Regulation of PD-L1 Expression by SAHA-Mediated Histone Deacetylase Inhibition in Lung Cancer Cells
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Lines and Reagents
2.2. Western Blotting
2.3. Immunostaining for Acetylation of Histones
2.4. RNA Extraction
2.5. cDNA Synthesis Using the RT2 First Strand Kit
2.6. Human Epigenetic Chromatin Modification Enzymes RT2 Profiler PCR Array
2.7. Biological Pathway Analysis Using STRING
2.8. Statistical Analysis
3. Results
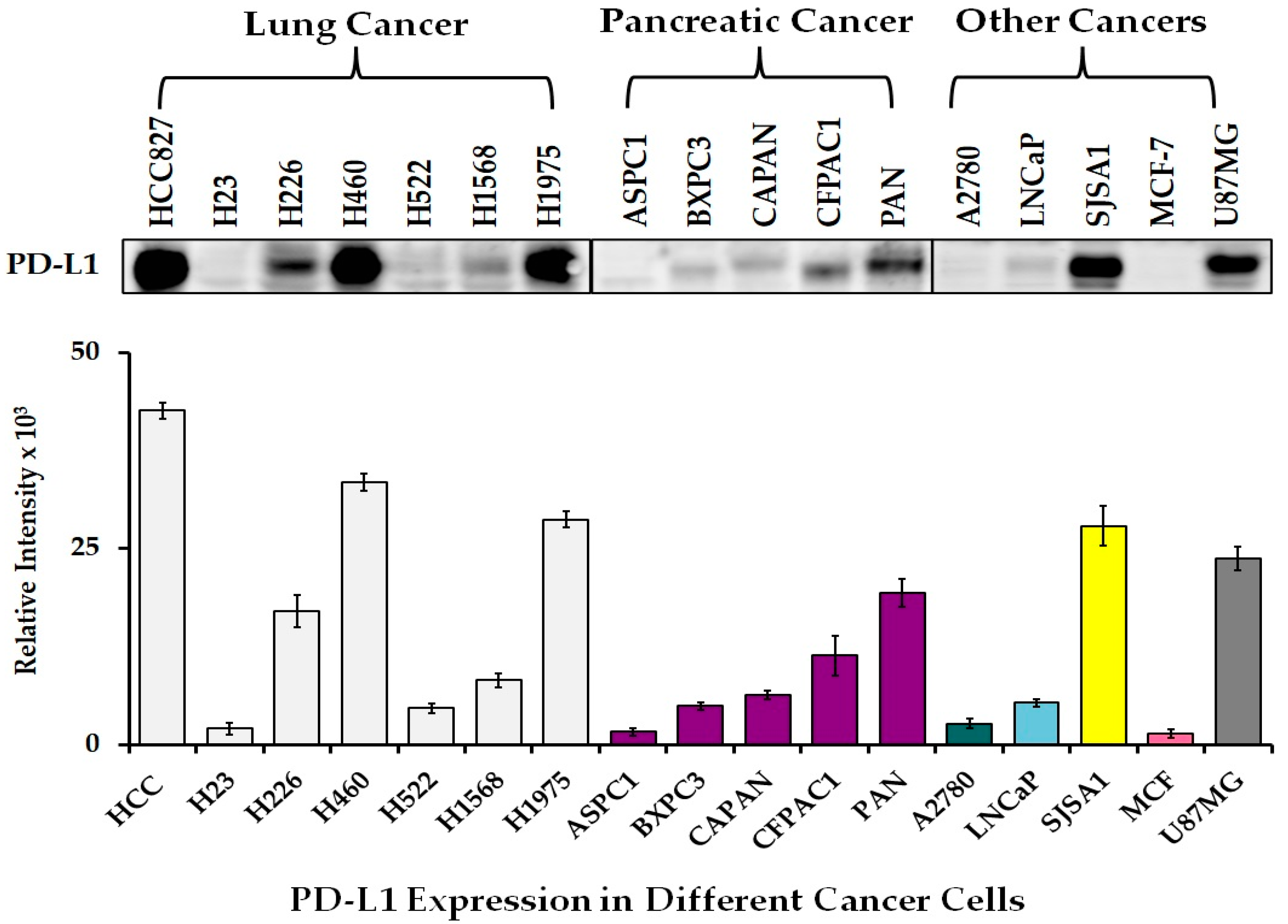
3.1. Expression of PD-L1 in Different Lung Cancer Cells and the Effects of SAHA
3.2. Expression of Cell Cycle Regulators
3.3. Effect of SAHA on STAT3, CDK4, CDK6, NF-κB, and pRB Levels
3.4. The Effect of SAHA on the Levels of Acetylated Histones
3.5. The Effect of SAHA on the Levels of DNMTs and Methylated Histones
3.6. Identification of Differential Gene Expression in Lung Cancer Cells Using the Heat Map Generated with the RT2 Profiler PCR Array Data
3.7. Network Analysis of Differentially Expressed Genes Using STRING
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Ac-H2A | Acetyl Histone H2A |
| Ac-H2B | Acetyl Histone H2B |
| Ac-H3 | Acetyl Histone H3 |
| Ac-H4 | Acetyl Histone H4 |
| ALK | Anaplastic Lymphoma Kinase |
| ATTC | American Type Culture Collection |
| BC | Breast Cancer |
| BCA | Bicinchoninic Acid |
| BSA | Bovine Serum Albumin |
| CTLA-4 | Cytotoxic T-Lymphocyte-Associated Protein 4 |
| CD80 | B7-1 |
| DNMTs | DNA Methyltransferases |
| ∆∆Ct | Double Delta Ct |
| EBV | Epstein–Barr Virus |
| FBS | Fetal Bovine Serum |
| FDR | False Discovery Rate |
| GC | Gastric Cancer |
| HATs | Histone Acetyltransferases |
| HDACIs | Histone Deacetylase Inhibitors |
| HC | Hodgkin’s Lymphoma |
| HCC | Hepatocellular Carcinoma |
| HIF-1α | Hypoxia-Inducible Factor Alpha |
| HNSCC | Head and Neck Squamous Cell Carcinoma |
| HRP | Horseradish Peroxidase |
| IC | Immune Checkpoint |
| ICC | Immunocytochemistry |
| MCC | Merkel cell carcinoma |
| MGMT | O-6-methylguanine-DNA methyltransferase |
| MLL1 | Mixed-Lineage Leukemia Protein |
| NSCLC | Non-Small Cell Lung Cancer |
| PBS | Phosphate-Buffered Saline |
| PD-L1 | Programmed Death Ligand-1 |
| PPI | Protein–Protein Interaction |
| pp53 | Phospho-p53 |
| pRB | Phospho-RB |
| pSTAT3 | Phosphorylated STAT3 |
| PMSF | Phenyl Methyl Sulfonyl Fluoride |
| qRT-PCR | Quantitative Reverse Transcription PCR |
| RCC | Renal Cell Carcinoma |
| RPMI | Roswell Park Memorial Institute |
| SAHA | Suberoylanilide Hydroxamic Acid |
| TCC | Transitional Cell Carcinoma |
| Tri Me-H3 | Tri-Methyl Histone H3 |
| Tri Me-H4 | Tri-Methyl Histone H4 |
References
- Siegel, R.L.; Giaquinto, A.N.; Jemal, A. Cancer statistics, 2024. CA Cancer J. Clin. 2024, 74, 12–49. [Google Scholar] [CrossRef]
- Megyesfalvi, Z.; Gay, C.M.; Popper, H.; Pirker, R.; Ostoros, G.; Heeke, S.; Lang, C.; Hoetzenecker, K.; Schwendenwein, A.; Boettiger, K.; et al. Clinical insights into small cell lung cancer: Tumor heterogeneity, diagnosis, therapy, and future directions. CA Cancer J. Clin. 2023, 73, 620–652. [Google Scholar] [CrossRef]
- Zappasodi, R.; Merghoub, T.; Wolchok, J.D. Emerging Concepts for Immune Checkpoint Blockade-Based Combination Therapies. Cancer Cell 2018, 33, 581–598. [Google Scholar] [CrossRef]
- Casey, S.C.; Tong, L.; Li, Y.; Do, R.; Walz, S.; Fitzgerald, K.N.; Gouw, A.M.; Baylot, V.; Gütgemann, I.; Eilers, M.; et al. MYC regulates the antitumor immune response through CD47 and PD-L1. Science 2016, 352, 227–231. [Google Scholar] [CrossRef]
- Butte, M.J.; Keir, M.E.; Phamduy, T.B.; Sharpe, A.H.; Freeman, G.J. Programmed death-1 ligand 1 interacts specifically with the B7-1 costimulatory molecule to inhibit T cell responses. Immunity 2007, 27, 111–122. [Google Scholar] [CrossRef]
- Chikuma, S.; Bluestone, J.A. CTLA-4 and tolerance: The biochemical point of view. Immunol. Res. 2003, 28, 241–253. [Google Scholar] [CrossRef]
- Rittmeyer, A.; Barlesi, F.; Waterkamp, D.; Park, K.; Ciardiello, F.; von Pawel, J.; Gadgeel, S.M.; Hida, T.; Kowalski, D.M.; Dols, M.C.; et al. OAK Study Group Atezolizumab versus docetaxel in patients with previously treated non-small-cell lung cancer (OAK): A phase 3, open-label, multicentre randomised controlled trial. Lancet 2017, 389, 255–265. [Google Scholar] [CrossRef]
- Topalian, S.L.; Hodi, F.S.; Brahmer, J.R.; Gettinger, S.N.; Smith, D.C.; McDermott, D.F.; Powderly, J.D.; Carvajal, R.D.; Sosman, J.A.; Atkins, M.B.; et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 2012, 366, 2443–2454. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; He, D.; Cheng, L.; Huang, C.; Zhang, Y.; Rao, X.; Kong, Y.; Li, C.; Zhang, Z.; Liu, J.; et al. p300/CBP inhibition enhances the efficacy of programmed death-ligand 1 blockade treatment in prostate cancer. Oncogene 2020, 39, 3939–3951. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, U.; Venkatesan, T.; Radhakrishnan, V.; Samuel, S.; Rasappan, P.; Rathinavelu, A. Cell Cycle Arrest and Cytotoxic Effects of SAHA and RG7388 Mediated through p21WAF1/CIP1 and p27KIP1 in Cancer Cells. Medicina 2019, 55, 30. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, U.; Venkatesan, T.; Radhakrishnan, V.; Samuel, S.; Rathinavelu, A. Differential Mechanisms of Cell Death Induced by HDAC Inhibitor SAHA and MDM2 Inhibitor RG7388 in MCF-7 Cells. Cells 2018, 8, 8. [Google Scholar] [CrossRef]
- Natarajan, U.; Venkatesan, T.; Das, J.; Rathinavelu, A. Regulation of PD-L1 Expression by Histone Deacetylase Inhibitor SAHA in Lung Cancer Cells. Biomed. J. Sci. Tech. Res. 2019, 19, 14519–14522. [Google Scholar] [CrossRef]
- Natarajan, U.; Venkatesan, T.; Radhakrishnan, V.; Samuel, S.; Rathinavelu, A. Effect of histone deacetylase inhibitor on PD-L1 expression in lung cancer cells. Cancer Res. 2019, 79, 5187. [Google Scholar] [CrossRef]
- Natarajan, U.; Venkatesan, T.; Mathuram, T.L.; Das, J.K.; Rathinavelu, A. Analysis of the regulation of PD-L1 expression and methyl transferases levels following inhibition of histone deacetylases in cancer cells. Cancer Res. 2020, 80, 3657. [Google Scholar] [CrossRef]
- Zhang, X.; Sun, J.; Canfrán-Duque, A.; Aryal, B.; Tellides, G.; Chang, Y.J.; Suárez, Y.; Osborne, T.F.; Fernández-Hernando, C. Deficiency of histone lysine methyltransferase SETDB2 in hematopoietic cells promotes vascular inflammation and accelerates atherosclerosis. JCI Insight 2021, 6, e147984. [Google Scholar] [CrossRef]
- Huisinga, K.L.; Brower-Toland, B.; Elgin, S.C. The contradictory definitions of heterochromatin: Transcription and silencing. Chromosoma 2006, 115, 110–122. [Google Scholar] [CrossRef] [PubMed]
- Karati, D.; Mukherjee, S.; Roy, S. Emerging therapeutic strategies in cancer therapy by HDAC inhibition as the chemotherapeutic potent and epigenetic regulator. Med. Oncol. 2024, 41, 84. [Google Scholar] [CrossRef]
- Natarajan, U.; Venkatesan, T.; Rathinavelu, A. Effect of the HDAC Inhibitor on Histone Acetylation and Methyltransferases in A2780 Ovarian Cancer Cells. Medicina 2021, 57, 456. [Google Scholar] [CrossRef]
- Shiozawa, K.; Nakanishi, T.; Tan, M.; Fang, H.B.; Wang, W.C.; Edelman, M.J.; Carlton, D.; Gojo, I.; Sausville, E.A.; Ross, D.D. Preclinical studies of vorinostat (suberoylanilide hydroxamic acid) combined with cytosine arabinoside and etoposide for treatment of acute leukemias. Clin. Cancer Res. 2009, 15, 1698–1707. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.K.; Cho, S.W. The Evasion Mechanisms of Cancer Immunity and Drug Intervention in the Tumor Microenvironment. Front. Pharmacol. 2022, 13, 868695. [Google Scholar] [CrossRef]
- Jiang, X.; Wang, J.; Deng, X.; Xiong, F.; Ge, J.; Xiang, B.; Wu, X.; Ma, J.; Zhou, M.; Li, X.; et al. Role of the tumor microenvironment in PD-L1/PD-1-mediated tumor immune escape. Mol. Cancer 2019, 18, 10. [Google Scholar] [CrossRef]
- Singh, D.; Khan, M.A.; Siddique, H.R. Role of epigenetic drugs in sensitizing cancers to anticancer therapies: Emerging trends and clinical advancements. Epigenomics 2023, 15, 517–537. [Google Scholar] [CrossRef]
- Zhang, T.; Forde, P.M.; Sullivan, R.J.; Sharon, E.; Barksdale, E.; Selig, W.; Ebbinghaus, S.; Fusaro, G.; Gunenc, D.; Battle, D.; et al. Addressing resistance to PD-1/PD-(L)1 pathway inhibition: Considerations for combinatorial clinical trial designs. J. Immunother. Cancer 2023, 11, e006555. [Google Scholar] [CrossRef]
- Wang, H.; Fu, C.; Du, J.; Wang, H.; He, R.; Yin, X.; Li, H.; Li, X.; Wang, H.; Li, K.; et al. Enhanced histone H3 acetylation of the PD-L1 promoter via the COP1/c-Jun/HDAC3 axis is required for PD-L1 expression in drug-resistant cancer cells. J. Exp. Clin. Cancer Res. 2020, 39, 29. [Google Scholar] [CrossRef]
- Natarajan, U.; Venkatesan Radhakrishnan, V.; Samuel, S.; Rathinavelu, A. Effect of Histone Deacetylase (HDAC) Inhibitor on Gene Expression in LNCaP-MST and MCF-7 Cells. Biomed. J. Sci. Tech. Res. 2018, 10, 7848–7855. [Google Scholar] [CrossRef]
- Natarajan, U.; Venkatesan Radhakrishnan, V.; Samuel, S.; Rathinavelu, A. Comparative Effects of HDAC Inhibitor SAHA and MDM2 Inhibitor RG7388 in LNCaP Prostate Cancer Cells. Biomed. J. Sci. Tech. Res. 2018, 8, 6. [Google Scholar] [CrossRef]
- Venkatesan, T.; Natarajan, U.; Rathinavelu, A. Effect of SAHA on epigenetic chromatin modification enzymes in LNCaP and MCF-7 cells. Cancer Res. 2020, 80, 4681. [Google Scholar] [CrossRef]
- Rathinavelu, A.; Natarajan, U. Chapter 5—HDAC inhibition in cancer. In Translational Epigenetics, Epigenetics in Organ Specific Disorders; Academic Press: Cambridge, MA, USA, 2023; Volume 34, pp. 63–97. [Google Scholar] [CrossRef]
- Natarajan, U.; Rathinavelu, A. A synergy between MDM2 and AURKB pathways in driving cell cycle and cell survival in lung cancer cells. Cancer Res. 2024, 84, 4593. [Google Scholar] [CrossRef]
- Woods, D.M.; Sodré, A.L.; Villagra, A.; Sarnaik, A.; Sotomayor, E.M.; Weber, J. HDAC Inhibition Upregulates PD-1 Ligands in Melanoma and Augments Immunotherapy with PD-1 Blockade. Cancer Immunol. Res. 2015, 3, 1375–1385. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Liu, X.; Zhang, J.; Rice, S.J.; Wagman, M.; Kong, Y.; Zhu, L.; Zhu, J.; Joshi, M.; Belani, C.P. Expression of PD-1 on CD4+ T cells in peripheral blood associates with poor clinical outcome in non-small cell lung cancer. Oncotarget 2016, 7, 56233–56240. [Google Scholar] [CrossRef]
- West, A.C.; Johnstone, R.W. New and emerging HDAC inhibitors for cancer treatment. J. Clin. Investig. 2014, 124, 30–39. [Google Scholar] [CrossRef]
- Deng, S.; Hu, Q.; Zhang, H.; Yang, F.; Peng, C.; Huang, C. HDAC3 Inhibition Upregulates PD-L1 Expression in B-Cell Lymphomas and Augments the Efficacy of Anti-PD-L1 Therapy. Mol. Cancer Ther. 2019, 18, 900–908. [Google Scholar] [CrossRef]
- Richon, V.M.; Sandhoff, T.W.; Rifkind, R.A.; Marks, P.A. Histone deacetylase inhibitor selectively induces p21WAF1 expression and gene-associated histone acetylation. Proc. Natl. Acad. Sci. USA 2000, 97, 10014–10019. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Li, S.; Li, J.; Wang, D.; Li, Q. Effect of microRNA-135a on Cell Proliferation, Migration, Invasion, Apoptosis and Tumor Angiogenesis Through the IGF-1/PI3K/Akt Signaling Pathway in Non-Small Cell Lung Cancer. Cell. Physiol. Biochem. 2017, 42, 1431–1446. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.Z.; Wu, S.C.; Chang, C.M.; Lin, C.L.; Kwan, A.L. Arctigenin, a Potent Ingredient of Arctium lappa L., Induces Endothelial Nitric Oxide Synthase and Attenuates Subarachnoid Hemorrhage-Induced Vasospasm through PI3K/Akt Pathway in a Rat Model. BioMed Res. Int. 2015, 2015, 490209. [Google Scholar] [CrossRef] [PubMed]
- Chin, Y.R.; Toker, A. The actin-bundling protein palladin is an Akt1-specific substrate that regulates breast cancer cell migration. Mol. Cell 2010, 38, 333–344. [Google Scholar] [CrossRef]
- Jafari, M.; Ghadami, E.; Dadkhah, T.; Akhavan-Niaki, H. PI3k/AKT signaling pathway: Erythropoiesis and beyond. J. Cell. Physiol. 2019, 234, 2373–2385. [Google Scholar] [CrossRef]
- Jiang, X.; Zhou, J.; Giobbie-Hurder, A.; Wargo, J.; Hodi, F.S. The activation of MAPK in melanoma cells resistant to BRAF inhibition promotes PD-L1 expression that is reversible by MEK and PI3K inhibition. Clin. Cancer Res. 2013, 19, 598–609. [Google Scholar] [CrossRef]
- Mittendorf, E.A.; Philips, A.V.; Meric-Bernstam, F.; Qiao, N.; Wu, Y.; Harrington, S.; Su, X.; Wang, Y.; Gonzalez-Angulo, A.M.; Akcakanat, A.; et al. PD-L1 expression in triple-negative breast cancer. Cancer Immunol. Res. 2014, 2, 361–370. [Google Scholar] [CrossRef]
- Song, M.; Chen, D.; Lu, B.; Wang, C.; Zhang, J.; Huang, L.; Wang, X.; Timmons, C.L.; Hu, J.; Liu, B.; et al. PTEN loss increases PD-L1 protein expression and affects the correlation between PD-L1 expression and clinical parameters in colorectal cancer. PLoS ONE 2013, 8, e65821. [Google Scholar] [CrossRef]
- Cheng, H.; Liu, P.; Zhang, F.; Xu, E.; Symonds, L.; Ohlson, C.E.; Bronson, R.T.; Maira, S.M.; Di Tomaso, E.; Li, J.; et al. A genetic mouse model of invasive endometrial cancer driven by concurrent loss of Pten and Lkb1 Is highly responsive to mTOR inhibition. Cancer Res. 2014, 74, 15–23. [Google Scholar] [CrossRef]
- Gowrishankar, K.; Gunatilake, D.; Gallagher, S.J.; Tiffen, J.; Rizos, H.; Hersey, P. Inducible but not constitutive expression of PD-L1 in human melanoma cells is dependent on activation of NF-κB. PLoS ONE 2015, 10, e0123410. [Google Scholar] [CrossRef]
- Antonangeli, F.; Natalini, A.; Garassino, M.C.; Sica, A.; Santoni, A.; Di Rosa, F. Regulation of PD-L1 Expression by NF-κB in Cancer. Front. Immunol. 2020, 11, 584626. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.; Zhang, Y.; Ma, N.; Shi, J.; Hou, Y. NF-κB role on tumor proliferation, migration, invasion and immune escape. Cancer Gene Ther. 2024, 31, 1599–1610. [Google Scholar] [CrossRef] [PubMed]
- Nihira, N.T.; Miki, Y. Regulation of Intrinsic Functions of PD-L1 by Post-Translational Modification in Tumors. Front. Oncol. 2022, 12, 825284. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Ma, C.; Zhang, Z.; Zhang, H.; Hu, H. NF-κB signaling in inflammation and cancer. MedComm 2021, 2, 618–653. [Google Scholar] [CrossRef]
- Betzler, A.C.; Theodoraki, M.N.; Schuler, P.J.; Döscher, J.; Laban, S.; Hoffmann, T.K.; Brunner, C. NF-κB and Its Role in Checkpoint Control. Int. J. Mol. Sci. 2020, 21, 3949. [Google Scholar] [CrossRef]
- Pollizzi, K.N.; Powell, J.D. Integrating canonical and metabolic signalling programmes in the regulation of T cell responses. Nat. Rev. Immunol. 2014, 14, 435–446. [Google Scholar] [CrossRef]
- Pawelec, G.; Derhovanessian, E.; Larbi, A. Immunosenescence and cancer. Crit. Rev. Oncol. Hematol. 2010, 75, 165–172. [Google Scholar] [CrossRef]
- Barsoum, I.B.; Koti, M.; Siemens, D.R.; Graham, C.H. Mechanisms of hypoxia-mediated immune escape in cancer. Cancer Res. 2014, 74, 7185–7190. [Google Scholar] [CrossRef]
- Shehade, H.; Oldenhove, G.; Moser, M. Hypoxia in the intestine or solid tumors: A beneficial or deleterious alarm signal? Eur. J. Immunol. 2014, 44, 2550–2557. [Google Scholar] [CrossRef]
- Barsoum, I.B.; Smallwood, C.A.; Siemens, D.R.; Graham, C.H. A mechanism of hypoxia-mediated escape from adaptive immunity in cancer cells. Cancer Res. 2014, 74, 665–674. [Google Scholar] [CrossRef]
- Noman, M.Z.; Desantis, G.; Janji, B.; Hasmim, M.; Karray, S.; Dessen, P.; Bronte, V.; Chouaib, S. PD-L1 is a novel direct target of HIF-1α, and its blockade under hypoxia enhanced MDSC-mediated T cell activation. J. Exp. Med. 2014, 211, 781–790. [Google Scholar] [CrossRef]
- Noman, M.Z.; Chouaib, S. Targeting hypoxia at the forefront of anticancer immune responses. Oncoimmunology 2015, 3, e954463. [Google Scholar] [CrossRef]
- Shurin, M.R.; Umansky, V. Cross-talk between HIF and PD-1/PD-L1 pathways in carcinogenesis and therapy. J. Clin. Investig. 2022, 132, e159473. [Google Scholar] [CrossRef]
- Zheng, Z.; Bian, C.; Wang, H.; Su, J.; Meng, L.; Xin, Y.; Jiang, X. Prediction of immunotherapy efficacy and immunomodulatory role of hypoxia in colorectal cancer. Ther. Adv. Med. Oncol. 2022, 14, 17588359221138383. [Google Scholar] [CrossRef]
- Chen, J.; Jiang, C.C.; Jin, L.; Zhang, X.D. Regulation of PD-L1: A novel role of pro-survival signalling in cancer. Ann. Oncol. 2016, 27, 409–416. [Google Scholar] [CrossRef] [PubMed]
- Bandopadhyay, S.; Patranabis, S. Mechanisms of HIF-driven immunosuppression in tumour microenvironment. J. Egypt. Natl. Cancer Inst. 2023, 35, 27. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Han, F.; Du, Y.; Shi, H.; Zhou, W. Hypoxic microenvironment in cancer: Molecular mechanisms and therapeutic interventions. Signal Transduct. Target. Ther. 2023, 8, 70. [Google Scholar] [CrossRef]
- Schito, L.; Semenza, G.L. Hypoxia-Inducible Factors: Master Regulators of Cancer Progression. Trends Cancer 2016, 2, 758–770. [Google Scholar] [CrossRef] [PubMed]
- Maheshwari, M.; Yadav, N.; Hasanain, M.; Pandey, P.; Sahai, R.; Choyal, K.; Singh, A.; Nengroo, M.A.; Saini, K.K.; Kumar, D.; et al. Inhibition of p21 activates Akt kinase to trigger ROS-induced autophagy and impacts on tumor growth rate. Cell Death Dis. 2022, 13, 1045. [Google Scholar] [CrossRef]
- Krishnan, R.; Murugiah, M.; Lakshmi, N.P.; Mahalingam, S. Guanine nucleotide binding protein like-1 (GNL1) promotes cancer cell proliferation and survival through AKT/p21 CIP1 signaling cascade. Mol. Biol. Cell 2020, 31, 2904–2919. [Google Scholar] [CrossRef] [PubMed]
- Shrestha, M.; Wang, D.Y.; Ben-David, Y.; Zacksenhaus, E. CDK4/6 inhibitors and the pRB-E2F1 axis suppress PVR and PD-L1 expression in triple-negative breast cancer. Oncogenesis 2023, 12, 29. [Google Scholar] [CrossRef]
- Chen, L.; Liu, S.; Tao, Y. Regulating tumor suppressor genes: Post-translational modifications. Signal Transduct. Target. Ther. 2020, 5, 90. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, R.; Zhao, L.; Zhou, Y.; Shirasawa, M.; Uchida, A.; Murakawa, H.; Fikriyanti, M.; Iwanaga, R.; Bradford, A.P.; Araki, K.; et al. Deregulated E2F Activity as a Cancer-Cell Specific Therapeutic Tool. Genes 2023, 14, 393. [Google Scholar] [CrossRef]
- Al Bitar, S.; Gali-Muhtasib, H. The Role of the Cyclin Dependent Kinase Inhibitor p21cip1/waf1 in Targeting Cancer: Molecular Mechanisms and Novel Therapeutics. Cancers 2019, 11, 1475. [Google Scholar] [CrossRef]
- Chang, B.D.; Watanabe, K.; Broude, E.V.; Fang, J.; Poole, J.C.; Kalinichenko, T.V.; Roninson, I.B. Effects of p21Waf1/Cip1/Sdi1 on cellular gene expression: Implications for carcinogenesis, senescence, and age-related diseases. Proc. Natl. Acad. Sci. USA 2000, 97, 4291–4296. [Google Scholar] [CrossRef]
- Nielsen, S.J.; Schneider, R.; Bauer, U.M.; Bannister, A.J.; Morrison, A.; O’Carroll, D.; Firestein, R.; Cleary, M.; Jenuwein, T.; Herrera, R.E.; et al. Rb targets histone H3 methylation and HP1 to promoters. Nature 2001, 412, 561–565. [Google Scholar] [CrossRef]
- Marzec, M.; Zhang, Q.; Goradia, A.; Raghunath, P.N.; Liu, X.; Paessler, M.; Wang, H.Y.; Wysocka, M.; Cheng, M.; Ruggeri, B.A.; et al. Oncogenic kinase NPM/ALK induces through STAT3 expression of immunosuppressive protein CD274 (PD-L1, B7-H1). Proc. Natl. Acad. Sci. USA 2008, 105, 20852–20857. [Google Scholar] [CrossRef]
- Pearson, J.D.; Lee, J.K.; Bacani, J.T.; Lai, R.; Ingham, R.J. NPM-ALK: The Prototypic Member of a Family of Oncogenic Fusion Tyrosine Kinases. J. Signal Transduct. 2012, 2012, 123253. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Li, S.; Medeiros, L.J.; Lin, P.; Wang, S.A.; Tang, G.; Yin, C.C.; You, M.J.; Khoury, J.D.; Iyer, S.P.; et al. PD-L1 expression is associated with ALK positivity and STAT3 activation, but not outcome in patients with systemic anaplastic large cell lymphoma. Mod. Pathol. 2020, 33, 324–333. [Google Scholar] [CrossRef]
- Kong, J.; Dasari, S.; Feldman, A.L. PD-L1 expression in anaplastic large cell lymphoma. Mod. Pathol. 2020, 33, 1232–1233. [Google Scholar] [CrossRef]
- Xu, J.; Li, S.; Medeiros, L.J. Reply to “PD-L1 expression in anaplastic large cell lymphoma”. Mod. Pathol. 2020, 33, 1234–1235. [Google Scholar] [CrossRef] [PubMed]
- Fang, W.; Zhang, J.; Hong, S.; Zhan, J.; Chen, N.; Qin, T.; Tang, Y.; Zhang, Y.; Kang, S.; Zhou, T.; et al. EBV-driven LMP1 and IFN-γ up-regulate PD-L1 in nasopharyngeal carcinoma: Implications for oncotargeted therapy. Oncotarget 2014, 5, 12189–12202. [Google Scholar] [CrossRef]
- Vadivel, C.K.; Gluud, M.; Torres-Rusillo, S.; Boding, L.; Willerslev-Olsen, A.; Buus, T.B.; Nielsen, T.K.; Persson, J.L.; Bonefeld, C.M.; Geisler, C.; et al. JAK3 Is Expressed in the Nucleus of Malignant T Cells in Cutaneous T Cell Lymphoma (CTCL). Cancers 2021, 13, 280. [Google Scholar] [CrossRef]
- Mielcarek, M.; Benn, C.L.; Franklin, S.A.; Smith, D.L.; Woodman, B.; Marks, P.A.; Bates, G.P. SAHA decreases HDAC 2 and 4 levels in vivo and improves molecular phenotypes in the R6/2 mouse model of Huntington’s disease. PLoS ONE 2011, 6, e27746. [Google Scholar] [CrossRef]
- Wang, Y.; Sun, Q.; Mu, N.; Sun, X.; Wang, Y.; Fan, S.; Su, L.; Liu, X. The deubiquitinase USP22 regulates PD-L1 degradation in human cancer cells. Cell Commun. Signal. 2020, 18, 112. [Google Scholar] [CrossRef] [PubMed]
- Sukowati, C.; Cabral, L.K.D.; Anfuso, B.; Dituri, F.; Negro, R.; Giannelli, G.; Tiribelli, C. PD-L1 Downregulation and DNA Methylation Inhibition for Molecular Therapy against Cancer Stem Cells in Hepatocellular Carcinoma. Int. J. Mol. Sci. 2023, 24, 13357. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Liu, Y.; Meng, L.; Liu, K.; Ji, B. Targeting the PD-L1/DNMT1 axis in acquired resistance to sorafenib in human hepatocellular carcinoma. Oncol. Rep. 2017, 38, 899–907. [Google Scholar] [CrossRef]
- Tarcic, O.; Pateras, I.S.; Cooks, T.; Shema, E.; Kanterman, J.; Ashkenazi, H.; Boocholez, H.; Hubert, A.; Rotkopf, R.; Baniyash, M.; et al. RNF20 Links Histone H2B Ubiquitylation with Inflammation and Inflammation-Associated Cancer. Cell Rep. 2016, 14, 1462–1476. [Google Scholar] [CrossRef]
- Kufrin, V.; Seiler, A.; Brilloff, S.; Rothfuß, H.; Küchler, S.; Schäfer, S.; Rahimian, E.; Baumgarten, J.; Ding, L.; Buchholz, F.; et al. The histone modifier KAT2A presents a selective target in a subset of well-differentiated microsatellite-stable colorectal cancers. Cell Death Differ. 2025, 32, 1259–1272. [Google Scholar] [CrossRef]
- Yu, Z.; Ding, M.; Cai, Y.; Lu, T.; Chen, X.; Zhou, X.; Wang, X. Histone regulator KAT2A acts as a potential biomarker related to tumor microenvironment and prognosis of diffuse large B cell lymphoma. BMC Cancer 2023, 23, 934. [Google Scholar] [CrossRef]
- Chiu, B.L.; Li, C.H.; Chang, C.C. Selective modulation of MHC class II chaperons by a novel IFN-γ-inducible class II transactivator variant in lung adenocarcinoma A549 cells. Biochem. Biophys. Res. Commun. 2013, 440, 190–195. [Google Scholar] [CrossRef]
- Giaimo, B.D.; Ferrante, F.; Herchenröther, A.; Hake, S.B.; Borggrefe, T. The histone variant H2A.Z in gene regulation. Epigenetics Chromatin 2019, 12, 37. [Google Scholar] [CrossRef] [PubMed]
- Anderson, L.; Gomes, M.R.; daSilva, L.F.; Pereira, A.D.S.A.; Mourão, M.M.; Romier, C.; Pierce, R.; Verjovski-Almeida, S. Histone deacetylase inhibition modulates histone acetylation at gene promoter regions and affects genome-wide gene transcription in Schistosoma mansoni. PLoS Neglected Trop. Dis. 2017, 11, e0005539. [Google Scholar] [CrossRef] [PubMed]
- Ling, R.; Wang, J.; Fang, Y.; Yu, Y.; Su, Y.; Sun, W.; Li, X.; Tang, X. HDAC-an important target for improving tumor radiotherapy resistance. Front. Oncol. 2023, 13, 1193637. [Google Scholar] [CrossRef] [PubMed]
- Slaughter, M.J.; Shanle, E.K.; Khan, A.; Chua, K.F.; Hong, T.; Boxer, L.D.; Allis, C.D.; Josefowicz, S.Z.; Garcia, B.A.; Rothbart, S.B.; et al. HDAC inhibition results in widespread alteration of the histone acetylation landscape and BRD4 targeting to gene bodies. Cell Rep. 2021, 34, 108638. [Google Scholar] [CrossRef]
- Huang, Q.; Zheng, Y.; Gao, Z.; Yuan, L.; Sun, Y.; Chen, H. Comparative Efficacy and Safety of PD-1/PD-L1 Inhibitors for Patients with Solid Tumors: A Systematic Review and Bayesian Network Meta-analysis. J. Cancer 2021, 12, 1133–1143. [Google Scholar] [CrossRef]
- Herbst, R.S.; Soria, J.C.; Kowanetz, M.; Fine, G.D.; Hamid, O.; Gordon, M.S.; Sosman, J.A.; McDermott, D.F.; Powderly, J.D.; Gettinger, S.N.; et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature 2014, 515, 563–567. [Google Scholar] [CrossRef]
- Dong, H.; Zhu, G.; Tamada, K.; Chen, L. B7-H1, a third member of the B7 family, co-stimulates T-cell proliferation and interleukin-10 secretion. Nat. Med. 1999, 5, 1365–1369. [Google Scholar] [CrossRef]
- Zhang, J.; Dang, F.; Ren, J.; Wei, W. Biochemical Aspects of PD-L1 Regulation in Cancer Immunotherapy. Trends Biochem. Sci. 2018, 43, 1014–1032. [Google Scholar] [CrossRef] [PubMed]











| Primary Antibodies | Vendor | Catalog No | WB/IF |
|---|---|---|---|
| PD-L1 | R&D Systems | AF1019 | 1:1000 |
| MDM2 | CST | sc-13161 | 2:1000 |
| p53 | CST | 32532 | 1:1000 |
| pp53 (Phospho-p53) | CST | 12571 | 1:1000 |
| p21 | CST | 2947 | 1:1000 |
| p27 | CST | 3686 | 1:1000 |
| STAT3 | CST | 9139 | 1:1000 |
| pSTAT3 (Phospho-Stat3) | CST | 9145 | 1:1000 |
| NFkB | CST | 4764 | 1:1000 |
| CDK4 | CST | 12790 | 1:1000 |
| CDK6 | CST | 13331 | 1:1000 |
| pRB (Phospho-RB) | CST | 8516 | 1:1000 |
| Ac-H2A (Acetyl Histone H2A) | CST | 2576 | 1:1000 |
| Ac-H2B (Acetyl Histone H2B) | CST | 34156 | 1:1000 |
| Ac-H3 (Acetyl Histone H3) | CST | 9677 | 1:1000/1:1500 |
| Ac-H4 (Acetyl Histone H4) | CST | 13944 | 1:1000/1:1500 |
| Tri Me-H3 (Tri-Methyl Histone H3) | CST | 4909 | 1:1000 |
| Tri Me-H4 (Tri-Methyl Histone H4) | CST | 5737 | 1:1000 |
| DNMT3B | CST | 67259 | 1:1000 |
| MGMT | CST | 2739 | 1:1000 |
| (A) H460—Up- and Down-Regulated Genes | |||
| Gene | Description | Fold Change | Accession Number |
| HDAC9 | Histone deacetylase 9 | 9.03 | NM_178425 |
| HDAC3 | Histone deacetylase 3 | 3.96 | NM_003883 |
| NCOA3 | Nuclear receptor coactivator 3 | 3.92 | NM_181659 |
| HDAC5 | Histone deacetylase 5 | 3.68 | NM_005474 |
| KDM5B | Lysine (K)-specific demethylase 5B | 3.06 | NM_006618 |
| KMT2C | Myeloid/lymphoid or mixed-lineage leukemia 3 | 3.02 | NM_170606 |
| DZIP3 | DAZ interacting protein 3, zinc finger | 2.54 | NM_014648 |
| HDAC11 | Histone deacetylase 11 | 2.42 | NM_024827 |
| SETDB2 | SET domain, bifurcated 2 | 2.42 | NM_031915 |
| RNF20 | Ring finger protein 20 | 2.08 | NM_019592 |
| AURKA | Aurora kinase A | −5.21 | NM_003600 |
| SETD7 | SET domain containing (lysine methyltransferase) 7 | −3.93 | NM_030648 |
| DNMT3A | DNA (cytosine-5-)-methyltransferase 3 alpha | −3.92 | NM_022552 |
| NSD1 | Nuclear receptor binding SET domain protein 1 | −3.50 | NM_022455 |
| HDAC7 | Histone deacetylase 7 | −3.21 | NM_001098416 |
| AURKB | Aurora kinase B | −2.75 | NM_004217 |
| SUV39H1 | Suppressor of variegation 3–9 homolog 1 (Drosophila) | −2.74 | NM_003173 |
| WHSC1 | Wolf-Hirschhorn syndrome candidate 1 | −2.71 | NM_007331 |
| SMYD3 | SET and MYND domain containing 3 | −2.69 | NM_022743 |
| SETD6 | SET domain containing 6 | −2.55 | NM_024860 |
| SETD1B | SET domain containing 1B | −2.38 | NM_015048 |
| KAT6B | K(lysine) acetyltransferase 6B | −2.33 | NM_012330 |
| PRMT1 | Protein arginine methyltransferase 1 | −2.23 | NM_001536 |
| KAT2A | K(lysine) acetyl transferase 2A | −2.09 | NM_021078 |
| KAT5 | K(lysine) acetyltransferase 5 | −2.01 | NM_006388 |
| USP21 | Ubiquitin-specific peptidase 21 | −2.01 | NM_012475 |
| (B) HCC827—Up- and Down-Regulated Genes | |||
| Gene | Description | Fold Change | Accession Number |
| HDAC3 | Histone deacetylase 3 | 7.05 | NM_003883 |
| KDM5C | Lysine (K)-specific demethylase 5C | 4.67 | NM_004187 |
| KMT2A | Myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila) | 4.11 | NM_005933 |
| KMT2C | Myeloid/lymphoid or mixed-lineage leukemia 3 | 4.09 | NM_170606 |
| KDM6B | Lysine (K)-specific demethylase 6B | 3.48 | NM_001080424 |
| HDAC9 | Histone deacetylase 9 | 2.90 | NM_178425 |
| RNF20 | Ring finger protein 20 | 2.25 | NM_019592 |
| NCOA3 | Nuclear receptor coactivator 3 | 2.14 | NM_181659 |
| DNMT3B | DNA (cytosine-5-)-methyltransferase 3 beta | −21.62 | NM_006892 |
| NSD1 | Nuclear receptor binding SET domain protein 1 | −6.06 | NM_022455 |
| AURKB | Aurora Kinase B | −4.65 | NM_004217 |
| DNMT3A | DNA (cytosine-5-)-methyltransferase 3 alpha | −3.79 | NM_022552 |
| NEK6 | NIMA (never in mitosis gene a)-related kinase 6 | −3.62 | NM_014397 |
| KAT6B | K(lysine) acetyltransferase 6B | −3.22 | NM_012330 |
| SUV39H1 | Suppressor of variegation 3–9 homolog 1 (Drosophila) | −2.82 | NM_003173 |
| PRMT1 | Protein arginine methyltransferase 1 | −2.62 | NM_001536 |
| WHSC1 | Wolf-Hirschhorn syndrome candidate 1 | −2.59 | NM_007331 |
| AURKA | Aurora kinase A | −2.51 | NM_003600 |
| SETD1B | SET domain containing 1B | −2.29 | NM_015048 |
| USP21 | Ubiquitin-specific peptidase 21 | −2.22 | NM_012475 |
| HDAC10 | Histone deacetylase 10 | −2.19 | NM_032019 |
| RPS6KA3 | Ribosomal protein S6 kinase, 90kDa, polypeptide 3 | −2.18 | NM_004586 |
| (A) | Gene Count | Protein Names | |
| Clusters 1 |  | 12 | KDM5B, KMT2C, NSD1, NSD2, ASH1L, PRMT1, RNF20, SETD1B, SETD6, SETD7, SETDB2, SMYD3 |
| Clusters 2 |  | 11 | DNMT3A, HDAC11, HDAC3, HDAC5, HDAC7, HDAC9, KAT2A, KAT5, KAT6B, NCOA3, SUV39H1 |
| Clusters 3 |  | 3 | AURKA, AURKB, USP21 |
| (B) | Gene Count | Protein Names | |
| Clusters 1 |  | 15 | AURKB, DNMT3A, DNMT3B, HDAC10, HDAC3, HDAC9, KAT6B, KDM5C, KDM6B, KMT2A, NSD1, NSD2, NCOA3, PRMT1, SUV39H1 |
| Clusters 2 |  | 2 | AURKA, USP21 |
| Clusters 3 |  | 2 | RNF20, SETD1B |
| Clusters 4 |  | 1 | KMT2C |
| p21- and PD-L1-Linked Genes | |
|---|---|
| p21 | AURKA, AURKB, DNMT3A, DNMT3B, DNMT1, KAT7, KAT2B |
| PD-L1 | USP22, DNMT1, RNF20, KAT2A, CIITA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Natarajan, U.; Rathinavelu, A. Regulation of PD-L1 Expression by SAHA-Mediated Histone Deacetylase Inhibition in Lung Cancer Cells. Cancers 2025, 17, 2919. https://doi.org/10.3390/cancers17172919
Natarajan U, Rathinavelu A. Regulation of PD-L1 Expression by SAHA-Mediated Histone Deacetylase Inhibition in Lung Cancer Cells. Cancers. 2025; 17(17):2919. https://doi.org/10.3390/cancers17172919
Chicago/Turabian StyleNatarajan, Umamaheswari, and Appu Rathinavelu. 2025. "Regulation of PD-L1 Expression by SAHA-Mediated Histone Deacetylase Inhibition in Lung Cancer Cells" Cancers 17, no. 17: 2919. https://doi.org/10.3390/cancers17172919
APA StyleNatarajan, U., & Rathinavelu, A. (2025). Regulation of PD-L1 Expression by SAHA-Mediated Histone Deacetylase Inhibition in Lung Cancer Cells. Cancers, 17(17), 2919. https://doi.org/10.3390/cancers17172919







