Role of Sirtuins in the Pathobiology of Onco-Hematological Diseases: A PROSPERO-Registered Study and In Silico Analysis
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Search Strategy
2.2. Search Database
2.3. Gene Expression Profile Using GEPIA Database
3. Results
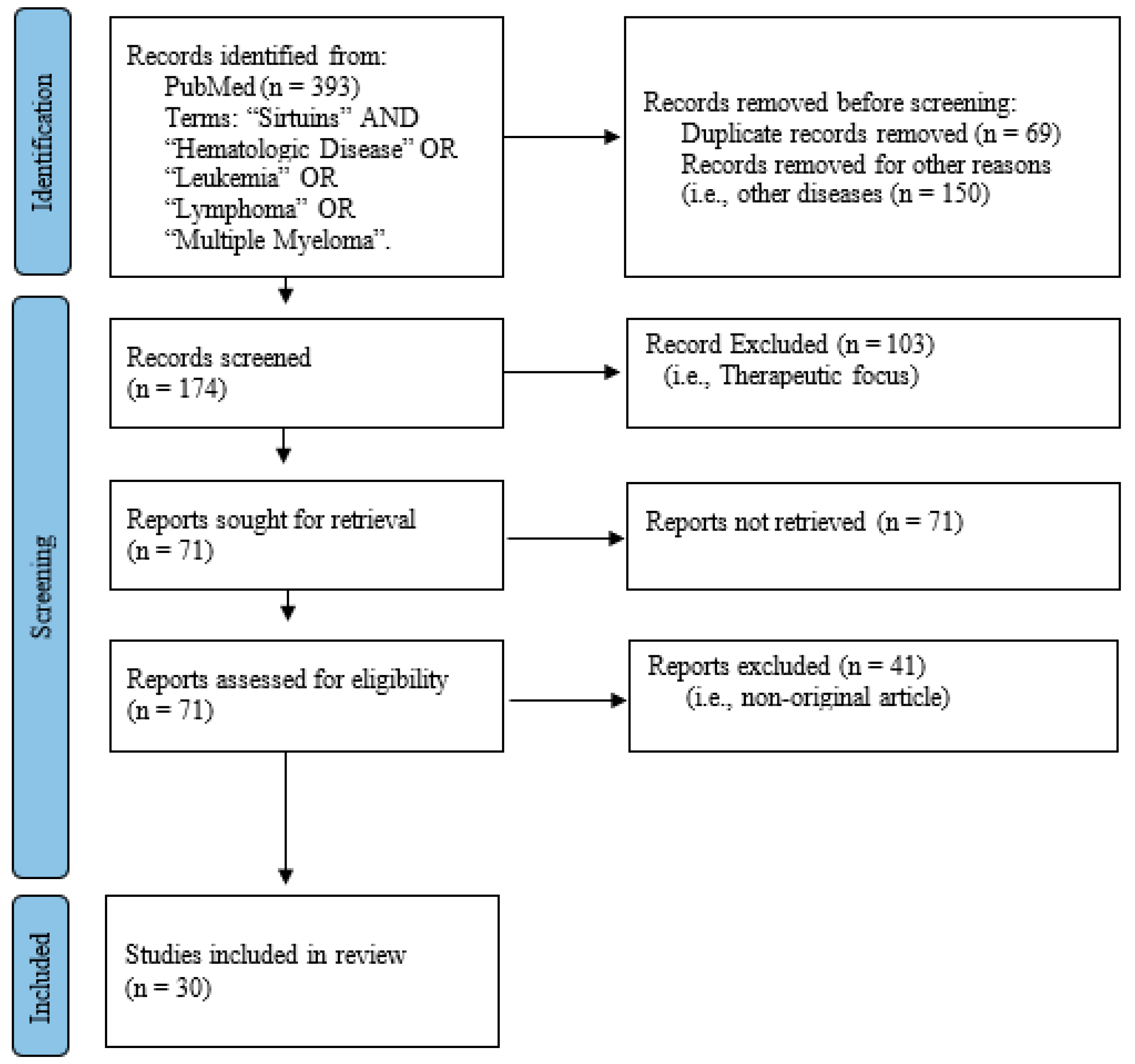
3.1. Search Results
3.2. Lymphoid Neoplasms
3.2.1. Acute Lymphocytic Leukemia (ALL)
| Target | Disease | Main Findings | References |
|---|---|---|---|
| SIRT7 | AML | Low expression in patients’ bone marrow cells | [25] |
| SIRT7 | CML | Low expression in patients’ bone marrow cells | |
| SIRT1 | CLL | High expression in the peripheral blood | [42] |
| SIRT1 | MM | High expression in MM cell line | [43] |
| SIRT1 | CML | High expression in primary human CML cells | [44] |
| SIRT1 | CML | High expression in patient CML cells | [45] |
| SIRT1 | CML | High expression in patient CML cells | [46] |
| SIRT1 | Lymphomas | High expression in patients with Hodgkin’s lymphoma | [47] |
| SIRT1 | AML | High expression of CD34+ and CD38- cells in the bone marrow | [48] |
| SIRT1 | Lymphomas | High expression in cutaneous t-cell lymphomas | [49] |
| SIRT1 | CML | High expression in patient CML cells | [50] |
| SIRT1 | ALL | High expression in patient ALL cells and in cultured ALL lineage cells | [51] |
| SIRT2 | High expression in patient ALL cells and in cultured ALL lineage cells | ||
| SIRT1 | CML | High expression in patient CML cells | [52] |
| SIRT1 | Lymphomas | High expression in patients with Hodgkin’s lymphoma | [53] |
| SIRT1 | Lymphomas | High expression in follicular hyperplasia | [54] |
| SIRT1 | Lymphomas | High expression in cultured lymphoma lineage cells | [55] |
| SIRT1 | MDS | Low expression in patient MDS cells | [56] |
| SIRT1 | CML | High expression in patient CML cells | [57] |
| SIRT1 | MM | High expression in MM cell line | [58] |
| SIRT1 | T-ALL | High expression on T-ALL cells in vivo and in vitro | [59] |
| SIRT1 | CLL | High expression in the peripheral blood | [60] |
| SIRT2 | High expression in the peripheral blood | ||
| SIRT3 | High expression in the peripheral blood | ||
| SIRT4 | Low expression in the peripheral blood | ||
| SIRT5 | Low expression in the peripheral blood | ||
| SIRT6 | High expression in the peripheral blood | ||
| SIRT7 | High expression in the peripheral blood | ||
| SIRT1 | CLL | High expression in cultured CLL lineage cells | [61] |
| SIRT2 | CLL | High expression in cultured CLL lineage cells | |
| SIRT2 | AML | High expression in patient AML cells | [62] |
| SIRT2 | MM | Low expression in patient MM cells | [63] |
| SIRT3 | MM | Low expression in patient MM cells | |
| SIRT2 | ALL | High expression on T-ALL cells both in vitro and in grafts | [64] |
| SIRT2 | AML | High expression in patient AML cells | [65] |
| SIRT3 | CLL | Low expression in patient CLL cells | [66] |
| SIRT3 | Lymphomas | Low expression in mantle cell lymphomas | |
| SIRT3 | AML | Low expression in patient AML cells | [67] |
| SIRT6 | MM | High expression in patient MM cells | [68] |
| SIRT6 | AML | High expression of CD34+ cells in the bone marrow | [69] |
| SIRT6 | Lymphomas | High expression in diffuse B-cell lymphomas | [70] |
3.2.2. Chronic Lymphocytic Leukemia (CLL)
3.2.3. Lymphomas
3.2.4. Multiple Myeloma
3.3. Myeloid Neoplasms
3.3.1. Acute Myeloid Leukemia (AML)
3.3.2. Chronic Myeloid Leukemia (CML)
3.3.3. Myelodysplastic Syndrome (MDS)
3.4. Sirtuin Gene Expression Profile in Hematological Malignances
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Roth, M.; Wang, Z.; Chen, W. Sirtuins in Hematological Aging and Malignancy. Crit. Rev. Oncog. 2013, 18, 531–547. [Google Scholar] [CrossRef] [PubMed]
- Lichtman, M. The Relationship of Patient Age to the Pathobiology of the Clonal Myeloid Diseases. Semin. Oncol. 2004, 31, 185–197. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.; Shilatifard, A. Epigenetics of Hematopoiesis and Hematological Malignancies. Genes Dev. 2016, 30, 2021–2041. [Google Scholar] [CrossRef] [PubMed]
- Zink, F.; Stacey, S.N.; Norddahl, G.L.; Frigge, M.L.; Magnusson, O.T.; Jonsdottir, I.; Thorgeirsson, T.E.; Sigurdsson, A.; Gudjonsson, S.A.; Gudmundsson, J.; et al. Clonal Hematopoiesis, with and without Candidate Driver Mutations, Is Common in the Elderly. Blood 2017, 130, 742–752. [Google Scholar] [CrossRef]
- Mattiucci, D.; Maurizi, G.; Leoni, P.; Poloni, A. Aging- and Senescence-Associated Changes of Mesenchymal Stromal Cells in Myelodysplastic Syndromes. Cell Transpl. 2018, 27, 754–764. [Google Scholar] [CrossRef]
- Chung, S.S.; Park, C.Y. Aging, Hematopoiesis, and the Myelodysplastic Syndromes. Hematol. Am. Soc Hematol. Educ. Program 2017, 2017, 73–78. [Google Scholar] [CrossRef]
- Kurosawa, S.; Iwama, A. Aging and Leukemic Evolution of Hematopoietic Stem Cells under Various Stress Conditions. Inflamm. Regen. 2020, 40, 29. [Google Scholar] [CrossRef]
- Verovskaya, E.V.; Dellorusso, P.V.; Passegué, E. Losing Sense of Self and Surroundings: Hematopoietic Stem Cell Aging and Leukemic Transformation. Trends Mol. Med. 2019, 25, 494–515. [Google Scholar] [CrossRef]
- Coppé, J.P.; Patil, C.K.; Rodier, F.; Sun, Y.; Muñoz, D.P.; Goldstein, J.; Nelson, P.S.; Desprez, P.Y.; Campisi, J. Senescence-Associated Secretory Phenotypes Reveal Cell-Nonautonomous Functions of Oncogenic RAS and the P53 Tumor Suppressor. PLoS Biol. 2008, 6, e301. [Google Scholar] [CrossRef]
- Lujambio, A. To Clear, or Not to Clear (Senescent Cells)? That Is the Question. Inside Cell 2016, 1, 87–95. [Google Scholar] [CrossRef]
- Corradi, G.; Baldazzi, C.; Očadlíková, D.; Marconi, G.; Parisi, S.; Testoni, N.; Finelli, C.; Cavo, M.; Curti, A.; Ciciarello, M. Mesenchymal Stromal Cells from Myelodysplastic and Acute Myeloid Leukemia Patients Display In Vitro Reduced Proliferative Potential and Similar Capacity to Support Leukemia Cell Survival. Stem Cell Res. Ther. 2018, 9, 271. [Google Scholar] [CrossRef] [PubMed]
- Goulard, M.; Dosquet, C.; Bonnet, D. Role of the Microenvironment in Myeloid Malignancies. Cell. Mol. Life Sci. 2017, 75, 1377–1391. [Google Scholar] [CrossRef] [PubMed]
- Valletta, S.; Thomas, A.; Meng, Y.; Ren, X.; Drissen, R.; Sengül, H.; di Genua, C.; Nerlov, C. Micro-Environmental Sensing by Bone Marrow Stroma Identifies IL-6 and TGFβ1 as Regulators of Hematopoietic Ageing. Nat. Commun. 2020, 11, 4075. [Google Scholar] [CrossRef] [PubMed]
- Kode, A.; Manavalan, J.S.; Mosialou, I.; Bhagat, G.; Rathinam, C.v.; Luo, N.; Khiabanian, H.; Lee, A.; Murty, V.v.; Friedman, R.; et al. Leukaemogenesis Induced by an Activating β-Catenin Mutation in Osteoblasts. Nature 2014, 506, 240–244. [Google Scholar] [CrossRef] [PubMed]
- Zhao, E.; Hou, J.; Ke, X.; Abbas, M.N.; Kausar, S.; Zhang, L.; Cui, H. The Roles of Sirtuin Family Proteins in Cancer Progression. Cancers 2019, 11, 1949. [Google Scholar] [CrossRef]
- Bedolla, M.A.H.; Espinosa, A.B.; Cortés, F.O.; Alonso, D.E.A.; Hernández, A.H.; Ostoa, Z.B.; Salguero, I.G.D. Papel Pleiotrópico y Homeostático de Las Sirtuinas En La Función Biológica Humana. Ciencia Huasteca Boletín Científico Esc. Super. Huejutla 2020, 8, 6–20. [Google Scholar] [CrossRef]
- Lee, S.H.; Lee, J.H.; Lee, H.Y.; Min, K.J. Sirtuin Signaling in Cellular Senescence and Aging. BMB Rep. 2019, 52, 24. [Google Scholar] [CrossRef]
- Martens, C.R.; Denman, B.A.; Mazzo, M.R.; Armstrong, M.L.; Reisdorph, N.; McQueen, M.B.; Chonchol, M.; Seals, D.R. Chronic Nicotinamide Riboside Supplementation Is Well-Tolerated and Elevates NAD + in Healthy Middle-Aged and Older Adults. Nat. Commun. 2018, 9, 1286. [Google Scholar] [CrossRef]
- Mendelsohn, A.R.; Larrick, J.W. Interacting NAD+ and Cell Senescence Pathways Complicate Antiaging Therapies. Rejuvenation Res. 2019, 22, 261–266. [Google Scholar] [CrossRef]
- Moon, J.; Kim, H.R.; Shin, M.G. Rejuvenating Aged Hematopoietic Stem Cells Through Improvement of Mitochondrial Function. Ann. Lab. Med. 2018, 38, 395–401. [Google Scholar] [CrossRef]
- O’Callaghan, C.; Vassilopoulos, A. Sirtuins at the Crossroads of Stemness, Aging, and Cancer. Aging Cell 2017, 16, 1208–1218. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Mu, W.C.; Karki, R.; Chiang, H.H.; Mohrin, M.; Shin, J.J.; Ohkubo, R.; Ito, K.; Kanneganti, T.D.; Chen, D. Mitochondrial Stress-Initiated Aberrant Activation of the NLRP3 Inflammasome Regulates the Functional Deterioration of Hematopoietic Stem Cell Aging. Cell Rep. 2019, 26, 945–954.e4. [Google Scholar] [CrossRef] [PubMed]
- Brown, K.; Xie, S.; Qiu, X.; Mohrin, M.; Shin, J.; Liu, Y.; Zhang, D.; Scadden, D.T.; Chen, D. SIRT3 Reverses Aging-Associated Degeneration. Cell Rep. 2013, 3, 319–327. [Google Scholar] [CrossRef] [PubMed]
- Mohrin, M.; Shin, J.; Liu, Y.; Brown, K.; Luo, H.; Xi, Y.; Haynes, C.M.; Chen, D. Stem Cell Aging. A Mitochondrial UPR-Mediated Metabolic Checkpoint Regulates Hematopoietic Stem Cell Aging. Science 2015, 347, 1374–1377. [Google Scholar] [CrossRef]
- Kaiser, A.; Schmidt, M.; Huber, O.; Frietsch, J.J.; Scholl, S.; Heidel, F.H.; Hochhaus, A.; Müller, J.P.; Ernst, T. SIRT7: An Influence Factor in Healthy Aging and the Development of Age-Dependent Myeloid Stem-Cell Disorders. Leukemia 2020, 34, 2206–2216. [Google Scholar] [CrossRef]
- Bi, S.; Liu, Z.; Wu, Z.; Wang, Z.; Liu, X.; Wang, S.; Ren, J.; Yao, Y.; Zhang, W.; Song, M.; et al. SIRT7 Antagonizes Human Stem Cell Aging as a Heterochromatin Stabilizer. Protein Cell 2020, 11, 483–504. [Google Scholar] [CrossRef]
- Tasselli, L.; Xi, Y.; Zheng, W.; Tennen, R.I.; Odrowaz, Z.; Simeoni, F.; Li, W.; Chua, K.F. SIRT6 Deacetylates H3K18ac at Pericentric Chromatin to Prevent Mitotic Errors and Cellular Senescence. Nat. Struct. Mol. Biol. 2016, 23, 434–440. [Google Scholar] [CrossRef]
- Minten, E.V.; Kapoor-Vazirani, P.; Li, C.; Zhang, H.; Balakrishnan, K.; Yu, D.S. SIRT2 Promotes BRCA1-BARD1 Heterodimerization through Deacetylation. Cell Rep 2021, 34, 108921. [Google Scholar] [CrossRef]
- Shi, Q.; Liu, T.; Zhang, X.; Geng, J.; He, X.; Nu, M.; Pang, D. Decreased Sirtuin 4 Expression Is Associated with Poor Prognosis in Patients with Invasive Breast Cancer. Oncol. Lett. 2016, 12, 2606–2612. [Google Scholar] [CrossRef]
- Tang, X.; Li, Y.; Liu, L.; Guo, R.; Zhang, P.; Zhang, Y.; Zhang, Y.; Zhao, J.; Su, J.; Sun, L.; et al. Sirtuin 3 Induces Apoptosis and Necroptosis by Regulating Mutant P53 Expression in Small-cell Lung Cancer. Oncol. Rep. 2019, 43, 591–600. [Google Scholar] [CrossRef]
- Wu, X.; Wang, S.; Zhao, X.; Lai, S.; Yuan, Z.; Zhan, Y.; Ni, K.; Liu, Z.; Liu, L.; Xin, R.; et al. Clinicopathological and Prognostic Value of SIRT6 in Patients with Solid Tumors: A Meta-Analysis and TCGA Data Review. Cancer Cell Int. 2022, 22, 84. [Google Scholar] [CrossRef] [PubMed]
- Grbesa, I.; Pajares, M.J.; Martínez-Terroba, E.; Agorreta, J.; Mikecin, A.-M.; Larráyoz, M.; Idoate, M.A.; Gall-Troselj, K.; Pio, R.; Montuenga, L.M. Expression of Sirtuin 1 and 2 Is Associated with Poor Prognosis in Non-Small Cell Lung Cancer Patients. PLoS ONE 2015, 10, e0124670. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Lockwood, C.; Munn, Z.; Porritt, K. Qualitative Research Synthesis: Methodological Guidance for Systematic Reviewers Utilizing Meta-Aggregation. Int. J. Evid. Based Healthc. 2015, 13, 179–187. [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 Statement: An Updated Guideline for Reporting Systematic Reviews. BMJ 2021, 372, n71. [Google Scholar] [CrossRef]
- Tang, Z.; Li, C.; Kang, B.; Gao, G.; Li, C.; Zhang, Z. GEPIA: A Web Server for Cancer and Normal Gene Expression Profiling and Interactive Analyses. Nucleic Acids Res 2017, 45, W98–W102. [Google Scholar] [CrossRef]
- O’Brien, M.M.; Seif, A.E.; Hunger, S.P. Acute lymphoblastic leukemia in children. In Wintrobe’s Clinical Hematology, 14th ed.; Lippincott Williams & Wilkins: Philadeplhia, PA, USA, 2018; pp. 4939–5015. [Google Scholar] [CrossRef]
- Bhojwani, D.; Yang, J.J.; Pui, C.H. Biology of Childhood Acute Lymphoblastic Leukemia. Pediatr. Clin. N. Am. 2015, 62, 47–60. [Google Scholar] [CrossRef]
- Onciu, M. Acute Lymphoblastic Leukemia. Hematol. Oncol. Clin. N. Am. 2009, 23, 655–674. [Google Scholar] [CrossRef]
- Aldoss, I.; Stein, A.S. Advances in Adult Acute Lymphoblastic Leukemia Therapy. Leuk Lymphoma 2018, 59, 1033–1050. [Google Scholar] [CrossRef]
- Aldoss, I.; Forman, S.J.; Pullarkat, V. Acute Lymphoblastic Leukemia in the Older Adult. J. Oncol. Pract. 2019, 15, 67–75. [Google Scholar] [CrossRef]
- Audrito, V.; Vaisitti, T.; Rossi, D.; Gottardi, D.; D’Arena, G.; Laurenti, L.; Gaidano, G.; Malavasi, F.; Deaglio, S. Nicotinamide Blocks Proliferation and Induces Apoptosis of Chronic Lymphocytic Leukemia Cells through Activation of the P53/MiR-34a/SIRT1 Tumor Suppressor Network. Cancer Res. 2011, 71, 4473–4483. [Google Scholar] [CrossRef] [PubMed]
- Chauhan, D.; Bandi, M.; Singh, A.V.; Ray, A.; Raje, N.; Richardson, P.; Anderson, K.C. Preclinical Evaluation of a Novel SIRT1 Modulator SRT1720 in Multiple Myeloma Cells. Br. J. Haematol. 2011, 155, 588–598. [Google Scholar] [CrossRef] [PubMed]
- Yuan, H.; Wang, Z.; Li, L.; Zhang, H.; Modi, H.; Horne, D.; Bhatia, R.; Chen, W.Y. Activation of Stress Response Gene SIRT1 by BCR-ABL Promotes Leukemogenesis. Blood 2012, 119, 1904–1914. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Wang, L.; Li, L.; Wang, Z.; Ho, Y.; McDonald, T.; Holyoake, T.L.; Chen, W.Y.; Bhatia, R. Activation of P53 by SIRT1 Inhibition Enhances Elimination of CML Leukemia Stem Cells in Combination with Imatinib. Cancer Cell 2012, 21, 266–281. [Google Scholar] [CrossRef] [PubMed]
- Abraham, A.; Qiu, S.; Chacko, B.K.; Li, H.; Paterson, A.; He, J.; Agarwal, P.; Shah, M.; Welner, R.; Darley-Usmar, V.M.; et al. SIRT1 Regulates Metabolism and Leukemogenic Potential in CML Stem Cells. J. Clin. Investig. 2019, 129, 2685–2701. [Google Scholar] [CrossRef]
- Frazzi, R.; Valli, R.; Tamagnini, I.; Casali, B.; Latruffe, N.; Merli, F. Resveratrol-Mediated Apoptosis of Hodgkin Lymphoma Cells Involves SIRT1 Inhibition and FOXO3a Hyperacetylation. Int. J. Cancer 2013, 132, 1013–1021. [Google Scholar] [CrossRef]
- Sasca, D.; Hähnel, P.S.; Szybinski, J.; Khawaja, K.; Kriege, O.; Pante, S.V.; Bullinger, L.; Strand, S.; Strand, D.; Theobald, M.; et al. SIRT1 Prevents Genotoxic Stress-Induced P53 Activation in Acute Myeloid Leukemia. Blood 2014, 124, 121–133. [Google Scholar] [CrossRef]
- Nihal, M.; Ahmad, N.; Wood, G.S. SIRT1 Is Upregulated in Cutaneous T-Cell Lymphoma, and Its Inhibition Induces Growth Arrest and Apoptosis. Cell Cycle 2013, 13, 632–640. [Google Scholar] [CrossRef]
- Kim, H.B.; Lee, S.H.; Um, J.H.; Kim, M.J.; Hyun, S.K.; Gong, E.J.; Oh, W.K.; Kang, C.D.; Kim, S.H. Sensitization of Chemo-Resistant Human Chronic Myeloid Leukemia Stem-like Cells to Hsp90 Inhibitor by SIRT1 Inhibition. Int. J. Biol. Sci. 2015, 11, 923–934. [Google Scholar] [CrossRef][Green Version]
- Jin, Y.; Cao, Q.; Chen, C.; Du, X.; Jin, B.; Pan, J. Tenovin-6-Mediated Inhibition of SIRT1/2 Induces Apoptosis in Acute Lymphoblastic Leukemia (ALL) Cells and Eliminates ALL Stem/Progenitor Cells. BMC Cancer 2015, 15, 226. [Google Scholar] [CrossRef]
- Zhang, W.; Wu, H.; Yang, M.; Ye, S.; Li, L.; Zhang, H.; Hu, J.; Wang, X.; Xu, J.; Liang, A.; et al. SIRT1 Inhibition Impairs Non-Homologous End Joining DNA Damage Repair by Increasing Ku70 Acetylation in Chronic Myeloid Leukemia Cells. Oncotarget 2015, 7, 13538–13550. [Google Scholar] [CrossRef]
- Quesada, A.E.; Assylbekova, B.; Jabcuga, C.E.; Zhang, R.; Covinsky, M.; Rios, A.; Nguyen, N.D.; Brown, R.E. Expression of Sirt1 and FoxP3 in Classical Hodgkin Lymphoma and Tumor Infiltrating Lymphocytes: Implications for Immune Dysregulation, Prognosis and Potential Therapeutic Targeting. Int. J. Clin. Exp. Pathol. 2015, 8, 13241–13248. [Google Scholar] [PubMed]
- Frazzi, R.; Zanetti, E.; Pistoni, M.; Tamagnini, I.; Valli, R.; Braglia, L.; Merli, F. Methylation Changes of SIRT1, KLF4, DAPK1 and SPG20 in B-Lymphocytes Derived from Follicular and Diffuse Large B-Cell Lymphoma. Leuk. Res. 2017, 57, 89–96. [Google Scholar] [CrossRef]
- He, M.; Tan, B.; Vasan, K.; Yuan, H.; Cheng, F.; Ramos da Silva, S.; Lu, C.; Gao, S.J. SIRT1 and AMPK Pathways Are Essential for the Proliferation and Survival of Primary Effusion Lymphoma Cells. J. Pathol. 2017, 242, 309–321. [Google Scholar] [CrossRef]
- Sun, J.; He, X.; Zhu, Y.; Ding, Z.; Dong, H.; Feng, Y.; Du, J.; Wang, H.; Wu, X.; Zhang, L.; et al. SIRT1 Activation Disrupts Maintenance of Myelodysplastic Syndrome Stem and Progenitor Cells by Restoring TET2 Function. Cell Stem Cell 2018, 23, 355–369.e9. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wang, M.; Dou, H.; Lin, W.; Zou, L. Sirtuin 1 Inhibits Lipopolysaccharide-Induced Inflammation in Chronic Myelogenous Leukemia K562 Cells through Interacting with the Toll-like Receptor 4-Nuclear Factor κ B-Reactive Oxygen Species Signaling Axis. Cancer Cell Int. 2020, 20, 73. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Liu, J.; Jiang, H.; Wang, J.; Li, X.; Wang, J.; Zhu, S.; Guo, J.; Li, T.; Zhong, Y.; et al. Proteasome Inhibitor Induced SIRT1 Deacetylates GLI2 to Enhance Hedgehog Signaling Activity and Drug Resistance in Multiple Myeloma. Oncogene 2019, 39, 922–934. [Google Scholar] [CrossRef]
- Wang, F.; Li, Z.; Zhou, J.; Wang, G.; Zhang, W.; Xu, J.; Liang, A. SIRT1 Regulates the Phosphorylation and Degradation of P27 by Deacetylating CDK2 to Promote T-Cell Acute Lymphoblastic Leukemia Progression. J. Exp. Clin. Cancer Res. 2021, 40, 259. [Google Scholar] [CrossRef]
- Van Damme, M.; Crompot, E.; Meuleman, N.; Mineur, P.; Bron, D.; Lagneaux, L.; Stamatopoulos, B. HDAC Isoenzyme Expression Is Deregulated in Chronic Lymphocytic Leukemia B-Cells and Has a Complex Prognostic Significance. Epigenetics 2012, 7, 1403. [Google Scholar] [CrossRef]
- Bhalla, S.; Gordon, L.I. Functional Characterization of NAD Dependent De-Acetylases SIRT1 and SIRT2 in B-Cell Chronic Lymphocytic Leukemia (CLL). Cancer Biol. Ther. 2016, 17, 300–309. [Google Scholar] [CrossRef]
- Dan, L.; Klimenkova, O.; Klimiankou, M.; Klusman, J.H.; van den Heuvel-Eibrink, M.M.; Reinhardt, D.; Welte, K.; Skokowa, J. The Role of Sirtuin 2 Activation by Nicotinamide Phosphoribosyltransferase in the Aberrant Proliferation and Survival of Myeloid Leukemia Cells. Haematologica 2012, 97, 551. [Google Scholar] [CrossRef] [PubMed]
- Allegra, A.; Innao, V.; Polito, F.; Oteri, R.; Alibrandi, A.; Allegra, A.G.; Oteri, G.; di Giorgio, R.M.; Musolino, C.; Aguennouz, M. SIRT2 and SIRT3 Expression Correlates with Redox Imbalance and Advanced Clinical Stage in Patients with Multiple Myeloma. Clin. Biochem. 2021, 93, 42–49. [Google Scholar] [CrossRef] [PubMed]
- Morishima, T.; Krahl, A.C.; Nasri, M.; Xu, Y.; Aghaallaei, N.; Findik, B.; Klimiankou, M.; Ritter, M.; Hartmann, M.D.; Gloeckner, C.J.; et al. LMO2 Activation by Deacetylation Is Indispensable for Hematopoiesis and T-ALL Leukemogenesis. Blood 2019, 134, 1159–1175. [Google Scholar] [CrossRef] [PubMed]
- Deng, A.; Ning, Q.; Zhou, L.; Liang, Y. SIRT2 Is an Unfavorable Prognostic Biomarker in Patients with Acute Myeloid Leukemia. Sci. Rep. 2016, 6, 27694. [Google Scholar] [CrossRef] [PubMed]
- Yu, W.; Denu, R.A.; Krautkramer, K.A.; Grindle, K.M.; Yang, D.T.; Asimakopoulos, F.; Hematti, P.; Denu, J.M. Loss of SIRT3 Provides Growth Advantage for B Cell Malignancies. J. Biol. Chem. 2016, 291, 3268. [Google Scholar] [CrossRef]
- Ma, J.; Liu, B.; Yu, D.; Zuo, Y.; Cai, R.; Yang, J.; Cheng, J. SIRT3 Deacetylase Activity Confers Chemoresistance in AML via Regulation of Mitochondrial Oxidative Phosphorylation. Br. J. Haematol. 2019, 187, 49. [Google Scholar] [CrossRef]
- Cea, M.; Cagnetta, A.; Adamia, S.; Acharya, C.; Tai, Y.T.; Fulciniti, M.; Ohguchi, H.; Munshi, A.; Acharya, P.; Bhasin, M.K.; et al. Evidence for a Role of the Histone Deacetylase SIRT6 in DNA Damage Response of Multiple Myeloma Cells. Blood 2016, 127, 1138–1150. [Google Scholar] [CrossRef]
- Cagnetta, A.; Soncini, D.; Orecchioni, S.; Talarico, G.; Minetto, P.; Guolo, F.; Retali, V.; Colombo, N.; Carminati, E.; Clavio, M.; et al. Depletion of SIRT6 Enzymatic Activity Increases Acute Myeloid Leukemia Cells’ Vulnerability to DNA-Damaging Agents. Haematologica 2018, 103, 80. [Google Scholar] [CrossRef]
- Yang, J.; Li, Y.; Zhang, Y.; Fang, X.; Chen, N.; Zhou, X.; Wang, X. Sirt6 Promotes Tumorigenesis and Drug Resistance of Diffuse Large B-Cell Lymphoma by Mediating PI3K/Akt Signaling. J. Exp. Clin. Cancer Res. 2020, 39, 142. [Google Scholar] [CrossRef]
- Rodrigues, C.A.; Gonçalves, M.V.; Ikoma, M.R.V.; Lorand-Metze, I.; Pereira, A.D.; de Farias, D.L.C.; de Chauffaille, M.L.L.F.; Schaffel, R.; Ribeiro, E.F.O.; da Rocha, T.S.; et al. Diagnosis and Treatment of Chronic Lymphocytic Leukemia: Recommendations from the Brazilian Group of Chronic Lymphocytic Leukemia. Rev. Bras. Hematol. Hemoter. 2016, 38, 346–357. [Google Scholar] [CrossRef]
- Burger, J.A. Treatment of Chronic Lymphocytic Leukemia. New Engl. J. Med. 2020, 383, 460–473. [Google Scholar] [CrossRef] [PubMed]
- Hodgson, K.; Ferrer, G.; Montserrat, E.; Moreno, C. Chronic Lymphocytic Leukemia and Autoimmunity: A Systematic Review. Haematologica 2011, 96, 752. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M.; Bennani, N.N.; Feldman, A.L. Lymphoma Classification Update: T-Cell Lymphomas, Hodgkin Lymphomas, and Histiocytic/Dendritic Cell Neoplasms. Expert Rev. Hematol. 2017, 10, 239–249. [Google Scholar] [CrossRef] [PubMed]
- De Paepe, P.; de Wolf-Peeters, C. Diffuse Large B-Cell Lymphoma: A Heterogeneous Group of Non-Hodgkin Lymphomas Comprising Several Distinct Clinicopathological Entities. Leukemia 2006, 21, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Van de Donk, N.W.C.J.; Pawlyn, C.; Yong, K.L. Multiple Myeloma. Lancet 2021, 397, 410–427. [Google Scholar] [CrossRef]
- Narayanan, D.; Weinberg, O.K. How I Investigate Acute Myeloid Leukemia. Int. J. Lab. Hematol. 2020, 42, 3–15. [Google Scholar] [CrossRef]
- Pourrajab, F.; Zare-Khormizi, M.R.; Hashemi, A.S.; Hekmatimoghaddam, S. Genetic Characterization and Risk Stratification of Acute Myeloid Leukemia. Cancer Manag. Res. 2020, 12, 2231–2253. [Google Scholar] [CrossRef]
- Arber, D.A.; Orazi, A.; Hasserjian, R.; Thiele, J.; Borowitz, M.J.; le Beau, M.M.; Bloomfield, C.D.; Cazzola, M.; Vardiman, J.W. The 2016 Revision to the World Health Organization Classification of Myeloid Neoplasms and Acute Leukemia. Blood 2016, 127, 2391–2405. [Google Scholar] [CrossRef]
- Papaemmanuil, E.; Gerstung, M.; Bullinger, L.; Gaidzik, V.I.; Paschka, P.; Roberts, N.D.; Potter, N.E.; Heuser, M.; Thol, F.; Bolli, N.; et al. Genomic Classification and Prognosis in Acute Myeloid Leukemia. New Engl. J. Med. 2016, 374, 2209–2221. [Google Scholar] [CrossRef]
- Deininger, M.W.N.; Goldman, J.M.; Melo, J.V. The Molecular Biology of Chronic Myeloid Leukemia. Blood 2000, 96, 3343–3356. [Google Scholar] [CrossRef]
- An, X.; Tiwari, A.K.; Sun, Y.; Ding, P.R.; Ashby, C.R.; Chen, Z.S. BCR-ABL Tyrosine Kinase Inhibitors in the Treatment of Philadelphia Chromosome Positive Chronic Myeloid Leukemia: A Review. Leuk. Res. 2010, 34, 1255–1268. [Google Scholar] [CrossRef] [PubMed]
- Vardiman, J.W. Chronic Myelogenous Leukemia, BCR-ABL1+. Am. J. Clin. Pathol. 2009, 132, 250–260. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Kao, Y.-R.; Sun, D.; Todorova, T.I.; Reynolds, D.; Narayanagari, S.-R.; Montagna, C.; Will, B.; Verma, A.; Steidl, U. Myelodysplastic Syndrome Progression to Acute Myeloid Leukemia at the Stem Cell Level. Nat. Med. 2019, 25, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Hasserjian, R.P. Myelodysplastic Syndrome Updated. Pathobiology 2019, 86, 53–61. [Google Scholar] [CrossRef]
- Bejar, R.; Stevenson, K.; Abdel-Wahab, O.; Galili, N.; Nilsson, B.; Garcia-Manero, G.; Kantarjian, H.; Raza, A.; Levine, R.L.; Neuberg, D.; et al. Clinical Effect of Point Mutations in Myelodysplastic Syndromes. N. Engl. J. Med. 2011, 364, 2496–2506. [Google Scholar] [CrossRef]
- Haferlach, T.; Nagata, Y.; Grossmann, V.; Okuno, Y.; Bacher, U.; Nagae, G.; Schnittger, S.; Sanada, M.; Kon, A.; Alpermann, T.; et al. Landscape of Genetic Lesions in 944 Patients with Myelodysplastic Syndromes. Leukemia 2013, 28, 241–247. [Google Scholar] [CrossRef]
- Bernard, E.; Nannya, Y.; Hasserjian, R.P.; Devlin, S.M.; Tuechler, H.; Medina-Martinez, J.S.; Yoshizato, T.; Shiozawa, Y.; Saiki, R.; Malcovati, L.; et al. Implications of TP53 Allelic State for Genome Stability, Clinical Presentation and Outcomes in Myelodysplastic Syndromes. Nat. Med. 2020, 26, 1549–1556. [Google Scholar] [CrossRef]
- Belli, C.B.; Pinheiro, R.F.; Bestach, Y.; Larripa, I.B.; da Silva Tanizawa, R.S.; Alfonso, G.; Gonzalez, J.; Rosenhain, M.; Watman, N.; Cavalcante de Andrade Silva, M.; et al. Myelodysplastic Syndromes in South America: A Multinational Study of 1080 Patients. Am. J. Hematol. 2015, 90, 851–858. [Google Scholar] [CrossRef]
- Yamakuchi, M.; Lowenstein, C.J. MiR-34, SIRT1, and P53: The Feedback Loop. Cell Cycle 2009, 8, 712–715. [Google Scholar] [CrossRef]
- Villalba, J.M.; Alcaín, F.J. Sirtuin Activators and Inhibitors. Biofactors 2012, 38, 349–359. [Google Scholar] [CrossRef]
- Hu, J.; Jing, H.; Lin, H. Sirtuin Inhibitors as Anticancer Agents. Future Med. Chem. 2014, 6, 945. [Google Scholar] [CrossRef]
- Bai, X.; Yao, L.; Ma, X.; Xu, X. Small Molecules as SIRT Modulators. Mini-Rev. Med. Chem. 2016, 18, 1151–1157. [Google Scholar] [CrossRef]
- Song, X.R.; Cheng, Y.Q.; Su, D.F.; Liu, A.J. The Sirt1 Activator Resveratrol Improved Hematopoiesis in Pancytopenia Mice Induced by Irradiation. J. Pharmacol. Sci. 2019, 140, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.M.; Galson, D.L.; Roodman, G.D.; Ouyang, H. Resveratrol Triggers the Pro-Apoptotic Endoplasmic Reticulum Stress Response and Represses pro-Survival XBP1 Signaling in Human Multiple Myeloma Cells. Exp. Hematol. 2011, 39, 999–1006. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.S.; Deater, M.; Schubert, K.; Marquez-Loza, L.; Pelz, C.; Sinclair, D.A.; Grompe, M. The Sirt1 Activator SRT3025 Expands Hematopoietic Stem and Progenitor Cells and Improves Hematopoiesis in Fanconi Anemia Mice. Stem Cell Res. 2015, 15, 130–140. [Google Scholar] [CrossRef]
- Wang, Z.; Yuan, H.; Roth, M.; Stark, J.M.; Bhatia, R.; Chen, W.Y. SIRT1 Deacetylase Promotes Acquisition of Genetic Mutations for Drug Resistance in CML Cells. Oncogene 2013, 32, 589. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Osdal, T.; Ho, Y.; Chun, S.; McDonald, T.; Agarwal, P.; Lin, A.; Chu, S.; Qi, J.; Li, L.; et al. SIRT1 Activation by a C-MYC Oncogenic Network Promotes the Maintenance and Drug Resistance of Human FLT3-ITD Acute Myeloid Leukemia Stem Cells. Cell Stem Cell 2014, 15, 431. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Goes, J.V.C.; Carvalho, L.G.; de Oliveira, R.T.G.; Melo, M.M.d.L.; Novaes, L.A.C.; Moreno, D.A.; Gonçalves, P.G.; Montefusco-Pereira, C.V.; Pinheiro, R.F.; Ribeiro Junior, H.L. Role of Sirtuins in the Pathobiology of Onco-Hematological Diseases: A PROSPERO-Registered Study and In Silico Analysis. Cancers 2022, 14, 4611. https://doi.org/10.3390/cancers14194611
Goes JVC, Carvalho LG, de Oliveira RTG, Melo MMdL, Novaes LAC, Moreno DA, Gonçalves PG, Montefusco-Pereira CV, Pinheiro RF, Ribeiro Junior HL. Role of Sirtuins in the Pathobiology of Onco-Hematological Diseases: A PROSPERO-Registered Study and In Silico Analysis. Cancers. 2022; 14(19):4611. https://doi.org/10.3390/cancers14194611
Chicago/Turabian StyleGoes, João Vitor Caetano, Luiz Gustavo Carvalho, Roberta Taiane Germano de Oliveira, Mayara Magna de Lima Melo, Lázaro Antônio Campanha Novaes, Daniel Antunes Moreno, Paola Gyuliane Gonçalves, Carlos Victor Montefusco-Pereira, Ronald Feitosa Pinheiro, and Howard Lopes Ribeiro Junior. 2022. "Role of Sirtuins in the Pathobiology of Onco-Hematological Diseases: A PROSPERO-Registered Study and In Silico Analysis" Cancers 14, no. 19: 4611. https://doi.org/10.3390/cancers14194611
APA StyleGoes, J. V. C., Carvalho, L. G., de Oliveira, R. T. G., Melo, M. M. d. L., Novaes, L. A. C., Moreno, D. A., Gonçalves, P. G., Montefusco-Pereira, C. V., Pinheiro, R. F., & Ribeiro Junior, H. L. (2022). Role of Sirtuins in the Pathobiology of Onco-Hematological Diseases: A PROSPERO-Registered Study and In Silico Analysis. Cancers, 14(19), 4611. https://doi.org/10.3390/cancers14194611






