Elevated GRHL2 Imparts Plasticity in ER-Positive Breast Cancer Cells
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Generation of Tetracycline-Inducible GRHL2-GFP Expression in MCF7 Cell Line
2.3. Drug Treatment
2.4. RNA Isolation and RT-qPCR
2.5. RNA Sequencing
2.6. Protein Preparation and Western Blotting
2.7. Chromatin Immunoprecipitation (ChIP) and Sequencing
2.8. Transient Transfection
2.9. Migration Assay
2.10. Flow Cytometry
2.11. Immunocytochemistry and Immunohistrochemistry
2.12. Soft Agar Assay
2.13. Mammosphere Assay
2.14. Tumor Xenograft
2.15. Statistical Analysis
3. Results
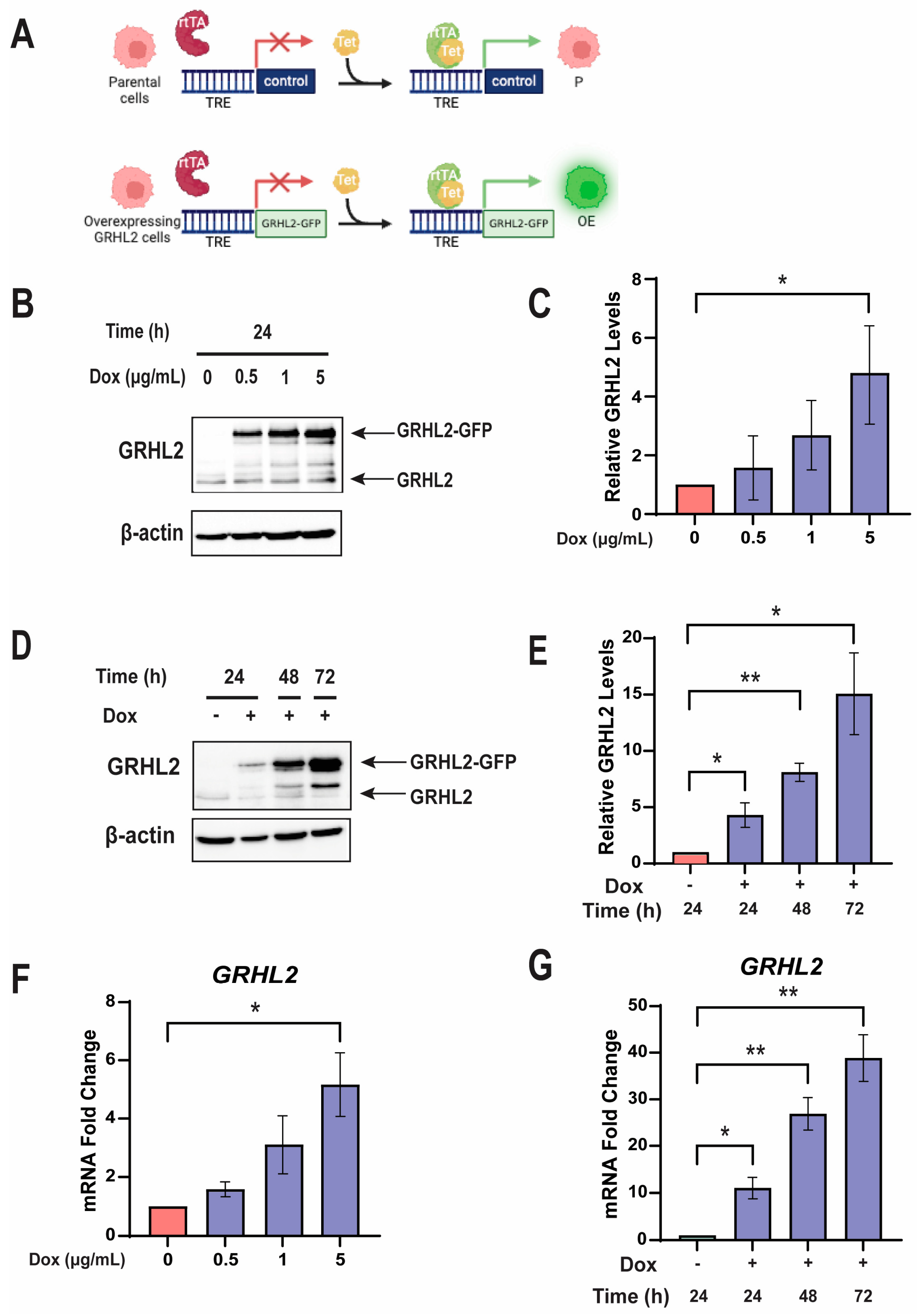
3.1. A Tetracycline-Inducible Model of High GRHL2 Expression in Breast Cancer Cells Expresses GRHL2 Protein and mRNA in a Dox Dose-Dependent Manner
3.2. High GRHL2 Expression Increases Epithelial Cell Phenotypes
3.3. GRHL2 Overexpression Alters Its Endogenous Transcriptional Activity and Gene Expression
3.4. GRHL2 Overexpression Regulates Growth and Developmental Pathways
3.5. GRHL2 Overexpression Inhibits Proliferation in Vivo
3.6. GRHL2 Overexpression Enriches Stem Cell-like Characteristics
3.7. GRHL2 Overexpression Leads to a Complex Epithelial–Mesenchymal Hybrid Phenotype
3.8. GRHL2 Overexpression Alters GRHL2 Genome Binding in a Dynamic Manner
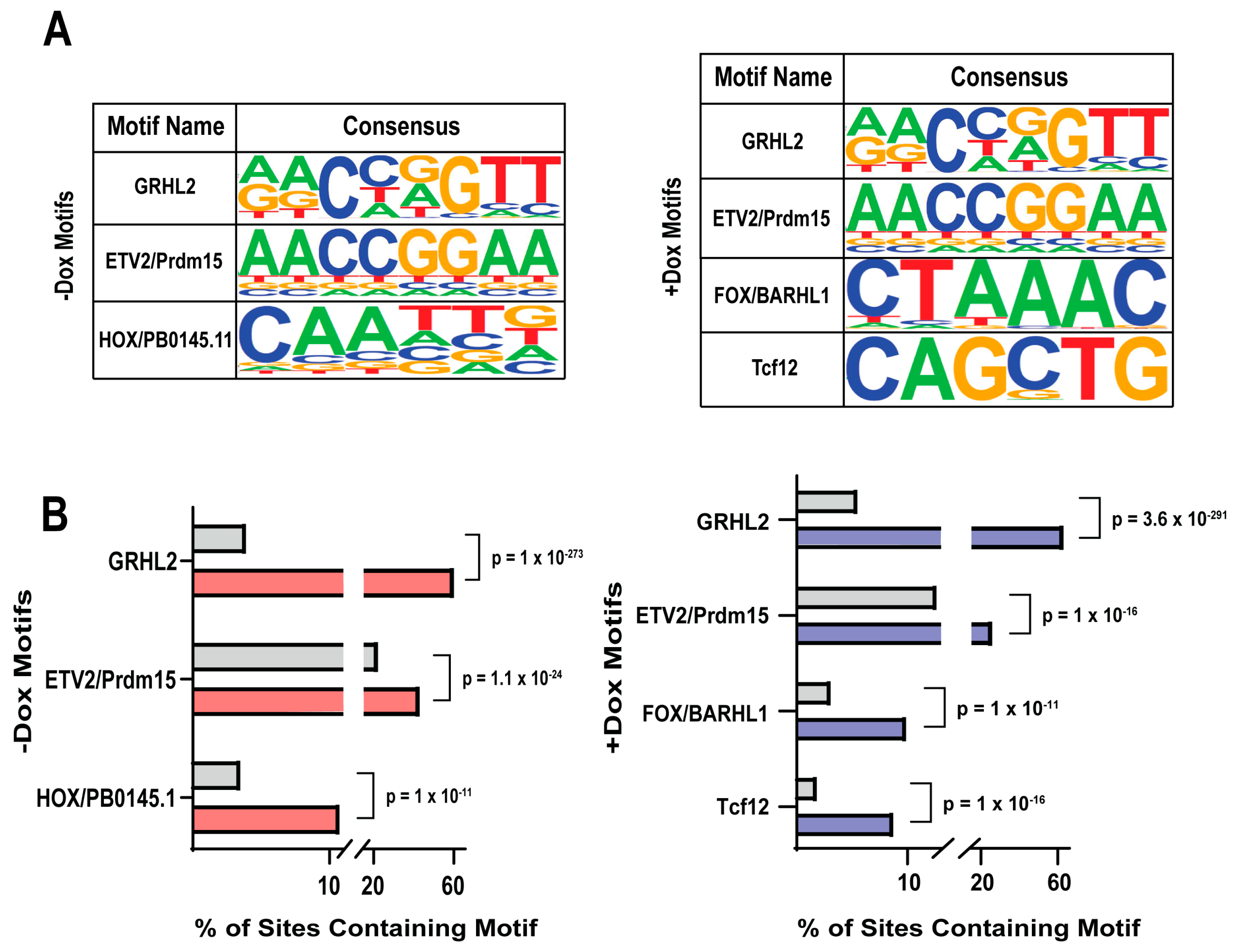
3.9. GRHL2 Overexpression Changes Motifs Found near GRHL2 Binding Sites
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Wagle, N.S.; Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 2023, 73, 17–48. [Google Scholar] [CrossRef] [PubMed]
- Giaquinto, A.N.; Sung, H.; Miller, K.D.; Kramer, J.L.; Newman, L.A.; Minihan, A.; Jemal, A.; Siegel, R.L. Breast Cancer Statistics, 2022. CA Cancer J. Clin. 2022, 72, 524–541. [Google Scholar] [CrossRef] [PubMed]
- Jayasekara, H.; MacInnis, R.J.; Chamberlain, J.A.; Dite, G.S.; Leoce, N.M.; Dowty, J.G.; Bickerstaffe, A.; Win, A.K.; Milne, R.L.; Giles, G.G.; et al. Mortality after breast cancer as a function of time since diagnosis by estrogen receptor status and age at diagnosis. Int. J. Cancer 2019, 145, 3207–3217. [Google Scholar] [CrossRef] [PubMed]
- Rueda, O.M.; Sammut, S.J.; Seoane, J.A.; Chin, S.F.; Caswell-Jin, J.L.; Callari, M.; Batra, R.; Pereira, B.; Bruna, A.; Ali, H.R.; et al. Dynamics of breast-cancer relapse reveal late-recurring ER-positive genomic subgroups. Nature 2019, 567, 399–404. [Google Scholar] [CrossRef]
- Pedersen, R.N.; Esen, B.; Mellemkjær, L.; Christiansen, P.; Ejlertsen, B.; Lash, T.L.; Nørgaard, M.; Cronin-Fenton, D. The Incidence of Breast Cancer Recurrence 10-32 Years After Primary Diagnosis. J. Natl. Cancer Inst. 2022, 114, 391–399. [Google Scholar] [CrossRef] [PubMed]
- Pan, H.; Gray, R.; Braybrooke, J.; Davies, C.; Taylor, C.; McGale, P.; Peto, R.; Pritchard, K.I.; Bergh, J.; Dowsett, M.; et al. 20-Year Risks of Breast-Cancer Recurrence after Stopping Endocrine Therapy at 5 Years. N. Engl. J. Med. 2017, 377, 1836–1846. [Google Scholar] [CrossRef]
- Reese, R.M.; Helzer, K.T.; Allen, K.O.; Zheng, C.; Solodin, N.; Alarid, E.T. GRHL2 Enhances Phosphorylated Estrogen Receptor (ER) Chromatin Binding and Regulates ER-Mediated Transcriptional Activation and Repression. Mol. Cell Biol. 2022, 42, e0019122. [Google Scholar] [CrossRef]
- Helzer, K.T.; Szatkowski Ozers, M.; Meyer, M.B.; Benkusky, N.A.; Solodin, N.; Reese, R.M.; Warren, C.L.; Pike, J.W.; Alarid, E.T. The Phosphorylated Estrogen Receptor α (ER) Cistrome Identifies a Subset of Active Enhancers Enriched for Direct ER-DNA Binding and the Transcription Factor GRHL2. Mol. Cell Biol. 2019, 39, e00417-18. [Google Scholar] [CrossRef] [PubMed]
- Reese, R.M.; Harrison, M.M.; Alarid, E.T. Grainyhead-like Protein 2: The Emerging Role in Hormone-Dependent Cancers and Epigenetics. Endocrinology 2019, 160, 1275–1288. [Google Scholar] [CrossRef]
- Hinze, C.; Ruffert, J.; Walentin, K.; Himmerkus, N.; Nikpey, E.; Tenstad, O.; Wiig, H.; Mutig, K.; Yurtdas, Z.Y.; Klein, J.D.; et al. GRHL2 Is Required for Collecting Duct Epithelial Barrier Function and Renal Osmoregulation. J. Am. Soc. Nephrol. 2018, 29, 857–868. [Google Scholar] [CrossRef]
- Mooney, S.M.; Talebian, V.; Jolly, M.K.; Jia, D.; Gromala, M.; Levine, H.; McConkey, B.J. The GRHL2/ZEB Feedback Loop-A Key Axis in the Regulation of EMT in Breast Cancer. J. Cell Biochem. 2017, 118, 2559–2570. [Google Scholar] [CrossRef] [PubMed]
- Crane-Smith, Z.; De Castro, S.C.P.; Nikolopoulou, E.; Wolujewicz, P.; Smedley, D.; Lei, Y.; Mather, E.; Santos, C.; Hopkinson, M.; Pitsillides, A.A.; et al. A non-coding insertional mutation of Grhl2 causes gene over-expression and multiple structural anomalies including cleft palate, spina bifida and encephalocele. Hum. Mol. Genet. 2023, 32, 2681–2692. [Google Scholar] [CrossRef]
- He, J.; Feng, C.; Zhu, H.; Wu, S.; Jin, P.; Xu, T. Grainyhead-like 2 as a double-edged sword in development and cancer. Am. J. Transl. Res. 2020, 12, 310–331. [Google Scholar]
- Werth, M.; Walentin, K.; Aue, A.; Schönheit, J.; Wuebken, A.; Pode-Shakked, N.; Vilianovitch, L.; Erdmann, B.; Dekel, B.; Bader, M.; et al. The transcription factor grainyhead-like 2 regulates the molecular composition of the epithelial apical junctional complex. Development 2010, 137, 3835–3845. [Google Scholar] [CrossRef]
- Cieply, B.; Farris, J.; Denvir, J.; Ford, H.L.; Frisch, S.M. Epithelial-mesenchymal transition and tumor suppression are controlled by a reciprocal feedback loop between ZEB1 and Grainyhead-like-2. Cancer Res. 2013, 73, 6299–6309. [Google Scholar] [CrossRef]
- Xiang, X.; Deng, Z.; Zhuang, X.; Ju, S.; Mu, J.; Jiang, H.; Zhang, L.; Yan, J.; Miller, D.; Zhang, H.G. Grhl2 determines the epithelial phenotype of breast cancers and promotes tumor progression. PLoS ONE 2012, 7, e50781. [Google Scholar] [CrossRef]
- Quan, Y.; Jin, R.; Huang, A.; Zhao, H.; Feng, B.; Zang, L.; Zheng, M. Downregulation of GRHL2 inhibits the proliferation of colorectal cancer cells by targeting ZEB1. Cancer Biol. Ther. 2014, 15, 878–887. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Wu, D.; Chen, Y.; Min, Z.; Quan, Y. GRHL2 inhibits colorectal cancer progression and metastasis via oppressing epithelial-mesenchymal transition. Cancer Biol. Ther. 2019, 20, 1195–1205. [Google Scholar] [CrossRef]
- Faddaoui, A.; Sheta, R.; Bachvarova, M.; Plante, M.; Gregoire, J.; Renaud, M.C.; Sebastianelli, A.; Gobeil, S.; Morin, C.; Ghani, K.; et al. Suppression of the grainyhead transcription factor 2 gene (GRHL2) inhibits the proliferation, migration, invasion and mediates cell cycle arrest of ovarian cancer cells. Cell Cycle 2017, 16, 693–706. [Google Scholar] [CrossRef]
- Holding, A.N.; Giorgi, F.M.; Donnelly, A.; Cullen, A.E.; Nagarajan, S.; Selth, L.A.; Markowetz, F. VULCAN integrates ChIP-seq with patient-derived co-expression networks to identify GRHL2 as a key co-regulator of ERa at enhancers in breast cancer. Genome Biol. 2019, 20, 91. [Google Scholar] [CrossRef]
- Riethdorf, S.; Frey, S.; Santjer, S.; Stoupiec, M.; Otto, B.; Riethdorf, L.; Koop, C.; Wilczak, W.; Simon, R.; Sauter, G.; et al. Diverse expression patterns of the EMT suppressor grainyhead-like 2 (GRHL2) in normal and tumour tissues. Int. J. Cancer 2016, 138, 949–963. [Google Scholar] [CrossRef]
- Chi, D.; Singhal, H.; Li, L.; Xiao, T.; Liu, W.; Pun, M.; Jeselsohn, R.; He, H.; Lim, E.; Vadhi, R.; et al. Estrogen receptor signaling is reprogrammed during breast tumorigenesis. Proc. Natl. Acad. Sci. USA 2019, 116, 11437–11443. [Google Scholar] [CrossRef]
- Dompe, N.; Rivers, C.S.; Li, L.; Cordes, S.; Schwickart, M.; Punnoose, E.A.; Amler, L.; Seshagiri, S.; Tang, J.; Modrusan, Z.; et al. A whole-genome RNAi screen identifies an 8q22 gene cluster that inhibits death receptor-mediated apoptosis. Proc. Natl. Acad. Sci. USA 2011, 108, E943–E951. [Google Scholar] [CrossRef]
- Cocce, K.J.; Jasper, J.S.; Desautels, T.K.; Everett, L.; Wardell, S.; Westerling, T.; Baldi, R.; Wright, T.M.; Tavares, K.; Yllanes, A.; et al. The Lineage Determining Factor GRHL2 Collaborates with FOXA1 to Establish a Targetable Pathway in Endocrine Therapy-Resistant Breast Cancer. Cell Rep. 2019, 29, 889–903.e10. [Google Scholar] [CrossRef]
- Wang, Z.; Coban, B.; Liao, C.Y.; Chen, Y.J.; Liu, Q.; Danen, E.H.J. GRHL2 Regulation of Growth/Motility Balance in Luminal versus Basal Breast Cancer. Int. J. Mol. Sci. 2023, 24, 2512. [Google Scholar] [CrossRef]
- Bai, X.; Li, Y.; Li, F.; Che, N.; Ni, C.; Zhao, N.; Zhao, X.; Liu, T. GRHL2 Expression Functions in Breast Cancer Aggressiveness and Could Serve as Prognostic and Diagnostic Biomarker for Breast Cancer. Clin. Med. Insights Oncol. 2022, 16, 11795549221109511. [Google Scholar] [CrossRef]
- Fowler, A.M.; Solodin, N.; Preisler-Mashek, M.T.; Zhang, P.; Lee, A.V.; Alarid, E.T. Increases in estrogen receptor-alpha concentration in breast cancer cells promote serine 118/104/106-independent AF-1 transactivation and growth in the absence of estrogen. FASEB J. 2004, 18, 81–93. [Google Scholar] [CrossRef]
- Gossen, M.; Freundlieb, S.; Bender, G.; Muller, G.; Hillen, W.; Bujard, H. Transcriptional activation by tetracyclines in mammalian cells. Science 1995, 268, 1766–1769. [Google Scholar] [CrossRef] [PubMed]
- Meyer, M.B.; Benkusky, N.A.; Lee, S.M.; Yoon, S.H.; Mannstadt, M.; Wein, M.N.; Pike, J.W. Rapid genomic changes by mineralotropic hormones and kinase SIK inhibition drive coordinated renal Cyp27b1 and Cyp24a1 expression via CREB modules. J. Biol. Chem. 2022, 298, 102559. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef]
- Heinz, S.; Benner, C.; Spann, N.; Bertolino, E.; Lin, Y.C.; Laslo, P.; Cheng, J.X.; Murre, C.; Singh, H.; Glass, C.K. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 2010, 38, 576–589. [Google Scholar] [CrossRef]
- Suarez-Arnedo, A.; Torres Figueroa, F.; Clavijo, C.; Arbeláez, P.; Cruz, J.C.; Muñoz-Camargo, C. An image J plugin for the high throughput image analysis of in vitro scratch wound healing assays. PLoS ONE 2020, 15, e0232565. [Google Scholar] [CrossRef]
- Powers, G.L.; Ellison-Zelski, S.J.; Casa, A.J.; Lee, A.V.; Alarid, E.T. Proteasome inhibition represses ERalpha gene expression in ER+ cells: A new link between proteasome activity and estrogen signaling in breast cancer. Oncogene 2010, 29, 1509–1518. [Google Scholar] [CrossRef]
- Kumar, M.; Salem, K.; Jeffery, J.J.; Yan, Y.; Mahajan, A.M.; Fowler, A.M. Longitudinal Molecular Imaging of Progesterone Receptor Reveals Early Differential Response to Endocrine Therapy in Breast Cancer with an Activating ESR1 Mutation. J. Nucl. Med. 2021, 62, 500–506. [Google Scholar] [CrossRef] [PubMed]
- Ahler, E.; Sullivan, W.J.; Cass, A.; Braas, D.; York, A.G.; Bensinger, S.J.; Graeber, T.G.; Christofk, H.R. Doxycycline alters metabolism and proliferation of human cell lines. PLoS ONE 2013, 8, e64561. [Google Scholar] [CrossRef]
- Chen, Y.F.; Yang, Y.N.; Chu, H.R.; Huang, T.Y.; Wang, S.H.; Chen, H.Y.; Li, Z.L.; Yang, Y.S.H.; Lin, H.Y.; Hercbergs, A.; et al. Role of Integrin αvβ3 in Doxycycline-Induced Anti-Proliferation in Breast Cancer Cells. Front. Cell Dev. Biol. 2022, 10, 829788. [Google Scholar] [CrossRef]
- Narasimha, M.; Uv, A.; Krejci, A.; Brown, N.H.; Bray, S.J. Grainy head promotes expression of septate junction proteins and influences epithelial morphogenesis. J. Cell Sci. 2008, 121, 747–752. [Google Scholar] [CrossRef]
- Cristo, I.; Carvalho, L.; Ponte, S.; Jacinto, A. Novel role for Grainy head in the regulation of cytoskeletal and junctional dynamics during epithelial repair. J. Cell Sci. 2018, 131, jcs213595. [Google Scholar] [CrossRef] [PubMed]
- Tunggal, J.A.; Helfrich, I.; Schmitz, A.; Schwarz, H.; Günzel, D.; Fromm, M.; Kemler, R.; Krieg, T.; Niessen, C.M. E-cadherin is essential for in vivo epidermal barrier function by regulating tight junctions. EMBO J. 2005, 24, 1146–1156. [Google Scholar] [CrossRef]
- Krause, G.; Winkler, L.; Mueller, S.L.; Haseloff, R.F.; Piontek, J.; Blasig, I.E. Structure and function of claudins. Biochim. Biophys. Acta 2008, 1778, 631–645. [Google Scholar] [CrossRef]
- Theodorou, V.; Stark, R.; Menon, S.; Carroll, J.S. GATA3 acts upstream of FOXA1 in mediating ESR1 binding by shaping enhancer accessibility. Genome Res. 2013, 23, 12–22. [Google Scholar] [CrossRef] [PubMed]
- Thiery, J.P. Epithelial-mesenchymal transitions in tumour progression. Nat. Rev. Cancer 2002, 2, 442–454. [Google Scholar] [CrossRef]
- Yang, J.; Weinberg, R.A. Epithelial-mesenchymal transition: At the crossroads of development and tumor metastasis. Dev. Cell 2008, 14, 818–829. [Google Scholar] [CrossRef]
- May, C.D.; Sphyris, N.; Evans, K.W.; Werden, S.J.; Guo, W.; Mani, S.A. Epithelial-mesenchymal transition and cancer stem cells: A dangerously dynamic duo in breast cancer progression. Breast Cancer Res. 2011, 13, 202. [Google Scholar] [CrossRef] [PubMed]
- Puisieux, A.; Brabletz, T.; Caramel, J. Oncogenic roles of EMT-inducing transcription factors. Nat. Cell Biol. 2014, 16, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Chung, V.Y.; Tan, T.Z.; Ye, J.; Huang, R.L.; Lai, H.C.; Kappei, D.; Wollmann, H.; Guccione, E.; Huang, R.Y. The role of GRHL2 and epigenetic remodeling in epithelial-mesenchymal plasticity in ovarian cancer cells. Commun. Biol. 2019, 2, 272. [Google Scholar] [CrossRef]
- Kawabe, N.; Matsuoka, K.; Komeda, K.; Muraki, N.; Takaba, M.; Togami, Y.; Ito, Y.; Yamada, M.; Sunaga, N.; Girard, L.; et al. Silencing of GRHL2 induces epithelial-to-mesenchymal transition in lung cancer cell lines with different effects on proliferation and clonogenic growth. Oncol. Lett. 2023, 26, 391. [Google Scholar] [CrossRef]
- Li, H.; Zhang, W.; Liu, Y.; Cai, Z.; Lan, A.; Shu, D.; Shen, M.; Li, K.; Pu, D.; Tan, W.; et al. UTRN as a potential biomarker in breast cancer: A comprehensive bioinformatics and in vitro study. Sci. Rep. 2024, 14, 7702. [Google Scholar] [CrossRef]
- Kikuchi, K.; Tsukamoto, H. Stearoyl-CoA desaturase and tumorigenesis. Chem. Biol. Interact. 2020, 316, 108917. [Google Scholar] [CrossRef]
- Park, J.; Tacam, M.J.; Chauhan, G.; Cohen, E.N.; Gagliardi, M.; Iles, L.R.; Ueno, N.T.; Battula, V.L.; Sohn, Y.K.; Wang, X.; et al. Nonphosphorylatable PEA15 mutant inhibits epithelial-mesenchymal transition in triple-negative breast cancer partly through the regulation of IL-8 expression. Breast Cancer Res. Treat. 2021, 189, 333–345. [Google Scholar] [CrossRef]
- Bartholomeusz, C.; Rosen, D.; Wei, C.; Kazansky, A.; Yamasaki, F.; Takahashi, T.; Itamochi, H.; Kondo, S.; Liu, J.; Ueno, N.T. PEA-15 induces autophagy in human ovarian cancer cells and is associated with prolonged overall survival. Cancer Res. 2008, 68, 9302–9310. [Google Scholar] [CrossRef] [PubMed]
- Bartholomeusz, C.; Gonzalez-Angulo, A.M.; Kazansky, A.; Krishnamurthy, S.; Liu, P.; Yuan, L.X.; Yamasaki, F.; Liu, S.; Hayashi, N.; Zhang, D.; et al. PEA-15 inhibits tumorigenesis in an MDA-MB-468 triple-negative breast cancer xenograft model through increased cytoplasmic localization of activated extracellular signal-regulated kinase. Clin. Cancer Res. 2010, 16, 1802–1811. [Google Scholar] [CrossRef] [PubMed]
- Hindupur, S.K.; Balaji, S.A.; Saxena, M.; Pandey, S.; Sravan, G.S.; Heda, N.; Kumar, M.V.; Mukherjee, G.; Dey, D.; Rangarajan, A. Identification of a novel AMPK-PEA15 axis in the anoikis-resistant growth of mammary cells. Breast Cancer Res. 2014, 16, 420. [Google Scholar] [CrossRef] [PubMed]
- Sulzmaier, F.; Opoku-Ansah, J.; Ramos, J.W. Phosphorylation is the switch that turns PEA-15 from tumor suppressor to tumor promoter. Small GTPases 2012, 3, 173–177. [Google Scholar] [CrossRef] [PubMed]
- Nobre, A.R.; Dalla, E.; Yang, J.; Huang, X.; Wullkopf, L.; Risson, E.; Razghandi, P.; Anton, M.L.; Zheng, W.; Seoane, J.A.; et al. ZFP281 drives a mesenchymal-like dormancy program in early disseminated breast cancer cells that prevents metastatic outgrowth in the lung. Nat. Cancer 2022, 3, 1165–1180. [Google Scholar] [CrossRef] [PubMed]
- Wu, R.; Roy, A.M.; Tokumaru, Y.; Gandhi, S.; Asaoka, M.; Oshi, M.; Yan, L.; Ishikawa, T.; Takabe, K. NR2F1, a Tumor Dormancy Marker, Is Expressed Predominantly in Cancer-Associated Fibroblasts and Is Associated with Suppressed Breast Cancer Cell Proliferation. Cancers 2022, 14, 2962. [Google Scholar] [CrossRef]
- Montagner, M.; Sahai, E. In vitro Models of Breast Cancer Metastatic Dormancy. Front. Cell Dev. Biol. 2020, 8, 37. [Google Scholar] [CrossRef]
- Borgen, E.; Rypdal, M.C.; Sosa, M.S.; Renolen, A.; Schlichting, E.; Lønning, P.E.; Synnestvedt, M.; Aguirre-Ghiso, J.A.; Naume, B. NR2F1 stratifies dormant disseminated tumor cells in breast cancer patients. Breast Cancer Res. 2018, 20, 120. [Google Scholar] [CrossRef]
- Truskowski, K.; Amend, S.R.; Pienta, K.J. Dormant cancer cells: Programmed quiescence, senescence, or both? Cancer Metastasis Rev. 2023, 42, 37–47. [Google Scholar] [CrossRef]
- Marescal, O.; Cheeseman, I.M. Cellular Mechanisms and Regulation of Quiescence. Dev. Cell 2020, 55, 259–271. [Google Scholar] [CrossRef]
- Morrison, S.J.; Shah, N.M.; Anderson, D.J. Regulatory mechanisms in stem cell biology. Cell 1997, 88, 287–298. [Google Scholar] [CrossRef] [PubMed]
- Dontu, G.; Abdallah, W.M.; Foley, J.M.; Jackson, K.W.; Clarke, M.F.; Kawamura, M.J.; Wicha, M.S. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev. 2003, 17, 1253–1270. [Google Scholar] [CrossRef]
- Senbanjo, L.T.; Chellaiah, M.A. CD44: A Multifunctional Cell Surface Adhesion Receptor Is a Regulator of Progression and Metastasis of Cancer Cells. Front. Cell Dev. Biol. 2017, 5, 18. [Google Scholar] [CrossRef]
- Vadhan, A.; Hou, M.F.; Vijayaraghavan, P.; Wu, Y.C.; Hu, S.C.; Wang, Y.M.; Cheng, T.L.; Wang, Y.Y.; Yuan, S.F. CD44 Promotes Breast Cancer Metastasis through AKT-Mediated Downregulation of Nuclear FOXA2. Biomedicines 2022, 10, 2488. [Google Scholar] [CrossRef] [PubMed]
- Jaggupilli, A.; Elkord, E. Significance of CD44 and CD24 as cancer stem cell markers: An enduring ambiguity. Clin. Dev. Immunol. 2012, 2012, 708036. [Google Scholar] [CrossRef]
- Tomita, H.; Tanaka, K.; Tanaka, T.; Hara, A. Aldehyde dehydrogenase 1A1 in stem cells and cancer. Oncotarget 2016, 7, 11018–11032. [Google Scholar] [CrossRef] [PubMed]
- Grosse-Wilde, A.; Fouquier d’Hérouël, A.; McIntosh, E.; Ertaylan, G.; Skupin, A.; Kuestner, R.E.; del Sol, A.; Walters, K.A.; Huang, S. Stemness of the hybrid Epithelial/Mesenchymal State in Breast Cancer and Its Association with Poor Survival. PLoS ONE 2015, 10, e0126522. [Google Scholar] [CrossRef]
- Pradella, D.; Naro, C.; Sette, C.; Ghigna, C. EMT and stemness: Flexible processes tuned by alternative splicing in development and cancer progression. Mol. Cancer 2017, 16, 8. [Google Scholar] [CrossRef]
- Harper, K.L.; Sosa, M.S.; Entenberg, D.; Hosseini, H.; Cheung, J.F.; Nobre, R.; Avivar-Valderas, A.; Nagi, C.; Girnius, N.; Davis, R.J.; et al. Mechanism of early dissemination and metastasis in Her2(+) mammary cancer. Nature 2016, 540, 588–592. [Google Scholar] [CrossRef]
- Gong, W.; Das, S.; Sierra-Pagan, J.E.; Skie, E.; Dsouza, N.; Larson, T.A.; Garry, M.G.; Luzete-Monteiro, E.; Zaret, K.S.; Garry, D.J. ETV2 functions as a pioneer factor to regulate and reprogram the endothelial lineage. Nat. Cell Biol. 2022, 24, 672–684. [Google Scholar] [CrossRef]
- Seifert, A.; Werheid, D.F.; Knapp, S.M.; Tobiasch, E. Role of Hox genes in stem cell differentiation. World J. Stem Cells 2015, 7, 583–595. [Google Scholar] [CrossRef]
- Castaneda, M.; Hollander, P.D.; Mani, S.A. Forkhead Box Transcription Factors: Double-Edged Swords in Cancer. Cancer Res. 2022, 82, 2057–2065. [Google Scholar] [CrossRef]
- Tang, X.; Hou, Y.; Yang, G.; Wang, X.; Tang, S.; Du, Y.E.; Yang, L.; Yu, T.; Zhang, H.; Zhou, M.; et al. Stromal miR-200s contribute to breast cancer cell invasion through CAF activation and ECM remodeling. Cell Death Differ. 2016, 23, 132–145. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.; Jeselsohn, R.; Pereira, R.; Hollingsworth, E.F.; Creighton, C.J.; Li, F.; Shea, M.; Nardone, A.; De Angelis, C.; Heiser, L.M.; et al. FOXA1 overexpression mediates endocrine resistance by altering the ER transcriptome and IL-8 expression in ER-positive breast cancer. Proc. Natl. Acad. Sci. USA 2016, 113, E6600–E6609. [Google Scholar] [CrossRef] [PubMed]
- Werner, S.; Frey, S.; Riethdorf, S.; Schulze, C.; Alawi, M.; Kling, L.; Vafaizadeh, V.; Sauter, G.; Terracciano, L.; Schumacher, U.; et al. Dual roles of the transcription factor grainyhead-like 2 (GRHL2) in breast cancer. J. Biol. Chem. 2013, 288, 22993–23008. [Google Scholar] [CrossRef]
- Paltoglou, S.; Das, R.; Townley, S.L.; Hickey, T.E.; Tarulli, G.A.; Coutinho, I.; Fernandes, R.; Hanson, A.R.; Denis, I.; Carroll, J.S.; et al. Novel Androgen Receptor Coregulator GRHL2 Exerts Both Oncogenic and Antimetastatic Functions in Prostate Cancer. Cancer Res. 2017, 77, 3417–3430. [Google Scholar] [CrossRef] [PubMed]
- Nishino, H.; Takano, S.; Yoshitomi, H.; Suzuki, K.; Kagawa, S.; Shimazaki, R.; Shimizu, H.; Furukawa, K.; Miyazaki, M.; Ohtsuka, M. Grainyhead-like 2 (GRHL2) regulates epithelial plasticity in pancreatic cancer progression. Cancer Med. 2017, 6, 2686–2696. [Google Scholar] [CrossRef]
- Jolly, M.K.; Somarelli, J.A.; Sheth, M.; Biddle, A.; Tripathi, S.C.; Armstrong, A.J.; Hanash, S.M.; Bapat, S.A.; Rangarajan, A.; Levine, H. Hybrid epithelial/mesenchymal phenotypes promote metastasis and therapy resistance across carcinomas. Pharmacol. Ther. 2019, 194, 161–184. [Google Scholar] [CrossRef]
- Jolly, M.K.; Tripathi, S.C.; Jia, D.; Mooney, S.M.; Celiktas, M.; Hanash, S.M.; Mani, S.A.; Pienta, K.J.; Ben-Jacob, E.; Levine, H. Stability of the hybrid epithelial/mesenchymal phenotype. Oncotarget 2016, 7, 27067–27084. [Google Scholar] [CrossRef]
- Jolly, M.K.; Boareto, M.; Huang, B.; Jia, D.; Lu, M.; Ben-Jacob, E.; Onuchic, J.N.; Levine, H. Implications of the Hybrid Epithelial/Mesenchymal Phenotype in Metastasis. Front. Oncol. 2015, 5, 155. [Google Scholar] [CrossRef]
- Tam, W.L.; Weinberg, R.A. The epigenetics of epithelial-mesenchymal plasticity in cancer. Nat. Med. 2013, 19, 1438–1449. [Google Scholar] [CrossRef]
- Sinha, D.; Saha, P.; Samanta, A.; Bishayee, A. Emerging Concepts of Hybrid Epithelial-to-Mesenchymal Transition in Cancer Progression. Biomolecules 2020, 10, 1561. [Google Scholar] [CrossRef]
- Brabletz, S.; Schuhwerk, H.; Brabletz, T.; Stemmler, M.P. Dynamic EMT: A multi-tool for tumor progression. EMBO J. 2021, 40, e108647. [Google Scholar] [CrossRef] [PubMed]
- Pastushenko, I.; Blanpain, C. EMT Transition States during Tumor Progression and Metastasis. Trends Cell Biol. 2019, 29, 212–226. [Google Scholar] [CrossRef]
- Morel, A.P.; Hinkal, G.W.; Thomas, C.; Fauvet, F.; Courtois-Cox, S.; Wierinckx, A.; Devouassoux-Shisheboran, M.; Treilleux, I.; Tissier, A.; Gras, B.; et al. EMT inducers catalyze malignant transformation of mammary epithelial cells and drive tumorigenesis towards claudin-low tumors in transgenic mice. PLoS Genet. 2012, 8, e1002723. [Google Scholar] [CrossRef]
- Kröger, C.; Afeyan, A.; Mraz, J.; Eaton, E.N.; Reinhardt, F.; Khodor, Y.L.; Thiru, P.; Bierie, B.; Ye, X.; Burge, C.B.; et al. Acquisition of a hybrid E/M state is essential for tumorigenicity of basal breast cancer cells. Proc. Natl. Acad. Sci. USA 2019, 116, 7353–7362. [Google Scholar] [CrossRef] [PubMed]
- Fischer, K.R.; Durrans, A.; Lee, S.; Sheng, J.; Li, F.; Wong, S.T.; Choi, H.; El Rayes, T.; Ryu, S.; Troeger, J.; et al. Epithelial-to-mesenchymal transition is not required for lung metastasis but contributes to chemoresistance. Nature 2015, 527, 472–476. [Google Scholar] [CrossRef]
- Catalanotto, M.; Vaz, J.M.; Abshire, C.; Youngblood, R.; Chu, M.; Levine, H.; Jolly, M.K.; Dragoi, A.M. Dual role of CASP8AP2/FLASH in regulating epithelial-to-mesenchymal transition plasticity (EMP). Transl. Oncol. 2024, 39, 101837. [Google Scholar] [CrossRef] [PubMed]
- Aouad, P.; Zhang, Y.; De Martino, F.; Stibolt, C.; Ali, S.; Ambrosini, G.; Mani, S.A.; Maggs, K.; Quinn, H.M.; Sflomos, G.; et al. Epithelial-mesenchymal plasticity determines estrogen receptor positive breast cancer dormancy and epithelial reconversion drives recurrence. Nat. Commun. 2022, 13, 4975. [Google Scholar] [CrossRef]
- Gooding, A.J.; Schiemann, W.P. Epithelial-Mesenchymal Transition Programs and Cancer Stem Cell Phenotypes: Mediators of Breast Cancer Therapy Resistance. Mol. Cancer Res. 2020, 18, 1257–1270. [Google Scholar] [CrossRef]
- Prasetyanti, P.R.; Medema, J.P. Intra-tumor heterogeneity from a cancer stem cell perspective. Mol. Cancer 2017, 16, 41. [Google Scholar] [CrossRef]
- Tedja, R.; Alvero, A.B.; Fox, A.; Cardenas, C.; Pitruzzello, M.; Chehade, H.; Bawa, T.; Adzibolosu, N.; Gogoi, R.; Mor, G. Generation of Stable Epithelial-Mesenchymal Hybrid Cancer Cells with Tumorigenic Potential. Cancers 2023, 15, 684. [Google Scholar] [CrossRef]
- Mani, S.A.; Guo, W.; Liao, M.J.; Eaton, E.N.; Ayyanan, A.; Zhou, A.Y.; Brooks, M.; Reinhard, F.; Zhang, C.C.; Shipitsin, M.; et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008, 133, 704–715. [Google Scholar] [CrossRef] [PubMed]
- Althobiti, M.; El Ansari, R.; Aleskandarany, M.; Joseph, C.; Toss, M.S.; Green, A.R.; Rakha, E.A. The prognostic significance of ALDH1A1 expression in early invasive breast cancer. Histopathology 2020, 77, 437–448. [Google Scholar] [CrossRef] [PubMed]
- Fluegen, G.; Avivar-Valderas, A.; Wang, Y.; Padgen, M.R.; Williams, J.K.; Nobre, A.R.; Calvo, V.; Cheung, J.F.; Bravo-Cordero, J.J.; Entenberg, D.; et al. Phenotypic heterogeneity of disseminated tumour cells is preset by primary tumour hypoxic microenvironments. Nat. Cell Biol. 2017, 19, 120–132. [Google Scholar] [CrossRef]
- Kim, R.S.; Avivar-Valderas, A.; Estrada, Y.; Bragado, P.; Sosa, M.S.; Aguirre-Ghiso, J.A.; Segall, J.E. Dormancy signatures and metastasis in estrogen receptor positive and negative breast cancer. PLoS ONE 2012, 7, e35569. [Google Scholar] [CrossRef]
- La, T.; Chen, S.; Guo, T.; Zhao, X.H.; Teng, L.; Li, D.; Carnell, M.; Zhang, Y.Y.; Feng, Y.C.; Cole, N.; et al. Visualization of endogenous p27 and Ki67 reveals the importance of a c-Myc-driven metabolic switch in promoting survival of quiescent cancer cells. Theranostics 2021, 11, 9605–9622. [Google Scholar] [CrossRef]
- Sanchez Calle, A.; Yamamoto, T.; Kawamura, Y.; Hironaka-Mitsuhashi, A.; Ono, M.; Tsuda, H.; Shimomura, A.; Tamura, K.; Takeshita, F.; Ochiya, T.; et al. Long non-coding NR2F1-AS1 is associated with tumor recurrence in estrogen receptor-positive breast cancers. Mol. Oncol. 2020, 14, 2271–2287. [Google Scholar] [CrossRef]
- Liukkonen, S.; Leidenius, M.; Saarto, T.; Sjöström-Mattson, J. Breast cancer in very young women. Eur. J. Surg. Oncol. 2011, 37, 1030–1037. [Google Scholar] [CrossRef]
- Llinas-Bertran, A.; Bellet-Ezquerra, M.; Seoane, J.A. Epigenetic Control of Cancer Cell Dormancy and Awakening in Endocrine Therapy Resistance. Cancer Discov. 2024, 14, 704–706. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.L.; Zhang, M.; Tang, Y.L.; Liang, X.H. Cancer cell dormancy: Mechanisms and implications of cancer recurrence and metastasis. OncoTargets Ther. 2017, 10, 5219–5228. [Google Scholar] [CrossRef]
- Giancotti, F.G. Mechanisms governing metastatic dormancy and reactivation. Cell 2013, 155, 750–764. [Google Scholar] [CrossRef] [PubMed]
- Weidenfeld, K.; Barkan, D. EMT and Stemness in Tumor Dormancy and Outgrowth: Are They Intertwined Processes? Front. Oncol. 2018, 8, 381. [Google Scholar] [CrossRef]
- Wang, Z.; Coban, B.; Wu, H.; Chouaref, J.; Daxinger, L.; Paulsen, M.T.; Ljungman, M.; Smid, M.; Martens, J.W.M.; Danen, E.H.J. GRHL2-controlled gene expression networks in luminal breast cancer. Cell Commun. Signal 2023, 21, 15. [Google Scholar] [CrossRef] [PubMed]
- Fowler, A.M.; Solodin, N.M.; Valley, C.C.; Alarid, E.T. Altered target gene regulation controlled by estrogen receptor-alpha concentration. Mol. Endocrinol. 2006, 20, 291–301. [Google Scholar] [CrossRef]
- Struhl, G.; Struhl, K.; Macdonald, P.M. The gradient morphogen bicoid is a concentration-dependent transcriptional activator. Cell 1989, 57, 1259–1273. [Google Scholar] [PubMed]
- Jiang, J.; Kosman, D.; Ip, Y.T.; Levine, M. The dorsal morphogen gradient regulates the mesoderm determinant twist in early Drosophila embryos. Genes Dev. 1991, 5, 1881–1891. [Google Scholar] [PubMed]
- Harbison, C.T.; Gordon, D.B.; Lee, T.I.; Rinaldi, N.J.; Macisaac, K.D.; Danford, T.W.; Hannett, N.M.; Tagne, J.-B.; Reynolds, D.B.; Yoo, J.; et al. Transcriptional regulatory code of a eukaryotic genome. Nature 2004, 431, 99–104. [Google Scholar]
- Sipeky, C.; Tammela, T.L.J.; Auvinen, A.; Schleutker, J. Novel prostate cancer susceptibility gene SP6 predisposes patients to aggressive disease. Prostate Cancer Prostatic Dis. 2021, 24, 1158–1166. [Google Scholar] [CrossRef]
- Jacobs, J.; Atkins, M.; Davie, K.; Imrichova, H.; Romanelli, L.; Christiaens, V.; Hulselmans, G.; Potier, D.; Wouters, J.; Taskiran, I.I.; et al. The transcription factor Grainy head primes epithelial enhancers for spatiotemporal activation by displacing nucleosomes. Nat. Genet. 2018, 50, 1011–1020. [Google Scholar] [CrossRef] [PubMed]
- Chen, A.F.; Liu, A.J.; Krishnakumar, R.; Freimer, J.W.; DeVeale, B.; Blelloch, R. GRHL2-Dependent Enhancer Switching Maintains a Pluripotent Stem Cell Transcriptional Subnetwork after Exit from Naive Pluripotency. Cell Stem Cell 2018, 23, 226–238.e4. [Google Scholar] [CrossRef] [PubMed]
- Potier, D.; Davie, K.; Hulselmans, G.; Naval Sanchez, M.; Haagen, L.; Huynh-Thu, V.A.; Koldere, D.; Celik, A.; Geurts, P.; Christiaens, V.; et al. Mapping gene regulatory networks in Drosophila eye development by large-scale transcriptome perturbations and motif inference. Cell Rep. 2014, 9, 2290–2303. [Google Scholar] [CrossRef] [PubMed]
- Gibson, T.J.; Larson, E.D.; Harrison, M.M. Protein-intrinsic properties and context-dependent effects regulate pioneer factor binding and function. Nat. Struct. Mol. Biol. 2024, 31, 548–558. [Google Scholar] [CrossRef] [PubMed]
- Kumegawa, K.; Takahashi, Y.; Saeki, S.; Yang, L.; Nakadai, T.; Osako, T.; Mori, S.; Noda, T.; Ohno, S.; Ueno, T.; et al. GRHL2 motif is associated with intratumor heterogeneity of cis-regulatory elements in luminal breast cancer. NPJ Breast Cancer 2022, 8, 70. [Google Scholar] [CrossRef] [PubMed]
- Khadri, F.Z.; Issac, M.S.M.; Gaboury, L.A. Impact of epithelial-mesenchymal transition on the immune landscape in breast cancer. Cancers 2021, 13, 99. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Liu, Y.; Zhang, Q.; Zhang, H.; Du, J. Tumor-derived extracellular vesicles containing microRNA-1290 promote immune escape of cancer cells through the Grhl2/ZEB1/PD-L1 axis in gastric cancer. Transl. Res. 2021, 231, 102–112. [Google Scholar] [CrossRef]
- Coban, B.; Wang, Z.; Liao, C.Y.; Beslmüller, K.; Timmermans, M.A.M.; Martens, J.W.M.; Hundscheid, J.H.M.; Slutter, B.; Zweemer, A.J.M.; Neubert, E.; et al. GRHL2 suppression of NT5E/CD73 in breast cancer cells modulates CD73-mediated adenosine production and T cell recruitment. iScience 2024, 27, 109738. [Google Scholar] [CrossRef]
- Shadbad, M.A.; Safaei, S.; Brunetti, O.; Derakhshani, A.; Lotfinejad, P.; Mokhtarzadeh, A.; Hemmat, N.; Racanelli, V.; Solimando, A.G.; Argentiero, A.; et al. A systematic review on the therapeutic potentiality of PD-L1-inhibiting microRNAs for triple-negative breast cancer: Toward single-cell sequencing-guided biomimetic delivery. Genes 2021, 12, 1206. [Google Scholar] [CrossRef]







Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zheng, C.; Allen, K.O.; Liu, T.; Solodin, N.M.; Meyer, M.B.; Salem, K.; Tsourkas, P.K.; McIlwain, S.J.; Vera, J.M.; Cromwell, E.R.; et al. Elevated GRHL2 Imparts Plasticity in ER-Positive Breast Cancer Cells. Cancers 2024, 16, 2906. https://doi.org/10.3390/cancers16162906
Zheng C, Allen KO, Liu T, Solodin NM, Meyer MB, Salem K, Tsourkas PK, McIlwain SJ, Vera JM, Cromwell ER, et al. Elevated GRHL2 Imparts Plasticity in ER-Positive Breast Cancer Cells. Cancers. 2024; 16(16):2906. https://doi.org/10.3390/cancers16162906
Chicago/Turabian StyleZheng, Christy, Kaelyn O. Allen, Tianrui Liu, Natalia M. Solodin, Mark B. Meyer, Kelley Salem, Phillipos K. Tsourkas, Sean J. McIlwain, Jessica M. Vera, Erika R. Cromwell, and et al. 2024. "Elevated GRHL2 Imparts Plasticity in ER-Positive Breast Cancer Cells" Cancers 16, no. 16: 2906. https://doi.org/10.3390/cancers16162906
APA StyleZheng, C., Allen, K. O., Liu, T., Solodin, N. M., Meyer, M. B., Salem, K., Tsourkas, P. K., McIlwain, S. J., Vera, J. M., Cromwell, E. R., Ozers, M. S., Fowler, A. M., & Alarid, E. T. (2024). Elevated GRHL2 Imparts Plasticity in ER-Positive Breast Cancer Cells. Cancers, 16(16), 2906. https://doi.org/10.3390/cancers16162906




