Deep Learning for Skin Melanocytic Tumors in Whole-Slide Images: A Systematic Review
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
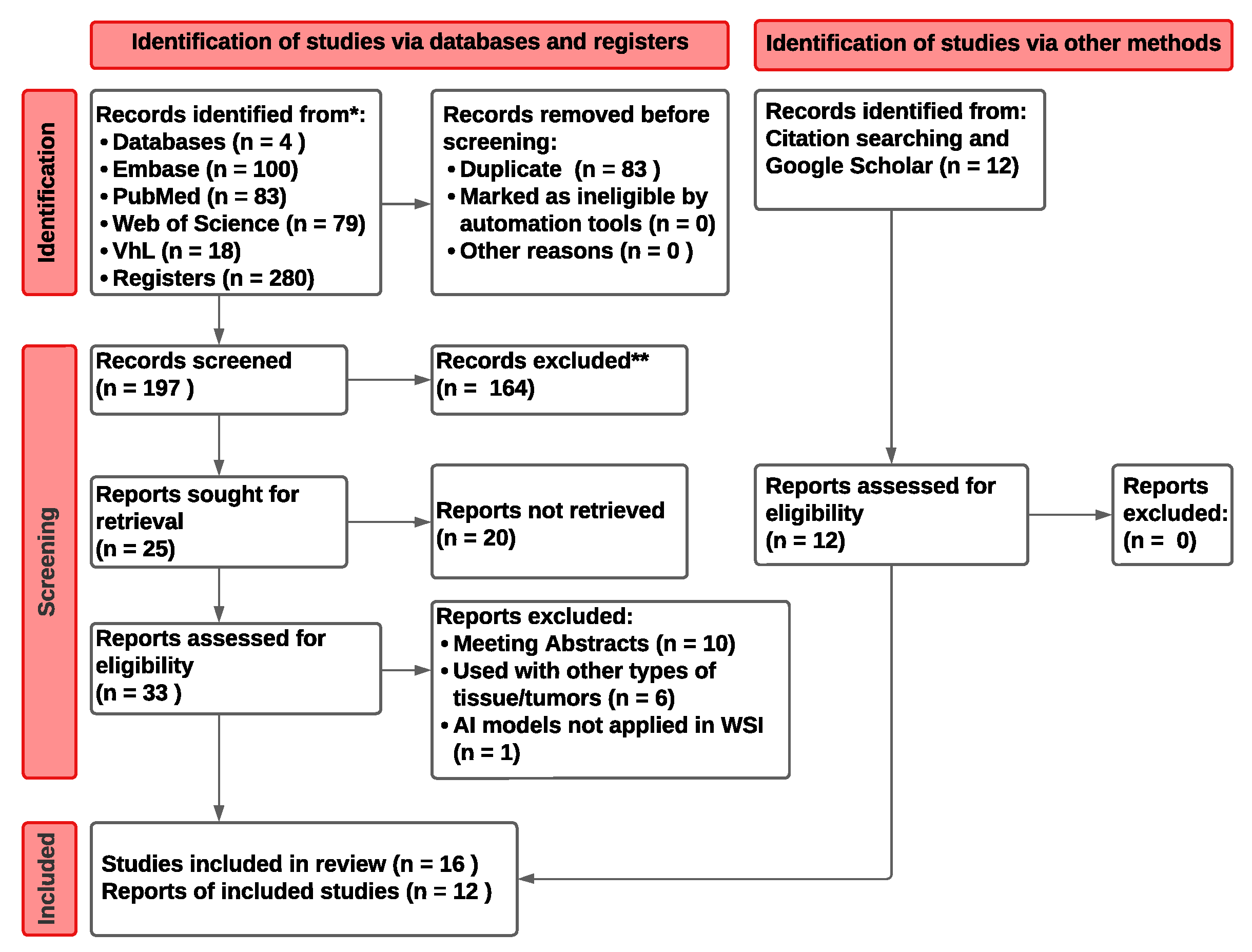
2.1. Literature Search Strategy
2.2. Study Eligibility and Selection
- i.
- No DL-based methods were used;
- ii.
- The writing language was different than English;
- iii.
- The analyzed tissues containing melanoma were other than skin (e.g., lymph node metastasis and uveal melanoma);
- iv.
- The used data sets were not of human origin.
2.3. Study Analysis and Performance Metrics
3. Results
- DL models vs pathologists (n = 10), where the algorithm is compared with a group of pathologists apart from those who were in charge of GT;
- Diagnostic prediction (n = 7), where the algorithm demonstrates its performance differentiating different groups of melanocytic lesions (e.g., melanoma and nevus);
- Prognosis (n = 5), where the algorithm recognizes important characteristics to determine the patient prognosis, i.e., lymph node metastasis and disease-specific survival (DSS), among others;
- Histological features and Regions Of Interest (ROIs) (n = 6), where the algorithm identifies key histopathological ROIs for further diagnosis (e.g., mitosis, tumor region, and epidermis).
| Study | Year | Studied Structures | Mag. | # WSIs | Patch Size | Pre-Processing | DL Method | GPU Used | # Sources | Metadata | xAI | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Comparison vs. pathologists | Ba et al. [42] | 2021 | Tumor | 40× | 781 | 256 × 256 | Image quality review | CNN and random forest | n/a | 2 | no | yes |
| Bao et al. [36] | 2022 | Tumor | 40× | 981 | 224 × 224 | Random patch selection, structure-preserving color normalization | ResNet-152 | NVIDIA GTX 2080Ti | 3 | no | no | |
| Brinker et al. [43] | 2022 | Tumor | n/a | 100 | n/a | n/a | ResNeXt50 | n/a | n/a | no | yes | |
| Hekler et al. [14,15] | 2019 | Tumor | 10× | 695 | n/a | n/a | ResNet50 | n/a | 1 | no | no | |
| Phillips et al. [27] | 2019 | Tumor, dermis, and epidermis | 40× | 50 | 512 × 512 | Subtraction | Modified FCN | NVIDIA GTX 1080 Ti | 10 † | no | yes | |
| Sturm et al. [16] | 2022 | Mitosis | 20× | 102 | n/a | n/a | n/a | n/a | 1 | yes | no | |
| Wang et al. [37] | 2020 | Tumor | 20× | 155 | 256 × 256 | Random cropping to 224 × 224, data enhancement, and augmentation | VGG16 | n/a | 2 | no | yes | |
| Xie et al. [28] | 2021 | Tumor, dermis, and epidermis | 20× | 701 | 224 × 224 | Discard blank patches (Otsu) | ResNet50 | n/a | 3 † | yes | no | |
| Xie et al. [17] | 2021 | Tumor | n/a | 841 | 256 × 256 | Discard blank patches (Otsu) | ResNet50 | NVIDIA TITAN RTX | 1 | no | yes | |
| Diagnosis | Del Amor et al. [19] | 2021 | Tumor | 10× | 51 | 512 × 512 | Discard blank patches (Otsu) | VGG16 with attention | NVIDIA DGX A100 | 1 | no | yes |
| Del Amor et al. [18] | 2022 | Tumor | 5×, 10×, 20× | 43 | 512 × 512 | Discard blank patches and with less than 20% of tissue (Otsu) | ResNet18 with late fusion of multiresolution feature maps | NVIDIA GP102 TITAN Xp | 1 | no | yes | |
| Hart el al. [34] | 2019 | Tumor | 40× | 300 | 299 × 299 | n/a | InceptionV3 | 4 NVIDIA GeForce GTX 1080 | n/a | no | yes | |
| Höhn et al. [38] | 2021 | Tumor | n/a | 431 | 512 × 512 | Remove patches with more than 50% of background, random selection of 100 tiles per slide | ResNeXt50 with fusion model to combine patient data and image features | NVIDIA GeForce GTX 745 | 2 | yes | yes | |
| Li et al. [29] | 2021 | Tumor, dermis, and epidermis | 20× | 701 | 224 × 224 | Discard blank patches (Otsu) | ResNet50 | n/a | 2 † | yes | yes | |
| Van Zon et al. [20] | 2020 | Tumor | 40× | 563 | 256 × 256 | Data augmentation | U-Net | NVIDIA 2080 | 1 | no | no | |
| Xie et al. [21] | 2021 | Tumor | 40× | 312 | 500 × 500 | Filter out background tiles | Transfer learning vs fully trained: InceptionV3, ResNet50, MobileNet | n/a | 1 | no | no | |
| Prognosis | Brinker et al. [13] | 2021 | Tumor | n/a | 415 | 256 × 256 | n/a | ResNeXt50 | n/a | 3 | yes | no |
| Kim et al. [30] | 2022 | Tumor, inflammatory cells, and other | 20× | 305 | 299 × 299 | n/a | Inception v3 with fivefold cross-validation | n/a | 2 † | yes | no | |
| Kulkarni et al. [40] | 2020 | Tumor, inflammatory cells, and other | 40× | n/a | 500 × 500 | Downsample to 100 × 100, nuclear segmentation with watershed cell detection | n/a | n/a | 2 | yes | no | |
| Moore et al. [41] | 2021 | Tumor, inflammatory cells, and other | 40×, 20× | n/a | 100 × 100 | n/a | QuIP TIL CNN [44] | NVIDIA GP102GL [Quadro P6000] | 2 | yes | no | |
| Zormpas-Petridis et al. [31] | 2019 | Tumor, inflammatory cells, and other | 20×, 5×, 1.25× | 105 | 2000 × 2000 (20× WSIs) | n/a | Spatially constrained CNN with spatial regression, neighboring ensemble with softmax | NVIDIA Tesla P100-PCIE-16GB | 1 † | yes | no | |
| ROI/histological features | Alheejawi et al. [22] | 2021 | Tumor, inflammatory cells, and epidermis | 40× | 4 | 960 × 960 | Divide patches into 64 × 64 blocks | ResNet50 | NVIDIA GeForce GTX 745 | 1 | no | no |
| De Logu et al. [39] | 2020 | Tumor and healthy tissues | 20× | 100 | 299 × 299 | Data augmentation, discard patches with more than 50% background | Inception-ResNet-v2 | n/a | 3 | no | yes | |
| Kucharski et al. [23] | 2020 | Tumor | 10× | 70 | 128 × 128 | Data augmentation, overlapping only for minority class to balance data set | Autoencoders | n/a | 1 | no | yes | |
| Liu et al. [24] | 2021 | Tumor | 10× | 227 ROIs ‡ | 1000 × 1000 | Downscale magnification to 5× | Mask R-CNN | 4 NVIDIA GeForce GTX 1080 | 1 | no | no | |
| Nofallah et al. [25] | 2021 | Mitosis | 40× | 22 | 101 × 101 | Data augmentation | ESPNet, DenseNet, ResNet, and ShuffleNet | NVIDIA GeForce GTX 1080 | 1 | no | no | |
| Zhang et al. [26] | 2021 | Tumor | n/a | 30 | 1024 × 1024 | Data augmentation, color analysis for tissue-contained patch selection, normalization of patches to a uniform size, resize patches to 512 × 512 | CNN, feature fusion | NVIDIA RTX 2080-12G | 1 | no | no |
3.1. Deep Learning Models vs. Pathologists
3.2. Diagnostic Prediction
3.3. Prognosis
3.4. Histological Features and ROIs
4. Discussion
4.1. Assistance Utility in Clinical Practice
4.2. The Rise of DL for WSI Analysis: Requirements and Promises
4.3. Making the Bridge between Pathologists and AI Developers
4.4. Limitations
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AI | Artificial Intelligence |
| AUC | Area Under the ROC Curve |
| CBIR | Content-Based Image Retrieval |
| CI | Confidence Interval |
| CLAIM | Checklist for Artificial Intelligence in Medical Imaging |
| CNN | Convolutional Neural Network |
| DL | Deep Learning |
| DNN | Deep Neural Network |
| DP | Digital Pathology |
| DSS | Disease-Specific Survival |
| FCN | Fully Convolutional Network |
| GDC | Genomic Data Commons |
| GT | Ground Truth |
| H&E | Hematoxylin and Eosin |
| HR | Hazard Ratio |
| MELTUMP | Melanocytic Tumors of Uncertain Malignant Potential |
| ML | Machine Learning |
| NCI | National Cancer Institute |
| PHH3 | Phosphohistone-H3 |
| RNN | Recurrent Neural Network |
| ROI | Region of Interest |
| SN | Sentinel Node |
| SR | Systematic Review |
| TCGA | The Cancer Genome Atlas |
| WSI | Whole-Slide Image |
| xAI | Explainable Artificial Intelligence |
References
- Elmore, J.G.; Barnhill, R.L.; Elder, D.E.; Longton, G.M.; Pepe, M.S.; Reisch, L.M.; Carney, P.A.; Titus, L.J.; Nelson, H.D.; Onega, T.; et al. Pathologists’ diagnosis of invasive melanoma and melanocytic proliferations: Observer accuracy and reproducibility study. BMJ 2017, 357. [Google Scholar] [CrossRef] [PubMed]
- Farmer, E.R.; Gonin, R.; Hanna, M.P. Discordance in the histopathologic diagnosis of melanoma and melanocytic nevi between expert pathologists. Hum. Pathol. 1996, 27, 528–531. [Google Scholar] [CrossRef] [PubMed]
- Shoo, B.A.; Sagebiel, R.W.; Kashani-Sabet, M. Discordance in the histopathologic diagnosis of melanoma at a melanoma referral center. J. Am. Acad. Dermatol. 2010, 62, 751–756. [Google Scholar] [CrossRef] [PubMed]
- Yeh, I. Recent advances in molecular genetics of melanoma progression: Implications for diagnosis and treatment. F1000Research 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Pantanowitz, L. Digital images and the future of digital pathology. J. Pathol. Inform. 2010, 1. [Google Scholar] [CrossRef]
- Mukhopadhyay, S.; Feldman, M.D.; Abels, E.; Ashfaq, R.; Beltaifa, S.; Cacciabeve, N.G.; Cathro, H.P.; Cheng, L.; Cooper, K.; Dickey, G.E.; et al. Whole slide imaging versus microscopy for primary diagnosis in surgical pathology: A multicenter blinded randomized noninferiority study of 1992 cases (pivotal study). Am. J. Surg. Pathol. 2018, 42, 39. [Google Scholar] [CrossRef]
- Tizhoosh, H.R.; Pantanowitz, L. Artificial intelligence and digital pathology: Challenges and opportunities. J. Pathol. Inform. 2018, 9, 38. [Google Scholar] [CrossRef]
- Harrison, J.H.; Gilbertson, J.R.; Hanna, M.G.; Olson, N.H.; Seheult, J.N.; Sorace, J.M.; Stram, M.N. Introduction to Artificial Intelligence and Machine Learning for Pathology. Arch. Pathol. Lab. Med. 2021, 145, 1228–1254. [Google Scholar] [CrossRef]
- Jiang, Y.; Yang, M.; Wang, S.; Li, X.; Sun, Y. Emerging role of deep learning-based artificial intelligence in tumor pathology. Cancer Commun. 2020, 40, 154–166. [Google Scholar] [CrossRef]
- Echle, A.; Rindtorff, N.T.; Brinker, T.J.; Luedde, T.; Pearson, A.T.; Kather, J.N. Deep learning in cancer pathology: A new generation of clinical biomarkers. Br. J. Cancer 2021, 124, 686–696. [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 statement: An updated guideline for reporting systematic reviews. Syst. Rev. 2021, 10, 1–11. [Google Scholar]
- Mongan, J.; Moy, L.; Kahn Jr, C.E. Checklist for artificial intelligence in medical imaging (CLAIM): A guide for authors and reviewers. Radiology. Artif. Intell. 2020, 2. [Google Scholar] [CrossRef] [PubMed]
- Brinker, T.J.; Kiehl, L.; Schmitt, M.; Jutzi, T.B.; Krieghoff-Henning, E.I.; Krahl, D.; Kutzner, H.; Gholam, P.; Haferkamp, S.; Klode, J.; et al. Deep learning approach to predict sentinel lymph node status directly from routine histology of primary melanoma tumours. Eur. J. Cancer 2021, 154, 227–234. [Google Scholar] [CrossRef] [PubMed]
- Hekler, A.; Utikal, J.S.; Enk, A.H.; Solass, W.; Schmitt, M.; Klode, J.; Schadendorf, D.; Sondermann, W.; Franklin, C.; Bestvater, F.; et al. Deep learning outperformed 11 pathologists in the classification of histopathological melanoma images. Eur. J. Cancer 2019, 118, 91–96. [Google Scholar] [CrossRef]
- Hekler, A.; Utikal, J.S.; Enk, A.H.; Berking, C.; Klode, J.; Schadendorf, D.; Jansen, P.; Franklin, C.; Holland-Letz, T.; Krahl, D.; et al. Pathologist-level classification of histopathological melanoma images with deep neural networks. Eur. J. Cancer 2019, 115, 79–83. [Google Scholar] [CrossRef]
- Sturm, B.; Creytens, D.; Smits, J.; Ooms, A.H.; Eijken, E.; Kurpershoek, E.; Küsters-Vandevelde, H.V.; Wauters, C.; Blokx, W.A.; van der Laak, J.A. Computer-Aided Assessment of Melanocytic Lesions by Means of a Mitosis Algorithm. Diagnostics 2022, 12, 436. [Google Scholar] [CrossRef]
- Xie, P.; Zuo, K.; Liu, J.; Chen, M.; Zhao, S.; Kang, W.; Li, F. Interpretable diagnosis for whole-slide melanoma histology images using convolutional neural network. J. Healthc. Eng. 2021, 2021. [Google Scholar] [CrossRef]
- Del Amor, R.; Curieses, F.J.; Launet, L.; Colomer, A.; Moscardó, A.; Mosquera-Zamudio, A.; Monteagudo, C.; Naranjo, V. Multi-Resolution Framework For Spitzoid Neoplasm Classification Using Histological Data. In Proceedings of the 2022 IEEE 14th Image, Video, and Multidimensional Signal Processing Workshop (IVMSP), Nafplio, Greece, 26–29 June 2022; pp. 1–5. [Google Scholar]
- Del Amor, R.; Launet, L.; Colomer, A.; Moscardó, A.; Mosquera-Zamudio, A.; Monteagudo, C.; Naranjo, V. An attention-based weakly supervised framework for spitzoid melanocytic lesion diagnosis in whole slide images. Artif. Intell. Med. 2021, 121, 102197. [Google Scholar] [CrossRef]
- Van Zon, M.; Stathonikos, N.; Blokx, W.A.; Komina, S.; Maas, S.L.; Pluim, J.P.; Van Diest, P.J.; Veta, M. Segmentation and classification of melanoma and nevus in whole slide images. In Proceedings of the 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI), Iowa City, IA, USA, 3–7 April 2020; pp. 263–266. [Google Scholar]
- Xie, P.; Li, T.; Liu, J.; Li, F.; Zhou, J.; Zuo, K. Analyze Skin Histopathology Images Using Multiple Deep Learning Methods. In Proceedings of the 2021 IEEE 3rd International Conference on Frontiers Technology of Information and Computer (ICFTIC), Virtual, 12–14 November 2021; pp. 374–377. [Google Scholar]
- Alheejawi, S.; Berendt, R.; Jha, N.; Maity, S.P.; Mandal, M. Detection of malignant melanoma in H&E-stained images using deep learning techniques. Tissue Cell 2021, 73, 101659. [Google Scholar]
- Kucharski, D.; Kleczek, P.; Jaworek-Korjakowska, J.; Dyduch, G.; Gorgon, M. Semi-supervised nests of melanocytes segmentation method using convolutional autoencoders. Sensors 2020, 20, 1546. [Google Scholar] [CrossRef]
- Liu, K.; Mokhtari, M.; Li, B.; Nofallah, S.; May, C.; Chang, O.; Knezevich, S.; Elmore, J.; Shapiro, L. Learning Melanocytic Proliferation Segmentation in Histopathology Images from Imperfect Annotations. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Nashville, TN, USA, 20–25 June 2021; pp. 3766–3775. [Google Scholar]
- Nofallah, S.; Mehta, S.; Mercan, E.; Knezevich, S.; May, C.J.; Weaver, D.; Witten, D.; Elmore, J.G.; Shapiro, L. Machine learning techniques for mitoses classification. Comput. Med Imaging Graph. 2021, 87, 101832. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Han, H.; Du, S.; Zhu, L.; Yang, J.; Wang, X.; Wang, L.; Xu, M. MPMR: Multi-Scale Feature and Probability Map for Melanoma Recognition. Front. Med. 2021, 8. [Google Scholar] [CrossRef] [PubMed]
- Phillips, A.; Teo, I.; Lang, J. Segmentation of prognostic tissue structures in cutaneous melanoma using whole slide images. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops, (CVPRW), Long Beach, CA, USA, 16–17 June 2019; pp. 2738–2747. [Google Scholar]
- Xie, P.; Li, T.; Li, F.; Liu, J.; Zhou, J.; Zuo, K. Automated Diagnosis of Melanoma Histopathological Images Based on Deep Learning Using Trust Counting Method. In Proceedings of the 2021 IEEE 3rd International Conference on Frontiers Technology of Information and Computer (ICFTIC), Virtual, 12–14 November 2021; pp. 26–29. [Google Scholar]
- Li, T.; Xie, P.; Liu, J.; Chen, M.; Zhao, S.; Kang, W.; Zuo, K.; Li, F. Automated Diagnosis and Localization of Melanoma from Skin Histopathology Slides Using Deep Learning: A Multicenter Study. J. Healthc. Eng. 2021, 2021. [Google Scholar] [CrossRef] [PubMed]
- Kim, R.H.; Nomikou, S.; Coudray, N.; Jour, G.; Dawood, Z.; Hong, R.; Esteva, E.; Sakellaropoulos, T.; Donnelly, D.; Moran, U.; et al. Deep learning and pathomics analyses reveal cell nuclei as important features for mutation prediction of BRAF-mutated melanomas. J. Investig. Dermatol. 2022, 142, 1650–1658. [Google Scholar] [CrossRef]
- Zormpas-Petridis, K.; Failmezger, H.; Raza, S.E.A.; Roxanis, I.; Jamin, Y.; Yuan, Y. Superpixel-based Conditional Random Fields (SuperCRF): Incorporating global and local context for enhanced deep learning in melanoma histopathology. Front. Oncol. 2019, 9, 1045. [Google Scholar] [CrossRef]
- Tomczak, K.; Czerwińska, P.; Wiznerowicz, M. Review The Cancer Genome Atlas (TCGA): An immeasurable source of knowledge. Contemp. Oncol. Onkol. 2015, 2015, 68–77. [Google Scholar] [CrossRef]
- Jensen, M.A.; Ferretti, V.; Grossman, R.L.; Staudt, L.M. The NCI Genomic Data Commons as an engine for precision medicine. Blood, J. Am. Soc. Hematol. 2017, 130, 453–459. [Google Scholar] [CrossRef]
- Hart, S.N.; Flotte, W.; Andrew, F.; Shah, K.K.; Buchan, Z.R.; Mounajjed, T.; Flotte, T.J. Classification of melanocytic lesions in selected and whole-slide images via convolutional neural networks. J. Pathol. Inform. 2019, 10, 5. [Google Scholar] [CrossRef]
- Xie, P.; Zuo, K.; Zhang, Y.; Li, F.; Yin, M.; Lu, K. Interpretable classification from skin cancer histology slides using deep learning: A retrospective multicenter study. arXiv 2019, arXiv:1904.06156. [Google Scholar]
- Bao, Y.; Zhang, J.; Zhao, X.; Zhou, H.; Chen, Y.; Jian, J.; Shi, T.; Gao, X. Deep learning-based fully automated diagnosis of melanocytic lesions by using whole slide images. J. Dermatol. Treat. 2022, 1–7. [Google Scholar] [CrossRef]
- Wang, L.; Ding, L.; Liu, Z.; Sun, L.; Chen, L.; Jia, R.; Dai, X.; Cao, J.; Ye, J. Automated identification of malignancy in whole-slide pathological images: Identification of eyelid malignant melanoma in gigapixel pathological slides using deep learning. Br. J. Ophthalmol. 2020, 104, 318–323. [Google Scholar] [CrossRef] [PubMed]
- Höhn, J.; Krieghoff-Henning, E.; Jutzi, T.B.; von Kalle, C.; Utikal, J.S.; Meier, F.; Gellrich, F.F.; Hobelsberger, S.; Hauschild, A.; Schlager, J.G.; et al. Combining CNN-based histologic whole slide image analysis and patient data to improve skin cancer classification. Eur. J. Cancer 2021, 149, 94–101. [Google Scholar] [CrossRef] [PubMed]
- De Logu, F.; Ugolini, F.; Maio, V.; Simi, S.; Cossu, A.; Massi, D.; Italian Association for Cancer Research (AIRC) Study Group; Nassini, R.; Laurino, M. Recognition of cutaneous melanoma on digitized histopathological slides via artificial intelligence algorithm. Front. Oncol. 2020, 10, 1559. [Google Scholar] [CrossRef] [PubMed]
- Kulkarni, P.M.; Robinson, E.J.; Sarin Pradhan, J.; Gartrell-Corrado, R.D.; Rohr, B.R.; Trager, M.H.; Geskin, L.J.; Kluger, H.M.; Wong, P.F.; Acs, B.; et al. Deep Learning Based on Standard H&E Images of Primary Melanoma Tumors Identifies Patients at Risk for Visceral Recurrence and DeathDeep Learning–based Prognostic Biomarker for Melanoma. Clin. Cancer Res. 2020, 26, 1126–1134. [Google Scholar] [PubMed]
- Moore, M.R.; Friesner, I.D.; Rizk, E.M.; Fullerton, B.T.; Mondal, M.; Trager, M.H.; Mendelson, K.; Chikeka, I.; Kurc, T.; Gupta, R.; et al. Automated digital TIL analysis (ADTA) adds prognostic value to standard assessment of depth and ulceration in primary melanoma. Sci. Rep. 2021, 11, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Ba, W.; Wang, R.; Yin, G.; Song, Z.; Zou, J.; Zhong, C.; Yang, J.; Yu, G.; Yang, H.; Zhang, L.; et al. Diagnostic assessment of deep learning for melanocytic lesions using whole-slide pathological images. Transl. Oncol. 2021, 14, 101161. [Google Scholar] [CrossRef]
- Brinker, T.J.; Schmitt, M.; Krieghoff-Henning, E.I.; Barnhill, R.; Beltraminelli, H.; Braun, S.A.; Carr, R.; Fernandez-Figueras, M.T.; Ferrara, G.; Fraitag, S.; et al. Diagnostic performance of artificial intelligence for histologic melanoma recognition compared to 18 international expert pathologists. J. Am. Acad. Dermatol. 2022, 86, 640–642. [Google Scholar] [CrossRef]
- Saltz, J.; Gupta, R.; Hou, L.; Kurc, T.; Singh, P.; Nguyen, V.; Samaras, D.; Shroyer, K.R.; Zhao, T.; Batiste, R.; et al. Spatial organization and molecular correlation of tumor-infiltrating lymphocytes using deep learning on pathology images. Cell Rep. 2018, 23, 181–193. [Google Scholar] [CrossRef]
- Araújo, T.; Aresta, G.; Castro, E.; Rouco, J.; Aguiar, P.; Eloy, C.; Polónia, A.; Campilho, A. Classification of breast cancer histology images using convolutional neural networks. PLoS ONE 2017, 12, e0177544. [Google Scholar] [CrossRef]
- Deniz, E.; Şengür, A.; Kadiroğlu, Z.; Guo, Y.; Bajaj, V.; Budak, Ü. Transfer learning based histopathologic image classification for breast cancer detection. Health Inf. Sci. Syst. 2018, 6, 1–7. [Google Scholar] [CrossRef]
- Tellez, D.; Balkenhol, M.; Otte-Höller, I.; van de Loo, R.; Vogels, R.; Bult, P.; Wauters, C.; Vreuls, W.; Mol, S.; Karssemeijer, N.; et al. Whole-slide mitosis detection in H&E breast histology using PHH3 as a reference to train distilled stain-invariant convolutional networks. IEEE Trans. Med Imaging 2018, 37, 2126–2136. [Google Scholar]
- Piepkorn, M.W.; Barnhill, R.L.; Elder, D.E.; Knezevich, S.R.; Carney, P.A.; Reisch, L.M.; Elmore, J.G. The MPATH-Dx reporting schema for melanocytic proliferations and melanoma. J. Am. Acad. Dermatol. 2014, 70, 131–141. [Google Scholar] [CrossRef] [PubMed]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar]
- Ilse, M.; Tomczak, J.; Welling, M. Attention-based deep multiple instance learning. In Proceedings of the International Conference on Machine Learning. PMLR, Stockholm, Sweden, 10–15 July 2018; pp. 2127–2136. [Google Scholar]
- Hu, J.; Shen, L.; Sun, G. Squeeze-and-excitation networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 7132–7141. [Google Scholar]
- Sirinukunwattana, K.; Raza, S.E.A.; Tsang, Y.W.; Snead, D.R.; Cree, I.A.; Rajpoot, N.M. Locality sensitive deep learning for detection and classification of nuclei in routine colon cancer histology images. IEEE Trans. Med Imaging 2016, 35, 1196–1206. [Google Scholar] [CrossRef] [PubMed]
- Lafferty, J.; McCallum, A.; Pereira, F.C. Conditional Random Fields: Probabilistic Models for Segmenting and Labeling Sequence Data. In Proceedings of the International Conference on Machine Learning 2001 (ICML 2001), Williamstown, MA, USA, 28 June–1 July 1 2001. [Google Scholar]
- Graham, S.; Vu, Q.D.; Raza, S.E.A.; Azam, A.; Tsang, Y.W.; Kwak, J.T.; Rajpoot, N. Hover-net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med Image Anal. 2019, 58, 101563. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, A.E.; Jones, T.R.; Lamprecht, M.R.; Clarke, C.; Kang, I.H.; Friman, O.; Guertin, D.A.; Chang, J.H.; Lindquist, R.A.; Moffat, J.; et al. CellProfiler: Image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 2006, 7, 1–11. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Berlin/Heidelberg, Germany, 2015; pp. 234–241. [Google Scholar]
- Mehta, S.; Rastegari, M.; Caspi, A.; Shapiro, L.; Hajishirzi, H. Espnet: Efficient spatial pyramid of dilated convolutions for semantic segmentation. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; pp. 552–568. [Google Scholar]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 4700–4708. [Google Scholar]
- Li, Z.; Koban, K.C.; Schenck, T.L.; Giunta, R.E.; Li, Q.; Sun, Y. Artificial Intelligence in Dermatology Image Analysis: Current Developments and Future Trends. J. Clin. Med. 2022, 11, 6826. [Google Scholar] [CrossRef] [PubMed]
- Goyal, M.; Knackstedt, T.; Yan, S.; Hassanpour, S. Artificial intelligence-based image classification methods for diagnosis of skin cancer: Challenges and opportunities. Comput. Biol. Med. 2020, 127, 104065. [Google Scholar] [CrossRef]
- Haggenmüller, S.; Maron, R.C.; Hekler, A.; Utikal, J.S.; Barata, C.; Barnhill, R.L.; Beltraminelli, H.; Berking, C.; Betz-Stablein, B.; Blum, A.; et al. Skin cancer classification via convolutional neural networks: Systematic review of studies involving human experts. Eur. J. Cancer 2021, 156, 202–216. [Google Scholar] [CrossRef]
- Zhang, S.; Wang, Y.; Zheng, Q.; Li, J.; Huang, J.; Long, X. Artificial intelligence in melanoma: A systematic review. J. Cosmet. Dermatol. 2022. [Google Scholar] [CrossRef]
- Popescu, D.; El-Khatib, M.; El-Khatib, H.; Ichim, L. New Trends in Melanoma Detection Using Neural Networks: A Systematic Review. Sensors 2022, 22, 496. [Google Scholar] [CrossRef]
- Cazzato, G.; Colagrande, A.; Cimmino, A.; Arezzo, F.; Loizzi, V.; Caporusso, C.; Marangio, M.; Foti, C.; Romita, P.; Lospalluti, L.; et al. Artificial intelligence in dermatopathology: New insights and perspectives. Dermatopathology 2021, 8, 418–425. [Google Scholar] [CrossRef] [PubMed]
- Homeyer, A.; Geißler, C.; Schwen, L.O.; Zakrzewski, F.; Evans, T.; Strohmenger, K.; Westphal, M.; Bülow, R.D.; Kargl, M.; Karjauv, A.; et al. Recommendations on compiling test datasets for evaluating artificial intelligence solutions in pathology. Mod. Pathol. 2022, 35, 1759–1769. [Google Scholar] [CrossRef] [PubMed]
- Wiesner, T.; Kutzner, H.; Cerroni, L.; Mihm Jr, M.C.; Busam, K.J.; Murali, R. Genomic aberrations in spitzoid melanocytic tumours and their implications for diagnosis, prognosis and therapy. Pathology 2016, 48, 113–131. [Google Scholar] [CrossRef] [PubMed]
- Romo-Bucheli, D.; Janowczyk, A.; Gilmore, H.; Romero, E.; Madabhushi, A. Automated tubule nuclei quantification and correlation with oncotype DX risk categories in ER+ breast cancer whole slide images. Sci. Rep. 2016, 6, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Ferrara, G.; Argenyi, Z.; Argenziano, G.; Cerio, R.; Cerroni, L.; Di Blasi, A.; Feudale, E.A.; Giorgio, C.M.; Massone, C.; Nappi, O.; et al. The influence of clinical information in the histopathologic diagnosis of melanocytic skin neoplasms. PLoS ONE 2009, 4, e5375. [Google Scholar] [CrossRef]
- Schmitt, M.; Maron, R.C.; Hekler, A.; Stenzinger, A.; Hauschild, A.; Weichenthal, M.; Tiemann, M.; Krahl, D.; Kutzner, H.; Utikal, J.S.; et al. Hidden variables in deep learning digital pathology and their potential to cause batch effects: Prediction model study. J. Med Internet Res. 2021, 23, e23436. [Google Scholar] [CrossRef] [PubMed]
- Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 1979, 9, 62–66. [Google Scholar] [CrossRef]
- Hauser, K.; Kurz, A.; Haggenmüller, S.; Maron, R.C.; von Kalle, C.; Utikal, J.S.; Meier, F.; Hobelsberger, S.; Gellrich, F.F.; Sergon, M.; et al. Explainable artificial intelligence in skin cancer recognition: A systematic review. Eur. J. Cancer 2022, 167, 54–69. [Google Scholar] [CrossRef]
- Tosun, A.B.; Pullara, F.; Becich, M.J.; Taylor, D.; Fine, J.L.; Chennubhotla, S.C. Explainable AI (xAI) for anatomic pathology. Adv. Anat. Pathol. 2020, 27, 241–250. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mosquera-Zamudio, A.; Launet, L.; Tabatabaei, Z.; Parra-Medina, R.; Colomer, A.; Oliver Moll, J.; Monteagudo, C.; Janssen, E.; Naranjo, V. Deep Learning for Skin Melanocytic Tumors in Whole-Slide Images: A Systematic Review. Cancers 2023, 15, 42. https://doi.org/10.3390/cancers15010042
Mosquera-Zamudio A, Launet L, Tabatabaei Z, Parra-Medina R, Colomer A, Oliver Moll J, Monteagudo C, Janssen E, Naranjo V. Deep Learning for Skin Melanocytic Tumors in Whole-Slide Images: A Systematic Review. Cancers. 2023; 15(1):42. https://doi.org/10.3390/cancers15010042
Chicago/Turabian StyleMosquera-Zamudio, Andrés, Laëtitia Launet, Zahra Tabatabaei, Rafael Parra-Medina, Adrián Colomer, Javier Oliver Moll, Carlos Monteagudo, Emiel Janssen, and Valery Naranjo. 2023. "Deep Learning for Skin Melanocytic Tumors in Whole-Slide Images: A Systematic Review" Cancers 15, no. 1: 42. https://doi.org/10.3390/cancers15010042
APA StyleMosquera-Zamudio, A., Launet, L., Tabatabaei, Z., Parra-Medina, R., Colomer, A., Oliver Moll, J., Monteagudo, C., Janssen, E., & Naranjo, V. (2023). Deep Learning for Skin Melanocytic Tumors in Whole-Slide Images: A Systematic Review. Cancers, 15(1), 42. https://doi.org/10.3390/cancers15010042





