MPscore: A Novel Predictive and Prognostic Scoring for Progressive Meningioma
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Preprocessing and Tissue Sample Validation
2.2. Consensus Clustering
2.3. Differentially Expressed Gene (DEG) Analysis
2.4. Stemness Index (SI) Prediction
2.5. Copy Number Alteration (CNA) and DNA Methylation Analysis
2.6. Immune Cell Infiltration Prediction
2.7. Fusion Genes Identification
2.8. Random Forest Model
2.9. Meningioma Progression Score (MPscore)
2.10. Statistical Analysis
3. Results
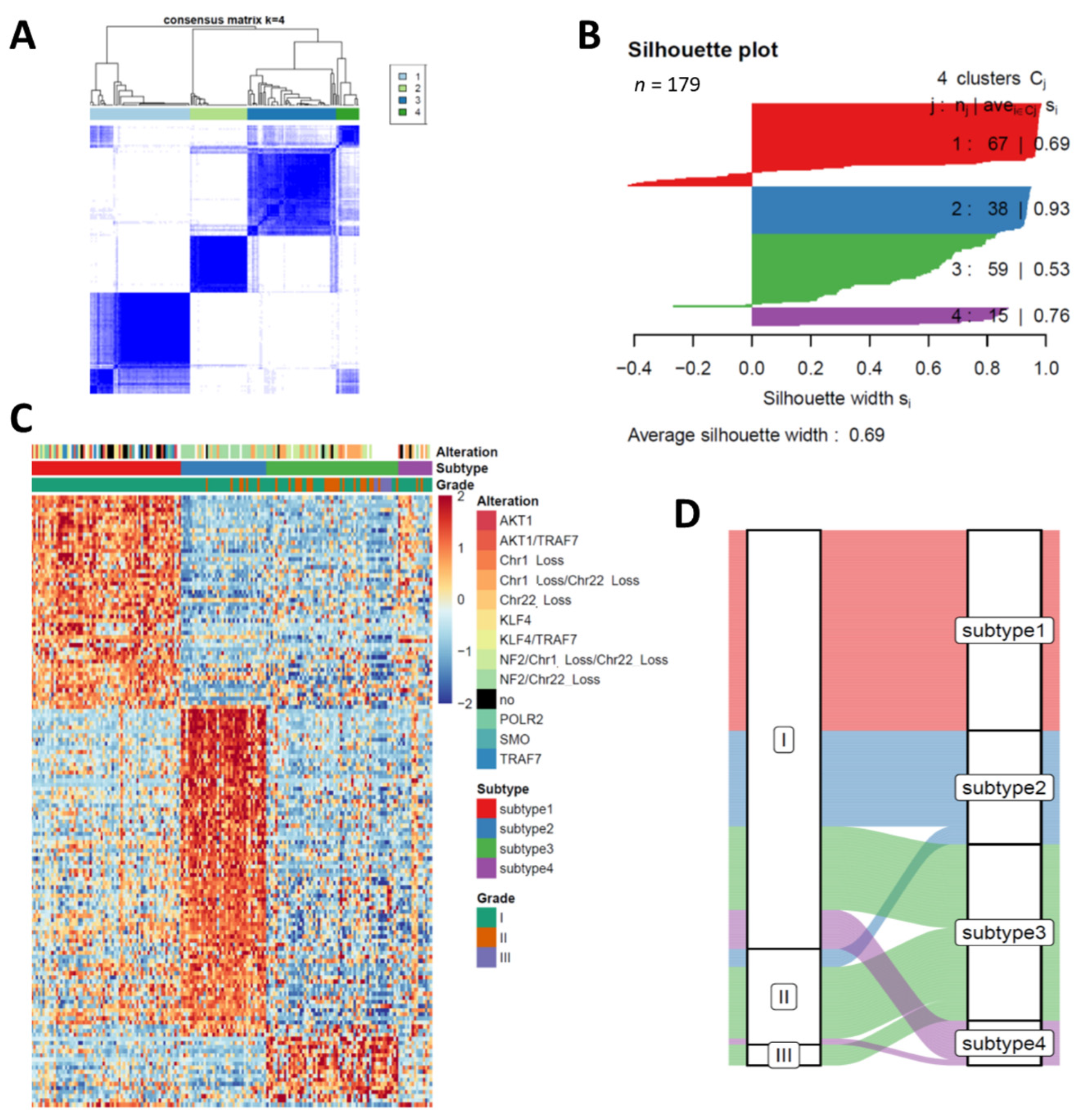
3.1. Transcriptome Profiling Unravels Four Distinct Subtypes in Meningiomas
3.2. Stemness Indexes Reveal the Different Progressive Potentials between Subtypes
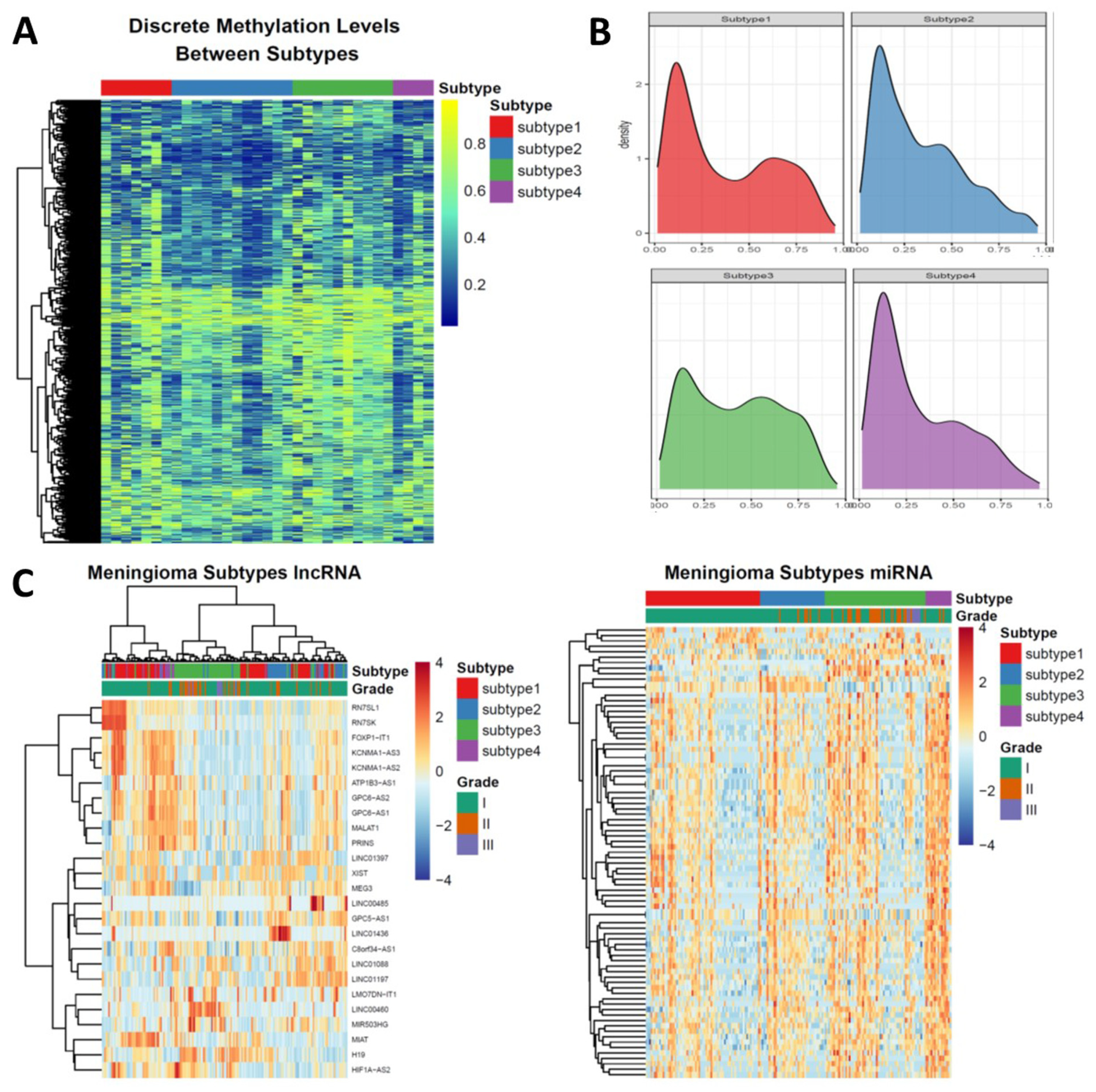
3.3. Epigenetic Alterations Recapitulate the Subtyping of Meningioma
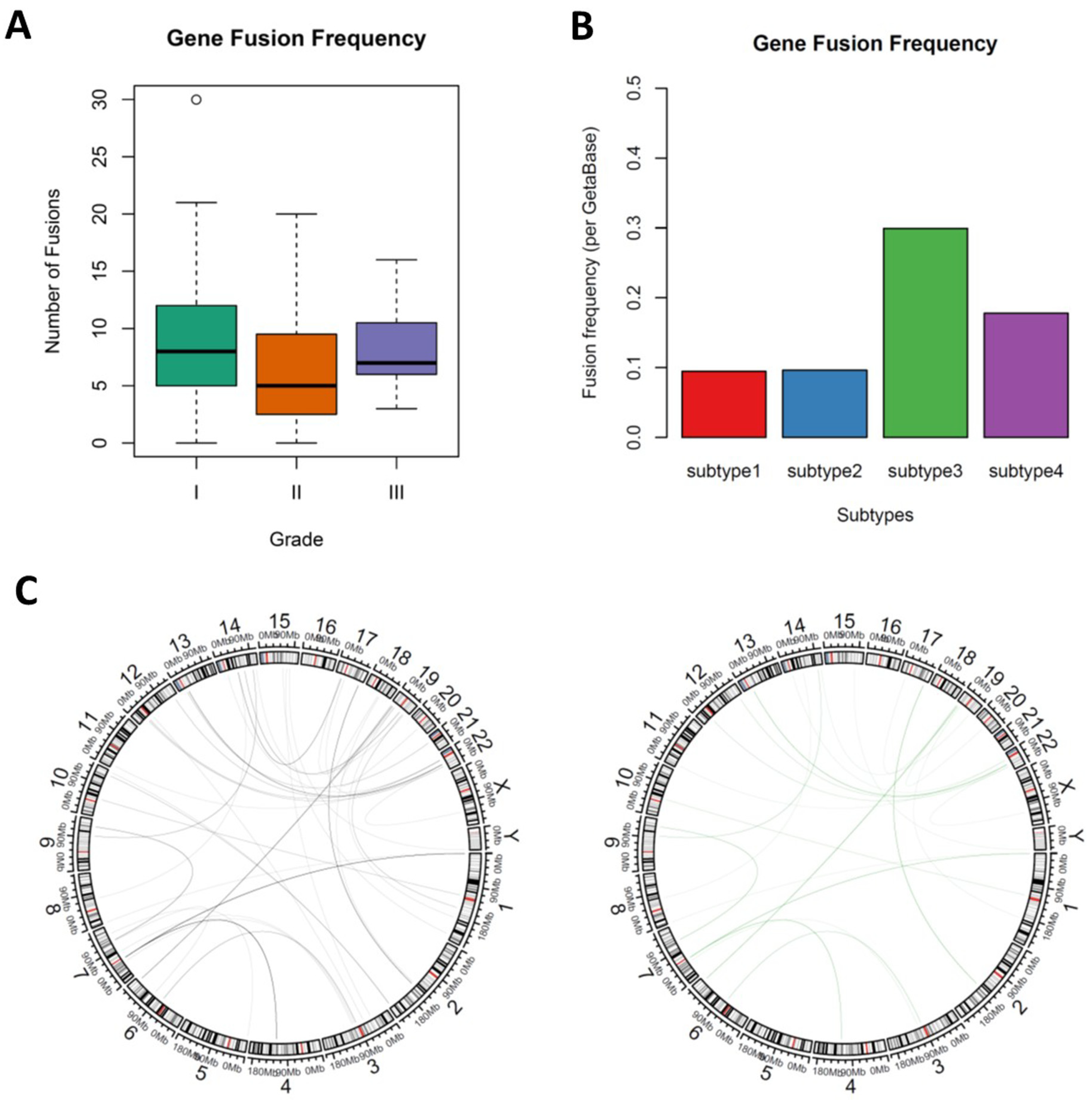
3.4. Novel Fusion Genes were Identified between Subtypes
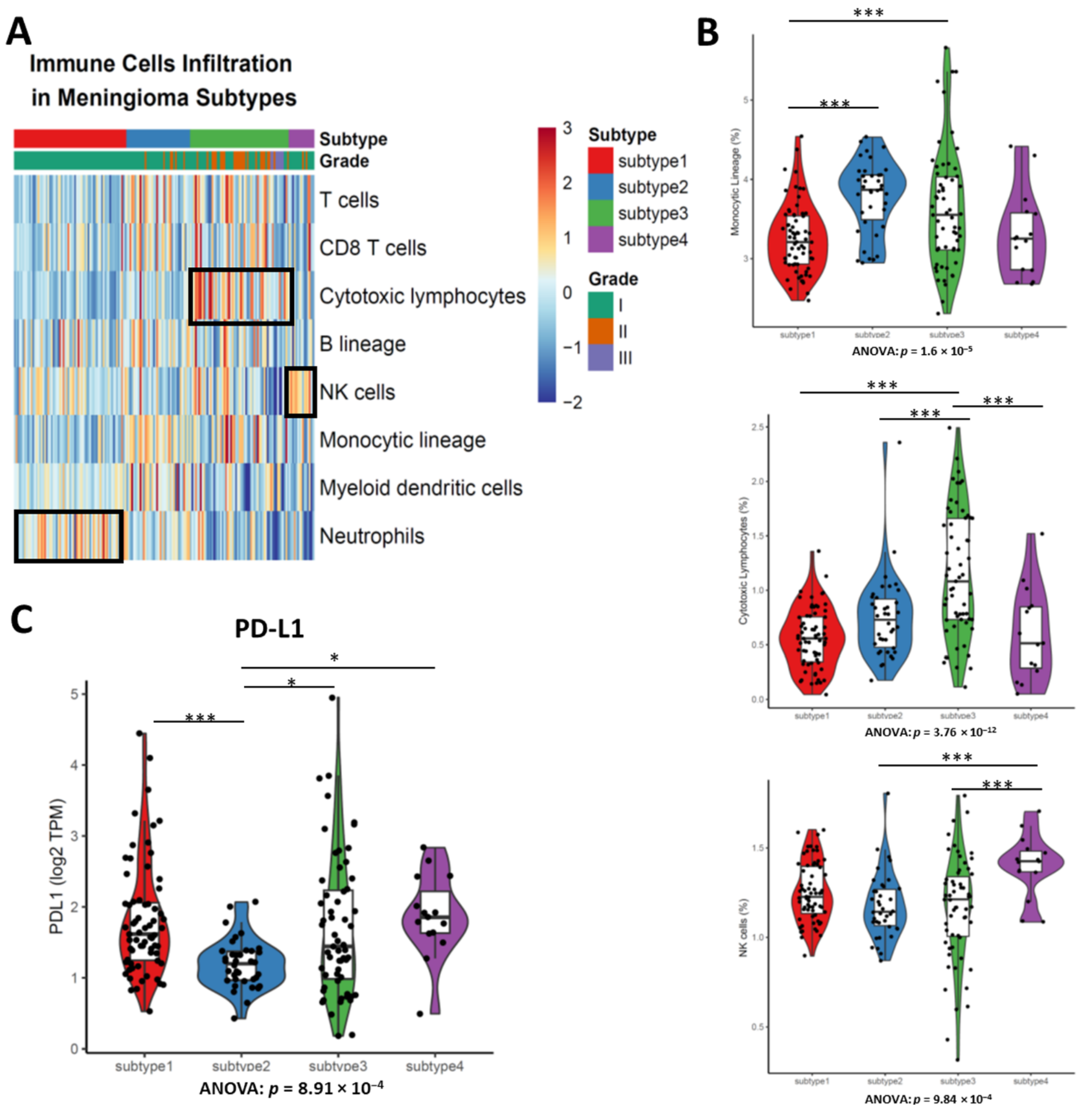
3.5. Different Types of Enriched Immune Cells between Subtypes Demonstrates the Disparate Level of Immune Cell Infiltration
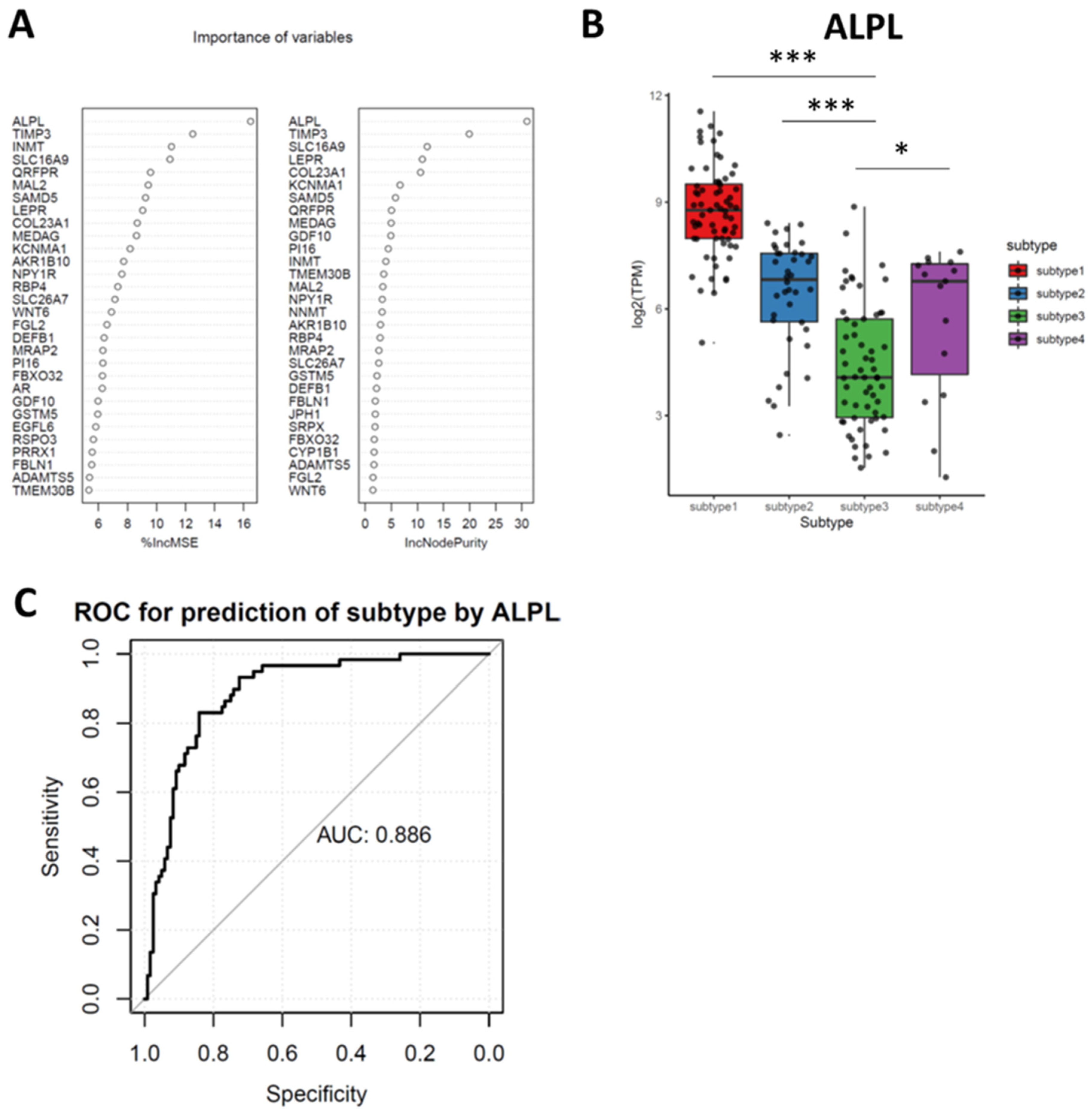
3.6. Random Forest (RF) Identifies Downregulated ALPL as the Feature Genes in Subtypes 3
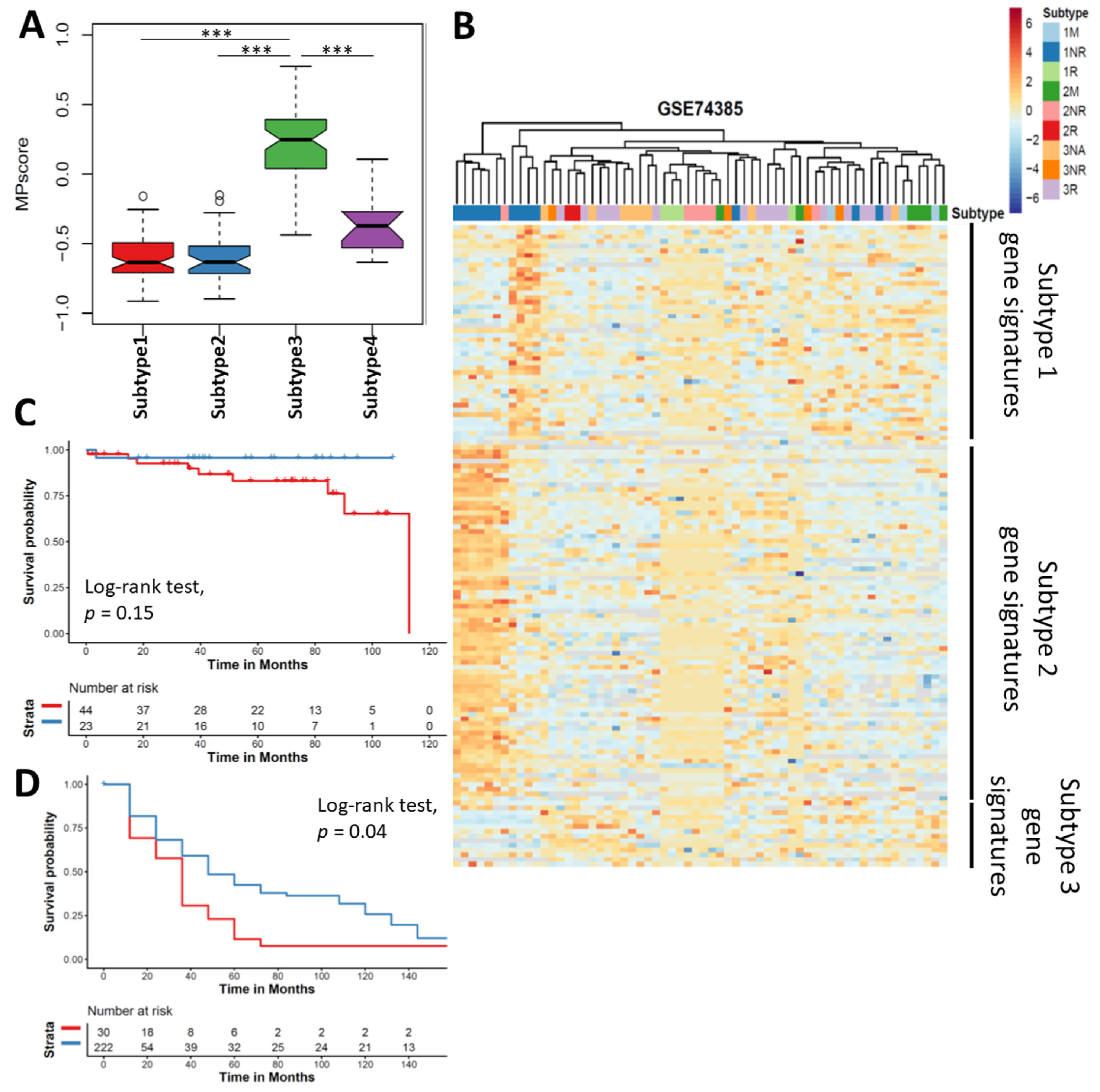
3.7. Meningioma Progression Score (MPscore) Discriminates the Progression of Meningioma
3.8. Small Molecules were Predicted to Target the Subtype 3 Meningioma
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ostrom, Q.T.; Cioffi, G.; Gittleman, H.; Patil, N.; Waite, K.; Kruchko, C.; Barnholtz-Sloan, J.S. CBTRUS Statistical Report: Primary Brain and Other Central Nervous System Tumors Diagnosed in the United States in 2012–2016. Neuro-Oncology 2019, 21, v1–v100. [Google Scholar] [CrossRef]
- Goldbrunner, R.; Minniti, G.; Preusser, M.; Jenkinson, M.D.; Sallabanda, K.; Houdart, E.; von Deimling, A.; Stavrinou, P.; Lefranc, F.; Lund-Johansen, M.; et al. EANO guidelines for the diagnosis and treatment of meningiomas. Lancet Oncol. 2016, 17, e383–e391. [Google Scholar] [CrossRef]
- Louis, D.N.; Perry, A.; Reifenberger, G.; von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef] [PubMed]
- Al-Rashed, M.; Foshay, K.; Abedalthagafi, M. Recent Advances in Meningioma Immunogenetics. Front. Oncol. 2019, 9, 1472. [Google Scholar] [CrossRef] [PubMed]
- Domingues, P.H.; Sousa, P.; Otero, A.; Goncalves, J.M.; Ruiz, L.; de Oliveira, C.; Lopes, M.C.; Orfao, A.; Tabernero, M.D. Proposal for a new risk stratification classification for meningioma based on patient age, WHO tumor grade, size, localization, and karyotype. Neuro-Oncology 2014, 16, 735–747. [Google Scholar] [CrossRef]
- Schmidt, M.; Mock, A.; Jungk, C.; Sahm, F.; Ull, A.T.; Warta, R.; Lamszus, K.; Gousias, K.; Ketter, R.; Roesch, S.; et al. Transcriptomic analysis of aggressive meningiomas identifies PTTG1 and LEPR as prognostic biomarkers independent of WHO grade. Oncotarget 2016, 7, 14551–14568. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Xiang, C.X.; Zhou, Y.; Ao, X.S.; Zhou, D.Q.; Peng, P.; Zhang, H.Q.; Liu, H.D.; Huang, X. Gene expression profile for predicting survival of patients with meningioma. Int. J. Oncol. 2015, 46, 791–797. [Google Scholar] [CrossRef] [PubMed]
- Olar, A.; Goodman, L.D.; Wani, K.M.; Boehling, N.S.; Sharma, D.S.; Mody, R.R.; Gumin, J.; Claus, E.B.; Lang, F.F.; Cloughesy, T.F.; et al. A gene expression signature predicts recurrence-free survival in meningioma. Oncotarget 2018, 9, 16087–16098. [Google Scholar] [CrossRef] [PubMed]
- Olar, A.; Wani, K.M.; Wilson, C.D.; Zadeh, G.; DeMonte, F.; Jones, D.T.; Pfister, S.M.; Sulman, E.P.; Aldape, K.D. Global epigenetic profiling identifies methylation subgroups associated with recurrence-free survival in meningioma. Acta Neuropathol. 2017, 133, 431–444. [Google Scholar] [CrossRef]
- Sahm, F.; Schrimpf, D.; Stichel, D.; Jones, D.T.W.; Hielscher, T.; Schefzyk, S.; Okonechnikov, K.; Koelsche, C.; Reuss, D.E.; Capper, D.; et al. DNA methylation-based classification and grading system for meningioma: A multicentre, retrospective analysis. Lancet Oncol. 2017, 18, 682–694. [Google Scholar] [CrossRef]
- Domingues, P.; Gonzalez-Tablas, M.; Otero, A.; Pascual, D.; Ruiz, L.; Miranda, D.; Sousa, P.; Goncalves, J.M.; Lopes, M.C.; Orfao, A.; et al. Genetic/molecular alterations of meningiomas and the signaling pathways targeted. Oncotarget 2015, 6, 10671–10688. [Google Scholar] [CrossRef] [PubMed]
- Verhaak, R.G.; Hoadley, K.A.; Purdom, E.; Wang, V.; Qi, Y.; Wilkerson, M.D.; Miller, C.R.; Ding, L.; Golub, T.; Mesirov, J.P.; et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 2010, 17, 98–110. [Google Scholar] [CrossRef] [PubMed]
- Taylor, M.D.; Northcott, P.A.; Korshunov, A.; Remke, M.; Cho, Y.J.; Clifford, S.C.; Eberhart, C.G.; Parsons, D.W.; Rutkowski, S.; Gajjar, A.; et al. Molecular subgroups of medulloblastoma: The current consensus. Acta Neuropathol. 2012, 123, 465–472. [Google Scholar] [CrossRef] [PubMed]
- Pajtler, K.W.; Witt, H.; Sill, M.; Jones, D.T.; Hovestadt, V.; Kratochwil, F.; Wani, K.; Tatevossian, R.; Punchihewa, C.; Johann, P.; et al. Molecular Classification of Ependymal Tumors across All CNS Compartments, Histopathological Grades, and Age Groups. Cancer Cell 2015, 27, 728–743. [Google Scholar] [CrossRef]
- Fang, L.; Lowther, D.E.; Meizlish, M.L.; Anderson, R.C.; Bruce, J.N.; Devine, L.; Huttner, A.J.; Kleinstein, S.H.; Lee, J.Y.; Stern, J.N.; et al. The immune cell infiltrate populating meningiomas is composed of mature, antigen-experienced T and B cells. Neuro-Oncology 2013, 15, 1479–1490. [Google Scholar] [CrossRef]
- Han, S.J.; Reis, G.; Kohanbash, G.; Shrivastav, S.; Magill, S.T.; Molinaro, A.M.; McDermott, M.W.; Theodosopoulos, P.V.; Aghi, M.K.; Berger, M.S.; et al. Expression and prognostic impact of immune modulatory molecule PD-L1 in meningioma. J. Neurooncol. 2016, 130, 543–552. [Google Scholar] [CrossRef]
- Proctor, D.T.; Ramachandran, S.; Lama, S.; Sutherland, G.R. Towards Molecular Classification of Meningioma: Evolving Treatment and Diagnostic Paradigms. World Neurosurg. 2018, 119, 366–373. [Google Scholar] [CrossRef]
- Clark, V.E.; Harmanci, A.S.; Bai, H.; Youngblood, M.W.; Lee, T.I.; Baranoski, J.F.; Ercan-Sencicek, A.G.; Abraham, B.J.; Weintraub, A.S.; Hnisz, D.; et al. Recurrent somatic mutations in POLR2A define a distinct subset of meningiomas. Nat. Genet. 2016, 48, 1253–1259. [Google Scholar] [CrossRef]
- Patel, A.J.; Wan, Y.W.; Al-Ouran, R.; Revelli, J.P.; Cardenas, M.F.; Oneissi, M.; Xi, L.; Jalali, A.; Magnotti, J.F.; Muzny, D.M.; et al. Molecular profiling predicts meningioma recurrence and reveals loss of DREAM complex repression in aggressive tumors. Proc. Natl. Acad. Sci. USA 2019, 116, 21715–21726. [Google Scholar] [CrossRef]
- Agnihotri, S.; Suppiah, S.; Tonge, P.D.; Jalali, S.; Danesh, A.; Bruce, J.P.; Mamatjan, Y.; Klironomos, G.; Gonen, L.; Au, K.; et al. Therapeutic radiation for childhood cancer drives structural aberrations of NF2 in meningiomas. Nat. Commun. 2017, 8, 186. [Google Scholar] [CrossRef]
- Malta, T.M.; Sokolov, A.; Gentles, A.J.; Burzykowski, T.; Poisson, L.; Weinstein, J.N.; Kaminska, B.; Huelsken, J.; Omberg, L.; Gevaert, O.; et al. Machine Learning Identifies Stemness Features Associated with Oncogenic Dedifferentiation. Cell 2018, 173, 338–354.e15. [Google Scholar] [CrossRef] [PubMed]
- Becht, E.; Giraldo, N.A.; Lacroix, L.; Buttard, B.; Elarouci, N.; Petitprez, F.; Selves, J.; Laurent-Puig, P.; Sautes-Fridman, C.; Fridman, W.H.; et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 2016, 17, 218. [Google Scholar] [CrossRef] [PubMed]
- Barbie, D.A.; Tamayo, P.; Boehm, J.S.; Kim, S.Y.; Moody, S.E.; Dunn, I.F.; Schinzel, A.C.; Sandy, P.; Meylan, E.; Scholl, C.; et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 2009, 462, 108–112. [Google Scholar] [CrossRef] [PubMed]
- Kamamoto, D.; Saga, I.; Ohara, K.; Yoshida, K.; Sasaki, H. Association Between CD133, CD44, and Nestin Expression and Prognostic Factors in High-Grade Meningioma. World Neurosurg. 2019, 124, e188–e196. [Google Scholar] [CrossRef]
- Shivapathasundram, G.; Wickremesekera, A.C.; Brasch, H.D.; Marsh, R.; Tan, S.T.; Itinteang, T. Expression of Embryonic Stem Cell Markers on the Microvessels of WHO Grade I Meningioma. Front. Surg. 2018, 5, 65. [Google Scholar] [CrossRef]
- Viaene, A.N.; Zhang, B.; Martinez-Lage, M.; Xiang, C.; Tosi, U.; Thawani, J.P.; Gungor, B.; Zhu, Y.; Roccograndi, L.; Zhang, L.; et al. Transcriptome signatures associated with meningioma progression. Acta Neuropathol. Commun. 2019, 7, 67. [Google Scholar] [CrossRef]
- Johnson, M.D. PD-L1 expression in meningiomas. J. Clin. Neurosci. 2018, 57, 149–151. [Google Scholar] [CrossRef]
- Vamathevan, J.; Clark, D.; Czodrowski, P.; Dunham, I.; Ferran, E.; Lee, G.; Li, B.; Madabhushi, A.; Shah, P.; Spitzer, M.; et al. Applications of machine learning in drug discovery and development. Nat. Rev. Drug Discov. 2019, 18, 463–477. [Google Scholar] [CrossRef]
- Huang, S.; Cai, N.; Pacheco, P.P.; Narrandes, S.; Wang, Y.; Xu, W. Applications of Support Vector Machine (SVM) Learning in Cancer Genomics. Cancer Genom. Proteom. 2018, 15, 41–51. [Google Scholar]
- Hulbert, A.; Jusue-Torres, I.; Stark, A.; Chen, C.; Rodgers, K.; Lee, B.; Griffin, C.; Yang, A.; Huang, P.; Wrangle, J.; et al. Early Detection of Lung Cancer Using DNA Promoter Hypermethylation in Plasma and Sputum. Clin Cancer Res. 2017, 23, 1998–2005. [Google Scholar] [CrossRef]
- Nordstrand, A.; Ylitalo, E.B.; Thysell, E.; Jernberg, E.; Crnalic, S.; Widmark, A.; Bergh, A.; Lerner, U.H.; Wikstrom, P. Bone Cell Activity in Clinical Prostate Cancer Bone Metastasis and Its Inverse Relation to Tumor Cell Androgen Receptor Activity. Int. J. Mol. Sci. 2018, 19, 1223. [Google Scholar] [CrossRef] [PubMed]
- Andreux, P.A.; Williams, E.G.; Koutnikova, H.; Houtkooper, R.H.; Champy, M.F.; Henry, H.; Schoonjans, K.; Williams, R.W.; Auwerx, J. Systems genetics of metabolism: The use of the BXD murine reference panel for multiscalar integration of traits. Cell 2012, 150, 1287–1299. [Google Scholar] [CrossRef]
- Bouvier, C.; Liprandi, A.; Colin, C.; Giorgi, R.; Quilichini, B.; Metellus, P.; Figarella-Branger, D. Lack of alkaline phosphatase activity predicts meningioma recurrence. Am. J. Clin. Pathol. 2005, 124, 252–258. [Google Scholar] [CrossRef][Green Version]
- Muller, P.; Henn, W.; Niedermayer, I.; Ketter, R.; Feiden, W.; Steudel, W.I.; Zang, K.D.; Steilen-Gimbel, H. Deletion of chromosome 1p and loss of expression of alkaline phosphatase indicate progression of meningiomas. Clin. Cancer Res. 1999, 5, 3569–3577. [Google Scholar]
- Alshabi, A.M.; Vastrad, B.; Shaikh, I.A.; Vastrad, C. Identification of Crucial Candidate Genes and Pathways in Glioblastoma Multiform by Bioinformatics Analysis. Biomolecules 2019, 9, 201. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, K.; Kho, D.H.; Yanagawa, T.; Harazono, Y.; Gao, X.; Hogan, V.; Raz, A. Galectin-3 inhibits osteoblast differentiation through notch signaling. Neoplasia 2014, 16, 939–949. [Google Scholar] [CrossRef]
- Wrobel, G.; Roerig, P.; Kokocinski, F.; Neben, K.; Hahn, M.; Reifenberger, G.; Lichter, P. Microarray-based gene expression profiling of benign, atypical and anaplastic meningiomas identifies novel genes associated with meningioma progression. Int. J. Cancer 2005, 114, 249–256. [Google Scholar] [CrossRef]
- Fevre-Montange, M.; Champier, J.; Durand, A.; Wierinckx, A.; Honnorat, J.; Guyotat, J.; Jouvet, A. Microarray gene expression profiling in meningiomas: Differential expression according to grade or histopathological subtype. Int. J. Oncol. 2009, 35, 1395–1407. [Google Scholar] [CrossRef] [PubMed]
- Menghi, F.; Orzan, F.N.; Eoli, M.; Farinotti, M.; Maderna, E.; Pisati, F.; Bianchessi, D.; Valletta, L.; Lodrini, S.; Galli, G.; et al. DNA microarray analysis identifies CKS2 and LEPR as potential markers of meningioma recurrence. Oncologist 2011, 16, 1440–1450. [Google Scholar] [CrossRef][Green Version]
- Nakane, Y.; Natsume, A.; Wakabayashi, T.; Oi, S.; Ito, M.; Inao, S.; Saito, K.; Yoshida, J. Malignant transformation-related genes in meningiomas: Allelic loss on 1p36 and methylation status of p73 and RASSF1A. J. Neurosurg. 2007, 107, 398–404. [Google Scholar] [CrossRef] [PubMed]
- Ohba, S.; Yoshida, K.; Hirose, Y.; Ikeda, E.; Kawase, T. Early malignant transformation of a petroclival meningothelial meningioma. Neurosurg. Rev. 2009, 32, 495–499. [Google Scholar] [CrossRef] [PubMed]
- Pico, C.; Sanchez, J.; Oliver, P.; Palou, A. Leptin production by the stomach is up-regulated in obese (fa/fa) Zucker rats. Obes. Res. 2002, 10, 932–938. [Google Scholar] [CrossRef]
- Aghi, M.K.; Eskandar, E.N.; Carter, B.S.; Curry, W.T., Jr.; Barker, F.G., 2nd. Increased prevalence of obesity and obesity-related postoperative complications in male patients with meningiomas. Neurosurgery 2007, 61, 754–760, discussion 760–761. [Google Scholar] [CrossRef] [PubMed]
- Schneider, B.; Pulhorn, H.; Rohrig, B.; Rainov, N.G. Predisposing conditions and risk factors for development of symptomatic meningioma in adults. Cancer Detect. Prev. 2005, 29, 440–447. [Google Scholar] [CrossRef]
- Du, Z.; Brewster, R.; Merrill, P.H.; Chmielecki, J.; Francis, J.; Aizer, A.; Abedalthagafi, M.; Sholl, L.M.; Geffers, L.; Alexander, B.; et al. Meningioma transcription factors link cell lineage with systemic metabolic cues. Neuro-Oncology 2018, 20, 1331–1343. [Google Scholar] [CrossRef] [PubMed]
- Domingues, P.H.; Teodosio, C.; Ortiz, J.; Sousa, P.; Otero, A.; Maillo, A.; Barcena, P.; Garcia-Macias, M.C.; Lopes, M.C.; de Oliveira, C.; et al. Immunophenotypic identification and characterization of tumor cells and infiltrating cell populations in meningiomas. Am. J. Pathol. 2012, 181, 1749–1761. [Google Scholar] [CrossRef]
- Du, Z.; Abedalthagafi, M.; Aizer, A.A.; McHenry, A.R.; Sun, H.H.; Bray, M.A.; Viramontes, O.; Machaidze, R.; Brastianos, P.K.; Reardon, D.A.; et al. Increased expression of the immune modulatory molecule PD-L1 (CD274) in anaplastic meningioma. Oncotarget 2015, 6, 4704–4716. [Google Scholar] [CrossRef]
- Giles, A.J.; Hao, S.; Padget, M.; Song, H.; Zhang, W.; Lynes, J.; Sanchez, V.; Liu, Y.; Jung, J.; Cao, X.; et al. Efficient ADCC killing of meningioma by avelumab and a high-affinity natural killer cell line, haNK. JCI Insight 2019, 4, e130688. [Google Scholar] [CrossRef]
- Lee, H.H.; Wang, Y.N.; Xia, W.; Chen, C.H.; Rau, K.M.; Ye, L.; Wei, Y.; Chou, C.K.; Wang, S.C.; Yan, M.; et al. Removal of N-Linked Glycosylation Enhances PD-L1 Detection and Predicts Anti-PD-1/PD-L1 Therapeutic Efficacy. Cancer Cell 2019, 36, 168–178.e4. [Google Scholar] [CrossRef]
- Mezzadra, R.; Sun, C.; Jae, L.T.; Gomez-Eerland, R.; de Vries, E.; Wu, W.; Logtenberg, M.E.W.; Slagter, M.; Rozeman, E.A.; Hofland, I.; et al. Identification of CMTM6 and CMTM4 as PD-L1 protein regulators. Nature 2017, 549, 106–110. [Google Scholar] [CrossRef] [PubMed]
- Barresi, V.; Lionti, S.; Caliri, S.; Caffo, M. Histopathological features to define atypical meningioma: What does really matter for prognosis? Brain Tumor. Pathol. 2018, 35, 168–180. [Google Scholar] [CrossRef] [PubMed]
| Items | Subtype 1 | Subtype 2 | Subtype 3 | Subtype 4 |
|---|---|---|---|---|
| Age (median, 25–75% quantile) | 61, 47.5–68 | 61, 50–66 | 61,49.5–72 | 61, 58–68.5 |
| Gender (Male:Female) | 53:12 | 24:14 | 20:27 | 10:4 |
| WHO Grade | ||||
| I | 67 | 32 | 28 | 13 |
| II | 0 | 6 | 24 | 2 |
| III | 0 | 0 | 7 | 0 |
| Variables | Sub-Variables | Coefficient | Lower 95% CI | Upper 95% CI | p Value |
|---|---|---|---|---|---|
| MPscore | 1.63762 | 2.00798 | 13.1721 | 0.000643 | |
| Location | |||||
| Convexity | −0.19415 | 0.18135 | 3.7398 | 0.801446 | |
| Falx | −0.44845 | 0.13599 | 2.9991 | 0.569863 | |
| Intraventricular | 0.59538 | 0.25013 | 13.1514 | 0.555854 | |
| Olfactory groove | 0.45279 | 0.32038 | 7.7202 | 0.576995 | |
| Optic nerve | −0.50801 | 0.08373 | 4.3237 | 0.613643 | |
| Parasagittal | −0.0126 | 0.2065 | 4.7222 | 0.987409 | |
| Posterior fossa | 0.55699 | 0.37301 | 8.1672 | 0.479293 | |
| Sphenoid wing | −0.22505 | 0.17501 | 3.643 | 0.771353 | |
| Supresellar | −1.27342 | 0.05222 | 1.5 | 0.137122 | |
| Gender | −0.34111 | 0.40969 | 1.2338 | 0.225178 | |
| MIB-1 | −0.06811 | 0.87729 | 0.9947 | 0.033544 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, F.; Qian, J.; Ma, C. MPscore: A Novel Predictive and Prognostic Scoring for Progressive Meningioma. Cancers 2021, 13, 1113. https://doi.org/10.3390/cancers13051113
Liu F, Qian J, Ma C. MPscore: A Novel Predictive and Prognostic Scoring for Progressive Meningioma. Cancers. 2021; 13(5):1113. https://doi.org/10.3390/cancers13051113
Chicago/Turabian StyleLiu, Feili, Jin Qian, and Chenkai Ma. 2021. "MPscore: A Novel Predictive and Prognostic Scoring for Progressive Meningioma" Cancers 13, no. 5: 1113. https://doi.org/10.3390/cancers13051113
APA StyleLiu, F., Qian, J., & Ma, C. (2021). MPscore: A Novel Predictive and Prognostic Scoring for Progressive Meningioma. Cancers, 13(5), 1113. https://doi.org/10.3390/cancers13051113






