Impact of Metastatic Microenvironment on Physiology and Metabolism of Small Cell Neuroendocrine Prostate Cancer Patient-Derived Xenografts
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal Models
2.2. Short Tandem Repeat (STR) Profiling
2.3. 1H MRI
2.4. 13C HP MRI
2.5. MRI Data Analysis
2.6. Stable Isotope Resolved Metabolomics (SIRM) of PDX Tumors
2.7. Biological Correlates
2.8. Immunohistochemistry (IHC)
2.9. Micro-Computed Tomography (CT) Image Acquisition and Analysis
2.10. RNA-Sequencing (RNA-Seq)
2.11. Statistical Analysis
3. Results
3.1. Growth Characteristics of SCNC PDX Propagated Under the Renal Capsule
3.2. MRI and Glycolytic Features of PDXs Propagated Under the Renal Capsule
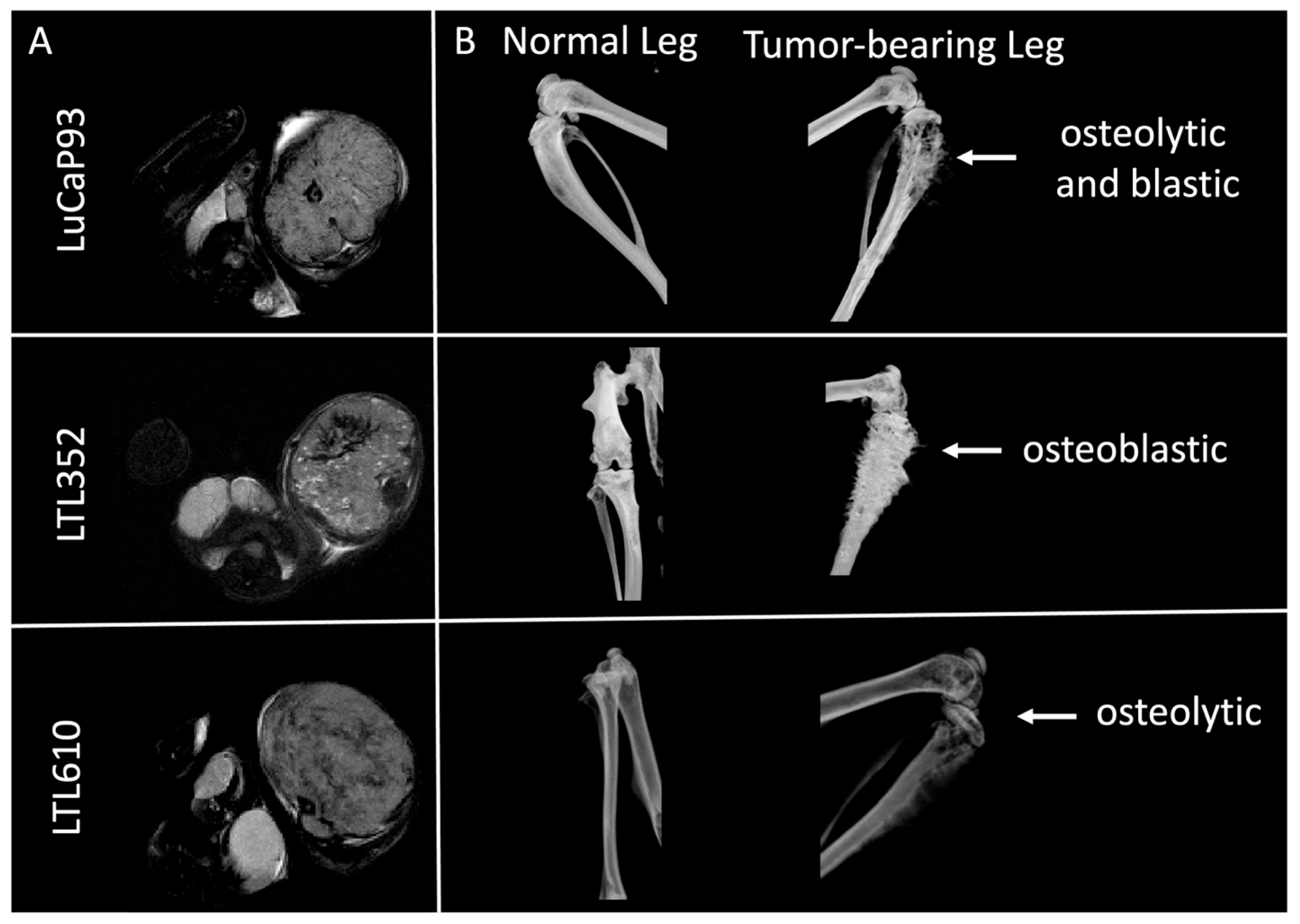
3.3. Growth Characteristics of SCNC PDXs Propagated in Bone
3.4. MRI and Glycolytic Features of PDXs Propagated in Bone
3.5. Growth Characteristics of SCNC PDXs Propagated in Liver
3.6. MRI and Glycolytic Features of SCNC PDXs Propagated in Liver
3.7. Microenvironmental Impact on Functional and Metabolic Parameters of SCNC PDXs
3.8. Steady-State Metabolism of PDXs in Bone Versus Liver
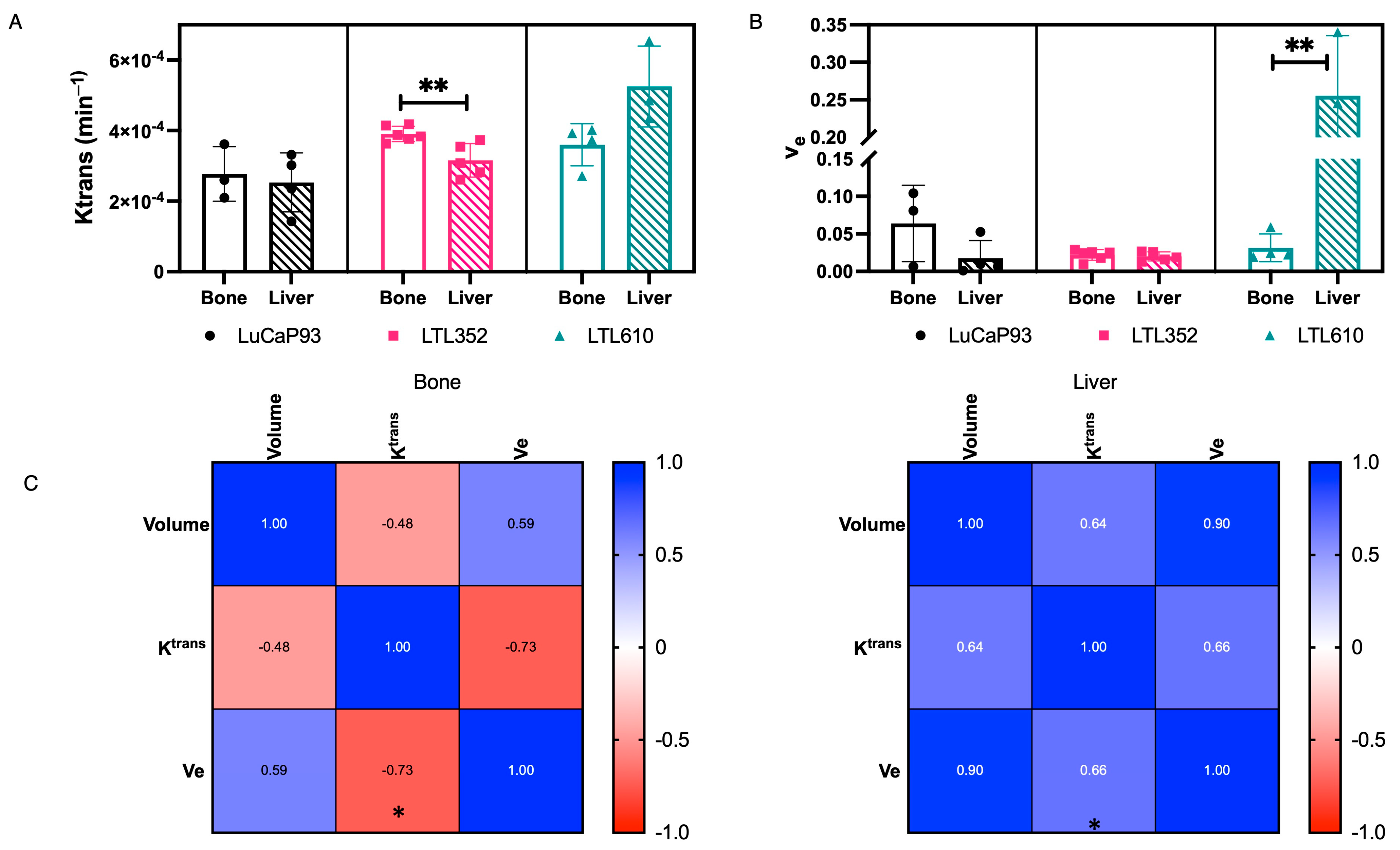
3.9. Characterization of Perfusion of SCNC PDXs in Bone and Liver by Multiparametric MRI
3.10. Comparison of Transcriptional Landscape of PDXs in the Bone and Liver
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kanayama, M.; Luo, J. Delineating the Molecular Events Underlying Development of Prostate Cancer Variants with Neuroendocrine/Small Cell Carcinoma Characteristics. Int. J. Mol. Sci. 2021, 22, 12742. [Google Scholar] [CrossRef] [PubMed]
- Beltran, H.; Prandi, D.; Mosquera, J.M.; Benelli, M.; Puca, L.; Cyrta, J.; Marotz, C.; Giannopoulou, E.; Chakravarthi, B.V.S.K.; Varambally, S.; et al. Divergent Clonal Evolution of Castration-Resistant Neuroendocrine Prostate Cancer. Nat. Med. 2016, 22, 298–305. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Xu, W.; Mierxiati, A.; Huang, Y.; Wei, Y.; Lin, G.; Dai, B.; Freedland, S.J.; Qin, X.; Zhu, Y.; et al. Low-serum Prostate-specific Antigen Level Predicts Poor Outcomes in Patients with Primary Neuroendocrine Prostate Cancer. Prostate 2019, 79, 1563–1571. [Google Scholar] [CrossRef] [PubMed]
- Epstein, J.I.; Amin, M.B.; Beltran, H.; Lotan, T.L.; Mosquera, J.-M.; Reuter, V.E.; Robinson, B.D.; Troncoso, P.; Rubin, M.A. Proposed Morphologic Classification of Prostate Cancer With Neuroendocrine Differentiation. Am. J. Surg. Pathol. 2014, 38, 756–767. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, R.; Huang, J.; Alumkal, J.J.; Zhang, L.; Feng, F.Y.; Thomas, G.V.; Weinstein, A.S.; Friedl, V.; Zhang, C.; Witte, O.N.; et al. Clinical and Genomic Characterization of Treatment-Emergent Small-Cell Neuroendocrine Prostate Cancer: A Multi-Institutional Prospective Study. JCO 2018, 36, 2492–2503. [Google Scholar] [CrossRef] [PubMed]
- Logotheti, S.; Papadaki, E.; Zolota, V.; Logothetis, C.; Vrahatis, A.G.; Soundararajan, R.; Tzelepi, V. Lineage Plasticity and Stemness Phenotypes in Prostate Cancer: Harnessing the Power of Integrated “Omics” Approaches to Explore Measurable Metrics. Cancers 2023, 15, 4357. [Google Scholar] [CrossRef] [PubMed]
- Abida, W.; Cyrta, J.; Heller, G.; Prandi, D.; Armenia, J.; Coleman, I.; Cieslik, M.; Benelli, M.; Robinson, D.; Van Allen, E.M.; et al. Genomic Correlates of Clinical Outcome in Advanced Prostate Cancer. Proc. Natl. Acad. Sci. USA 2019, 116, 11428–11436. [Google Scholar] [CrossRef] [PubMed]
- Bluemn, E.G.; Coleman, I.M.; Lucas, J.M.; Coleman, R.T.; Hernandez-Lopez, S.; Tharakan, R.; Bianchi-Frias, D.; Dumpit, R.F.; Kaipainen, A.; Corella, A.N.; et al. Androgen Receptor Pathway-Independent Prostate Cancer Is Sustained through FGF Signaling. Cancer Cell 2017, 32, 474–489.e6. [Google Scholar] [CrossRef] [PubMed]
- Dondi, F.; Antonelli, A.; Suardi, N.; Guerini, A.E.; Albano, D.; Lucchini, S.; Camoni, L.; Treglia, G.; Bertagna, F. PET/CT and Conventional Imaging for the Assessment of Neuroendocrine Prostate Cancer: A Systematic Review. Cancers 2023, 15, 4404. [Google Scholar] [CrossRef] [PubMed]
- Iravani, A.; Mitchell, C.; Akhurst, T.; Sandhu, S.; Hofman, M.S.; Hicks, R.J. Molecular Imaging of Neuroendocrine Differentiation of Prostate Cancer: A Case Series. Clin. Genitourin. Cancer 2021, 19, e200–e205. [Google Scholar] [CrossRef] [PubMed]
- Eule, C.J.; Hu, J.; Al-Saad, S.; Collier, K.; Boland, P.; Lewis, A.R.; McKay, R.R.; Narayan, V.; Bosse, D.; Mortazavi, A.; et al. Outcomes of Second-Line Therapies in Patients With Metastatic de Novo and Treatment-Emergent Neuroendocrine Prostate Cancer: A Multi-Institutional Study. Clin. Genitourin. Cancer 2023, 21, 483–490. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Alabi, B.R.; Yin, Q.; Stoyanova, T. Molecular Mechanisms Underlying the Development of Neuroendocrine Prostate Cancer. Semin. Cancer Biol. 2022, 86, 57–68. [Google Scholar] [CrossRef] [PubMed]
- Davies, A.; Zoubeidi, A.; Selth, L.A. The Epigenetic and Transcriptional Landscape of Neuroendocrine Prostate Cancer. Endocr.-Relat. Cancer 2020, 27, R35–R50. [Google Scholar] [CrossRef] [PubMed]
- Shi, M.; Wang, Y.; Lin, D.; Wang, Y. Patient-Derived Xenograft Models of Neuroendocrine Prostate Cancer. Cancer Lett. 2022, 525, 160–169. [Google Scholar] [CrossRef] [PubMed]
- Corella, A.N.; Cabiliza Ordonio, M.V.A.; Coleman, I.; Lucas, J.M.; Kaipainen, A.; Nguyen, H.M.; Sondheim, D.; Brown, L.G.; True, L.D.; Lee, J.K.; et al. Identification of Therapeutic Vulnerabilities in Small-Cell Neuroendocrine Prostate Cancer. Clin. Cancer Res. 2020, 26, 1667–1677. [Google Scholar] [CrossRef] [PubMed]
- Kaarijärvi, R.; Kaljunen, H.; Ketola, K. Molecular and Functional Links between Neurodevelopmental Processes and Treatment-Induced Neuroendocrine Plasticity in Prostate Cancer Progression. Cancers 2021, 13, 692. [Google Scholar] [CrossRef] [PubMed]
- Elia, I.; Haigis, M.C. Metabolites and the Tumour Microenvironment: From Cellular Mechanisms to Systemic Metabolism. Nat. Metab. 2021, 3, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Xu, J.; Liu, J.; Wang, N.; Zhou, L.; Guo, J. Liver Metastasis in Cancer: Molecular Mechanisms and Management. MedComm 2025, 6, e70119. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Zhang, G.; Liu, Y.; Liu, Y.; Tian, A.; Che, J.; Zhang, Z. Molecular Mechanisms and Targeted Therapy for the Metastasis of Prostate Cancer to the Bones (Review). Int. J. Oncol. 2024, 65, 104. [Google Scholar] [CrossRef] [PubMed]
- Ni, X.; Wei, Y.; Li, X.; Pan, J.; Fang, B.; Zhang, T.; Lu, Y.; Ye, D.; Zhu, Y. From Biology to the Clinic—Exploring Liver Metastasis in Prostate Cancer. Nat. Rev. Urol. 2024, 21, 593–614. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, H.M.; Vessella, R.L.; Morrissey, C.; Brown, L.G.; Coleman, I.M.; Higano, C.S.; Mostaghel, E.A.; Zhang, X.; True, L.D.; Lam, H.-M.; et al. LuCaP Prostate Cancer Patient-Derived Xenografts Reflect the Molecular Heterogeneity of Advanced Disease and Serve as Models for Evaluating Cancer Therapeutics: Prostate Cancer PDXs: LuCaP Series. Prostate 2017, 77, 654–671. [Google Scholar] [CrossRef] [PubMed]
- Lin, D.; Wyatt, A.W.; Xue, H.; Wang, Y.; Dong, X.; Haegert, A.; Wu, R.; Brahmbhatt, S.; Mo, F.; Jong, L.; et al. High Fidelity Patient-Derived Xenografts for Accelerating Prostate Cancer Discovery and Drug Development. Cancer Res. 2014, 74, 1272–1283. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, S.; Peehl, D.M.; Sriram, R. Subrenal Capsule Implantation of Tumor Tissue. 2021. Available online: https://www.protocols.io/view/subrenal-capsule-src-implantation-of-tumor-tissue-8epv5zk75v1b/v1 (accessed on 20 December 2021).
- Agarwal, S.; Peehl, D.M.; Sriram, R. Single Cell Digestion of Tumor Tissue. 2021. Available online: https://www.protocols.io/view/single-cell-digestion-of-tumor-tissue-kxygxp4kdl8j/v1 (accessed on 20 December 2021).
- Agarwal, S.; Peehl, D.M.; Sriram, R. Intratibial Implantation of Tumor Cells. 2021. Available online: https://www.protocols.io/view/intratibial-implantation-of-tumor-cells-261ge4kp7v47/v1 (accessed on 20 December 2021).
- Agarwal, S.; Bok, R.; Peehl, D.M.; Sriram, R. Intrahepatic Implantation of Tumor Cells. 2021. Available online: https://www.protocols.io/view/intrahepatic-implantation-of-tumor-cells-q26g78nzklwz/v1 (accessed on 20 December 2021).
- Rohrer, M.; Bauer, H.; Mintorovitch, J.; Requardt, M.; Weinmann, H.-J. Comparison of Magnetic Properties of MRI Contrast Media Solutions at Different Magnetic Field Strengths. Investig. Radiol. 2005, 40, 715–724. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, S.; Decavel-Bueff, E.; Wang, Y.-H.; Qin, H.; Santos, R.D.; Evans, M.J.; Sriram, R. Defining the Magnetic Resonance Features of Renal Lesions and Their Response to Everolimus in a Transgenic Mouse Model of Tuberous Sclerosis Complex. Front. Oncol. 2022, 12, 851192. [Google Scholar] [CrossRef] [PubMed]
- Agudelo, J.P.; Upadhyay, D.; Zhang, D.; Zhao, H.; Nolley, R.; Sun, J.; Agarwal, S.; Bok, R.A.; Vigneron, D.B.; Brooks, J.D.; et al. Multiparametric Magnetic Resonance Imaging and Metabolic Characterization of Patient-Derived Xenograft Models of Clear Cell Renal Cell Carcinoma. Metabolites 2022, 12, 1117. [Google Scholar] [CrossRef] [PubMed]
- Qin, H.; Zhang, V.; Bok, R.A.; Santos, R.D.; Cunha, J.A.; Hsu, I.-C.; Santos, J.D.; Lee, J.E.; Sukumar, S.; Larson, P.E.Z.; et al. Simultaneous Metabolic and Perfusion Imaging Using Hyperpolarized 13C MRI Can Evaluate Early and Dose-Dependent Response to Radiation Therapy in a Prostate Cancer Mouse Model. Int. J. Radiat. Oncol. Biol. Phys. 2020, 107, 887–896. [Google Scholar] [CrossRef] [PubMed]
- Crane, J.C.; Olson, M.P.; Nelson, S.J. SIVIC: Open-Source, Standards-Based Software for DICOM MR Spectroscopy Workflows. Int. J. Biomed. Imaging 2013, 2013, 169526. [Google Scholar] [CrossRef] [PubMed]
- Larson, P.E.Z.; Chen, H.-Y.; Gordon, J.W.; Korn, N.; Maidens, J.; Arcak, M.; Tang, S.; Criekinge, M.; Carvajal, L.; Mammoli, D.; et al. Investigation of Analysis Methods for Hyperpolarized 13C-Pyruvate Metabolic MRI in Prostate Cancer Patients. NMR Biomed. 2018, 31, e3997. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Bok, R.A.; DeLos Santos, J.; Upadhyay, D.; DeLos Santos, R.; Agarwal, S.; Van Criekinge, M.; Vigneron, D.B.; Aggarwal, R.; Peehl, D.M.; et al. Resistance to Androgen Deprivation Leads to Altered Metabolism in Human and Murine Prostate Cancer Cell and Tumor Models. Metabolites 2021, 11, 139. [Google Scholar] [CrossRef] [PubMed]
- Sriram, R.; Gordon, J.; Baligand, C.; Ahamed, F.; Delos Santos, J.; Qin, H.; Bok, R.; Vigneron, D.; Kurhanewicz, J.; Larson, P.; et al. Non-Invasive Assessment of Lactate Production and Compartmentalization in Renal Cell Carcinomas Using Hyperpolarized 13C Pyruvate MRI. Cancers 2018, 10, 313. [Google Scholar] [CrossRef] [PubMed]
- Bankhead, P.; Loughrey, M.B.; Fernández, J.A.; Dombrowski, Y.; McArt, D.G.; Dunne, P.D.; McQuaid, S.; Gray, R.T.; Murray, L.J.; Coleman, H.G.; et al. QuPath: Open Source Software for Digital Pathology Image Analysis. Sci. Rep. 2017, 7, 16878. [Google Scholar] [CrossRef] [PubMed]
- Loening, A.M.; Gambhir, S.S. AMIDE: A Free Software Tool for Multimodality Medical Image Analysis. Mol. Imaging 2003, 2, 153535002003031. [Google Scholar] [CrossRef] [PubMed]
- Labrecque, M.P.; Coleman, I.M.; Brown, L.G.; True, L.D.; Kollath, L.; Lakely, B.; Nguyen, H.M.; Yang, Y.C.; da Costa, R.M.G.; Kaipainen, A.; et al. Molecular Profiling Stratifies Diverse Phenotypes of Treatment-Refractory Metastatic Castration-Resistant Prostate Cancer. J. Clin. Investig. 2019, 129, 4492–4505. [Google Scholar] [CrossRef] [PubMed]
- Flores-Morales, A.; Bergmann, T.B.; Lavallee, C.; Batth, T.S.; Lin, D.; Lerdrup, M.; Friis, S.; Bartels, A.; Kristensen, G.; Krzyzanowska, A.; et al. Proteogenomic Characterization of Patient-Derived Xenografts Highlights the Role of REST in Neuroendocrine Differentiation of Castration-Resistant Prostate Cancer. Clin. Cancer Res. 2019, 25, 595–608. [Google Scholar] [CrossRef] [PubMed]
- Grashei, M.; Biechl, P.; Schilling, F.; Otto, A.M. Conversion of Hyperpolarized [1-13C]Pyruvate in Breast Cancer Cells Depends on Their Malignancy, Metabolic Program and Nutrient Microenvironment. Cancers 2022, 14, 1845. [Google Scholar] [CrossRef] [PubMed]
- Maleki, Z.; Nadella, A.; Nadella, M.; Patel, G.; Patel, S.; Kholová, I. INSM1, a Novel Biomarker for Detection of Neuroendocrine Neoplasms: Cytopathologists’ View. Diagnostics 2021, 11, 2172. [Google Scholar] [CrossRef] [PubMed]
- Xin, Z.; Zhang, Y.; Jiang, Z.; Zhao, L.; Fan, L.; Wang, Y.; Xie, S.; Shangguan, X.; Zhu, Y.; Pan, J.; et al. Insulinoma-Associated Protein 1 Is a Novel Sensitive and Specific Marker for Small Cell Carcinoma of the Prostate. Hum. Pathol. 2018, 79, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Park, J.W.; Lee, J.K.; Witte, O.N.; Huang, J. FOXA2 Is a Sensitive and Specific Marker for Small Cell Neuroendocrine Carcinoma of the Prostate. Mod. Pathol. 2017, 30, 1262–1272. [Google Scholar] [CrossRef] [PubMed]
- Go, S.; Kramer, T.T.; Verhoeven, A.J.; Oude Elferink, R.P.J.; Chang, J.-C. The Extracellular Lactate-to-Pyruvate Ratio Modulates the Sensitivity to Oxidative Stress-Induced Apoptosis via the Cytosolic NADH/NAD+ Redox State. Apoptosis 2021, 26, 38–51. [Google Scholar] [CrossRef] [PubMed]
- Rao, Y.; Gammon, S.; Zacharias, N.M.; Liu, T.; Salzillo, T.; Xi, Y.; Wang, J.; Bhattacharya, P.; Piwnica-Worms, D. Hyperpolarized [1-13C]Pyruvate-to-[1-13C]Lactate Conversion Is Rate-Limited by Monocarboxylate Transporter-1 in the Plasma Membrane. Proc. Natl. Acad. Sci. USA 2020, 117, 22378–22389. [Google Scholar] [CrossRef] [PubMed]
- Labrecque, M.P.; Brown, L.G.; Coleman, I.M.; Lakely, B.; Brady, N.J.; Lee, J.K.; Nguyen, H.M.; Li, D.; Hanratty, B.; Haffner, M.C.; et al. RNA Splicing Factors SRRM3 and SRRM4 Distinguish Molecular Phenotypes of Castration-Resistant Neuroendocrine Prostate Cancer. Cancer Res. 2021, 81, 4736–4750. [Google Scholar] [CrossRef] [PubMed]
- Cejas, P.; Xie, Y.; Font-Tello, A.; Lim, K.; Syamala, S.; Qiu, X.; Tewari, A.K.; Shah, N.; Nguyen, H.M.; Patel, R.A.; et al. Subtype Heterogeneity and Epigenetic Convergence in Neuroendocrine Prostate Cancer. Nat. Commun. 2021, 12, 5775. [Google Scholar] [CrossRef] [PubMed]
- Feng, E.; Rydzewski, N.R.; Zhang, M.; Lundberg, A.; Bootsma, M.; Helzer, K.T.; Lang, J.M.; Aggarwal, R.; Small, E.J.; Quigley, D.A.; et al. Intrinsic Molecular Subtypes of Metastatic Castration-Resistant Prostate Cancer. Clin. Cancer Res. 2022, 28, 5396–5404. [Google Scholar] [CrossRef] [PubMed]
- Vandergrift, L.A.; Decelle, E.A.; Kurth, J.; Wu, S.; Fuss, T.L.; DeFeo, E.M.; Halpern, E.F.; Taupitz, M.; McDougal, W.S.; Olumi, A.F.; et al. Metabolomic Prediction of Human Prostate Cancer Aggressiveness: Magnetic Resonance Spectroscopy of Histologically Benign Tissue. Sci. Rep. 2018, 8, 4997. [Google Scholar] [CrossRef] [PubMed]







Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Agarwal, S.; Upadhyay, D.; Sun, J.; Decavel-Bueff, E.; Bok, R.A.; Santos, R.D.; Al Muzhahimi, S.; Nolley, R.; Crane, J.; Kurhanewicz, J.; et al. Impact of Metastatic Microenvironment on Physiology and Metabolism of Small Cell Neuroendocrine Prostate Cancer Patient-Derived Xenografts. Cancers 2025, 17, 2385. https://doi.org/10.3390/cancers17142385
Agarwal S, Upadhyay D, Sun J, Decavel-Bueff E, Bok RA, Santos RD, Al Muzhahimi S, Nolley R, Crane J, Kurhanewicz J, et al. Impact of Metastatic Microenvironment on Physiology and Metabolism of Small Cell Neuroendocrine Prostate Cancer Patient-Derived Xenografts. Cancers. 2025; 17(14):2385. https://doi.org/10.3390/cancers17142385
Chicago/Turabian StyleAgarwal, Shubhangi, Deepti Upadhyay, Jinny Sun, Emilie Decavel-Bueff, Robert A. Bok, Romelyn Delos Santos, Said Al Muzhahimi, Rosalie Nolley, Jason Crane, John Kurhanewicz, and et al. 2025. "Impact of Metastatic Microenvironment on Physiology and Metabolism of Small Cell Neuroendocrine Prostate Cancer Patient-Derived Xenografts" Cancers 17, no. 14: 2385. https://doi.org/10.3390/cancers17142385
APA StyleAgarwal, S., Upadhyay, D., Sun, J., Decavel-Bueff, E., Bok, R. A., Santos, R. D., Al Muzhahimi, S., Nolley, R., Crane, J., Kurhanewicz, J., Peehl, D. M., & Sriram, R. (2025). Impact of Metastatic Microenvironment on Physiology and Metabolism of Small Cell Neuroendocrine Prostate Cancer Patient-Derived Xenografts. Cancers, 17(14), 2385. https://doi.org/10.3390/cancers17142385





