Diagnostic, Predictive and Prognostic Molecular Biomarkers in Pancreatic Cancer: An Overview for Clinicians
Abstract
Simple Summary
Abstract
1. Introduction
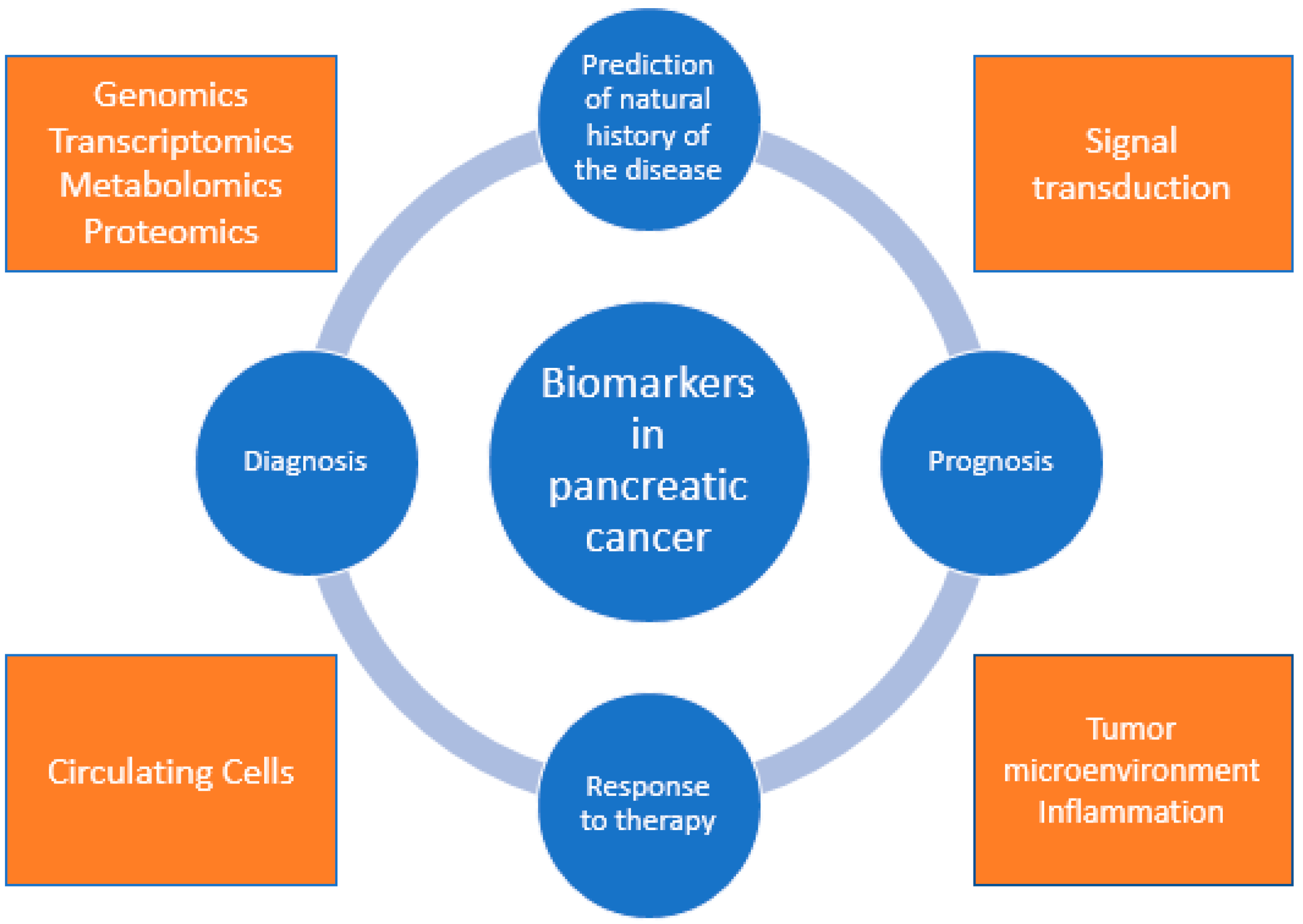
2. Biomarkers
2.1. CA 19-9: Diagnostic and Prognostic Role
2.2. microRNAs: Diagnostic and Prognostic Role
2.3. DNA Methylation Patterns: Diagnostic and Prognostic Role
2.4. Mismatch Repair Genes and Microsatellite Instability: Diagnostic and Predictive Role
2.5. KRAS Mutations: Diagnostic and Prognostic Role
2.6. Exosomes: Diagnostic and Prognostic Role
2.7. Circulating Tumor Cells: Diagnostic and Prognostic Role
2.8. PAM4/MUC5AC: Diagnostic Role
2.9. Osteopontin: Diagnostic and Prognostic Role
2.10. SMAD4/DPC4: Diagnostic, Prognostic and Predictive Role
2.11. Immune Response and Inflammatory Markers: Prognostic and Predictive Role
2.12. Human Equilibrative Nucleoside Transporter 1 (hENT1): Prognostic and Predictive Role
2.13. Human Concentrative Nucleoside Transporters 1 and 3 (hCNT1 and hCNT3): Prognostic and Predictive Role
2.14. BRCA1 and BRCA2: Prognostic and Predictive Role
3. Hypoxia as the Hallmark of Pancreatic Cancer Pathogenesis
4. Framing Proteomic Work in Pancreatic Cancer
5. Challenges and Future Directions
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer Statistics, 2019. CA Cancer J. Clin. 2019, 69, 7–34. [Google Scholar] [CrossRef] [PubMed]
- Raimondi, S.; Maisonneuve, P.; Lowenfels, A.B. Epidemiology of Pancreatic Cancer: An Overview. Nat. Rev. Gastroenterol. Hepatol. 2009, 6, 699–708. [Google Scholar] [CrossRef]
- Cancer of the Pancreas—Cancer Stat Facts. Available online: https://seer.cancer.gov/statfacts/html/pancreas.html (accessed on 26 April 2020).
- Hidalgo, M. Pancreatic Cancer. N. Engl. J. Med. 2010, 362, 1605–1617. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Wei, M.; Xu, J.; Hua, J.; Liang, C.; Meng, Q.; Zhang, Y.; Liu, J.; Zhang, B.; Yu, X.; et al. PARP Inhibitors in Pancreatic Cancer: Molecular Mechanisms and Clinical Applications. Mol. Cancer 2020, 19. [Google Scholar] [CrossRef] [PubMed]
- Bengtsson, A.; Andersson, R.; Ansari, D. The Actual 5-Year Survivors of Pancreatic Ductal Adenocarcinoma Based on Real-World Data. Sci. Rep. 2020, 10, 16425. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, E.A.; Beal, E.W.; Chakedis, J.; Paredes, A.Z.; Moris, D.; Pawlik, T.M.; Schmidt, C.R.; Dillhoff, M.E. Exosomes in Pancreatic Cancer: From Early Detection to Treatment. J. Gastrointest. Surg. 2018, 22, 737–750. [Google Scholar] [CrossRef]
- Chari, S.T.; Kelly, K.; Hollingsworth, M.A.; Thayer, S.P.; Ahlquist, D.A.; Andersen, D.K.; Batra, S.K.; Brentnall, T.A.; Canto, M.; Cleeter, D.F.; et al. Early Detection of Sporadic Pancreatic Cancer: Summative Review. Pancreas 2015, 44, 693–712. [Google Scholar] [CrossRef] [PubMed]
- Goonetilleke, K.S.; Siriwardena, A.K. Systematic Review of Carbohydrate Antigen (CA 19-9) as a Biochemical Marker in the Diagnosis of Pancreatic Cancer. Eur. J. Surg. Oncol. 2007, 33, 266–270. [Google Scholar] [CrossRef] [PubMed]
- Tessler, D.A.; Catanzaro, A.; Velanovich, V.; Havstad, S.; Goel, S. Predictors of Cancer in Patients with Suspected Pancreatic Malignancy without a Tissue Diagnosis. Am. J. Surg. 2006, 191, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Van Manen, L.; Groen, J.V.; Putter, H.; Vahrmeijer, A.L.; Swijnenburg, R.-J.; Bonsing, B.A.; Mieog, J.S.D. Elevated CEA and CA19-9 Serum Levels Independently Predict Advanced Pancreatic Cancer at Diagnosis. Biomarkers 2020, 25, 186–193. [Google Scholar] [CrossRef]
- Palmquist, C.; Dehlendorff, C.; Calatayud, D.; Hansen, C.P.; Hasselby, J.P.; Johansen, J.S. Prediction of Unresectability and Prognosis in Patients Undergoing Surgery on Suspicion of Pancreatic Cancer Using Carbohydrate Antigen 19-9, Interleukin 6, and YKL-40. Pancreas 2020, 49, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Ferrone, C.R.; Finkelstein, D.M.; Thayer, S.P.; Muzikansky, A.; Fernandez-delCastillo, C.; Warshaw, A.L. Perioperative CA19-9 Levels Can Predict Stage and Survival in Patients with Resectable Pancreatic Adenocarcinoma. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2006, 24, 2897–2902. [Google Scholar] [CrossRef]
- Berardi, R.; Mandolesi, A.; Pellei, C.; Maccaroni, E.; Onofri, A.; Lucarelli, A.; Biagetti, S.; Alfonsi, S.; Caramanti, M.; Savini, A.; et al. Prognostic Factors in Pancreatic Cancer: The Role of Perineural, Vascular and Lymphatic Invasion and of Ca19-9. J. Gastrointest. Dig. Syst. 2013, 03. [Google Scholar] [CrossRef]
- Kim, J.-E.; Lee, K.T.; Lee, J.K.; Paik, S.W.; Rhee, J.C.; Choi, K.W. Clinical Usefulness of Carbohydrate Antigen 19-9 as a Screening Test for Pancreatic Cancer in an Asymptomatic Population. J. Gastroenterol. Hepatol. 2004, 19, 182–186. [Google Scholar] [CrossRef]
- Chang, C.-Y.; Huang, S.-P.; Chiu, H.-M.; Lee, Y.-C.; Chen, M.-F.; Lin, J.-T. Low Efficacy of Serum Levels of CA 19-9 in Prediction of Malignant Diseases in Asymptomatic Population in Taiwan. Hepatogastroenterology 2006, 53, 1–4. [Google Scholar]
- Kim, M.S.; Jeon, T.J.; Park, J.Y.; Choi, J.; Shin, W.C.; Park, S.E.; Seo, J.Y.; Kim, Y.M. Clinical Interpretation of Elevated CA 19-9 Levels in Obstructive Jaundice Following Benign and Malignant Pancreatobiliary Disease. Korean J. Gastroenterol. Taehan Sohwagi Hakhoe Chi 2017, 70, 96–102. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Binicier, O.B.; Pakoz, Z.B. CA 19-9 Levels in Patients with Acute Pancreatitis Due to Gallstone and Metabolic/Toxic Reasons. Rev. Assoc. Med. Bras. 2019, 65, 965–970. [Google Scholar] [CrossRef] [PubMed]
- Pandey, D.; Sharma, R.; Sharma, S.; Salhan, S. Unusually High Serum Levels of CA 19-9 in an Ovarian Tumour: Malignant or Benign? J. Clin. Diagn. Res. JCDR 2017, 11, QD08–QD10. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.-Y.; Lin, M.; Zhang, H.-B. Diagnostic Value of Carcinoembryonic Antigen and Carcinoma Antigen 19-9 for Colorectal Carcinoma. Int. J. Clin. Exp. Pathol. 2015, 8, 9404–9409. [Google Scholar] [PubMed]
- Lumachi, F.; Lo Re, G.; Tozzoli, R.; D’Aurizio, F.; Facomer, F.; Chiara, G.B.; Basso, S.M.M. Measurement of Serum Carcinoembryonic Antigen, Carbohydrate Antigen 19-9, Cytokeratin-19 Fragment and Matrix Metalloproteinase-7 for Detecting Cholangiocarcinoma: A Preliminary Case-Control Study. Anticancer Res. 2014, 34, 6663–6667. [Google Scholar]
- Bertino, G.; Ardiri, A.M.; Calvagno, G.S.; Malaguarnera, G.; Interlandi, D.; Vacante, M.; Bertino, N.; Lucca, F.; Madeddu, R.; Motta, M. Carbohydrate 19.9 Antigen Serum Levels in Liver Disease. BioMed Res. Int. 2013, 2013, 531640. [Google Scholar] [CrossRef]
- Luo, G.; Liu, C.; Guo, M.; Cheng, H.; Lu, Y.; Jin, K.; Liu, L.; Long, J.; Xu, J.; Lu, R.; et al. Potential Biomarkers in Lewis Negative Patients with Pancreatic Cancer. Ann. Surg. 2017, 265, 800–805. [Google Scholar] [CrossRef]
- Kannagi, R. Carbohydrate Antigen Sialyl Lewis A--Its Pathophysiological Significance and Induction Mechanism in Cancer Progression. Chang. Gung Med. J. 2007, 30, 189–209. [Google Scholar] [PubMed]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402. [Google Scholar] [CrossRef] [PubMed]
- Lekchnov, E.A.; Zaporozhchenko, I.A.; Morozkin, E.S.; Bryzgunova, O.E.; Vlassov, V.V.; Laktionov, P.P. Protocol for MiRNA Isolation from Biofluids. Anal. Biochem. 2016, 499, 78–84. [Google Scholar] [CrossRef] [PubMed]
- Bloomston, M.; Frankel, W.L.; Petrocca, F.; Volinia, S.; Alder, H.; Hagan, J.P.; Liu, C.-G.; Bhatt, D.; Taccioli, C.; Croce, C.M. MicroRNA Expression Patterns to Differentiate Pancreatic Adenocarcinoma from Normal Pancreas and Chronic Pancreatitis. JAMA 2007, 297, 1901–1908. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.-Y.; Sun, Y.-W.; Liu, D.-J.; Zhang, J.-F.; Li, J.; Hua, R. MicroRNAs in Stool Samples as Potential Screening Biomarkers for Pancreatic Ductal Adenocarcinoma Cancer. Am. J. Cancer Res. 2014, 4, 663–673. [Google Scholar]
- Debernardi, S.; Massat, N.J.; Radon, T.P.; Sangaralingam, A.; Banissi, A.; Ennis, D.P.; Dowe, T.; Chelala, C.; Pereira, S.P.; Kocher, H.M.; et al. Noninvasive Urinary MiRNA Biomarkers for Early Detection of Pancreatic Adenocarcinoma. Am. J. Cancer Res. 2015, 5, 3455–3466. [Google Scholar] [PubMed]
- Brancaccio, M.; Natale, F.; Falco, G.; Angrisano, T. Cell-Free DNA Methylation: The New Frontiers of Pancreatic Cancer Biomarkers’ Discovery. Genes 2019, 11, 14. [Google Scholar] [CrossRef] [PubMed]
- Koch, A.; Joosten, S.C.; Feng, Z.; de Ruijter, T.C.; Draht, M.X.; Melotte, V.; Smits, K.M.; Veeck, J.; Herman, J.G.; Van Neste, L.; et al. Analysis of DNA Methylation in Cancer: Location Revisited. Nat. Rev. Clin. Oncol. 2018, 15, 459–466. [Google Scholar] [CrossRef]
- Baylin, S.B.; Jones, P.A. Epigenetic Determinants of Cancer. Cold Spring Harb. Perspect. Biol. 2016, 8. [Google Scholar] [CrossRef] [PubMed]
- Kisiel, J.B.; Raimondo, M.; Taylor, W.R.; Yab, T.C.; Mahoney, D.W.; Sun, Z.; Middha, S.; Baheti, S.; Zou, H.; Smyrk, T.C.; et al. New DNA Methylation Markers for Pancreatic Cancer: Discovery, Tissue Validation, and Pilot Testing in Pancreatic Juice. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2015, 21, 4473–4481. [Google Scholar] [CrossRef] [PubMed]
- Matsubayashi, H.; Canto, M.; Sato, N.; Klein, A.; Abe, T.; Yamashita, K.; Yeo, C.J.; Kalloo, A.; Hruban, R.; Goggins, M. DNA Methylation Alterations in the Pancreatic Juice of Patients with Suspected Pancreatic Disease. Cancer Res. 2006, 66, 1208–1217. [Google Scholar] [CrossRef] [PubMed]
- Parsi, M.A.; Li, A.; Li, C.-P.; Goggins, M. DNA methylation alterations in ERCP brush samples of patients with suspected pancreaticobiliary disease. Clin. Gastroenterol. Hepatol. Off. Clin. Pract. J. Am. Gastroenterol. Assoc. 2008, 6, 1270–1278. [Google Scholar] [CrossRef]
- Henriksen, S.D.; Madsen, P.H.; Larsen, A.C.; Johansen, M.B.; Pedersen, I.S.; Krarup, H.; Thorlacius-Ussing, O. Cell-Free DNA Promoter Hypermethylation in Plasma as a Predictive Marker for Survival of Patients with Pancreatic Adenocarcinoma. Oncotarget 2017, 8, 93942–93956. [Google Scholar] [CrossRef]
- Lupinacci, R.M.; Bachet, J.-B.; André, T.; Duval, A.; Svrcek, M. Pancreatic Ductal Adenocarcinoma Harboring Microsatellite Instability/DNA Mismatch Repair Deficiency. Towards Personalized Medicine. Surg. Oncol. 2019, 28, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Grant, R.C.; Denroche, R.; Jang, G.H.; Nowak, K.M.; Zhang, A.; Borgida, A.; Holter, S.; Topham, J.T.; Wilson, J.; Dodd, A.; et al. Clinical and Genomic Characterisation of Mismatch Repair Deficient Pancreatic Adenocarcinoma. Gut 2020. [Google Scholar] [CrossRef] [PubMed]
- Singhi, A.D.; George, B.; Greenbowe, J.R.; Chung, J.; Suh, J.; Maitra, A.; Klempner, S.J.; Hendifar, A.; Milind, J.M.; Golan, T.; et al. Real-Time Targeted Genome Profile Analysis of Pancreatic Ductal Adenocarcinomas Identifies Genetic Alterations That Might Be Targeted with Existing Drugs or Used as Biomarkers. Gastroenterology 2019, 156, 2242–2253.e4. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.I.; Shia, J.; Stadler, Z.K.; Varghese, A.M.; Capanu, M.; Salo-Mullen, E.; Lowery, M.A.; Diaz, L.A.; Mandelker, D.; Yu, K.H.; et al. Evaluating Mismatch Repair Deficiency in Pancreatic Adenocarcinoma: Challenges and Recommendations. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2018, 24, 1326–1336. [Google Scholar] [CrossRef] [PubMed]
- Lupinacci, R.M.; Goloudina, A.; Buhard, O.; Bachet, J.-B.; Maréchal, R.; Demetter, P.; Cros, J.; Bardier-Dupas, A.; Collura, A.; Cervera, P.; et al. Prevalence of Microsatellite Instability in Intraductal Papillary Mucinous Neoplasms of the Pancreas. Gastroenterology 2018, 154, 1061–1065. [Google Scholar] [CrossRef]
- Humphris, J.L.; Patch, A.-M.; Nones, K.; Bailey, P.J.; Johns, A.L.; McKay, S.; Chang, D.K.; Miller, D.K.; Pajic, M.; Kassahn, K.S.; et al. Hypermutation In Pancreatic Cancer. Gastroenterology 2017, 152, 68–74.e2. [Google Scholar] [CrossRef]
- Nakata, B.; Wang, Y.Q.; Yashiro, M.; Nishioka, N.; Tanaka, H.; Ohira, M.; Ishikawa, T.; Nishino, H.; Hirakawa, K. Prognostic Value of Microsatellite Instability in Resectable Pancreatic Cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2002, 8, 2536–2540. [Google Scholar]
- Ottenhof, N.A.; Morsink, F.H.M.; ten Kate, F.; van Noorden, C.J.F.; Offerhaus, G.J.A. Multivariate Analysis of Immunohistochemical Evaluation of Protein Expression in Pancreatic Ductal Adenocarcinoma Reveals Prognostic Significance for Persistent Smad4 Expression Only. Cell. Oncol. Dordr. 2012, 35, 119–126. [Google Scholar] [CrossRef]
- Marabelle, A.; Le, D.T.; Ascierto, P.A.; Di Giacomo, A.M.; De Jesus-Acosta, A.; Delord, J.-P.; Geva, R.; Gottfried, M.; Penel, N.; Hansen, A.R.; et al. Efficacy of Pembrolizumab in Patients With Noncolorectal High Microsatellite Instability/Mismatch Repair-Deficient Cancer: Results From the Phase II KEYNOTE-158 Study. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2020, 38, 1–10. [Google Scholar] [CrossRef]
- Cloyd, J.M.; Katz, M.H.G.; Wang, H.; Cuddy, A.; You, Y.N. Clinical and Genetic Implications of DNA Mismatch Repair Deficiency in Patients With Pancreatic Ductal Adenocarcinoma. JAMA Surg. 2017, 152, 1086–1088. [Google Scholar] [CrossRef] [PubMed]
- Riazy, M.; Kalloger, S.E.; Sheffield, B.S.; Peixoto, R.D.; Li-Chang, H.H.; Scudamore, C.H.; Renouf, D.J.; Schaeffer, D.F. Mismatch Repair Status May Predict Response to Adjuvant Chemotherapy in Resectable Pancreatic Ductal Adenocarcinoma. Mod. Pathol. 2015, 28, 1383–1389. [Google Scholar] [CrossRef]
- Australian Pancreatic Cancer Genome Initiative; Biankin, A.V.; Waddell, N.; Kassahn, K.S.; Gingras, M.-C.; Muthuswamy, L.B.; Johns, A.L.; Miller, D.K.; Wilson, P.J.; Patch, A.-M.; et al. Pancreatic Cancer Genomes Reveal Aberrations in Axon Guidance Pathway Genes. Nature 2012, 491, 399–405. [Google Scholar] [CrossRef]
- Gall, T.M.H.; Belete, S.; Khanderia, E.; Frampton, A.E.; Jiao, L.R. Circulating Tumor Cells and Cell-Free DNA in Pancreatic Ductal Adenocarcinoma. Am. J. Pathol. 2019, 189, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Gharibi, A.; Adamian, Y.; Kelber, J.A. Cellular and Molecular Aspects of Pancreatic Cancer. Acta Histochem. 2016, 118, 305–316. [Google Scholar] [CrossRef]
- Collins, M.A.; Bednar, F.; Zhang, Y.; Brisset, J.-C.; Galbán, S.; Galbán, C.J.; Rakshit, S.; Flannagan, K.S.; Adsay, N.V.; Pasca di Magliano, M. Oncogenic Kras Is Required for Both the Initiation and Maintenance of Pancreatic Cancer in Mice. J. Clin. Investig. 2012, 122, 639–653. [Google Scholar] [CrossRef]
- Ying, H.; Kimmelman, A.C.; Lyssiotis, C.A.; Hua, S.; Chu, G.C.; Fletcher-Sananikone, E.; Locasale, J.W.; Son, J.; Zhang, H.; Coloff, J.L.; et al. Oncogenic Kras Maintains Pancreatic Tumors through Regulation of Anabolic Glucose Metabolism. Cell 2012, 149, 656–670. [Google Scholar] [CrossRef]
- Kinugasa, H.; Nouso, K.; Miyahara, K.; Morimoto, Y.; Dohi, C.; Tsutsumi, K.; Kato, H.; Matsubara, T.; Okada, H.; Yamamoto, K. Detection of K-Ras Gene Mutation by Liquid Biopsy in Patients with Pancreatic Cancer. Cancer 2015, 121, 2271–2280. [Google Scholar] [CrossRef]
- Inga, E.; Perdomo Zaldivar, E.; Gómez, M.; Cano, T.; Rodriguez Alonso, B.; Ortiz, M.; Rodriguez Alonso, R.; Toledano Fonseca, M.; Rodriguez Ariza, A.; Aranda, E. Impact of KRAS Mutations in Clinical Features and Survival in Pancreatic Cancer Patients: A Single Institution Experience. Ann. Oncol. 2019, 30, iv94. [Google Scholar] [CrossRef]
- Cheng, H.; Luo, G.; Jin, K.; Fan, Z.; Huang, Q.; Gong, Y.; Xu, J.; Yu, X.; Liu, C. Kras Mutation Correlating with Circulating Regulatory T Cells Predicts the Prognosis of Advanced Pancreatic Cancer Patients. Cancer Med. 2020, 9, 2153–2159. [Google Scholar] [CrossRef]
- Lee, B.; Lipton, L.; Cohen, J.; Tie, J.; Javed, A.A.; Li, L.; Goldstein, D.; Burge, M.; Cooray, P.; Nagrial, A.; et al. Circulating Tumor DNA as a Potential Marker of Adjuvant Chemotherapy Benefit Following Surgery for Localized Pancreatic Cancer. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2019, 30, 1472–1478. [Google Scholar] [CrossRef] [PubMed]
- Chan-Seng-Yue, M.; Kim, J.C.; Wilson, G.W.; Ng, K.; Figueroa, E.F.; O’Kane, G.M.; Connor, A.A.; Denroche, R.E.; Grant, R.C.; McLeod, J.; et al. Transcription Phenotypes of Pancreatic Cancer Are Driven by Genomic Events during Tumor Evolution. Nat. Genet. 2020, 52, 231–240. [Google Scholar] [CrossRef]
- Ge, R.; Tan, E.; Sharghi-Namini, S.; Asada, H.H. Exosomes in Cancer Microenvironment and Beyond: Have We Overlooked These Extracellular Messengers? Cancer Microenviron. Off. J. Int. Cancer Microenviron. Soc. 2012, 5, 323–332. [Google Scholar] [CrossRef]
- Lorenzon, L.; Blandino, G. Glypican-1 Exosomes: Do They Initiate a New Era for Early Pancreatic Cancer Diagnosis? Transl. Gastroenterol. Hepatol. 2016, 1, 8. [Google Scholar] [CrossRef] [PubMed]
- Cazzoli, R.; Buttitta, F.; Di Nicola, M.; Malatesta, S.; Marchetti, A.; Rom, W.N.; Pass, H.I. MicroRNAs Derived from Circulating Exosomes as Noninvasive Biomarkers for Screening and Diagnosing Lung Cancer. J. Thorac. Oncol. 2013, 8, 1156–1162. [Google Scholar] [CrossRef] [PubMed]
- Moris, D.; Beal, E.W.; Chakedis, J.; Burkhart, R.A.; Schmidt, C.; Dillhoff, M.; Zhang, X.; Theocharis, S.; Pawlik, T.M. Role of Exosomes in Treatment of Hepatocellular Carcinoma. Surg. Oncol. 2017, 26, 219–228. [Google Scholar] [CrossRef]
- Wu, C.-Y.; Du, S.-L.; Zhang, J.; Liang, A.-L.; Liu, Y.-J. Exosomes and Breast Cancer: A Comprehensive Review of Novel Therapeutic Strategies from Diagnosis to Treatment. Cancer Gene Ther. 2017, 24, 6–12. [Google Scholar] [CrossRef] [PubMed]
- Ogata-Kawata, H.; Izumiya, M.; Kurioka, D.; Honma, Y.; Yamada, Y.; Furuta, K.; Gunji, T.; Ohta, H.; Okamoto, H.; Sonoda, H.; et al. Circulating Exosomal MicroRNAs as Biomarkers of Colon Cancer. PLoS ONE 2014, 9, e92921. [Google Scholar] [CrossRef] [PubMed]
- Allenson, K.; Castillo, J.; Lucas, F.A.S.; Scelo, G.; Kim, D.U.; Bernard, V.; Davis, G.; Kumar, T.; Katz, M.; Overman, M.J.; et al. High Prevalence of MutantKRAS in Circulating Exosome-Derived DNA from Early-Stage Pancreatic Cancer Patients. Ann. Oncol. 2017, 28, 741–747. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Che, S.P.Y.; Kurywchak, P.; Tavormina, J.L.; Gansmo, L.B.; Correa de Sampaio, P.; Tachezy, M.; Bockhorn, M.; Gebauer, F.; Haltom, A.R.; et al. Detection of Mutant KRAS and TP53 DNA in Circulating Exosomes from Healthy Individuals and Patients with Pancreatic Cancer. Cancer Biol. Ther. 2017, 18, 158–165. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Kleeff, J.; Kayed, H.; Felix, K.; Penzel, R.; Büchler, M.W.; Korc, M.; Friess, H. Glypican-1 Antisense Transfection Modulates TGF-Beta-Dependent Signaling in Colo-357 Pancreatic Cancer Cells. Biochem. Biophys. Res. Commun. 2004, 320, 1148–1155. [Google Scholar] [CrossRef] [PubMed]
- Ding, K.; Lopez-Burks, M.; Sánchez-Duran, J.A.; Korc, M.; Lander, A.D. Growth Factor-Induced Shedding of Syndecan-1 Confers Glypican-1 Dependence on Mitogenic Responses of Cancer Cells. J. Cell Biol. 2005, 171, 729–738. [Google Scholar] [CrossRef] [PubMed]
- Kleeff, J.; Wildi, S.; Kumbasar, A.; Friess, H.; Lander, A.D.; Korc, M. Stable Transfection of a Glypican-1 Antisense Construct Decreases Tumorigenicity in PANC-1 Pancreatic Carcinoma Cells. Pancreas 1999, 19, 281–288. [Google Scholar] [CrossRef] [PubMed]
- Duan, L.; Hu, X.; Feng, D.; Lei, S.; Hu, G. GPC-1 May Serve as a Predictor of Perineural Invasion and a Prognosticator of Survival in Pancreatic Cancer. Asian J. Surg. 2013, 36, 7–12. [Google Scholar] [CrossRef] [PubMed]
- Melo, S.A.; Luecke, L.B.; Kahlert, C.; Fernandez, A.F.; Gammon, S.T.; Kaye, J.; LeBleu, V.S.; Mittendorf, E.A.; Weitz, J.; Rahbari, N.; et al. Glypican-1 Identifies Cancer Exosomes and Detects Early Pancreatic Cancer. Nature 2015, 523, 177–182. [Google Scholar] [CrossRef]
- Que, R.; Ding, G.; Chen, J.; Cao, L. Analysis of Serum Exosomal MicroRNAs and Clinicopathologic Features of Patients with Pancreatic Adenocarcinoma. World J. Surg. Oncol. 2013, 11, 219. [Google Scholar] [CrossRef]
- Machida, T.; Tomofuji, T.; Maruyama, T.; Yoneda, T.; Ekuni, D.; Azuma, T.; Miyai, H.; Mizuno, H.; Kato, H.; Tsutsumi, K.; et al. MiR-1246 and MiR-4644 in Salivary Exosome as Potential Biomarkers for Pancreatobiliary Tract Cancer. Oncol. Rep. 2016, 36, 2375–2381. [Google Scholar] [CrossRef] [PubMed]
- Martini, V.; Timme-Bronsert, S.; Fichtner-Feigl, S.; Hoeppner, J.; Kulemann, B. Circulating Tumor Cells in Pancreatic Cancer: Current Perspectives. Cancers 2019, 11, 1659. [Google Scholar] [CrossRef]
- Rhim, A.D.; Mirek, E.T.; Aiello, N.M.; Maitra, A.; Bailey, J.M.; McAllister, F.; Reichert, M.; Beatty, G.L.; Rustgi, A.K.; Vonderheide, R.H.; et al. EMT and Dissemination Precede Pancreatic Tumor Formation. Cell 2012, 148, 349–361. [Google Scholar] [CrossRef] [PubMed]
- Kulemann, B.; Pitman, M.B.; Liss, A.S.; Valsangkar, N.; Fernández-Del Castillo, C.; Lillemoe, K.D.; Hoeppner, J.; Mino-Kenudson, M.; Warshaw, A.L.; Thayer, S.P. Circulating Tumor Cells Found in Patients with Localized and Advanced Pancreatic Cancer. Pancreas 2015, 44, 547–550. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Zhang, H.; Chen, N.; Hao, J.; Jin, H.; Ma, X. Diagnostic Value of Various Liquid Biopsy Methods for Pancreatic Cancer: A Systematic Review and Meta-Analysis. Medicine 2020, 99, e18581. [Google Scholar] [CrossRef]
- Wang, Y.; Yu, X.; Hartmann, D.; Zhou, J. Circulating Tumor Cells in Peripheral Blood of Pancreatic Cancer Patients and Their Prognostic Role: A Systematic Review and Meta-Analysis. HPB 2019. [Google Scholar] [CrossRef] [PubMed]
- Gold, D.V.; Gaedcke, J.; Ghadimi, B.M.; Goggins, M.; Hruban, R.H.; Liu, M.; Newsome, G.; Goldenberg, D.M. PAM4 Enzyme Immunoassay Alone and in Combination with CA 19-9 for the Detection of Pancreatic Adenocarcinoma. Cancer 2013, 119, 522–528. [Google Scholar] [CrossRef]
- Gold, D.V.; Newsome, G.; Liu, D.; Goldenberg, D.M. Mapping PAM4 (Clivatuzumab), a Monoclonal Antibody in Clinical Trials for Early Detection and Therapy of Pancreatic Ductal Adenocarcinoma, to MUC5AC Mucin. Mol. Cancer 2013, 12, 143. [Google Scholar] [CrossRef]
- Liu, D.; Chang, C.-H.; Gold, D.V.; Goldenberg, D.M. Identification of PAM4 (Clivatuzumab)-Reactive Epitope on MUC5AC: A Promising Biomarker and Therapeutic Target for Pancreatic Cancer. Oncotarget 2015, 6, 4274–4285. [Google Scholar] [CrossRef]
- Shi, C.; Merchant, N.; Newsome, G.; Goldenberg, D.M.; Gold, D.V. Differentiation of Pancreatic Ductal Adenocarcinoma from Chronic Pancreatitis by PAM4 Immunohistochemistry. Arch. Pathol. Lab. Med. 2014, 138, 220–228. [Google Scholar] [CrossRef] [PubMed]
- Koopmann, J.; Fedarko, N.S.; Jain, A.; Maitra, A.; Iacobuzio-Donahue, C.; Rahman, A.; Hruban, R.H.; Yeo, C.J.; Goggins, M. Evaluation of Osteopontin as Biomarker for Pancreatic Adenocarcinoma. Cancer Epidemiol. Biomark. Prev. Publ. Am. Assoc. Cancer Res. Cosponsored Am. Soc. Prev. Oncol. 2004, 13, 487–491. [Google Scholar] [CrossRef]
- O’Brien, E.R.; Garvin, M.R.; Stewart, D.K.; Hinohara, T.; Simpson, J.B.; Schwartz, S.M.; Giachelli, C.M. Osteopontin Is Synthesized by Macrophage, Smooth Muscle, and Endothelial Cells in Primary and Restenotic Human Coronary Atherosclerotic Plaques. Arterioscler. Thromb. J. Vasc. Biol. 1994, 14, 1648–1656. [Google Scholar] [CrossRef] [PubMed]
- Poruk, K.E.; Firpo, M.A.; Scaife, C.L.; Adler, D.G.; Emerson, L.L.; Boucher, K.M.; Mulvihill, S.J. Serum Osteopontin and Tissue Inhibitor of Metalloproteinase 1 as Diagnostic and Prognostic Biomarkers for Pancreatic Adenocarcinoma. Pancreas 2013, 42, 193–197. [Google Scholar] [CrossRef] [PubMed]
- Rychlíková, J.; Vecka, M.; Jáchymová, M.; Macášek, J.; Hrabák, P.; Zeman, M.; Vávrová, L.; Řoupal, J.; Krechler, T.; Ák, A. Osteopontin as a Discriminating Marker for Pancreatic Cancer and Chronic Pancreatitis. Cancer Biomark. Sect. Dis. Markers 2016, 17, 55–65. [Google Scholar] [CrossRef] [PubMed]
- Hahn, S.A.; Schutte, M.; Hoque, A.T.; Moskaluk, C.A.; da Costa, L.T.; Rozenblum, E.; Weinstein, C.L.; Fischer, A.; Yeo, C.J.; Hruban, R.H.; et al. DPC4, a Candidate Tumor Suppressor Gene at Human Chromosome 18q21.1. Science 1996, 271, 350–353. [Google Scholar] [CrossRef]
- Tascilar, M.; Skinner, H.G.; Rosty, C.; Sohn, T.; Wilentz, R.E.; Offerhaus, G.J.A.; Adsay, V.; Abrams, R.A.; Cameron, J.L.; Kern, S.E.; et al. The SMAD4 Protein and Prognosis of Pancreatic Ductal Adenocarcinoma. Clin. Cancer Res. 2001, 7, 4115–4121. [Google Scholar] [PubMed]
- Wang, Z.; Li, Y.; Zhan, S.; Zhang, L.; Zhang, S.; Tang, Q.; Li, M.; Tan, Z.; Liu, S.; Xing, X. SMAD4 Y353C Promotes the Progression of PDAC. BMC Cancer 2019, 19, 1037. [Google Scholar] [CrossRef] [PubMed]
- Bazzichetto, C.; Conciatori, F.; Luchini, C.; Simionato, F.; Santoro, R.; Vaccaro, V.; Corbo, V.; Falcone, I.; Ferretti, G.; Cognetti, F.; et al. From Genetic Alterations to Tumor Microenvironment: The Ariadne’s String in Pancreatic Cancer. Cells 2020, 9, 309. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, S. Molecular Pathogenesis and Targeted Therapy of Pancreatic Cancer. Ann. Surg. Oncol. 2016, 23 (Suppl. 2), S197–S205. [Google Scholar] [CrossRef] [PubMed]
- Bardeesy, N.; Cheng, K.-H.; Berger, J.H.; Chu, G.C.; Pahler, J.; Olson, P.; Hezel, A.F.; Horner, J.; Lauwers, G.Y.; Hanahan, D.; et al. Smad4 Is Dispensable for Normal Pancreas Development yet Critical in Progression and Tumor Biology of Pancreas Cancer. Genes Dev. 2006, 20, 3130–3146. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, A.J.; Chetty, R. Smad4/DPC4. J. Clin. Pathol. 2018, 71, 661–664. [Google Scholar] [CrossRef] [PubMed]
- Wilentz, R.E.; Iacobuzio-Donahue, C.A.; Argani, P.; McCarthy, D.M.; Parsons, J.L.; Yeo, C.J.; Kern, S.E.; Hruban, R.H. Loss of Expression of Dpc4 in Pancreatic Intraepithelial Neoplasia: Evidence That DPC4 Inactivation Occurs Late in Neoplastic Progression. Cancer Res. 2000, 60, 2002–2006. [Google Scholar]
- Hsieh, Y.-Y.; Liu, T.-P.; Chou, C.-J.; Chen, H.-Y.; Lee, K.-H.; Yang, P.-M. Integration of Bioinformatics Resources Reveals the Therapeutic Benefits of Gemcitabine and Cell Cycle Intervention in SMAD4-Deleted Pancreatic Ductal Adenocarcinoma. Genes 2019, 10, 766. [Google Scholar] [CrossRef]
- Shugang, X.; Hongfa, Y.; Jianpeng, L.; Xu, Z.; Jingqi, F.; Xiangxiang, L.; Wei, L. Prognostic Value of SMAD4 in Pancreatic Cancer: A Meta-Analysis. Transl. Oncol. 2016, 9, 1–7. [Google Scholar] [CrossRef]
- Wang, F.; Xia, X.; Yang, C.; Shen, J.; Mai, J.; Kim, H.-C.; Kirui, D.; Kang, Y.; Fleming, J.B.; Koay, E.J.; et al. SMAD4 Gene Mutation Renders Pancreatic Cancer Resistance to Radiotherapy through Promotion of Autophagy. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2018, 24, 3176–3185. [Google Scholar] [CrossRef]
- Schlick, K.; Magnes, T.; Huemer, F.; Ratzinger, L.; Weiss, L.; Pichler, M.; Melchardt, T.; Greil, R.; Egle, A. C-Reactive Protein and Neutrophil/Lymphocytes Ratio: Prognostic Indicator for Doubling Overall Survival Prediction in Pancreatic Cancer Patients. J. Clin. Med. 2019, 8, 1791. [Google Scholar] [CrossRef] [PubMed]
- Hoshimoto, S.; Hishinuma, S.; Shirakawa, H.; Tomikawa, M.; Ozawa, I.; Ogata, Y. Validation and Clinical Usefulness of Pre- and Postoperative Systemic Inflammatory Parameters as Prognostic Markers in Patients with Potentially Resectable Pancreatic Cancer. Pancreatol. Off. J. Int. Assoc. Pancreatol. IAP Al 2020, 20, 239–246. [Google Scholar] [CrossRef] [PubMed]
- Giakoustidis, A.; Neofytou, K.; Costa Neves, M.; Giakoustidis, D.; Louri, E.; Cunningham, D.; Mudan, S. Identifying the Role of Neutrophil-to-Lymphocyte Ratio and Platelets-to-Lymphocyte Ratio as Prognostic Markers in Patients Undergoing Resection of Pancreatic Ductal Adenocarcinoma. Ann. Hepato-Biliary-Pancreat. Surg. 2018, 22, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Recio-Boiles, A.; Nallagangula, A.; Veeravelli, S.; Vondrak, J.; Saboda, K.; Roe, D.; Elquza, E.; McBride, A.; Babiker, H.M. Neutrophil-to-Lymphocyte and Platelet-to-Lymphocyte Ratios Inversely Correlate to Clinical and Pathologic Stage in Patients with Resectable Pancreatic Ductal Adenocarcinoma. Ann. Pancreat. Cancer 2019, 2. [Google Scholar] [CrossRef]
- Sakamoto, T.; Saito, H.; Uchinaka, E.I.; Morimoto, M.; Amisaki, M.; Tokuyasu, N.; Honjo, S.; Ashida, K.; Fujiwara, Y. The Combination of Neutrophil-to-Lymphocyte Ratio and Serum Carbohydrate Antigen 19-9 Level as a Prognostic Indicator in Patients with Recurrent Pancreatic Cancer. Anticancer Res. 2018, 38, 5497–5503. [Google Scholar] [CrossRef]
- Fan, Z.; Luo, G.; Gong, Y.; Xu, H.; Qian, Y.; Deng, S.; Huang, Q.; Yang, C.; Cheng, H.; Jin, K.; et al. Prognostic Value of the C-Reactive Protein/Lymphocyte Ratio in Pancreatic Cancer. Ann. Surg. Oncol. 2020. [Google Scholar] [CrossRef]
- Onoe, S.; Maeda, A.; Takayama, Y.; Fukami, Y.; Takahashi, T.; Uji, M.; Kaneoka, Y. The Prognostic Impact of the Lymphocyte-to-Monocyte Ratio in Resected Pancreatic Head Adenocarcinoma. Med. Princ. Pract. Int. J. Kuwait Univ. Health Sci. Cent. 2019, 28, 517–525. [Google Scholar] [CrossRef] [PubMed]
- Greenhalf, W.; Ghaneh, P.; Neoptolemos, J.P.; Palmer, D.H.; Cox, T.F.; Lamb, R.F.; Garner, E.; Campbell, F.; Mackey, J.R.; Costello, E.; et al. Pancreatic Cancer HENT1 Expression and Survival from Gemcitabine in Patients from the ESPAC-3 Trial. J. Natl. Cancer Inst. 2014, 106, djt347. [Google Scholar] [CrossRef] [PubMed]
- Aoyama, T.; Kazama, K.; Miyagi, Y.; Murakawa, M.; Yamaoku, K.; Atsumi, Y.; Shiozawa, M.; Ueno, M.; Morimoto, M.; Oshima, T.; et al. Predictive Role of Human Equilibrative Nucleoside Transporter 1 in Patients with Pancreatic Cancer Treated by Curative Resection and Gemcitabine-Only Adjuvant Chemotherapy. Oncol. Lett. 2017, 14, 599–606. [Google Scholar] [CrossRef] [PubMed]
- Bird, N.T.E.; Elmasry, M.; Jones, R.; Psarelli, E.; Dodd, J.; Malik, H.; Greenhalf, W.; Kitteringham, N.; Ghaneh, P.; Neoptolemos, J.P.; et al. Immunohistochemical HENT1 Expression as a Prognostic Biomarker in Patients with Resected Pancreatic Ductal Adenocarcinoma Undergoing Adjuvant Gemcitabine-Based Chemotherapy. Br. J. Surg. 2017, 104, 328–336. [Google Scholar] [CrossRef] [PubMed]
- Chhetri, P.; Giri, A.; Shakya, S.; Shakya, S.; Sapkota, B.; Pramod, K. Current Development of Anti-Cancer Drug S-1. J. Clin. Diagn. Res. JCDR 2016, 10, XE01–XE05. [Google Scholar] [CrossRef] [PubMed]
- Okamura, Y.; Yasukawa, S.; Narimatsu, H.; Boku, N.; Fukutomi, A.; Konishi, M.; Morinaga, S.; Toyama, H.; Kaneoka, Y.; Shimizu, Y.; et al. Human Equilibrative Nucleoside Transporter-1 Expression Is a Predictor in Patients with Resected Pancreatic Cancer Treated with Adjuvant S-1 Chemotherapy. Cancer Sci. 2020, 111, 548–560. [Google Scholar] [CrossRef] [PubMed]
- Xi, Y.; Yuan, P.; Li, T.; Zhang, M.; Liu, M.-F.; Li, B. HENT1 Reverses Chemoresistance by Regulating Glycolysis in Pancreatic Cancer. Cancer Lett. 2020. [Google Scholar] [CrossRef]
- Triantafyllou, E.-A.; Georgatsou, E.; Mylonis, I.; Simos, G.; Paraskeva, E. Expression of AGPAT2, an Enzyme Involved in the Glycerophospholipid/Triacylglycerol Biosynthesis Pathway, Is Directly Regulated by HIF-1 and Promotes Survival and Etoposide Resistance of Cancer Cells under Hypoxia. Biochim. Biophys. Acta BBA Mol. Cell Biol. Lipids 2018, 1863, 1142–1152. [Google Scholar] [CrossRef] [PubMed]
- Sierzega, M.; Pach, R.; Kulig, P.; Legutko, J.; Kulig, J. Prognostic Implications of Expression Profiling for Gemcitabine-Related Genes (HENT1, DCK, RRM1, RRM2) in Patients With Resectable Pancreatic Adenocarcinoma Receiving Adjuvant Chemotherapy. Pancreas 2017, 46, 684–689. [Google Scholar] [CrossRef]
- Raffenne, J.; Nicolle, R.; Puleo, F.; Le Corre, D.; Boyez, C.; Marechal, R.; Emile, J.F.; Demetter, P.; Bardier, A.; Laurent-Puig, P.; et al. HENT1 Testing in Pancreatic Ductal Adenocarcinoma: Are We Ready? A Multimodal Evaluation of HENT1 Status. Cancers 2019, 11, 1808. [Google Scholar] [CrossRef]
- Bhutia, Y.D.; Hung, S.W.; Patel, B.; Lovin, D.; Govindarajan, R. CNT1 Expression Influences Proliferation and Chemosensitivity in Drug-Resistant Pancreatic Cancer Cells. Cancer Res. 2011, 71, 1825–1835. [Google Scholar] [CrossRef]
- Paproski, R.J.; Yao, S.Y.M.; Favis, N.; Evans, D.; Young, J.D.; Cass, C.E.; Zemp, R.J. Human Concentrative Nucleoside Transporter 3 Transfection with Ultrasound and Microbubbles in Nucleoside Transport Deficient HEK293 Cells Greatly Increases Gemcitabine Uptake. PLoS ONE 2013, 8, e56423. [Google Scholar] [CrossRef][Green Version]
- Hesler, R.A.; Huang, J.J.; Starr, M.D.; Treboschi, V.M.; Bernanke, A.G.; Nixon, A.B.; McCall, S.J.; White, R.R.; Blobe, G.C. TGF-β-Induced Stromal CYR61 Promotes Resistance to Gemcitabine in Pancreatic Ductal Adenocarcinoma through Downregulation of the Nucleoside Transporters HENT1 and HCNT3. Carcinogenesis 2016, 37, 1041–1051. [Google Scholar] [CrossRef]
- Wang, C.; Buolamwini, J.K. A Novel RNA Variant of Human Concentrative Nucleoside Transporter 1 (HCNT1) That Is a Potential Cancer Biomarker. Exp. Hematol. Oncol. 2019, 8, 18. [Google Scholar] [CrossRef]
- Skrypek, N.; Vasseur, R.; Vincent, A.; Duchêne, B.; Van Seuningen, I.; Jonckheere, N. The Oncogenic Receptor ErbB2 Modulates Gemcitabine and Irinotecan/SN-38 Chemoresistance of Human Pancreatic Cancer Cells via HCNT1 Transporter and Multidrug-Resistance Associated Protein MRP-2. Oncotarget 2015, 6, 10853–10867. [Google Scholar] [CrossRef] [PubMed]
- Skrypek, N.; Duchêne, B.; Hebbar, M.; Leteurtre, E.; van Seuningen, I.; Jonckheere, N. The MUC4 Mucin Mediates Gemcitabine Resistance of Human Pancreatic Cancer Cells via the Concentrative Nucleoside Transporter Family. Oncogene 2013, 32, 1714–1723. [Google Scholar] [CrossRef] [PubMed]
- Maréchal, R.; Mackey, J.R.; Lai, R.; Demetter, P.; Peeters, M.; Polus, M.; Cass, C.E.; Young, J.; Salmon, I.; Devière, J.; et al. Human Equilibrative Nucleoside Transporter 1 and Human Concentrative Nucleoside Transporter 3 Predict Survival after Adjuvant Gemcitabine Therapy in Resected Pancreatic Adenocarcinoma. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2009, 15, 2913–2919. [Google Scholar] [CrossRef] [PubMed]
- Lowery, M.A.; Kelsen, D.P.; Stadler, Z.K.; Yu, K.H.; Janjigian, Y.Y.; Ludwig, E.; D’Adamo, D.R.; Salo-Mullen, E.; Robson, M.E.; Allen, P.J.; et al. An Emerging Entity: Pancreatic Adenocarcinoma Associated with a Known BRCA Mutation: Clinical Descriptors, Treatment Implications, and Future Directions. Oncologist 2011, 16, 1397–1402. [Google Scholar] [CrossRef]
- Goggins, M.; Hruban, R.H.; Kern, S.E. BRCA2 Is Inactivated Late in the Development of Pancreatic Intraepithelial Neoplasia: Evidence and Implications. Am. J. Pathol. 2000, 156, 1767–1771. [Google Scholar] [CrossRef]
- Rowley, M.; Ohashi, A.; Mondal, G.; Mills, L.; Yang, L.; Zhang, L.; Sundsbak, R.; Shapiro, V.; Muders, M.H.; Smyrk, T.; et al. Inactivation of Brca2 Promotes Trp53-Associated but Inhibits KrasG12D-Dependent Pancreatic Cancer Development in Mice. Gastroenterology 2011, 140, 1303–1313.e3. [Google Scholar] [CrossRef] [PubMed]
- Skoulidis, F.; Cassidy, L.D.; Pisupati, V.; Jonasson, J.G.; Bjarnason, H.; Eyfjord, J.E.; Karreth, F.A.; Lim, M.; Barber, L.M.; Clatworthy, S.A.; et al. Germline Brca2 Heterozygosity Promotes Kras(G12D) -Driven Carcinogenesis in a Murine Model of Familial Pancreatic Cancer. Cancer Cell 2010, 18, 499–509. [Google Scholar] [CrossRef] [PubMed]
- Moynahan, M.E.; Jasin, M. Mitotic Homologous Recombination Maintains Genomic Stability and Suppresses Tumorigenesis. Nat. Rev. Mol. Cell Biol. 2010, 11, 196–207. [Google Scholar] [CrossRef] [PubMed]
- Grant, R.C.; Denroche, R.E.; Borgida, A.; Virtanen, C.; Cook, N.; Smith, A.L.; Connor, A.A.; Wilson, J.M.; Peterson, G.; Roberts, N.J.; et al. Exome-Wide Association Study of Pancreatic Cancer Risk. Gastroenterology 2018, 154, 719–722.e3. [Google Scholar] [CrossRef]
- Brose, M.S.; Rebbeck, T.R.; Calzone, K.A.; Stopfer, J.E.; Nathanson, K.L.; Weber, B.L. Cancer Risk Estimates for BRCA1 Mutation Carriers Identified in a Risk Evaluation Program. J. Natl. Cancer Inst. 2002, 94, 1365–1372. [Google Scholar] [CrossRef]
- Farmer, H.; McCabe, N.; Lord, C.J.; Tutt, A.N.J.; Johnson, D.A.; Richardson, T.B.; Santarosa, M.; Dillon, K.J.; Hickson, I.; Knights, C.; et al. Targeting the DNA Repair Defect in BRCA Mutant Cells as a Therapeutic Strategy. Nature 2005, 434, 917–921. [Google Scholar] [CrossRef]
- Golan, T.; Hammel, P.; Reni, M.; Van Cutsem, E.; Macarulla, T.; Hall, M.J.; Park, J.-O.; Hochhauser, D.; Arnold, D.; Oh, D.-Y.; et al. Maintenance Olaparib for Germline BRCA-Mutated Metastatic Pancreatic Cancer. N. Engl. J. Med. 2019, 381, 317–327. [Google Scholar] [CrossRef]
- Wattenberg, M.M.; Asch, D.; Yu, S.; O’Dwyer, P.J.; Domchek, S.M.; Nathanson, K.L.; Rosen, M.A.; Beatty, G.L.; Siegelman, E.S.; Reiss, K.A. Platinum Response Characteristics of Patients with Pancreatic Ductal Adenocarcinoma and a Germline BRCA1, BRCA2 or PALB2 Mutation. Br. J. Cancer 2020, 122, 333–339. [Google Scholar] [CrossRef]
- Golan, T.; Barenboim, A.; Lahat, G.; Nachmany, I.; Goykhman, Y.; Shacham-Shmueli, E.; Halpern, N.; Brazowski, E.; Geva, R.; Wolf, I.; et al. Increased Rate of Complete Pathologic Response After Neoadjuvant FOLFIRINOX for BRCA Mutation Carriers with Borderline Resectable Pancreatic Cancer. Ann. Surg. Oncol. 2020. [Google Scholar] [CrossRef]
- Ballehaninna, U.K.; Chamberlain, R.S. The Clinical Utility of Serum CA 19-9 in the Diagnosis, Prognosis and Management of Pancreatic Adenocarcinoma: An Evidence Based Appraisal. J. Gastrointest. Oncol. 2012, 3, 15. [Google Scholar]
- Tao, J.; Yang, G.; Zhou, W.; Qiu, J.; Chen, G.; Luo, W.; Zhao, F.; You, L.; Zheng, L.; Zhang, T.; et al. Targeting Hypoxic Tumor Microenvironment in Pancreatic Cancer. J. Hematol. Oncol. Hematol. Oncol. 2021, 14, 14. [Google Scholar] [CrossRef]
- Geismann, C.; Arlt, A. Coming in the Air: Hypoxia Meets Epigenetics in Pancreatic Cancer. Cells 2020, 9, 2353. [Google Scholar] [CrossRef]
- Ebrahimi, S.; Hosseini, M.; Shahidsales, S.; Maftouh, M.; Ferns, G.A.; Ghayour-Mobarhan, M.; Hassanian, S.M.; Avan, A. Targeting the Akt/PI3K Signaling Pathway as a Potential Therapeutic Strategy for the Treatment of Pancreatic Cancer. Curr. Med. Chem. 2017, 24, 1321–1331. [Google Scholar] [CrossRef]
- Zhang, Z.; Fang, E.; Rong, Y.; Han, H.; Gong, Q.; Xiao, Y.; Li, H.; Mei, P.; Li, H.; Zhu, Z.; et al. Hypoxia-Induced LncRNA CASC9 Enhances Glycolysis and the Epithelial-Mesenchymal Transition of Pancreatic Cancer by a Positive Feedback Loop with AKT/HIF-1α Signaling. Am. J. Cancer Res. 2021, 11, 123–137. [Google Scholar]
- Tan, Z.; Xu, J.; Zhang, B.; Shi, S.; Yu, X.; Liang, C. Hypoxia: A Barricade to Conquer the Pancreatic Cancer. Cell. Mol. Life Sci. CMLS 2020, 77, 3077–3083. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Peng, C.; Zhu, C.; Nie, S.; Qian, X.; Shi, Z.; Shi, M.; Liang, Y.; Ding, X.; Zhang, S.; et al. Hypoxia Promotes the Metastasis of Pancreatic Cancer through Regulating NOX4/KDM5A-Mediated Histone Methylation Modification Changes in a HIF1A-Independent Manner. Clin. Epigenetics 2021, 13, 18. [Google Scholar] [CrossRef] [PubMed]
- Deutsch, O.; Haviv, Y.; Krief, G.; Keshet, N.; Westreich, R.; Stemmer, S.M.; Zaks, B.; Navat, S.P.; Yanko, R.; Lahav, O.; et al. Possible Proteomic Biomarkers for the Detection of Pancreatic Cancer in Oral Fluids. Sci. Rep. 2020, 10, 21995. [Google Scholar] [CrossRef] [PubMed]
- Ansari, D.; Torén, W.; Zhou, Q.; Hu, D.; Andersson, R. Proteomic and Genomic Profiling of Pancreatic Cancer. Cell Biol. Toxicol. 2019, 35, 333–343. [Google Scholar] [CrossRef] [PubMed]
- Pan, S.; Chen, R.; Tamura, Y.; Crispin, D.A.; Lai, L.A.; May, D.H.; McIntosh, M.W.; Goodlett, D.R.; Brentnall, T.A. Quantitative Glycoproteomics Analysis Reveals Changes in N-Glycosylation Level Associated with Pancreatic Ductal Adenocarcinoma. J. Proteome Res. 2014, 13, 1293–1306. [Google Scholar] [CrossRef]
- Bauden, M.; Kristl, T.; Sasor, A.; Andersson, B.; Marko-Varga, G.; Andersson, R.; Ansari, D. Histone Profiling Reveals the H1.3 Histone Variant as a Prognostic Biomarker for Pancreatic Ductal Adenocarcinoma. BMC Cancer 2017, 17, 810. [Google Scholar] [CrossRef]
- Grønborg, M.; Kristiansen, T.Z.; Iwahori, A.; Chang, R.; Reddy, R.; Sato, N.; Molina, H.; Jensen, O.N.; Hruban, R.H.; Goggins, M.G.; et al. Biomarker Discovery from Pancreatic Cancer Secretome Using a Differential Proteomic Approach. Mol. Cell. Proteom. MCP 2006, 5, 157–171. [Google Scholar] [CrossRef] [PubMed]
- Cintas, C.; Douché, T.; Therville, N.; Arcucci, S.; Ramos-Delgado, F.; Basset, C.; Thibault, B.; Guillermet-Guibert, J. Signal-Targeted Therapies and Resistance Mechanisms in Pancreatic Cancer: Future Developments Reside in Proteomics. Cancers 2018, 10, 174. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, Y.; Takadate, T.; Mizuma, M.; Shima, H.; Suzuki, T.; Tachibana, T.; Shimura, M.; Hata, T.; Iseki, M.; Kawaguchi, K.; et al. Stromal Expression of Hemopexin Is Associated with Lymph-Node Metastasis in Pancreatic Ductal Adenocarcinoma. PLoS ONE 2020, 15, e0235904. [Google Scholar] [CrossRef]
- Allgöwer, C.; Kretz, A.-L.; von Karstedt, S.; Wittau, M.; Henne-Bruns, D.; Lemke, J. Friend or Foe: S100 Proteins in Cancer. Cancers 2020, 12, 2037. [Google Scholar] [CrossRef] [PubMed]
- Pan, S.; Brentnall, T.A.; Chen, R. Proteomics Analysis of Bodily Fluids in Pancreatic Cancer. Proteomics 2015, 15, 2705–2715. [Google Scholar] [CrossRef] [PubMed]
- Brar, G.; Blais, E.M.; Joseph Bender, R.; Brody, J.R.; Sohal, D.; Madhavan, S.; Picozzi, V.J.; Hendifar, A.E.; Chung, V.M.; Halverson, D.; et al. Multi-Omic Molecular Comparison of Primary versus Metastatic Pancreatic Tumours. Br. J. Cancer 2019, 121, 264–270. [Google Scholar] [CrossRef] [PubMed]
- Moris, D.; Damaskos, C.; Spartalis, E.; Papalampros, A.; Vernadakis, S.; Dimitroulis, D.; Griniatsos, J.; Felekouras, E.; Nikiteas, N. Updates and Critical Evaluation on Novel Biomarkers for the Malignant Progression of Intraductal Papillary Mucinous Neoplasms of the Pancreas. Anticancer Res. 2017, 37, 2185–2194. [Google Scholar] [CrossRef]
- Moris, D.; Tsaroucha, A.K.; Dimitrokallis, N.; Papanikolaou, I.S.; Schizas, D. Biomarkers in the Differential Diagnosis of Pancreatic Diseases: Looking for a Compass. Pancreas 2017, 46, e38–e39. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.-H.; Wang, W.-Q.; Han, X.; Gao, H.-L.; Li, T.-J.; Xu, S.-S.; Li, S.; Xu, H.-X.; Li, H.; Ye, L.-Y.; et al. Advances on Diagnostic Biomarkers of Pancreatic Ductal Adenocarcinoma: A Systems Biology Perspective. Comput. Struct. Biotechnol. J. 2020, 18, 3606–3614. [Google Scholar] [CrossRef] [PubMed]
| Biomarker | Diagnostic | Prognostic | Predictive | Clinical Specimen | References |
|---|---|---|---|---|---|
| CA 19-9 | X | X | Serum | [9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24] | |
| microRNAs | X | X | Pancreatic tissue, pancreatic fluid, and urine | [25,26,27,28,29] | |
| DNA methylation | X | X | Pancreatic fluid, endoscopic brush samples, and serum (cfDNA) | [30,31,32,33,34,35,36] | |
| Mismatch repair genes/microsatellite instability | X | X | Pancreatic tissue | [37,38,39,40,41,42,43,44,45,46,47] | |
| KRAS | X | X | Pancreatic tissue and serum (cfDNA) | [48,49,50,51,52,53,54,55,56,57] | |
| Exosomes | X | X | Serum and saliva | [58,59,60,61,62,63,64,65,66,67,68,69,70,71,72] | |
| Circulating tumor cells | X | X | Peripheral blood | [73,74,75,76,77] | |
| PAM4/MUC5AC | X | Pancreatic tissue | [78,79,80,81] | ||
| Osteopontin | X | X | Serum | [82,83,84,85] | |
| SMAD4/DPC4 | X | X | X | Pancreatic tissue | [86,87,88,89,90,91,92,93,94,95,96] |
| Immune response and inflammatory markers | X | X | Serum and blood | [97,98,99,100,101,102,103] | |
| hENT1 | X | X | Pancreatic tissue | [104,105,106,107,108,109,110,111,112] | |
| hCNT1/hCNT3 | X | X | Pancreatic tissue | [113,114,115,116,117,118,119] | |
| BRCA1/BRCA2 | X | X | Pancreatic tissue | [120,121,122,123,124,125,126,127,128,129,130] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Giannis, D.; Moris, D.; Barbas, A.S. Diagnostic, Predictive and Prognostic Molecular Biomarkers in Pancreatic Cancer: An Overview for Clinicians. Cancers 2021, 13, 1071. https://doi.org/10.3390/cancers13051071
Giannis D, Moris D, Barbas AS. Diagnostic, Predictive and Prognostic Molecular Biomarkers in Pancreatic Cancer: An Overview for Clinicians. Cancers. 2021; 13(5):1071. https://doi.org/10.3390/cancers13051071
Chicago/Turabian StyleGiannis, Dimitrios, Dimitrios Moris, and Andrew S. Barbas. 2021. "Diagnostic, Predictive and Prognostic Molecular Biomarkers in Pancreatic Cancer: An Overview for Clinicians" Cancers 13, no. 5: 1071. https://doi.org/10.3390/cancers13051071
APA StyleGiannis, D., Moris, D., & Barbas, A. S. (2021). Diagnostic, Predictive and Prognostic Molecular Biomarkers in Pancreatic Cancer: An Overview for Clinicians. Cancers, 13(5), 1071. https://doi.org/10.3390/cancers13051071







