Prognostic Gene Expression, Stemness and Immune Microenvironment in Pediatric Tumors
Abstract
Simple Summary
Abstract
1. Introduction
2. Results
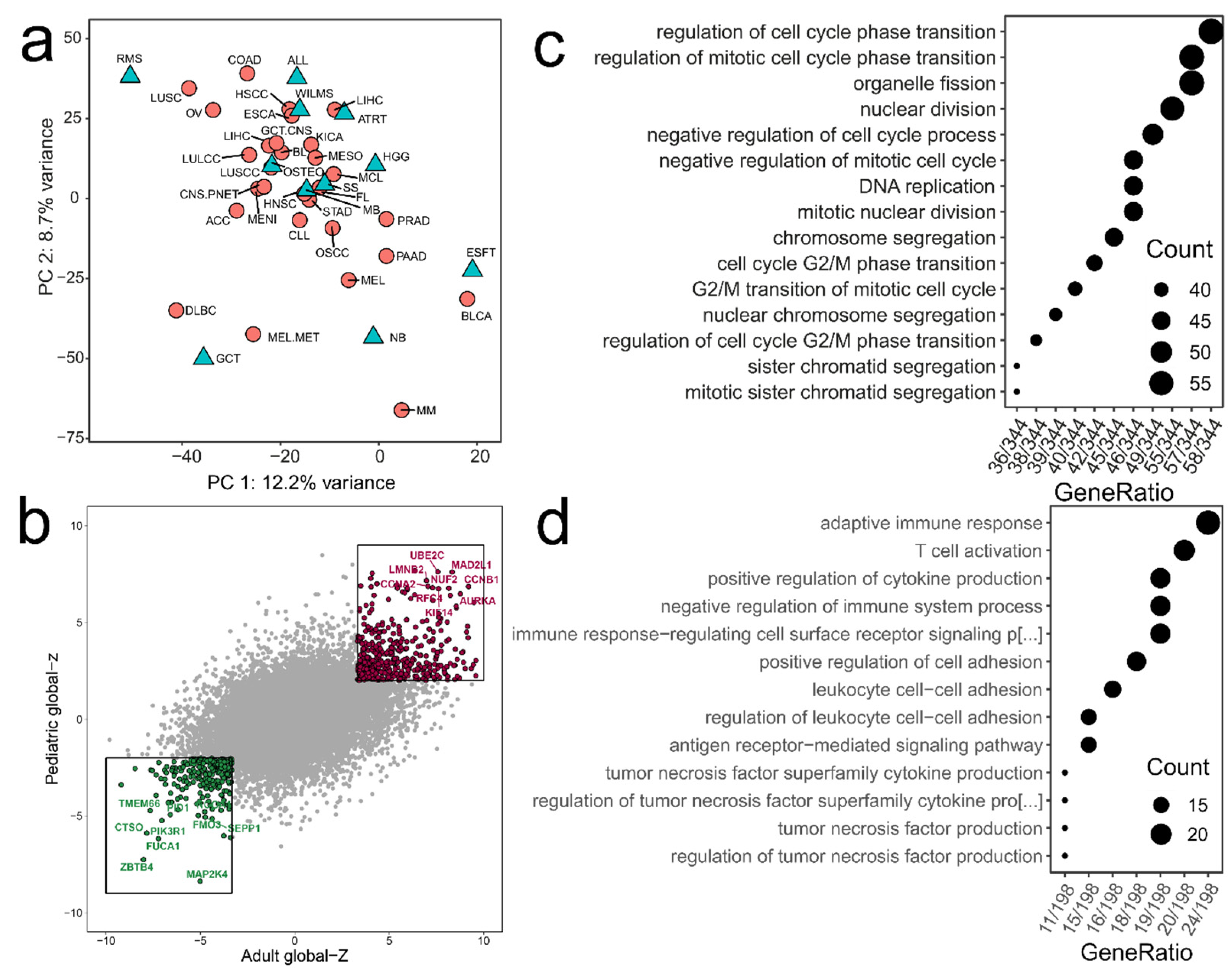
2.1. The Prognostic Landscape of Gene Expression
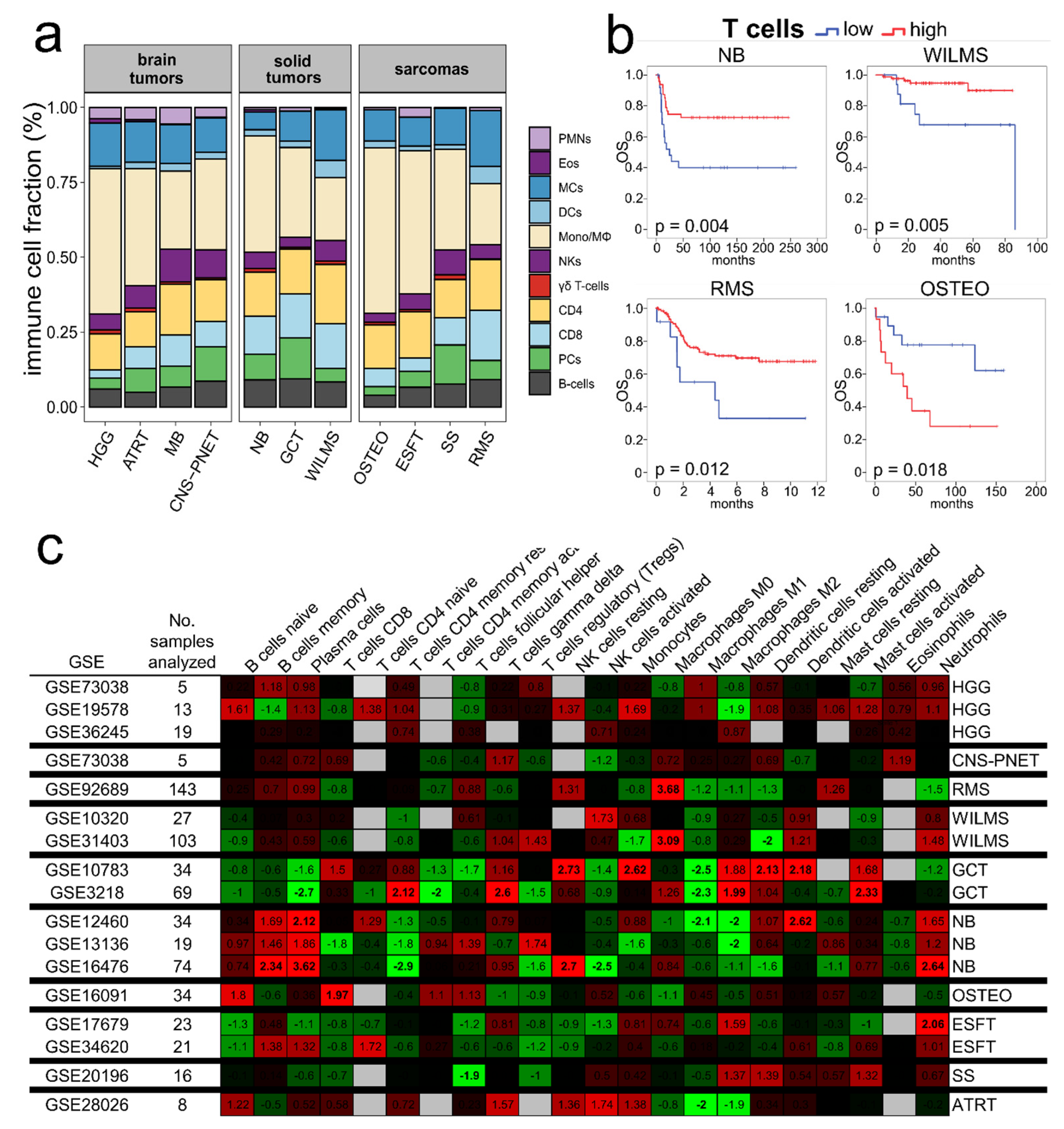
2.2. Immune Cell Infiltration—Composition and Prognostication
2.3. Stemness Correlates with Immature Histology, Clinic-Pathological Features and Survival
2.4. Immune TME and Stemness in WILMS
3. Discussion
4. Materials and Methods
4.1. Gene Expression Data and Study Design
4.2. Assessment of Prognostic Landscape by PRECOG
4.3. Assessment of Immune Infiltration by CIBERSORT
4.4. Calculation of Gene Expression-Based Stemness Index (mRNAsi)
4.5. Tissue Microarray (TMA) Cohort, Immunohistochemistry and Scoring
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Smith, A. A glossary for stem-cell biology. Nature 2006, 441, 1060. [Google Scholar] [CrossRef]
- Marshall, G.M.; Carter, D.R.; Cheung, B.B.; Liu, T.; Mateos, M.K.; Meyerowitz, J.G.; Weiss, W.A. The prenatal origins of cancer. Nat. Rev. Cancer 2014, 14, 277–289. [Google Scholar] [CrossRef] [PubMed]
- Miranda, A.; Hamilton, P.T.; Zhang, A.W.; Pattnaik, S.; Becht, E.; Mezheyeuski, A.; Bruun, J.; Micke, P.; de Reynies, A.; Nelson, B.H. Cancer stemness, intratumoral heterogeneity, and immune response across cancers. Proc. Natl. Acad. Sci. USA 2019, 116, 9020–9029. [Google Scholar] [CrossRef] [PubMed]
- Malta, T.M.; Sokolov, A.; Gentles, A.J.; Burzykowski, T.; Poisson, L.; Weinstein, J.N.; Kamińska, B.; Huelsken, J.; Omberg, L.; Gevaert, O.; et al. Machine Learning Identifies Stemness Features Associated with Oncogenic Dedifferentiation. Cell 2018, 173, 338–354.e15. [Google Scholar] [CrossRef]
- Shats, I.; Gatza, M.L.; Chang, J.T.; Mori, S.; Wang, J.; Rich, J.; Nevins, J.R. Using a stem cell-based signature to guide therapeutic selection in cancer. Cancer Res. 2011, 71, 1772–1780. [Google Scholar] [CrossRef] [PubMed]
- Smith, B.A.; Balanis, N.G.; Nanjundiah, A.; Sheu, K.M.; Tsai, B.L.; Zhang, Q.; Park, J.W.; Thompson, M.; Huang, J.; Witte, O.N.; et al. A Human Adult Stem Cell Signature Marks Aggressive Variants across Epithelial Cancers. Cell Rep. 2018, 24, 3353–3366.e5. [Google Scholar] [CrossRef] [PubMed]
- Zeng, H.; Ji, J.; Song, X.; Huang, Y.; Li, H.; Huang, J.; Ma, X. Stemness Related Genes Revealed by Network Analysis Associated With Tumor Immune Microenvironment and the Clinical Outcome in Lung Adenocarcinoma. Front. Genet. 2020, 11, 549213. [Google Scholar] [CrossRef] [PubMed]
- Agudo, J.; Park, E.S.; Rose, S.A.; Alibo, E.; Sweeney, R.; Dhainaut, M.; Kobayashi, K.S.; Sachidanandam, R.; Baccarini, A.; Merad, M.; et al. Quiescent Tissue Stem Cells Evade Immune Surveillance. Immunity 2018, 48, 271–285.e5. [Google Scholar] [CrossRef] [PubMed]
- Clara, J.A.; Monge, C.; Yang, Y.; Takebe, N. Targeting signalling pathways and the immune microenvironment of cancer stem cells—A clinical update. Nat. Rev. Clin. Oncol. 2020, 17, 204–232. [Google Scholar] [CrossRef]
- Bruttel, V.S.; Wischhusen, J. Cancer stem cell immunology: Key to understanding tumorigenesis and tumor immune escape? Front. Immunol. 2014, 5, 360. [Google Scholar] [CrossRef] [PubMed]
- Maccalli, C.; Volontè, A.; Cimminiello, C.; Parmiani, G. Immunology of cancer stem cells in solid tumours. A review. Eur. J. Cancer 2014, 50, 649–655. [Google Scholar] [CrossRef] [PubMed]
- Meis-Kindblom, J.M.; Stenman, G.; Kindblom, L.G. Differential diagnosis of small round cell tumors. Semin. Diagn. Pathol. 1996, 13, 213–241. [Google Scholar]
- Rajwanshi, A.; Srinivas, R.; Upasana, G. Malignant small round cell tumors. J. Cytol. 2009, 26, 1–10. [Google Scholar] [CrossRef]
- Popov, S.D.; Sebire, N.J.; Vujanic, G.M. Wilms’ Tumour—Histology and Differential Diagnosis; Codon Publications: Brisbane, Australia, 2016; ISBN 9780994438119. [Google Scholar]
- Rivera, M.N.; Haber, D.A. Wilms’ tumour: Connecting tumorigenesis and organ development in the kidney. Nat. Rev. Cancer 2005, 5, 699–712. [Google Scholar] [CrossRef]
- Pinto, N.; Park, J.R.; Murphy, E.; Yearley, J.; McClanahan, T.; Annamalai, L.; Hawkins, D.S.; Rudzinski, E.R. Patterns of PD-1, PD-L1, and PD-L2 expression in pediatric solid tumors. Pediatr. Blood Cancer 2017, 64. [Google Scholar] [CrossRef]
- Aoki, T.; Hino, M.; Koh, K.; Kyushiki, M.; Kishimoto, H.; Arakawa, Y.; Hanada, R.; Kawashima, H.; Kurihara, J.; Shimojo, N.; et al. Low Frequency of Programmed Death Ligand 1 Expression in Pediatric Cancers. Pediatr. Blood Cancer 2016, 63, 1461–1464. [Google Scholar] [CrossRef]
- Stahl, D.; Gentles, A.J.; Thiele, R.; Gütgemann, I. Prognostic profiling of the immune cell microenvironment in Ewing´s Sarcoma Family of Tumors. Oncoimmunology 2019, 8, e1674113. [Google Scholar] [CrossRef] [PubMed]
- Petitprez, F.; de Reyniès, A.; Keung, E.Z.; Chen, T.W.-W.; Sun, C.-M.; Calderaro, J.; Jeng, Y.-M.; Hsiao, L.-P.; Lacroix, L.; Bougoüin, A.; et al. B cells are associated with survival and immunotherapy response in sarcoma. Nature 2020, 577, 556–560. [Google Scholar] [CrossRef] [PubMed]
- Gröbner, S.N.; Worst, B.C.; Weischenfeldt, J.; Buchhalter, I.; Kleinheinz, K.; Rudneva, V.A.; Johann, P.D.; Balasubramanian, G.P.; Segura-Wang, M.; Brabetz, S.; et al. The landscape of genomic alterations across childhood cancers. Nature 2018, 555, 321–327. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Liu, Y.; Liu, Y.; Alexandrov, L.B.; Edmonson, M.N.; Gawad, C.; Zhou, X.; Li, Y.; Rusch, M.C.; Easton, J.; et al. Pan-cancer genome and transcriptome analyses of 1699 paediatric leukaemias and solid tumours. Nature 2018, 555, 371–376. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Walsh, M.F.; Wu, G.; Edmonson, M.N.; Gruber, T.A.; Easton, J.; Hedges, D.; Ma, X.; Zhou, X.; Yergeau, D.A.; et al. Germline Mutations in Predisposition Genes in Pediatric Cancer. N. Engl. J. Med. 2015, 373, 2336–2346. [Google Scholar] [CrossRef] [PubMed]
- Kattner, P.; Strobel, H.; Khoshnevis, N.; Grunert, M.; Bartholomae, S.; Pruss, M.; Fitzel, R.; Halatsch, M.-E.; Schilberg, K.; Siegelin, M.D.; et al. Compare and contrast: Pediatric cancer versus adult malignancies. Cancer Metastasis Rev. 2019, 38, 673–682. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Hsu, W.-H.; Han, J.; Xia, Y.; DePinho, R.A. Cancer Stemness Meets Immunity: From Mechanism to Therapy. Cell Rep. 2021, 34, 108597. [Google Scholar] [CrossRef]
- Grabovska, Y.; Mackay, A.; O’Hare, P.; Crosier, S.; Finetti, M.; Schwalbe, E.C.; Pickles, J.C.; Fairchild, A.R.; Avery, A.; Cockle, J.; et al. Pediatric pan-central nervous system tumor analysis of immune-cell infiltration identifies correlates of antitumor immunity. Nat. Commun. 2020, 11, 4324. [Google Scholar] [CrossRef] [PubMed]
- Lian, H.; Han, Y.-P.; Zhang, Y.-C.; Zhao, Y.; Yan, S.; Li, Q.-F.; Wang, B.-C.; Wang, J.-J.; Meng, W.; Yang, J.; et al. Integrative analysis of gene expression and DNA methylation through one-class logistic regression machine learning identifies stemness features in medulloblastoma. Mol. Oncol. 2019, 13, 2227–2245. [Google Scholar] [CrossRef] [PubMed]
- Gentles, A.J.; Newman, A.M.; Liu, C.L.; Bratman, S.V.; Feng, W.; Kim, D.; Nair, V.S.; Xu, Y.; Khuong, A.; Hoang, C.D.; et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat. Med. 2015, 21, 938–945. [Google Scholar] [CrossRef]
- Newman, A.M.; Liu, C.L.; Green, M.R.; Gentles, A.J.; Feng, W.; Xu, Y.; Hoang, C.D.; Diehn, M.; Alizadeh, A.A. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 2015, 12, 453–457. [Google Scholar] [CrossRef] [PubMed]
- Budczies, J.; Klauschen, F.; Sinn, B.V.; Győrffy, B.; Schmitt, W.D.; Darb-Esfahani, S.; Denkert, C. Cutoff Finder: A comprehensive and straightforward Web application enabling rapid biomarker cutoff optimization. PLoS ONE 2012, 7, e51862. [Google Scholar] [CrossRef] [PubMed]
- Campbell, K.; Gastier-Foster, J.M.; Mann, M.; Naranjo, A.H.; van Ryn, C.; Bagatell, R.; Matthay, K.K.; London, W.B.; Irwin, M.S.; Shimada, H.; et al. Association of MYCN copy number with clinical features, tumor biology, and outcomes in neuroblastoma: A report from the Children’s Oncology Group. Cancer 2017, 123, 4224–4235. [Google Scholar] [CrossRef]
- Goltsev, Y.; Samusik, N.; Kennedy-Darling, J.; Bhate, S.; Hale, M.; Vazquez, G.; Black, S.; Nolan, G.P. Deep Profiling of Mouse Splenic Architecture with CODEX Multiplexed Imaging. Cell 2018, 174, 968–981.e15. [Google Scholar] [CrossRef]
- Maturu, P.; Overwijk, W.W.; Hicks, J.; Ekmekcioglu, S.; Grimm, E.A.; Huff, V. Characterization of the inflammatory microenvironment and identification of potential therapeutic targets in wilms tumors. Transl. Oncol. 2014, 7, 484–492. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.-H.; Yeh, D.-W.; Lai, C.-Y.; Liu, Y.-L.; Huang, L.-R.; Lee, A.Y.-L.; Jin, S.-L.C.; Chuang, T.-H. USP17 mediates macrophage-promoted inflammation and stemness in lung cancer cells by regulating TRAF2/TRAF3 complex formation. Oncogene 2018, 37, 6327–6340. [Google Scholar] [CrossRef] [PubMed]
- Barrett, T.; Suzek, T.O.; Troup, D.B.; Wilhite, S.E.; Ngau, W.-C.; Ledoux, P.; Rudnev, D.; Lash, A.E.; Fujibuchi, W.; Edgar, R. NCBI GEO: Mining millions of expression profiles--database and tools. Nucleic Acids Res. 2005, 33, D562–D566. [Google Scholar] [CrossRef]
- Irizarry, R.A.; Hobbs, B.; Collin, F.; Beazer-Barclay, Y.D.; Antonellis, K.J.; Scherf, U.; Speed, T.P. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 2003, 4, 249–264. [Google Scholar] [CrossRef]
- Daily, K.; Ho Sui, S.J.; Schriml, L.M.; Dexheimer, P.J.; Salomonis, N.; Schroll, R.; Bush, S.; Keddache, M.; Mayhew, C.; Lotia, S.; et al. Molecular, phenotypic, and sample-associated data to describe pluripotent stem cell lines and derivatives. Sci. Data 2017, 4, 170030. [Google Scholar] [CrossRef] [PubMed]
- Salomonis, N.; Dexheimer, P.J.; Omberg, L.; Schroll, R.; Bush, S.; Huo, J.; Schriml, L.; Ho Sui, S.; Keddache, M.; Mayhew, C.; et al. Integrated Genomic Analysis of Diverse Induced Pluripotent Stem Cells from the Progenitor Cell Biology Consortium. Stem Cell Rep. 2016, 7, 110–125. [Google Scholar] [CrossRef]
- Kononen, J.; Bubendorf, L.; Kallioniemi, A.; Bärlund, M.; Schraml, P.; Leighton, S.; Torhorst, J.; Mihatsch, M.J.; Sauter, G.; Kallioniemi, O.P. Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat. Med. 1998, 4, 844–847. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B (Methodol.) 1995, 57, 289–300. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stahl, D.; Knoll, R.; Gentles, A.J.; Vokuhl, C.; Buness, A.; Gütgemann, I. Prognostic Gene Expression, Stemness and Immune Microenvironment in Pediatric Tumors. Cancers 2021, 13, 854. https://doi.org/10.3390/cancers13040854
Stahl D, Knoll R, Gentles AJ, Vokuhl C, Buness A, Gütgemann I. Prognostic Gene Expression, Stemness and Immune Microenvironment in Pediatric Tumors. Cancers. 2021; 13(4):854. https://doi.org/10.3390/cancers13040854
Chicago/Turabian StyleStahl, David, Rainer Knoll, Andrew J. Gentles, Christian Vokuhl, Andreas Buness, and Ines Gütgemann. 2021. "Prognostic Gene Expression, Stemness and Immune Microenvironment in Pediatric Tumors" Cancers 13, no. 4: 854. https://doi.org/10.3390/cancers13040854
APA StyleStahl, D., Knoll, R., Gentles, A. J., Vokuhl, C., Buness, A., & Gütgemann, I. (2021). Prognostic Gene Expression, Stemness and Immune Microenvironment in Pediatric Tumors. Cancers, 13(4), 854. https://doi.org/10.3390/cancers13040854





