PIRE: Interoperable Platform for Electronic Records
Abstract
1. Introduction
2. Related Work
3. Colombian Regulatory and Technical Context
4. Materials and Methods
4.1. Materials
4.1.1. Technology Stack
4.1.2. HAPI FHIR R4 Server
4.1.3. AWS EC2 Configuration
4.1.4. Integration with the Universidad Militar Nueva Granada
4.1.5. Biomedical Sensing Device
4.2. Methods
4.2.1. Phase 1: Regulatory and Normative Foundation
4.2.2. Phase 2: Semantic Data Modeling
4.2.3. Phase 3: Architectural Design and Implementation
4.2.4. Phase 4: Security, Governance, and User-Centered Design
4.2.5. Validation Approach
5. Development
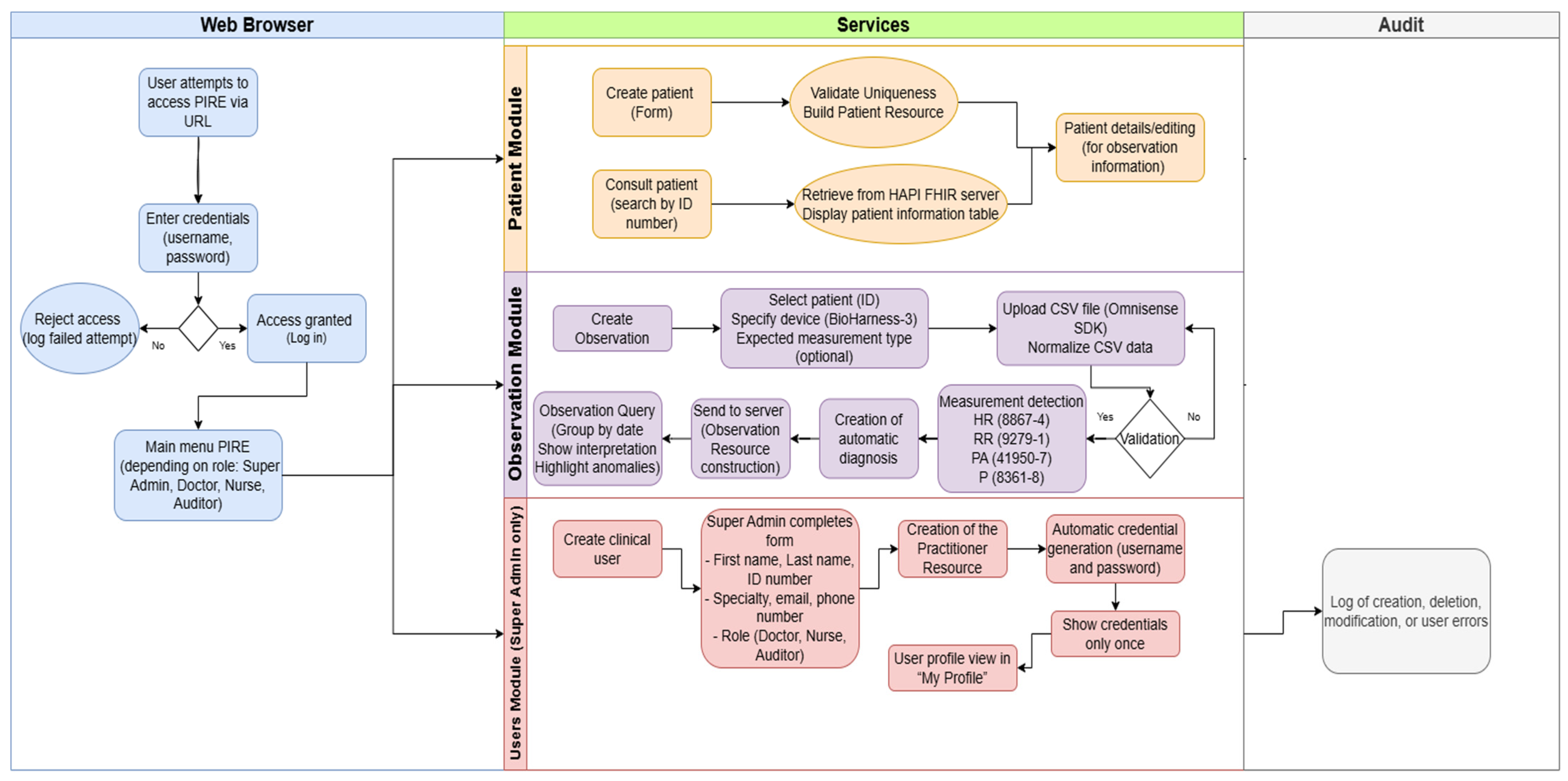
5.1. Operational Flow
5.2. Technical Architecture of the Platform
5.2.1. General System Topology
5.2.2. Functional Components and Modules
5.3. Authentication, Authorization, and Access Control System
5.3.1. Model of Roles Implemented
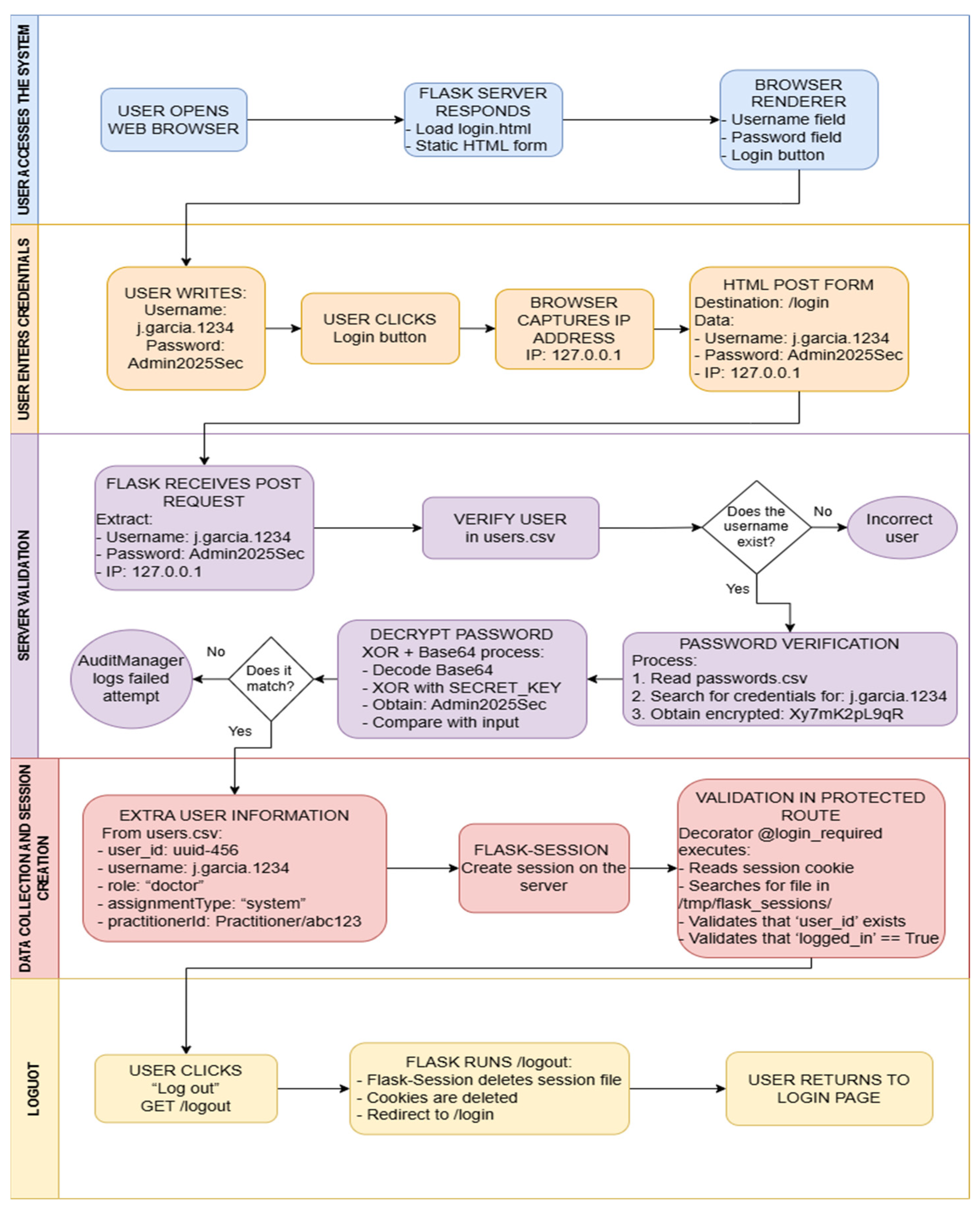
5.3.2. Authentication Process and Session Management
5.3.3. Authorization Mechanisms and Permission Matrix
5.3.4. Credential Security and Automatic User Generation
5.3.5. Auditing Authentication and Access Events
5.4. Patient Management and Resource Generation FHIR Patient
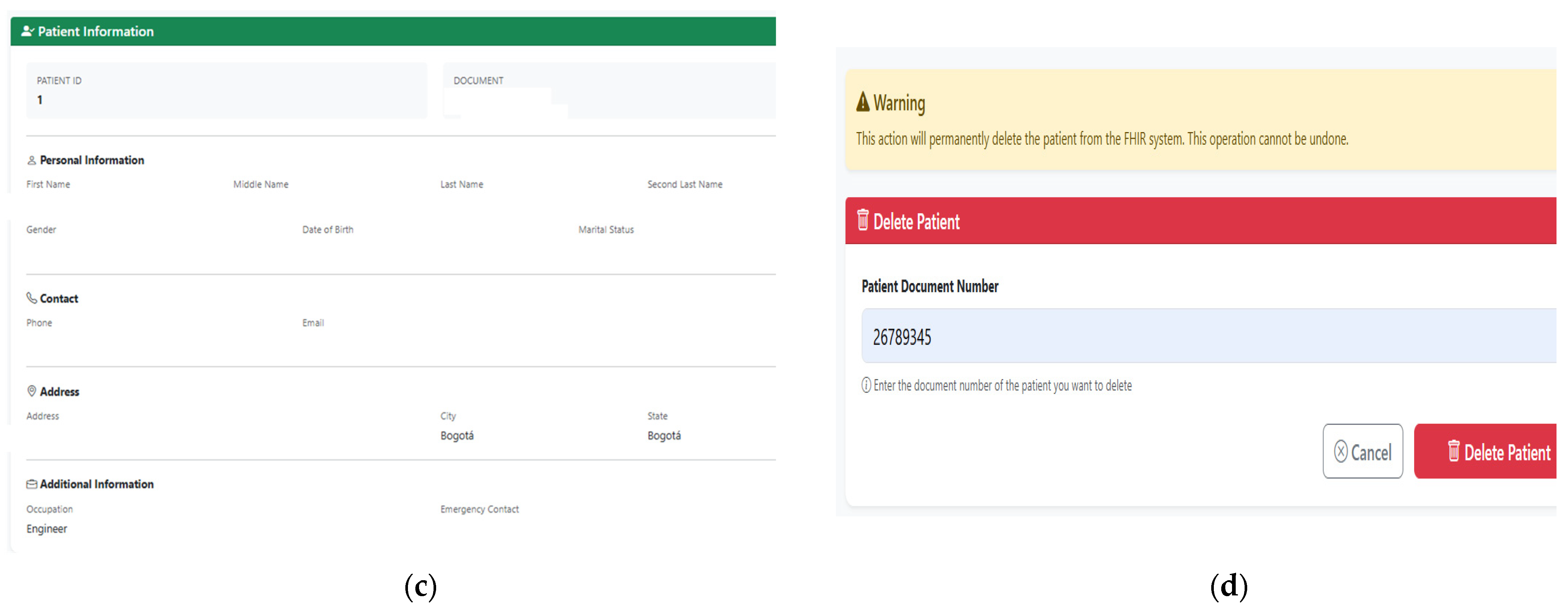
5.4.1. Processes for Creating, Reading, Updating, and Deleting Patients (CRUD)
5.4.2. FHIR Patient Resource Structure
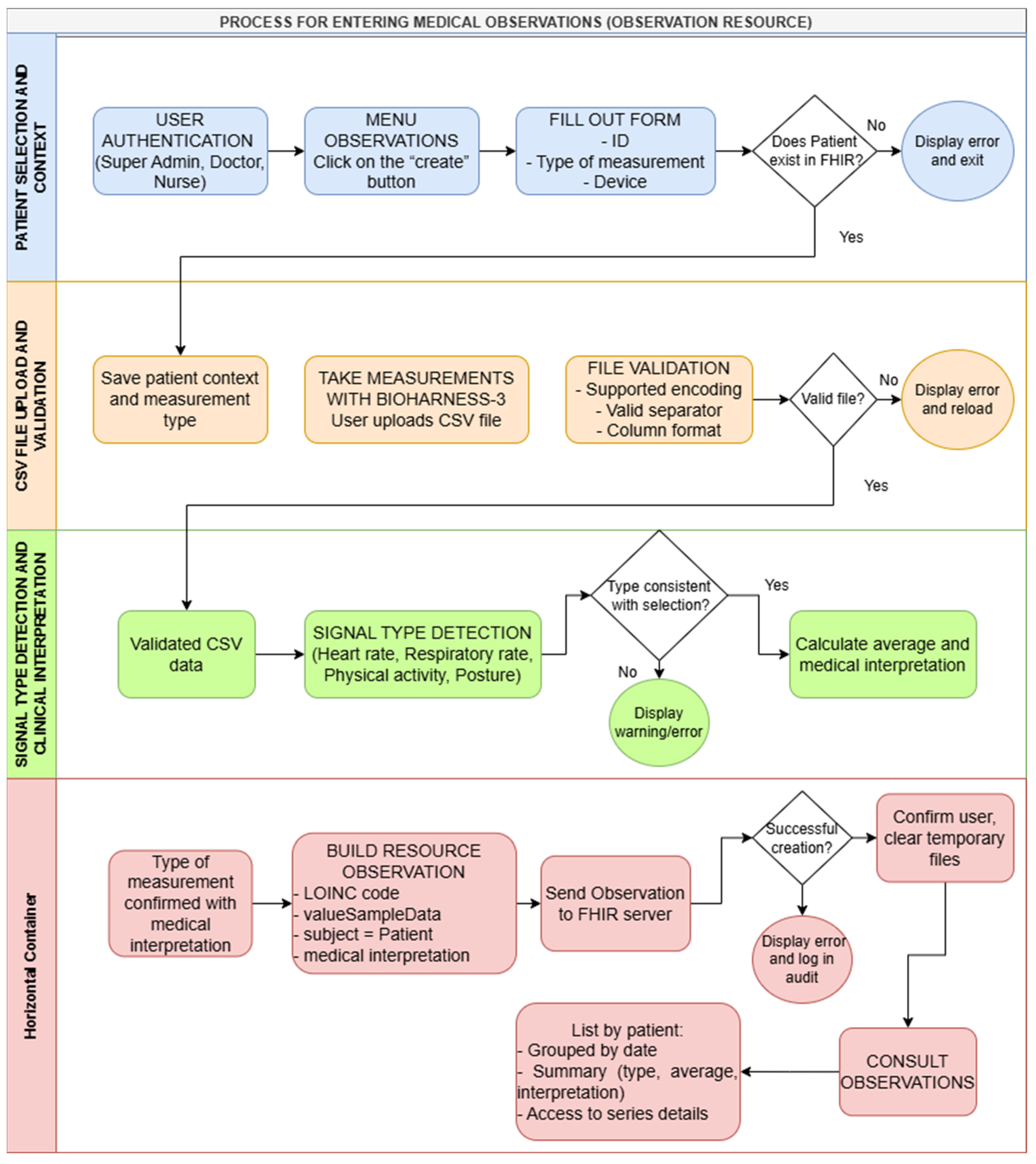
5.5. Biomedical Observation Taking and Resource Generation FHIR Observation
5.5.1. CSV File Upload and Data Validation Process
5.5.2. Automatic Detection of Measurement Type
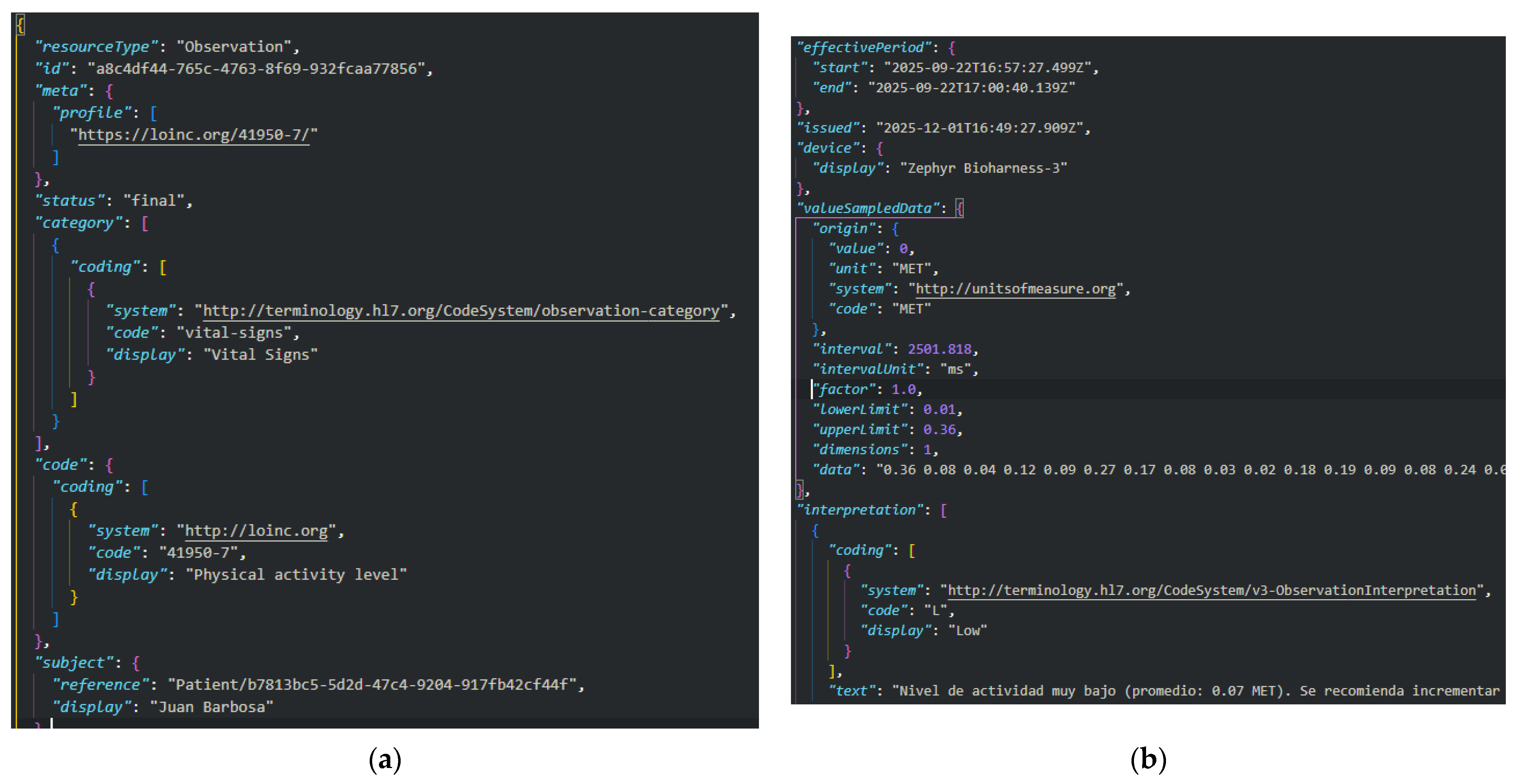
5.5.3. Construction of the FHIR Observation Resource
5.5.4. Clinical Interpretation Calculation and Use of Ranges
5.5.5. Error Handling
5.6. FHIR Practitioner Health Professional and Resource Management
5.6.1. Creation of the Practitioner Resource
5.6.2. Integration with the User System
5.7. System Auditing and Logging
5.7.1. Logging of Critical Actions
5.7.2. Transaction Traceability and Application Logging
6. Results
6.1. Function Validation
6.1.1. Patient Management and FHIR Patient Resource Generation
6.1.2. Biomedical Observation Management
6.1.3. User Management and Practitioner Resource Generation
6.1.4. Audit and Logging System
6.2. Performance Evaluation
6.2.1. End-to-End Latency
6.2.2. Concurrency Behavior
6.2.3. Data Integrity of the CSV-to-FHIR Transformation
6.2.4. FHIR Semantic Conformance Validation
7. Discussion
7.1. Convergence with Historical Evolution and the Colombian Regulatory Framework
7.2. Technical Architecture and Biosignal Management
7.3. Security Models and Implementation Lessons
7.4. Limitations and Operational Responsibilities
8. Future Work
8.1. Advanced Analytics and Data Exploitation
8.2. Enhanced Security and Governance
8.3. Usability and Data Transformation
9. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| ABAC | Attribute-Based Access Control |
| API | Application Programming Interface |
| AWS | Amazon Web Services |
| CDA | Clinical Document Architecture |
| CIE-10 | Clasificación Internacional de Enfermedades, 10a Edición (International Classification of Diseases, 10th Edition) |
| CRUD | Create, Read, Update, Delete |
| CSV | Comma-Separated Values |
| EHDS | European Health Data Space |
| ELGA | Elektronische Gesundheitsakte |
| ENHIS | European Network of Health Information Systems |
| ESP32 | Microcontroller Unit |
| ETL | Extract, Transform Load |
| FHIR | Fast Healthcare Interoperability Resources |
| FHIR CORE CO | FHIR Core Colombia Implementation Guide |
| FHIR-Former | FHIR-bases Transformer with Large Language Models |
| FHIRPath | FHIR Path Navigation Syntax |
| FML | FHIR Mapping Language |
| Flask | Flask Web Framework |
| Flask-Session | Flask Session Management Extension |
| FL | Federated Learning |
| GIT | Version Control System |
| HABEAS DATA | Constitutional Right to Data Protection |
| HAPI | Healthcare API |
| HAPI FHIR | HAPI FHIR server |
| HL7 | Health Level 7 |
| HL7 CDA | HL7 Clinical Document Architecture |
| HL7 FHIR R4 | HL7 FHIR Release 4 |
| HTML | Hyper Text Markup Language |
| HTTP | Hyper Text Transfer Protocol |
| HTTPS | HTTP Secure |
| IHCE | Interoperable Electronic Health Record (Colombia) |
| IP | Internet Protocol |
| IoMT | Internet of Medical Things |
| IoT | Internet of Things |
| JSON | JavaScript Object Notation |
| JPA | Java Persistence API |
| LOINC | Logical Observation Identifiers Names and Codes |
| LLM | Large Language Model |
| Maven | Apache Maven |
| ONC | Office of the National Coordination for Health Information Technology |
| OpenEHR | Open Electronic Health Record |
| Omnisense | Zephyr Omnisense SDK |
| Portable Document Format | |
| PGHD | Patient-Generated Health Data |
| PIRE | Plataforma Interoperable de Registros Electrónicos |
| POMR | Problem-Oriented Medical Record |
| PPG | Photoplenthysmography |
| RBAC | Role-Based Access Control |
| RDA | Resumen Digital de Atención (Digital Care Summary) |
| REDCap | Research Electronic Data Capture |
| REST | Representational State Transfer |
| RMRS | Regenstrief Medical Record System |
| SDK | Software Development Kit |
| SFPBRF | SMART on FHIR Platform-Based Reference Framework |
| SMART | Substitutable Medical Applications and Reusable Technology |
| SNOMED CT | Systematized Nomenclature of Medicine Clinical Terms |
| SSH | Secure Shell |
| SQL | Structured Query Language |
| SQL Server | Microsoft SQL Server |
| SURA | Seguros Unidos de Reaseguro Americano |
| TermX | Terminology Exchange Tool |
| Ubuntu | Linux Operating System Distribution |
| VPC | Virtual Private Cloud |
| WHO | World Health Organization |
| XOR | Exclusive OR |
| Zephyr | Zephyr Performance System |
References
- Shen, Y.; Yu, J.; Zhou, J.; Hu, G. Twenty-Five Years of Evolution and Hurdles in Electronic Health Records and Interoperability in Medical Research: Comprehensive Review. J. Med. Internet Res. 2025, 27, e59024. [Google Scholar] [CrossRef] [PubMed]
- Finnegan, H.; Mountford, N. 25 Years of Electronic Health Record Implementation Processes: Scoping Review. J. Med. Internet Res. 2025, 27, e60077. [Google Scholar] [CrossRef] [PubMed]
- El-Yafouri, R.; Klieb, L. A scoping review of electronic health records interoperability levels, expectations, approaches, and problems. Health Inform. J. 2025, 31, 14604582251385986. [Google Scholar] [CrossRef]
- Ministerio de Salud y Protección Social. de Colombia Ley 2015 de 2020; Ministerio de Salud y Protección Social: Bogota, Colombia, 2020.
- Ministerio de Salud y Protección Social. de Colombia Resolución 866 de 2021; Ministerio de Salud y Protección Social: Bogota, Colombia, 2021.
- Ministerio de Salud y Protección Social. de Colombia Resolución 1888 de 2025; Ministerio de Salud y Protección Social: Bogota, Colombia, 2025.
- Castaño, C.D.A.; Cardenas, J.F.Z. Interoperability Project Brings Better Healthcare; HL7 International: Ann Arbor, MI, USA, 2024. [Google Scholar]
- Zephyr Technology. BioHarness 3 User Manual; Zephyr Technology Corp.: Annapolis, MD, USA, 2012. [Google Scholar]
- Johnstone, J.A.; Ford, P.A.; Hughes, G.; Watson, T.; Garrett, A.T. Bioharness™ multivariable monitoring device part I: Validity. J. Sports Sci. Med. 2012, 11, 400–408. [Google Scholar]
- Duftschmid, G.; Katsch, F.; Ciortuz, G.; Kalra, D.; Rinner, C. Reusing data from HL7 CDA-based shared EHR systems for clinical trial conduct: A method for analyzing feasibility. BMC Med. Inform. Decis. Mak. 2025, 25, 155. [Google Scholar] [CrossRef] [PubMed]
- Palojoki, S.; Lehtonen, L.; Vuokko, R. Semantic Interoperability of Electronic Health Records: Systematic Review of Alternative Approaches for Enhancing Patient Information Availability. JMIR Med. Inform. 2024, 12, e53535. [Google Scholar] [CrossRef]
- Bossenko, I.; Randmaa, R.; Piho, G.; Ross, P. Interoperability of health data using FHIR Mapping Language: Transforming HL7 CDA to FHIR with reusable visual components. Front. Digit. Health 2024, 6, 1480600. [Google Scholar] [CrossRef]
- Gabriel, M.H.; Richwine, C.; Strawley, C.; Barker, W.; Everson, J. Interoperable Exchange of Patient Health Information Among U.S. Hospitals: 2023. Available online: https://www.healthit.gov/data/quickstats/electronic-health-information-exchange-hospitals (accessed on 15 January 2026).
- Adib, K.; Salama, N.; Davia, S.; Martínez-Millana, A.; Traver, V.; Davtyan, K.; Tornero-Costa, R. Electronic health records and data exchange in the WHO European region: A subregional analysis of achievements, challenges, and prospects. Int. J. Med. Inform. 2025, 194, 105687. [Google Scholar] [CrossRef]
- Weistra, W. The State of FHIR in 2025: Growing Adoption and Evolving Maturity. Available online: https://fire.ly/blog/the-state-of-fhir-in-2025/ (accessed on 15 January 2026).
- Hornback, A.; Marteau, B.; Tan, S.Q.Y.; Kim, K.; Patil, O.; Traynelis, J.; Zhu, Y.; Giuste, F.; Wang, M.D. FHIR in Focus: Enabling Biomedical Data Harmonization for Intelligent Healthcare Systems. IEEE Rev. Biomed. Eng. 2025, 19, 305–336. [Google Scholar] [CrossRef]
- Ahmad, A.; Azam, F.; Anwar, M.W. Implementation of SMART on FHIR in developing countries through SFPBRF. In ICBBE ’18: Proceedings of the 2018 5th International Conference on Biomedical and Bioinformatics Engineering; ACM International Conference Proceeding Series; Association for Computing Machinery: New York, NY, USA, 2018; pp. 137–144. [Google Scholar] [CrossRef]
- Cheng, A.C.; Shyr, C.; Lewis, A.; Delacqua, F.; Bosler, T.; Banasiewicz, M.K.; Taylor, R.; Lindsell, C.J.; Harris, P.A. Opportunities, barriers, and remedies for implementing REDCap integration with electronic health records via Fast Healthcare Interoperability Resources (FHIR). JAMIA Open 2025, 8, ooaf111. [Google Scholar] [CrossRef]
- Khvastova, M.; Witt, M.; Essenwanger, A.; Sass, J.; Thun, S.; Krefting, D. Towards Interoperability in Clinical Research—Enabling FHIR on the Open-Source Research Platform XNAT. J. Med. Syst. 2020, 44, 137. [Google Scholar] [CrossRef]
- Kruse, J.; Wiedekopf, J.; Kock-Schoppenhauer, A.K.; Essenwanger, A.; Ingenerf, J.; Ulrich, H. A Generic Transformation Approach for Complex Laboratory Data Using the Fast Healthcare Interoperability Resources Mapping Language: Method Development and Implementation. JMIR Med. Inform. 2024, 12, e57569. [Google Scholar] [CrossRef]
- Engelke, M.; Baldini, G.; Kleesiek, J.; Nensa, F.; Dada, A. FHIR-Former: Enhancing clinical predictions through Fast Healthcare Interoperability Resources and large language models. J. Am. Med. Inform. Assoc. 2025, 32, 1793–1801. [Google Scholar] [CrossRef]
- Hou, Z.; Jiang, M.; Liu, H.; Zhuang, Y. LLM-Integrated Normalization and Knowledge for FHIR (LINK-FHIR). In Studies in Health Technology and Informatics; IOS Press BV: Amsterdam, Holland, 2025; pp. 17–21. [Google Scholar] [CrossRef]
- Grimes, J.; Brush, R.; Rhyzhikov, N.; Szul, P.; Mandel, J.; Gottlieb, D.; Grieve, G.; Sadjad, B.; Sanyal, A. SQL on FHIR—Tabular views of FHIR data using FHIRPath. npj Digit. Med. 2025, 8, 342. [Google Scholar] [CrossRef]
- Mukherjee, S.; Ray, I.; Ray, I.; Shirazi, H.; Ong, T.; Kahn, M.G. Attribute based access control for healthcare resources. In ABAC 2017—Proceedings of the 2nd ACM Workshop on Attribute-Based Access Control, Co-Located with CODASPY 2017; Association for Computing Machinery, Inc.: New York, NY, USA, 2017; pp. 29–40. [Google Scholar] [CrossRef]
- Martínez, A.L.; Naghmouchi, M.; Laurent, M.; Alfaro, J.G.; Pérez, M.G.; Martínez, A.R. Breaking barriers in healthcare: A secure identity framework for seamless access. Comput. Stand. Interfaces 2026, 95, 104020. [Google Scholar] [CrossRef]
- Haffer, N.; Stellmach, C.; Sass, J.; Muzoora, M.R.; Graefe, A.S.L.; Thun, S.; Vorisek, C.N. Genomics on FHIR—A feasibility study to support a National Strategy for Genomic Medicine. npj Genom. Med. 2025, 10, 57. [Google Scholar] [CrossRef]
- Schmidt, C.S.; Wutzkowsky, J.; Schäfer, H.; Reinartz, L.; Chahin, H.; Winnekens, P.; Alsulaiman, J.A.; Temme, C.; Lenz, V.; Hosch, R.; et al. A real-time dashboard for optimizing blood product utilization through a FHIR-based data integration pipeline. Transfus. Apher. Sci. 2026, 65, 104301. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.; Algorri, M.; Abernathy, M.J. Real-time algorithmic exchange and processing of pharmaceutical quality data and information. Int. J. Pharm. 2023, 645, 123342. [Google Scholar] [CrossRef] [PubMed]
- Fecho, K.; Garcia, J.J.; Yi, H.; Roupe, G.; Krishnamurthy, A. FHIR PIT: A geospatial and spatiotemporal data integration pipeline to support subject-level clinical research. BMC Med. Inform. Decis. Mak. 2025, 25, 24. [Google Scholar] [CrossRef]
- Akhmetov, A.; Latif, Z.; Tyler, B.; Yazici, A. Enhancing healthcare data privacy and interoperability with federated learning. PeerJ Comput. Sci. 2025, 11, e2870. [Google Scholar] [CrossRef] [PubMed]
- Seixas-Lopes, F.A.; Lopes, C.; Marques, M.; Agostinho, C.; Jardim-Goncalves, R. Musculoskeletal Disorder (MSD) Health Data Collection, Personalized Management and Exchange Using Fast Healthcare Interoperability Resources (FHIR). Sensors 2024, 24, 5175. [Google Scholar] [CrossRef] [PubMed]
- Randine, P.; Årsand, E. FHIR-Based Information System for Type 1 Diabetes Consultations Based on Patient-Generated Health Data. In Studies in Health Technology and Informatics; IOS Press BV: Amsterdam, Holland, 2025; pp. 189–193. [Google Scholar] [CrossRef]
- Urbauer, P.; David, V.; Frohner, M.; Sauermann, S. Wearable activity trackers supporting elderly living independently: A standards based approach for data integration to health information systems. In DSAI ‘18: Proceedings of the 8th International Conference on Software Development and Technologies for Enhancing Accessibility and Fighting Info-Exclusion; ACM International Conference Proceeding Series; Association for Computing Machinery: New York, NY, USA, 2018; pp. 302–309. [Google Scholar] [CrossRef]
- Abedian, S.; Yesakov, E.; Ostrovskiy, S.; Hussein, R. Streamlining wearable data integration for EHDS: A case study on advancing healthcare interoperability using Garmin devices and FHIR. Front. Digit. Health 2025, 7, 1636775. [Google Scholar] [CrossRef] [PubMed]
- Adochiei, F.C.; Țoi, F.A.; Adochiei, I.R.; Argatu, F.C.; Serițan, G.; Petroiu, G.G. HL7 FHIR-Based Open-Source Framework for Real-Time Biomedical Signal Acquisition and IoMT Interoperability. Appl. Sci. 2025, 15, 12803. [Google Scholar] [CrossRef]
- Ministerio de Salud y Protección Social. de Colombia Resumen Digital de Atención en Salud (RDA). Available online: https://vulcano.ihcecol.gov.co/ (accessed on 15 January 2026).
- Consejo Nacional de Política Económica y Social (CONPES). “Política de Salud Digital,” Departamento Nacional de Planeación, Bogotá, Colombia, Doc. CONPES 4070, dic. 2021. Available online: https://colaboracion.dnp.gov.co/CDT/Conpes/Económicos/4070.pdf (accessed on 15 January 2026).
- MEDIFOLIOS. Asistene de Registros Médicos. Available online: https://www.medifolios.net/modulos/integracion-ecopetrol.php (accessed on 15 January 2026).
- MEDIFOLIOS. HCE: Normativas y Beneficios Colombia. Available online: https://medifolios.net/articulos/historia-clinica-normativas-beneficios.php (accessed on 15 January 2026).
- ISO 8601-1:2019; Date and Time—Representations for Information Interchange—Part 1: Basic Rules. International Organization for Standardization (ISO): Geneva, Switzerland, 2019.
- Lee, J.; Jin, J.; Kim, Y. Voting-based real-time monitoring scheme for healthcare data on medical record clouds. In 2024 Research in Adaptive and Convergent Systems—Proceedings of the 2024 International Conference on Research in Adaptive and Convergent Systems; RACS 2024; Association for Computing Machinery, Inc.: New York, NY, USA, 2025; pp. 157–159. [Google Scholar] [CrossRef]
- Vogel, C.; Pryss, R.; Heuschmann, P.; Rücker, V.; Winter, M. Assessing a visual editor for healthcare questionnaires based on the fast healthcare interoperability resources (FHIR) standard: Protocol for a cross-sectional, mixed-methods usability evaluation using eyetracking and retrospective think-aloud. BMJ Open 2025, 15, e099744. [Google Scholar] [CrossRef] [PubMed]







| Layer | Component | Technology | Version |
|---|---|---|---|
| Web application | Framework | Flask (Python) | 2.x |
| Web application | Sessions | Flask-Session | - |
| FHIR server | Implementation | HAPI FHIR JPA Server | 8.2.0 |
| FHIR server | Runtime | OpenJDK | 17.0.15 |
| FHIR server | Build | Maven | 3.x |
| Database | Engine | PostgreSQL | 16.9 |
| Operating system | Distribution | Ubuntu Linux | 24.04.2 LTS |
| Infrastructure | Cloud | AWS EC2 | - |
| Bioharness CSV Fields | Conversion and Normalization Logic | FHIR Element Mapping |
|---|---|---|
| Timestamp | Parsed to standard ISO 8601 [40] format (UTC). The minimum and maximum timestamps define the period. | Observation.effectivePeriod.start Observation.effectivePeriod.end |
| Time Differences | Calculates the average time elapsed between consecutive samples in milliseconds (ms). | Observation.valueSampledData.interval Observation.valueSampledData.intervalUnit = “ms” |
| HR_BPM | Filters valid physiological ranges (30–200 bpm). Converts the filtered numeric array into a single space-separated string. | Observation.code.coding.code = “8867-4” Observation.code.coding.display = “Heart rate” Observation.valueSampledData.data = [string] Observation.valueSampledData.origin.unit = “beats/min” |
| BR_BPM | Filters valid ranges (2–40 breaths/min). Converts the floating-point array into a space-separated string. | Observation.code.coding.code = “9279-1” Observation.code.coding.display = “Respiratory rate” Observation.valueSampledData.data = [string] Observation.valueSampledData.origin.unit = “breaths/min” |
| Activity | Filters valid ranges (0–10 METs). | Observation.code.coding.code = “41950-7” Observation.code.coding.display = “Physical activity level” Observation.valueSampledData.data = [string] Observation.valueSampledData.origin.unit = “MET” |
| Posture | Filters valid angular ranges (−90° to 90°). | Observation.code.coding.code = “8361-8” Observation.code.coding.display = “Body position” Observation.valueSampledData.data = [string] Observation.valueSampledData.origin.unit = “position_code” |
| Array Mean Value | Evaluates the mean of the array against clinical thresholds to determine an interpretation code (N = Normal, H = High, L = Low). | Observation.interpretation.coding.code = [Code] Observation.interpretation.coding.system = “…v3-ObservationInterpretation” |
| Module | Components | Functional Responsibilities |
|---|---|---|
| auth/ | Authentication, user manager, cryptography, security decorators, auditing | Identity validation; session creation and destruction; permission evaluation; secure encryption; credential generation; path protection; event auditing |
| services/ | Patients, observations, professionals | Patient lifecycle (CRUD); CSV processing; automatic measurement detection; clinical validation; FHIR construction; remote synchronization |
| utils/ | Logging, validators | Patient lifecycle (CRUD); CSV processing; automatic measurement detection; clinical validation; FHIR construction; remote synchronization |
| Resource/Action | Super Admin | Doctor | Nurse | Auditor |
|---|---|---|---|---|
| Create patients | ✓ | ✓ | ✕ | ✕ |
| List patients | ✓ | ✓ | ✓ | ✓ |
| Consult patients | ✓ | ✓ | ✓ | ✓ |
| Edit patients | ✓ | ✓ | ✕ | ✕ |
| Delete patients | ✓ | ✕ | ✕ | ✕ |
| Create observations | ✓ | ✓ | ✓ | ✕ |
| Consult observations | ✓ | ✓ | ✓ | ✓ |
| See details of observations | ✓ | ✓ | ✓ | ✓ |
| Delete observations | ✓ | ✓ | ✕ | ✕ |
| Create user | ✓ | ✕ | ✕ | ✕ |
| List user | ✓ | ✕ | ✕ | ✕ |
| View user details | ✓ | ✕ | ✕ | ✕ |
| View passwords | ✓ | ✕ | ✕ | ✕ |
| Delete user | ✓ | ✕ | ✕ | ✕ |
| View audit | ✓ | ✕ | ✕ | ✓ |
| View my profile | ✓ | ✓ | ✓ | ✓ |
| Test Type | File Size | N Requests | Simultaneous Users | Mean (ms) | Std. Dev. (ms) | Min (ms) | Max (ms) | Status Code | Success Rate |
|---|---|---|---|---|---|---|---|---|---|
| LATENCY | 1 MB | 10 | 1 | 104.0 | 9.9 | 62.4 | 375.1 | 201 | 100% |
| LATENCY | 5 MB | 10 | 1 | 228.4 | 7.7 | 213.7 | 240.5 | 201 | 100% |
| LATENCY | 10 MB | 10 | 1 | 438.1 | 14.7 | 400.7 | 453.8 | 201 | 100% |
| Test Type | File Size | N Requests | Simultaneous Users | Mean (ms) | Std. Dev. (ms) | Min (ms) | Max (ms) | Status Code | Success Rate |
|---|---|---|---|---|---|---|---|---|---|
| CONCURRENCY | 5 MB | 10 | 10 | 113.0 | 14.6 | 92.0 | 133.1 | 201 | 100% |
| CONCURRENCY | 5 MB | 100 | 100 | 653.4 | 320.2 | 113.2 | 1239.5 | 201 | 100% |
| Test Type | File Size | N Requests | Simultaneous Users | Status Code | Success Rate | Heart Rate Error | Respiratory Rate Error | Activity Error | Posture Error |
|---|---|---|---|---|---|---|---|---|---|
| INTEGRITY | 5 MB | 200 | 1 | 201 | 100% | 0% (200/200) | 0% (200/200) | 0% (200/200) | 0% (200/200) |
| Validation Category | Validation Mechanism | Resources Evaluated | Result | Pass Rate |
|---|---|---|---|---|
| Structural conformance (FHIR R4 core) | HAPI FhirInstanceValidator interceptor | 200 | 200/200 | 100% (200/200) |
| LOINC code correctness | Flask middleware pre-submission check | 200 | 200/200 | 100% (200/200) |
| LOINC code correctness | Flask middleware pre-submission check | 200 | 200/200 | 100% (200/200) |
| Patient reference linkage | Flask middleware pre-submission check | 200 | 200/200 | 100% (200/200) |
| HTTP transaction success (201 Created) | HAPI FHIR server response | 200 | 200/200 | 100% (200/200) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Ramirez Lopez, L.J.; Jaimes Salazar, N.E.; Barbosa Posada, J.E. PIRE: Interoperable Platform for Electronic Records. Computers 2026, 15, 162. https://doi.org/10.3390/computers15030162
Ramirez Lopez LJ, Jaimes Salazar NE, Barbosa Posada JE. PIRE: Interoperable Platform for Electronic Records. Computers. 2026; 15(3):162. https://doi.org/10.3390/computers15030162
Chicago/Turabian StyleRamirez Lopez, Leonardo Juan, Norman Eduardo Jaimes Salazar, and Juan Esteban Barbosa Posada. 2026. "PIRE: Interoperable Platform for Electronic Records" Computers 15, no. 3: 162. https://doi.org/10.3390/computers15030162
APA StyleRamirez Lopez, L. J., Jaimes Salazar, N. E., & Barbosa Posada, J. E. (2026). PIRE: Interoperable Platform for Electronic Records. Computers, 15(3), 162. https://doi.org/10.3390/computers15030162






