Chemosensitivity of 3D Pancreatic Cancer Organoids Is Not Affected by Transformation to 2D Culture or Switch to Physiological Culture Medium
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Maintenance and Passaging of 3D Organoids
2.2. Transformation of 3D Organoids into 2D Cell Cultures
2.3. Maintenance and Passaging of 2D Transformed Cell Cultures
2.4. Switching from Conventional to Physiological Culture Medium
2.5. Chemosensitivity Assay
2.6. Quality Control
2.7. Dose–Response Curves
2.8. Doubling Time
2.9. Growth Rate 50
2.10. Proliferation Assay
2.11. RNA Isolation
2.12. RNA Sequencing
2.13. Statistics
3. Results
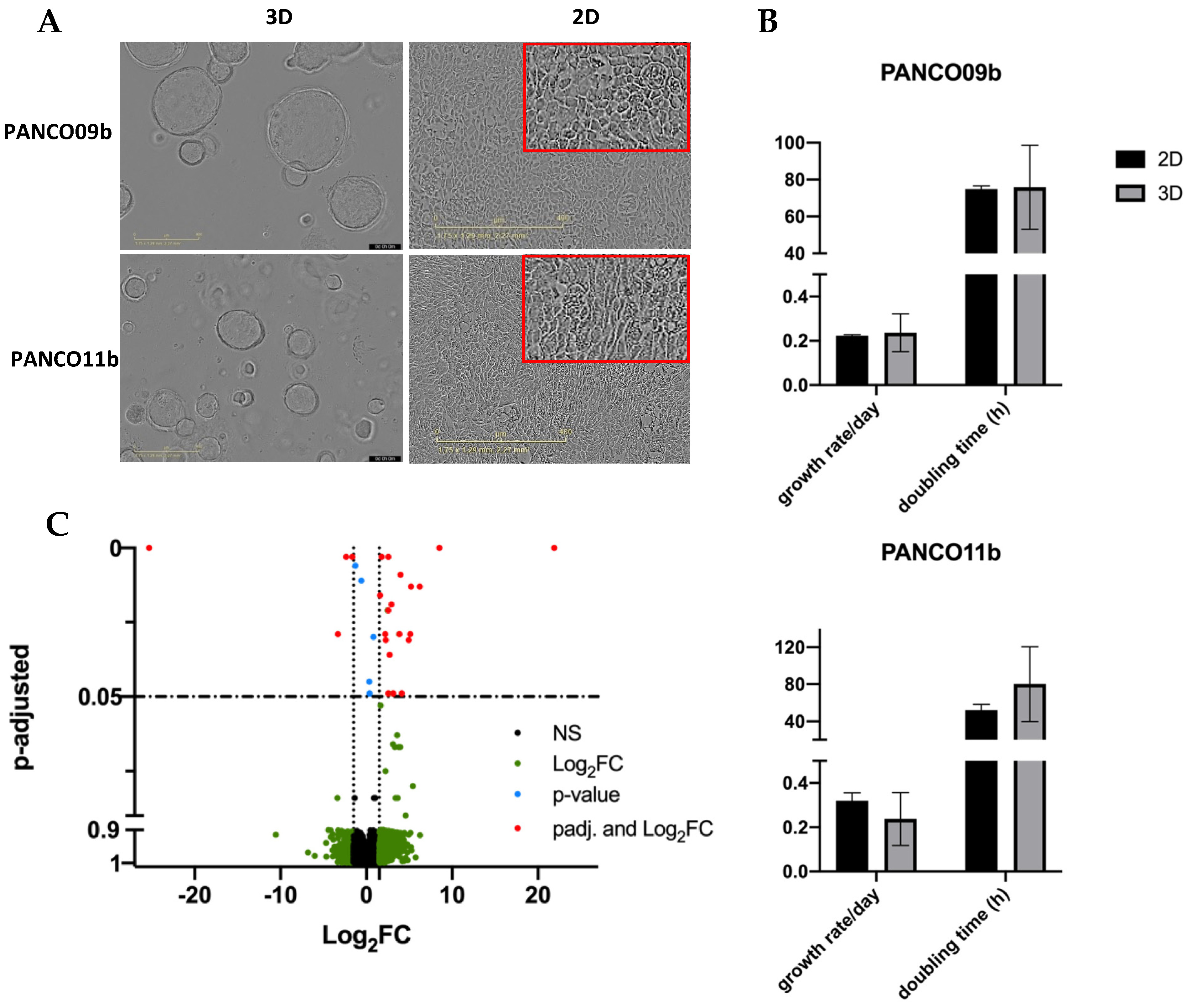
3.1. Establishing 2D Transformed Cell Cultures from 3D Organoids
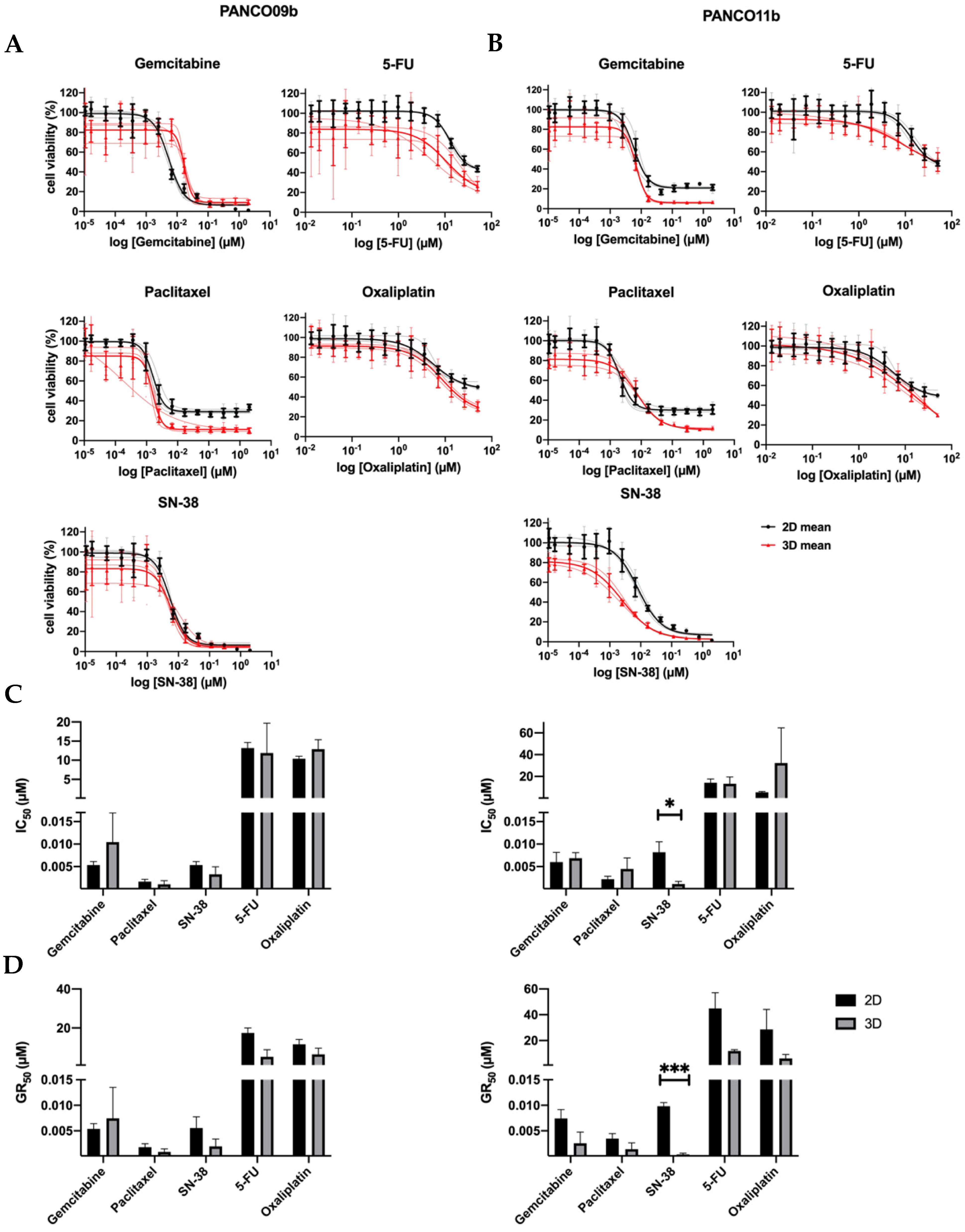
3.2. Similar Chemosensitivity of 3D Organoids and Corresponding 2D Transformed Cell Cultures
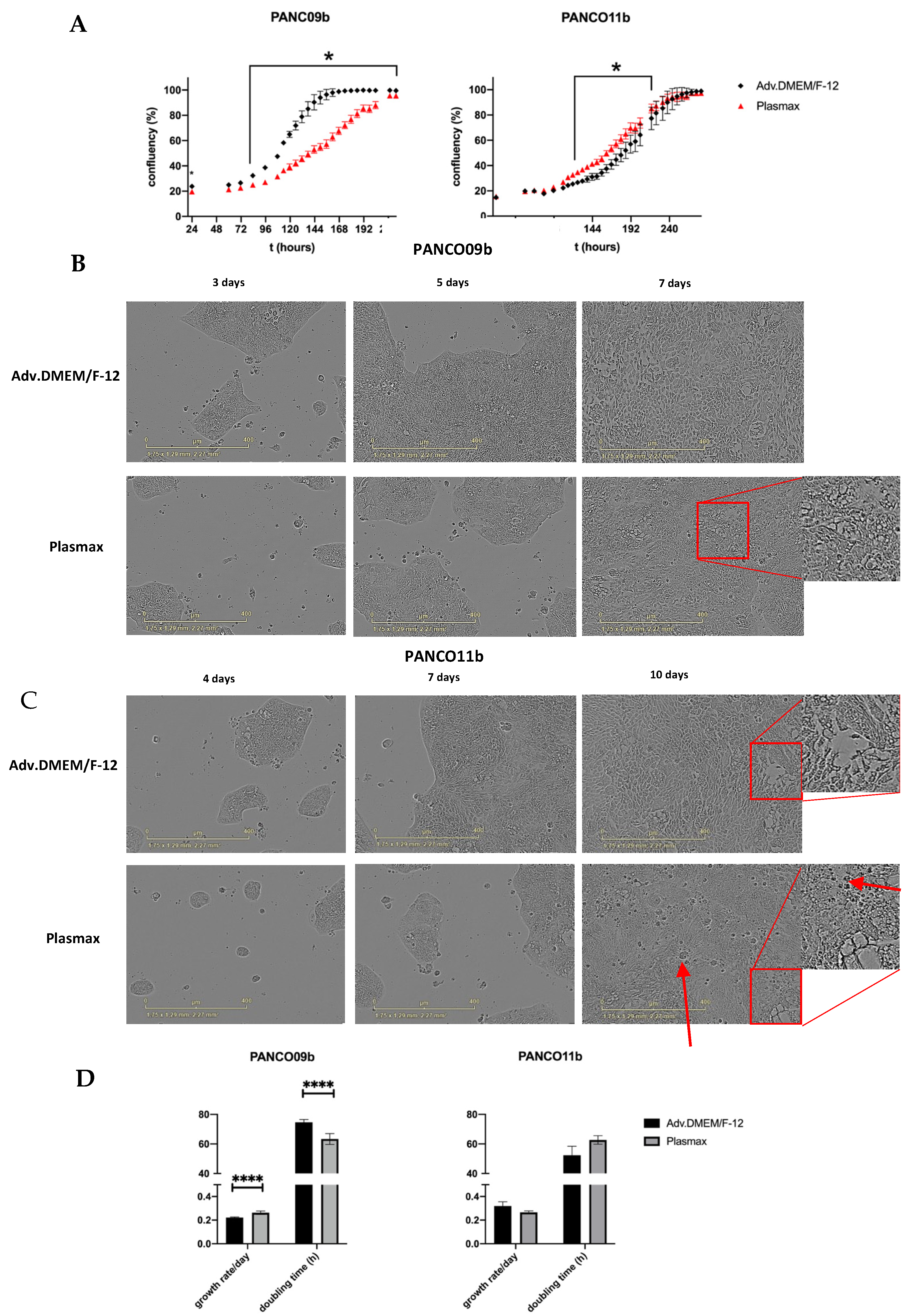
3.3. Impact of Physiological Cell Culture Medium on Proliferation and Morphology of 2D Transformed Cell Cultures
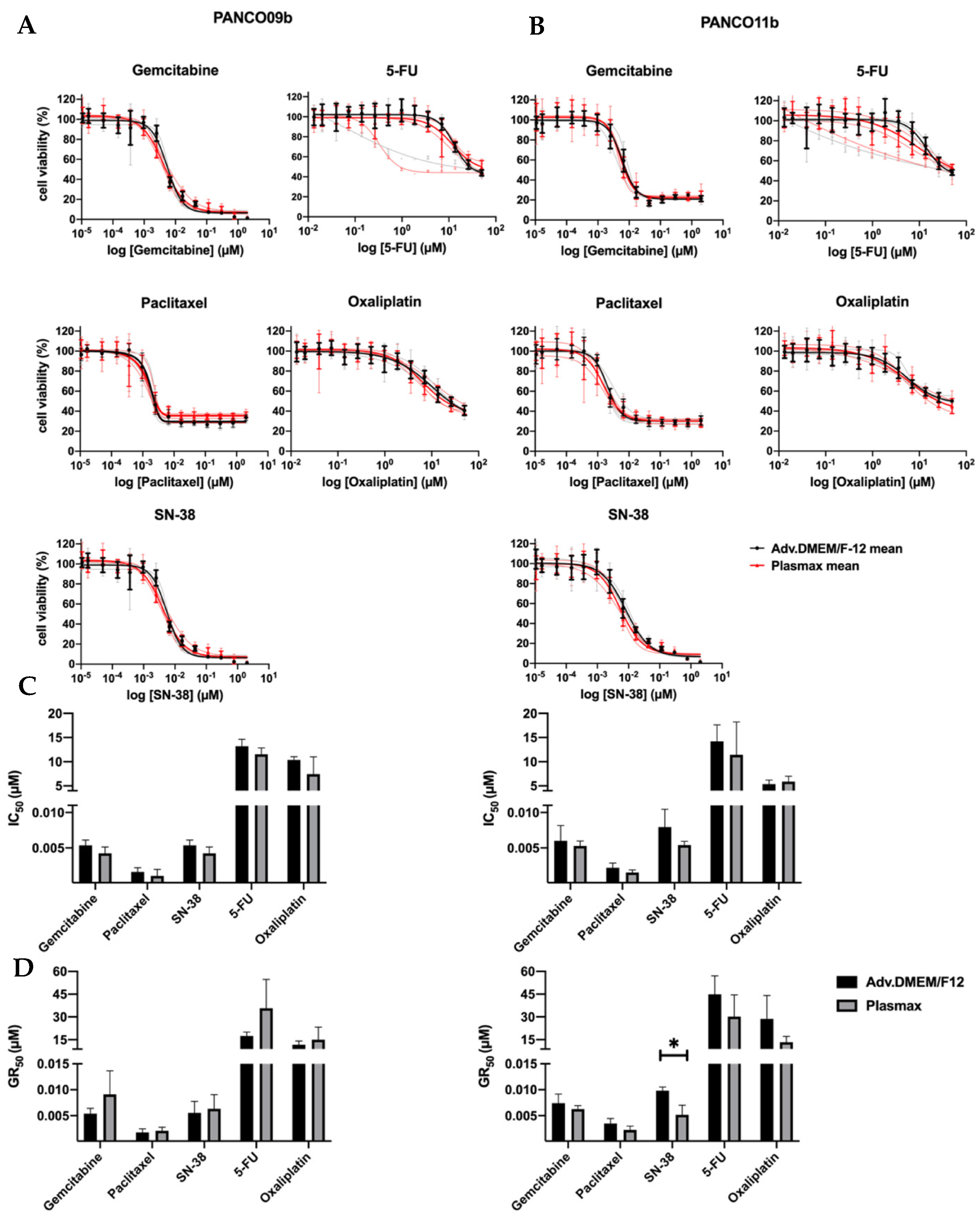
3.4. Physiological Cell Culture Medium Does Not Affect Chemosensitivity of 2D Transformed Cell Cultures
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Rawla, P.; Sunkara, T.; Gaduputi, V. Epidemiology of Pancreatic Cancer: Global Trends, Etiology and Risk Factors. Rev. World J. Oncol. 2019, 10, 10–27. [Google Scholar] [CrossRef] [PubMed]
- Saung, M.T.; Zheng, L. Current Standards of Chemotherapy for Pancreatic Cancer. Clin. Ther. 2017, 39, 2125–2134. [Google Scholar] [CrossRef] [PubMed]
- Springfeld, C.; Jäger, D.; Büchler, M.W.; Strobel, O.; Hackert, T.; Palmer, D.H.; Neoptolemos, J.P. Chemotherapy for pancreatic cancer. Press. Med. 2019, 48, e159–e174. [Google Scholar] [CrossRef] [PubMed]
- Tiriac, H.; Plenker, D.; Baker, L.A.; Tuveson, D.A. Organoid models for translational pancreatic cancer research. Curr. Opin. Genet. Dev. 2019, 54, 7–11. [Google Scholar] [CrossRef]
- Kim, J.; Koo, B.K.; Knoblich, J.A. Human organoids: Model systems for human biology and medicine. Nat. Rev. Mol. Cell Biol. 2020, 21, 571–584. [Google Scholar] [CrossRef]
- Aboulkheyr, E.H.; Montazeri, L.; Aref, A.R.; Vosough, M.; Baharvand, H. Personalized Cancer Medicine: An Organoid Approach. Trends Biotechnol. 2018, 36, 358–371. [Google Scholar] [CrossRef]
- Vaes, R.D.W.; van Dijk, D.P.J.; Welbers, T.T.J.; Blok, M.J.; Aberle, M.R.; Heij, L.; Boj, S.F.; Olde Damink, S.W.M.; Rensen, S.S. Generation and initial characterization of novel tumour organoid models to study human pancreatic cancer-induced cachexia. J. Cachexia. Sarcopenia Muscle 2020, 11, 1509–1524. [Google Scholar] [CrossRef]
- Tiriac, H.; Belleau, P.; Engle, D.D.; Plenker, D.; Deschênes, A.; Somerville, T.D.D.; Froeling, F.E.M.; Burkhart, R.A.; Denroche, R.E.; Jang, G.H.; et al. Organoid profiling identifies common responders to chemotherapy in pancreatic cancer. Cancer Discov. 2018, 8, 1112–1129. [Google Scholar] [CrossRef]
- Kapałczyńska, M.; Kolenda, T.; Przybyła, W.; Zajączkowska, M.; Teresiak, A.; Filas, V.; Ibbs, M.; Bliźniak, R.; Łuczewski, Ł.; Lamperska, K. 2D and 3D cell cultures—A comparison of different types of cancer cell cultures. Arch. Med. Sci. 2018, 14, 910–919. [Google Scholar] [CrossRef]
- Abugomaa, A.; Elbadawy, M.; Yamanaka, M.; Goto, Y.; Hayashi, K.; Mori, T.; Uchide, T.; Azakami, D.; Fukushima, R.; Yoshida, T.; et al. Establishment of 2.5D organoid culture model using 3D bladder cancer organoid culture. Sci. Rep. 2020, 10, 3–5. [Google Scholar] [CrossRef]
- Baker, L.A.; Tiriac, H.; Tuveson, D.A. Generation and culture of human pancreatic ductal adenocarcinoma organoids from resected tumor specimens. Methods Mol. Biol. 2019, 1882, 97–115. [Google Scholar] [PubMed]
- Vande Voorde, J.; Ackermann, T.; Pfetzer, N.; Sumpton, D.; Mackay, G.; Kalna, G.; Nixon, C.; Blyth, K.; Gottlieb, E.; Tardito, S. Improving the metabolic fidelity of cancer models with a physiological cell culture medium. Sci. Adv. 2019, 5, eaau7314. [Google Scholar] [CrossRef] [PubMed]
- Cantor, J.R.; Abu-Remaileh, M.; Kanarek, N.; Freinkman, E.; Gao, X.; Louissaint, A.; Lewis, C.A.; Sabatini, D.M. Physiologic Medium Rewires Cellular Metabolism and Reveals Uric Acid as an Endogenous Inhibitor of UMP Synthase. Cell 2017, 169, 258–272.e17. [Google Scholar] [CrossRef] [PubMed]
- Baghban, R.; Roshangar, L.; Jahanban-Esfahlan, R.; Seidi, K.; Ebrahimi-Kalan, A.; Jaymand, M.; Kolahian, S.; Javaheri, T.; Zare, P. Tumor microenvironment complexity and therapeutic implications at a glance. Cell Commun. Signal. 2020, 18, 59. [Google Scholar] [CrossRef] [PubMed]
- Muir, A.; Vander Heiden, M.G. The nutrient microenvironment affects cancer therapy. Science 2018, 360, 962–963. [Google Scholar] [CrossRef] [PubMed]
- Ackermann, T.; Tardito, S. Cell Culture Medium Formulation and Its Implications in Cancer Metabolism. Trends Cancer 2019, 5, 329–332. [Google Scholar] [CrossRef]
- Online GR Calculator. Available online: http://www.grcalculator.org/grcalculator/ (accessed on 25 May 2022).
- Clark, N.A.; Hafner, M.; Kouril, M.; Williams, E.H.; Muhlich, J.L.; Pilarczyk, M.; Niepel, M.; Sorger, P.K.; Medvedovic, M. GRcalculator: An online tool for calculating and mining dose-response data. BMC Cancer 2017, 17, 698. [Google Scholar] [CrossRef]
- Navani, S. The Human Protein Atlas. J. Obstet. Gynecol. India 2011, 27–31. [Google Scholar] [CrossRef]
- GeneCards. Gene Search: Human Genes. Genes Database. Available online: https://www.genecards.org/ (accessed on 1 September 2022).
- UniProt. Available online: https://www.uniprot.org/ (accessed on 1 September 2022).
- Sun, Y.; Liu, Y.; Ma, X.; Hu, H. Molecular Sciences The Influence of Cell Cycle Regulation on Chemotherapy. Int. J. Mol. Sci. 2021, 22, 6923. [Google Scholar] [CrossRef]
- De Sousa Cavalcante, L.; Monteiro, G. Gemcitabine: Metabolism and molecular mechanisms of action, sensitivity and chemoresistance in pancreatic cancer. Eur. J. Pharmacol. 2014, 741, 8–16. [Google Scholar] [CrossRef]
- Maurya, D.K.; Ayuzawa, R.; Doi, C.; Troyer, D.; Tamura, M. Topoisomerase i inhibitor SN-38 effectively attenuates growth of human non-small-cell lung cancer cell lines in vitro and in vivo. J. Environ. Pathol. Toxicol. Oncol. 2011, 30, 1–10. [Google Scholar] [CrossRef]
- Kampan, N.C.; Madondo, M.T.; McNally, O.M.; Quinn, M.; Plebanski, M. Paclitaxel and its evolving role in the management of ovarian cancer. Bio. Med. Res. Int. 2015, 2015, 413076. [Google Scholar] [CrossRef]
- Zhang, N.; Yin, Y.; Xu, S.J.; Chen, W.S. 5-Fluorouracil: Mechanisms of resistance and reversal strategies. Molecules 2008, 13, 1551–1569. [Google Scholar] [CrossRef]
- Arango, D.; Wilson, A.J.; Shi, Q.; Corner, G.A.; Arañes, M.J.; Nicholas, C.; Lesser, M.; Mariadason, J.M.; Augenlicht, L.H. Molecular mechanisms of action and prediction of response to oxaliplatin in colorectal cancer cells. Br. J. Cancer 2004, 91, 1931–1946. [Google Scholar] [CrossRef]
- Minami, F.; Sasaki, N.; Shichi, Y.; Gomi, F.; Michishita, M.; Ohkusu-Tsukada, K.; Toyoda, M.; Takahashi, K.; Ishiwata, T. Morphofunctional analysis of human pancreatic cancer cell lines in 2- and 3-dimensional cultures. Sci. Rep. 2021, 11, 6775. [Google Scholar] [CrossRef] [PubMed]
- Ben-David, U.; Siranosian, B.; Ha, G.; Tang, H.; Oren, Y.; Hinohara, K.; Strathdee, C.A.; Dempster, J.; Lyons, N.J.; Burns, R.; et al. Genetic and transcriptional evolution alters cancer cell line drug response. Nature 2018, 560, 325–330. [Google Scholar] [CrossRef]
- Zhou, Z.; Cong, L.; Cong, X. Patient-Derived Organoids in Precision Medicine: Drug Screening, Organoid-on-a-Chip and Living Organoid Biobank. Front. Oncol. 2021, 11, 762184. [Google Scholar] [CrossRef] [PubMed]
- Fontoura, J.C.; Viezzer, C.; dos Santos, F.G.; Ligabue, R.A.; Weinlich, R.; Puga, R.D.; Antonow, D.; Severino, P.; Bonorino, C. Comparison of 2D and 3D cell culture models for cell growth, gene expression and drug resistance. Mater. Sci. Eng. C 2020, 107, 110264. [Google Scholar] [CrossRef]
- Melissaridou, S.; Wiechec, E.; Magan, M.; Jain, M.V.; Chung, M.K.; Farnebo, L.; Roberg, K. The effect of 2D and 3D cell cultures on treatment response, EMT profile and stem cell features in head and neck cancer 11 Medical and Health Sciences 1112 Oncology and Carcinogenesis. Cancer Cell Int. 2019, 19, 16. [Google Scholar] [CrossRef] [PubMed]
- Flörkemeier, I.; Steinhauer, T.N.; Hedemann, N.; Weimer, J.P.; Rogmans, C.; van Mackelenbergh, M.T.; Maass, N.; Clement, B.; Bauerschlag, D.O. High antitumor activity of the dual topoisomerase inhibitor p8-d6 in breast cancer. Cancers 2022, 14, 2. [Google Scholar] [CrossRef]
- Mittal, B.; Tulsyan, S.; Kumar, S.; Mittal, R.D.; Agarwal, G. Cytochrome P450 in Cancer Susceptibility and Treatment. In Advances in Clinical Chemistry; Academic Press Inc.: Cambridge, MA, USA, 2015; Volume 71, pp. 77–139. [Google Scholar]
- Koh, B.; Jeon, H.; Kim, D.; Kang, D.; Kim, K.R. Effect of Fibroblast Co-Culture on the Proliferation, Viability and Drug Response of Colon Cancer Cells. Oncol. Lett. 2019, 17, 2409–2417. [Google Scholar] [CrossRef] [PubMed]
- Palam, L.R.; Gore, J.; Craven, K.E.; Wilson, J.L.; Korc, M. Integrated Stress Response Is Critical for Gemcitabine Resistance in Pancreatic Ductal Adenocarcinoma. Cell Death Dis. 2015, 6, e1913. [Google Scholar] [CrossRef] [PubMed]
- Kumbul, Y.Ç.; Nazıroğlu, M. Paclitaxel Promotes Oxidative Stress–Mediated Human Laryngeal Squamous Tumor Cell Death through the Stimulation of Calcium and Zinc Signaling Pathways: No Synergic Action of Melatonin. Biol. Trace Elem. Res. 2022, 200, 2084–2098. [Google Scholar] [CrossRef] [PubMed]
- Di Cesare Mannelli, L.; Zanardelli, M.; Failli, P.; Ghelardini, C. Oxaliplatin-Induced Oxidative Stress in Nervous System-Derived Cellular Models: Could It Correlate with in Vivo Neuropathy? Free Radic. Biol. Med. 2013, 61, 143–150. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Peralta, R.M.; Avina-Ochoa, N.; Delgoffe, G.M.; Kaech, S.M. Metabolic Regulation of T Cells in the Tumor Microenvironment by Nutrient Availability and Diet. Semin. Immunol. 2021, 52, 101485. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gassl, V.; Aberle, M.R.; Boonen, B.; Vaes, R.D.W.; Olde Damink, S.W.M.; Rensen, S.S. Chemosensitivity of 3D Pancreatic Cancer Organoids Is Not Affected by Transformation to 2D Culture or Switch to Physiological Culture Medium. Cancers 2022, 14, 5617. https://doi.org/10.3390/cancers14225617
Gassl V, Aberle MR, Boonen B, Vaes RDW, Olde Damink SWM, Rensen SS. Chemosensitivity of 3D Pancreatic Cancer Organoids Is Not Affected by Transformation to 2D Culture or Switch to Physiological Culture Medium. Cancers. 2022; 14(22):5617. https://doi.org/10.3390/cancers14225617
Chicago/Turabian StyleGassl, Vincent, Merel R. Aberle, Bas Boonen, Rianne D. W. Vaes, Steven W. M. Olde Damink, and Sander S. Rensen. 2022. "Chemosensitivity of 3D Pancreatic Cancer Organoids Is Not Affected by Transformation to 2D Culture or Switch to Physiological Culture Medium" Cancers 14, no. 22: 5617. https://doi.org/10.3390/cancers14225617
APA StyleGassl, V., Aberle, M. R., Boonen, B., Vaes, R. D. W., Olde Damink, S. W. M., & Rensen, S. S. (2022). Chemosensitivity of 3D Pancreatic Cancer Organoids Is Not Affected by Transformation to 2D Culture or Switch to Physiological Culture Medium. Cancers, 14(22), 5617. https://doi.org/10.3390/cancers14225617




