Liquid Biopsy-Based Exo-oncomiRNAs Can Predict Prostate Cancer Aggressiveness
Abstract
Simple Summary
Abstract
1. Introduction
2. Results
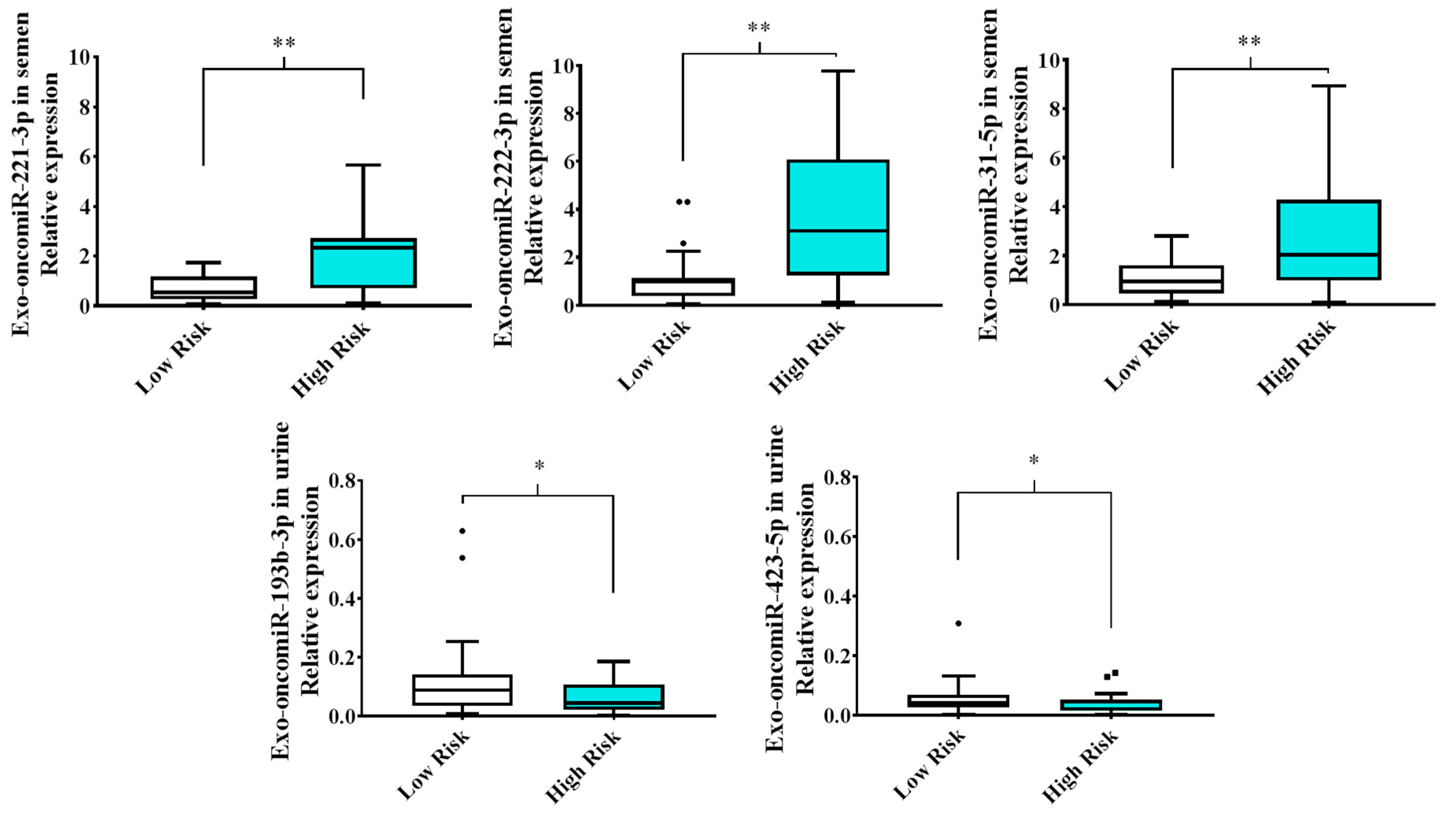
2.1. Extracellular Vesicle-Derived Exo-oncomiRNAs Are Differentially Expressed in Liquid Biopsies from Patients with Prostate Cancer Based on the Degree of Cancer Aggressiveness
2.2. Semen Levels of Exo-oncomiR-221-3p May Help Identify an Aggressive Prostate Cancer Phenotype
2.3. TWEAK Modulates Potential Predicted Targets for oncomiR-221-3p
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Extracellular Vesicle Isolation from Cell Culture Media and Exo-oncomiRNA Expression Profile Using TaqMan Low-Density Arrays
4.3. Extracellular Vesicle Analysis
4.4. Transmission Electron Microscopy Analysis
4.5. Patients
4.6. Analytical Methods
4.7. Sample Processing
4.8. Extracellular Vesicles Extraction from Liquid Biopsy and Exo-onocomiRNA Quantitative Real-Time PCR Profiling
4.9. Target Search by Bioinformatic Analysis
4.10. Functional Studies
4.11. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA. Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Markozannes, G.; Tzoulaki, I.; Karli, D.; Evangelou, E.; Ntzani, E.; Gunter, M.J.; Norat, T.; Ioannidis, J.P.; Tsilidis, K.K. Diet, body size, physical activity and risk of prostate cancer: An umbrella review of the evidence. Eur. J. Cancer 2016, 69, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Epstein, J.I.; Egevad, L.; Amin, M.B.; Delahunt, B.; Srigley, J.R.; Humphrey, P.A. The 2014 International Society of Urological Pathology (ISUP) Consensus Conference on Gleason Grading of Prostatic Carcinoma. Am. J. Surg. Pathol. 2015, 40, 1. [Google Scholar] [CrossRef] [PubMed]
- Saini, S. PSA and beyond: Alternative prostate cancer biomarkers. Cell. Oncol. 2016, 39, 97–106. [Google Scholar] [CrossRef]
- Di Meo, A.; Bartlett, J.; Cheng, Y.; Pasic, M.D.; Yousef, G.M. Liquid biopsy: A step forward towards precision medicine in urologic malignancies. Mol. Cancer 2017, 16, 1–14. [Google Scholar] [CrossRef]
- Greten, F.R.; Grivennikov, S.I. Inflammation and Cancer: Triggers, Mechanisms, and Consequences. Immunity 2019, 51, 27–41. [Google Scholar] [CrossRef]
- Winkles, J.A. The TWEAK-Fn14 cytokine-receptor axis: Discovery, biology and therapeutic targeting. Nat. Rev. Drug Discov. 2008, 7, 411–425. [Google Scholar] [CrossRef]
- Maecker, H.; Varfolomeev, E.; Kischkel, F.; Lawrence, D.; LeBlanc, H.; Lee, W.; Hurst, S.; Danilenko, D.; Li, J.; Filvaroff, E.; et al. TWEAK attenuates the transition from innate to adaptive immunity. Cell 2005, 123, 931–944. [Google Scholar] [CrossRef]
- Terra, X.; Gómez, D. External validation of sTWEAK as a prognostic noninvasive biomarker for head and neck squamous cell carcinoma. Head Neck 2014, 36, 1391. [Google Scholar] [CrossRef]
- Avilés-Jurado, F.X.; Terra, X.; Gómez, D.; Flores, J.C.; Raventós, A.; Maymó-Masip, E.; León, X.; Serrano-Gonzalvo, V.; Vendrell, J.; Figuerola, E.; et al. Low blood levels of sTWEAK are related to locoregional failure in head and neck cancer. Eur. Arch. Oto-Rhino-Laryngol. 2015, 272, 1733–1741. [Google Scholar] [CrossRef]
- Ruiz-Plazas, X.; Rodríguez-Gallego, E.; Alves, M.; Altuna-Coy, A.; Lozano-Bartolomé, J.; Portero-Otin, M.; García-Fontgivell, J.F.; Martínez-González, S.; Segarra, J.; Chacón, M.R. Biofluid quantification of TWEAK/Fn14 axis in combination with a selected biomarker panel improves assessment of prostate cancer aggressiveness. J. Transl. Med. 2019, 17, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Enderle, D.; Spiel, A.; Coticchia, C.M.; Berghoff, E.; Mueller, R.; Schlumpberger, M.; Sprenger-Haussels, M.; Shaffer, J.M.; Lader, E.; Skog, J.; et al. Characterization of RNA from exosomes and other extracellular vesicles isolated by a novel spin column-based method. Wiley Interdiscip. Rev. RNA 2012, 3, 286–293. [Google Scholar] [CrossRef] [PubMed]
- Ruivo, C.F.; Adem, B.; Silva, M.; Melo, S.A. The biology of cancer exosomes: Insights and new perspectives. Cancer Res. 2017, 77, 6480–6488. [Google Scholar] [CrossRef] [PubMed]
- Tkach, M.; Théry, C. Communication by Extracellular Vesicles: Where We Are and Where We Need to Go. Cell 2016, 164, 1226–1232. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Ding, M.; Xu, K.; Yang, C.; Mao, L.J. Exosomes in diagnosis and therapy of prostate cancer. Oncotarget 2017, 8, 97693–97700. [Google Scholar] [CrossRef]
- Huang, C.; Luo, W.-F.; Ye, Y.-F.; Lin, L.; Wang, Z.; Luo, M.-H.; Song, Q.-D.; He, X.-P.; Chen, H.-W.; Kong, Y.; et al. Characterization of Inflammatory Factor-Induced Changes in Mesenchymal Stem Cell Exosomes and Sequencing Analysis of Exosomal microRNAs. World J. Stem Cells 2019, 11, 859–890. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Narita, S.; Tsuchiya, N.; Ma, Z.; Numakura, K.; Obara, T.; Tsuruta, H.; Saito, M.; Inoue, T.; Horikawa, Y.; et al. Overexpression of Fn14 promotes androgen-independent prostate cancer progression through MMP-9 and correlates with poor treatment outcome. Carcinogenesis 2011, 32, 1589–1596. [Google Scholar] [CrossRef]
- Yin, J.; Morrissey, C.; Barrett, B.; Corey, E.; Ylaya, K.; Hewitt, S.; Fang, L.; Tillman, H.; Lake, R.; Vessella, R.; et al. AR-Regulated TWEAK-FN14 Pathway Promotes Prostate Cancer Bone Metastasis. Cancer Res. 2014, 74, 4306–4317. [Google Scholar] [CrossRef]
- Hu, Y.; Li, D.; Wu, A.; Qiu, X.; Di, W.; Huang, L.; Qiu, L. TWEAK-stimulated macrophages inhibit metastasis of epithelial ovarian cancer via exosomal shuttling of microRNA. Cancer Lett. 2017, 393, 60–67. [Google Scholar] [CrossRef]
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef]
- Svoronos, A.A.; Engelman, D.M.; Slack, F.J. OncomiR or tumor suppressor? The duplicity of MicroRNAs in cancer. Cancer Res. 2016, 76, 3666–3670. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Li, D.; Hou, S.; Zhu, X. The cancer exosomes: Clinical implications, applications and challenges. Int. J. Cancer 2020, 146, 2946–2959. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Ni, J.; Beretov, J.; Thompson, J.; Graham, P.; Li, Y. Exosomal microRNAs as liquid biopsy biomarkers in prostate cancer. Crit. Rev. Oncol. Hematol. 2020, 145, 4–10. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Yuan, T.; Liang, M.; Du, M.; Xia, S.; Dittmar, R.; Wang, D.; See, W.; Costello, B.A.; Quevedo, F.; et al. Exosomal miR-1290 and miR-375 as Prognostic Markers in Castration-resistant Prostate Cancer. Eur. Urol. 2015, 67, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Endzelinš, E.; Berger, A.; Melne, V.; Bajo-Santos, C.; Sobolevska, K.; Abols, A.; Rodriguez, M.; Šantare, D.; Rudnickiha, A.; Lietuvietis, V.; et al. Detection of circulating miRNAs: Comparative analysis of extracellular vesicle-incorporated miRNAs and cell-free miRNAs in whole plasma of prostate cancer patients. BMC Cancer 2017, 17, 1–13. [Google Scholar] [CrossRef]
- Foj, L.; Ferrer, F.; Serra, M.; Arévalo, A.; Gavagnach, M.; Giménez, N.; Filella, X. Exosomal and Non-Exosomal Urinary miRNAs in Prostate Cancer Detection and Prognosis. Prostate 2017, 77, 573–583. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Ma, Y.Y.; Wang, J.; Zeng, X.F.; Li, R.; Kang, W.; Hao, X.K. Exosomal microRNA-141 is upregulated in the serum of prostate cancer patients. OncoTargets Ther. 2015, 9, 139–148. [Google Scholar] [CrossRef]
- Wani, S.; Kaul, D.; Mavuduru, R.S.; Kakkar, N.; Bhatia, A. Urinary-exosomal miR-2909: A novel pathognomonic trait of prostate cancer severity. J. Biotechnol. 2017, 259, 135–139. [Google Scholar] [CrossRef]
- Barceló, M.; Castells, M.; Bassas, L.; Vigués, F.; Larriba, S. Semen miRNAs Contained in Exosomes as Non-Invasive Biomarkers for Prostate Cancer Diagnosis. Sci. Rep. 2019, 9, 1–16. [Google Scholar] [CrossRef]
- Lee, J.M.; Im, G.-I. PTHrP isoforms have differing effect on chondrogenic differentiation and hypertrophy of mesenchymal stem cells. Biochem. Biophys. Res. Commun. 2012, 421, 819–824. [Google Scholar] [CrossRef]
- Konoshenko, M.Y.; Lekchnov, E.A.; Vlassov, A.V.; Laktionov, P.P. Isolation of Extracellular Vesicles: General Methodologies and Latest Trends. Biomed. Res. Int. 2018, 2018, 8545347. [Google Scholar] [CrossRef] [PubMed]
- Fay, M.P.; Shih, J.H. Permutation tests using estimated distribution functions. J. Am. Stat. Assoc. 1998, 93, 387–396. [Google Scholar] [CrossRef]
- Song, J.; Ouyang, Y.; Che, J.; Li, X.; Zhao, Y.; Yang, K.; Zhao, X.; Chen, Y.; Fan, C.; Yuan, W. Potential value of miR-221/222 as diagnostic, prognostic, and therapeutic biomarkers for diseases. Front. Immunol. 2017, 8, 1–9. [Google Scholar] [CrossRef]
- Goto, Y.; Kojima, S.; Nishikawa, R.; Kurozumi, A.; Kato, M.; Enokida, H.; Matsushita, R.; Yamazaki, K.; Ishida, Y.; Nakagawa, M.; et al. MicroRNA expression signature of castration-resistant prostate cancer: The microRNA-221/222 cluster functions as a tumour suppressor and disease progression marker. Br. J. Cancer 2015, 113, 1055–1065. [Google Scholar] [CrossRef]
- Coarfa, C.; Fiskus, W.; Eedunuri, V.K.; Rajapakshe, K.; Foley, C.; Chew, S.A.; Shah, S.S.; Geng, C.; Shou, J.; Mohamed, J.S.; et al. Comprehensive proteomic profiling identifies the androgen receptor axis and other signaling pathways as targets of microRNAs suppressed in metastatic prostate cancer. Oncogene 2016, 35, 2345–2356. [Google Scholar] [CrossRef]
- Lobo, J.; Gillis, A.J.; van den Berg, A.; Dorssers, L.C.; Belge, G.; Dieckmann, K.P.; Roest, H.P.; van der Laan, L.J.W.; Gietema, J.; Robert, J. Hamilton Looijenga Identification and Validation Model for Informative Liquid Biopsy-Based microRNA Biomarkers: Insights from Germ Cell Tumor In Vitro, In Vivo and Patient-Derived Data João. Cells 2019, 8, 1637. [Google Scholar] [CrossRef]
- Peng, Y.; Croce, C.M. The role of microRNAs in human cancer. Signal Transduct. Target. Ther. 2016, 1, 15004. [Google Scholar] [CrossRef]
- Agaoglu, F.Y.; Kovancilar, M.; Dizdar, Y.; Darendeliler, E.; Holdenrieder, S.; Dalay, N.; Gezer, U. Investigation of miR-21, miR-141, and miR-221 in blood circulation of patients with prostate cancer. Tumor Biol. 2011, 32, 583–588. [Google Scholar] [CrossRef]
- Drabovich, A.P.; Saraon, P.; Jarvi, K.; Diamandis, E.P. Seminal plasma as a diagnostic fluid for male reproductive system disorders. Nat. Rev. Urol. 2014, 11, 278–288. [Google Scholar] [CrossRef]
- Eskra, J.N.; Rabizadeh, D.; Pavlovich, C.P.; Catalona, W.J.; Luo, J. Approaches to urinary detection of prostate cancer. Prostate Cancer Prostatic Dis. 2019, 22, 362–381. [Google Scholar] [CrossRef]
- Song, Q.; An, Q.; Niu, B.; Lu, X.; Zhang, N.; Cao, X. Role of miR-221/222 in Tumor Development and the Underlying Mechanism. J. Oncol. 2019, 2, 7252013. [Google Scholar] [CrossRef]
- Mercatelli, N.; Coppola, V.; Bonci, D.; Miele, F.; Costantini, A.; Guadagnoli, M.; Bonanno, E.; Muto, G.; Frajese, G.V.; De Maria, R.; et al. The inhibition of the highly expressed mir-221 and mir-222 impairs the growth of prostate carcinoma xenografts in mice. PLoS ONE 2008, 3, e4029. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.F.; Ma, J.; Huang, L.; Yi, H.Y.; Zhang, Y.M.; Wu, X.G.; Yan, R.M.; Liang, L.; Zhong, M.; Yu, Y.H.; et al. Cervical squamous cell carcinoma-secreted exosomal miR-221-3p promotes lymphangiogenesis and lymphatic metastasis by targeting VASH1. Oncogene 2019, 38, 1256–1268. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.; Hong, X.; Lai, J.; Cheng, L.; Cheng, Y.; Yao, M.; Wang, R.; Hu, N. Exosomal MicroRNA-221-3p Confers Adriamycin Resistance in Breast Cancer Cells by Targeting PIK3R1. Front. Oncol. 2020, 10, 441. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Bhowmik, S.; Majumdar, S.; Goswami, A.; Chakraborty, J.; Gupta, S.; Aggarwal, S.; Ray, S.; Chatterjee, R.; Bhattacharyya, S.; et al. The exosome encapsulated microRNAs as circulating diagnostic marker for hepatocellular carcinoma with low alpha-fetoprotein. Int. J. Cancer 2020, 147, 2934–2947. [Google Scholar] [CrossRef]
- Larrabeiti-Etxebarria, A.; Lopez-Santillan, M.; Santos-Zorrozua, B.; Lopez-Lopez, E.; Garcia-Orad, A. Systematic review of the potential of MicroRNAs in diffuse large B cell lymphoma. Cancers 2019, 11, 144. [Google Scholar] [CrossRef]
- Wu, Y.; Wei, J.; Zhang, W.; Xie, M.; Wang, X.; Xu, J. Serum exosomal miR-1290 is a potential biomarker for lung adenocarcinoma. OncoTargets Ther. 2020, 13, 7809–7818. [Google Scholar] [CrossRef]
- Kristensen, H.; Thomsen, A.R.; Haldrup, C.; Dyrskjøt, L.; Høyer, S.; Borre, M.; Mouritzen, P.; Ørntoft, T.F.; Sørensen, K.D. Novel diagnostic and prognostic classifiers for prostate cancer identified by genome-wide microRNA profiling. Oncotarget 2016, 7, 30760–30771. [Google Scholar] [CrossRef]
- Kurul, N.O.; Ates, F.; Yilmaz, I.; Narli, G.; Yesildal, C.; Senkul, T. The association of let-7c, miR-21, miR-145, miR-182, and miR-221 with clinicopathologic parameters of prostate cancer in patients diagnosed with low-risk disease. Prostate 2019, 79, 1125–1132. [Google Scholar] [CrossRef]
- Galardi, S.; Mercatelli, N.; Giorda, E.; Massalini, S.; Frajese, G.V.; Ciafrè, S.A.; Farace, M.G. miR-221 and miR-222 expression affects the proliferation potential of human prostate carcinoma cell lines by targeting p27Kip1. J. Biol. Chem. 2007, 282, 23716–23724. [Google Scholar] [CrossRef]
- He, X.Y.; Tan, Z.L.; Mou, Q.; Liu, F.J.; Liu, S.; Yu, C.W.; Zhu, J.; Lv, L.Y.; Zhang, J.; Wang, S.; et al. microRNA-221 enhances MYCN via targeting nemo-like kinase and functions as an oncogene related to poor prognosis in neuroblastoma. Clin. Cancer Res. 2017, 23, 2905–2918. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Yang, Y.; He, Y.; Li, J. The emerging role of Nemo-like kinase (NLK) in the regulation of cancers. Tumor Biol. 2015, 36, 9147–9152. [Google Scholar] [CrossRef] [PubMed]
- Emami, K.H.; Brown, L.G.; Pitts, T.E.M.; Sun, X.; Vessella, R.L.; Corey, E. Nemo-like kinase induces apoptosis and inhibits androgen receptor signaling in prostate cancer cells. Prostate 2009, 69, 1481–1492. [Google Scholar] [CrossRef] [PubMed]
- Kishore, S.; Jaskiewicz, L.; Burger, L.; Hausser, J.; Khorshid, M.; Zavolan, M. A quantitative analysis of CLIP methods for identifying binding sites of RNA-binding proteins. Nat. Methods 2011, 8, 559–567. [Google Scholar] [CrossRef]
- Mullany, L.E.; Herrick, J.S.; Wolff, R.K.; Stevens, J.R.; Samowitz, W.; Slattery, M.L. Transcription factor-microRNA associations and their impact on colorectal cancer survival. Mol. Carcinog. 2017, 56, 2512–2526. [Google Scholar] [CrossRef]
- Chen, W.S.; Chen, C.C.; Chen, L.L.; Lee, C.C.; Huang, T.S. Secreted heat shock protein 90α (HSP90α) induces nuclear factor-κB-mediated TCF12 protein expression to down-regulate E-cadherin and to enhance colorectal cancer cell migration and invasion. J. Biol. Chem. 2013, 288, 9001–9010. [Google Scholar] [CrossRef]
- Chen, Q.B.; Liang, Y.K.; Zhang, Y.Q.; Jiang, M.Y.; Han, Z.D.; Liang, Y.X.; Wan, Y.P.; Yin, J.; He, H.C.; Zhong, W. De Decreased expression of TCF12 contributes to progression and predicts biochemical recurrence in patients with prostate cancer. Tumor Biol. 2017, 39. [Google Scholar] [CrossRef]
- Zhang, M.; Xiao, X.; Xiong, D.; Liu, Q. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. J. Artif. Intell. Res. 2014, 50, 1–30. [Google Scholar] [CrossRef]
- Vendrell, J.; Maymó-Masip, E.; Tinahones, F.; García-España, A.; Megia, A.; Caubet, E.; García-Fuentes, E.; Chacón, M.R. Tumor necrosis-like weak inducer of apoptosis as a proinflammatory cytokine in human adipocyte cells: Up-regulation in severe obesity is mediated by inflammation but not hypoxia. J. Clin. Endocrinol. Metab. 2010, 95, 2983–2992. [Google Scholar] [CrossRef]
- Riffo-Campos, Á.L.; Riquelme, I.; Brebi-Mieville, P. Tools for sequence-based miRNA target prediction: What to choose? Int. J. Mol. Sci. 2016, 17, 1987. [Google Scholar] [CrossRef]
- Willkomm, S.; Zander, A.; Grohmann, D. Drug Target miRNA. Methods Mol. Biol. 2017, 1517, 291–304. [Google Scholar] [CrossRef] [PubMed]
- Sticht, C.; De La Torre, C.; Parveen, A.; Gretz, N. Mirwalk: An online resource for prediction of microrna binding sites. PLoS ONE 2018, 13, e0206239. [Google Scholar] [CrossRef] [PubMed]



| Patient’s Characteristics | Mean ± SD | N |
|---|---|---|
| Age (years) | 63.5 ± 6.35 | 97 |
| Prostatic Volume (c.c) | 47.49 ± 23.09 | 97 |
| Testosterone (nmol/L) | 14.37 ± 5.07 | 97 |
| Total PSA (ng/mL) | 9.57 ± 7.92 | 97 |
| N (%) | ||
| BMI (kg/m2) | <25 | 25 (25.8) |
| 25 ≤ x ≤ 29.99 | 50 (51.5) | |
| ≥30 | 19 (19.6) | |
| Total PSA (ng/mL) | ||
| <4 | 8 (8.2) | |
| 4 ≤ x < 10 | 60 (61.9) | |
| ≥ 10 | 29 (29.9) | |
| ISUP-GG | ||
| Low Risk | Group I | 32 (33.0) |
| Group II | 25 (25.8) | |
| High Risk | Group III | 23 (23.7) |
| Group IV | 10 (10.3) | |
| Group V | 7 (7.2) | |
| T pathological stage | ||
| ≤T2a | 68 (70.1) | |
| T3,T4 | 29 (29.9) | |
| N pathological stage | ||
| NX | 57 (58.8) | |
| N0 | 34 (35.1) | |
| N1 | 6 (6.2) | |
| ISUP GG Classification | |||
|---|---|---|---|
| Patient’s Stratification | Low-Risk | High-Risk | |
| (Group I and II) | (Group III, IV and V) | ||
| N = 57 | N = 40 | ||
| Mean ± SD | Mean ± SD | p-Value | |
| Anthropometric parameters | |||
| Age (years) | 62.46 ± 6.74 | 64.96 ± 5.52 | 0.066 |
| BMI (kg/m2) | 27.97 ± 4.07 | 27.64 ± 3.46 | 0.718 |
| Prostatic volume (c.c) | 48.68 ± 24.56 | 45.81 ± 21 | 0.687 |
| Glycemic profile | |||
| Glucose (mmol/L) | 5.82 ± 1.1 | 6.29 ± 2.26 | 0.388 |
| Insulin (pmol/L) | 89.36 ± 58.39 | 87.42 ± 47.09 | 0.841 |
| HOMA-IR | 3.46 ± 2.58 | 3.67 ± 2.71 | 0.841 |
| HbA1c (%) | 5.74 ± 0.64 | 5.92 ± 0.84 | 0.364 |
| Lipid profile | |||
| Cholesterol (mmol/L) | 5.03 ± 1.06 | 5.03 ± 1.1 | 0.957 |
| HDL cholesterol (mmol/L) | 1.49 ± 0.73 | 1.42 ± 0.39 | 0.672 |
| LDL cholesterol (mmol/L) | 3.28 ± 1.3 | 3 ± 0.88 | 0.503 |
| Triglycerides (mmol/L) | 1.36 ± 0.74 | 1.55 ± 0.96 | 0.711 |
| Hepatic profile | |||
| AST (µkat/L) | 0.39 ± 0.19 | 0.33 ± 0.07 | 0.171 |
| ALT (µkat/L) | 0.42 ± 0.22 | 0.36 ± 0.11 | 0.402 |
| GGT (µkat/L) | 0.7 ± 0.85 | 0.65 ± 0.48 | 0.887 |
| Renal profile | |||
| Uric acid (µmol/L) | 368.05 ± 83.1 | 456.2 ± 529.55 | 0.376 |
| Urea (mmol/L) | 14.26 ± 3.23 | 14.8 ± 5.12 | 0.808 |
| Creatinine (μmol/L) | 85.83 ± 18.44 | 80.05 ± 13.97 | 0.072 |
| Hormonal profile | |||
| SHBG (nmol/L) | 46.13 ± 52.46 | 40.02 ± 16.36 | 0.814 |
| Testosterone (nmol/L) | 14.93 ± 4.62 | 13.55 ± 5.63 | 0.101 |
| Tumoral marker | |||
| Total PSA (μg/L) | 7.71 ± 4.8 | 12.24 ± 10.43 | 0.007 |
| Biofluid Biomarker profile | |||
| Semen cytokines (pg/mg of total protein) | |||
| sTWEAK | 989.62 ± 685.75 | 617.25 ± 447.57 | 0.009 |
| Exo-oncomiRNAs in semen—Relative expression levels | |||
| miR-221-3p | 0.75 ± 0.6 | 2.17 ± 1.7 | 0.002 |
| miR-222-3p | 2.01 ± 2.79 | 3.79 ± 2.92 | 0.006 |
| miR-31-5p | 1.05 ± 0.73 | 2.75 ± 2.27 | 0.004 |
| Exo-oncomiRNAs in urine—Relative expression levels | |||
| miR-193-3p | 0.12 ± 0.12 | 0.06 ± 0.05 | 0.037 |
| miR-423-5p | 0.05 ± 0.05 | 0.04 ± 0.03 | 0.034 |
| 95% CI | ||||||||
|---|---|---|---|---|---|---|---|---|
| ROC Model | AUC | Error | p-Value | Lower | Upper | Sensivity (%) | Specificity (%) | % Correct Diagnosis |
| Age | 0.610 | 0.058 | 0.066 | 0.496 | 0.724 | 85 | 75.4 | 62.9 |
| Total PSA | 0.662 | 0.056 | 0.007 | 0.552 | 0.772 | 85 | 31.6 | 63.9 |
| sTWEAK | 0.708 | 0.072 | 0.009 | 0.567 | 0.848 | 85.7 | 52.8 | 71.9 |
| exo-oncomiR-221-3p | 0.79 | 0.078 | 0.002 | 0.638 | 0.943 | 86.7 | 55.6 | 78.6 |
| exo-oncomiR-222-3p | 0.758 | 0.08 | 0.006 | 0.601 | 0.915 | 86.7 | 74.1 | 66.7 |
| exo-oncomiR-31-5p | 0.768 | 0.082 | 0.004 | 0.607 | 0.929 | 86.7 | 48.1 | 76.2 |
| Total PSA + Age | 0.704 | 0.054 | 0.001 | 0.597 | 0.810 | 85 | 70.2 | 67 |
| Total PSA + sTWEAK | 0.738 | 0.072 | 0.003 | 0.597 | 0.879 | 85.7 | 47.2 | 71.9 |
| Total PSA + exo-oncomiR-221-3p | 0.864 | 0.063 | <0.001 | 0.74 | 0.998 | 86.7 | 55.6 | 83.3 |
| Total PSA + exo-oncomiR-222-3p | 0.78 | 0.071 | 0.003 | 0.641 | 0.919 | 86.7 | 55.6 | 73.8 |
| Total PSA + exo-oncomiR-31-5p | 0.832 | 0.07 | <0.001 | 0.695 | 0.969 | 86.7 | 51.9 | 81 |
| sTWEAK + Age | 0.709 | 0.069 | 0.009 | 0.574 | 0.844 | 85.7 | 50 | 66.7 |
| sTWEAK + exo-oncomiR-221-3p | 0.841 | 0.073 | <0.001 | 0.698 | 0.983 | 85.7 | 69.2 | 82.5 |
| sTWEAK + exo-oncomiR-222-3p | 0.745 | 0.086 | 0.012 | 0.576 | 0.913 | 85.7 | 42.3 | 70 |
| sTWEAK + exo-oncomiR-31-5p | 0.808 | 0.077 | 0.001 | 0.657 | 0.958 | 85.7 | 61.5 | 77.5 |
| exo-oncomiR-221-3p + Age | 0.802 | 0.077 | 0.001 | 0.651 | 0.954 | 86.7 | 33.3 | 76.2 |
| exo-oncomiR-221-3p + exo-oncomiR-222-3p | 0.802 | 0.078 | 0.001 | 0.65 | 0.955 | 86.7 | 63 | 76.2 |
| exo-oncomiR-221-3p + exo-oncomiR-31-5p | 0.8 | 0.079 | 0.001 | 0.646 | 0.954 | 86.7 | 55.6 | 81 |
| exo-oncomiR-222-3p + Age | 0.751 | 0.081 | 0.008 | 0.592 | 0.909 | 86.7 | 66.7 | 73.8 |
| exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.8 | 0.077 | 0.001 | 0.649 | 0.951 | 86.7 | 55.6 | 81 |
| exo-oncomiR-31-5p + Age | 0.778 | 0.078 | 0.003 | 0.625 | 0.930 | 86.7 | 44.4 | 73.8 |
| Total PSA + sTWEAK + Age | 0.746 | 0.067 | 0.002 | 0.614 | 0.878 | 85.7 | 44.4 | 73.7 |
| Total PSA + sTWEAK + exo-oncomiR-221-3p | 0.863 | 0.068 | <0.001 | 0.73 | 0.996 | 85.7 | 69.2 | 85 |
| Total PSA + sTWEAK + exo-oncomiR-222-3p | 0.758 | 0.086 | 0.008 | 0.59 | 0.926 | 85.7 | 46.2 | 75 |
| Total PSA + sTWEAK + exo-oncomiR-31-5p | 0.824 | 0.076 | 0.001 | 0.675 | 0.974 | 85.7 | 73.1 | 77.5 |
| Total PSA + exo-oncomiR-221-3p + Age | 0.889 | 0.056 | <0.001 | 0.780 | 0.998 | 85.7 | 37 | 83.3 |
| Total PSA + exo-oncomiR-221-3p + exo-oncomiR-222-3p | 0.872 | 0.06 | <0.001 | 0.755 | 0.988 | 86.7 | 59.3 | 83.3 |
| Total PSA + exo-oncomiR-221-3p + exo-oncomiR-31-5p | 0.854 | 0.067 | <0.001 | 0.724 | 0.985 | 86.7 | 51.9 | 83.3 |
| Total PSA + exo-oncomiR-222-3p + Age | 0.840 | 0.064 | <0.001 | 0.714 | 0.965 | 86.7 | 37 | 83.3 |
| Total PSA + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.849 | 0.069 | <0.001 | 0.713 | 0.985 | 86.7 | 59.3 | 83.3 |
| Total PSA + exo-oncomiR-31-5p + Age | 0.862 | 0.061 | <0.001 | 0.743 | 0.981 | 86.7 | 37 | 83.3 |
| sTWEAK + exo-oncomiR-221-3p + Age | 0.854 | 0.067 | <0.001 | 0.723 | 0.986 | 85.7 | 23.7 | 77.5 |
| sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-222-3p | 0.857 | 0.069 | <0.001 | 0.721 | 0.993 | 85.7 | 76.9 | 87.5 |
| sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-31-5p | 0.841 | 0.073 | <0.001 | 0.698 | 0.983 | 85.7 | 69.2 | 82.5 |
| sTWEAK + exo-oncomiR-222-3p + Age | 0.764 | 0.078 | 0.006 | 0.611 | 0.917 | 85.7 | 50 | 72.5 |
| sTWEAK + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.83 | 0.073 | 0.001 | 0.687 | 0.972 | 85.7 | 53.8 | 82.5 |
| sTWEAK + exo-oncomiR-31-5p + Age | 0.821 | 0.074 | 0.001 | 0.677 | 0.966 | 85.7 | 34.6 | 75 |
| exo-oncomiR-221-3p + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.807 | 0.076 | 0.001 | 0.658 | 0.956 | 86.7 | 51.9 | 83.3 |
| exo-oncomiR-221-3p + exo-oncomiR-222-3p + Age | 0.820 | 0.074 | 0.001 | 0.675 | 0.965 | 86.7 | 25.9 | 76.2 |
| exo-oncomiR-221-3p + exo-oncomiR-31-5p + Age | 0.812 | 0.074 | 0.001 | 0.668 | 0.957 | 86.7 | 37 | 78.6 |
| exo-oncomiR-222-3p + exo-oncomiR-31-5p + Age | 0.802 | 0.075 | 0.001 | 0.655 | 0.950 | 86.7 | 44 | 78.6 |
| Total PSA + sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-222-3p | 0.86 | 0.071 | <0.001 | 0.721 | 0.999 | 85.7 | 69.2 | 85 |
| Total PSA + sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-31-5p | 0.86 | 0.069 | <0.001 | 0.724 | 0.995 | 85.7 | 69.2 | 85 |
| Total PSA + sTWEAK + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.83 | 0.076 | 0.001 | 0.682 | 0.978 | 85.7 | 65.4 | 82.5 |
| Age + Total PSA + sTWEAK + exo-oncomiR-221-3p | 0.879 | 0.62 | <0.001 | 0.757 | 1 | 85.7 | 23.1 | 85 |
| Age + Total PSA + sTWEAK + exo-oncomiR-222-3p | 0.808 | 0.074 | 0.001 | 0.662 | 0.953 | 85.7 | 50 | 82.5 |
| Age + Total PSA + sTWEAK + exo-oncomiR-31-5p | 0.849 | 0.069 | <0.001 | 0.715 | 0.983 | 85.7 | 53.2 | 82.5 |
| Age + Total PSA + exo-oncomiR-221-3p + exo-oncomiR-222-3p | 0.894 | 0.053 | <0.001 | 0.789 | 0.998 | 86.7 | 56.7 | 83.3 |
| Age + Total PSA + exo-oncomiR-221-3p + exo-oncomiR-31-5p | 0.879 | 0.059 | <0.001 | 0.764 | 0.994 | 86.7 | 54.3 | 83.3 |
| Age + Total PSA + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.867 | 0.059 | <0.001 | 0.752 | 0.982 | 86.7 | 49.2 | 81 |
| Age + sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-222-3p | 0.868 | 0.061 | <0.001 | 0.748 | 0.988 | 86.7 | 46.5 | 80 |
| Age + sTWEAK + exo-onxomiR-221-3p + exo-oncomiR-31-5p | 0.857 | 0.067 | <0.001 | 0.726 | 0.988 | 85.7 | 76.9 | 80 |
| Age + sTWEAK + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.832 | 0.070 | 0.001 | 0.695 | 0.969 | 86.7 | 46.5 | 80 |
| Age + exo-oncomiR-221-3p + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.820 | 0.072 | 0.001 | 0.678 | 0.962 | 86.7 | 48.1 | 81 |
| Total PSA + exo-oncomiR-221-3p + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.874 | 0.061 | <0.001 | 0.754 | 0.995 | 86.7 | 55.6 | 83.3 |
| sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.86 | 0.068 | <0.001 | 0.726 | 0.994 | 85.7 | 73.1 | 85 |
| Total PSA + sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.865 | 0.069 | <0.001 | 0.73 | 1 | 85.7 | 69.2 | 85 |
| Age + Total PSA + sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-222-3p | 0.879 | 0.062 | <0.001 | 0.757 | 1 | 86.7 | 70.4 | 87.5 |
| Age + Total PSA + sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-31-5p | 0.879 | 0.062 | <0.001 | 0.758 | 1 | 86.7 | 57.9 | 85 |
| Age + Total PSA + sTWEAK + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.857 | 0.065 | <0.001 | 0.729 | 0.985 | 85.7 | 58.3 | 85 |
| Age + Total PSA + exo-oncomiR-221-3p + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.896 | 0.053 | <0.001 | 0.793 | 0.999 | 86.7 | 57.9 | 83.3 |
| Age + sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.874 | 0.061 | <0.001 | 0.755 | 0.992 | 85.7 | 63.8 | 82.5 |
| Age + Total PSA + sTWEAK + exo-oncomiR-221-3p + exo-oncomiR-222-3p + exo-oncomiR-31-5p | 0.879 | 0.062 | <0.001 | 0.757 | 1 | 85.7 | 68.9 | 87.5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruiz-Plazas, X.; Altuna-Coy, A.; Alves-Santiago, M.; Vila-Barja, J.; García-Fontgivell, J.F.; Martínez-González, S.; Segarra-Tomás, J.; Chacón, M.R. Liquid Biopsy-Based Exo-oncomiRNAs Can Predict Prostate Cancer Aggressiveness. Cancers 2021, 13, 250. https://doi.org/10.3390/cancers13020250
Ruiz-Plazas X, Altuna-Coy A, Alves-Santiago M, Vila-Barja J, García-Fontgivell JF, Martínez-González S, Segarra-Tomás J, Chacón MR. Liquid Biopsy-Based Exo-oncomiRNAs Can Predict Prostate Cancer Aggressiveness. Cancers. 2021; 13(2):250. https://doi.org/10.3390/cancers13020250
Chicago/Turabian StyleRuiz-Plazas, Xavier, Antonio Altuna-Coy, Marta Alves-Santiago, José Vila-Barja, Joan Francesc García-Fontgivell, Salomé Martínez-González, José Segarra-Tomás, and Matilde R. Chacón. 2021. "Liquid Biopsy-Based Exo-oncomiRNAs Can Predict Prostate Cancer Aggressiveness" Cancers 13, no. 2: 250. https://doi.org/10.3390/cancers13020250
APA StyleRuiz-Plazas, X., Altuna-Coy, A., Alves-Santiago, M., Vila-Barja, J., García-Fontgivell, J. F., Martínez-González, S., Segarra-Tomás, J., & Chacón, M. R. (2021). Liquid Biopsy-Based Exo-oncomiRNAs Can Predict Prostate Cancer Aggressiveness. Cancers, 13(2), 250. https://doi.org/10.3390/cancers13020250







