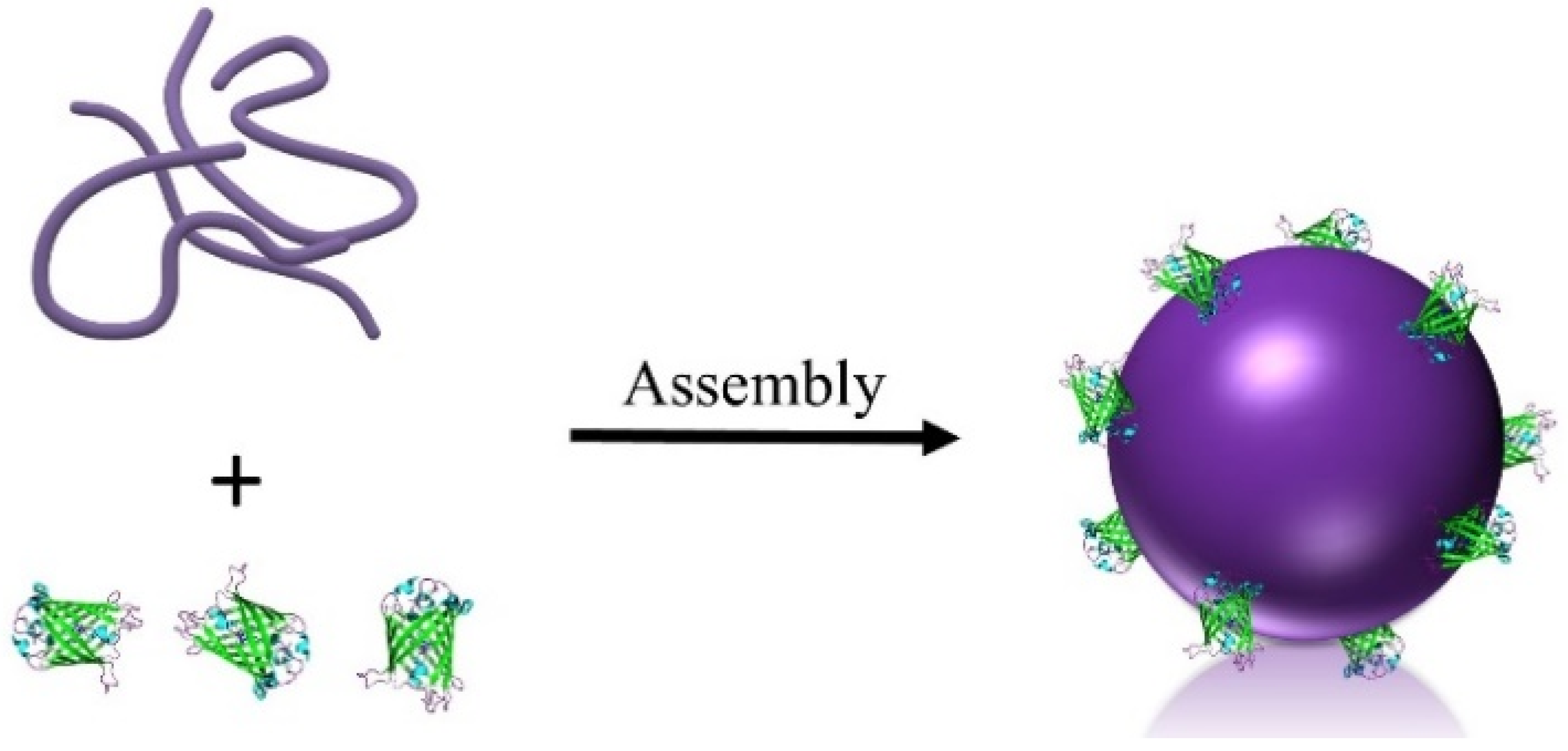
Synthesis and Characterization of Bio-Active GFP-P4VP Core–Shell Nanoparticles
Abstract
1. Introduction
2. Results and Discussion
2.1. Synthesis and Purification
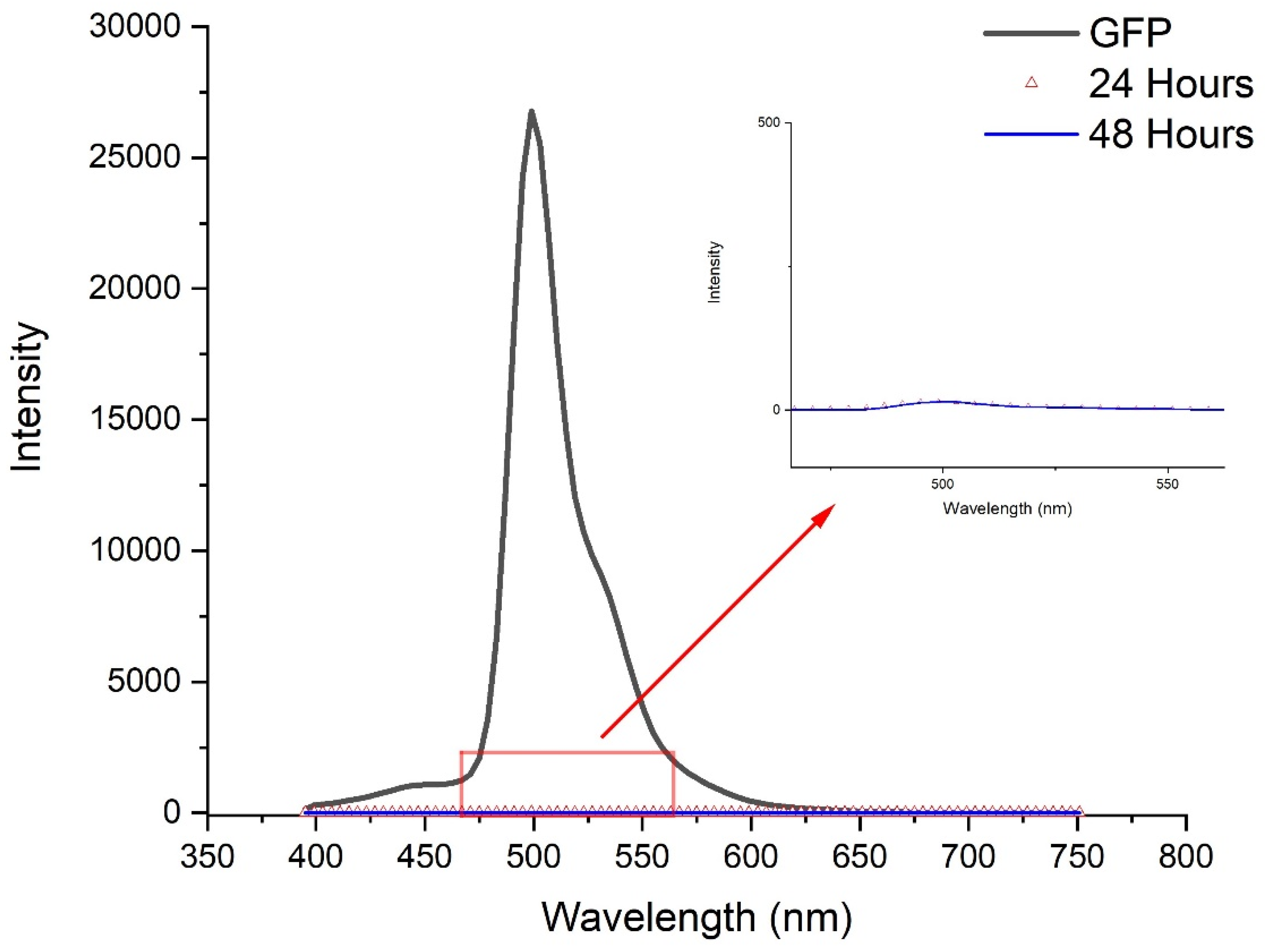
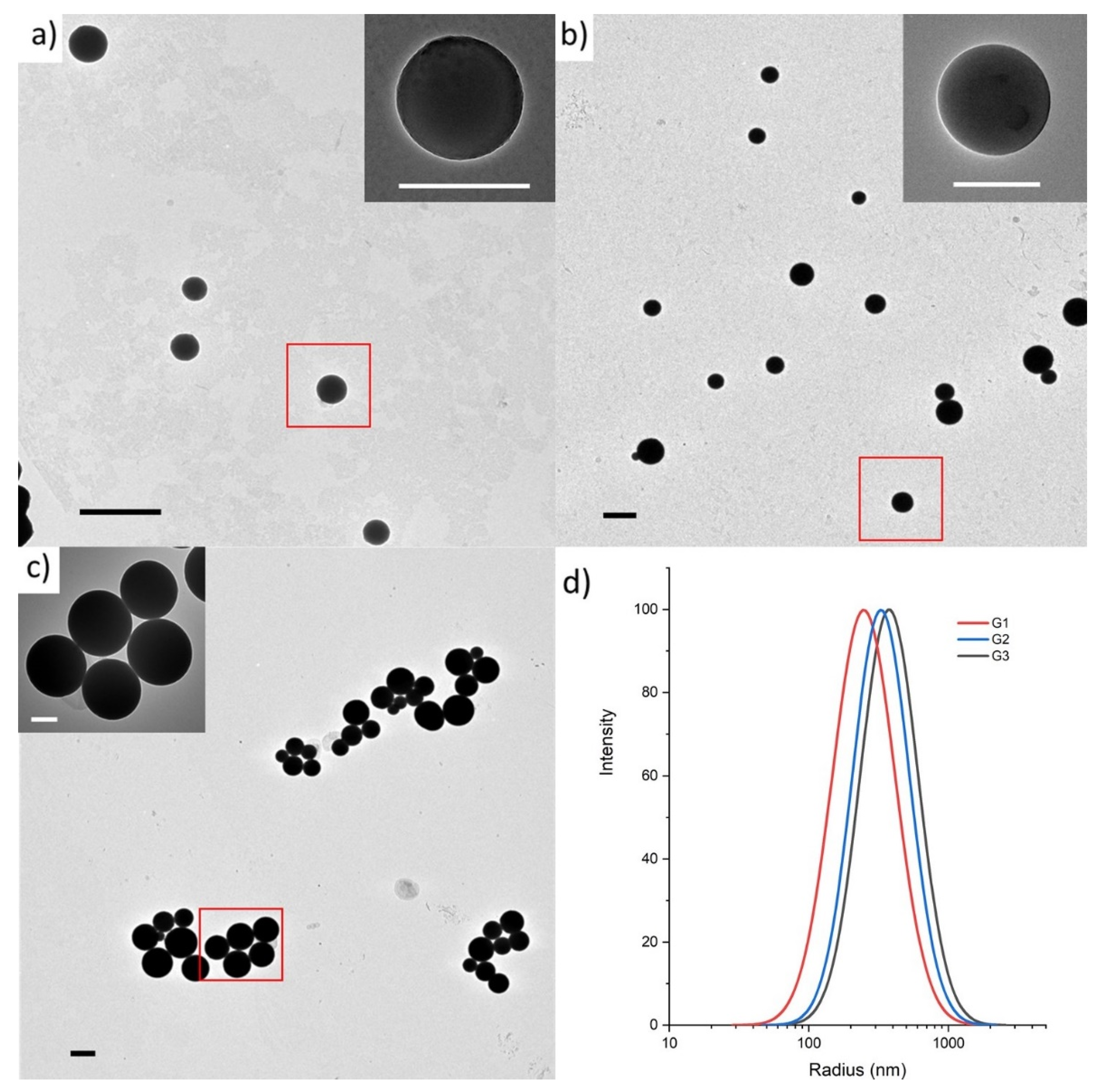
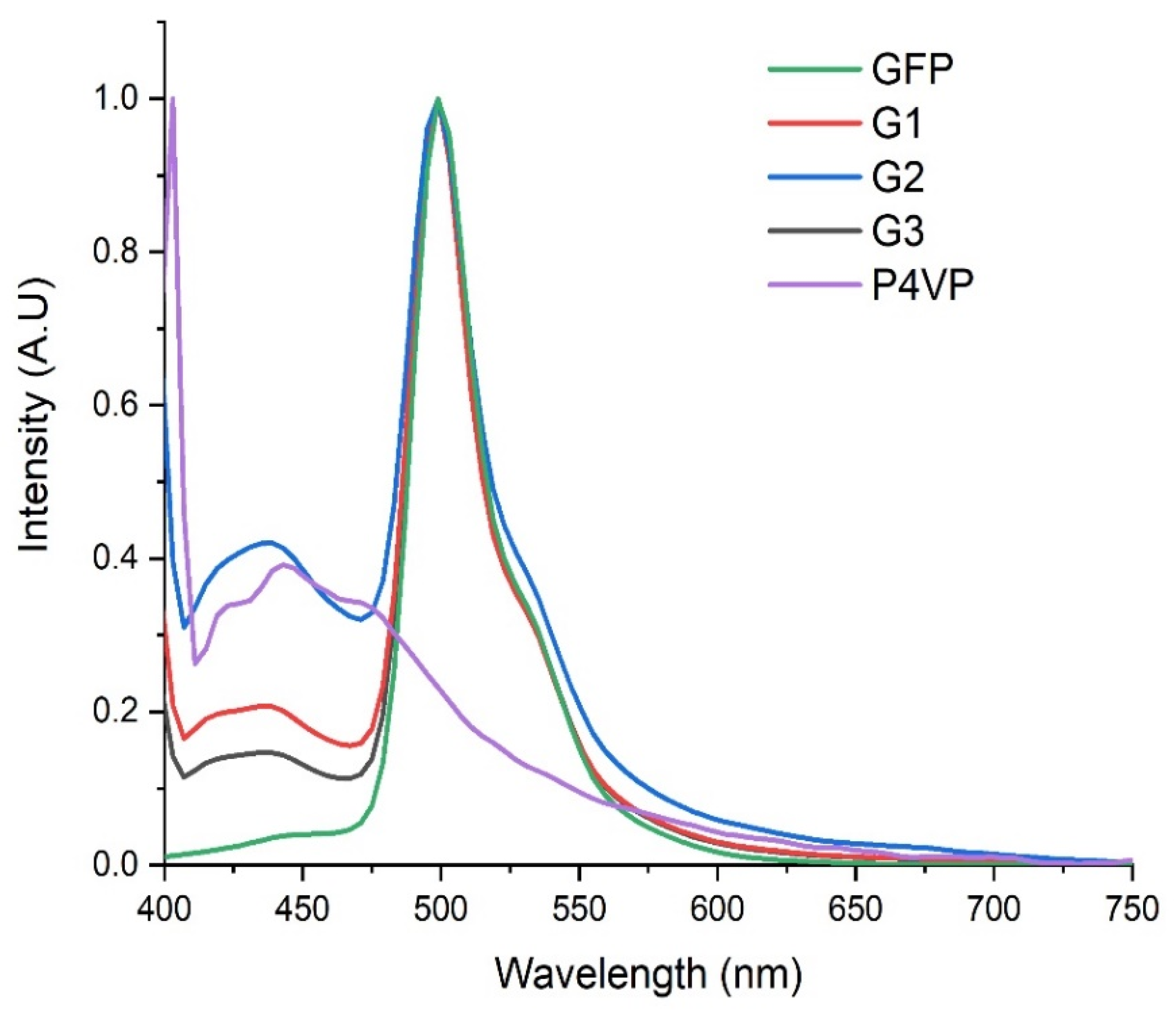
2.2. Size and Surface Characterization
3. Experimental Section
3.1. Chemicals and Materials
3.2. Sample Preparation
3.3. Characterization
3.4. Gold–GFP Conjugation
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Li, S.; Yang, X.; Yang, S.; Zhu, M.; Wang, X. Technology Prospecting on Enzymes: Application, Marketing And Engineering. Comput. Struct. Biotechnol. J. 2012, 2, e201209017. [Google Scholar] [CrossRef] [PubMed]
- Kirk, O.; Borchert, T.V.; Fuglsang, C.C. Industrial enzyme applications. Curr. Opin. Biotechnol. 2002, 13, 345–351. [Google Scholar] [CrossRef]
- Morgenstern, J.; Gil Alvaradejo, G.; Bluthardt, N.; Beloqui, A.; Delaittre, G.; Hubbuch, J. Impact of Polymer Bioconjugation on Protein Stability and Activity Investigated with Discrete Conjugates: Alternatives to PEGylation. Biomacromolecules 2018, 19, 4250–4262. [Google Scholar] [CrossRef] [PubMed]
- Wan, L.; Chen, Q.; Liu, J.; Yang, X.; Huang, J.; Li, L.; Guo, X.; Zhang, J.; Wang, K. Programmable Self-Assembly of DNA–Protein Hybrid Hydrogel for Enzyme Encapsulation with Enhanced Biological Stability. Biomacromolecules 2016, 17, 1543–1550. [Google Scholar] [CrossRef] [PubMed]
- Murata, H.; Cummings, C.S.; Koepsel, R.R.; Russell, A.J. Polymer-Based Protein Engineering Can Rationally Tune Enzyme Activity, pH-Dependence, and Stability. Biomacromolecules 2013, 14, 1919–1926. [Google Scholar] [CrossRef]
- Hoener, C.F.; Allan, K.A.; Bard, A.J.; Campion, A.; Fox, M.A.; Mallouk, T.E.; Webber, S.E.; White, J.M. Demonstration of a shell-core structure in layered cadmium selenide-zinc selenide small particles by x-ray photoelectron and Auger spectroscopies. J. Phys. Chem. 1992, 96, 3812–3817. [Google Scholar] [CrossRef]
- Ghosh Chaudhuri, R.; Paria, S. Core/Shell Nanoparticles: Classes, Properties, Synthesis Mechanisms, Characterization, and Applications. Chem. Rev. 2012, 112, 2373–2433. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wu, H.-C.; Chhuan, M.; Terrell, J.L.; Tsao, C.-Y.; Bentley, W.E.; Payne, G.F. Functionalizing Soft Matter for Molecular Communication. Acs Biomater. Sci. Eng. 2015, 1, 320–328. [Google Scholar] [CrossRef]
- Chen, X.; Gerasopoulos, K.; Guo, J.; Brown, A.; Wang, C.; Ghodssi, R.; Culver, J.N. Virus-Enabled Silicon Anode for Lithium-Ion Batteries. Acs Nano 2010, 4, 5366–5372. [Google Scholar] [CrossRef] [PubMed]
- Gallat, F.-X.; Brogan, A.P.S.; Fichou, Y.; McGrath, N.; Moulin, M.; Härtlein, M.; Combet, J.; Wuttke, J.; Mann, S.; Zaccai, G.; et al. A Polymer Surfactant Corona Dynamically Replaces Water in Solvent-Free Protein Liquids and Ensures Macromolecular Flexibility and Activity. J. Am. Chem. Soc. 2012, 134, 13168–13171. [Google Scholar] [CrossRef] [PubMed]
- Gebauer, J.S.; Malissek, M.; Simon, S.; Knauer, S.K.; Maskos, M.; Stauber, R.H.; Peukert, W.; Treuel, L. Impact of the Nanoparticle–Protein Corona on Colloidal Stability and Protein Structure. Langmuir 2012, 28, 9673–9679. [Google Scholar] [CrossRef] [PubMed]
- Klein, J. Probing the interactions of proteins and nanoparticles. Proc. Natl. Acad. Sci. USA 2007, 104, 2029–2030. [Google Scholar] [CrossRef] [PubMed]
- Mahmoudi, M.; Lynch, I.; Ejtehadi, M.R.; Monopoli, M.P.; Bombelli, F.B.; Laurent, S. Protein−Nanoparticle Interactions: Opportunities and Challenges. Chem. Rev. 2011, 111, 5610–5637. [Google Scholar] [CrossRef] [PubMed]
- Tao, L.; Mantovani, G.; Lecolley, F.; Haddleton, D.M. α-Aldehyde Terminally Functional Methacrylic Polymers from Living Radical Polymerization: Application in Protein Conjugation “Pegylation”. J. Am. Chem. Soc. 2004, 126, 13220–13221. [Google Scholar] [CrossRef] [PubMed]
- Bartczak, D.; Kanaras, A.G. Preparation of Peptide-Functionalized Gold Nanoparticles Using One Pot EDC/Sulfo-NHS Coupling. Langmuir 2011, 27, 10119–10123. [Google Scholar] [CrossRef] [PubMed]
- Liu, E.Y.; Jung, S.; Yi, H. Improved Protein Conjugation with Uniform, Macroporous Poly(acrylamide-co-acrylic acid) Hydrogel Microspheres via EDC/NHS Chemistry. Langmuir 2016, 32, 11043–11054. [Google Scholar] [CrossRef]
- Totaro, K.A.; Liao, X.; Bhattacharya, K.; Finneman, J.I.; Sperry, J.B.; Massa, M.A.; Thorn, J.; Ho, S.V.; Pentelute, B.L. Systematic Investigation of EDC/sNHS-Mediated Bioconjugation Reactions for Carboxylated Peptide Substrates. Bioconjugate Chem. 2016, 27, 994–1004. [Google Scholar] [CrossRef]
- Heredia, K.L.; Bontempo, D.; Ly, T.; Byers, J.T.; Halstenberg, S.; Maynard, H.D. In Situ Preparation of Protein−“Smart” Polymer Conjugates with Retention of Bioactivity. J. Am. Chem. Soc. 2005, 127, 16955–16960. [Google Scholar] [CrossRef]
- Vertegel, A.A.; Siegel, R.W.; Dordick, J.S. Silica Nanoparticle Size Influences the Structure and Enzymatic Activity of Adsorbed Lysozyme. Langmuir 2004, 20, 6800–6807. [Google Scholar] [CrossRef]
- Kittler, S.; Greulich, C.; Gebauer, J.S.; Diendorf, J.; Treuel, L.; Ruiz, L.; Gonzalez-Calbet, J.M.; Vallet-Regi, M.; Zellner, R.; Köller, M.; et al. The influence of proteins on the dispersability and cell-biological activity of silver nanoparticles. J. Mater. Chem. 2010, 20, 512–518. [Google Scholar] [CrossRef]
- Lynch, I.; Salvati, A.; Dawson, K.A. What does the cell see? Nat. Nanotechnol. 2009, 4, 546–547. [Google Scholar] [CrossRef] [PubMed]
- Monopoli, M.P.; Walczyk, D.; Campbell, A.; Elia, G.; Lynch, I.; Baldelli Bombelli, F.; Dawson, K.A. Physical−Chemical Aspects of Protein Corona: Relevance to in Vitro and in Vivo Biological Impacts of Nanoparticles. J. Am. Chem. Soc. 2011, 133, 2525–2534. [Google Scholar] [CrossRef] [PubMed]
- Suthiwangcharoen, N.; Li, T.; Wu, L.; Reno, H.B.; Thompson, P.; Wang, Q. Facile Co-Assembly Process to Generate Core–Shell Nanoparticles with Functional Protein Corona. Biomacromolecules 2014, 15, 948–956. [Google Scholar] [CrossRef] [PubMed]
- Shu, J.Y.; Huang, Y.-J.; Tan, C.; Presley, A.D.; Chang, J.; Xu, T. Amphiphilic Peptide−Polymer Conjugates Based on the Coiled-Coil Helix Bundle. Biomacromolecules 2010, 11, 1443–1452. [Google Scholar] [CrossRef] [PubMed]
- Parajuli, O.; Gupta, A.; Kumar, N.; Hahm, J.-i. Evaluation of Enzymatic Activity on Nanoscale Polystyrene-block-Polymethylmethacrylate Diblock Copolymer Domains. J. Phys. Chem. B 2007, 111, 14022–14027. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K.P.; Collins, A.M.; Perriman, A.W.; Mann, S. Enzymatically Active Self-Standing Protein-Polymer Surfactant Films Prepared by Hierarchical Self-Assembly. Adv. Mater. 2013, 25, 2005–2010. [Google Scholar] [CrossRef]
- Ventura, J.; Eron, S.J.; González-Toro, D.C.; Raghupathi, K.; Wang, F.; Hardy, J.A.; Thayumanavan, S. Reactive Self-Assembly of Polymers and Proteins to Reversibly Silence a Killer Protein. Biomacromolecules 2015, 16, 3161–3171. [Google Scholar] [CrossRef] [PubMed]
- Russell, J.T.; Lin, Y.; Böker, A.; Su, L.; Carl, P.; Zettl, H.; He, J.; Sill, K.; Tangirala, R.; Emrick, T.; et al. Self-Assembly and Cross-Linking of Bionanoparticles at Liquid–Liquid Interfaces. Angew. Chem. Int. Ed. 2005, 44, 2420–2426. [Google Scholar] [CrossRef]
- Rijn, P.; C Mougin, N.; Franke, D.; Park, H.; Böker, A. Pickering emulsion templated soft capsules by self-assembling cross-linkable ferritin-polymer conjugates. Chem. Commun. 2011, 47, 8376–8378. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Wu, L.; Suthiwangcharoen, N.; Bruckman, M.A.; Cash, D.; Hudson, J.S.; Ghoshroy, S.; Wang, Q. Controlled assembly of rodlike viruses with polymers. Chem. Commun. 2009, 20, 2869–2871. [Google Scholar] [CrossRef]
- Zhang, X.; Zhao, X.; Luckanagul, J.A.; Yan, J.; Nie, Y.; Lee, L.A.; Wang, Q. Polymer–Protein Core–Shell Nanoparticles for Enhanced Antigen Immunogenicity. Acs Macro Lett. 2017, 6, 442–446. [Google Scholar] [CrossRef]
- Prendergast, F.G.; Mann, K.G. Chemical and physical properties of aequorin and the green fluorescent protein isolated from Aequorea forskalea. Biochemistry 1978, 17, 3448–3453. [Google Scholar] [CrossRef] [PubMed]
- Shimomura, O.; Johnson Frank, H.; Saiga, Y. Extraction, Purification and Properties of Aequorin, a Bioluminescent Protein from the Luminous Hydromedusan, Aequorea. J. Cell. Comp. Physiol. 1962, 59, 223–239. [Google Scholar] [CrossRef] [PubMed]
- Kneen, M.; Farinas, J.; Li, Y.; Verkman, A.S. Green fluorescent protein as a noninvasive intracellular pH indicator. Biophys. J. 1998, 74, 1591–1599. [Google Scholar] [CrossRef]
- Tsien, R.Y. The Green Fluorescent Protein. Annu. Rev. Biochem. 1998, 67, 509–544. [Google Scholar] [CrossRef] [PubMed]
- Cody, C.W.; Prasher, D.C.; Westler, W.M.; Prendergast, F.G.; Ward, W.W. Chemical structure of the hexapeptide chromophore of the Aequorea green-fluorescent protein. Biochemistry 1993, 32, 1212–1218. [Google Scholar] [CrossRef] [PubMed]
- Chalfie, M.; Tu, Y.; Euskirchen, G.; Ward, W.W.; Prasher, D.C. Green fluorescent protein as a marker for gene expression. Science 1994, 263, 802–805. [Google Scholar] [CrossRef] [PubMed]
- Heim, R.; Prasher, D.C.; Tsien, R.Y. Wavelength mutations and posttranslational autoxidation of green fluorescent protein. Proc. Natl. Acad. Sci. USA 1994, 91, 12501–12504. [Google Scholar] [CrossRef]
- Tamres, M.; Searles, S.; Leighly, E.M.; Mohrman, D.W. Hydrogen Bond Formation with Pyridines and Aliphatic Amines1. J. Am. Chem. Soc. 1954, 76, 3983–3985. [Google Scholar] [CrossRef]
- Li, T.; Zhou, P.; Mattei, A. Electronic origin of pyridinyl N as a better hydrogen-bonding acceptor than carbonyl O. CrystEngComm 2011, 13, 6356–6360. [Google Scholar] [CrossRef]
- Vyhnalkova, R.; Müller, A.H.E.; Eisenberg, A. Control of Morphology and Corona Composition in Aggregates of Mixtures of PS-b-PAA and PS-b-P4VP Diblock Copolymers: Effects of Solvent, Water Content, and Mixture Composition. Langmuir 2014, 30, 13152–13163. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Niu, Z.; Emrick, T.; Russell, T.P.; Wang, Q. Core/Shell Biocomposites from the Hierarchical Assembly of Bionanoparticles and Polymer. Small 2008, 4, 1624–1629. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Jiang, M. Strategies for Constructing Polymeric Micelles and Hollow Spheres in Solution via Specific Intermolecular Interactions. Acc. Chem. Res. 2005, 38, 494–502. [Google Scholar] [CrossRef] [PubMed]
- Nie, L.; Liu, S.; Shen, W.; Chen, D.; Jiang, M. One-Pot Synthesis of Amphiphilic Polymeric Janus Particles and Their Self-Assembly into Supermicelles with a Narrow Size Distribution. Angew. Chem. Int. Ed. 2007, 46, 6321–6324. [Google Scholar] [CrossRef] [PubMed]
- Panganiban, B.; Qiao, B.; Jiang, T.; DelRe, C.; Obadia, M.M.; Nguyen, T.D.; Smith, A.A.A.; Hall, A.; Sit, I.; Crosby, M.G.; et al. Random heteropolymers preserve protein function in foreign environments. Science 2018, 359, 1239–1243. [Google Scholar] [CrossRef] [PubMed]
- Page Faulk, W.; Malcolm Taylor, G. Communication to the editors: An immunocolloid method for the electron microscope. Immunochemistry 1971, 8, 1081–1083. [Google Scholar] [CrossRef]
- Horisberger, M.; Rosset, J. Colloidal gold, a useful marker for transmission and scanning electron microscopy. J. Histochem. Cytochem. 1977, 25, 295–305. [Google Scholar] [CrossRef]
- Kübel, C.; Voigt, A.; Schoenmakers, R.; Otten, M.; Su, D.; Lee, T.-C.; Carlsson, A.; Bradley, J. Recent Advances in Electron Tomography: TEM and HAADF-STEM Tomography for Materials Science and Semiconductor Applications. Microsc. Microanal. 2005, 11, 378–400. [Google Scholar] [CrossRef]
- Bals, S.; Van Tendeloo, G.; Kisielowski, C. A New Approach for Electron Tomography: Annular Dark-Field Transmission Electron Microscopy. Adv. Mater. 2006, 18, 892–895. [Google Scholar] [CrossRef]
- Morise, H.; Shimomura, O.; Johnson, F.H.; Winant, J. Intermolecular energy transfer in the bioluminescent system of Aequorea. Biochemistry 1974, 13, 2656–2662. [Google Scholar] [CrossRef]
- Kutrowska, B.W.; Narczyk, M.; Buszko, A.; Bzowska, A.; Clark, P.L. Folding and unfolding of a non-fluorescent mutant of green fluorescent protein. J. Phys. Condens. Matter Inst. Phys. J. 2007, 28, 285223. [Google Scholar] [CrossRef] [PubMed]
- Markova, S.V.; Burakova, L.P.; Frank, L.A.; Golz, S.; Korostileva, K.A.; Vysotski, E.S. Green-fluorescent protein from the bioluminescent jellyfish Clytia gregaria: cDNA cloning, expression, and characterization of novel recombinant protein. Photochem. Photobiol. Sci. 2010, 9, 757–765. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Hu, J.; Li, M.; Wang, W.; Liu, Y.; Winans, R.; Li, T.; Liu, T.; Yin, P. Hydrogen Bonding Directed Co-Assembly of Polyoxometalates and Polymer to Core-Shell Nanoparticles. Mater. Chem. Front. 2018, 2, 2070–2075. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sarnello, E.; Liu, Y.; Palen, B.; Sun, E.; Zuo, X.; Xu, T.; Li, T. Synthesis and Characterization of Bio-Active GFP-P4VP Core–Shell Nanoparticles. Catalysts 2020, 10, 627. https://doi.org/10.3390/catal10060627
Sarnello E, Liu Y, Palen B, Sun E, Zuo X, Xu T, Li T. Synthesis and Characterization of Bio-Active GFP-P4VP Core–Shell Nanoparticles. Catalysts. 2020; 10(6):627. https://doi.org/10.3390/catal10060627
Chicago/Turabian StyleSarnello, Erik, Yuzi Liu, Bethany Palen, Elaine Sun, Xiaobing Zuo, Tao Xu, and Tao Li. 2020. "Synthesis and Characterization of Bio-Active GFP-P4VP Core–Shell Nanoparticles" Catalysts 10, no. 6: 627. https://doi.org/10.3390/catal10060627
APA StyleSarnello, E., Liu, Y., Palen, B., Sun, E., Zuo, X., Xu, T., & Li, T. (2020). Synthesis and Characterization of Bio-Active GFP-P4VP Core–Shell Nanoparticles. Catalysts, 10(6), 627. https://doi.org/10.3390/catal10060627







