Extracellular Vesicle-Mediated Metastasis Suppressors NME1 and NME2 Modify Lipid Metabolism in Fibroblasts
Abstract
Simple Summary
Abstract
1. Introduction
2. Results
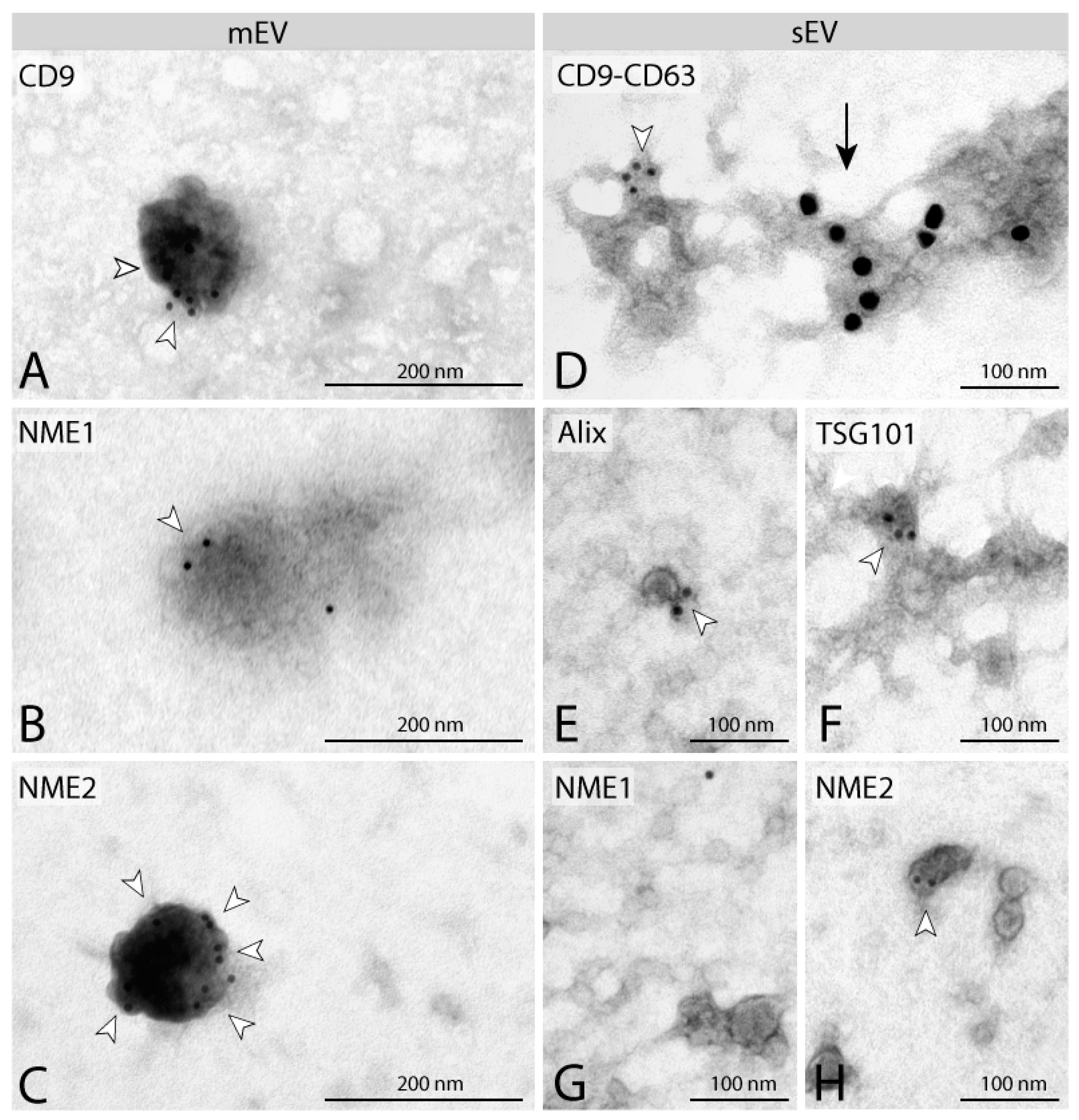
2.1. NME1 and NME2 Are Present in EVs Derived from NME1/NME2 Overexpressing Human Breast Carcinoma Cell Lines
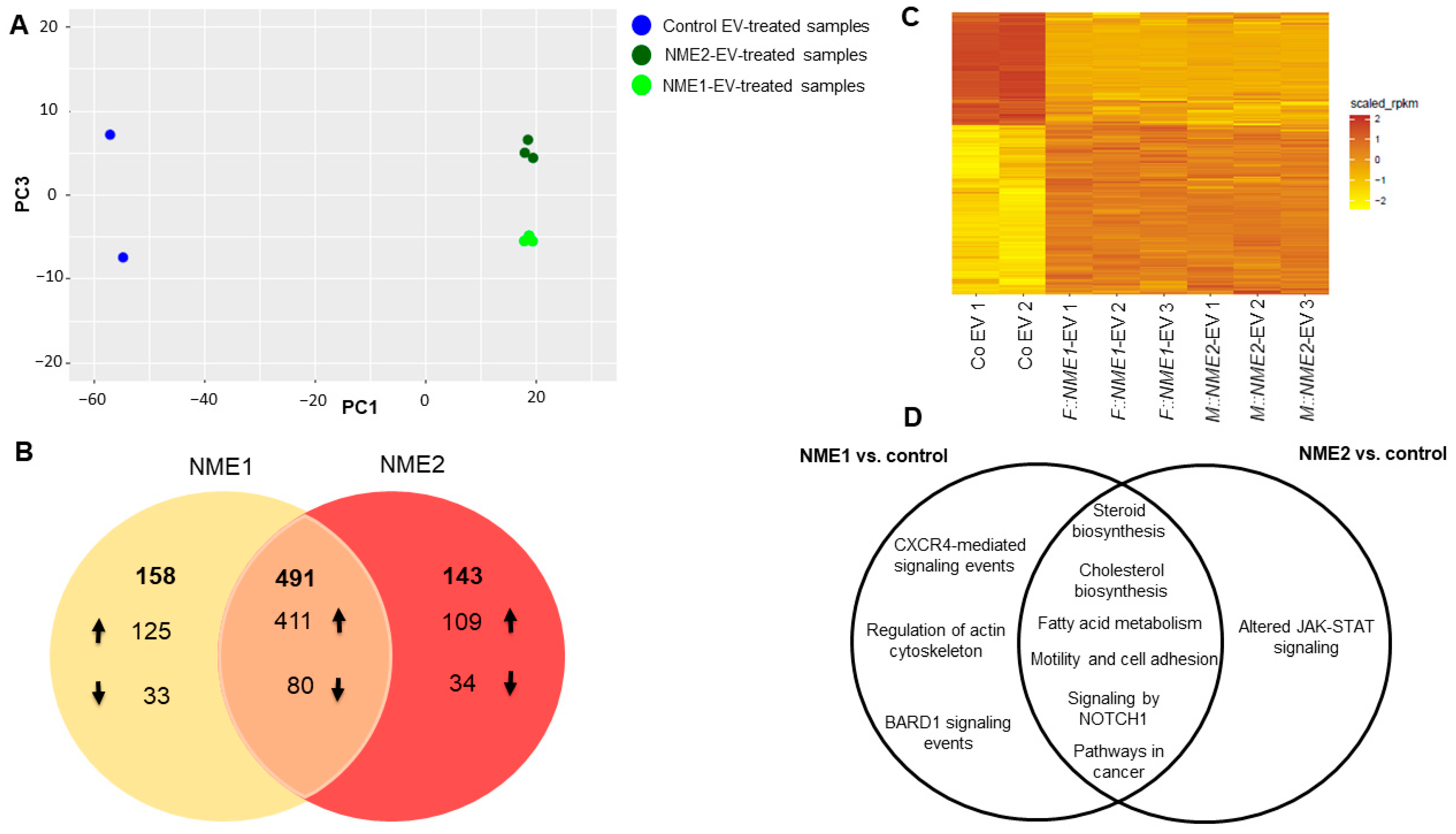
2.2. Analysing the Transcriptomic Effect of NME1/NME2-Containing Microvesicles and Exosomes Exerted on Fibroblasts
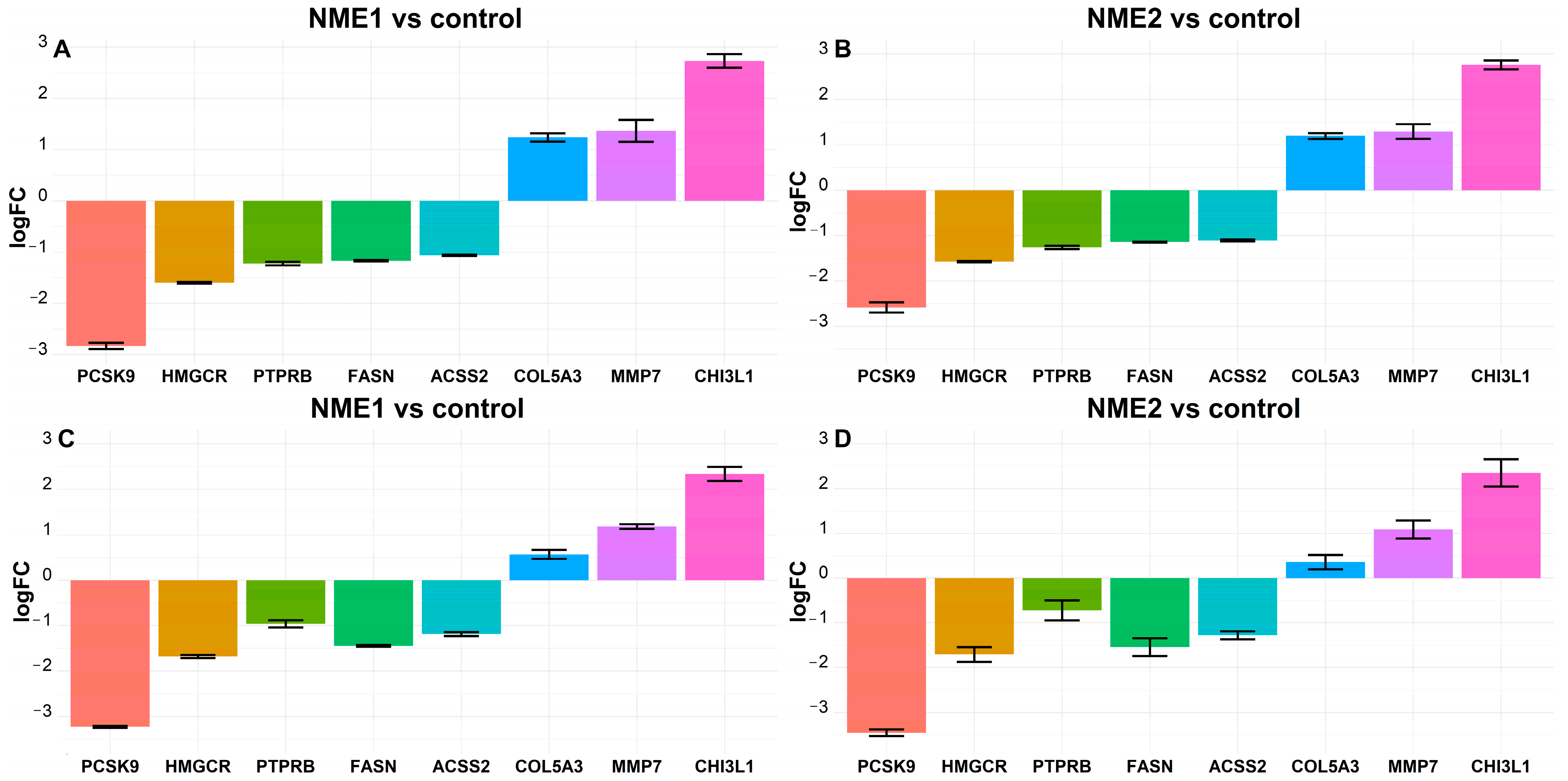
2.3. Fatty Acid Metabolism-Related Genes FASN and ACSS2 Are Downregulated in Fibroblasts Treated by NME1- and NME2-Containing EVs
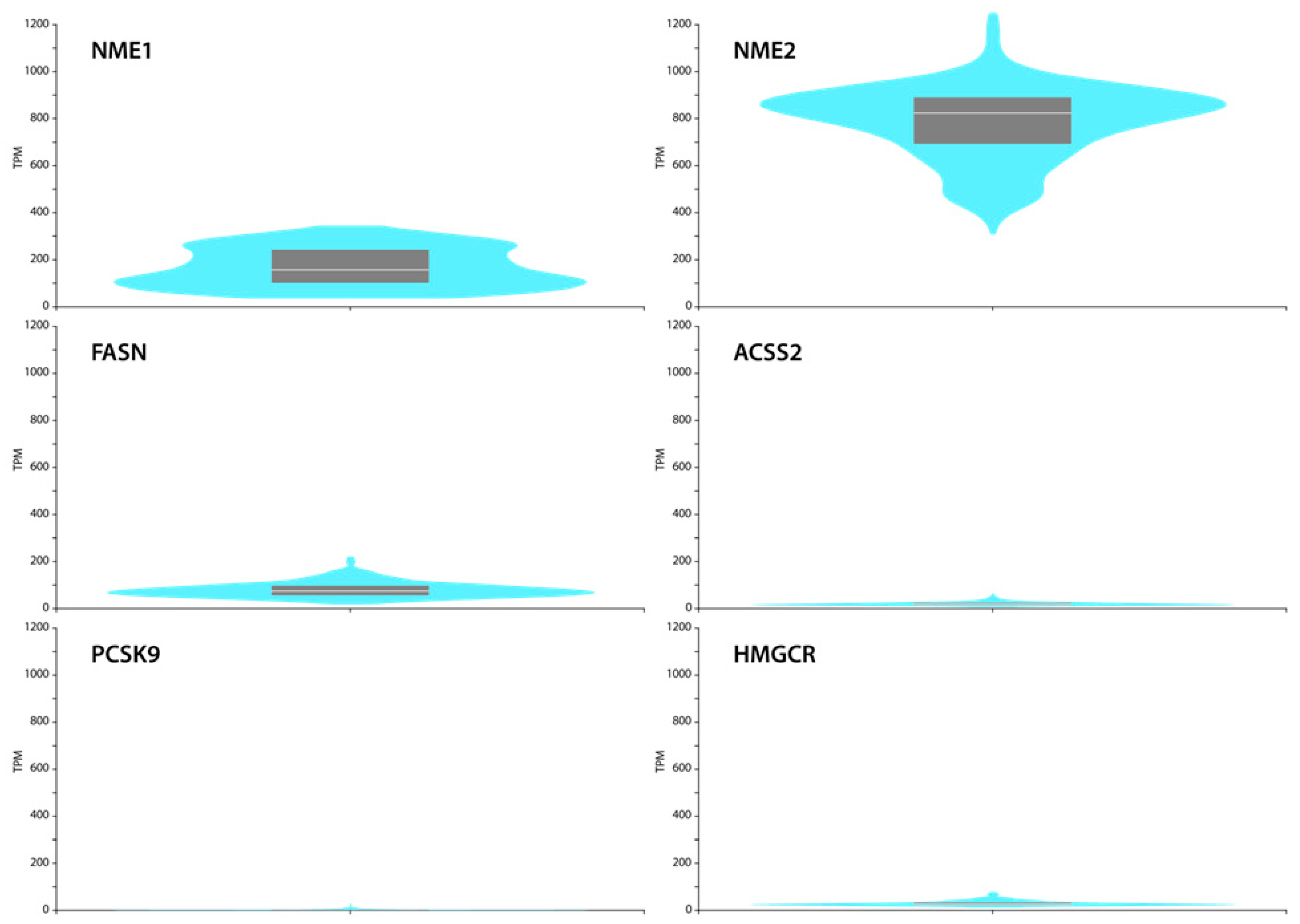
2.4. GTEx Portal Data Show Low FASN, ACSS2, PCSK9 and HMGCR Levels but Relatively High NME1 and NME2 Expression in Normal Skin Fibroblasts
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Cell Lines
5.2. Extracellular Vesicle Isolation from Conditioned Cell Culture Medium
5.3. Nanoparticle Tracking Analysis (NTA) of EVs
5.4. Detection of EVs Content by Immunoelectron Microscopy
5.5. EV Treatment of Fibroblasts and RNA Isolation
5.6. Transcriptome Analysis
5.6.1. Transcriptome Sequencing
5.6.2. Genome Sequence and Gene Annotations
5.6.3. Pre-Processing of RNAseq Data
5.6.4. Alignment of Obtained Reads to Genomes
5.6.5. Testing for DE Genes
5.6.6. Bioinformatic Analysis
5.7. qPCR Experiments
5.8. EV Treatment of Fibroblasts and Protein Isolation
5.9. Capillary Western Immunoassay (WESTM)
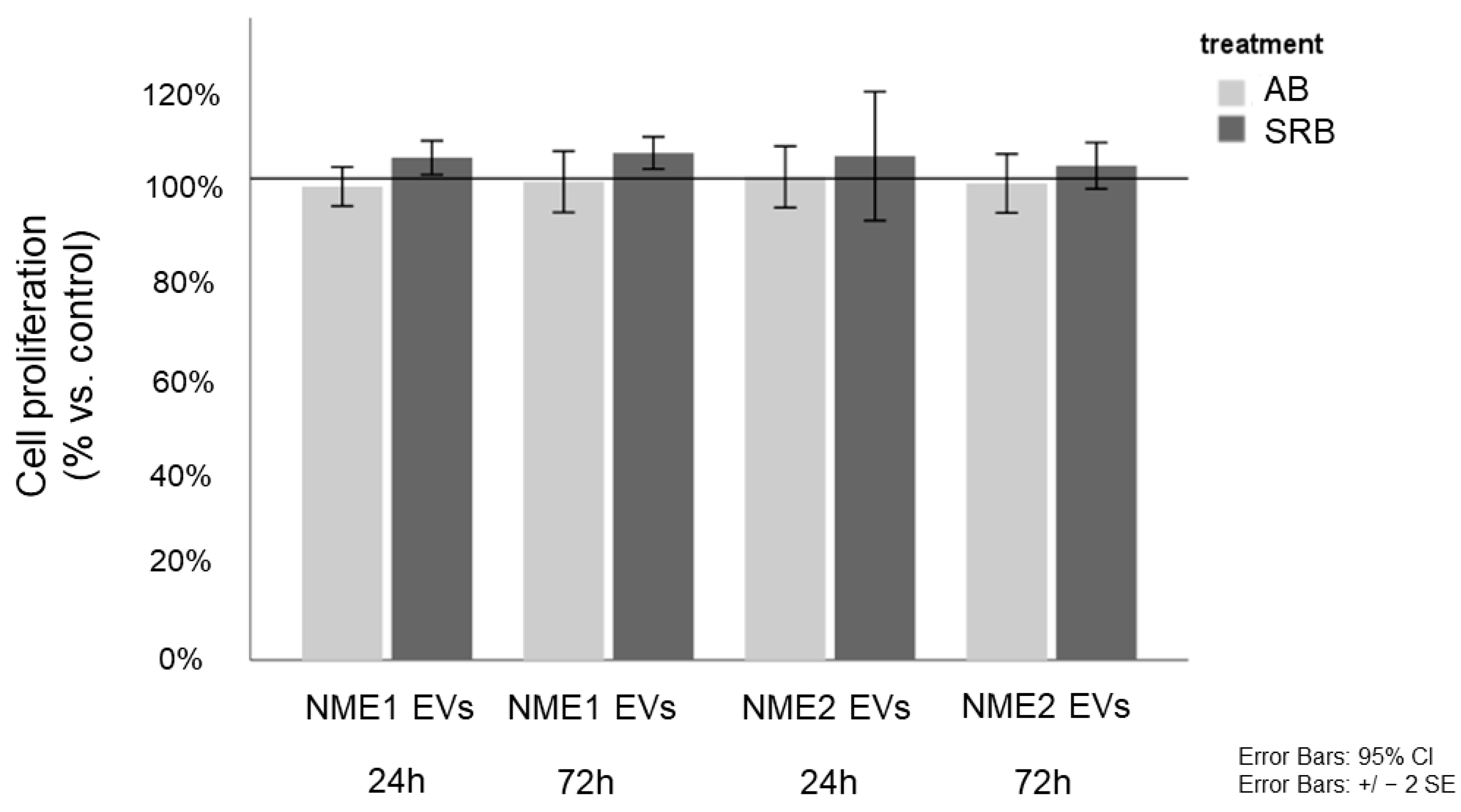
5.10. In Vitro Proliferation Assays—Alamar Blue and Sulforhodamine B Assays
5.11. Free Fatty Acid Measurements
5.12. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chaffer, C.L.; Weinberg, R.A. A Perspective on Cancer Cell Metastasis. Science 2011, 331, 1559–1564. [Google Scholar] [CrossRef] [PubMed]
- Rinker-Schaeffer, C.W.; O’Keefe, J.P.; Welch, D.R.; Theodorescu, D. Metastasis Suppressor Proteins: Discovery, Molecular Mechanisms, and Clinical Application. Clin. Cancer Res. 2006, 12, 3882–3889. [Google Scholar] [CrossRef] [PubMed]
- Steeg, P.S.; Bevilacqua, G.; Kopper, L.; Thorgeirsson, U.P.; Talmadge, J.E.; Liotta, L.A.; Sobep, M.E. Evidence for a Novel Gene Associated with Low Tumor Metastatic Potential. JNCI J. Natl. Cancer Inst. 1988, 80, 200–204. [Google Scholar] [CrossRef]
- Boissan, M.; Dabernat, S.; Peuchant, E.; Schlattner, U.; Lascu, I.; Lacombe, M.-L. The Mammalian Nm23/NDPK Family: From Metastasis Control to Cilia Movement. Mol. Cell. Biochem. 2009, 329, 51–62. [Google Scholar] [CrossRef] [PubMed]
- Desvignes, T.; Pontarotti, P.; Fauvel, C.; Bobe, J. Nme Protein Family Evolutionary History, a Vertebrate Perspective. BMC Evol. Biol. 2009, 9, 256. [Google Scholar] [CrossRef]
- Bevilacqua, G.; Sobel, M.E.; Liotta, L.A.; Steeg, P.S. Association of Low Nm23 RNA Levels in Human Primary Infiltrating Ductal Breast Carcinomas with Lymph Node Involvement and Other Histopathological Indicators of High Metastatic Potential. Cancer Res. 1989, 49, 5185–5190. [Google Scholar]
- Flørenes, V.A.; Aamdal, S.; Myklebost, O.; Maelandsmo, G.M.; Bruland, O.S.; Fodstad, O. Levels of Nm23 Messenger RNA in Metastatic Malignant Melanomas: Inverse Correlation to Disease Progression. Cancer Res. 1992, 52, 6088–6091. [Google Scholar]
- Xerri, L.; Grob, J.J.; Battyani, Z.; Gouvernet, J.; Hassoun, J.; Bonerandi, J.J. NM23 Expression in Metastasis of Malignant Melanoma Is a Predictive Prognostic Parameter Correlated with Survival. Br. J. Cancer 1994, 70, 1224–1228. [Google Scholar] [CrossRef][Green Version]
- Boissan, M.; Lacombe, M.-L. Nm23/NDP Kinases in Hepatocellular Carcinoma. J. Bioenerg. Biomembr. 2006, 38, 169–175. [Google Scholar] [CrossRef]
- An, R.; Meng, J.; Shi, Q.; Dai, X.-X.; Chen, J.-H.; Lei, Y.-J.; Shan, B.; Gao, C.; Chu, Y.-L.; Dong, X.-P. Expressions of Nucleoside Diphosphate Kinase (Nm23) in Tumor Tissues Are Related with Metastasis and Length of Survival of Patients with Hepatocellular Carcinoma. Biomed. Environ. Sci. 2010, 23, 267–272. [Google Scholar] [CrossRef]
- Hartsough, M.T.; Steeg, P.S. Nm23/Nucleoside Diphosphate Kinase in Human Cancers. J. Bioenerg. Biomembr. 2000, 32, 301–308. [Google Scholar] [CrossRef]
- Lacombe, M.; Boissan, M. NME1 (NME/NM23 Nucleoside Diphosphate Kinase 1). Atlas Genet. Cytogenet. Oncol. Haematol. 2013, 17, 526–538. [Google Scholar] [CrossRef][Green Version]
- Liu, L.; Li, M.; Zhang, C.; Zhang, J.; Li, G.; Zhang, Z.; He, X.; Fan, M. Prognostic Value and Clinicopathologic Significance of Nm23 in Various Cancers: A Systematic Review and Meta-Analysis. Int. J. Surg. 2018, 60, 257–265. [Google Scholar] [CrossRef]
- Leonard, M.K.; McCorkle, J.R.; Snyder, D.E.; Novak, M.; Zhang, Q.; Shetty, A.C.; Mahurkar, A.A.; Kaetzel, D.M. Identification of a Gene Expression Signature Associated with the Metastasis Suppressor Function of NME1: Prognostic Value in Human Melanoma. Lab. Investig. 2018, 98, 327–338. [Google Scholar] [CrossRef]
- Howlett, A.R.; Petersen, O.W.; Steeg, P.S.; Bissell, M.J. A Novel Function for the Nm23-H1 Gene: Overexpression in Human Breast Carcinoma Cells Leads to the Formation of Basement Membrane and Growth Arrest. J. Natl. Cancer Inst. 1994, 86, 1838–1844. [Google Scholar] [CrossRef] [PubMed]
- Harłozińska, A.; Bar, J.K.; Gerber, J. Nm23 Expression in Tissue Sections and Tumor Effusion Cells of Ovarian Neoplasms. Int. J. Cancer 1996, 69, 415–419. [Google Scholar] [CrossRef]
- Niitsu, N.; Honma, Y.; Lijima, K.; Takagi, T.; Higashihara, M.; Sawada, U.; Okabe-Kado, J. Clinical Significance of Nm23-H1 Proteins Expressed on Cell Surface in Non-Hodgkin’s Lymphoma. Leukemia 2003, 17, 196–202. [Google Scholar] [CrossRef] [PubMed]
- Andolfo, I.; De Martino, D.; Liguori, L.; Petrosino, G.; Troncone, G.; Tata, N.; Galasso, A.; Roma, C.; Chiancone, F.; Zarrilli, S.; et al. Correlation of NM23-H1 Cytoplasmic Expression with Metastatic Stage in Human Prostate Cancer Tissue. Naunyn. Schmiedebergs. Arch. Pharmacol. 2011, 384, 489–498. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chang, C.L.; Zhu, X.X.; Thoraval, D.H.; Ungar, D.; Rawwas, J.; Hora, N.; Strahler, J.R.; Hanash, S.M.; Radany, E. Nm23-H1 Mutation in Neuroblastoma. Nature 1994, 370, 335–336. [Google Scholar] [CrossRef]
- Garcia, I.; Mayol, G.; Rios, J.; Domenech, G.; Cheung, N.-K.V.; Oberthuer, A.; Fischer, M.; Maris, J.M.; Brodeur, G.M.; Hero, B.; et al. A Three-Gene Expression Signature Model for Risk Stratification of Patients with Neuroblastoma. Clin. Cancer Res. 2012, 18, 2012–2023. [Google Scholar] [CrossRef] [PubMed]
- Ouatas, T.; Salerno, M.; Palmieri, D.; Steeg, P.S. Basic and Translational Advances in Cancer Metastasis: Nm23. J. Bioenerg. Biomembr. 2003, 35, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Salerno, M.; Ouatas, T.; Palmieri, D.; Steeg, P.S. Inhibition of Signal Transduction by the Nm23 Metastasis Suppressor: Possible Mechanisms. Clin. Exp. Metastasis 2003, 20, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Thakur, R.K.; Yadav, V.K.; Kumar, P.; Chowdhury, S. Mechanisms of Non-Metastatic 2 (NME2)-Mediated Control of Metastasis across Tumor Types. Naunyn. Schmiedebergs. Arch. Pharmacol. 2011, 384, 397–406. [Google Scholar] [CrossRef]
- Niitsu, N.; Okabe-Kado, J.; Nakayama, M.; Wakimoto, N.; Sakashita, A.; Maseki, N.; Motoyoshi, K.; Umeda, M.; Honma, Y. Plasma Levels of the Differentiation Inhibitory Factor Nm23-H1 Protein and Their Clinical Implications in Acute Myelogenous Leukemia. Blood 2000, 96, 1080–1086. [Google Scholar] [CrossRef] [PubMed]
- Niitsu, N.; Okabe-Kado, J.; Okamoto, M.; Takagi, T.; Yoshida, T.; Aoki, S.; Hirano, M.; Honma, Y. Serum Nm23-H1 Protein as a Prognostic Factor in Aggressive Non-Hodgkin Lymphoma. Blood 2001, 97, 1202–1210. [Google Scholar] [CrossRef]
- Okabe-Kado, J.; Kasukabe, T.; Honma, Y.; Hanada, R.; Nakagawara, A.; Kaneko, Y. Clinical Significance of Serum NM23-H1 Protein in Neuroblastoma. Cancer Sci. 2005, 96, 653–660. [Google Scholar] [CrossRef] [PubMed]
- Yokdang, N.; Nordmeier, S.; Speirs, K.; Burkin, H.R.; Buxton, I.L.O. Blockade of Extracellular NM23 or Its Endothelial Target Slows Breast Cancer Growth and Metastasis. Integr. Cancer Sci. Ther. 2015, 2, 192–200. [Google Scholar] [CrossRef] [PubMed]
- Bunce, C.M.; Khanim, F.L. The “Known-Knowns”, and “Known-Unknowns” of Extracellular Nm23-H1/NDPK Proteins. Lab. Investig. 2018, 98, 602–608. [Google Scholar] [CrossRef]
- Álvarez-Chaver, P.; Rodríguez-Piñeiro, A.M.; Rodríguez-Berrocal, F.J.; García-Lorenzo, A.; Páez de la Cadena, M.; Martínez-Zorzano, V.S. Selection of Putative Colorectal Cancer Markers by Applying PCA on the Soluble Proteome of Tumors: NDK A as a Promising Candidate. J. Proteom. 2011, 74, 874–886. [Google Scholar] [CrossRef]
- Okabe-Kado, J.; Kasukabe, T.; Honma, Y.; Hayashi, M.; Henzel, W.J.; Hozumi, M. Identity of a Differentiation Inhibiting Factor for Mouse Myeloid Leukemia Cells with NM23/Nucleoside Diphosphate Kinase. Biochem. Biophys. Res. Commun. 1992, 182, 987–994. [Google Scholar] [CrossRef]
- Anzinger, J.; Malmquist, N.A.; Gould, J.; Buxton, I.L.O. Secretion of a Nucleoside Diphosphate Kinase (Nm23-H2) by Cells from Human Breast, Colon, Pancreas and Lung Tumors. Proc. West. Pharmacol. Soc. 2001, 44, 61–63. [Google Scholar] [PubMed]
- Romani, P.; Ignesti, M.; Gargiulo, G.; Hsu, T.; Cavaliere, V. Extracellular NME Proteins: A Player or a Bystander? Lab. Investig. 2018, 98, 248–257. [Google Scholar] [CrossRef] [PubMed]
- Burnstock, G. A Basis for Distinguishing Two Types of Purinergic Receptor; Raven Press: New York, NY, USA, 1978; p. 556. [Google Scholar]
- Burnstock, G.; Kennedy, C. Is There a Basis for Distinguishing Two Types of P2-Purinoceptor? Gen. Pharmacol. 1985, 16, 433–440. [Google Scholar] [CrossRef]
- Abbracchio, M.P.; Burnstock, G. Purinoceptors: Are There Families of P2X and P2Y Purinoceptors? Pharmacol. Ther. 1994, 64, 445–475. [Google Scholar] [CrossRef]
- Rumjahn, S.M.; Javed, M.A.; Wong, N.; Law, W.E.; Buxton, I.L.O. Purinergic Regulation of Angiogenesis by Human Breast Carcinoma-Secreted Nucleoside Diphosphate Kinase. Br. J. Cancer 2007, 97, 1372–1380. [Google Scholar] [CrossRef]
- Buxton, I.L.O.; Rumjahn, S.M.; Yokdang, N.; Baldwin, K.A.; Thai, J. Purinergic Regulation of Vascular Endothelial Growth Factor Signaling in Angiogenesis. Br. J. Cancer 2009, 100, 1465–1470. [Google Scholar] [CrossRef]
- Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef]
- Dai, J.; Su, Y.; Zhong, S.; Cong, L.; Liu, B.; Yang, J.; Tao, Y.; He, Z.; Chen, C.; Jiang, Y. Exosomes: Key Players in Cancer and Potential Therapeutic Strategy. Signal Transduct. Target. Ther. 2020, 5, 145. [Google Scholar] [CrossRef]
- Ćetković, H.; Bosnar, M.H.; Perina, D.; Mikoč, A.; Deželjin, M.; Belužić, R.; Bilandžija, H.; Ruiz-Trillo, I.; Harcet, M. Characterization of a Group I Nme Protein of Capsaspora Owczarzaki-a Close Unicellular Relative of Animals. Lab. Investig. 2018, 98, 304–314. [Google Scholar] [CrossRef]
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K.; et al. Minimal Information for Studies of Extracellular Vesicles 2018 (MISEV2018): A Position Statement of the International Society for Extracellular Vesicles and Update of the MISEV2014 Guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef]
- Escola, J.M.; Kleijmeer, M.J.; Stoorvogel, W.; Griffith, J.M.; Yoshie, O.; Geuze, H.J. Selective Enrichment of Tetraspan Proteins on the Internal Vesicles of Multivesicular Endosomes and on Exosomes Secreted by Human B-Lymphocytes. J. Biol. Chem. 1998, 273, 20121–20127. [Google Scholar] [CrossRef] [PubMed]
- Simons, M.; Raposo, G. Exosomes—Vesicular Carriers for Intercellular Communication. Curr. Opin. Cell Biol. 2009, 21, 575–581. [Google Scholar] [CrossRef] [PubMed]
- van der Pol, E.; Böing, A.N.; Harrison, P.; Sturk, A.; Nieuwland, R. Classification, Functions, and Clinical Relevance of Extracellular Vesicles. Pharmacol. Rev. 2012, 64, 676–705. [Google Scholar] [CrossRef] [PubMed]
- Boissan, M.; Wendum, D.; Arnaud-Dabernat, S.; Munier, A.; Debray, M.; Lascu, I.; Daniel, J.-Y.; Lacombe, M.-L. Increased Lung Metastasis in Transgenic NM23-Null/SV40 Mice with Hepatocellular Carcinoma. J. Natl. Cancer Inst. 2005, 97, 836–845. [Google Scholar] [CrossRef]
- Fhu, C.W.; Ali, A. Fatty Acid Synthase: An Emerging Target in Cancer. Molecules 2020, 25, 3935. [Google Scholar] [CrossRef]
- Huang, Z.; Zhang, M.; Plec, A.A.; Estill, S.J.; Cai, L.; Repa, J.J.; McKnight, S.L.; Tu, B.P. ACSS2 Promotes Systemic Fat Storage and Utilization through Selective Regulation of Genes Involved in Lipid Metabolism. Proc. Natl. Acad. Sci. USA 2018, 115, E9499–E9506. [Google Scholar] [CrossRef]
- Li, C.-J.; Chiu, Y.-H.; Chang, C.; Chang, Y.-C.I.; Sheu, J.J.-C.; Chiang, A.-J. Acetyl Coenzyme A Synthase 2 Acts as a Prognostic Biomarker Associated with Immune Infiltration in Cervical Squamous Cell Carcinoma. Cancers 2021, 13, 3125. [Google Scholar] [CrossRef]
- Atanasova, K.; Lee, J.; Roberts, J.; Lee, K.; Ojcius, D.M.; Yilmaz, Ö. Nucleoside-Diphosphate-Kinase of P. Gingivalis Is Secreted from Epithelial Cells in the Absence of a Leader Sequence through a Pannexin-1 Interactome. Sci. Rep. 2016, 6, 37643. [Google Scholar] [CrossRef]
- Maisano, D.; Mimmi, S.; Dattilo, V.; Marino, F.; Gentile, M.; Vecchio, E.; Fiume, G.; Nisticò, N.; Aloisio, A.; de Santo, M.P.; et al. A Novel Phage Display Based Platform for Exosome Diversity Characterization. Nanoscale 2022, 14, 2998–3003. [Google Scholar] [CrossRef]
- Liu, Q.; Zhang, X.; Zhang, J. Exosome-Based Nanoplatforms: The Emerging Tools for Breast Cancer Therapy. Front. Oncol. 2022, 12, 898605. [Google Scholar] [CrossRef]
- Zhang, S.; Nelson, O.D.; Price, I.R.; Zhu, C.; Lu, X.; Fernandez, I.R.; Weiss, R.S.; Lin, H. Long-Chain Fatty Acyl Coenzyme A Inhibits NME1/2 and Regulates Cancer Metastasis. Proc. Natl. Acad. Sci. USA 2022, 119, e2117013119. [Google Scholar] [CrossRef] [PubMed]
- Asif, P.J.; Longobardi, C.; Hahne, M.; Medema, J.P. The Role of Cancer-Associated Fibroblasts in Cancer Invasion and Metastasis. Cancers 2021, 13, 4720. [Google Scholar] [CrossRef] [PubMed]
- Sebestyén, A.; Dankó, T.; Sztankovics, D.; Moldvai, D.; Raffay, R.; Cervi, C.; Krencz, I.; Zsiros, V.; Jeney, A.; Petővári, G. The Role of Metabolic Ecosystem in Cancer Progression—Metabolic Plasticity and MTOR Hyperactivity in Tumor Tissues; Springer: New York, NY, USA, 2021; Volume 40, ISBN 0123456789. [Google Scholar]
- Yang, L.; Achreja, A.; Yeung, T.-L.; Mangala, L.S.; Jiang, D.; Han, C.; Baddour, J.; Marini, J.C.; Ni, J.; Nakahara, R.; et al. Targeting Stromal Glutamine Synthetase in Tumors Disrupts Tumor Microenvironment-Regulated Cancer Cell Growth. Cell Metab. 2016, 24, 685–700. [Google Scholar] [CrossRef] [PubMed]
- Gong, J.; Lin, Y.; Zhang, H.; Liu, C.; Cheng, Z.; Yang, X.; Zhang, J.; Xiao, Y.; Sang, N.; Qian, X.; et al. Reprogramming of Lipid Metabolism in Cancer-Associated Fibroblasts Potentiates Migration of Colorectal Cancer Cells. Cell Death Dis. 2020, 11, 267. [Google Scholar] [CrossRef] [PubMed]
- Butler, L.M.; Perone, Y.; Dehairs, J.; Lupien, L.E.; de Laat, V.; Talebi, A.; Loda, M.; Kinlaw, W.B.; Swinnen, J.V. Lipids and Cancer: Emerging Roles in Pathogenesis, Diagnosis and Therapeutic Intervention. Adv. Drug Deliv. Rev. 2020, 159, 245–293. [Google Scholar] [CrossRef]
- Zhang, Y.; Kurupati, R.; Liu, L.; Zhou, X.Y.; Zhang, G.; Hudaihed, A.; Filisio, F.; Giles-Davis, W.; Xu, X.; Karakousis, G.C.; et al. Enhancing CD8+ T Cell Fatty Acid Catabolism within a Metabolically Challenging Tumor Microenvironment Increases the Efficacy of Melanoma Immunotherapy. Cancer Cell 2017, 32, 377–391.e9. [Google Scholar] [CrossRef]
- Manzo, T.; Prentice, B.M.; Anderson, K.G.; Raman, A.; Schalck, A.; Codreanu, G.S.; Nava Lauson, C.B.; Tiberti, S.; Raimondi, A.; Jones, M.A.; et al. Accumulation of Long-Chain Fatty Acids in the Tumor Microenvironment Drives Dysfunction in Intrapancreatic Cd8+ t Cells. J. Exp. Med. 2020, 217, e20191920. [Google Scholar] [CrossRef]
- Prendeville, H.; Lynch, L. Diet, Lipids, and Antitumor Immunity. Cell. Mol. Immunol. 2022, 19, 432–444. [Google Scholar] [CrossRef]
- Lopes-Coelho, F.; André, S.; Félix, A.; Serpa, J. Breast Cancer Metabolic Cross-Talk: Fibroblasts Are Hubs and Breast Cancer Cells Are Gatherers of Lipids. Mol. Cell. Endocrinol. 2018, 462, 93–106. [Google Scholar] [CrossRef]
- Petővári, G.; Dankó, T.; Tőkés, A.-M.; Vetlényi, E.; Krencz, I.; Raffay, R.; Hajdu, M.; Sztankovics, D.; Németh, K.; Vellai-Takács, K.; et al. In Situ Metabolic Characterisation of Breast Cancer and Its Potential Impact on Therapy. Cancers 2020, 12, 2492. [Google Scholar] [CrossRef]
- Antalis, C.J.; Uchida, A.; Buhman, K.K.; Siddiqui, R.A. Migration of MDA-MB-231 Breast Cancer Cells Depends on the Availability of Exogenous Lipids and Cholesterol Esterification. Clin. Exp. Metastasis 2011, 28, 733–741. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt Removes Adapter Sequences from High-Throughput Sequencing Reads. EMBnet. J. 2011, 17, 10. [Google Scholar] [CrossRef]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate Alignment of Transcriptomes in the Presence of Insertions, Deletions and Gene Fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and Memory-Efficient Alignment of Short DNA Sequences to the Human Genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Smyth, G.K.; Shi, W. FeatureCounts: An Efficient General Purpose Program for Assigning Sequence Reads to Genomic Features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. EdgeR: A Bioconductor Package for Differential Expression Analysis of Digital Gene Expression Data. Bioinformatics 2009, 26, 139–140. [Google Scholar] [CrossRef]
- McCarthy, D.J.; Chen, Y.; Smyth, G.K. Differential Expression Analysis of Multifactor RNA-Seq Experiments with Respect to Biological Variation. Nucleic Acids Res. 2012, 40, 4288–4297. [Google Scholar] [CrossRef]
- Chen, J.; Bardes, E.E.; Aronow, B.J.; Jegga, A.G. ToppGene Suite for Gene List Enrichment Analysis and Candidate Gene Prioritization. Nucleic Acids Res. 2009, 37, W305–W311. [Google Scholar] [CrossRef]


| Primary Antibody | WES | Electron Microscopy | Size (kDa) | ||
|---|---|---|---|---|---|
| Cat. No. | Dilution | Cat. No. | Dilution | ||
| NME1 | OriGene #TA801264 | 1:50 | OriGene #TA801264 | 1:100 | 17 |
| NME2 | [45] | 1:50 | [45] | 1:100 | 17 |
| FLAG (biotin) | Sigma #F9291 | 1:50 | - | - | 17 |
| MYC | Sigma #M4439 | 1:50 | - | - | 17, 62 |
| β-Actin | Sigma #A2228 | 1:100 | - | - | 42 |
| CD63 | Santa Cruz #sc-15363 | 1:50 | Abcam #ab134045 | 1:50 | 26/30, 60 |
| CD81 | Sigma #SAB3500454 | 1:50 | - | - | 26 |
| CD9 | - | - | Sigma #C9993 | 1:50 | 24 |
| Alix | Sigma #SAB4200477 | 1:50 | Abcam #ab186429 | 1:100 | 95 |
| TSG101 | Sigma #T5701 | 1:50 | Sigma #T5701 | 1:100 | 46 |
| FASN | Cell signaling CS #3180 | 1:50 | - | - | 241 |
| ACSS2 | Cell signaling CS #3658 | 1:50 | - | - | 79 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mátyási, B.; Petővári, G.; Dankó, T.; Butz, H.; Likó, I.; Lőw, P.; Petit, I.; Bittar, R.; Bonnefont-Rousselot, D.; Farkas, Z.; et al. Extracellular Vesicle-Mediated Metastasis Suppressors NME1 and NME2 Modify Lipid Metabolism in Fibroblasts. Cancers 2022, 14, 3913. https://doi.org/10.3390/cancers14163913
Mátyási B, Petővári G, Dankó T, Butz H, Likó I, Lőw P, Petit I, Bittar R, Bonnefont-Rousselot D, Farkas Z, et al. Extracellular Vesicle-Mediated Metastasis Suppressors NME1 and NME2 Modify Lipid Metabolism in Fibroblasts. Cancers. 2022; 14(16):3913. https://doi.org/10.3390/cancers14163913
Chicago/Turabian StyleMátyási, Barbara, Gábor Petővári, Titanilla Dankó, Henriett Butz, István Likó, Péter Lőw, Isabelle Petit, Randa Bittar, Dominique Bonnefont-Rousselot, Zsolt Farkas, and et al. 2022. "Extracellular Vesicle-Mediated Metastasis Suppressors NME1 and NME2 Modify Lipid Metabolism in Fibroblasts" Cancers 14, no. 16: 3913. https://doi.org/10.3390/cancers14163913
APA StyleMátyási, B., Petővári, G., Dankó, T., Butz, H., Likó, I., Lőw, P., Petit, I., Bittar, R., Bonnefont-Rousselot, D., Farkas, Z., Szeniczey, T., Molnár, K., Pálóczi, K., Buzás, E. I., Boissan, M., Sebestyén, A., & Takács-Vellai, K. (2022). Extracellular Vesicle-Mediated Metastasis Suppressors NME1 and NME2 Modify Lipid Metabolism in Fibroblasts. Cancers, 14(16), 3913. https://doi.org/10.3390/cancers14163913








