Large B-Cell Lymphomas in the 5th Edition of the WHO-Classification of Haematolymphoid Neoplasms—Updated Classification and New Concepts
Abstract
Simple Summary
Abstract
1. Introduction
2. Diffuse Large B-Cell Lymphoma, Not Otherwise Specified
3. The Molecular Pathogenesis of DLBCL
4. T-Cell/Histiocyte-Rich Large B-Cell Lymphoma
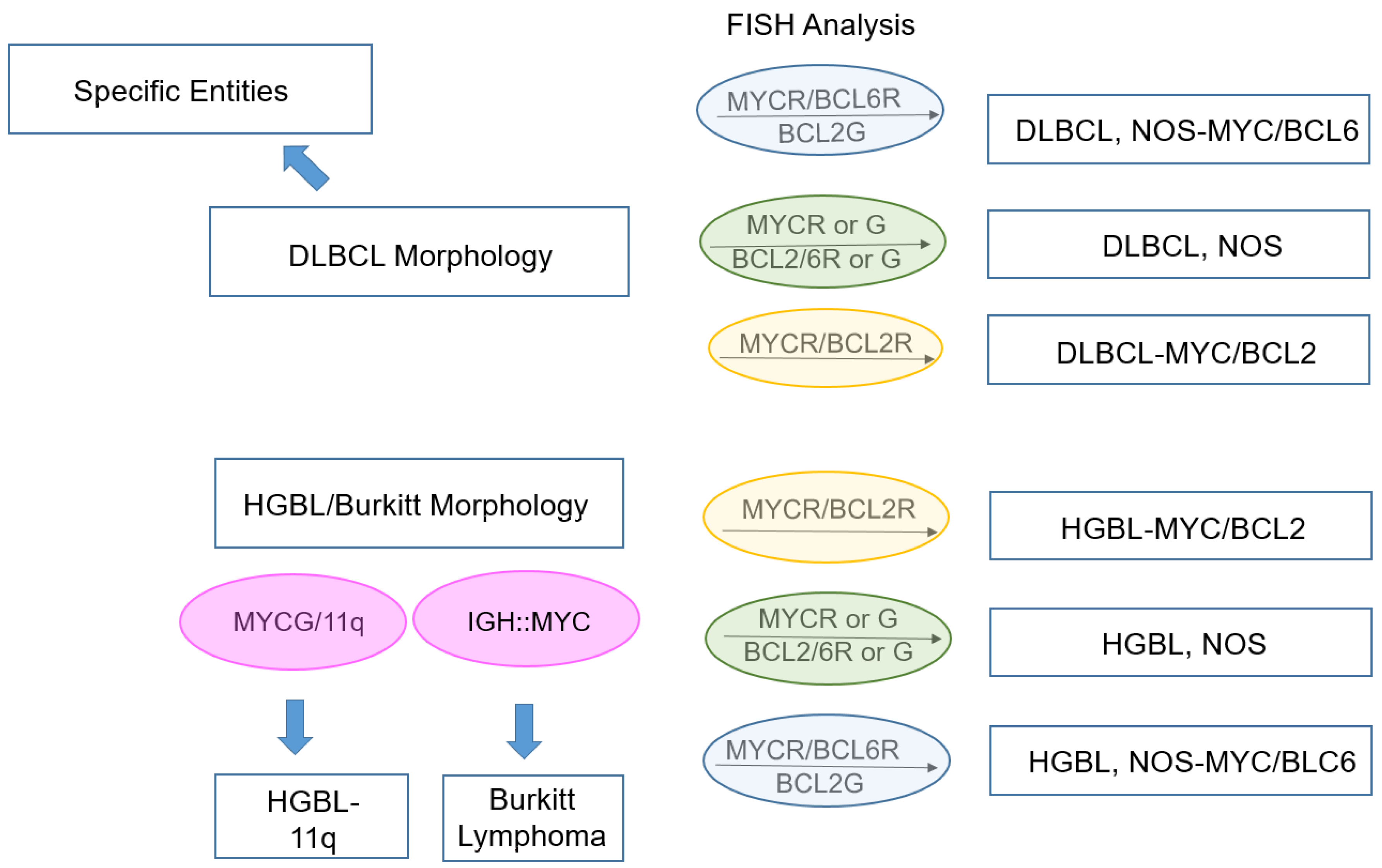
5. Diffuse Large B-Cell Lymphoma/High-Grade B-Cell Lymphoma with MYC and BCL2 Rearrangements
6. ALK-Positive Large B-Cell Lymphoma
7. IRF4-Rearranged Large B-Cell Lymphoma
8. High-Grade B-Cell Lymphoma with 11q Aberrations
9. Lymphomatoid Granulomatosis
10. EBV-Positive Diffuse Large B-Cell Lymphoma
11. Diffuse Large B-Cell Lymphoma Associated with Chronic Inflammation
12. Fibrin-Associated Large B-Cell Lymphoma
13. Fluid Overload-Associated Large B-Cell Lymphoma
14. Plasmablastic Lymphoma
15. Primary Large B-Cell Lymphomas of Immune-Privileged Sites
16. Primary Cutaneous Diffuse Large B-Cell Lymphoma, Leg Type
17. Intravascular Large B-Cell Lymphoma
18. Primary Mediastinal Large B-Cell Lymphoma
19. Mediastinal Grey Zone Lymphoma
20. High-Grade-B-Cell-Lymphoma, NOS
21. Diffuse Large B-Cell Lymphomas Arising in Immune Deficiency/Dysregulation
22. Perspective
23. Conclusions and Future Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- The Non-Hodgkin’s Lymphoma Classification Project. A clinical evaluation of the International Lymphoma Study Group classification of non-Hodgkin’s lymphoma. Blood 1997, 89, 3909–3918. [Google Scholar] [CrossRef]
- Sehn, L.H.; Salles, G. Diffuse Large B-Cell Lymphoma. N. Engl. J. Med. 2021, 384, 842–858. [Google Scholar] [CrossRef] [PubMed]
- Yoo, K.H.; Lee, H.; Suh, C. Lymphoma epidemiology in Korea and the real clinical field including the Consortium for Improving Survival of Lymphoma (CISL) trial. Int. J. Hematol. 2018, 107, 395–404. [Google Scholar] [CrossRef] [PubMed]
- Alaggio, R.; Amador, C.; Anagnostopoulos, I.; Attygalle, A.D.; Araujo, I.B.d.O.; Berti, E.; Bhagat, G.; Borges, A.M.; Boyer, D.; Calaminici, M.; et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Lymphoid Neoplasms. Leukemia 2022, 36, 1720–1748. [Google Scholar] [CrossRef]
- Cerhan, J.R.; Kricker, A.; Paltiel, O.; Flowers, C.R.; Wang, S.S.; Monnereau, A.; Blair, A.; Dal Maso, L.; Kane, E.V.; Nieters, A.; et al. Medical history, lifestyle, family history, and occupational risk factors for diffuse large B-cell lymphoma: The InterLymph Non-Hodgkin Lymphoma Subtypes Project. J. Natl. Cancer Inst. Monogr. 2014, 2014, 15–25. [Google Scholar] [CrossRef]
- Anderson, J.R.; Armitage, J.O.; Weisenburger, D.D. Epidemiology of the non-Hodgkin’s lymphomas: Distribution of the major subtypes differ by geographic locations. Non-Hodgkin’s Lymphoma Classification Project. Ann. Oncol. 1998, 9, 717–720. [Google Scholar] [CrossRef]
- Rosenwald, A.; Wright, G.; Chan, W.C.; Connors, J.M.; Campo, E.; Fisher, R.I.; Gascoyne, R.D.; Muller-Hermelink, H.K.; Smeland, E.B.; Giltnane, J.M.; et al. The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N. Engl. J. Med. 2002, 346, 1937–1947. [Google Scholar] [CrossRef]
- Campo, E.; Jaffe, E.S.; Cook, J.R.; Quintanilla-Martinez, L.; Swerdlow, S.H.; Anderson, K.C.; Brousset, P.; Cerroni, L.; de Leval, L.; Dirnhofer, S.; et al. The International Consensus Classification of Mature Lymphoid Neoplasms: A report from the Clinical Advisory Committee. Blood 2022, 140, 1229–1253. [Google Scholar] [CrossRef]
- Dunleavy, K.; Roschewski, M.; Wilson, W.H. Precision treatment of distinct molecular subtypes of diffuse large B-cell lymphoma: Ascribing treatment based on the molecular phenotype. Clin. Cancer Res. 2014, 20, 5182–5193. [Google Scholar] [CrossRef]
- Staiger, A.M.; Ziepert, M.; Horn, H.; Scott, D.W.; Barth, T.F.E.; Bernd, H.-W.; Feller, A.C.; Klapper, W.; Szczepanowski, M.; Hummel, M.; et al. Clinical Impact of the Cell-of-Origin Classification and the MYC/BCL2 Dual Expresser Status in Diffuse Large B-Cell Lymphoma Treated within Prospective Clinical Trials of the German High-Grade Non-Hodgkin’s Lymphoma Study Group. J. Clin. Oncol. 2017, 35, 2515–2526. [Google Scholar] [CrossRef]
- Hans, C.P.; Weisenburger, D.D.; Greiner, T.C.; Gascoyne, R.D.; Delabie, J.; Ott, G.; Muller-Hermelink, H.K.; Campo, E.; Braziel, R.M.; Jaffe, E.S.; et al. Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood 2004, 103, 275–282. [Google Scholar] [CrossRef]
- Phang, K.-C.; Akhter, A.; Tizen, N.M.S.; Rahman, F.A.; Zahratul Azma, R.; Elyamany, G.; Shabani-Rad, M.-T.; Masir, N.; Mansoor, A. Comparison of protein-based cell-of-origin classification to the Lymph2Cx RNA assay in a cohort of diffuse large B-cell lymphomas in Malaysia. J. Clin. Pathol. 2018, 71, 215–220. [Google Scholar] [CrossRef]
- Pasqualucci, L.; Dalla-Favera, R. The genetic landscape of diffuse large B-cell lymphoma. Semin. Hematol. 2015, 52, 67–76. [Google Scholar] [CrossRef]
- Ennishi, D.; Jiang, A.; Boyle, M.; Collinge, B.; Grande, B.M.; Ben-Neriah, S.; Rushton, C.; Tang, J.; Thomas, N.; Slack, G.W.; et al. Double-Hit Gene Expression Signature Defines a Distinct Subgroup of Germinal Center B-Cell-Like Diffuse Large B-Cell Lymphoma. J. Clin. Oncol. 2018, 37, 190. [Google Scholar] [CrossRef]
- Sha, C.; Barrans, S.; Cucco, F.; Bentley, M.A.; Care, M.A.; Cummin, T.; Kennedy, H.; Thompson, J.S.; Uddin, R.; Worrillow, L.; et al. Molecular High-Grade B-Cell Lymphoma: Defining a Poor-Risk Group That Requires Different Approaches to Therapy. J. Clin. Oncol. 2019, 37, 202–212. [Google Scholar] [CrossRef]
- Ennishi, D.; Mottok, A.; Ben-Neriah, S.; Shulha, H.P.; Farinha, P.; Chan, F.C.; Meissner, B.; Boyle, M.; Hother, C.; Kridel, R.; et al. Genetic profiling of MYC and BCL2 in diffuse large B-cell lymphoma determines cell-of-origin-specific clinical impact. Blood 2017, 129, 2760–2770. [Google Scholar] [CrossRef]
- Iqbal, J.; Greiner, T.C.; Patel, K.; Dave, B.J.; Smith, L.; Ji, J.; Wright, G.; Sanger, W.G.; Pickering, D.L.; Jain, S.; et al. Distinctive patterns of BCL6 molecular alterations and their functional consequences in different subgroups of diffuse large B-cell lymphoma. Leukemia 2007, 21, 2332–2343. [Google Scholar] [CrossRef]
- Rosenwald, A.; Bens, S.; Advani, R.; Barrans, S.; Copie-Bergman, C.; Elsensohn, M.-H.; Natkunam, Y.; Calaminici, M.; Sander, B.; Baia, M.; et al. Prognostic Significance of MYC Rearrangement and Translocation Partner in Diffuse Large B-Cell Lymphoma: A Study by the Lunenburg Lymphoma Biomarker Consortium. J. Clin. Oncol. 2019, 37, 3359–3368. [Google Scholar] [CrossRef]
- Bertrand, P.; Bastard, C.; Maingonnat, C.; Jardin, F.; Maisonneuve, C.; Courel, M.-N.; Ruminy, P.; Picquenot, J.-M.; Tilly, H. Mapping of MYC breakpoints in 8q24 rearrangements involving non-immunoglobulin partners in B-cell lymphomas. Leukemia 2007, 21, 515–523. [Google Scholar] [CrossRef]
- Chong, L.C.; Ben-Neriah, S.; Slack, G.W.; Freeman, C.; Ennishi, D.; Mottok, A.; Collinge, B.; Abrisqueta, P.; Farinha, P.; Boyle, M.; et al. High-resolution architecture and partner genes of MYC rearrangements in lymphoma with DLBCL morphology. Blood Adv. 2018, 2, 2755–2765. [Google Scholar] [CrossRef]
- Johnson, S.M.; Umakanthan, J.M.; Yuan, J.; Fedoriw, Y.; Bociek, R.G.; Kaiser-Rogers, K.; Sanmann, J.N.; Montgomery, N.D. Lymphomas with pseudo-double-hit BCL6-MYC translocations due to t(3;8)(q27;q24) are associated with a germinal center immunophenotype, extranodal involvement, and frequent BCL2 translocations. Hum. Pathol. 2018, 80, 192–200. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.; Zhang, J.; Davis, N.S.; Moffitt, A.B.; Love, C.L.; Waldrop, A.; Leppa, S.; Pasanen, A.; Meriranta, L.; Karjalainen-Lindsberg, M.-L.; et al. Genetic and Functional Drivers of Diffuse Large B Cell Lymphoma. Cell 2017, 171, 481–494.e15. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, R.; Wright, G.W.; Huang, D.W.; Johnson, C.A.; Phelan, J.D.; Wang, J.Q.; Roulland, S.; Kasbekar, M.; Young, R.M.; Shaffer, A.L.; et al. Genetics and Pathogenesis of Diffuse Large B-Cell Lymphoma. N. Engl. J. Med. 2018, 378, 1396–1407. [Google Scholar] [CrossRef] [PubMed]
- Chapuy, B.; Stewart, C.; Dunford, A.J.; Kim, J.; Kamburov, A.; Redd, R.A.; Lawrence, M.S.; Roemer, M.G.M.; Li, A.J.; Ziepert, M.; et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat. Med. 2018, 24, 679–690. [Google Scholar] [CrossRef]
- Lacy, S.E.; Barrans, S.L.; Beer, P.A.; Painter, D.; Smith, A.G.; Roman, E.; Cooke, S.L.; Ruiz, C.; Glover, P.; van Hoppe, S.J.L.; et al. Targeted sequencing in DLBCL, molecular subtypes, and outcomes: A Haematological Malignancy Research Network report. Blood 2020, 135, 1759–1771. [Google Scholar] [CrossRef]
- Achten, R.; Verhoef, G.; Vanuytsel, L.; de Wolf-Peeters, C. T-cell/histiocyte-rich large B-cell lymphoma: A distinct clinicopathologic entity. J. Clin. Oncol. 2002, 20, 1269–1277. [Google Scholar] [CrossRef]
- Fan, Z.; Natkunam, Y.; Bair, E.; Tibshirani, R.; Warnke, R.A. Characterization of variant patterns of nodular lymphocyte predominant hodgkin lymphoma with immunohistologic and clinical correlation. Am. J. Surg. Pathol. 2003, 27, 1346–1356. [Google Scholar] [CrossRef]
- Moore, E.M.; Swerdlow, S.H.; Gibson, S.E. J chain and myocyte enhancer factor 2B are useful in differentiating classical Hodgkin lymphoma from nodular lymphocyte predominant Hodgkin lymphoma and primary mediastinal large B-cell lymphoma. Hum. Pathol. 2017, 68, 47–53. [Google Scholar] [CrossRef]
- Hartmann, S.; Schuhmacher, B.; Rausch, T.; Fuller, L.; Döring, C.; Weniger, M.; Lollies, A.; Weiser, C.; Thurner, L.; Rengstl, B.; et al. Highly recurrent mutations of SGK1, DUSP2 and JUNB in nodular lymphocyte predominant Hodgkin lymphoma. Leukemia 2016, 30, 844–853. [Google Scholar] [CrossRef]
- Hartmann, S.; Tousseyn, T.; Doring, C.; Fluchter, P.; Hackstein, H.; Herreman, A.; Ponzoni, M.; de Wolf-Peeters, C.; Facchetti, F.; Gascoyne, R.D.; et al. Macrophages in T cell/histiocyte rich large B cell lymphoma strongly express metal-binding proteins and show a bi-activated phenotype. Int. J. Cancer 2013, 133, 2609–2618. [Google Scholar] [CrossRef]
- Macon, W.R.; Cousar, J.B.; Waldron, J.A.; Hsu, S.M. Interleukin-4 may contribute to the abundant T-cell reaction and paucity of neoplastic B cells in T-cell-rich B-cell lymphomas. Am. J. Pathol. 1992, 141, 1031–1036. [Google Scholar]
- Chetaille, B.; Bertucci, F.; Finetti, P.; Esterni, B.; Stamatoullas, A.; Picquenot, J.M.; Copin, M.C.; Morschhauser, F.; Casasnovas, O.; Petrella, T.; et al. Molecular profiling of classical Hodgkin lymphoma tissues uncovers variations in the tumor microenvironment and correlations with EBV infection and outcome. Blood 2009, 113, 2765–3775. [Google Scholar] [CrossRef]
- Van Loo, P.; Tousseyn, T.; Vanhentenrijk, V.; Dierickx, D.; Malecka, A.; Vanden Bempt, I.; Verhoef, G.; Delabie, J.; Marynen, P.; Matthys, P.; et al. T-cell/histiocyte-rich large B-cell lymphoma shows transcriptional features suggestive of a tolerogenic host immune response. Haematologica 2010, 95, 440–448. [Google Scholar] [CrossRef]
- Kunder, C.; Cascio, M.J.; Bakke, A.; Venkataraman, G.; O’Malley, D.P.; Ohgami, R.S. Predominance of CD4+ T Cells in T-Cell/Histiocyte-Rich Large B-Cell Lymphoma and Identification of a Subset of Patients with Peripheral B-Cell Lymphopenia. Am. J. Clin. Pathol. 2017, 147, 596–603. [Google Scholar] [CrossRef]
- Wu, D.; Thomas, A.; Fromm, J.R. Reactive T cells by flow cytometry distinguish Hodgkin lymphomas from T cell/histiocyte-rich large B cell lymphoma. Cytometry B Clin. Cytom. 2016, 90, 424–432. [Google Scholar] [CrossRef]
- Collinge, B.; Ben-Neriah, S.; Chong, L.; Boyle, M.; Jiang, A.; Miyata-Takata, T.; Farinha, P.; Craig, J.W.; Slack, G.W.; Ennishi, D.; et al. The impact of MYC and BCL2 structural variants in tumors of DLBCL morphology and mechanisms of false-negative MYC IHC. Blood 2021, 137, 2196–2208. [Google Scholar] [CrossRef]
- Li, S.; Lin, P.; Fayad, L.E.; Lennon, P.A.; Miranda, R.N.; Yin, C.C.; Lin, E.; Medeiros, L.J. B-cell lymphomas with MYC/8q24 rearrangements and IGH@BCL2/t(14;18)(q32;q21): An aggressive disease with heterogeneous histology, germinal center B-cell immunophenotype and poor outcome. Mod. Pathol. 2012, 25, 145–156. [Google Scholar] [CrossRef]
- Laude, M.-C.; Lebras, L.; Sesques, P.; Ghesquieres, H.; Favre, S.; Bouabdallah, K.; Croizier, C.; Guieze, R.; La Drieu Rochelle, L.; Gyan, E.; et al. First-line treatment of double-hit and triple-hit lymphomas: Survival and tolerance data from a retrospective multicenter French study. Am. J. Hematol. 2021, 96, 302–311. [Google Scholar] [CrossRef]
- Scott, D.W.; King, R.L.; Staiger, A.M.; Ben-Neriah, S.; Jiang, A.; Horn, H.; Mottok, A.; Farinha, P.; Slack, G.W.; Ennishi, D.; et al. High-grade B-cell lymphoma with MYC and BCL2 and/or BCL6 rearrangements with diffuse large B-cell lymphoma morphology. Blood 2018, 131, 2060–2064. [Google Scholar] [CrossRef]
- Aukema, S.M.; Kreuz, M.; Kohler, C.W.; Rosolowski, M.; Hasenclever, D.; Hummel, M.; Kuppers, R.; Lenze, D.; Ott, G.; Pott, C.; et al. Biological characterization of adult MYC-translocation-positive mature B-cell lymphomas other than molecular Burkitt lymphoma. Haematologica 2014, 99, 726–735. [Google Scholar] [CrossRef]
- Krull, J.E.; Wenzl, K.; Hartert, K.T.; Manske, M.K.; Sarangi, V.; Maurer, M.J.; Larson, M.C.; Nowakowski, G.S.; Ansell, S.M.; McPhail, E.; et al. Somatic copy number gains in MYC, BCL2, and BCL6 identifies a subset of aggressive alternative-DH/TH DLBCL patients. Blood Cancer J. 2020, 10, 117. [Google Scholar] [CrossRef]
- Aukema, S.M.; Siebert, R.; Schuuring, E.; van Imhoff, G.W.; Kluin-Nelemans, H.C.; Boerma, E.-J.; Kluin, P.M. Double-hit B-cell lymphomas. Blood 2011, 117, 2319–2331. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Medeiros, L.J.; Lin, P.; Wang, W.; Tang, G.; Khoury, J.; Konoplev, S.; Yin, C.C.; Xu, J.; Oki, Y.; et al. MYC/BCL2/BCL6 triple hit lymphoma: A study of 40 patients with a comparison to MYC/BCL2 and MYC/BCL6 double hit lymphomas. Mod. Pathol. 2018, 31, 1470–1478. [Google Scholar] [CrossRef] [PubMed]
- McPhail, E.D.; Maurer, M.J.; Macon, W.R.; Feldman, A.L.; Kurtin, P.J.; Ketterling, R.P.; Vaidya, R.; Cerhan, J.R.; Ansell, S.M.; Porrata, L.F.; et al. Inferior survival in high-grade B-cell lymphoma with MYC and BCL2 and/or BCL6 rearrangements is not associated with MYC/IG gene rearrangements. Haematologica 2018, 103, 1899–1907. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, M.Ø.; Gang, A.O.; Clasen-Linde, E.; Breinholt, M.F.; Knudsen, H.; Nielsen, S.L.; Poulsen, T.S.; Klausen, T.W.; Høgdall, E.; Nørgaard, P. Stratification by MYC expression has prognostic impact in MYC translocated B-cell lymphoma-Identifies a subgroup of patients with poor outcome. Eur. J. Haematol. 2019, 102, 395–406. [Google Scholar] [CrossRef]
- Cucco, F.; Barrans, S.; Sha, C.; Clipson, A.; Crouch, S.; Dobson, R.; Chen, Z.; Thompson, J.S.; Care, M.A.; Cummin, T.; et al. Distinct genetic changes reveal evolutionary history and heterogeneous molecular grade of DLBCL with MYC/BCL2 double-hit. Leukemia 2020, 34, 1329–1341. [Google Scholar] [CrossRef]
- Vogelsberg, A.; Steinhilber, J.; Mankel, B.; Federmann, B.; Schmidt, J.; Montes-Mojarro, I.A.; Hüttl, K.; Rodriguez-Pinilla, M.; Baskaran, P.; Nahnsen, S.; et al. Genetic evolution of in situ follicular neoplasia to aggressive B-cell lymphoma of germinal center subtype. Haematologica 2021, 106, 2673–2681. [Google Scholar] [CrossRef]
- Wright, G.W.; Da Huang, W.; Phelan, J.D.; Coulibaly, Z.A.; Roulland, S.; Young, R.M.; Wang, J.Q.; Schmitz, R.; Morin, R.D.; Tang, J.; et al. A Probabilistic Classification Tool for Genetic Subtypes of Diffuse Large B Cell Lymphoma with Therapeutic Implications. Cancer Cell 2020, 37, 551–568.e14. [Google Scholar] [CrossRef]
- Hilton, L.K.; Tang, J.; Ben-Neriah, S.; Alcaide, M.; Jiang, A.; Grande, B.M.; Rushton, C.K.; Boyle, M.; Meissner, B.; Scott, D.W.; et al. The double-hit signature identifies double-hit diffuse large B-cell lymphoma with genetic events cryptic to FISH. Blood 2019, 134, 1528–1532. [Google Scholar] [CrossRef]
- Hummel, M.; Bentink, S.; Berger, H.; Klapper, W.; Wessendorf, S.; Barth, T.F.E.; Bernd, H.-W.; Cogliatti, S.B.; Dierlamm, J.; Feller, A.C.; et al. A biologic definition of Burkitt’s lymphoma from transcriptional and genomic profiling. N. Engl. J. Med. 2006, 354, 2419–2430. [Google Scholar] [CrossRef]
- Bhavsar, S.; Liu, Y.-C.; Gibson, S.E.; Moore, E.M.; Swerdlow, S.H. Mutational Landscape of TdT+ Large B-cell Lymphomas Supports Their Distinction from B-lymphoblastic Neoplasms: A Multiparameter Study of a Rare and Aggressive Entity. Am. J. Surg. Pathol. 2022, 46, 71–82. [Google Scholar] [CrossRef]
- Qiu, L.; Xu, J.; Lin, P.; Cohen, E.N.; Tang, G.; Wang, S.A.; Khanlari, M.; Wang, W.; Khoury, J.D.; Konoplev, S.; et al. Unique pathologic features and gene expression signatures distinguish blastoid high-grade B-cell lymphoma from B acute lymphoblastic leukemia/lymphoma. Haematologica 2022, 3, 895–899. [Google Scholar] [CrossRef]
- Wagener, R.; López, C.; Kleinheinz, K.; Bausinger, J.; Aukema, S.M.; Nagel, I.; Toprak, U.H.; Seufert, J.; Altmüller, J.; Thiele, H.; et al. IG-MYC+ neoplasms with precursor B-cell phenotype are molecularly distinct from Burkitt lymphomas. Blood 2018, 132, 2280–2285. [Google Scholar] [CrossRef]
- Liu, W.; Hu, S.; Konopleva, M.; Khoury, J.D.; Kalhor, N.; Tang, G.; Bueso-Ramos, C.E.; Jorgensen, J.L.; Lin, P.; Medeiros, L.J.; et al. De Novo MYC and BCL2 Double-hit B-Cell Precursor Acute Lymphoblastic Leukemia (BCP-ALL) in Pediatric and Young Adult Patients Associated with Poor Prognosis. Pediatr. Hematol. Oncol. 2015, 32, 535–547. [Google Scholar] [CrossRef]
- Khanlari, M.; Medeiros, L.J.; Lin, P.; Xu, J.; You, M.J.; Tang, G.; Yin, C.C.; Wang, W.; Qiu, L.; Miranda, R.N.; et al. Blastoid high-grade B-cell lymphoma initially presenting in bone marrow: A diagnostic challenge. Mod. Pathol. 2022, 35, 419–426. [Google Scholar] [CrossRef]
- Geyer, J.T.; Subramaniyam, S.; Jiang, Y.; Elemento, O.; Ferry, J.A.; de Leval, L.; Nakashima, M.O.; Liu, Y.-C.; Martin, P.; Mathew, S.; et al. Lymphoblastic transformation of follicular lymphoma: A clinicopathologic and molecular analysis of 7 patients. Hum. Pathol. 2015, 46, 260–271. [Google Scholar] [CrossRef]
- Pan, Z.; Hu, S.; Li, M.; Zhou, Y.; Kim, Y.S.; Reddy, V.; Sanmann, J.N.; Smith, L.M.; Chen, M.; Gao, Z.; et al. ALK-positive Large B-cell Lymphoma: A Clinicopathologic Study of 26 Cases with Review of Additional 108 Cases in the Literature. Am. J. Surg. Pathol. 2017, 41, 25–38. [Google Scholar] [CrossRef]
- Laurent, C.; Do, C.; Gascoyne, R.D.; Lamant, L.; Ysebaert, L.; Laurent, G.; Delsol, G.; Brousset, P. Anaplastic lymphoma kinase-positive diffuse large B-cell lymphoma: A rare clinicopathologic entity with poor prognosis. J. Clin. Oncol. 2009, 27, 4211–4216. [Google Scholar] [CrossRef]
- Valera, A.; Colomo, L.; Martinez, A.; de Jong, D.; Balague, O.; Matheu, G.; Martinez, M.; Taddesse-Heath, L.; Jaffe, E.S.; Bacchi, C.E.; et al. ALK-positive large B-cell lymphomas express a terminal B-cell differentiation program and activated STAT3 but lack MYC rearrangements. Mod. Pathol. 2013, 26, 1329–1337. [Google Scholar] [CrossRef]
- Sakamoto, K.; Nakasone, H.; Togashi, Y.; Sakata, S.; Tsuyama, N.; Baba, S.; Dobashi, A.; Asaka, R.; Tsai, C.-C.; Chuang, S.-S.; et al. ALK-positive large B-cell lymphoma: Identification of EML4-ALK and a review of the literature focusing on the ALK immunohistochemical staining pattern. Int. J. Hematol. 2016, 103, 399–408. [Google Scholar] [CrossRef]
- Castillo, J.J.; Beltran, B.E.; Malpica, L.; Marques-Piubelli, M.L.; Miranda, R.N. Anaplastic lymphoma kinase-positive large B-cell lymphoma (ALK + LBCL): A systematic review of clinicopathological features and management. Leuk. Lymphoma 2021, 62, 2845–2853. [Google Scholar] [CrossRef] [PubMed]
- Onciu, M.; Behm, F.G.; Downing, J.R.; Shurtleff, S.A.; Raimondi, S.C.; Ma, Z.; Morris, S.W.; Kennedy, W.; Jones, S.C.; Sandlund, J.T. ALK-positive plasmablastic B-cell lymphoma with expression of the NPM-ALK fusion transcript: Report of 2 cases. Blood 2003, 102, 2642–2644. [Google Scholar] [CrossRef] [PubMed]
- Rossky, P.J.; Walker, G.C. Retrospective. Paul F. Barbara (1953–2010). Science 2010, 330, 1191. [Google Scholar] [CrossRef]
- Takeuchi, K.; Soda, M.; Togashi, Y.; Ota, Y.; Sekiguchi, Y.; Hatano, S.; Asaka, R.; Noguchi, M.; Mano, H. Identification of a novel fusion, SQSTM1-ALK, in ALK-positive large B-cell lymphoma. Haematologica 2011, 96, 464–467. [Google Scholar] [CrossRef]
- Lee, S.E.; Kang, S.Y.; Takeuchi, K.; Ko, Y.H. Identification of RANBP2-ALK fusion in ALK positive diffuse large B-cell lymphoma. Hematol. Oncol. 2014, 32, 221–224. [Google Scholar] [CrossRef] [PubMed]
- Ise, M.; Kageyama, H.; Araki, A.; Itami, M. Identification of a novel GORASP2-ALK fusion in an ALK-positive large B-cell lymphoma. Leuk. Lymphoma 2019, 60, 493–497. [Google Scholar] [CrossRef]
- Salaverria, I.; Philipp, C.; Oschlies, I.; Kohler, C.W.; Kreuz, M.; Szczepanowski, M.; Burkhardt, B.; Trautmann, H.; Gesk, S.; Andrusiewicz, M.; et al. Translocations activating IRF4 identify a subtype of germinal center-derived B-cell lymphoma affecting predominantly children and young adults. Blood 2011, 118, 139–147. [Google Scholar] [CrossRef]
- Au-Yeung, R.K.H.; Arias Padilla, L.; Zimmermann, M.; Oschlies, I.; Siebert, R.; Woessmann, W.; Burkhardt, B.; Klapper, W. Experience with provisional WHO-entities large B-cell lymphoma with IRF4-rearrangement and Burkitt-like lymphoma with 11q aberration in paediatric patients of the NHL-BFM group. Br. J. Haematol. 2020, 190, 753–763. [Google Scholar] [CrossRef]
- De Leval, L.; Bonnet, C.; Copie-Bergman, C.; Seidel, L.; Baia, M.; Brière, J.; Molina, T.J.; Fabiani, B.; Petrella, T.; Bosq, J.; et al. Diffuse large B-cell lymphoma of Waldeyer’s ring has distinct clinicopathologic features: A GELA study. Ann. Oncol. 2012, 23, 3143–3151. [Google Scholar] [CrossRef]
- Chisholm, K.M.; Mohlman, J.; Liew, M.; Termuhlen, A.; Cairo, M.S.; Gross, T.G.; Perkins, S.L.; Miles, R.R. IRF4 translocation status in pediatric follicular and diffuse large B-cell lymphoma patients enrolled in Children’s Oncology Group trials. Pediatr. Blood Cancer 2019, 66, e27770. [Google Scholar] [CrossRef]
- Ramis-Zaldivar, J.E.; Gonzalez-Farré, B.; Balagué, O.; Celis, V.; Nadeu, F.; Salmerón-Villalobos, J.; Andrés, M.; Martin-Guerrero, I.; Garrido-Pontnou, M.; Gaafar, A.; et al. Distinct molecular profile of IRF4-rearranged large B-cell lymphoma. Blood 2020, 135, 274–286. [Google Scholar] [CrossRef]
- Montes-Moreno, S.; King, R.L.; Oschlies, I.; Ponzoni, M.; Goodlad, J.R.; Dotlic, S.; Traverse-Glehen, A.; Ott, G.; Ferry, J.A.; Calaminici, M. Update on lymphoproliferative disorders of the gastrointestinal tract: Disease spectrum from indolent lymphoproliferations to aggressive lymphomas. Virchows Arch. 2020, 476, 667–681. [Google Scholar] [CrossRef]
- Salaverria, I.; Martin-Guerrero, I.; Burkhardt, B.; Kreuz, M.; Zenz, T.; Oschlies, I.; Arnold, N.; Baudis, M.; Bens, S.; García-Orad, A.; et al. High resolution copy number analysis of IRF4 translocation-positive diffuse large B-cell and follicular lymphomas. Genes Chromosomes. Cancer 2013, 52, 150–155. [Google Scholar] [CrossRef]
- Grygalewicz, B.; Woroniecka, R.; Rymkiewicz, G.; Rygier, J.; Borkowska, K.; Kotyl, A.; Blachnio, K.; Bystydzienski, Z.; Nowakowska, B.; Pienkowska-Grela, B. The 11q-Gain/Loss Aberration Occurs Recurrently in MYC-Negative Burkitt-like Lymphoma with 11q Aberration, as Well as MYC-Positive Burkitt Lymphoma and MYC-Positive High-Grade B-Cell Lymphoma, NOS. Am. J. Clin. Pathol. 2017, 149, 17–28. [Google Scholar] [CrossRef]
- Gonzalez-Farre, B.; Ramis-Zaldivar, J.E.; Salmeron-Villalobos, J.; Balagué, O.; Celis, V.; Verdu-Amoros, J.; Nadeu, F.; Sábado, C.; Ferrández, A.; Garrido, M.; et al. Burkitt-like lymphoma with 11q aberration: A germinal center derived lymphoma genetically unrelated to Burkitt lymphoma. Haematologica 2019, 104, 1822. [Google Scholar] [CrossRef] [PubMed]
- Ferreiro, J.F.; Morscio, J.; Dierickx, D.; Marcelis, L.; Verhoef, G.; Vandenberghe, P.; Tousseyn, T.; Wlodarska, I. Post-transplant molecularly defined Burkitt lymphomas are frequently MYC-negative and characterized by the 11q-gain/loss pattern. Haematologica 2015, 100, e275–e279. [Google Scholar] [CrossRef]
- Salaverria, I.; Martin-Guerrero, I.; Wagener, R.; Kreuz, M.; Kohler, C.W.; Richter, J.; Pienkowska-Grela, B.; Adam, P.; Burkhardt, B.; Claviez, A.; et al. A recurrent 11q aberration pattern characterizes a subset of MYC-negative high-grade B-cell lymphomas resembling Burkitt lymphoma. Blood 2014, 123, 1187–1198. [Google Scholar] [CrossRef] [PubMed]
- Horn, H.; Kalmbach, S.; Wagener, R.; Staiger, A.M.; Hüttl, K.; Mottok, A.; Bens, S.; Traverse-Glehen, A.; Fontaine, J.; Siebert, R.; et al. A Diagnostic Approach to the Identification of Burkitt-like Lymphoma with 11q Aberration in Aggressive B-Cell Lymphomas. Am. J. Surg. Pathol. 2021, 45, 356–364. [Google Scholar] [CrossRef]
- Wagener, R.; Seufert, J.; Raimondi, F.; Bens, S.; Kleinheinz, K.; Nagel, I.; Altmüller, J.; Thiele, H.; Hübschmann, D.; Kohler, C.W.; et al. The mutational landscape of Burkitt-like lymphoma with 11q aberration is distinct from that of Burkitt lymphoma. Blood 2018, 133, 962–966. [Google Scholar] [CrossRef] [PubMed]
- Katzenstein, A.L.; Doxtader, E.; Narendra, S. Lymphomatoid granulomatosis: Insights gained over 4 decades. Am. J. Surg. Pathol. 2010, 34, e35–e48. [Google Scholar] [CrossRef]
- Song, J.Y.; Pittaluga, S.; Dunleavy, K.; Grant, N.; White, T.; Jiang, L.; Vies-Hill, T.; Raffeld, M.; Wilson, W.H.; Jaffe, E.S. Lymphomatoid granulomatosis—A single institute experience: Pathologic findings and clinical correlations. Am. J. Surg. Pathol. 2015, 39, 141–156. [Google Scholar] [CrossRef] [PubMed]
- Chavez, J.C.; Sandoval-Sus, J.; Horna, P.; Dalia, S.; Bello, C.; Chevernick, P.; Sotomayor, E.M.; Sokol, L.; Shah, B. Lymphomatoid Granulomatosis: A Single Institution Experience and Review of the Literature. Clin. Lymphoma Myeloma Leuk. 2016, 16, S170–S174. [Google Scholar] [CrossRef] [PubMed]
- Sordillo, P.P.; Epremian, B.; Koziner, B.; Lacher, M.; Lieberman, P. Lymphomatoid granulomatosis: An analysis of clinical and immunologic characteristics. Cancer 1982, 49, 2070–2076. [Google Scholar] [CrossRef]
- Wilson, W.H.; Kingma, D.W.; Raffeld, M.; Wittes, R.E.; Jaffe, E.S. Association of lymphomatoid granulomatosis with Epstein-Barr viral infection of B lymphocytes and response to interferon-alpha 2b. Blood 1996, 87, 4531–4537. [Google Scholar] [CrossRef] [PubMed]
- Teruya-Feldstein, J.; Jaffe, E.S.; Burd, P.R.; Kanegane, H.; Kingma, D.W.; Wilson, W.H.; Longo, D.L.; Tosato, G. The role of Mig, the monokine induced by interferon-gamma, and IP-10, the interferon-gamma-inducible protein-10, in tissue necrosis and vascular damage associated with Epstein-Barr virus-positive lymphoproliferative disease. Blood 1997, 90, 4099–4105. [Google Scholar] [CrossRef]
- Nicolae, A.; Pittaluga, S.; Abdullah, S.; Steinberg, S.M.; Pham, T.A.; vies-Hill, T.; Xi, L.; Raffeld, M.; Jaffe, E.S. EBV-positive large B-cell lymphomas in young patients: A nodal lymphoma with evidence for a tolerogenic immune environment. Blood 2015, 126, 863–872. [Google Scholar] [CrossRef]
- Park, S.; Lee, J.; Ko, Y.H.; Han, A.; Jun, H.J.; Lee, S.C.; Hwang, I.G.; Park, Y.H.; Ahn, J.S.; Jung, C.W.; et al. The impact of Epstein-Barr virus status on clinical outcome in diffuse large B-cell lymphoma. Blood 2007, 110, 972–978. [Google Scholar] [CrossRef]
- Dojcinov, S.D.; Venkataraman, G.; Pittaluga, S.; Wlodarska, I.; Schrager, J.A.; Raffeld, M.; Hills, R.K.; Jaffe, E.S. Age-related EBV-associated lymphoproliferative disorders in the Western population: A spectrum of reactive lymphoid hyperplasia and lymphoma. Blood 2011, 117, 4726–4735. [Google Scholar] [CrossRef]
- Oyama, T.; Ichimura, K.; Suzuki, R.; Suzumiya, J.; Ohshima, K.; Yatabe, Y.; Yokoi, T.; Kojima, M.; Kamiya, Y.; Taji, H.; et al. Senile EBV+ B-cell lymphoproliferative disorders: A clinicopathologic study of 22 patients. Am. J. Surg. Pathol. 2003, 27, 16–26. [Google Scholar] [CrossRef]
- Ok, C.Y.; Papathomas, T.G.; Medeiros, L.J.; Young, K.H. EBV-positive diffuse large B-cell lymphoma of the elderly. Blood 2013, 122, 328–340. [Google Scholar] [CrossRef]
- Tracy, S.I.; Habermann, T.M.; Feldman, A.L.; Maurer, M.J.; Dogan, A.; Perepu, U.S.; Syrbu, S.; Ansell, S.M.; Thompson, C.A.; Weiner, G.J.; et al. Outcomes among North American patients with diffuse large B-cell lymphoma are independent of tumor Epstein-Barr virus positivity or immunosuppression. Haematologica 2018, 103, 297–303. [Google Scholar] [CrossRef] [PubMed]
- Bourbon, E.; Maucort-Boulch, D.; Fontaine, J.; Mauduit, C.; Sesques, P.; Safar, V.; Ferrant, E.; Golfier, C.; Ghergus, D.; Karlin, L.; et al. Clinicopathological features and survival in EBV-positive diffuse large B-cell lymphoma not otherwise specified. Blood Adv. 2021, 5, 3227–3239. [Google Scholar] [CrossRef]
- Oyama, T.; Yamamoto, K.; Asano, N.; Oshiro, A.; Suzuki, R.; Kagami, Y.; Morishima, Y.; Takeuchi, K.; Izumo, T.; Mori, S.; et al. Age-related EBV-associated B-cell lymphoproliferative disorders constitute a distinct clinicopathologic group: A study of 96 patients. Clin. Cancer Res. 2007, 13, 5124–5132. [Google Scholar] [CrossRef] [PubMed]
- Asano, N.; Yamamoto, K.; Tamaru, J.-I.; Oyama, T.; Ishida, F.; Ohshima, K.; Yoshino, T.; Nakamura, N.; Mori, S.; Yoshie, O.; et al. Age-related Epstein-Barr virus (EBV)-associated B-cell lymphoproliferative disorders: Comparison with EBV-positive classic Hodgkin lymphoma in elderly patients. Blood 2009, 113, 2629–2636. [Google Scholar] [CrossRef]
- Montes-Moreno, S.; Odqvist, L.; Diaz-Perez, J.A.; Lopez, A.B.; de Villambrosía, S.G.; Mazorra, F.; Castillo, M.E.; Lopez, M.; Pajares, R.; García, J.F.; et al. EBV-positive diffuse large B-cell lymphoma of the elderly is an aggressive post-germinal center B-cell neoplasm characterized by prominent nuclear factor-kB activation. Mod. Pathol. 2012, 25, 968–982. [Google Scholar] [CrossRef]
- Chen, B.J.; Chapuy, B.; Ouyang, J.; Sun, H.H.; Roemer, M.G.; Xu, M.L.; Yu, H.; Fletcher, C.D.; Freeman, G.J.; Shipp, M.A.; et al. PD-L1 expression is characteristic of a subset of aggressive B-cell lymphomas and virus-associated malignancies. Clin. Cancer Res. 2013, 19, 3462–3473. [Google Scholar] [CrossRef] [PubMed]
- Gebauer, N.; Künstner, A.; Ketzer, J.; Witte, H.M.; Rausch, T.; Benes, V.; Zimmermann, J.; Gebauer, J.; Merz, H.; Bernard, V.; et al. Genomic insights into the pathogenesis of Epstein-Barr virus-associated diffuse large B-cell lymphoma by whole-genome and targeted amplicon sequencing. Blood Cancer J. 2021, 11, 102. [Google Scholar] [CrossRef]
- Zhou, Y.; Xu, Z.; Lin, W.; Duan, Y.; Lu, C.; Liu, W.; Su, W.; Yan, Y.; Liu, H.; Liu, L.; et al. Comprehensive Genomic Profiling of EBV-Positive Diffuse Large B-cell Lymphoma and the Expression and Clinicopathological Correlations of Some Related Genes. Front. Oncol. 2019, 9, 683. [Google Scholar] [CrossRef]
- Takahara, T.; Satou, A.; Ishikawa, E.; Kohno, K.; Kato, S.; Suzuki, Y.; Takahashi, E.; Ohashi, A.; Asano, N.; Tsuzuki, T.; et al. Clinicopathological analysis of neoplastic PD-L1-positive EBV+ diffuse large B cell lymphoma, not otherwise specified, in a Japanese cohort. Virchows Arch. 2021, 478, 541–552. [Google Scholar] [CrossRef]
- Cheuk, W.; Chan, A.C.L.; Chan, J.K.C.; Lau, G.T.C.; Chan, V.N.H.; Yiu, H.H.Y. Metallic implant-associated lymphoma: A distinct subgroup of large B-cell lymphoma related to pyothorax-associated lymphoma? Am. J. Surg. Pathol. 2005, 29, 832–836. [Google Scholar] [CrossRef]
- Petitjean, B.; Jardin, F.; Joly, B.; Martin-Garcia, N.; Tilly, H.; Picquenot, J.M.; Briere, J.; Danel, C.; Mehaut, S.; bd-Al-Samad, I.; et al. Pyothorax-associated lymphoma: A peculiar clinicopathologic entity derived from B cells at late stage of differentiation and with occasional aberrant dual B- and T-cell phenotype. Am. J. Surg. Pathol. 2002, 26, 724–732. [Google Scholar] [CrossRef] [PubMed]
- Nakatsuka, S.; Yao, M.; Hoshida, Y.; Yamamoto, S.; Iuchi, K.; Aozasa, K. Pyothorax-associated lymphoma: A review of 106 cases. J. Clin. Oncol. 2002, 20, 4255–4260. [Google Scholar] [CrossRef] [PubMed]
- Narimatsu, H.; Ota, Y.; Kami, M.; Takeuchi, K.; Suzuki, R.; Matsuo, K.; Matsumura, T.; Yuji, K.; Kishi, Y.; Hamaki, T.; et al. Clinicopathological features of pyothorax-associated lymphoma; a retrospective survey involving 98 patients. Ann. Oncol. 2007, 18, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Aozasa, K.; Takakuwa, T.; Nakatsuka, S. Pyothorax-associated lymphoma: A lymphoma developing in chronic inflammation. Adv. Anat. Pathol. 2005, 12, 324–331. [Google Scholar] [CrossRef]
- Kanno, H.; Yasunaga, Y.; Iuchi, K.; Yamauchi, S.; Tatekawa, T.; Sugiyama, H.; Aozasa, K. Interleukin-6-mediated growth enhancement of cell lines derived from pyothorax-associated lymphoma. Lab. Investig. 1996, 75, 167–173. [Google Scholar]
- Kanno, H.; Naka, N.; Yasunaga, Y.; Aozasa, K. Role of an immunosuppressive cytokine, interleukin-10, in the development of pyothorax-associated lymphoma. Leukemia 1997, 11 (Suppl. 3), 525–526. [Google Scholar]
- Copie-Bergman, C.; Niedobitek, G.; Mangham, D.C.; Selves, J.; Baloch, K.; Diss, T.C.; Knowles, D.N.; Delsol, G.; Isaacson, P.G. Epstein-Barr virus in B-cell lymphomas associated with chronic suppurative inflammation. J. Pathol. 1997, 183, 287–292. [Google Scholar] [CrossRef]
- Fujimoto, M.; Haga, H.; Okamoto, M.; Obara, E.; Ishihara, M.; Mizuta, N.; Nishimura, K.; Manabe, T. EBV-associated diffuse large B-cell lymphoma arising in the chest wall with surgical mesh implant. Pathol. Int. 2008, 58, 668–671. [Google Scholar] [CrossRef]
- Nishiu, M.; Tomita, Y.; Nakatsuka, S.; Takakuwa, T.; Iizuka, N.; Hoshida, Y.; Ikeda, J.; Iuchi, K.; Yanagawa, R.; Nakamura, Y.; et al. Distinct pattern of gene expression in pyothorax-associated lymphoma (PAL), a lymphoma developing in long-standing inflammation. Cancer Sci. 2004, 95, 828–834. [Google Scholar] [CrossRef]
- Kanno, H.; Ohsawa, M.; Hashimoto, M.; Iuchi, K.; Nakajima, Y.; Aozasa, K. HLA-A alleles of patients with pyothorax-associated lymphoma: Anti-Epstein-Barr virus (EBV) host immune responses during the development of EBV latent antigen-positive lymphomas. Int. J. Cancer 1999, 82, 630–634. [Google Scholar] [CrossRef]
- Kanno, H.; Nakatsuka, S.; Iuchi, K.; Aozasa, K. Sequences of cytotoxic T-lymphocyte epitopes in the Epstein-Barr virus (EBV) nuclear antigen-3B gene in a Japanese population with or without EBV-positive lymphoid malignancies. Int. J. Cancer 2000, 88, 626–632. [Google Scholar] [CrossRef] [PubMed]
- Ando, M.; Sato, Y.; Takata, K.; Nomoto, J.; Nakamura, S.; Ohshima, K.; Takeuchi, T.; Orita, Y.; Kobayashi, Y.; Yoshino, T. A20 (TNFAIP3) deletion in Epstein-Barr virus-associated lymphoproliferative disorders/lymphomas. PLoS ONE 2013, 8, e56741. [Google Scholar] [CrossRef]
- Hongyo, T.; Kurooka, M.; Taniguchi, E.; Iuchi, K.; Nakajima, Y.; Aozasa, K.; Nomura, T. Frequent p53 mutations at dipyrimidine sites in patients with pyothorax-associated lymphoma. Cancer Res. 1998, 58, 1105–1107. [Google Scholar] [PubMed]
- Boroumand, N.; Ly, T.L.; Sonstein, J.; Medeiros, L.J. Microscopic diffuse large B-cell lymphoma (DLBCL) occurring in pseudocysts: Do these tumors belong to the category of DLBCL associated with chronic inflammation? Am. J. Surg. Pathol. 2012, 36, 1074–1080. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Pinilla, S.M.; García, F.J.S.; Balagué, O.; Rodríguez-Justo, M.; Piris, M.Á. Breast implant-associated Epstein-Barr virus-positive large B-cell lymphomas: A report of three cases. Haematologica 2020, 105, e412–e414. [Google Scholar] [CrossRef]
- Zanelli, M.; Zizzo, M.; Montanaro, M.; Gomes, V.; Martino, G.; de Marco, L.; Fraternali Orcioni, G.; Martelli, M.P.; Ascani, S. Fibrin-associated large B-cell lymphoma: First case report within a cerebral artery aneurysm and literature review. BMC Cancer 2019, 19, 916. [Google Scholar] [CrossRef]
- Boyer, D.F.; McKelvie, P.A.; de Leval, L.; Edlefsen, K.L.; Ko, Y.-H.; Aberman, Z.A.; Kovach, A.E.; Masih, A.; Nishino, H.T.; Weiss, L.M.; et al. Fibrin-associated EBV-positive Large B-Cell Lymphoma: An Indolent Neoplasm with Features Distinct from Diffuse Large B-Cell Lymphoma Associated with Chronic Inflammation. Am. J. Surg. Pathol. 2017, 41, 299–312. [Google Scholar] [CrossRef]
- Miller, D.V.; Firchau, D.J.; McClure, R.F.; Kurtin, P.J.; Feldman, A.L. Epstein-Barr virus-associated diffuse large B-cell lymphoma arising on cardiac prostheses. Am. J Surg. Pathol. 2010, 34, 377–384. [Google Scholar] [CrossRef]
- Loong, F.; Chan, A.C.L.; Ho, B.C.S.; Chau, Y.-P.; Lee, H.-Y.; Cheuk, W.; Yuen, W.-K.; Ng, W.-S.; Cheung, H.-L.; Chan, J.K.C. Diffuse large B-cell lymphoma associated with chronic inflammation as an incidental finding and new clinical scenarios. Mod. Pathol. 2010, 23, 493–501. [Google Scholar] [CrossRef]
- King, R.L.; Goodlad, J.R.; Calaminici, M.; Dotlic, S.; Montes-Moreno, S.; Oschlies, I.; Ponzoni, M.; Traverse-Glehen, A.; Ott, G.; Ferry, J.A. Lymphomas arising in immune-privileged sites: Insights into biology, diagnosis, and pathogenesis. Virchows Arch. 2020, 476, 647–665. [Google Scholar] [CrossRef]
- Baugh, L.; Brown, N.; Song, J.Y.; Pandya, S.; Montoya, V.; Perry, A.M. Fibrin-Associated, EBV-Negative Diffuse Large B-Cell Lymphoma Arising in Atrial Myxoma: Expanding the Spectrum of the Entity. Int. J. Surg. Pathol. 2022, 30, 39–45. [Google Scholar] [CrossRef]
- Kaji, D.; Ota, Y.; Sato, Y.; Nagafuji, K.; Ueda, Y.; Okamoto, M.; Terasaki, Y.; Tsuyama, N.; Matsue, K.; Kinoshita, T.; et al. Primary human herpesvirus 8-negative effusion-based lymphoma: A large B-cell lymphoma with favorable prognosis. Blood Adv. 2020, 4, 4442–4450. [Google Scholar] [CrossRef]
- Alexanian, S.; Said, J.; Lones, M.; Pullarkat, S.T. KSHV/HHV8-negative effusion-based lymphoma, a distinct entity associated with fluid overload states. Am. J. Surg. Pathol. 2013, 37, 241–249. [Google Scholar] [CrossRef]
- Wu, W.; Youm, W.; Rezk, S.A.; Zhao, X. Human herpesvirus 8-unrelated primary effusion lymphoma-like lymphoma: Report of a rare case and review of 54 cases in the literature. Am. J. Clin. Pathol. 2013, 140, 258–273. [Google Scholar] [CrossRef]
- Kubota, T.; Sasaki, Y.; Shiozawa, E.; Takimoto, M.; Hishima, T.; Chong, J.-M. Age and CD20 Expression Are Significant Prognostic Factors in Human Herpes Virus-8-negative Effusion-based Lymphoma. Am. J. Surg. Pathol. 2018, 42, 1607–1616. [Google Scholar] [CrossRef]
- Mendeville, M.; Roemer, M.G.M.; van den Hout, M.F.C.M.; Los-de Vries, G.T.; Bladergroen, R.; Stathi, P.; Hijmering, N.J.; Rosenwald, A.; Ylstra, B.; de Jong, D. Aggressive genomic features in clinically indolent primary HHV8-negative effusion-based lymphoma. Blood 2019, 133, 377–380. [Google Scholar] [CrossRef]
- Delecluse, H.J.; Anagnostopoulos, I.; Dallenbach, F.; Hummel, M.; Marafioti, T.; Schneider, U.; Huhn, D.; Schmidt-Westhausen, A.; Reichart, P.A.; Gross, U.; et al. Plasmablastic lymphomas of the oral cavity: A new entity associated with the human immunodeficiency virus infection. Blood 1997, 89, 1413–1420. [Google Scholar] [CrossRef]
- Montes-Moreno, S.; Gonzalez-Medina, A.R.; Rodriguez-Pinilla, S.M.; Maestre, L.; Sanchez-Verde, L.; Roncador, G.; Mollejo, M.; Garcia, J.F.; Menarguez, J.; Montalban, C.; et al. Aggressive large B-cell lymphoma with plasma cell differentiation: Immunohistochemical characterization of plasmablastic lymphoma and diffuse large B-cell lymphoma with partial plasmablastic phenotype. Haematologica 2010, 95, 1342–1349. [Google Scholar] [CrossRef]
- Montes-Moreno, S.; Martinez-Magunacelaya, N.; Zecchini-Barrese, T.; de Villambrosía, S.G.; Linares, E.; Ranchal, T.; Rodriguez-Pinilla, M.; Batlle, A.; Cereceda-Company, L.; Revert-Arce, J.B.; et al. Plasmablastic lymphoma phenotype is determined by genetic alterations in MYC and PRDM1. Mod. Pathol. 2017, 30, 85–94. [Google Scholar] [CrossRef]
- Laurent, C.; Fabiani, B.; Do, C.; Tchernonog, E.; Cartron, G.; Gravelle, P.; Amara, N.; Malot, S.; Palisoc, M.M.; Copie-Bergman, C.; et al. Immune-checkpoint expression in Epstein-Barr virus positive and negative plasmablastic lymphoma: A clinical and pathological study in 82 patients. Haematologica 2016, 101, 976–984. [Google Scholar] [CrossRef]
- Castillo, J.J.; Bibas, M.; Miranda, R.N. The biology and treatment of plasmablastic lymphoma. Blood 2015, 125, 2323–2330. [Google Scholar] [CrossRef] [PubMed]
- Morscio, J.; Dierickx, D.; Nijs, J.; Verhoef, G.; Bittoun, E.; Vanoeteren, X.; Wlodarska, I.; Sagaert, X.; Tousseyn, T. Clinicopathologic comparison of plasmablastic lymphoma in HIV-positive, immunocompetent, and posttransplant patients: Single-center series of 25 cases and meta-analysis of 277 reported cases. Am. J. Surg. Pathol. 2014, 38, 875–886. [Google Scholar] [CrossRef] [PubMed]
- Loghavi, S.; Alayed, K.; Aladily, T.N.; Zuo, Z.; Ng, S.-B.; Tang, G.; Hu, S.; Yin, C.C.; Miranda, R.N.; Medeiros, L.J.; et al. Stage, age, and EBV status impact outcomes of plasmablastic lymphoma patients: A clinicopathologic analysis of 61 patients. J. Hematol. Oncol. 2015, 8, 65. [Google Scholar] [CrossRef]
- Boy, S.C.; van Heerden, M.B.; Raubenheimer, E.J.; van Heerden, W.F.P. Plasmablastic lymphomas with light chain restriction-plasmablastic extramedullary plasmacytomas? J. Oral Pathol. Med. 2010, 39, 435–439. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, F.P.; Robinson, L.; van Heerden, M.B.; van Heerden, W.F.P. Oral plasmablastic lymphoma: A clinicopathological study of 113 cases. J. Oral Pathol. Med. 2021, 50, 594–602. [Google Scholar] [CrossRef]
- Frontzek, F.; Staiger, A.M.; Zapukhlyak, M.; Xu, W.; Bonzheim, I.; Borgmann, V.; Sander, P.; Baptista, M.J.; Heming, J.-N.; Berning, P.; et al. Molecular and functional profiling identifies therapeutically targetable vulnerabilities in plasmablastic lymphoma. Nat. Commun. 2021, 12, 1413. [Google Scholar] [CrossRef]
- Chapman, J.; Gentles, A.J.; Sujoy, V.; Vega, F.; Dumur, C.I.; Blevins, T.L.; Bernal-Mizrachi, L.; Mosunjac, M.; Pimentel, A.; Zhu, D.; et al. Gene expression analysis of plasmablastic lymphoma identifies downregulation of B-cell receptor signaling and additional unique transcriptional programs. Leukemia 2015, 29, 2270–2273. [Google Scholar] [CrossRef]
- Ramis-Zaldivar, J.E.; Gonzalez-Farre, B.; Nicolae, A.; Pack, S.; Clot, G.; Nadeu, F.; Mottok, A.; Horn, H.; Song, J.Y.; Fu, K.; et al. MAPK and JAK-STAT pathways dysregulation in plasmablastic lymphoma. Haematologica 2021, 106, 2682–2693. [Google Scholar] [CrossRef]
- Vega, F.; Chang, C.-C.; Medeiros, L.J.; Udden, M.M.; Cho-Vega, J.H.; Lau, C.-C.; Finch, C.J.; Vilchez, R.A.; McGregor, D.; Jorgensen, J.L. Plasmablastic lymphomas and plasmablastic plasma cell myelomas have nearly identical immunophenotypic profiles. Mod. Pathol. 2005, 18, 806–815. [Google Scholar] [CrossRef]
- Ambrosio, M.R.; de Falco, G.; Gozzetti, A.; Rocca, B.J.; Amato, T.; Mourmouras, V.; Gazaneo, S.; Mundo, L.; Candi, V.; Piccaluga, P.P.; et al. Plasmablastic transformation of a pre-existing plasmacytoma: A possible role for reactivation of Epstein Barr virus infection. Haematologica 2014, 99, e235–e237. [Google Scholar] [CrossRef]
- Liu, Y.; Jelloul, F.; Zhang, Y.; Bhavsar, T.; Ho, C.; Rao, M.; Lewis, N.E.; Cimera, R.; Baik, J.; Sigler, A.; et al. Genetic Basis of Extramedullary Plasmablastic Transformation of Multiple Myeloma. Am. J. Surg. Pathol. 2020, 44, 838–848. [Google Scholar] [CrossRef]
- Riemersma, S.A.; Jordanova, E.S.; Schop, R.F.; Philippo, K.; Looijenga, L.H.; Schuuring, E.; Kluin, P.M. Extensive genetic alterations of the HLA region, including homozygous deletions of HLA class II genes in B-cell lymphomas arising in immune-privileged sites. Blood 2000, 96, 3569–3577. [Google Scholar] [CrossRef]
- Alame, M.; Cornillot, E.; Cacheux, V.; Rigau, V.; Costes-Martineau, V.; Lacheretz-Szablewski, V.; Colinge, J. The immune contexture of primary central nervous system diffuse large B cell lymphoma associates with patient survival and specific cell signaling. Theranostics 2021, 11, 3565–3579. [Google Scholar] [CrossRef]
- Küker, W.; Nägele, T.; Korfel, A.; Heckl, S.; Thiel, E.; Bamberg, M.; Weller, M.; Herrlinger, U. Primary central nervous system lymphomas (PCNSL): MRI features at presentation in 100 patients. J. Neurooncol. 2005, 72, 169–177. [Google Scholar] [CrossRef]
- Coupland, S.E.; Damato, B. Understanding intraocular lymphomas. Clin. Exp. Ophthalmol. 2008, 36, 564–578. [Google Scholar] [CrossRef]
- Montesinos-Rongen, M.; Küppers, R.; Schlüter, D.; Spieker, T.; van Roost, D.; Schaller, C.; Reifenberger, G.; Wiestler, O.D.; Deckert-Schlüter, M. Primary central nervous system lymphomas are derived from germinal-center B cells and show a preferential usage of the V4-34 gene segment. Am. J. Pathol. 1999, 155, 2077–2086. [Google Scholar] [CrossRef]
- Montesinos-Rongen, M.; Purschke, F.; Küppers, R.; Deckert, M. Immunoglobulin repertoire of primary lymphomas of the central nervous system. J. Neuropathol. Exp. Neurol. 2014, 73, 1116–1125. [Google Scholar] [CrossRef]
- Belhouachi, N.; Xochelli, A.; Boudjoghra, M.; Lesty, C.; Cassoux, N.; Fardeau, C.; Tran, T.H.C.; Choquet, S.; Sarker, B.; Houillier, C.; et al. Primary vitreoretinal lymphomas display a remarkably restricted immunoglobulin gene repertoire. Blood Adv. 2020, 4, 1357–1366. [Google Scholar] [CrossRef] [PubMed]
- Coupland, S.E.; Hummel, M.; Müller, H.-H.; Stein, H. Molecular analysis of immunoglobulin genes in primary intraocular lymphoma. Investig. Ophthalmol. Vis. Sci. 2005, 46, 3507–3514. [Google Scholar] [CrossRef]
- Montesinos-Rongen, M.; Godlewska, E.; Brunn, A.; Wiestler, O.D.; Siebert, R.; Deckert, M. Activating L265P mutations of the MYD88 gene are common in primary central nervous system lymphoma. Acta Neuropathol. 2011, 122, 791–792. [Google Scholar] [CrossRef]
- Montesinos-Rongen, M.; Schafer, E.; Siebert, R.; Deckert, M. Genes regulating the B cell receptor pathway are recurrently mutated in primary central nervous system lymphoma. Acta Neuropathol. 2012, 124, 905–906. [Google Scholar] [CrossRef]
- Cheah, C.Y.; Wirth, A.; Seymour, J.F. Primary testicular lymphoma. Blood 2014, 123, 486–493. [Google Scholar] [CrossRef] [PubMed]
- Vater, I.; Montesinos-Rongen, M.; Schlesner, M.; Haake, A.; Purschke, F.; Sprute, R.; Mettenmeyer, N.; Nazzal, I.; Nagel, I.; Gutwein, J.; et al. The mutational pattern of primary lymphoma of the central nervous system determined by whole-exome sequencing. Leukemia 2015, 29, 677–685. [Google Scholar] [CrossRef]
- Chapuy, B.; Roemer, M.G.M.; Stewart, C.; Tan, Y.; Abo, R.P.; Zhang, L.; Dunford, A.J.; Meredith, D.M.; Thorner, A.R.; Jordanova, E.S.; et al. Targetable genetic features of primary testicular and primary central nervous system lymphomas. Blood 2016, 127, 869–881. [Google Scholar] [CrossRef]
- Fontanilles, M.; Marguet, F.; Bohers, É.; Viailly, P.-J.; Dubois, S.; Bertrand, P.; Camus, V.; Mareschal, S.; Ruminy, P.; Maingonnat, C.; et al. Non-invasive detection of somatic mutations using next-generation sequencing in primary central nervous system lymphoma. Oncotarget 2017, 8, 48157–48168. [Google Scholar] [CrossRef]
- Bödör, C.; Alpár, D.; Marosvári, D.; Galik, B.; Rajnai, H.; Bátai, B.; Nagy, Á.; Kajtár, B.; Burján, A.; Deák, B.; et al. Molecular Subtypes and Genomic Profile of Primary Central Nervous System Lymphoma. J. Neuropathol. Exp. Neurol. 2020, 79, 176–183. [Google Scholar] [CrossRef]
- Bonzheim, I.; Giese, S.; Deuter, C.; Süsskind, D.; Zierhut, M.; Waizel, M.; Szurman, P.; Federmann, B.; Schmidt, J.; Quintanilla-Martinez, L.; et al. High frequency of MYD88 mutations in vitreoretinal B-cell lymphoma: A valuable tool to improve diagnostic yield of vitreous aspirates. Blood 2015, 126, 76–79. [Google Scholar] [CrossRef] [PubMed]
- Hübschmann, D.; Kleinheinz, K.; Wagener, R.; Bernhart, S.H.; López, C.; Toprak, U.H.; Sungalee, S.; Ishaque, N.; Kretzmer, H.; Kreuz, M.; et al. Mutational mechanisms shaping the coding and noncoding genome of germinal center derived B-cell lymphomas. Leukemia 2021, 35, 2002–2016. [Google Scholar] [CrossRef]
- Montesinos-Rongen, M.; van Roost, D.; Schaller, C.; Wiestler, O.D.; Deckert, M. Primary diffuse large B-cell lymphomas of the central nervous system are targeted by aberrant somatic hypermutation. Blood 2004, 103, 1869–1875. [Google Scholar] [CrossRef]
- Cobbers, J.M.; Wolter, M.; Reifenberger, J.; Ring, G.U.; Jessen, F.; An, H.X.; Niederacher, D.; Schmidt, E.E.; Ichimura, K.; Floeth, F.; et al. Frequent inactivation of CDKN2A and rare mutation of TP53 in PCNSL. Brain Pathol. 1998, 8, 263–276. [Google Scholar] [CrossRef] [PubMed]
- Schwindt, H.; Vater, I.; Kreuz, M.; Montesinos-Rongen, M.; Brunn, A.; Richter, J.; Gesk, S.; Ammerpohl, O.; Wiestler, O.D.; Hasenclever, D.; et al. Chromosomal imbalances and partial uniparental disomies in primary central nervous system lymphoma. Leukemia 2009, 23, 1875–1884. [Google Scholar] [CrossRef] [PubMed]
- Bonzheim, I.; Sander, P.; Salmerón-Villalobos, J.; Süsskind, D.; Szurman, P.; Gekeler, F.; Spitzer, M.S.; Steinhilber, J.; Kohler, E.; Büssgen, M.; et al. The molecular hallmarks of primary and secondary vitreoretinal lymphoma. Blood Adv. 2022, 6, 1598–1607. [Google Scholar] [CrossRef] [PubMed]
- Kodama, K.; Massone, C.; Chott, A.; Metze, D.; Kerl, H.; Cerroni, L. Primary cutaneous large B-cell lymphomas: Clinicopathologic features, classification, and prognostic factors in a large series of patients. Blood 2005, 106, 2491–2497. [Google Scholar] [CrossRef]
- Zinzani, P.L.; Quaglino, P.; Pimpinelli, N.; Berti, E.; Baliva, G.; Rupoli, S.; Martelli, M.; Alaibac, M.; Borroni, G.; Chimenti, S.; et al. Prognostic factors in primary cutaneous B-cell lymphoma: The Italian Study Group for Cutaneous Lymphomas. J. Clin. Oncol. 2006, 24, 1376–1382. [Google Scholar] [CrossRef]
- Senff, N.J.; Hoefnagel, J.J.; Jansen, P.M.; Vermeer, M.H.; van Baarlen, J.; Blokx, W.A.; Canninga-van Dijk, M.R.; Geerts, M.-L.; Hebeda, K.M.; Kluin, P.M.; et al. Reclassification of 300 primary cutaneous B-Cell lymphomas according to the new WHO-EORTC classification for cutaneous lymphomas: Comparison with previous classifications and identification of prognostic markers. J. Clin. Oncol. 2007, 25, 1581–1587. [Google Scholar] [CrossRef] [PubMed]
- Mareschal, S.; Pham-Ledard, A.; Viailly, P.J.; Dubois, S.; Bertrand, P.; Maingonnat, C.; Fontanilles, M.; Bohers, E.; Ruminy, P.; Tournier, I.; et al. Identification of Somatic Mutations in Primary Cutaneous Diffuse Large B-Cell Lymphoma, Leg Type by Massive Parallel Sequencing. J. Investig. Dermatol. 2017, 137, 1984–1994. [Google Scholar] [CrossRef]
- Vermeer, M.H.; Geelen, F.A.; van Haselen, C.W.; Van, V.; Geerts, M.L.; van Vloten, W.A.; Willemze, R. Primary cutaneous large B-cell lymphomas of the legs. A distinct type of cutaneous B-cell lymphoma with an intermediate prognosis. Dutch Cutaneous Lymphoma Working Group. Arch Dermatol. 1996, 132, 1304–1308. [Google Scholar] [CrossRef]
- Dobos, G.; de Masson, A.; Ram-Wolff, C.; Beylot-Barry, M.; Pham-Ledard, A.; Ortonne, N.; Ingen-Housz-Oro, S.; Battistella, M.; d’Incan, M.; Rouanet, J.; et al. Epidemiological changes in cutaneous lymphomas: An analysis of 8593 patients from the French Cutaneous Lymphoma Registry. Br. J. Dermatol. 2021, 184, 1059–1067. [Google Scholar] [CrossRef]
- Goodlad, J.R.; Krajewski, A.S.; Batstone, P.J.; McKay, P.; White, J.M.; Benton, E.C.; Kavanagh, G.M.; Lucraft, H.H. Primary cutaneous diffuse large B-cell lymphoma: Prognostic significance of clinicopathological subtypes. Am. J. Surg. Pathol. 2003, 27, 1538–1545. [Google Scholar] [CrossRef]
- Hoefnagel, J.J.; Vermeer, M.H.; Jansen, P.M.; Fleuren, G.J.; Meijer, C.J.L.M.; Willemze, R. Bcl-2, Bcl-6 and CD10 expression in cutaneous B-cell lymphoma: Further support for a follicle centre cell origin and differential diagnostic significance. Br. J. Dermatol. 2003, 149, 1183–1191. [Google Scholar] [CrossRef]
- Grange, F.; Petrella, T.; Beylot-Barry, M.; Joly, P.; D’Incan, M.; Delaunay, M.; Machet, L.; Avril, M.-F.; Dalac, S.; Bernard, P.; et al. Bcl-2 protein expression is the strongest independent prognostic factor of survival in primary cutaneous large B-cell lymphomas. Blood 2004, 103, 3662–3668. [Google Scholar] [CrossRef]
- Koens, L.; Vermeer, M.H.; Willemze, R.; Jansen, P.M. IgM expression on paraffin sections distinguishes primary cutaneous large B-cell lymphoma, leg type from primary cutaneous follicle center lymphoma. Am. J. Surg. Pathol. 2010, 34, 1043–1048. [Google Scholar] [CrossRef]
- Demirkesen, C.; Tüzüner, N.; Esen, T.; Lebe, B.; Ozkal, S. The expression of IgM is helpful in the differentiation of primary cutaneous diffuse large B cell lymphoma and follicle center lymphoma. Leuk. Res. 2011, 35, 1269–1272. [Google Scholar] [CrossRef]
- Hallermann, C.; Kaune, K.M.; Gesk, S.; Martin-Subero, J.I.; Gunawan, B.; Griesinger, F.; Vermeer, M.H.; Santucci, M.; Pimpinelli, N.; Willemze, R.; et al. Molecular cytogenetic analysis of chromosomal breakpoints in the IGH, MYC, BCL6, and MALT1 gene loci in primary cutaneous B-cell lymphomas. J. Investig. Dermatol. 2004, 123, 213–219. [Google Scholar] [CrossRef]
- Pham-Ledard, A.; Prochazkova-Carlotti, M.; Andrique, L.; Cappellen, D.; Vergier, B.; Martinez, F.; Grange, F.; Petrella, T.; Beylot-Barry, M.; Merlio, J.P. Multiple genetic alterations in primary cutaneous large B-cell lymphoma, leg type support a common lymphomagenesis with activated B-cell-like diffuse large B-cell lymphoma. Mod. Pathol. 2014, 27, 402–411. [Google Scholar] [CrossRef]
- Schrader, A.M.R.; Jansen, P.M.; Vermeer, M.H.; Kleiverda, J.K.; Vermaat, J.S.P.; Willemze, R. High Incidence and Clinical Significance of MYC Rearrangements in Primary Cutaneous Diffuse Large B-Cell Lymphoma, Leg Type. Am. J. Surg. Pathol. 2018, 42, 1488–1494. [Google Scholar] [CrossRef]
- Menguy, S.; Laharanne, E.; Prochazkova-Carlotti, M.; Gros, A.; Vergier, B.; Parrens, M.; Beylot-Barry, M.; Pham-Ledard, A.; Merlio, J.-P. Challenges in Assessing MYC Rearrangement in Primary Cutaneous Diffuse Large B-Cell Lymphoma, Leg-Type. Am. J. Surg. Pathol. 2020, 44, 424–427. [Google Scholar] [CrossRef]
- Lucioni, M.; Pescia, C.; Bonometti, A.; Fraticelli, S.; Moltrasio, C.; Ramponi, A.; Riboni, R.; Roccio, S.; Ferrario, G.; Arcaini, L.; et al. Double expressor and double/triple hit status among primary cutaneous diffuse large B-cell lymphoma: A comparison between leg type and not otherwise specified subtypes. Hum. Pathol. 2021, 111, 1–9. [Google Scholar] [CrossRef]
- Dijkman, R.; Tensen, C.P.; Jordanova, E.S.; Knijnenburg, J.; Hoefnagel, J.J.; Mulder, A.A.; Rosenberg, C.; Raap, A.K.; Willemze, R.; Szuhai, K.; et al. Array-based comparative genomic hybridization analysis reveals recurrent chromosomal alterations and prognostic parameters in primary cutaneous large B-cell lymphoma. J. Clin. Oncol. 2006, 24, 296–305. [Google Scholar] [CrossRef]
- Senff, N.J.; Zoutman, W.H.; Vermeer, M.H.; Assaf, C.; Berti, E.; Cerroni, L.; Espinet, B.; de Misa Cabrera, R.F.; Geerts, M.-L.; Kempf, W.; et al. Fine-mapping chromosomal loss at 9p21: Correlation with prognosis in primary cutaneous diffuse large B-cell lymphoma, leg type. J. Investig. Dermatol. 2009, 129, 1149–1155. [Google Scholar] [CrossRef]
- Koens, L.; Zoutman, W.H.; Ngarmlertsirichai, P.; Przybylski, G.K.; Grabarczyk, P.; Vermeer, M.H.; Willemze, R.; Jansen, P.M.; Schmidt, C.A.; Tensen, C.P. Nuclear factor-κB pathway-activating gene aberrancies in primary cutaneous large B-cell lymphoma, leg type. J. Investig. Dermatol. 2014, 134, 290–292. [Google Scholar] [CrossRef] [PubMed]
- Pham-Ledard, A.; Beylot-Barry, M.; Barbe, C.; Leduc, M.; Petrella, T.; Vergier, B.; Martinez, F.; Cappellen, D.; Merlio, J.-P.; Grange, F. High frequency and clinical prognostic value of MYD88 L265P mutation in primary cutaneous diffuse large B-cell lymphoma, leg-type. JAMA Dermatol. 2014, 150, 1173–1179. [Google Scholar] [CrossRef]
- Zhou, X.A.; Louissaint, A.; Wenzel, A.; Yang, J.; Martinez-Escala, M.E.; Moy, A.P.; Morgan, E.A.; Paxton, C.N.; Hong, B.; Andersen, E.F.; et al. Genomic Analyses Identify Recurrent Alterations in Immune Evasion Genes in Diffuse Large B-Cell Lymphoma, Leg Type. J. Investig. Dermatol. 2018, 138, 2365–2376. [Google Scholar] [CrossRef] [PubMed]
- Ducharme, O.; Beylot-Barry, M.; Pham-Ledard, A.; Bohers, E.; Viailly, P.-J.; Bandres, T.; Faur, N.; Frison, E.; Vergier, B.; Jardin, F.; et al. Mutations of the B-Cell Receptor Pathway Confer Chemoresistance in Primary Cutaneous Diffuse Large B-Cell Lymphoma Leg Type. J. Investig. Dermatol. 2019, 139, 2334–2342.e8. [Google Scholar] [CrossRef] [PubMed]
- Ferreri, A.J.M.; Dognini, G.P.; Campo, E.; Willemze, R.; Seymour, J.F.; Bairey, O.; Martelli, M.; de Renz, A.O.; Doglioni, C.; Montalban, C.; et al. Variations in clinical presentation, frequency of hemophagocytosis and clinical behavior of intravascular lymphoma diagnosed in different geographical regions. Haematologica 2007, 92, 486–492. [Google Scholar] [CrossRef]
- Ponzoni, M.; Campo, E.; Nakamura, S. Intravascular large B-cell lymphoma: A chameleon with multiple faces and many masks. Blood 2018, 132, 1561–1567. [Google Scholar] [CrossRef]
- Liu, Z.; Zhang, Y.; Zhu, Y.; Zhang, W. Prognosis of Intravascular Large B Cell Lymphoma (IVLBCL): Analysis of 182 Patients from Global Case Series. Cancer Manag. Res. 2020, 12, 10531–10540. [Google Scholar] [CrossRef]
- Sakakibara, A.; Inagaki, Y.; Imaoka, E.; Sakai, Y.; Ito, M.; Ishikawa, E.; Shimada, S.; Shimada, K.; Suzuki, Y.; Nakamura, S.; et al. Divergence and heterogeneity of neoplastic PD-L1 expression: Two autopsy case reports of intravascular large B-cell lymphoma. Pathol. Int. 2019, 69, 148–154. [Google Scholar] [CrossRef]
- Gupta, G.K.; Jaffe, E.S.; Pittaluga, S. A study of PD-L1 expression in intravascular large B cell lymphoma: Correlation with clinical and pathological features. Histopathology 2019, 75, 282–286. [Google Scholar] [CrossRef]
- Murase, T.; Yamaguchi, M.; Suzuki, R.; Okamoto, M.; Sato, Y.; Tamaru, J.-I.; Kojima, M.; Miura, I.; Mori, N.; Yoshino, T.; et al. Intravascular large B-cell lymphoma (IVLBCL): A clinicopathologic study of 96 cases with special reference to the immunophenotypic heterogeneity of CD5. Blood 2007, 109, 478–485. [Google Scholar] [CrossRef]
- Ponzoni, M.; Arrigoni, G.; Gould, V.E.; Del Curto, B.; Maggioni, M.; Scapinello, A.; Paolino, S.; Cassisa, A.; Patriarca, C. Lack of CD 29 (beta1 integrin) and CD 54 (ICAM-1) adhesion molecules in intravascular lymphomatosis. Hum. Pathol. 2000, 31, 220–226. [Google Scholar] [CrossRef]
- Kinoshita, M.; Izumoto, S.; Hashimoto, N.; Kishima, H.; Kagawa, N.; Hashiba, T.; Chiba, Y.; Yoshimine, T. Immunohistochemical analysis of adhesion molecules and matrix metalloproteinases in malignant CNS lymphomas: A study comparing primary CNS malignant and CNS intravascular lymphomas. Brain Tumor Pathol. 2008, 25, 73–78. [Google Scholar] [CrossRef]
- Alon, R.; Shulman, Z. Chemokine triggered integrin activation and actin remodeling events guiding lymphocyte migration across vascular barriers. Exp. Cell Res. 2011, 317, 632–641. [Google Scholar] [CrossRef]
- Kasuya, A.; Fujiyama, T.; Shirahama, S.; Hashizume, H.; Tokura, Y. Decreased expression of homeostatic chemokine receptors in intravascular large B-cell lymphoma. Eur. J. Dermatol. 2012, 22, 272–273. [Google Scholar] [CrossRef]
- Shimada, K.; Shimada, S.; Sugimoto, K.; Nakatochi, M.; Suguro, M.; Hirakawa, A.; Hocking, T.D.; Takeuchi, I.; Tokunaga, T.; Takagi, Y.; et al. Development and analysis of patient-derived xenograft mouse models in intravascular large B-cell lymphoma. Leukemia 2016, 30, 1568–1579. [Google Scholar] [CrossRef]
- Schrader, A.M.R.; Jansen, P.M.; Willemze, R.; Vermeer, M.H.; Cleton-Jansen, A.-M.; Somers, S.F.; Veelken, H.; van Eijk, R.; Kraan, W.; Kersten, M.J.; et al. High prevalence of MYD88 and CD79B mutations in intravascular large B-cell lymphoma. Blood 2018, 131, 2086–2089. [Google Scholar] [CrossRef]
- Suehara, Y.; Sakata-Yanagimoto, M.; Hattori, K.; Nanmoku, T.; Itoh, T.; Kaji, D.; Yamamoto, G.; Abe, Y.; Narita, K.; Takeuchi, M.; et al. Liquid biopsy for the identification of intravascular large B-cell lymphoma. Haematologica 2018, 103, e241–e244. [Google Scholar] [CrossRef]
- Shimada, K.; Yoshida, K.; Suzuki, Y.; Iriyama, C.; Inoue, Y.; Sanada, M.; Kataoka, K.; Yuge, M.; Takagi, Y.; Kusumoto, S.; et al. Frequent genetic alterations in immune checkpoint-related genes in intravascular large B-cell lymphoma. Blood 2021, 137, 1491–1502. [Google Scholar] [CrossRef]
- Gonzalez-Farre, B.; Ramis-Zaldivar, J.E.; Castrejón de Anta, N.; Rivas-Delgado, A.; Nadeu, F.; Salmeron-Villalobos, J.; Enjuanes, A.; Karube, K.; Balagué, O.; Cobo, F.; et al. Intravascular Large B-Cell Lymphoma Genomic Profile Is Characterized by Alterations in Genes Regulating NF-κB and Immune Checkpoints. Am. J. Surg. Pathol. 2022, 47, 202–211. [Google Scholar] [CrossRef]
- Savage, K.J. Primary mediastinal large B-cell lymphoma. Blood 2022, 140, 955–970. [Google Scholar] [CrossRef]
- Yousem, S.A.; Weiss, L.M.; Warnke, R.A. Primary mediastinal non-Hodgkin’s lymphomas: A morphologic and immunologic study of 19 cases. Am. J. Clin. Pathol. 1985, 83, 676–680. [Google Scholar] [CrossRef] [PubMed]
- Möller, P.; Lämmler, B.; Herrmann, B.; Otto, H.F.; Moldenhauer, G.; Momburg, F. The primary mediastinal clear cell lymphoma of B-cell type has variable defects in MHC antigen expression. Immunology 1986, 59, 411–417. [Google Scholar] [PubMed]
- Paulli, M.; Strater, J.; Gianelli, U.; Rousset, M.T.; Gambacorta, M.; Orlandi, E.; Klersy, C.; Lavabre-Bertrand, T.; Morra, E.; Manegold, C.; et al. Mediastinal B-cell lymphoma: A study of its histomorphologic spectrum based on 109 cases. Hum. Pathol. 1999, 30, 178–187. [Google Scholar] [CrossRef] [PubMed]
- Higgins, J.P.; Warnke, R.A. CD30 expression is common in mediastinal large B-cell lymphoma. Am. J. Clin. Pathol. 1999, 112, 241–247. [Google Scholar] [CrossRef]
- Pileri, S.A.; Gaidano, G.; Zinzani, P.L.; Falini, B.; Gaulard, P.; Zucca, E.; Pieri, F.; Berra, E.; Sabattini, E.; Ascani, S.; et al. Primary mediastinal B-cell lymphoma: High frequency of BCL-6 mutations and consistent expression of the transcription factors OCT-2, BOB.1, and PU.1 in the absence of immunoglobulins. Am. J. Pathol. 2003, 162, 243–253. [Google Scholar] [CrossRef]
- Kim, H.-J.; Kim, H.K.; Park, G.; Min, S.K.; Cha, H.J.; Lee, H.; Choi, S.J.; Na, H.Y.; Choe, J.-Y.; Kim, J.E. Comparative pathologic analysis of mediastinal B-cell lymphomas: Selective expression of p63 but no GATA3 optimally differentiates primary mediastinal large B-cell lymphoma from classic Hodgkin lymphoma. Diagn. Pathol. 2019, 14, 133. [Google Scholar] [CrossRef]
- Copie-Bergman, C.; Gaulard, P.; Maouche-Chrétien, L.; Brière, J.; Haioun, C.; Alonso, M.A.; Roméo, P.H.; Leroy, K. The MAL gene is expressed in primary mediastinal large B-cell lymphoma. Blood 1999, 94, 3567–3575. [Google Scholar] [CrossRef]
- Calaminici, M.; Piper, K.; Lee, A.M.; Norton, A.J. CD23 expression in mediastinal large B-cell lymphomas. Histopathology 2004, 45, 619–624. [Google Scholar] [CrossRef]
- Dorfman, D.M.; Shahsafaei, A.; Alonso, M.A. Utility of CD200 immunostaining in the diagnosis of primary mediastinal large B cell lymphoma: Comparison with MAL, CD23, and other markers. Mod. Pathol. 2012, 25, 1637–1643. [Google Scholar] [CrossRef]
- Shi, M.; Roemer, M.G.M.; Chapuy, B.; Liao, X.; Sun, H.; Pinkus, G.S.; Shipp, M.A.; Freeman, G.J.; Rodig, S.J. Expression of programmed cell death 1 ligand 2 (PD-L2) is a distinguishing feature of primary mediastinal (thymic) large B-cell lymphoma and associated with PDCD1LG2 copy gain. Am. J. Surg. Pathol. 2014, 38, 1715–1723. [Google Scholar] [CrossRef]
- Gentry, M.; Bodo, J.; Durkin, L.; Hsi, E.D. Performance of a Commercially Available MAL Antibody in the Diagnosis of Primary Mediastinal Large B-Cell Lymphoma. Am. J. Surg. Pathol. 2017, 41, 189–194. [Google Scholar] [CrossRef]
- Rosenwald, A.; Wright, G.; Leroy, K.; Yu, X.; Gaulard, P.; Gascoyne, R.D.; Chan, W.C.; Zhao, T.; Haioun, C.; Greiner, T.C.; et al. Molecular diagnosis of primary mediastinal B cell lymphoma identifies a clinically favorable subgroup of diffuse large B cell lymphoma related to Hodgkin lymphoma. J. Exp. Med. 2003, 198, 851–862. [Google Scholar] [CrossRef]
- Savage, K.J.; Monti, S.; Kutok, J.L.; Cattoretti, G.; Neuberg, D.; de Leval, L.; Kurtin, P.; Dal Cin, P.; Ladd, C.; Feuerhake, F.; et al. The molecular signature of mediastinal large B-cell lymphoma differs from that of other diffuse large B-cell lymphomas and shares features with classical Hodgkin lymphoma. Blood 2003, 102, 3871–3879. [Google Scholar] [CrossRef]
- Mottok, A.; Wright, G.; Rosenwald, A.; Ott, G.; Ramsower, C.; Campo, E.; Braziel, R.M.; Delabie, J.; Weisenburger, D.D.; Song, J.Y.; et al. Molecular classification of primary mediastinal large B-cell lymphoma using routinely available tissue specimens. Blood 2018, 132, 2401–2405. [Google Scholar] [CrossRef]
- Weniger, M.A.; Gesk, S.; Ehrlich, S.; Martin-Subero, J.I.; Dyer, M.J.; Siebert, R.; Moller, P.; Barth, T.F. Gains of REL in primary mediastinal B-cell lymphoma coincide with nuclear accumulation of REL protein. Genes Chromosomes Cancer 2007, 46, 406–415. [Google Scholar] [CrossRef]
- Green, M.R.; Monti, S.; Rodig, S.J.; Juszczynski, P.; Currie, T.; O’Donnell, E.; Chapuy, B.; Takeyama, K.; Neuberg, D.; Golub, T.R.; et al. Integrative analysis reveals selective 9p24.1 amplification, increased PD-1 ligand expression, and further induction via JAK2 in nodular sclerosing Hodgkin lymphoma and primary mediastinal large B-cell lymphoma. Blood 2010, 116, 3268–3277. [Google Scholar] [CrossRef]
- Twa, D.D.; Chan, F.C.; Ben-Neriah, S.; Woolcock, B.W.; Mottok, A.; Tan, K.L.; Slack, G.W.; Gunawardana, J.; Lim, R.S.; McPherson, A.W.; et al. Genomic rearrangements involving programmed death ligands are recurrent in primary mediastinal large B-cell lymphoma. Blood 2014, 123, 2062–2065. [Google Scholar] [CrossRef]
- Steidl, C.; Shah, S.P.; Woolcock, B.W.; Rui, L.; Kawahara, M.; Farinha, P.; Johnson, N.A.; Zhao, Y.; Telenius, A.; Neriah, S.B.; et al. MHC class II transactivator CIITA is a recurrent gene fusion partner in lymphoid cancers. Nature 2011, 471, 377–381. [Google Scholar] [CrossRef]
- Traverse-Glehen, A.; Pittaluga, S.; Gaulard, P.; Sorbara, L.; Alonso, M.A.; Raffeld, M.; Jaffe, E.S. Mediastinal gray zone lymphoma: The missing link between classic Hodgkin’s lymphoma and mediastinal large B-cell lymphoma. Am. J. Surg. Pathol. 2005, 29, 1411–1421. [Google Scholar] [CrossRef]
- Pilichowska, M.; Pittaluga, S.; Ferry, J.A.; Hemminger, J.; Chang, H.; Kanakry, J.A.; Sehn, L.H.; Feldman, T.; Abramson, J.S.; Kritharis, A.; et al. Clinicopathologic consensus study of gray zone lymphoma with features intermediate between DLBCL and classical HL. Blood Adv. 2017, 1, 2600–2609. [Google Scholar] [CrossRef]
- Sarkozy, C.; Copie-Bergman, C.; Damotte, D.; Ben-Neriah, S.; Burroni, B.; Cornillon, J.; Lemal, R.; Golfier, C.; Fabiani, B.; Chassagne-Clément, C.; et al. Gray-zone Lymphoma Between cHL and Large B-Cell Lymphoma: A Histopathologic Series from the LYSA. Am. J. Surg. Pathol. 2019, 43, 341–351. [Google Scholar] [CrossRef] [PubMed]
- Sarkozy, C.; Hung, S.S.; Chavez, E.A.; Duns, G.; Takata, K.; Chong, L.C.; Aoki, T.; Jiang, A.; Miyata-Takata, T.; Telenius, A.; et al. Mutational landscape of gray zone lymphoma. Blood 2021, 137, 1765–1776. [Google Scholar] [CrossRef] [PubMed]
- Rimsza, L.; Pittaluga, S.; Dirnhofer, S.; Copie-Bergman, C.; de Leval, L.; Facchetti, F.; Pileri, S.; Rosenwald, A.; Wotherspoon, A.; Fend, F. The clinicopathologic spectrum of mature aggressive B cell lymphomas. Virchows Arch. 2017, 471, 453–466. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Liu, X.; Yao, Z.; Zhang, M. High-Grade B-Cell Lymphomas, Not Otherwise Specified: A Study of 41 Cases. Cancer Manag. Res. 2020, 12, 1903–1912. [Google Scholar] [CrossRef]
- Kanagal-Shamanna, R.; Medeiros, L.J.; Lu, G.; Wang, S.A.; Manning, J.T.; Lin, P.; Penn, G.M.; Young, K.H.; You, M.J.; Vega, F.; et al. High-grade B cell lymphoma, unclassifiable, with blastoid features: An unusual morphological subgroup associated frequently with BCL2 and/or MYC gene rearrangements and a poor prognosis. Histopathology 2012, 61, 945–954. [Google Scholar] [CrossRef]
- Moore, E.M.; Aggarwal, N.; Surti, U.; Swerdlow, S.H. Further Exploration of the Complexities of Large B-Cell Lymphomas with MYC Abnormalities and the Importance of a Blastoid Morphology. Am. J. Surg. Pathol. 2017, 41, 1155–1166. [Google Scholar] [CrossRef]
- Ott, G. Aggressive B-cell lymphomas in the update of the 4th edition of the World Health Organization classification of haematopoietic and lymphatic tissues: Refinements of the classification, new entities and genetic findings. Br. J. Haematol. 2017, 178, 871–887. [Google Scholar] [CrossRef]
- Hüttl, K.S.; Staiger, A.M.; Richter, J.; Ott, M.M.; Kalmbach, S.; Klapper, W.; Biesdorf, A.-S.; Trümper, L.; Rosenwald, A.; Ziepert, M.; et al. The “Burkitt-like” immunophenotype and genotype is rarely encountered in diffuse large B cell lymphoma and high-grade B cell lymphoma, NOS. Virchows Arch. 2021, 479, 575–583. [Google Scholar] [CrossRef]
- Gonzalez de Villambrosia, S.; Bastos, M.; Palanca, J.M.; Cruz, J.G.; Navarro, J.-T.; Tapia, G.; Alonso, S.A.; Martin, A.; Blanco, O.; Abrisqueta, P.; et al. BCL2 translocation in high grade B cell lymphoma (NOS, DH/TH) is associated with reduced progression free survival. Leuk. Lymphoma 2022, 63, 101–108. [Google Scholar] [CrossRef]
- Menter, T.; Juskevicius, D.; Alikian, M.; Steiger, J.; Dirnhofer, S.; Tzankov, A.; Naresh, K.N. Mutational landscape of B-cell post-transplant lymphoproliferative disorders. Br. J. Haematol. 2017, 178, 48–56. [Google Scholar] [CrossRef]
- Chapman, J.R.; Bouska, A.C.; Zhang, W.; Alderuccio, J.P.; Lossos, I.S.; Rimsza, L.M.; Maguire, A.; Yi, S.; Chan, W.C.; Vega, F.; et al. EBV-positive HIV-associated diffuse large B cell lymphomas are characterized by JAK/STAT (STAT3) pathway mutations and unique clinicopathologic features. Br. J. Haematol. 2021, 194, 870–878. [Google Scholar] [CrossRef]
- Morscio, J.; Dierickx, D.; Ferreiro, J.F.; Herreman, A.; van Loo, P.; Bittoun, E.; Verhoef, G.; Matthys, P.; Cools, J.; Wlodarska, I.; et al. Gene expression profiling reveals clear differences between EBV-positive and EBV-negative posttransplant lymphoproliferative disorders. Am. J. Transplant. 2013, 13, 1305–1316. [Google Scholar] [CrossRef]
- Ferreiro, J.F.; Morscio, J.; Dierickx, D.; Vandenberghe, P.; Gheysens, O.; Verhoef, G.; Zamani, M.; Tousseyn, T.; Wlodarska, I. EBV-Positive and EBV-Negative Posttransplant Diffuse Large B Cell Lymphomas Have Distinct Genomic and Transcriptomic Features. Am. J. Transplant. 2016, 16, 414–425. [Google Scholar] [CrossRef]
- Engels, E.A.; Pittaluga, S.; Whitby, D.; Rabkin, C.; Aoki, Y.; Jaffe, E.S.; Goedert, J.J. Immunoblastic lymphoma in persons with AIDS-associated Kaposi’s sarcoma: A role for Kaposi’s sarcoma-associated herpesvirus. Mod. Pathol. 2003, 16, 424–429. [Google Scholar] [CrossRef]
- Courville, E.L.; Sohani, A.R.; Hasserjian, R.P.; Zukerberg, L.R.; Harris, N.L.; Ferry, J.A. Diverse clinicopathologic features in human herpesvirus 8-associated lymphomas lead to diagnostic problems. Am. J. Clin. Pathol. 2014, 142, 816–829. [Google Scholar] [CrossRef]
- Gonzalez-Farre, B.; Martinez, D.; Lopez-Guerra, M.; Xipell, M.; Monclus, E.; Rovira, J.; Garcia, F.; Lopez-Guillermo, A.; Colomo, L.; Campo, E.; et al. HHV8-related lymphoid proliferations: A broad spectrum of lesions from reactive lymphoid hyperplasia to overt lymphoma. Mod. Pathol. 2017, 30, 745–760. [Google Scholar] [CrossRef]








| Diffuse large B-cell lymphoma, NOS |
| T-cell/histiocyte-rich large B-cell lymphoma |
| Diffuse large B-cell lymphoma/high-grade B-cell lymphoma with MYC and BCL2 rearrangements |
| ALK-positive large B-cell lymphoma |
| Large B-cell lymphoma with IRF4 rearrangement |
| High-grade B-cell lymphoma with 11q aberrations |
| Lymphomatoid granulomatosis |
| EBV-positive diffuse large B-cell lymphoma |
| Diffuse large B-cell lymphoma associated with chronic inflammation |
| Fibrin-associated large B-cell lymphoma |
| Fluid overload-associated large B-cell lymphoma |
| Plasmablastic lymphoma |
| Primary large B-cell lymphoma of immune-privileged sites |
| Primary cutaneous diffuse large B-cell lymphoma, leg type |
| Intravascular large B-cell lymphoma |
| Primary mediastinal large B-cell lymphoma |
| Mediastinal grey zone lymphoma |
| High-grade B-cell lymphoma, NOS |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kurz, K.S.; Ott, M.; Kalmbach, S.; Steinlein, S.; Kalla, C.; Horn, H.; Ott, G.; Staiger, A.M. Large B-Cell Lymphomas in the 5th Edition of the WHO-Classification of Haematolymphoid Neoplasms—Updated Classification and New Concepts. Cancers 2023, 15, 2285. https://doi.org/10.3390/cancers15082285
Kurz KS, Ott M, Kalmbach S, Steinlein S, Kalla C, Horn H, Ott G, Staiger AM. Large B-Cell Lymphomas in the 5th Edition of the WHO-Classification of Haematolymphoid Neoplasms—Updated Classification and New Concepts. Cancers. 2023; 15(8):2285. https://doi.org/10.3390/cancers15082285
Chicago/Turabian StyleKurz, Katrin S., Michaela Ott, Sabrina Kalmbach, Sophia Steinlein, Claudia Kalla, Heike Horn, German Ott, and Annette M. Staiger. 2023. "Large B-Cell Lymphomas in the 5th Edition of the WHO-Classification of Haematolymphoid Neoplasms—Updated Classification and New Concepts" Cancers 15, no. 8: 2285. https://doi.org/10.3390/cancers15082285
APA StyleKurz, K. S., Ott, M., Kalmbach, S., Steinlein, S., Kalla, C., Horn, H., Ott, G., & Staiger, A. M. (2023). Large B-Cell Lymphomas in the 5th Edition of the WHO-Classification of Haematolymphoid Neoplasms—Updated Classification and New Concepts. Cancers, 15(8), 2285. https://doi.org/10.3390/cancers15082285






