Systemic Inflammation and Tumour-Infiltrating T-Cell Receptor Repertoire Diversity Are Predictive of Clinical Outcome in High-Grade B-Cell Lymphoma with MYC and BCL2 and/or BCL6 Rearrangements
Abstract
Simple Summary
Abstract
1. Introduction
2. Results
2.1. Baseline Clinical Characteristics
2.2. Laboratory Assessment
2.3. Treatment Modalities
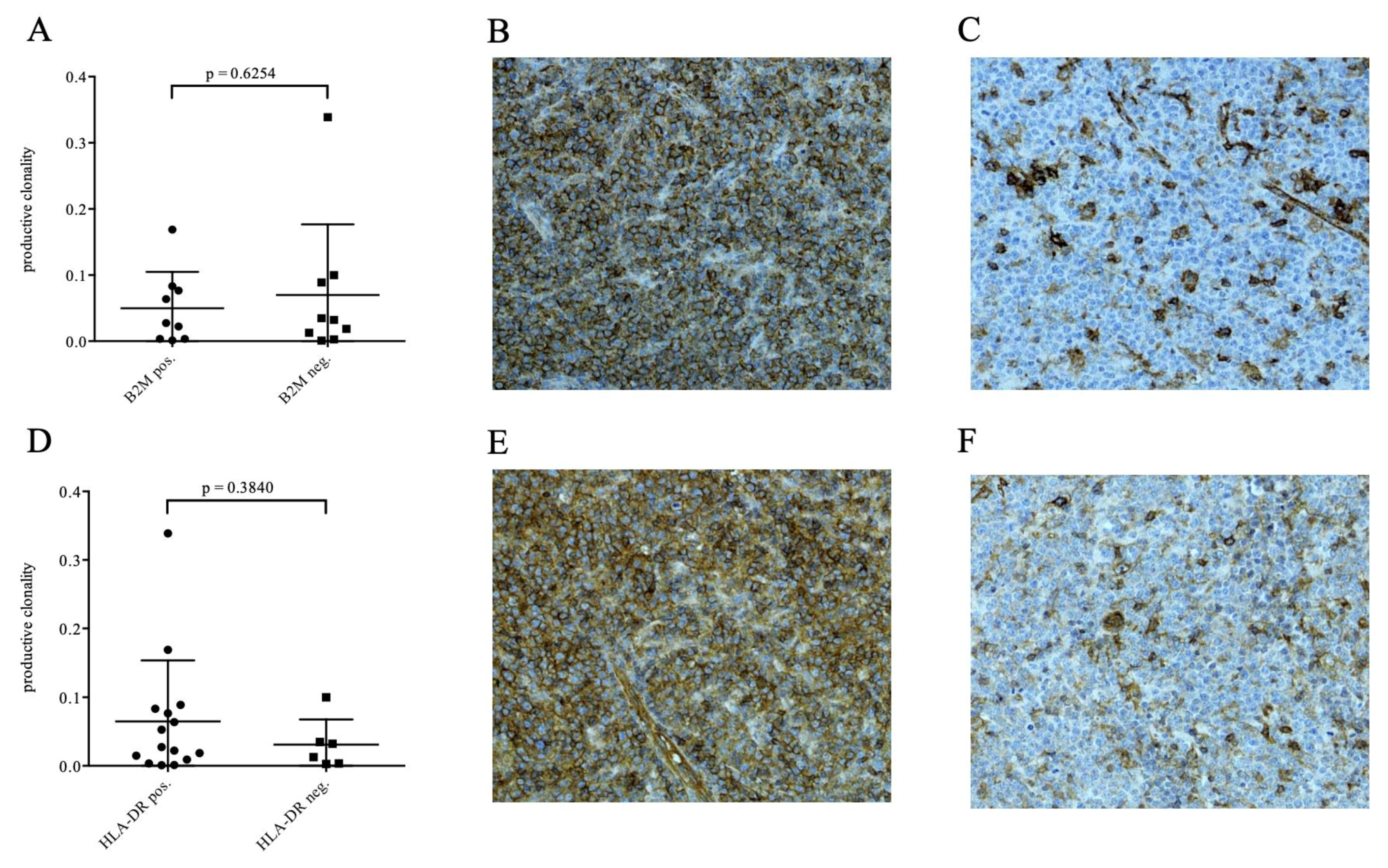
2.4. Histopathological Assessment, Immunophenotyping and FisH
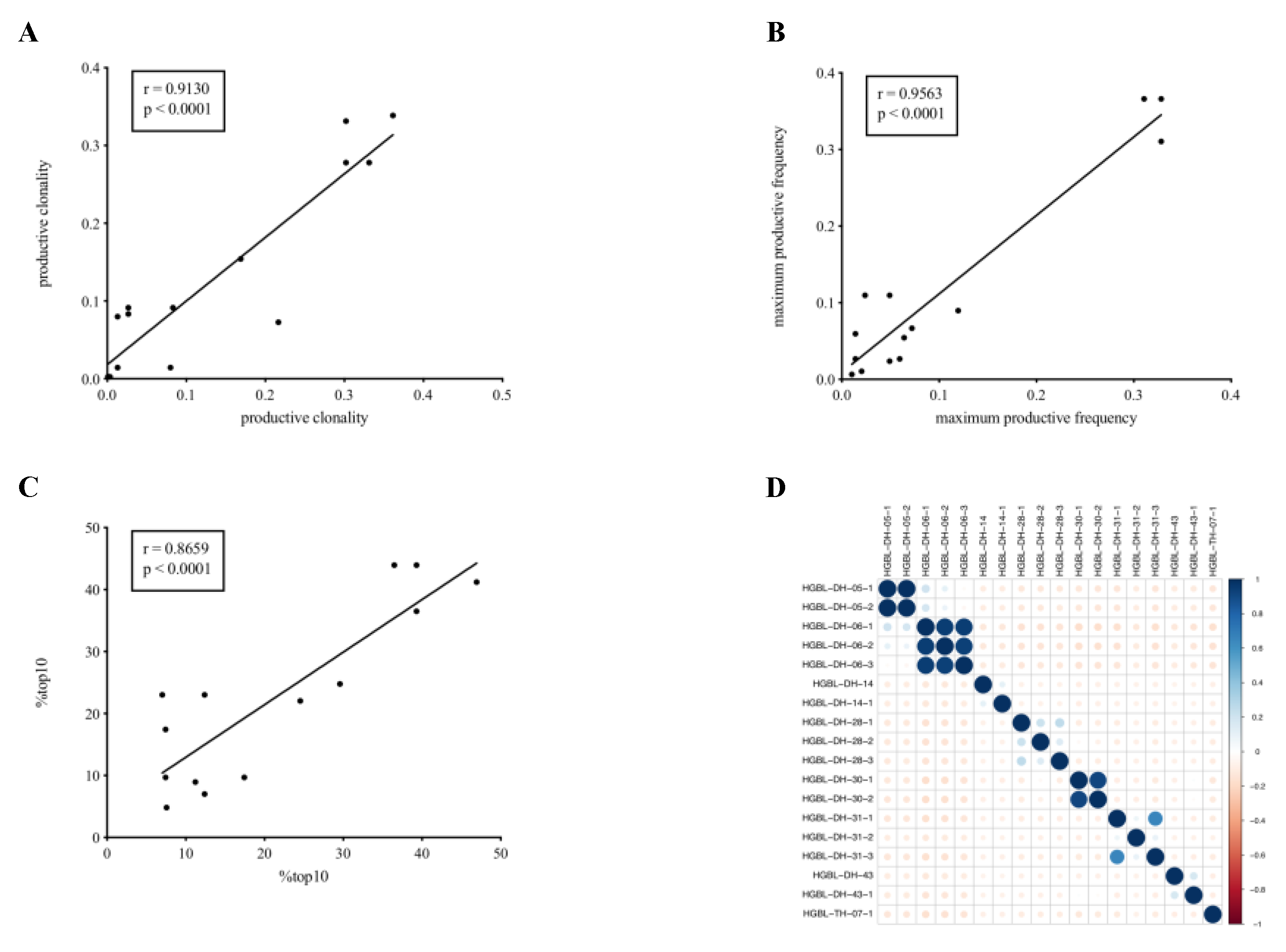
2.5. High Throughput TCR Sequencing
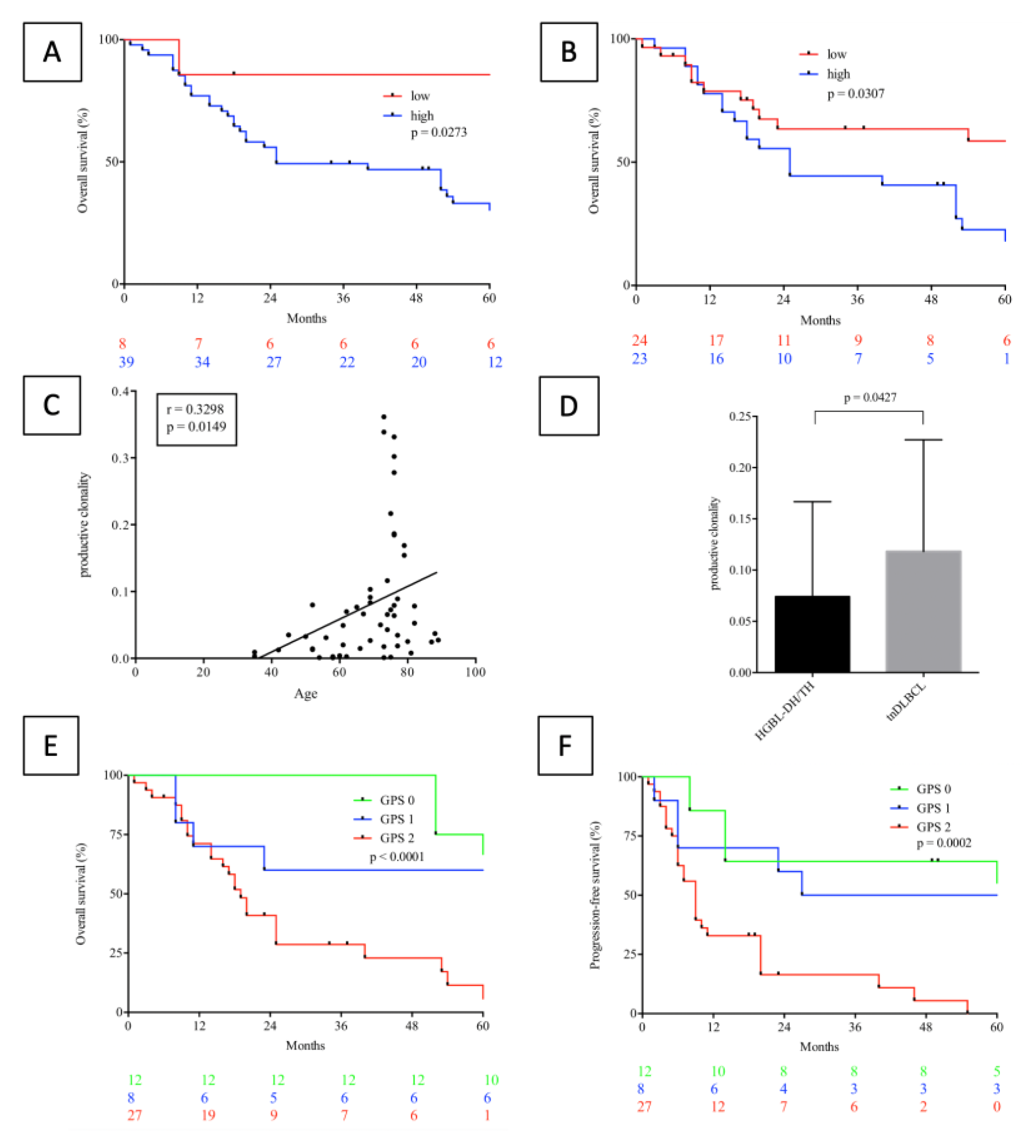
2.6. Comparison of the TCR Repertoire between HGBL and DLBCL
2.7. The Impact of TCR-Diversity and Inflammation on Survival Outcome
3. Discussion
4. Materials and Methods
4.1. Samples
4.2. Clinical Assessment
4.3. Immunophenotyping, Tumor Infiltrating T-Cell Quantification and FisH
4.4. High-Throughput Sequencing of the CDR3 Region of the TCRβ Chain
4.5. TCR-Sequencing Data Analysis
4.6. Comparative Analysis with DLBCL
4.7. Statistical Analysis
4.8. Ethics Statement
4.9. Availability of Data and Materials
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Swerdlow, S.H.; Campo, E.; Pileri, S.A.; Harris, N.L.; Stein, H.; Siebert, R.; Advani, R.; Ghielmini, M.; Salles, G.A.; Zelenetz, A.D.; et al. The 2016 Revision of the World Health Organization Classification of Lymphoid Neoplasms. Blood 2016, 127, 2375–2390. [Google Scholar] [CrossRef]
- Perry, A.M.; Crockett, D.; Dave, B.J.; Althof, P.; Winkler, L.; Smith, L.M.; Aoun, P.; Chan, W.C.; Fu, K.; Greiner, T.C.; et al. B-Cell Lymphoma, Unclassifiable, with Features Intermediate between Diffuse Large B-Cell Lymphoma and Burkitt Lymphoma: Study of 39 Cases. Br. J. Haematol. 2013, 162, 40–49. [Google Scholar] [CrossRef]
- Rosenwald, A.; Bens, S.; Advani, R.; Barrans, S.; Copie-Bergman, C.; Elsensohn, M.H.; Natkunam, Y.; Calaminici, M.; Sander, B.; Baia, M.; et al. Prognostic Significance of Myc Rearrangement and Translocation Partner in Diffuse Large B-Cell Lymphoma: A Study by the Lunenburg Lymphoma Biomarker Consortium. J. Clin. Oncol. 2019, 37, 3359–3368. [Google Scholar] [CrossRef] [PubMed]
- Landsburg, D.J.; Falkiewicz, M.K.; Maly, J.; Blum, K.A.; Howlett, C.; Feldman, T.; Mato, A.R.; Hill, B.T.; Li, S.; Medeiros, L.J.; et al. Outcomes of Patients with Double-Hit Lymphoma Who Achieve First Complete Remission. J. Clin. Oncol. 2017, 35, 2260–2267. [Google Scholar] [CrossRef] [PubMed]
- Keane, C.; Vari, F.; Hertzberg, M.; Cao, K.A.; Green, M.R.; Han, E.; Seymour, J.F.; Hicks, R.J.; Gill, D.; Crooks, P.; et al. Ratios of T-Cell Immune Effectors and Checkpoint Molecules as Prognostic Biomarkers in Diffuse Large B-Cell Lymphoma: A Population-Based Study. Lancet Haematol. 2015, 2, e445–e455. [Google Scholar] [CrossRef]
- Ansell, S.M.; Stenson, M.; Habermann, T.M.; Jelinek, D.F.; Witzig, T.E. Cd4+ T-Cell Immune Response to Large B-Cell Non-Hodgkin’s Lymphoma Predicts Patient Outcome. J. Clin. Oncol. 2001, 19, 720–726. [Google Scholar] [CrossRef]
- Gooden, M.J.; de Bock, G.H.; Leffers, N.; Daemen, T.; Nijman, H.W. The Prognostic Influence of Tumour-Infiltrating Lymphocytes in Cancer: A Systematic Review with Meta-Analysis. Br. J. Cancer 2011, 105, 93–103. [Google Scholar] [CrossRef]
- Nikolich-Zugich, J.; Slifka, M.K.; Messaoudi, I. The Many Important Facets of T-Cell Repertoire Diversity. Nat. Rev. Immunol. 2004, 4, 123–132. [Google Scholar] [CrossRef]
- Messaoudi, I.; Guevara Patino, J.A.; Dyall, R.; LeMaoult, J.; Nikolich-Zugich, J. Direct Link between Mhc Polymorphism, T Cell Avidity, and Diversity in Immune Defense. Science 2002, 298, 1797–1800. [Google Scholar] [CrossRef]
- Rieken, J.; Bernard, V.; Witte, H.M.; Peter, W.; Merz, H.; Olschewski, V.; Hertel, L.; Lehnert, H.; Biersack, H.; von Bubnoff, N.; et al. Exhaustion of tumour-infiltrating T-cell receptor repertoire diversity is an age-dependent indicator of immunological fitness independently predictive of clinical outcome in Burkitt lymphoma. Br. J. Haematol. 2020. [Google Scholar] [CrossRef]
- Keane, C.; Gould, C.; Jones, K.; Hamm, D.; Talaulikar, D.; Ellis, J.; Vari, F.; Birch, S.; Han, E.; Wood, P.; et al. The T-Cell Receptor Repertoire Influences the Tumor Microenvironment and Is Associated with Survival in Aggressive B-Cell Lymphoma. Clin. Cancer Res. 2017, 23, 1820–1828. [Google Scholar] [CrossRef]
- Li, X.; Zhang, Y.; Zhao, W.; Liu, Z.; Shen, Y.; Li, J.; Shen, Z. The Glasgow Prognostic Score as a Significant Predictor of Diffuse Large B Cell Lymphoma Treated with R-Chop in China. Ann. Hematol. 2015, 94, 57–63. [Google Scholar] [CrossRef]
- Hans, C.P.; Weisenburger, D.D.; Greiner, T.C.; Gascoyne, R.D.; Delabie, J.; Ott, G.; Muller-Hermelink, H.K.; Campo, E.; Braziel, R.M.; Jaffe, E.S.; et al. Confirmation of the Molecular Classification of Diffuse Large B-Cell Lymphoma by Immunohistochemistry Using a Tissue Microarray. Blood 2004, 103, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Cludts, I.; Farrell, P.J. Multiple Functions within the Epstein-Barr Virus Ebna-3a Protein. J. Virol. 1998, 72, 1862–1869. [Google Scholar] [CrossRef] [PubMed]
- Budczies, J.; Klauschen, F.; Sinn, B.V.; Gyorffy, B.; Schmitt, W.D.; Darb-Esfahani, S.; Denkert, C. Cutoff Finder: A Comprehensive and Straightforward Web Application Enabling Rapid Biomarker Cutoff Optimization. PLoS ONE 2012, 12, e51862. [Google Scholar] [CrossRef] [PubMed]
- Simnica, D.; Akyuz, N.; Schliffke, S.; Mohme, M.; Wenserski, V.; Mahrle, L.T.; Fanchi, L.F.; Lamszus, K.; Binder, M. T Cell Receptor Next-Generation Sequencing Reveals Cancer-Associated Repertoire Metrics and Reconstitution after Chemotherapy in Patients with Hematological and Solid Tumors. Oncoimmunology 2019, 8, e1644110. [Google Scholar] [CrossRef]
- Morin, A.; Kwan, T.; Ge, B.; Letourneau, L.; Ban, M.; Tandre, K.; Caron, M.; Sandling, J.K.; Carlsson, J.; Bourque, G.; et al. Immunoseq: The Identification of Functionally Relevant Variants through Targeted Capture and Sequencing of Active Regulatory Regions in Human Immune Cells. BMC Med. Genom. 2016, 9, 59. [Google Scholar] [CrossRef]
- Steinert, E.M.; Schenkel, J.M.; Fraser, K.A.; Beura, L.K.; Manlove, L.S.; Igyarto, B.Z.; Southern, P.J.; Masopust, D. Quantifying Memory Cd8 T Cells Reveals Regionalization of Immunosurveillance. Cell 2015, 161, 737–749. [Google Scholar] [CrossRef]
- Rosenwald, A.; Wright, G.; Chan, W.C.; Connors, J.M.; Campo, E.; Fisher, R.I.; Gascoyne, R.D.; Muller-Hermelink, H.K.; Smeland, E.B.; Giltnane, J.M.; et al. The Use of Molecular Profiling to Predict Survival after Chemotherapy for Diffuse Large-B-Cell Lymphoma. N. Engl. J. Med. 2002, 346, 1937–1947. [Google Scholar] [CrossRef]
- Sha, C.; Barrans, S.; Cucco, F.; Bentley, M.; Care, M.A.; Cummin, T.; Kennedy, H.; Thompson, J.S.; Uddin, R.; Worrillow, L.; et al. Molecular High-Grade B-Cell Lymphoma: Defining a Poor-Risk Group That Requires Different Approaches to Therapy. J. Clin. Oncol. 2019, 37, 202–212. [Google Scholar] [CrossRef] [PubMed]
- Alizadeh, A.A.; Eisen, M.B.; Davis, R.E.; Ma, C.; Lossos, I.S.; Rosenwald, A.; Boldrick, J.C.; Sabet, H.; Tran, T.; Yu, X.; et al. Distinct Types of Diffuse Large B-Cell Lymphoma Identified by Gene Expression Profiling. Nature 2000, 403, 503–511. [Google Scholar] [CrossRef]
- Yang, C.Y.; Lin, M.W.; Chang, Y.L.; Wu, C.T.; Yang, P.C. Programmed Cell Death-Ligand 1 Expression Is Associated with a Favourable Immune Microenvironment and Better Overall Survival in Stage I Pulmonary Squamous Cell Carcinoma. Eur. J. Cancer 2016, 57, 91–103. [Google Scholar] [CrossRef]
- Mitchem, J.B.; Brennan, D.J.; Knolhoff, B.L.; Belt, B.A.; Zhu, Y.; Sanford, D.E.; Belaygorod, L.; Carpenter, D.; Collins, L.; Piwnica-Worms, D.; et al. Targeting Tumor-Infiltrating Macrophages Decreases Tumor-Initiating Cells, Relieves Immunosuppression, and Improves Chemotherapeutic Responses. Cancer Res. 2013, 73, 128–141. [Google Scholar] [CrossRef]
- Bernd, H.W.; Ziepert, M.; Thorns, C.; Klapper, W.; Wacker, H.H.; Hummel, M.; Stein, H.; Hansmann, M.L.; Ott, G.; Rosenwald, A.; et al. Loss of Hla-Dr Expression and Immunoblastic Morphology Predict Adverse Outcome in Diffuse Large B-Cell Lymphoma—Analyses of Cases from Two Prospective Randomized Clinical Trials. Haematologica 2009, 94, 1569–1580. [Google Scholar] [CrossRef] [PubMed]
- Tada, K.; Maeshima, A.M.; Hiraoka, N.; Yamauchi, N.; Maruyama, D.; Kim, S.W.; Watanabe, T.; Katayama, N.; Heike, Y.; Tobinai, K.; et al. Prognostic Significance of Hla Class I and Ii Expression in Patients with Diffuse Large B Cell Lymphoma Treated with Standard Chemoimmunotherapy. Cancer Immunol. Immunother. 2016, 65, 1213–1222. [Google Scholar] [CrossRef] [PubMed]
- Nijland, M.; Veenstra, R.N.; Visser, L.; Xu, C.; Kushekhar, K.; van Imhoff, G.W.; Kluin, P.M.; van den Berg, A.; Diepstra, A. Hla Dependent Immune Escape Mechanisms in B-Cell Lymphomas: Implications for Immune Checkpoint Inhibitor Therapy? Oncoimmunology 2017, 6, e1295202. [Google Scholar] [CrossRef]
- Wilkinson, S.T.; Vanpatten, K.A.; Fernandez, D.R.; Brunhoeber, P.; Garsha, K.E.; Glinsmann-Gibson, B.J.; Grogan, T.M.; Teruya-Feldstein, J.; Rimsza, L.M. Partial Plasma Cell Differentiation as a Mechanism of Lost Major Histocompatibility Complex Class Ii Expression in Diffuse Large B-Cell Lymphoma. Blood 2012, 119, 1459–1467. [Google Scholar] [CrossRef]
- Robert, L.; Tsoi, J.; Wang, X.; Emerson, R.; Homet, B.; Chodon, T.; Mok, S.; Huang, R.R.; Cochran, A.J.; Comin-Anduix, B.; et al. Ctla4 Blockade Broadens the Peripheral T-Cell Receptor Repertoire. Clin. Cancer Res. 2014, 20, 2424–2432. [Google Scholar] [CrossRef]
- Costa, A.I.; Koning, D.; Ladell, K.; McLaren, J.E.; Grady, B.P.; Schellens, I.M.; van Ham, P.; Nijhuis, M.; Borghans, J.A.; Kesmir, C.; et al. Complex T-Cell Receptor Repertoire Dynamics Underlie the Cd8+ T-Cell Response to Hiv-1. J. Virol. 2015, 89, 110–119. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Tumeh, P.C.; Harview, C.L.; Yearley, J.H.; Shintaku, I.P.; Taylor, E.J.; Robert, L.; Chmielowski, B.; Spasic, M.; Henry, G.; Ciobanu, V.; et al. Pd-1 Blockade Induces Responses by Inhibiting Adaptive Immune Resistance. Nature 2014, 515, 568–571. [Google Scholar] [CrossRef] [PubMed]
- Baum, P.D.; Young, J.J.; Schmidt, D.; Zhang, Q.; Hoh, R.; Busch, M.; Martin, J.; Deeks, S.; McCune, J.M. Blood T-Cell Receptor Diversity Decreases During the Course of Hiv Infection, but the Potential for a Diverse Repertoire Persists. Blood 2012, 119, 3469–3477. [Google Scholar] [CrossRef][Green Version]
- McGranahan, N.; Furness, A.J.; Rosenthal, R.; Ramskov, S.; Lyngaa, R.; Saini, S.K.; Jamal-Hanjani, M.; Wilson, G.A.; Birkbak, N.J.; Hiley, C.T.; et al. Clonal Neoantigens Elicit T Cell Immunoreactivity and Sensitivity to Immune Checkpoint Blockade. Science 2016, 351, 1463–1469. [Google Scholar] [CrossRef]
- Evrard, S.M.; Pericart, S.; Grand, D.; Amara, N.; Escudie, F.; Gilhodes, J.; Bories, P.; Traverse-Glehen, A.; Dubois, R.; Brousset, P.; et al. Targeted Next Generation Sequencing Reveals High Mutation Frequency of Crebbp, Bcl2 and Kmt2d in High-Grade B-Cell Lymphoma with Myc and Bcl2 and/or Bcl6 Rearrangements. Haematologica 2019, 104, e154–e157. [Google Scholar] [CrossRef]
- Linnemann, C.; van Buuren, M.M.; Bies, L.; Verdegaal, E.M.; Schotte, R.; Calis, J.J.; Behjati, S.; Velds, A.; Hilkmann, H.; Atmioui, D.E.; et al. High-Throughput Epitope Discovery Reveals Frequent Recognition of Neo-Antigens by Cd4+ T Cells in Human Melanoma. Nat. Med. 2015, 21, 81–85. [Google Scholar] [CrossRef]
- Tamura, K.; Hazama, S.; Yamaguchi, R.; Imoto, S.; Takenouchi, H.; Inoue, Y.; Kanekiyo, S.; Shindo, Y.; Miyano, S.; Nakamura, Y.; et al. Characterization of the T Cell Repertoire by Deep T Cell Receptor Sequencing in Tissues and Blood from Patients with Advanced Colorectal Cancer. Oncol. Lett. 2016, 11, 3643–3649. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.D.; Madireddi, S.; de Almeida, P.E.; Banchereau, R.; Chen, Y.J.; Chitre, A.S.; Chiang, E.Y.; Iftikhar, H.; O’Gorman, W.E.; Au-Yeung, A.; et al. Peripheral T Cell Expansion Predicts Tumour Infiltration and Clinical Response. Nature 2020, 579, 274–278. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Xiong, D.; Wang, X.; Liu, H.; Wang, T. Mapping the Functional Landscape of T Cell Receptor Repertoires by Single-T Cell Transcriptomics. Nat. Methods 2021, 18, 92–99. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.N.; Meyerson, M. Antigen Identification for Hla Class I- and Hla Class Ii-Restricted T Cell Receptors Using Cytokine-Capturing Antigen-Presenting Cells. Sci. Immunol. 2021, 6, 55. [Google Scholar] [CrossRef]
- Kula, T.; Dezfulian, M.H.; Wang, C.I.; Abdelfattah, N.S.; Hartman, Z.C.; Wucherpfennig, K.W.; Lyerly, H.K.; Elledge, S.J. T-Scan: A Genome-Wide Method for the Systematic Discovery of T Cell Epitopes. Cell 2019, 178, 1016–1028.e13. [Google Scholar] [CrossRef] [PubMed]
- Mundo, L.; Del Porro, L.; Granai, M.; Siciliano, M.C.; Mancini, V.; Santi, R.; Marcar, L.; Vrzalikova, K.; Vergoni, F.; Di Stefano, G.; et al. Frequent Traces of Ebv Infection in Hodgkin and Non-Hodgkin Lymphomas Classified as Ebv-Negative by Routine Methods: Expanding the Landscape of Ebv-Related Lymphomas. Mod. Pathol. 2020, 33, 2407–2421. [Google Scholar] [CrossRef] [PubMed]
- Witte, H.; Biersack, H.; Kopelke, S.; Rades, D.; Merz, H.; Bernard, V.; Lehnert, H.; Fetscher, S.; Gebauer, N. The Glasgow prognostic score at diagnosis is an independent predictor of survival in advanced stage classical Hodgkin lymphoma. Br. J. Haematol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Witte, H.M.; Bonorden, B.; Riecke, A.; Biersack, H.; Steinestel, K.; Merz, H.; Feller, A.C.; Bernard, V.; Fetscher, S.; von Bubnoff, N.; et al. The Glasgow Prognostic Score at Diagnosis Is a Predictor of Clinical Outcome in Patients with Multiple Myeloma Undergoing Autologous Haematopoietic Stem Cell Transplantation. Cancers 2020, 12, 921. [Google Scholar] [CrossRef]
- Sehn, L.H.; Berry, B.; Chhanabhai, M.; Fitzgerald, C.; Gill, K.; Hoskins, P.; Klasa, R.; Savage, K.J.; Shenkier, T.; Sutherland, J.; et al. The Revised International Prognostic Index (R-Ipi) Is a Better Predictor of Outcome Than the Standard Ipi for Patients with Diffuse Large B-Cell Lymphoma Treated with R-Chop. Blood 2007, 109, 1857–1861. [Google Scholar] [CrossRef]
- Lister, T.A.; Crowther, D.; Sutcliffe, S.B.; Glatstein, E.; Canellos, G.P.; Young, R.C.; Rosenberg, S.A.; Coltman, C.A.; Tubiana, M. Report of a Committee Convened to Discuss the Evaluation and Staging of Patients with Hodgkin’s Disease: Cotswolds Meeting. J. Clin. Oncol. 1989, 7, 1630–1636. [Google Scholar] [CrossRef]
- Hao, X.; Wei, Y.; Wei, X.; Zhou, L.; Wei, Q.; Zhang, Y.; Huang, W.; Feng, R. Glasgow Prognostic Score Is Superior to Other Inflammation-Based Scores in Predicting Survival of Diffuse Large B-Cell Lymphoma. Oncotarget 2017, 8, 76740–76748. [Google Scholar] [CrossRef] [PubMed]
- Carlson, C.S.; Emerson, R.O.; Sherwood, A.M.; Desmarais, C.; Chung, M.W.; Parsons, J.M.; Steen, M.S.; LaMadrid-Herrmannsfeldt, M.A.; Williamson, D.W.; Livingston, R.J.; et al. Using Synthetic Templates to Design an Unbiased Multiplex Pcr Assay. Nat. Commun. 2013, 4, 2680. [Google Scholar] [CrossRef]
- Robins, H.S.; Campregher, P.V.; Srivastava, S.K.; Wacher, A.; Turtle, C.J.; Kahsai, O.; Riddell, S.R.; Warren, E.H.; Carlson, C.S. Comprehensive Assessment of T-Cell Receptor Beta-Chain Diversity in Alphabeta T Cells. Blood 2009, 114, 4099–4107. [Google Scholar] [CrossRef]
- Shugay, M.; Bagaev, D.V.; Zvyagin, I.V.; Vroomans, R.M.; Crawford, J.C.; Dolton, G.; Komech, E.A.; Sycheva, A.L.; Koneva, A.E.; Egorov, E.S.; et al. Vdjdb: A Curated Database of T-Cell Receptor Sequences with Known Antigen Specificity. Nucleic Acids Res. 2018, 46, D419–D427. [Google Scholar] [CrossRef] [PubMed]

| Characteristics | DHL (n = 40) | THL (n = 7) | Triple-Negative DLBCL (n = 27) |
|---|---|---|---|
| Age (yrs.; median + range) | 73.0 (35–89) | 69.0 (42–79) | 70 (18–87) |
| Sex | |||
| Female | 19 (47.5%) | 2 (28.6%) | 13 (48.1%) |
| Male | 21 (52.5%) | 5 (71.4%) | 14 (51.9%) |
| R-IPI | |||
| 0 | 1 (2.5%) | - | 1 (3.7%) |
| 1–2 | 16 (40.0%) | 2 (28.6%) | 15 (55.6%) |
| >2 | 23 (57.5%) | 5 (71.4%) | 11 (40.7%) |
| Stage (Ann Arbor) | |||
| I | 7 (17.5%) | - | 2 (7.4%) |
| II | 8 (20.0%) | 3 (42.8%) | 11 (40.7%) |
| III | 5 (12.5%) | 2 (28.6%) | 8 (29.6%) |
| IV | 20 (50.0%) | 2 (28.6%) | 6 (22.2%) |
| B-Symptoms | |||
| Yes | 22 (55.0%) | 4 (57.2%) | 12 (44.4%) |
| No | 18 (45.0%) | 3 (42.8%) | 15 (55.6%) |
| Extranodal sites | |||
| 0 | 10 (25.0%) | 3 (42.8%) | 10 (37.0%) |
| 1–2 | 28 (70.0%) | 4 (57.2%) | 17 (63.0%) |
| >2 | 2 (5.0%) | - | - |
| ECOG PS | |||
| 0–2 | 37 (92.5%) | 7 (100.0%) | 24 (88.9%) |
| >2 | 3 (7.5%) | - | 3 (11.1%) |
| LDH | |||
| Normal | 11 (27.5%) | 1 (14.3%) | 14 (51.9%) |
| Elevated | 29 (72.5%) | 6 (85.7%) | 13 (48.1%) |
| Glasgow prognostic score (GPS) | |||
| GPS 0 | 12 (30.0%) | 1 (14.3%) | 17 (63.0%) |
| GPS 1 | 8 (20.0%) | 1 (14.3%) | 7 (25.9%) |
| GPS 2 | 20 (50.0%) | 5 (71.4%) | 3 (11.1%) |
| CNS involvement at diagnosis | |||
| Yes | 1 (2.5%) | - | - |
| No | 45 (97.5%) | 7 (100.0%) | 27 (100.0%) |
| Characteristics | DHL (n = 40) | THL (n = 7) | Triple-Negative DLBCL (n = 27) |
|---|---|---|---|
| Frontline Therapy regimen | |||
| R-CHOP-like | 27 (67.5%) | 5 (71.4%) | 23 (85.2%) |
| DR | 7 (25.9%) | 3 (60.0%) | 10 (43.5%) |
| Others | 12 (30.0%) | 2 (28.6%) | 3 (11.1%) |
| Refusal of treatment | 1 (2.5%) | - | 1 (3.7%) |
| Response rates | |||
| CR | 15 (37.5%) | 1 (14.3%) | 14 (51.9%) |
| PR | 17 (42.5%) | 3 (42.9%) | 11 (40.7%) |
| SD | 3 (7.5%) | 1 (14.3%) | - |
| PD | 5 (12.5%) | 2 (28.6%) | 2 (7.4%) |
| Relapse rate | 21 (52.5%) | 6 (85.7%) | 4 (14.8%) |
| 2nd line treatment regimens (n = 35) | |||
| R-DHAP | 7 (33.3%) | 2 (33.3%) | 1 (25.0%) |
| R-Bendamustine | 3 (14.3%) | 2 (33.3%) | 1 (25.0%) |
| Auto/Allo HSCT | 4 (19.4%) | 1 (16.7%) | - |
| Others | 5 (23.8%) | - | 2 (50.0%) |
| Refusal of treatment | 2 (9.5%) | 1 (16.7%) | - |
| Toxicity profile (1st line) | |||
| Cytopenia grade III/IV | 13 (32.5%) | 3 (42.9%) | 11 (40.7%) |
| Polyneuropathy | 5 (12.5%) | 1 (14.3%) | 7 (25.9%) |
| GI toxicity | 2 (5.0%) | - | - |
| Mucositis | 2 (5.0%) | 2 (28.6%) | - |
| Pneumonia | 3 (7.5%) | - | 1 (3.7%) |
| Acute kidney injury | 5 (12.5%) | - | - |
| Sepsis | 12 (30.0%) | - | 7 (25.9%) |
| Characteristics | DHL (n = 40) | THL (n = 7) | Triple-Negative DLBCL (n = 27) |
|---|---|---|---|
| Hans classifier and IHC | |||
| GCB | 34 (85.0%) | 6 (85.7%) | 2 (7.4%) |
| Non-GCB | 6 (15.0%) | 1 (14.3%) | 25 (92.6%) |
| BCL2 | 28 (70.0%) | 5 (71.4%) | 1 (3.7%) |
| BCL6 | 33 (82.5%) | 6 (85.7%) | 3 (11.1%) |
| MYC | 25 (62.5%) | 4 (57.1%) | - |
| CD10 | 31 (77.5%) | 5 (71.4%) | 3 (11.1%) |
| MUM1/IRF4 | 9 (22.5%) | 2 (28.6%) | 3 (11.1%) |
| Ki-67 (median, range) | 85% (40–100%) | 75% (50–95%) | 85% (55–95%) |
| T-cell infiltration | |||
| Percentage (median, range) | 15% (1–30%) | 10% (3–30%) | 20% (5–30%) |
| Analyzed Parameter | HGBL-DH/TH (n = 47) | tnDLBCL (n = 27) | p-Value |
|---|---|---|---|
| Productive TCR rearrangements | 1504.5 (136–24,680) | 3447 (164–21,745) | 0.0050 |
| Productive Clonality | 0.036 (0.0009–0.3614) | 0.08785 (0.0013–0.4437) | 0.0427 |
| % Maximum frequency clone | 0.407 (0.205–1.822) | 4.1884579 (0.4035117–44.8000014) | 0.2473 |
| Amino Acid | Present in HGBL-DH/TH | Mean Frequency (%) | Epitope Gene | Epitope SPECIES | V-Gene | J-Gene |
|---|---|---|---|---|---|---|
| CASSPLGETQYF | 4 | 0.419 | - | - | TCRBV18-01*01 | TCRBJ02-05*01 |
| CASRDTEAFF | 4 | 1.822 | - | - | TCRBV06 | TCRBJ01-01*01 |
| CASSPTRTGDYEQYF | 3 | 1.593 | - | - | TCRBV07-09 | TCRBJ02-07*01 |
| CASSDSSLTEAFF | 3 | 0.201 | EBNA3A | EBV | TCRBV06-04 | TCRBJ01-01*01 |
| CASSQGGSGEQFF | 3 | 0.705 | - | - | TCRBV04-03*01 | TCRBJ02-01*01 |
| CASSLGGSPEAFF | 3 | 0.524 | - | - | TCRBV07-02*01 | TCRBJ01-01*01 |
| CASSQGGSGELFF | 3 | 0.468 | - | - | TCRBV04-03*01 | TCRBJ02-02*01 |
| CASSLESGANVLTF | 3 | 0.428 | - | - | TCRBV07-06*01 | TCRBJ02-06*01 |
| CASSLGGAGANVLTF | 3 | 0.395 | - | - | TCRBV05-06*01 | TCRBJ02-06*01 |
| CASSLAGGGTEAFF | 3 | 0.388 | - | - | TCRBV28-01*01 | TCRBJ01-01*01 |
| CSARDRVSYNEQFF | 3 | 0.302 | - | - | TCRBV20 | TCRBJ02-01*01 |
| CSARDPGSSYEQYF | 3 | 0.279 | - | - | TCRBV20 | TCRBJ02-07*01 |
| CASSGTEAFF | 3 | 0.216 | - | - | TCRBV28-01*01 | TCRBJ01-01*01 |
| CSARSGNTEAFF | 3 | 0.205 | - | - | TCRBV20 | TCRBJ01-01*01 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Olschewski, V.; Witte, H.M.; Bernard, V.; Steinestel, K.; Peter, W.; Merz, H.; Rieken, J.; Biersack, H.; von Bubnoff, N.; Feller, A.C.; et al. Systemic Inflammation and Tumour-Infiltrating T-Cell Receptor Repertoire Diversity Are Predictive of Clinical Outcome in High-Grade B-Cell Lymphoma with MYC and BCL2 and/or BCL6 Rearrangements. Cancers 2021, 13, 887. https://doi.org/10.3390/cancers13040887
Olschewski V, Witte HM, Bernard V, Steinestel K, Peter W, Merz H, Rieken J, Biersack H, von Bubnoff N, Feller AC, et al. Systemic Inflammation and Tumour-Infiltrating T-Cell Receptor Repertoire Diversity Are Predictive of Clinical Outcome in High-Grade B-Cell Lymphoma with MYC and BCL2 and/or BCL6 Rearrangements. Cancers. 2021; 13(4):887. https://doi.org/10.3390/cancers13040887
Chicago/Turabian StyleOlschewski, Vito, Hanno M. Witte, Veronica Bernard, Konrad Steinestel, Wolfgang Peter, Hartmut Merz, Johannes Rieken, Harald Biersack, Nikolas von Bubnoff, Alfred C. Feller, and et al. 2021. "Systemic Inflammation and Tumour-Infiltrating T-Cell Receptor Repertoire Diversity Are Predictive of Clinical Outcome in High-Grade B-Cell Lymphoma with MYC and BCL2 and/or BCL6 Rearrangements" Cancers 13, no. 4: 887. https://doi.org/10.3390/cancers13040887
APA StyleOlschewski, V., Witte, H. M., Bernard, V., Steinestel, K., Peter, W., Merz, H., Rieken, J., Biersack, H., von Bubnoff, N., Feller, A. C., & Gebauer, N. (2021). Systemic Inflammation and Tumour-Infiltrating T-Cell Receptor Repertoire Diversity Are Predictive of Clinical Outcome in High-Grade B-Cell Lymphoma with MYC and BCL2 and/or BCL6 Rearrangements. Cancers, 13(4), 887. https://doi.org/10.3390/cancers13040887








