XPS Studies of the Initial Oxidation of Polycrystalline Rh Surface
Abstract
1. Introduction
2. Results and Discussion
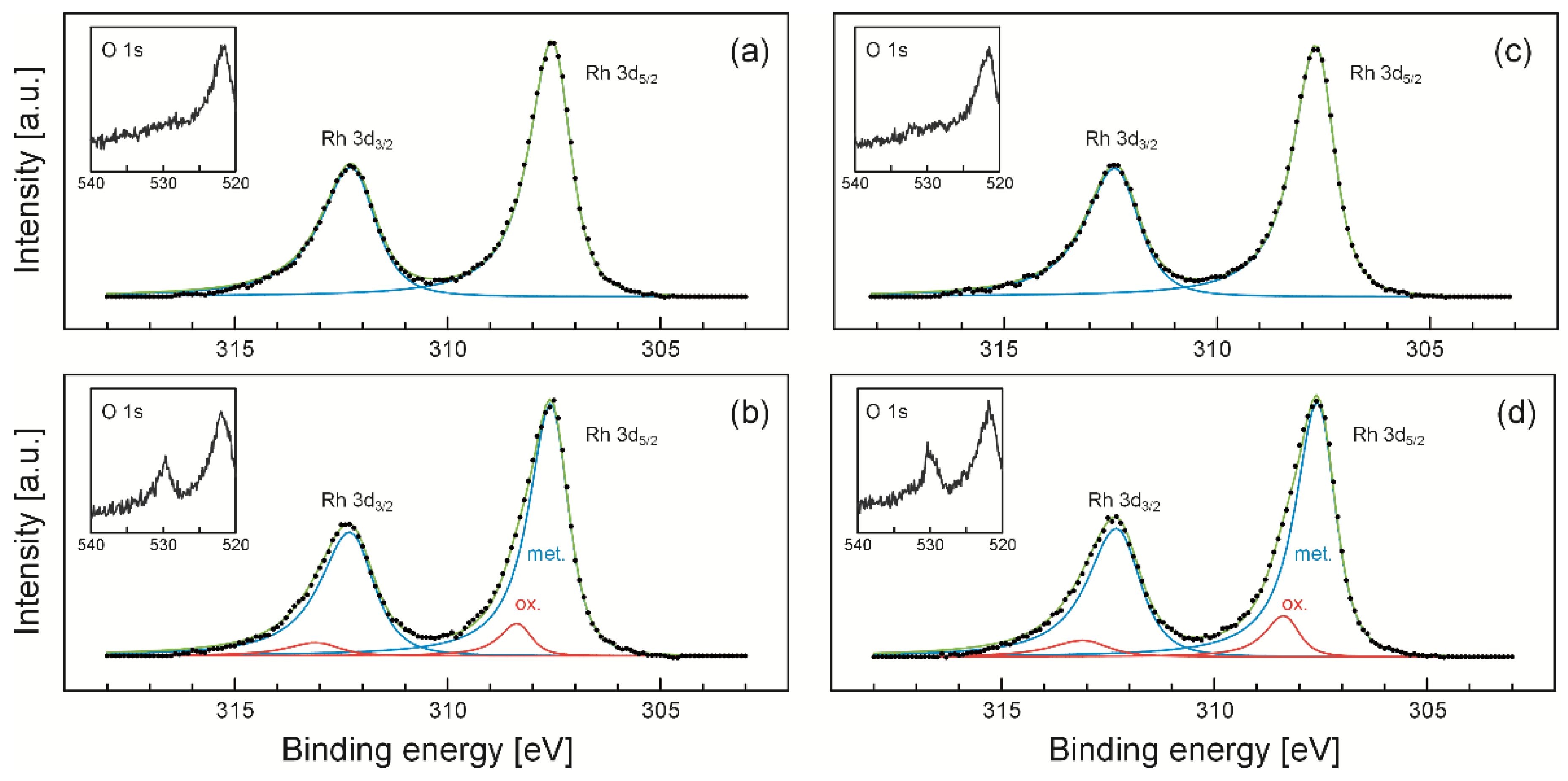
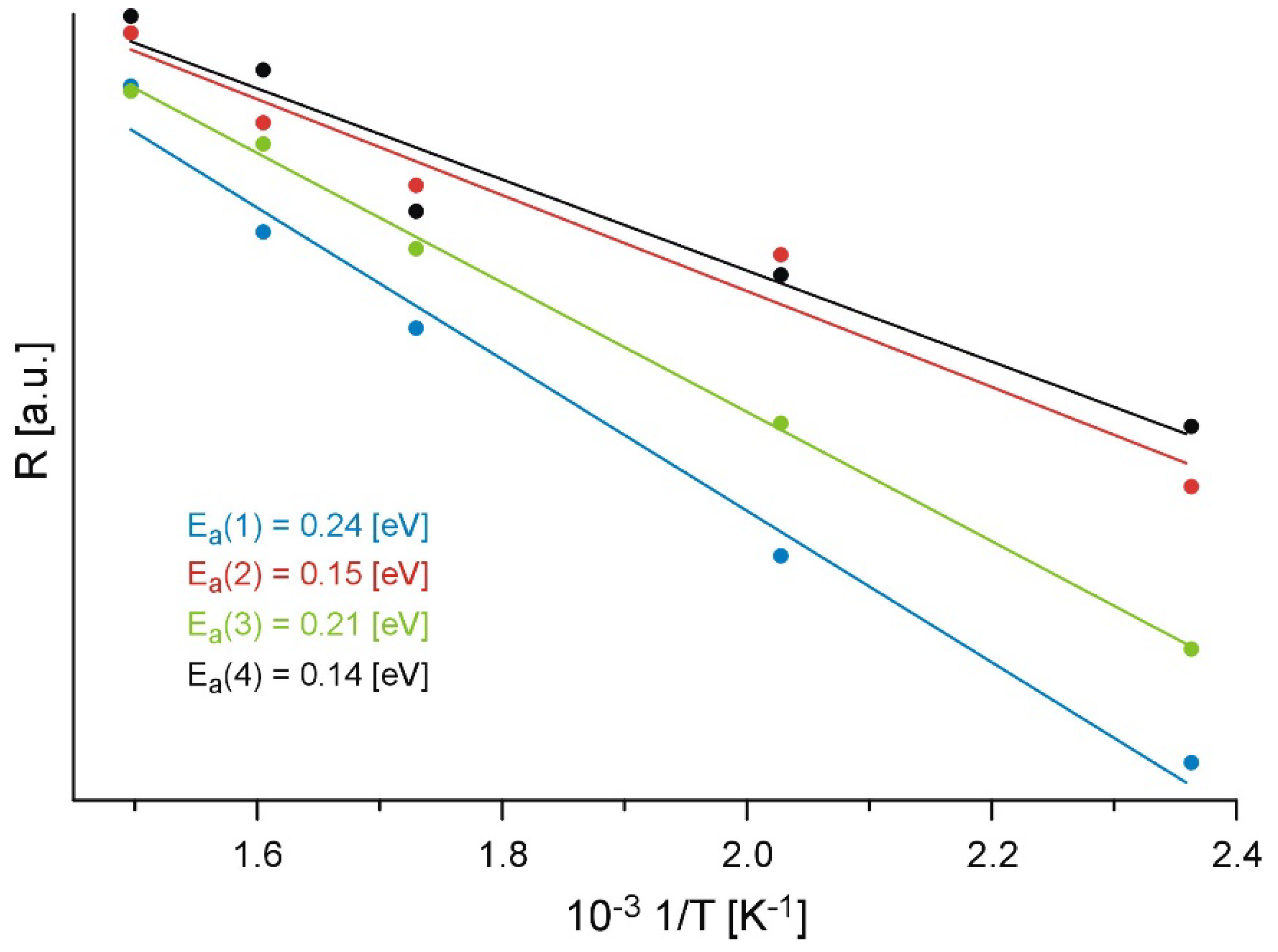
2.1. X-ray Photoelectron Spectroscopy (XPS) Results
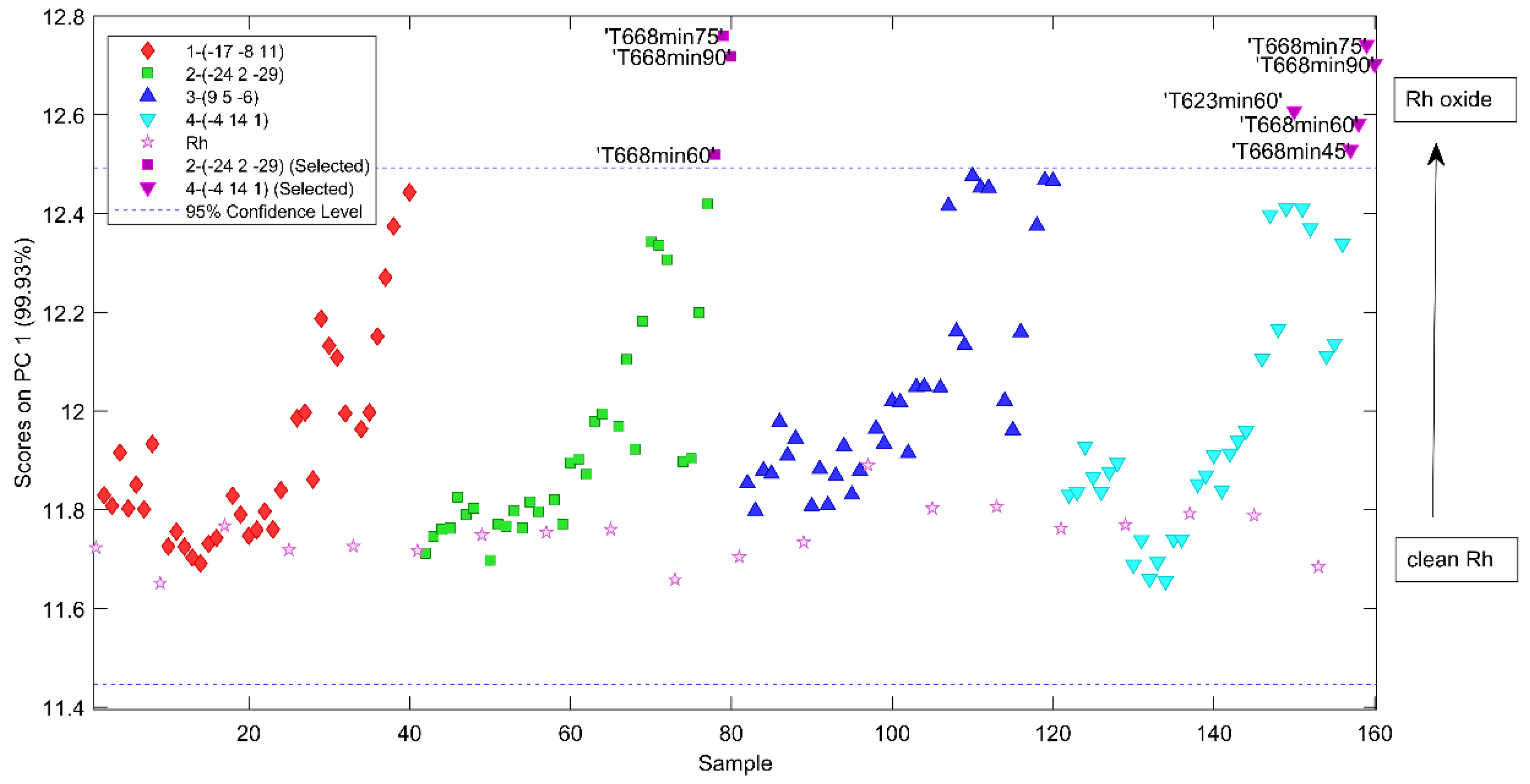
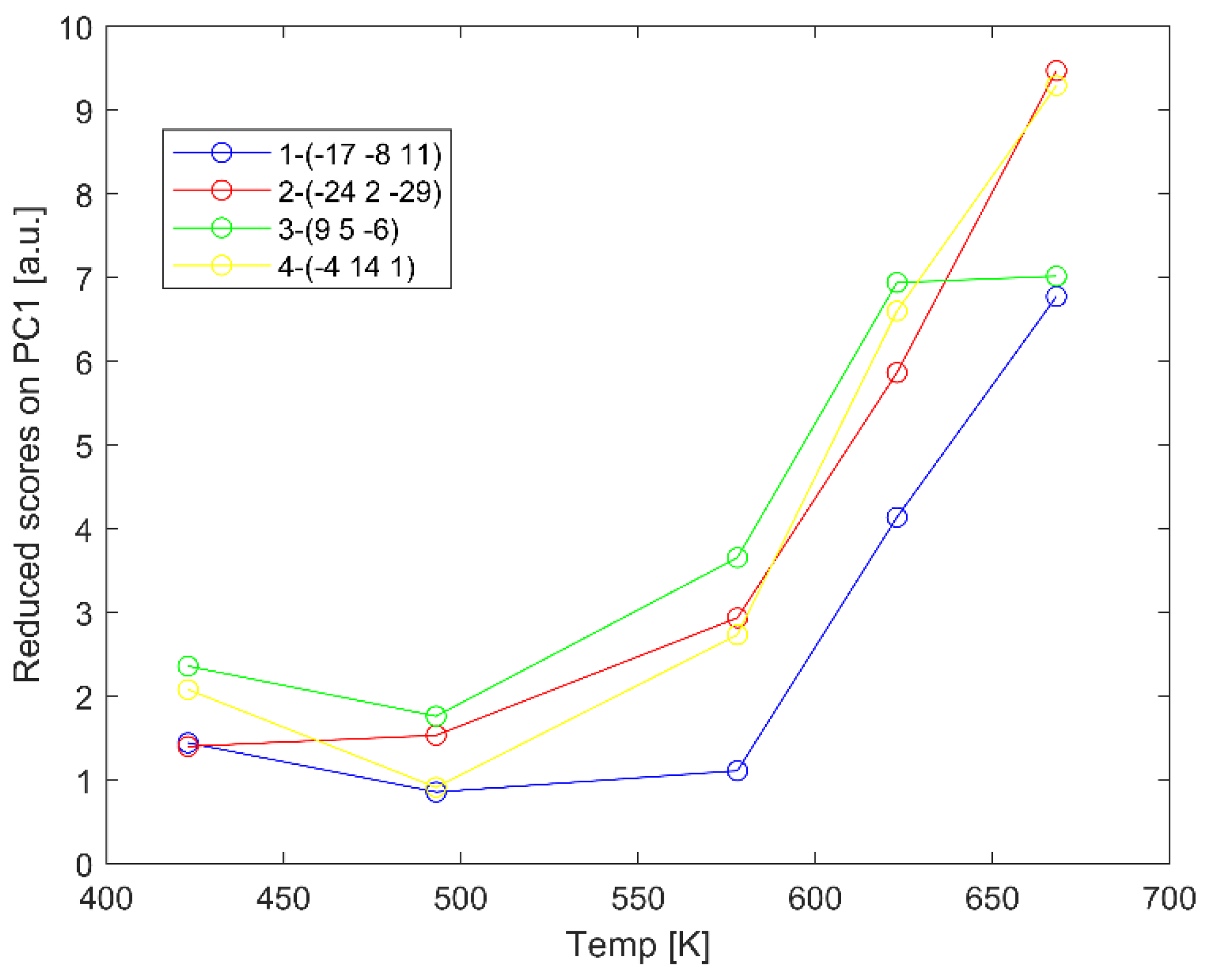
2.2. Diagnostics of Data Collected from Various Rh Crystallites by Principal Component Analysis
3. Materials and Methods
3.1. Polycrystalline Rhodium Sample
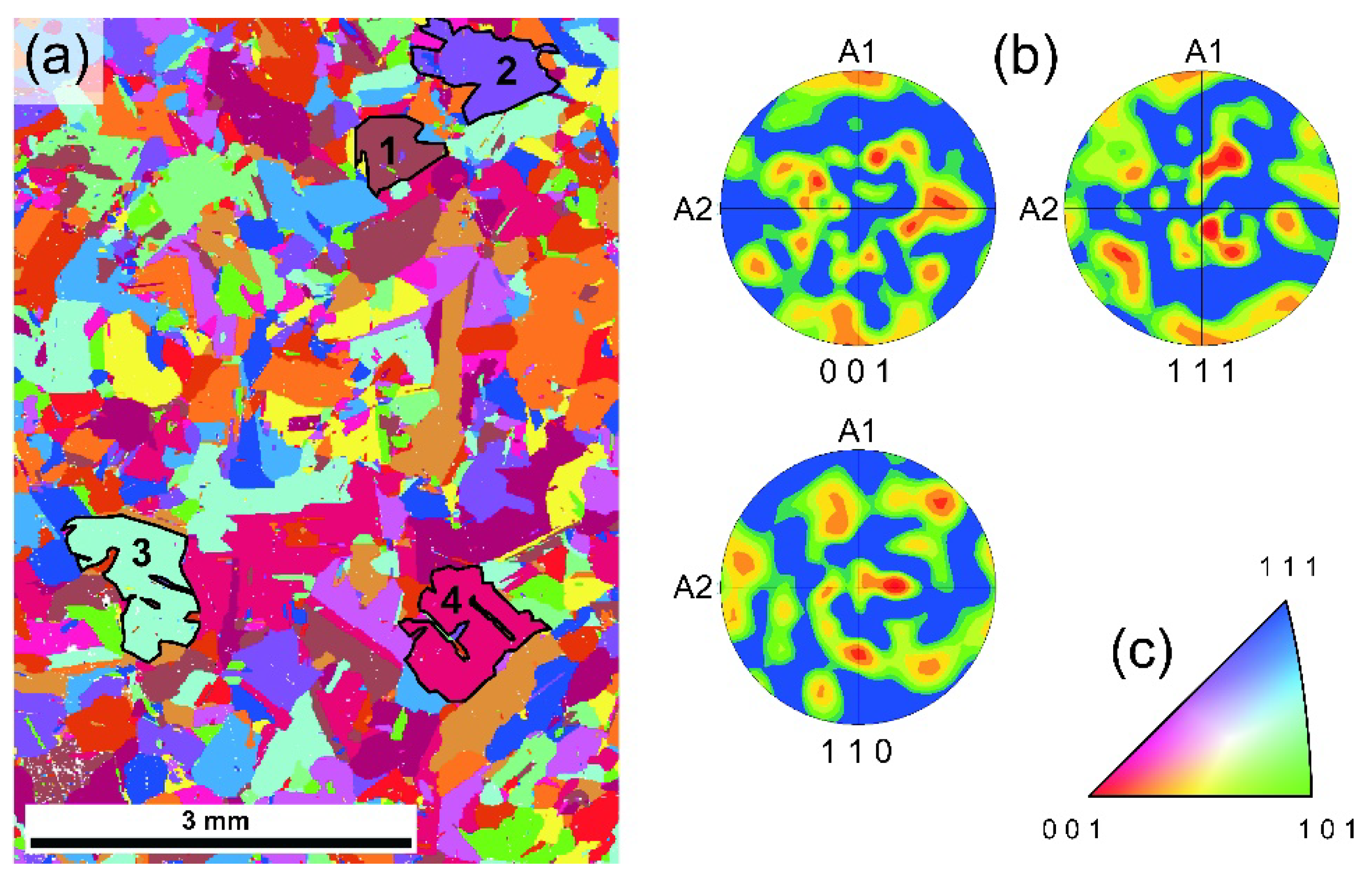
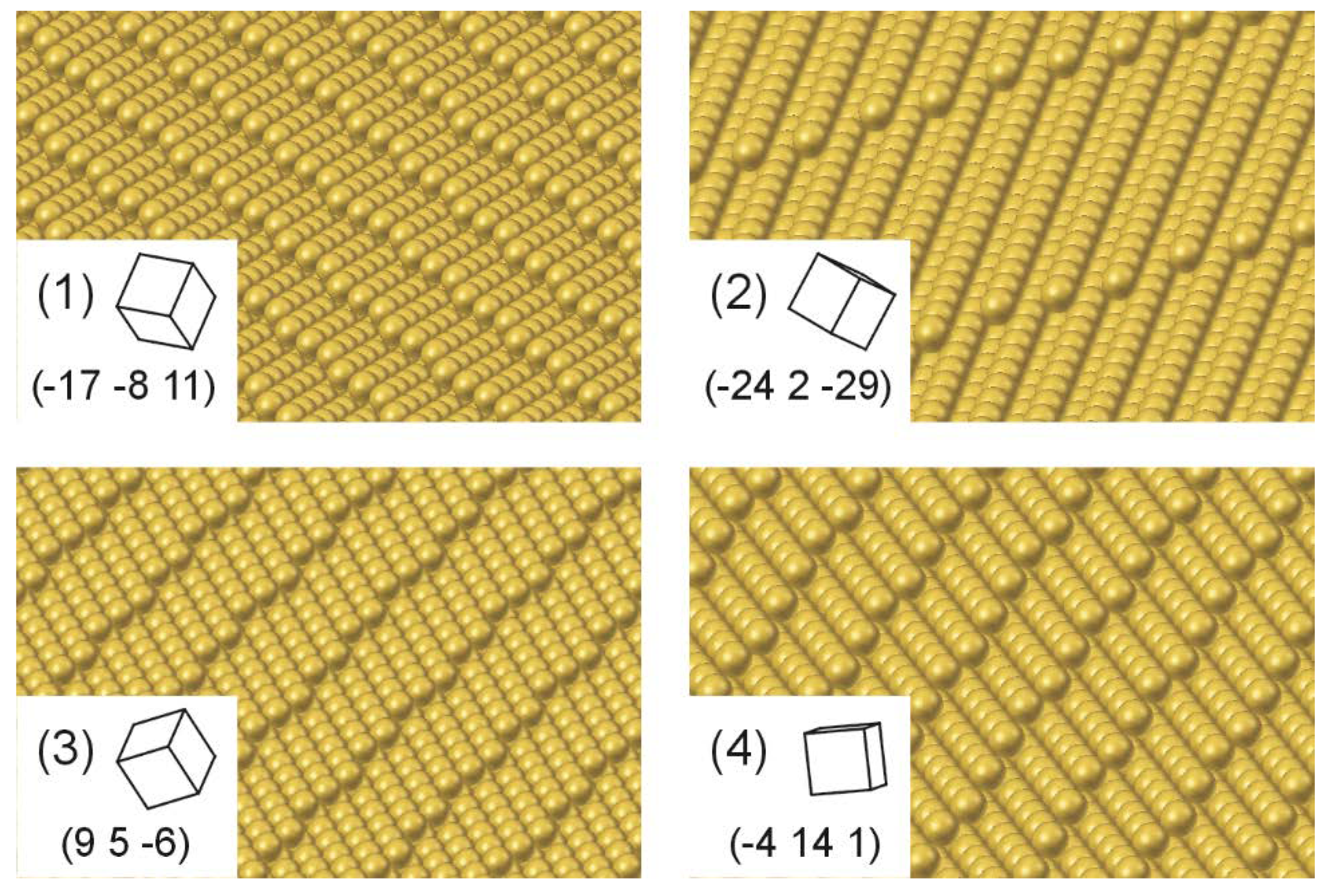
3.2. Crystallographic Orientation of Rh Foil Domains
3.3. Spatially-Resolved X-ray Photoelectron Spectroscopy (XPS) Analysis
3.4. Statistical Multivariate Analysis to Examine XPS Data for Chosen Rh Crystallites
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ertl, G.; Knözinger, H.; Weitkamp, J.; Schüth, F. Handbook of Heterogenous Catalysis; Wiley: Hoboken, NJ, USA, 2008. [Google Scholar]
- Ertl, G. Reactions at Solid Surfaces; Wiley: Hoboken, NJ, USA, 2009. [Google Scholar]
- Ertl, G. Nobel Lecture. Angew. Chem. Int. Ed. 2008, 47, 3524. [Google Scholar] [CrossRef] [PubMed]
- Chaston, J.C. The oxidation of the Platinum Metals. Platin. Met. Rev. 1975, 19, 135–140. [Google Scholar]
- Castner, D.; Sexton, B.; Somorjai, G. LEED and thermal desorption studies of small molecules (H2, O2, CO, CO2, NO, C2H4, C2H2 and C) chemisorbed on the rhodium (111) and (100) surfaces. Surf. Sci. 1978, 71, 519–540. [Google Scholar] [CrossRef]
- Derouin, J.; Farber, R.G.; Killelea, D.R. Combined STM and TPD Study of Rh(111) Under Conditions of High Oxygen Coverage. J. Phys. Chem. C 2015, 119, 14748–14755. [Google Scholar] [CrossRef]
- Castner, D.; Somorjai, G. LEED, AES and thermal desorption studies of the oxidation of the rhodium(111) surface. Appl. Surf. Sci. 1980, 6, 29–38. [Google Scholar] [CrossRef]
- Thiel, P.A.; Yates, J.T.; Weinberg, W.H. The interaction of oxygen with the Rh(111) surface. Surf. Sci. 1979, 82, 22–44. [Google Scholar] [CrossRef]
- Gibson, K.D.; Viste, M.; Sanchez, E.C.; Sibener, S.J. High density adsorbed oxygen on Rh(111) and enhanced routes to metallic oxidation using atomic oxygen. J. Chem. Phys. 1999, 110, 2757–2760. [Google Scholar] [CrossRef]
- Smirnov, M.Y.; Vovk, E.I.; Kalikin, A.V.; Pashis, A.V.; Bukhtiyarov, V.I. An XPS study of the oxidation of noble metal particles evaporated onto the surface of an oxide support in their reaction with NOx. Kinet. Catal. 2012, 53, 117–124. [Google Scholar] [CrossRef]
- Kim, S.M.; Qadir, K.; Seo, B.; Jeong, H.Y.; Joo, S.H.; Terasaki, O.; Park, J.Y. Nature of Rh Oxide on Rh Nanoparticles and Its Effect on the Catalytic Activity of CO Oxidation. Catal. Lett. 2013, 143, 1153–1161. [Google Scholar] [CrossRef]
- Gustafson, J.; Balmes, O.; Zhang, C.; Shipilin, M.; Schaefer, A.; Hagman, B.; Merte, L.R.; Martin, N.M.; Carlsson, P.A.; Jankowski, M.; et al. The Role of Oxides in Catalytic CO Oxidation over Rhodium and Palladium. ACS Catal. 2018, 8, 4438–4445. [Google Scholar] [CrossRef]
- Suchorski, Y.; Kozlov, M.; Bespalov, I.; Datler, M.; Vogel, D.; Budinska, Z.; Neyman, K.M.; Rupprechter, G. The role of metal/oxide interfaces for long-range metal particle activation during CO oxidation. Nat. Mater. 2018, 17, 519–522. [Google Scholar] [CrossRef] [PubMed]
- Berlowitz, P.J.; Peden, C.H.; Goodman, D.W. Kinetics of carbon monoxide oxidation on single-crystal palladium, platinum, and iridium. J. Phys. Chem. 1988, 92, 5213–5221. [Google Scholar] [CrossRef]
- Peden, C.H.; Goodman, D.W.; Blair, D.S.; Berlowitz, P.J.; Fisher, G.B.; Oh, S.H. Kinetics of carbon monoxide oxidation by oxygen or nitric oxide on rhodium(111) and rhodium(100) single crystals. J. Phys. Chem. 1988, 92, 1563–1567. [Google Scholar] [CrossRef]
- Gelius, U.; Wannberg, B.; Baltzer, P.; Fellner-Feldegg, H.; Carlsson, G.; Johansson, C.G.; Larsson, J.; Münger, P.; Vegerfors, G. A new ESCA instrument with improved surface sensitivity, fast imaging properties and excellent energy resolution. J. Electron. Spectrosc. 1990, 52, 747–785. [Google Scholar] [CrossRef]
- Todorova, M.; Li, W.X.; Baltzer, P.; Ganduglia-Pirovano, M.V.; Stampfl, C.; Reuter, K.; Scheffler, M. Role of Subsurface Oxygen in Oxide Formation at Transition Metal Surfaces. Phys. Rev. Lett. 2002, 89, 96103. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, A.J.; Kumar, M.; Adams, B.L.; Field, D.P. Electron Backscatter Diffraction in Materials Science; Springer: Boston, MA, USA, 2009. [Google Scholar]
- Geladi, P. Chemometrics in spectroscopy. Part 1. Classical chemometrics. Spectrochim. Acta B 2003, 58, 767–782. [Google Scholar] [CrossRef]
- Jolliffe, I.T. Principal Component Analysis; Springer: New York, NY, USA, 2002. [Google Scholar]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Trzcinski, M.; Balcerowska-Czerniak, G.; Bukaluk, A. XPS Studies of the Initial Oxidation of Polycrystalline Rh Surface. Catalysts 2020, 10, 617. https://doi.org/10.3390/catal10060617
Trzcinski M, Balcerowska-Czerniak G, Bukaluk A. XPS Studies of the Initial Oxidation of Polycrystalline Rh Surface. Catalysts. 2020; 10(6):617. https://doi.org/10.3390/catal10060617
Chicago/Turabian StyleTrzcinski, Marek, Grażyna Balcerowska-Czerniak, and Antoni Bukaluk. 2020. "XPS Studies of the Initial Oxidation of Polycrystalline Rh Surface" Catalysts 10, no. 6: 617. https://doi.org/10.3390/catal10060617
APA StyleTrzcinski, M., Balcerowska-Czerniak, G., & Bukaluk, A. (2020). XPS Studies of the Initial Oxidation of Polycrystalline Rh Surface. Catalysts, 10(6), 617. https://doi.org/10.3390/catal10060617




