Explainable CAD System for Classification of Acute Lymphoblastic Leukemia Based on a Robust White Blood Cell Segmentation
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials
2.1. Datasets
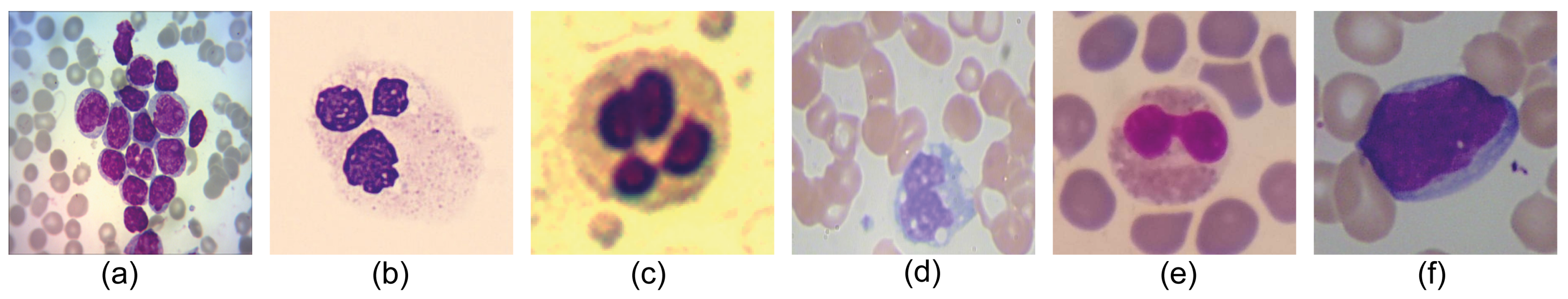
- Leukemia Dataset [16] is formed by 651 classified images of Acute Lymphoblastic Leukemia according to FAB classification (217-ALL1, 217-ALL2, 217-ALL3), with dimensions of pixels. This dataset is the only one in the state-of-the-art that labels the different types of Acute Lymphoblastic Leukemia with reliability through cytogenetic tests.
- CellaVision [22] is made up of 100 blood samples, and each image has dimensions of pixels and a bit depth of 24 bits. This dataset usually consists of a single cell, and the core color is violet, while the background has pinkish and yellowish tints.
- JTSC [22] is made by the Jiangxi Telecom Science Corporation in China. This dataset consists of 300 images of pixels containing the GT of the nucleus and cytoplasm for comparative analysis. It contains a wide variability among its samples since there are cells in which the nucleus has a highly saturated coloration, while in others, the nucleus is almost translucent. Furthermore, the image’s background varies from an intense yellow to a pinkish white.
- SMC_ID (Blood_Seg) [23] is composed of 367 images of WBC with a size of pixels. Each sample characterizes by the GT of the nucleus, which facilitates its analysis. Commonly, the images that integrate this dataset have a cell nucleus with low color saturation. Additionally, the WBC is located in diverse positions over the image.
- Raabin_WBC [24]. It provides 1145 images of blood samples, with dimensions of pixels, where white blood cells are subdivided into 242 lymphocytes, 242 monocytes, 242 neutrophils, 201 eosinophils, and 218 basophils. Each of these 1145 samples also contains a ground truth, both whole cell and nucleus. This is one of the best databases by now, as it has numerous samples of different cell types classified and annotated with ground truth for analysis and comparison of results.
- ALL_IDB2 [25]. It consists of 260 images of pixels. This dataset derives from the ALL-IDB1 dataset, where individual cells have been cropped to obtain the region of interest.
2.2. Metrics
- Accuracy value measures the appropriate classification over the total elements.
- Precision metric estimates the number of elements correctly classified among all the positive elements to evaluate.
- Recall also known as sensitivity, is used to denote the number of positive elements that are correctly classified.
- Specificity measures the proportion of true negatives that are successfully identified by the model.
- Dice Similarity Coefficient or DSC can be considered to be a harmonic mean of precision and recall. Furthermore, known as F1-Score.
- Intersection over Union also known as Jaccard Index is the most important metric in image segmentation tasks since it measures the magnitude of overlap between the GT and the segmented image.
3. Proposed Method
3.1. Handcrafted WBC Nuclei Segmentation
| Algorithm 1 Proposed Handcrafted WBC Nuclei Segmentation. |
|
3.2. Deep Learning WBC Nuclei Segmentation
3.3. Ensemble Segmentation
3.4. ALL Classification
3.5. Visual Explainability
3.6. Clustering Space Analysis
4. Results and Discussion
4.1. Segmentation Results
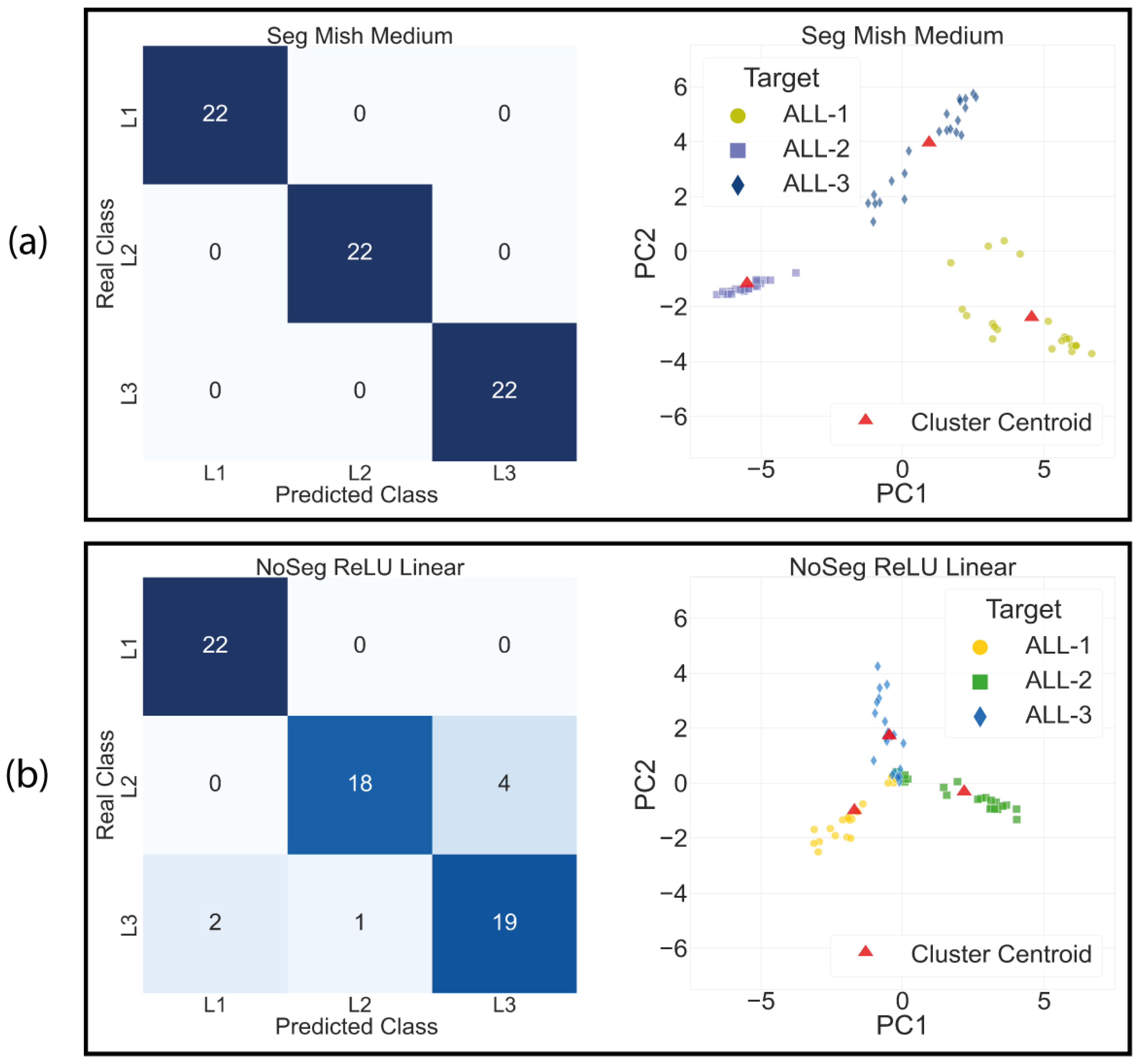
4.2. Leukemia Classification
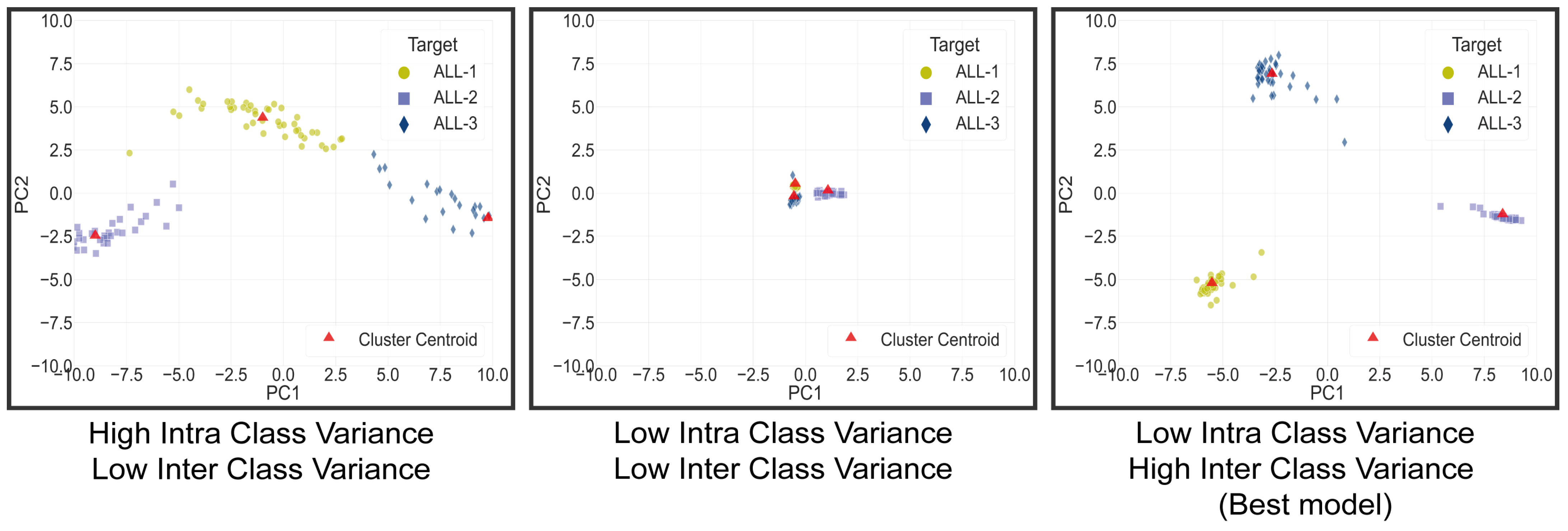
4.3. Clustering Space Analysis Results
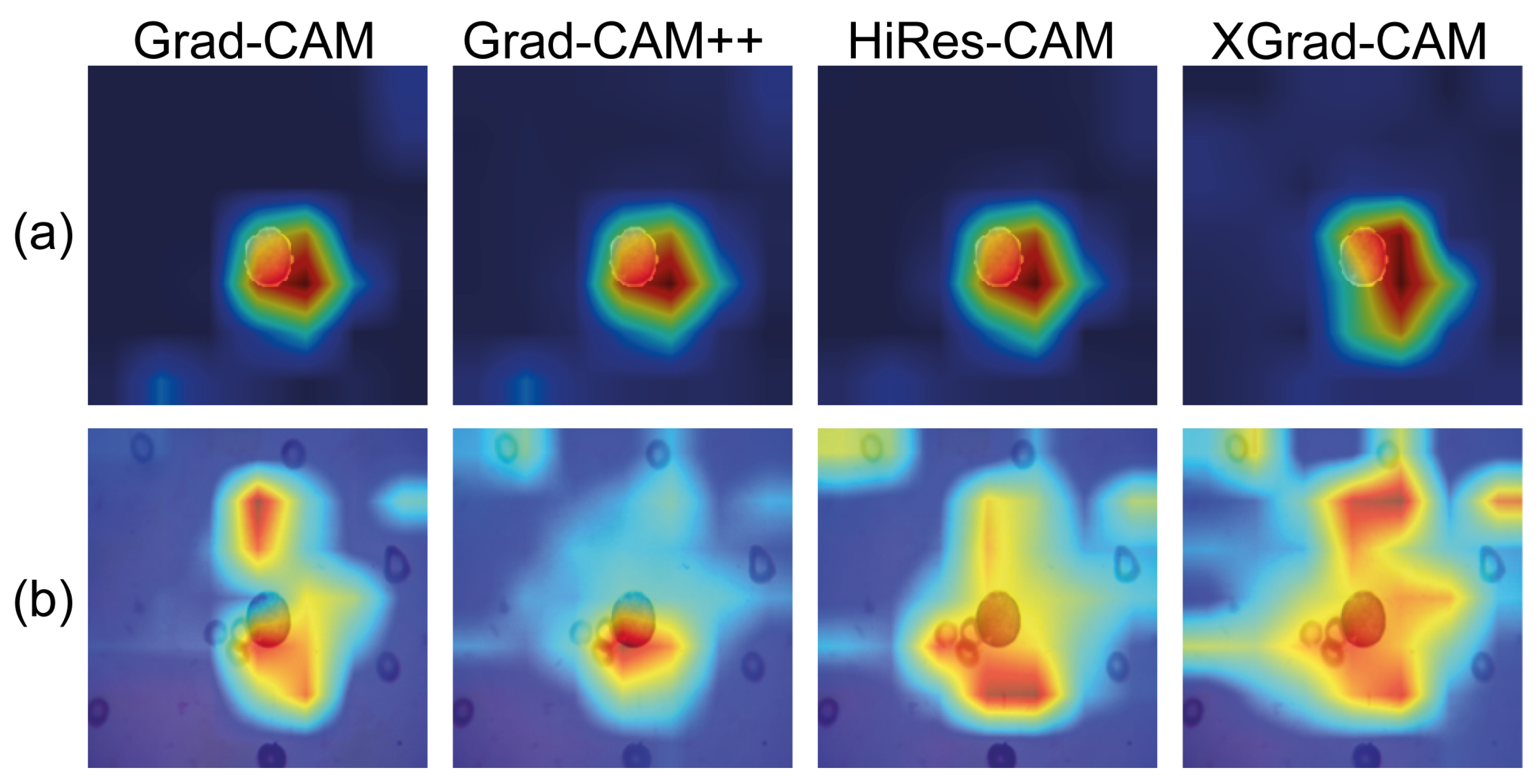
4.4. Class Activation Maps
4.5. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A. Segmentation and Classification Parameter’s and Extra Results
Appendix A.1. WBC Nuclei Segmentation
| Technique | Parameter |
|---|---|
| Bilateral Filter (Kernel size) | 9 |
| Bilateral Filter (radial, spatial sigma) | 50 |
| Closing Kernel | 3 |
| Area Filter | 150 |
| Dilation | 2 |
| Dilation (Raabin-Basophil) | 4 |
Appendix A.2. WBC Deep Learning Segmentation
| U-Net Train Parameters | ||||||
|---|---|---|---|---|---|---|
| Dataset | Lambda | Delta | Gamma | Learning Rate | Dropout | Weight Decay |
| Cellavision | 0.4 | 0.7 | 1.0 | 1 × 10−4 | 0.1 | 1 × 10−3 |
| JTSC | 0.5 | 0.8 | 1.0 | 1 × 10−4 | 0.05 | 1 × 10−4 |
| SMC_IDB | 0.6 | 0.7 | 1.0 | 1 × 10−4 | 0.05 | 1 × 10−3 |
| Raabin_WBC | 0.4 | 0.7 | 1.0 | 1 × 10−4 | 0.5 | 1 × 10−2 |
| ALL-IDB2 | 0.4 | 0.7 | 1.0 | 1 × 10−4 | 0.1 | 1 × 10−4 |
| Dataset Leukemia | 0.4 | 0.4 | 1.0 | 1 × 10−4 | 0.05 | 1 × 10−5 |

Appendix A.3. Deep Learning Classification
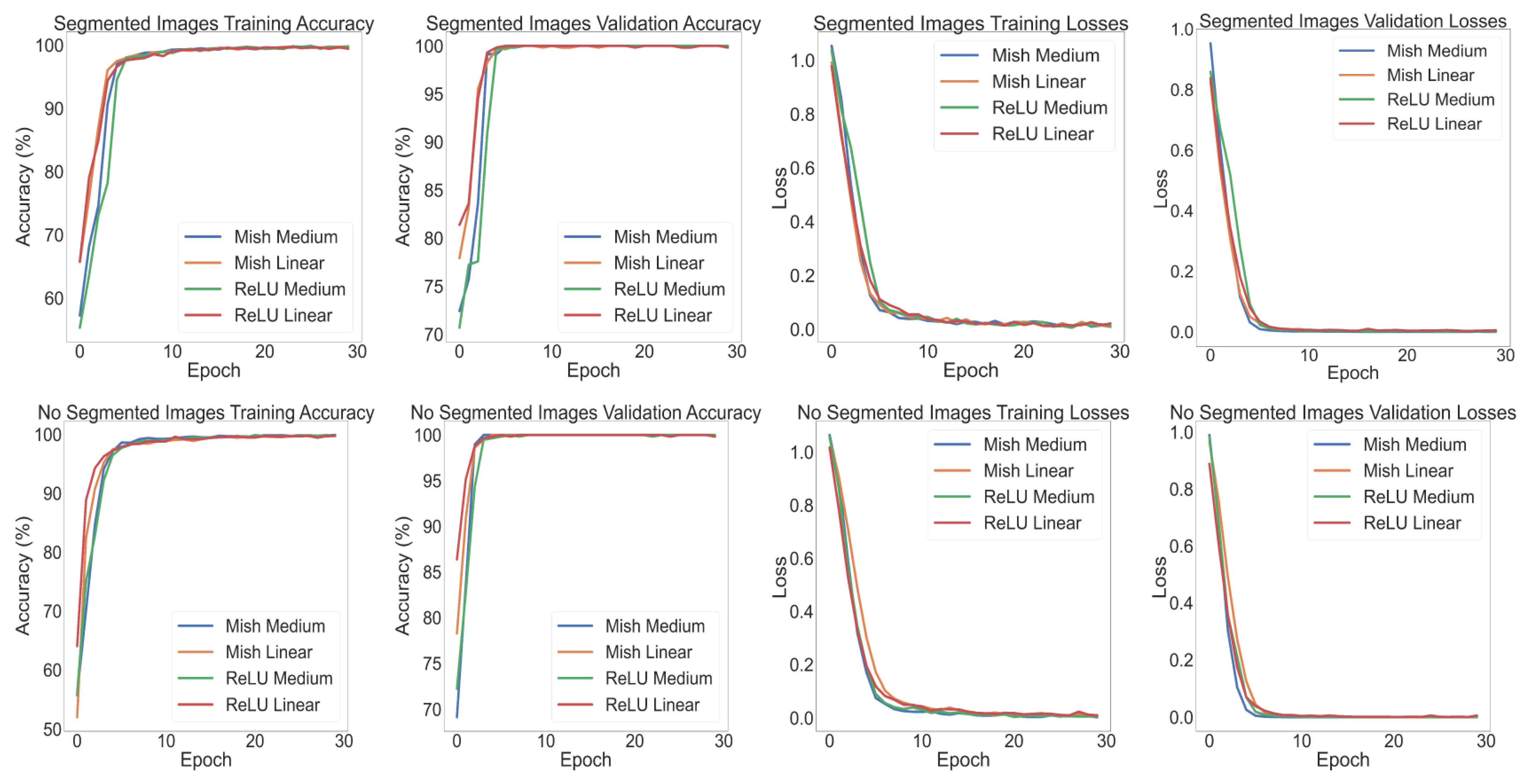
| Class Prediction Probability | |||||
|---|---|---|---|---|---|
| Image | Mish Linear-S | Mish Linear-NS | Mish Medium-S | Mish Medium-NS | |
| ALL-1 | 1_1_7 | 0.998 | 0.426 | 1.000 | 0.945 |
| 1_3_132 | 0.992 | 0.430 | 0.999 | 0.993 | |
| 1_3_158 | 0.998 | 0.464 | 1.000 | 0.993 | |
| ALL-2 | 2_1_33 | 0.998 | 0.750 | 1.000 | 0.991 |
| 2_2_127 | 0.999 | 0.734 | 1.000 | 0.990 | |
| 2_3_202 | 0.999 | 0.778 | 1.000 | 0.970 | |
| ALL-3 | 3_1_3 | 0.973 | 0.570 | 0.997 | 0.846 |
| 3_1_32 | 0.993 | 0.439 | 0.999 | 0.806 | |
| 3_2_11 | 0.998 | 0.492 | 1.000 | 0.838 | |
| Average | 0.994 | 0.565 | 0.999 | 0.930 | |
References
- Guyton, A.C.; Hall, J.E. Resistencia del organismo a la infección: I. Leucocitos, granulocitos, sistema monocitomacrofágico e inflamación. In Tratado de Fisiología Médica, 12th ed.; Elsevier: Barcelona, Spain, 2011; Chapter 34; pp. 1118–1139. [Google Scholar]
- Kumar, V.; Abul, A.; Jon, C. Hematopoietic and Lymphoid Systems. In Robins Basic Pathology; Elsevier: Amsterdam, The Netherlands, 2018; Chapter 12; pp. 459–467. [Google Scholar]
- Secretaria de Salud de México. Diagnóstico Oportuno de la Leucemia Aguda en Pediatría en Primer y Segundo Nivel de Atención; Technical Report; Secretaria de Salud: Ciudad de México, Mexico, 2017.
- Brereton, M.; De La Salle, B.; Ardern, J.; Hyde, K.; Burthem, J. Do We Know Why We Make Errors in Morphological Diagnosis? An Analysis of Approach and Decision-Making in Haematological Morphology. EBioMedicine 2015, 2, 1224–1234. [Google Scholar] [CrossRef] [PubMed]
- Loddo, A.; Putzu, L. On the Reliability of CNNs in Clinical Practice: A Computer-Aided Diagnosis System Case Study. Appl. Sci. 2022, 12, 3269. [Google Scholar] [CrossRef]
- Andrade, A.R.; Vogado, L.H.; Veras, R.d.M.; Silva, R.R.; Araujo, F.H.; Medeiros, F.N. Recent computational methods for white blood cell nuclei segmentation: A comparative study. Comput. Methods Programs Biomed. 2019, 173, 1–14. [Google Scholar] [CrossRef]
- Loddo, A.; Putzu, L. On the Effectiveness of Leukocytes Classification Methods in a Real Application Scenario. AI 2021, 2, 25. [Google Scholar] [CrossRef]
- Mahbod, A.; Tschandl, P.; Langs, G.; Ecker, R.; Ellinger, I. The effects of skin lesion segmentation on the performance of dermatoscopic image classification. Comput. Methods Programs Biomed. 2020, 197, 105725. [Google Scholar] [CrossRef] [PubMed]
- Al-masni, M.A.; Kim, D.H.; Kim, T.S. Multiple skin lesions diagnostics via integrated deep convolutional networks for segmentation and classification. Comput. Methods Programs Biomed. 2020, 190, 105351. [Google Scholar] [CrossRef]
- Vogado, L.H.; Veras, R.D.M.; Andrade, A.R.; De Araujo, F.H.; E Silva, R.R.; De Medeiros, F.N. Unsupervised leukemia cells segmentation based on multi-space color channels. In Proceedings of the 2016 IEEE International Symposium on Multimedia, ISM 2016, San Jose, CA, USA, 11–13 December 2016; pp. 451–456. [Google Scholar] [CrossRef]
- Makem, M.; Tiedeu, A. An efficient algorithm for detection of white blood cell nuclei using adaptive three stage PCA-based fusion. Inform. Med. Unlocked 2020, 20, 100416. [Google Scholar] [CrossRef]
- Mousavi, K.; Tavakoli, S.; Alipanah, A. Easy-GT: Open-Source Software to Facilitate Making the Ground Truth for White Blood Cells’ Nucleus. arXiv 2021, arXiv:2101.11654. [Google Scholar]
- Tavakoli, S.; Ghaffari, A.; Kouzehkanan, Z.M.; Hosseini, R. New segmentation and feature extraction algorithm for classification of white blood cells in peripheral smear images. Sci. Rep. 2021, 11, 19428. [Google Scholar] [CrossRef]
- Makem, M.; Tiedeu, A.; Kom, G.; Nkandeu, Y.P.K. A robust algorithm for white blood cell nuclei segmentation. Multimed. Tools Appl. 2022, 81, 17849–17874. [Google Scholar] [CrossRef]
- Mayala, S.; Haugsøen, J.B. Threshold estimation based on local minima for nucleus and cytoplasm segmentation. BMC Med. Imaging 2022, 22, 77. [Google Scholar] [CrossRef] [PubMed]
- Ochoa-montiel, R.; Ibarra, L.M.; Sossa, H.; Olague, G.; Polit, I. Handcraft and Automatic Approaches for the Recognition of Leukemia Images. Res. Comput. Sci. 2020, 149, 271–280. [Google Scholar]
- Haider, A.; Arsalan, M.; Lee, Y.W.; Park, K.R. Deep features aggregation-based joint segmentation of cytoplasm and nuclei in white blood cells. IEEE J. Biomed. Health Inform. 2022, 26, 3685–3696. [Google Scholar] [CrossRef]
- García-Lamont, F.; Alvarado, M.; López-Chau, A.; Cervantes, J. Efficient nucleus segmentation of white blood cells mimicking the human perception of color. Color Res. Appl. 2022, 47, 657–675. [Google Scholar] [CrossRef]
- Zhou, Z.; Siddiquee, M.R.; Tajbakhsh, N. UNet++: A Nested U-Net Architecture. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer International Publishing: Berlin/Heidelberg, Germany, 2018; pp. 3–11. [Google Scholar] [CrossRef]
- Oktay, O.; Schlemper, J.; Folgoc, L.L.; Lee, M.; Heinrich, M.; Misawa, K.; Mori, K.; McDonagh, S.; Hammerla, N.Y.; Kainz, B.; et al. Attention U-Net: Learning Where to Look for the Pancreas. arXiv 2018, arXiv:1804.03999. [Google Scholar]
- He, K.; Gkioxari, G.; Dollár, P.; Girshick, R. Mask R-CNN. IEEE Trans. Pattern Anal. Mach. Intell. 2020, 42, 386–397. [Google Scholar] [CrossRef]
- Zheng, X.; Wang, Y.; Wang, G.; Liu, J. Fast and robust segmentation of white blood cell images by self-supervised learning. Micron 2018, 107, 55–71. [Google Scholar] [CrossRef]
- Mohamed, M.; Far, B.; Guaily, A. An efficient technique for white blood cells nuclei automatic segmentation. In Proceedings of the IEEE International Conference on Systems, Man and Cybernetics, Seoul, Republic of Korea, 14–17 October 2012; pp. 220–225. [Google Scholar] [CrossRef]
- Kouzehkanan, Z.M.; Saghari, S.; Tavakoli, S.; Rostami, P.; Abaszadeh, M.; Mirzadeh, F.; Satlsar, E.S.; Gheidishahran, M.; Gorgi, F.; Mohammadi, S.; et al. A large dataset of white blood cells containing cell locations and types, along with segmented nuclei and cytoplasm. Sci. Rep. 2022, 12, 1123. [Google Scholar] [CrossRef]
- Labati, R.D.; Piuri, V.; Scotti, F. All-IDB: The Acute Lymphoblastic Leukemia Image Database for Image Processing. In Proceedings of the IEEE International Conference On Image Processing, Brussels, Belgium, 11–14 September 2011; pp. 2089–2092. [Google Scholar]
- Müller, D.; Soto-Rey, I.; Kramer, F. Towards a guideline for evaluation metrics in medical image segmentation. BMC Res. Notes 2022, 15, 1–7. [Google Scholar] [CrossRef]
- Hegde, R.B.; Prasad, K.; Hebbar, H.; Singh, B.M.K. Comparison of traditional image processing and deep learning approaches for classification of white blood cells in peripheral blood smear images. Biocybern. Biomed. Eng. 2019, 39, 382–392. [Google Scholar] [CrossRef]
- Paris, S.; Kornprobst, P.; Tumblin, J.; Durand, F. Bilateral filtering: Theory and applications. Found. Trends Comput. Graph. Vis. 2009, 4, 1–73. [Google Scholar] [CrossRef]
- Otsu, N. A Threshold Selection Method from Gray-Level Histograms. IEEE Trans. Syst. Man Cybern. 1979, 20, 62–66. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015, Munich, Germany, 5–9 October 2015; Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F., Eds.; Springer International Publishing: Cham, Switzerland, 2015; pp. 234–241. [Google Scholar]
- Misra, D. Mish: A Self Regularized Non-Monotonic Activation Function. arXiv 2019, arXiv:1908.08681. [Google Scholar]
- Yeung, M.; Sala, E.; Schönlieb, C.B.; Rundo, L. Unified Focal loss: Generalising Dice and cross entropy-based losses to handle class imbalanced medical image segmentation. Comput. Med. Imaging Graph. 2022, 95, 102026. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar] [CrossRef]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. Int. J. Comput. Vis. 2020, 128, 336–359. [Google Scholar] [CrossRef]
- Chattopadhay, A.; Sarkar, A.; Howlader, P.; Balasubramanian, V.N. Grad-CAM++: Generalized gradient-based visual explanations for deep convolutional networks. In Proceedings of the 2018 IEEE Winter Conference on Applications of Computer Vision, WACV 2018, Lake Tahoe, NV, USA, 12–15 March 2018; pp. 839–847. [Google Scholar] [CrossRef]
- Draelos, R.L.; Carin, L. Use HiResCAM instead of Grad-CAM for faithful explanations of convolutional neural networks. arXiv 2020, arXiv:2011.08891. [Google Scholar]
- Fu, R.; Hu, Q.; Dong, X.; Guo, Y.; Gao, Y.; Li, B. Axiom-based Grad-CAM: Towards Accurate Visualization and Explanation of CNNs. arXiv 2020, arXiv:2008.02312. [Google Scholar]
- Van der Walt, S.; Schönberger, J.L.; Nunez-Iglesias, J.; Boulogne, F.; Warner, J.D.; Yager, N.; Gouillart, E.; Yu, T. scikit-image: Image processing in Python. PeerJ 2014, 2, e453. [Google Scholar] [CrossRef] [PubMed]
- Bradski, G. The OpenCV Library. Dr. Dobb’S J. Softw. Tools 2000, 11, 120–123. [Google Scholar]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; et al. PyTorch: An Imperative Style, High-Performance Deep Learning Library. In Advances in Neural Information Processing Systems 32; Curran Associates Inc.: Red Hook, NY, USA, 2019; pp. 8024–8035. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Cardoso, M.J.; Li, W.; Brown, R.; Ma, N.; Kerfoot, E.; Wang, Y.; Murrey, B.; Myronenko, A.; Zhao, C.; Yang, D.; et al. MONAI: An open-source framework for deep learning in healthcare. arXiv 2022, arXiv:cs.LG/2211.02701. [Google Scholar]
- Szeghalmy, S.; Fazekas, A. A Comparative Study of the Use of Stratified Cross-Validation and Distribution-Balanced Stratified Cross-Validation in Imbalanced Learning. Sensors 2023, 23, 2333. [Google Scholar] [CrossRef] [PubMed]
- Vakharia, V.; Shah, M.; Suthar, V.; Patel, V.K.; Solanki, A. Hybrid perovskites thin films morphology identification by adapting multiscale-SinGAN architecture, heat transfer search optimized feature selection and machine learning algorithms. Phys. Scr. 2023, 98, 025203. [Google Scholar] [CrossRef]
- Buslaev, A.; Parinov, A.; Khvedchenya, E.; Iglovikov, V.I.; Kalinin, A.A. Albumentations: Fast and flexible image augmentations. arXiv 2018, arXiv:1809.06839. [Google Scholar] [CrossRef]
- Motlagh, M.H.; Science, C. Automatic Segmentation and Classification of Red and White Blood Cells in Thin Blood; Concordia University: Montreal, QC, USA, 2015. [Google Scholar]






| Input Image | |
|---|---|
| Segmented | Train the model with the previously Segmented Images |
| NoSegmented (Ablation) | Train the model with the original images (No Segmented Images) (Traditional manner) |
| MLP Classifier | |
| Linear | Modify the MLP classifier from 2048 to 3 neurons |
| Medium | Modify the MLP classifier from 2048 to 1024-512-3 neurons |
| Activation Function | |
| Mish | Change all the activation functions of the model to Mish, including MLP classifier |
| ReLU | Change all the activation functions of the model to ReLU, including MLP classifier |
| Dataset | Acc (%) | Pre (%) | Rec (%) | Spec (%) | DSC (%) | IoU |
|---|---|---|---|---|---|---|
| Leukemia Dataset | 98.50 | 88.32 | 95.03 | 98.59 | 91.16 | 0.840 |
| CellaVision | 99.32 | 97.08 | 97.88 | 99.57 | 97.40 | 0.951 |
| JTSC | 99.03 | 96.38 | 96.09 | 99.50 | 96.10 | 0.926 |
| SMC_IDB | 99.62 | 95.57 | 96.30 | 99.81 | 95.78 | 0.920 |
| Raabin_WBC | 98.99 | 97.38 | 94.71 | 99.65 | 94.83 | 0.923 |
| ALL_IDB2 | 98.51 | 93.45 | 97.14 | 98.60 | 95.14 | 0.910 |
| AVERAGE | 99.00 | 94.77 | 96.19 | 99.28 | 95.69 | 0.917 |
| Leukemia Dataset | ||||||
|---|---|---|---|---|---|---|
| Method | Acc (%) | Pre (%) | Rec (%) | Spec (%) | DSC (%) | IoU |
| Proposed HM | 97.96 | 82.43 | 97.63 | 97.70 | 89.01 | 0.806 |
| Proposed DEEP | 98.30 | 85.82 | 95.81 | 98.30 | 90.02 | 0.823 |
| Proposed Ensemble | 98.50 | 88.32 | 95.03 | 98.59 | 91.16 | 0.840 |
| Cellavision | |||||||
|---|---|---|---|---|---|---|---|
| Method | Acc (%) | Pre (%) | Rec (%) | Spec (%) | DSC (%) | IoU | # Test Images |
| Vogado et al. [10] | 98.77 | 97.88 | 99.75 | 89.39 | 93.22 | 0.873 | 100 |
| Makem & Tiedeu [11] | 99.37 | 97.37 | 96.97 | - | 97.06 | 0.945 | 100 |
| CPNNHSV [18] | 99.2 | 94.86 | 97.31 | 99.41 | 96.31 | 0.929 | 100 |
| Makem et al. [14] | 99.43 | 97.31 | 97.60 | 99.61 | 97.35 | 0.950 | 100 |
| LDS-NET [17] | - | 98.48 | 95.91 | - | 97.18 | 0.945 | 20 |
| LDAS-NET [17] | - | 99.09 | 97.11 | - | 98.09 | 0.963 | 20 |
| Proposed Ensemble | 99.32 | 97.08 | 97.88 | 99.57 | 97.40 | 0.951 | 100 |
| JTSC | |||||||
| Method | Acc (%) | Pre (%) | Rec (%) | Spec (%) | DSC (%) | IoU | # Test Images |
| Vogado et al. [10] | 97.13 | 93.55 | 98.99 | 83.18 | 87.68 | 0.781 | 300 |
| Makem & Tiedeu [11] | 97.29 | 91.01 | 93.12 | - | 90.79 | 0.843 | 300 |
| VarRGB [18] | 98.38 | 91.10 | 96.29 | 98.68 | 93.88 | 0.885 | 300 |
| Makem et al. [14] | 97.79 | 93.64 | 97.60 | 98.43 | 93.17 | 0.884 | 300 |
| Mayala & Haugsøen [15] | - | 94.89 | 95.30 | 99.31 | 94.81 | 0.903 | 300 |
| LDS-NET [17] | - | 98.85 | 92.39 | - | 95.56 | 0.917 | 60 |
| LDAS-NET [17] | - | 94.42 | 98.36 | - | 96.35 | 0.931 | 60 |
| Proposed Ensemble | 99.03 | 96.38 | 96.09 | 99.50 | 96.10 | 0.926 | 300 |
| SMC_IDB (BloodSeg) | |||||||
| Method | Acc (%) | Pre (%) | Rec (%) | Spec (%) | DSC (%) | IoU | # Test Images |
| Vogado et al. [10] | 99.15 | 80.51 | 94.51 | 99.30 | 86.46 | 0.761 | 367 |
| Makem & Tiedeu. [11] | 99.63 | 92.99 | 97.06 | - | 94.75 | 0.902 | 367 |
| Makem et al. [14] | 97.67 | 91.27 | 96.93 | 97.82 | 93.48 | 0.883 | 367 |
| Proposed Ensemble | 99.62 | 95.57 | 96.30 | 99.81 | 95.78 | 0.920 | 367 |
| Raabin | |||||||
| Method | Acc (%) | Pre (%) | Rec (%) | Spec (%) | DSC (%) | IoU | # Test Images |
| U-Net ++ [19] | - | 95.98 | 98.73 | - | 97.19 | 0.945 | 250 |
| Attention U-Net [20] | - | 94.78 | 98.50 | - | 96.33 | 0.929 | 250 |
| Mask R-CNN [21] | - | 8.59 | 96.80 | - | 91.98 | 0.852 | 250 |
| Mousavi et al. [12]. | - | 93.62 | 98.27 | - | 95.42 | 0.912 | 250 |
| Tavakoli et al. [13]. | - | 99.72 | 95.26 | - | 96.75 | 0.936 | 250 |
| Proposed Ensemble | 98.99 | 97.38 | 94.71 | 99.65 | 94.83 | 0.923 | 1145 |
| ALL_IDB2 | |||||||
| Method | Acc (%) | Pre (%) | Rec (%) | Spec (%) | DSC (%) | IoU | # Test Images |
| Vogado et al. [10] | 98.59 | 91.24 | 98.09 | 98.62 | 94.17 | 0.890 | 300 |
| CPNNHSV.[18] | 98.32 | 91.59 | 96.11 | 98.66 | 93.42 | 0.877 | 300 |
| Proposed Ensemble | 98.51 | 93.45 | 97.14 | 98.60 | 95.14 | 0.910 | 300 |
| Model | Acc (%) |
|---|---|
| Segmented Mish Medium | 99.99 |
| NoSegmented ReLU Medium | 99.97 |
| Segmented Mish Linear | 99.97 |
| No Segmented ReLU Linear | 99.97 |
| Method | Validation Accuracy (%) |
|---|---|
| Random_Forest Set_Full. [16] | 97.08 |
| LeNet. [16] | 98.36 |
| AlexNet. [16] | 99.98 |
| Proposed Method | 99.99 |
| Test Dataset | ||||
|---|---|---|---|---|
| Architecture | Acc (%) | Pre (%) | Rec (%) | F1 (%) |
| Segmented Mish Medium | 100 | 100 | 100 | 100 |
| Segmented ReLU Medium | 98.50 | 98.60 | 98.50 | 98.50 |
| No Segmented ReLU Linear | 89.40 | 89.70 | 89.40 | 89.40 |
| No Segmented Mish Medium | 80.30 | 87.60 | 80.30 | 80.30 |
| Dist L1–L2 | Dist L2–L3 | Dist L1–L3 | Dist Total | SD Cluster L1 | SD Cluster L2 | SD Cluster L3 | SD Total | Ratio Dist/SD | |
|---|---|---|---|---|---|---|---|---|---|
| Segmented Mish Medium | 10.12 | 8.22 | 7.31 | 25.65 | 2.05 | 0.68 | 2.06 | 2.99 | 25.78 |
| Segmented ReLU Medium | 6.60 | 5.17 | 3.65 | 15.41 | 1.09 | 1.43 | 0.66 | 1.92 | 24.12 |
| NoSegmented ReLU Linear | 3.94 | 3.34 | 2.99 | 10.27 | 1.32 | 1.60 | 1.33 | 2.46 | 12.52 |
| NoSegmented Mish Linear | 3.15 | 2.08 | 1.88 | 7.11 | 0.94 | 1.49 | 0.50 | 1.83 | 11.63 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Diaz Resendiz, J.L.; Ponomaryov, V.; Reyes Reyes, R.; Sadovnychiy, S. Explainable CAD System for Classification of Acute Lymphoblastic Leukemia Based on a Robust White Blood Cell Segmentation. Cancers 2023, 15, 3376. https://doi.org/10.3390/cancers15133376
Diaz Resendiz JL, Ponomaryov V, Reyes Reyes R, Sadovnychiy S. Explainable CAD System for Classification of Acute Lymphoblastic Leukemia Based on a Robust White Blood Cell Segmentation. Cancers. 2023; 15(13):3376. https://doi.org/10.3390/cancers15133376
Chicago/Turabian StyleDiaz Resendiz, Jose Luis, Volodymyr Ponomaryov, Rogelio Reyes Reyes, and Sergiy Sadovnychiy. 2023. "Explainable CAD System for Classification of Acute Lymphoblastic Leukemia Based on a Robust White Blood Cell Segmentation" Cancers 15, no. 13: 3376. https://doi.org/10.3390/cancers15133376
APA StyleDiaz Resendiz, J. L., Ponomaryov, V., Reyes Reyes, R., & Sadovnychiy, S. (2023). Explainable CAD System for Classification of Acute Lymphoblastic Leukemia Based on a Robust White Blood Cell Segmentation. Cancers, 15(13), 3376. https://doi.org/10.3390/cancers15133376






