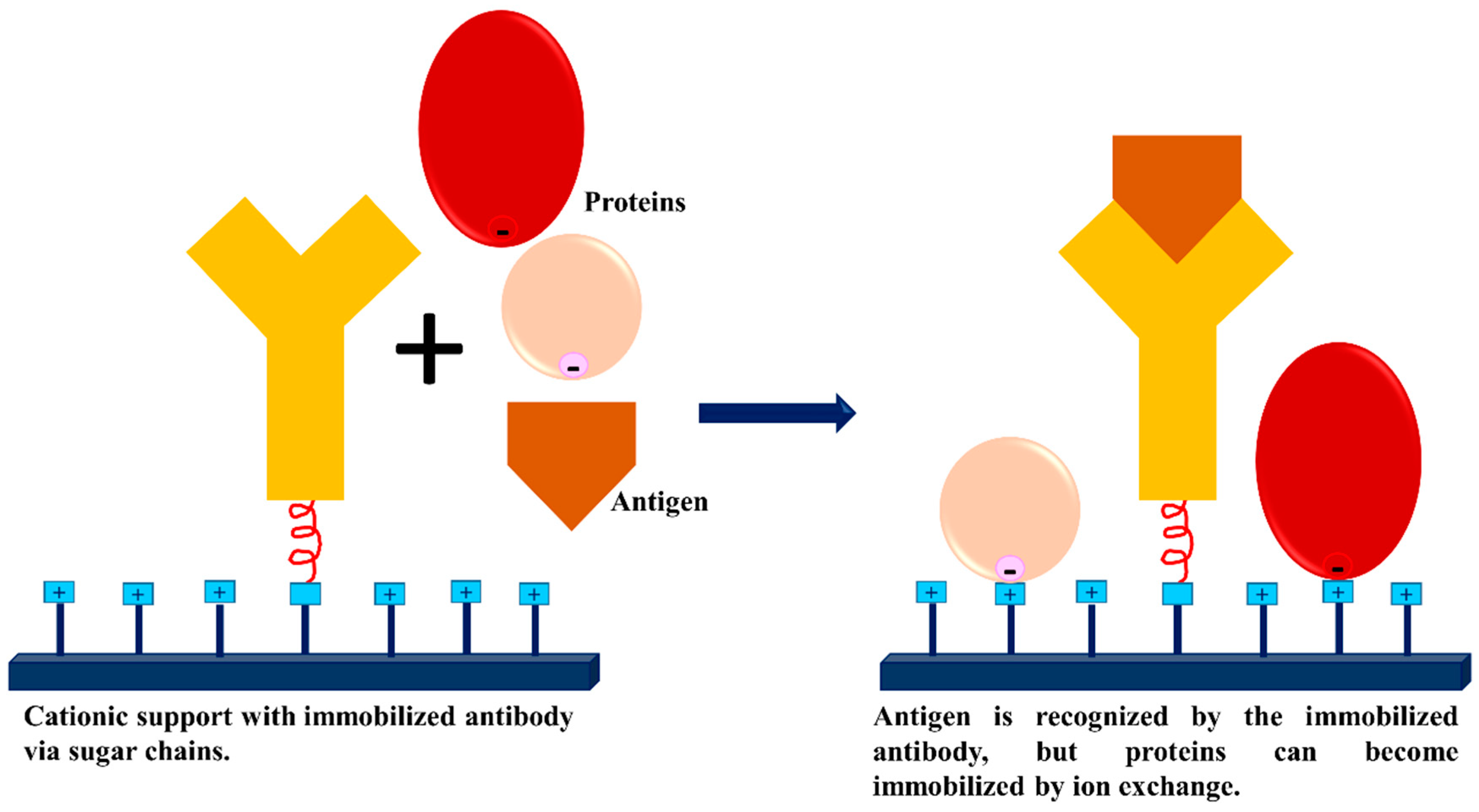
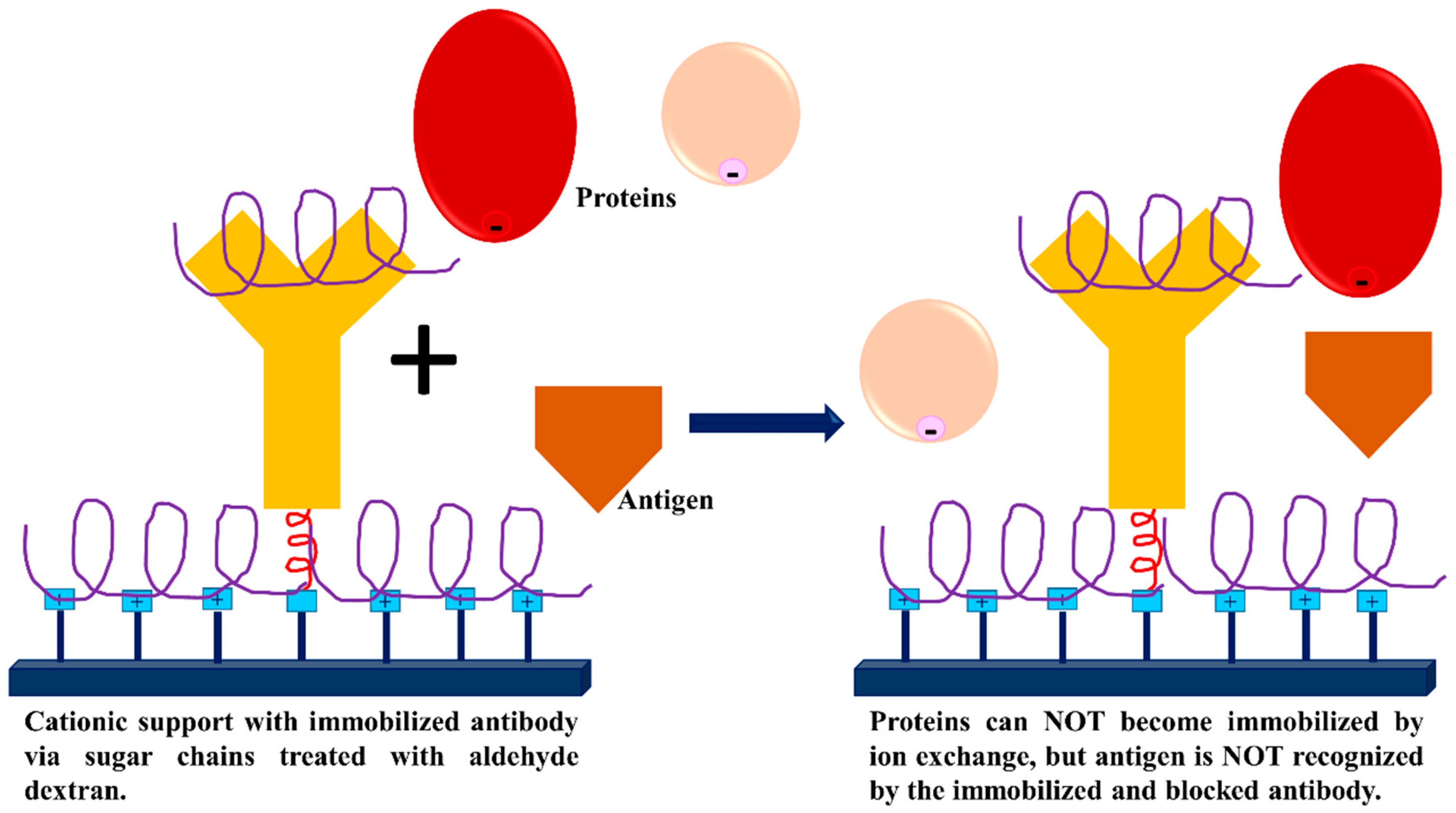
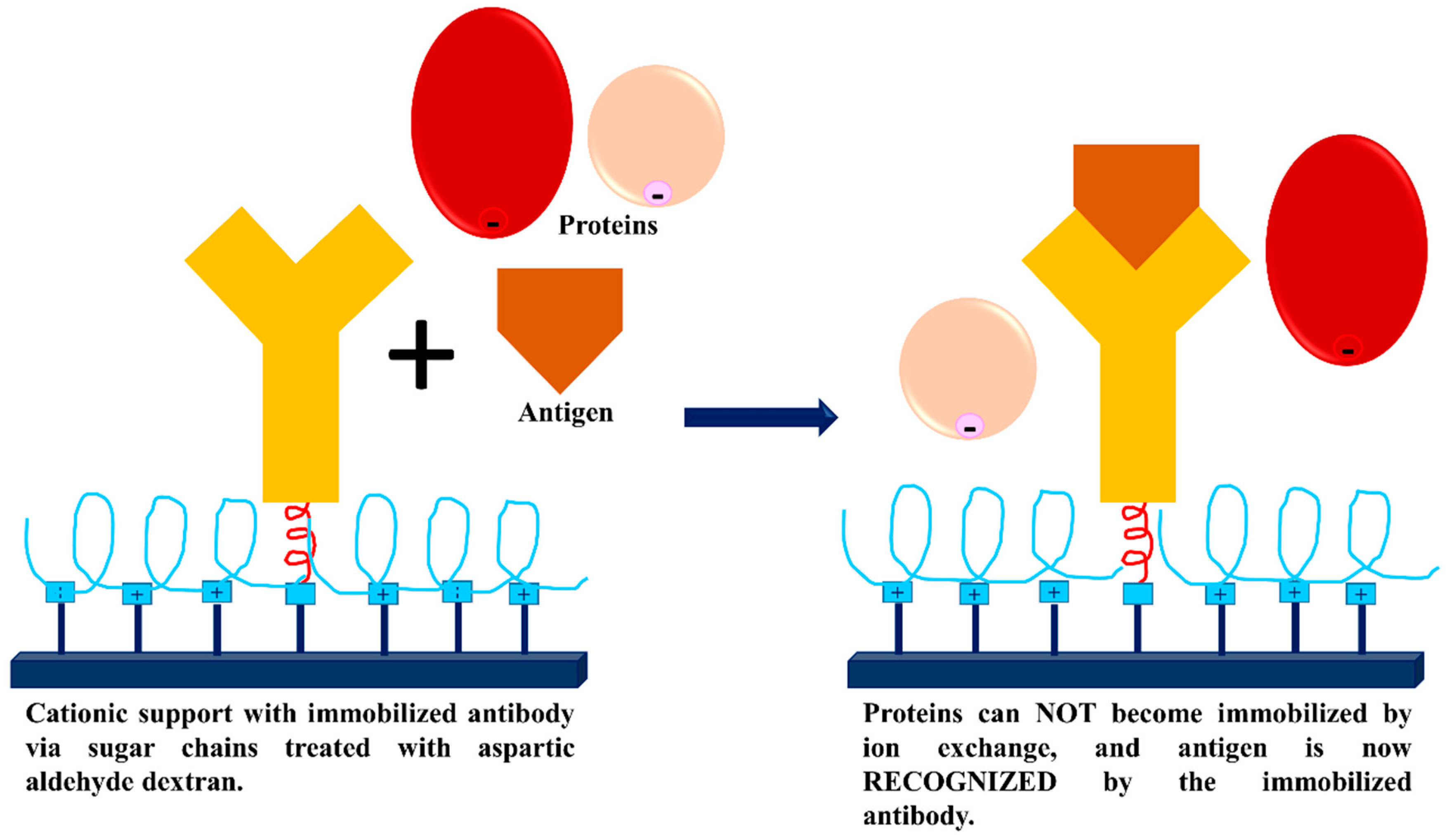

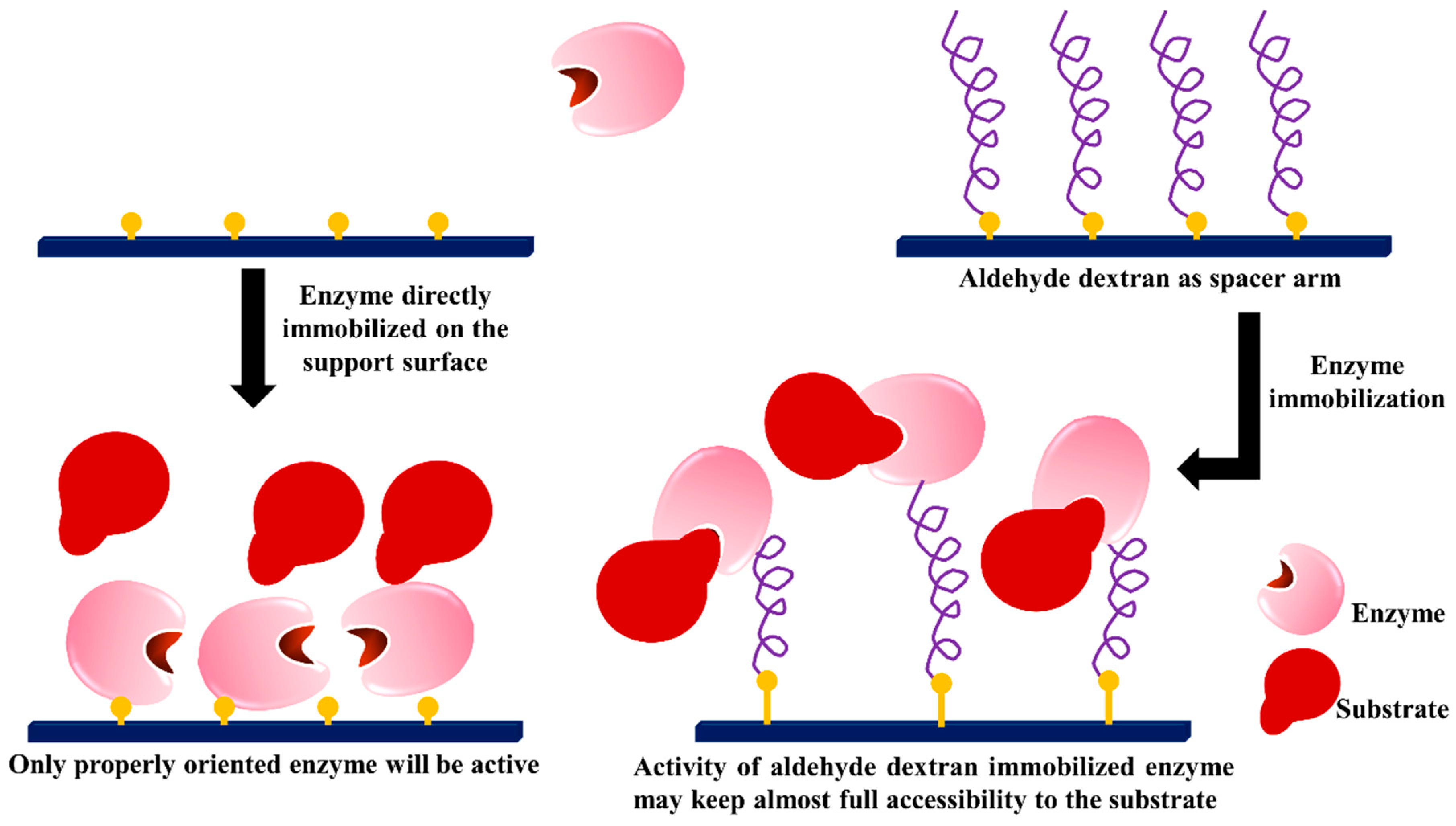
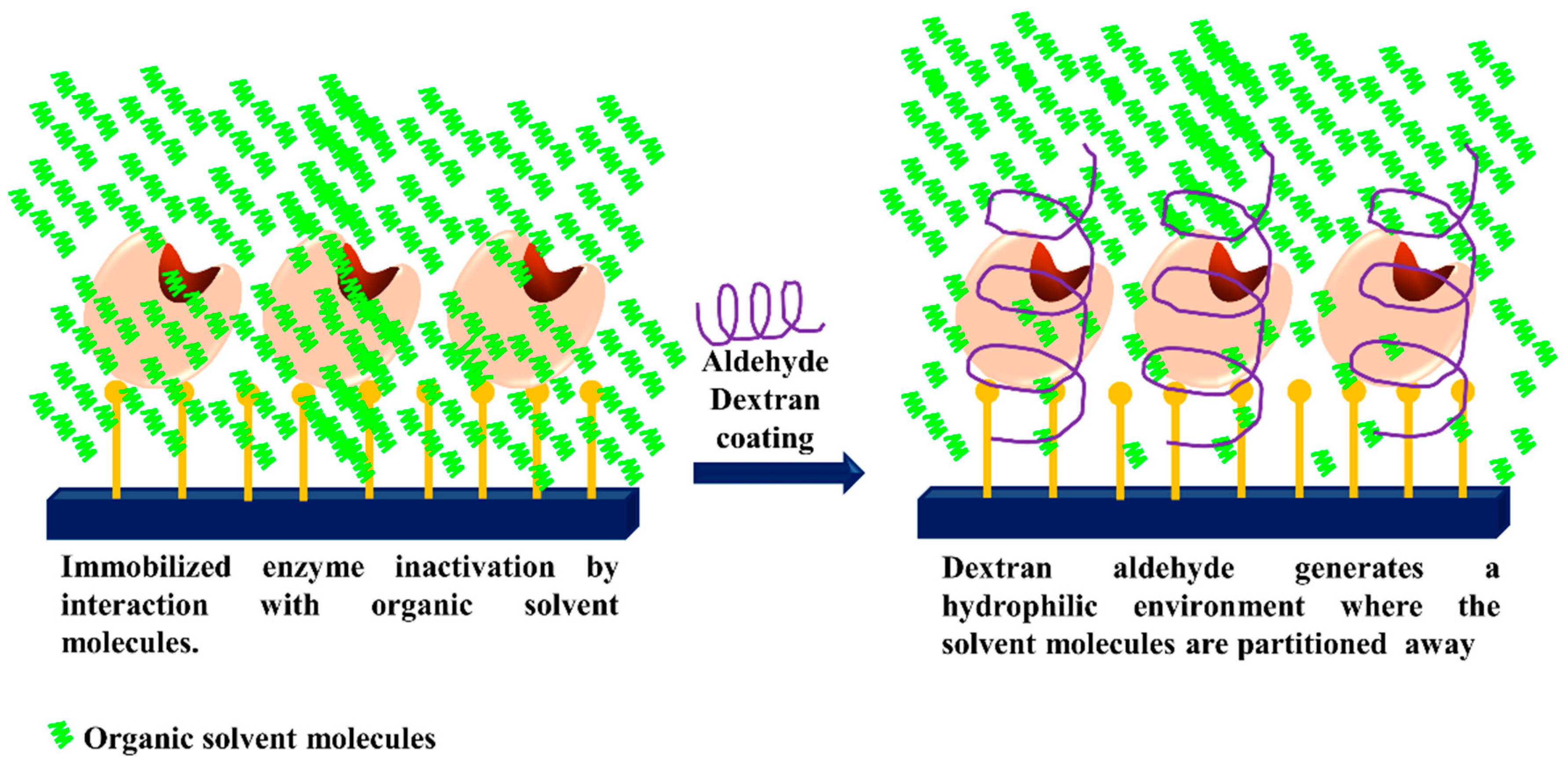
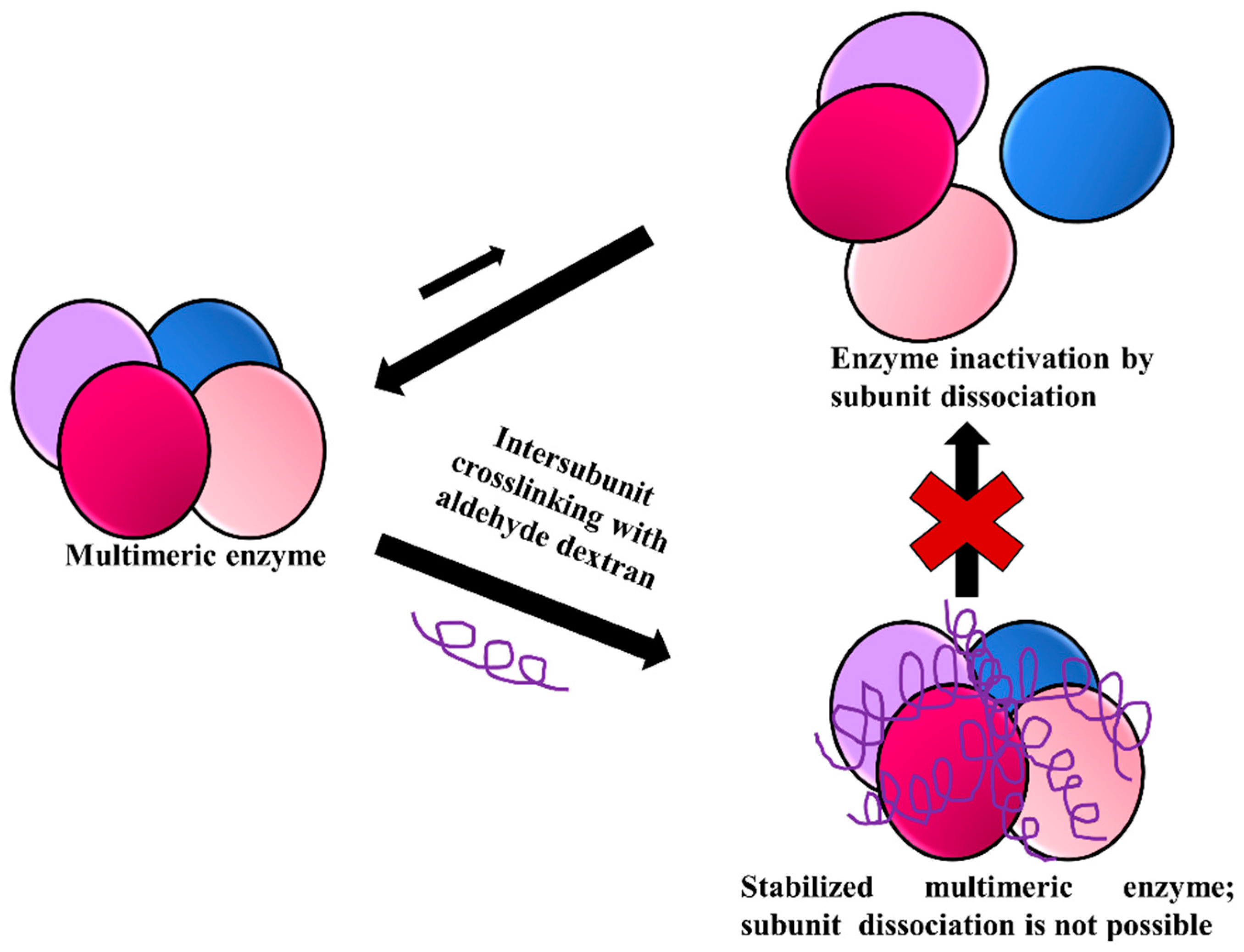
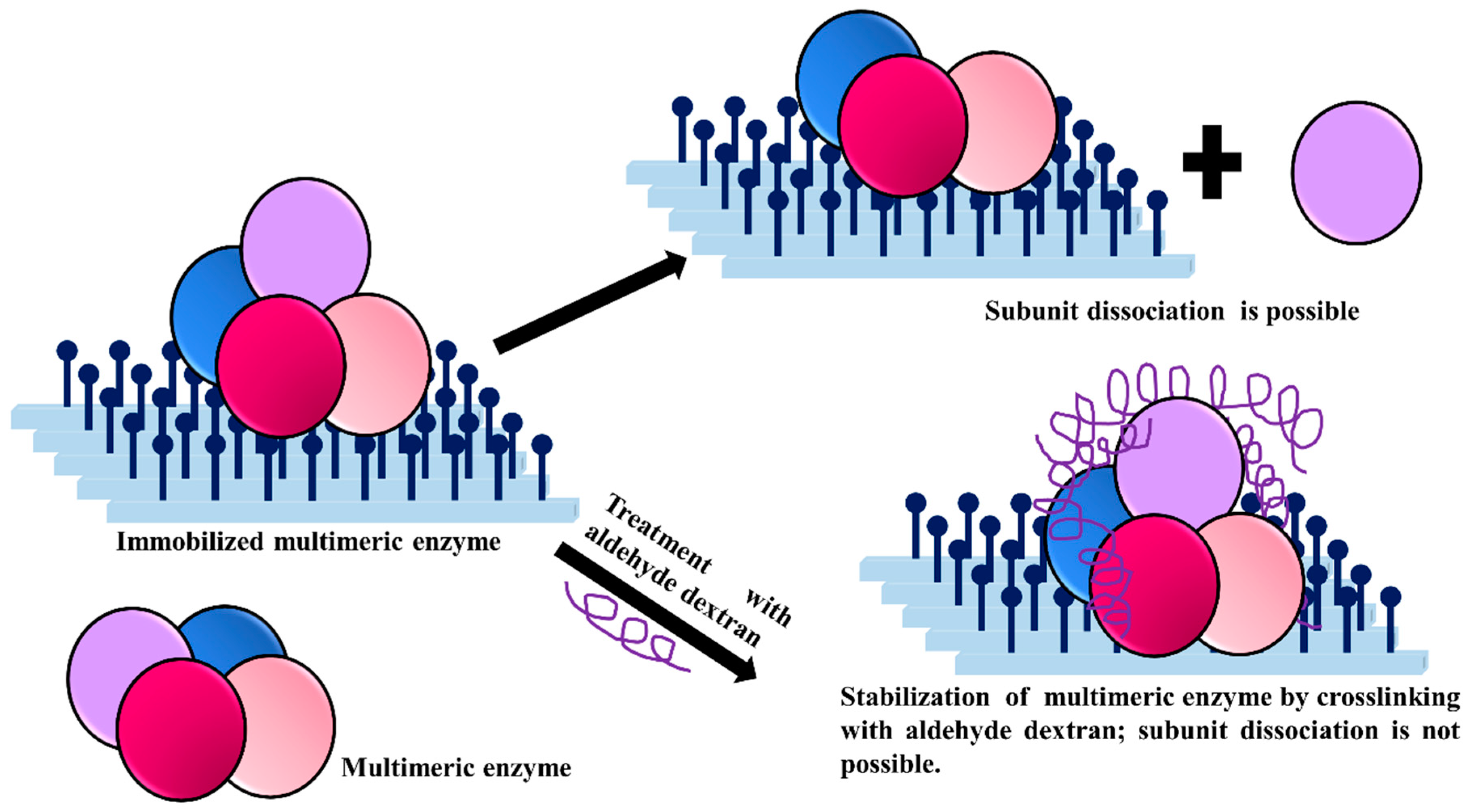


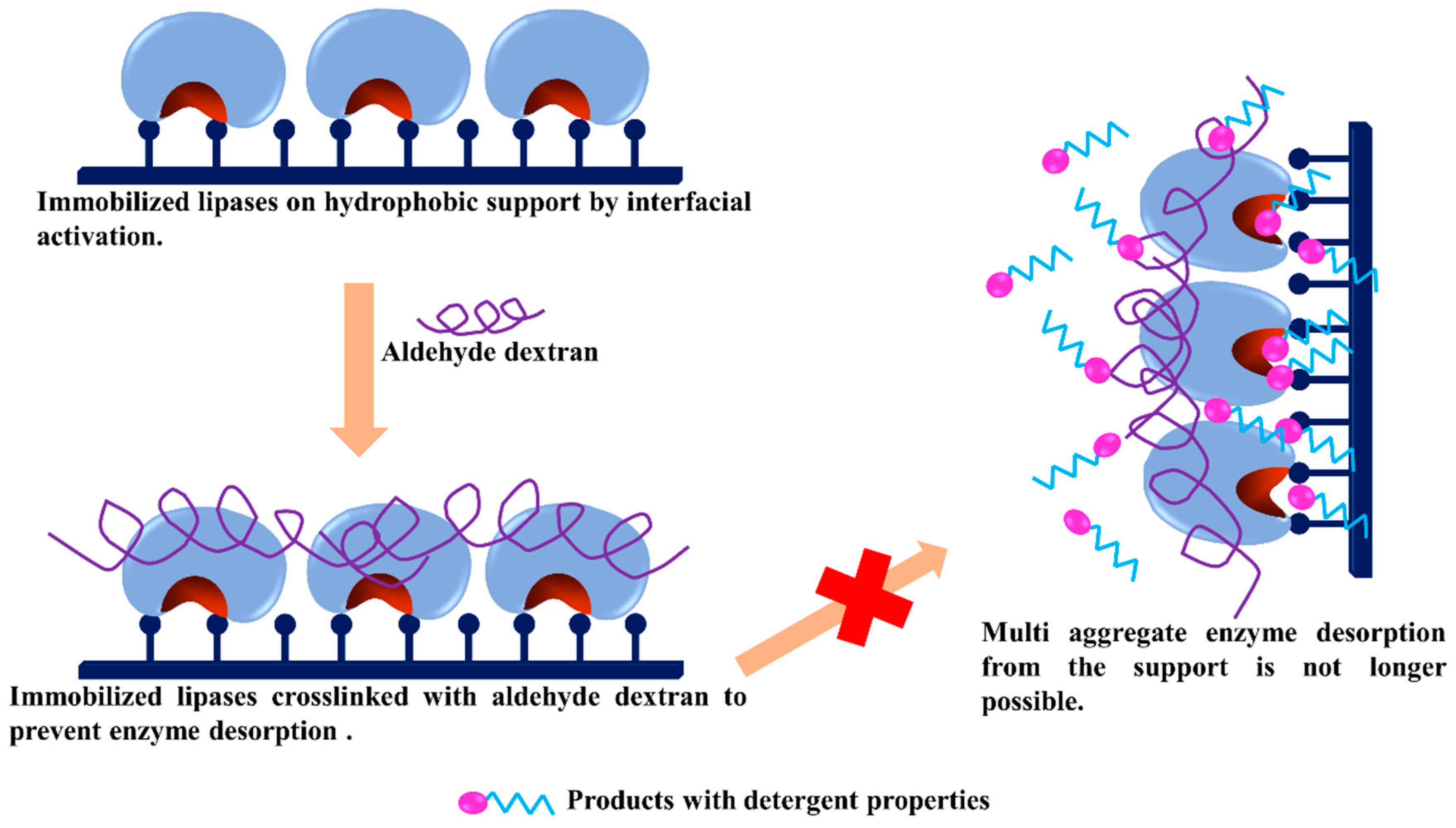
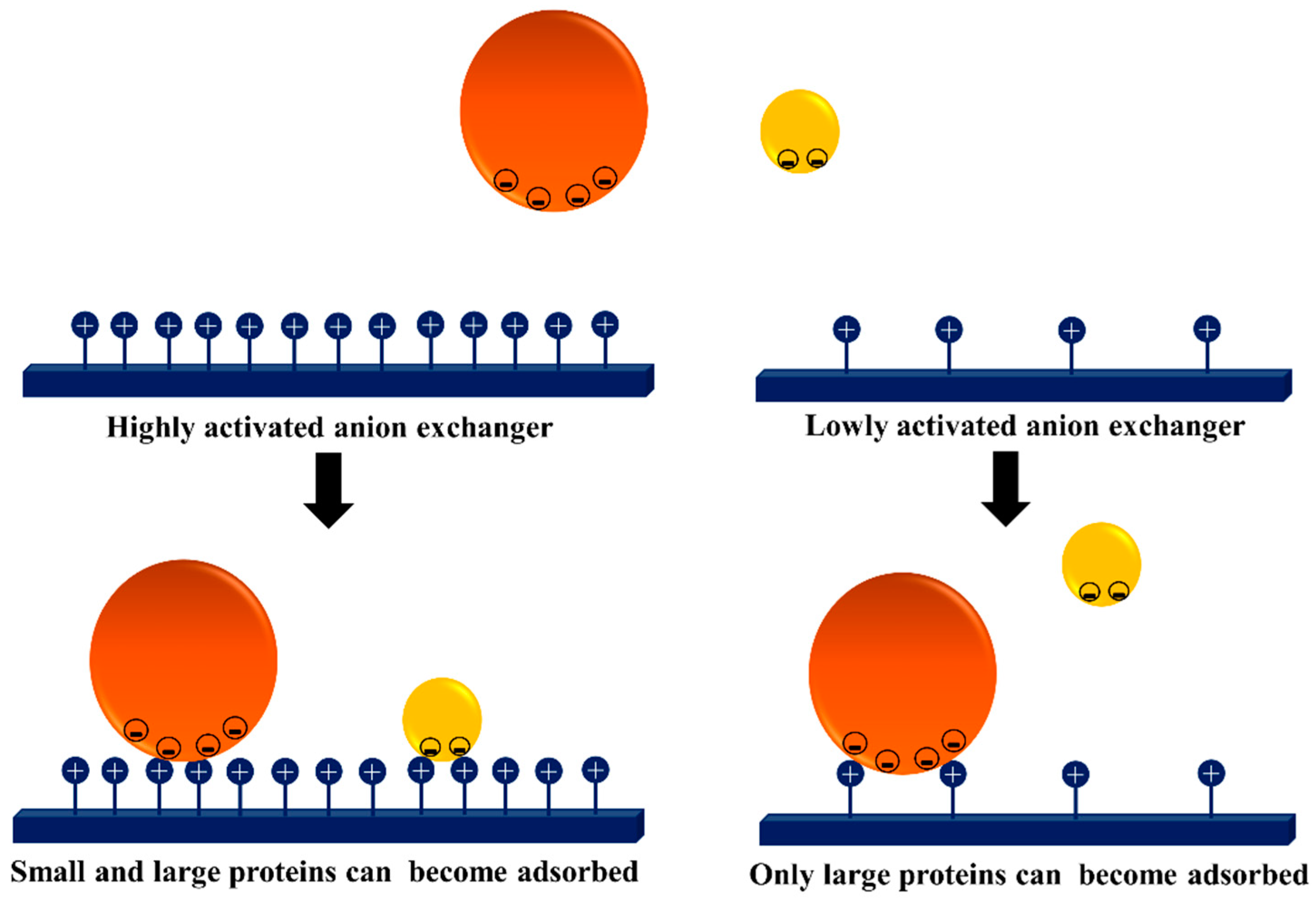

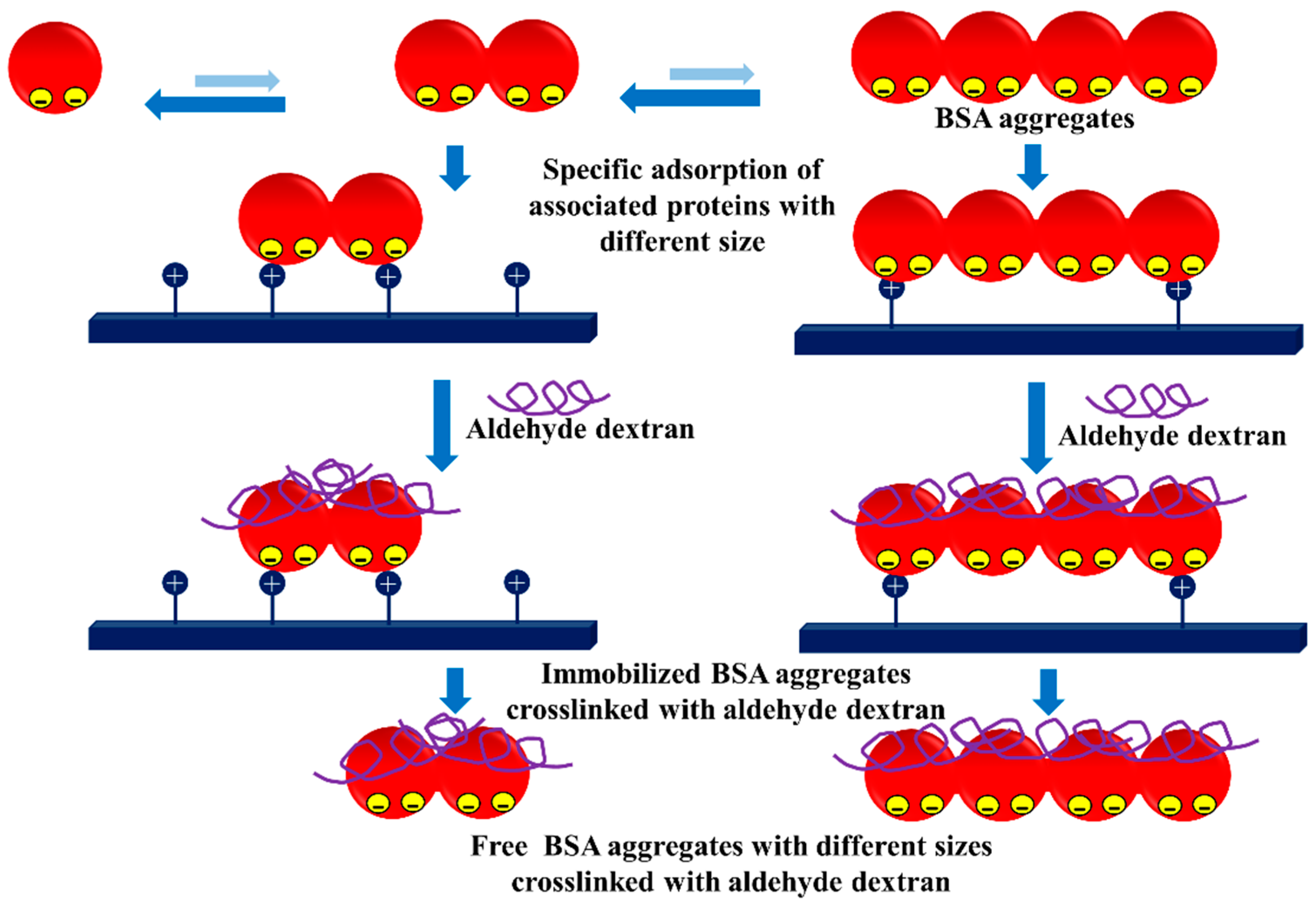
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tacias-Pascacio, V.G.; Ortiz, C.; Rueda, N.; Berenguer-Murcia, Á.; Acosta, N.; Aranaz, I.; Civera, C.; Fernandez-Lafuente, R.; Alcántara, A.R. Dextran Aldehyde in Biocatalysis: More Than a Mere Immobilization System. Catalysts 2019, 9, 622. https://doi.org/10.3390/catal9070622
Tacias-Pascacio VG, Ortiz C, Rueda N, Berenguer-Murcia Á, Acosta N, Aranaz I, Civera C, Fernandez-Lafuente R, Alcántara AR. Dextran Aldehyde in Biocatalysis: More Than a Mere Immobilization System. Catalysts. 2019; 9(7):622. https://doi.org/10.3390/catal9070622
Chicago/Turabian StyleTacias-Pascacio, Veymar G., Claudia Ortiz, Nazzoly Rueda, Ángel Berenguer-Murcia, Niuris Acosta, Inmaculada Aranaz, Concepción Civera, Roberto Fernandez-Lafuente, and Andrés R. Alcántara. 2019. "Dextran Aldehyde in Biocatalysis: More Than a Mere Immobilization System" Catalysts 9, no. 7: 622. https://doi.org/10.3390/catal9070622
APA StyleTacias-Pascacio, V. G., Ortiz, C., Rueda, N., Berenguer-Murcia, Á., Acosta, N., Aranaz, I., Civera, C., Fernandez-Lafuente, R., & Alcántara, A. R. (2019). Dextran Aldehyde in Biocatalysis: More Than a Mere Immobilization System. Catalysts, 9(7), 622. https://doi.org/10.3390/catal9070622