Reproducibility of mRNA-Based Testing of ESR1, PGR, ERBB2, and MKI67 Expression in Invasive Breast Cancer—A Europe-Wide External Quality Assessment
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Cohort
2.2. Immunohistochemical (IHC) Expression of ER, PgR, HER2, and Ki-67 and IHC Subtyping
- Samples with ER or PgR IHC ≥1% positive stained cells, regardless of staining intensity, were classified as positive (<10% as low positive); samples with <1% cells staining at any intensity were classified as negative [8,9,20,21]. ER and/or PgR positivity were summarized as hormone receptor positivity (HR+).
2.3. mRNA-Based Testing of ESR1, PGR, ERBB2, and MKI67 in the Coordinating Center
2.4. EQA Implementation
2.5. Re-Evaluation of Discrepant Cases
2.6. Statistical Analysis
3. Results
3.1. Comparison of Binary STRAT4 Results
3.2. Re-Evaluation of Discrepant Cases
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Perou, C.M.; Sorlie, T.; Eisen, M.B.; van de Rijn, M.; Jeffrey, S.S.; Rees, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A.; et al. Molecular portraits of human breast tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef] [PubMed]
- Sorlie, T.; Perou, C.M.; Tibshirani, R.; Aas, T.; Geisler, S.; Johnsen, H.; Hastie, T.; Eisen, M.B.; van de Rijn, M.; Jeffrey, S.S.; et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl. Acad. Sci. USA 2001, 98, 10869–10874. [Google Scholar] [CrossRef] [Green Version]
- Paik, S.; Shak, S.; Tang, G.; Kim, C.; Baker, J.; Cronin, M.; Baehner, F.L.; Walker, M.G.; Watson, D.; Park, T.; et al. A Multigene Assay to Predict Recurrence of Tamoxifen-Treated, Node-Negative Breast Cancer. N. Engl. J. Med. 2004, 351, 2817–2826. [Google Scholar] [CrossRef] [Green Version]
- Curigliano, G.; Burstein, H.J.; Winer, E.P.; Gnant, M.; Dubsky, P.; Loibl, S.; Colleoni, M.; Regan, M.M.; Piccart-Gebhart, M.; Senn, H.J.; et al. De-escalating and escalating treatments for early-stage breast cancer: The St. Gallen International Expert Consensus Conference on the Primary Therapy of Early Breast Cancer 2017. Ann. Oncol. 2017, 28, 1700–1712. [Google Scholar] [CrossRef] [PubMed]
- Goldhirsch, A.; Winer, E.P.; Coates, A.S.; Gelber, R.D.; Piccart-Gebhart, M.; Thurlimann, B.; Senn, H.J.; Panel, M. Personalizing the treatment of women with early breast cancer: Highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann. Oncol. Off. J. Eur. Soc. Med Oncol. 2013, 24, 2206–2223. [Google Scholar] [CrossRef]
- Dowsett, M.; Nielsen, T.O.; A’Hern, R.; Bartlett, J.; Coombes, R.C.; Cuzick, J.; Ellis, M.; Henry, N.L.; Hugh, J.C.; Lively, T.; et al. Assessment of Ki67 in breast cancer: Recommendations from the International Ki67 in Breast Cancer working group. J. Natl. Cancer Inst. 2011, 103, 1656–1664. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gonzalez-Ericsson, P.I.; Stovgaard, E.S.; Sua, L.F.; Reisenbichler, E.; Kos, Z.; Carter, J.M.; Michiels, S.; Le Quesne, J.; Nielsen, T.O.; Lænkholm, A.-V.; et al. The path to a better biomarker: Application of a risk management framework for the implementation of PD-L1 and TILs as immuno-oncology biomarkers into breast cancer clinical trials and daily practice. J. Pathol. 2020, 250, 667–684. [Google Scholar] [CrossRef] [Green Version]
- Hammond, M.E.; Hayes, D.F.; Dowsett, M.; Allred, D.C.; Hagerty, K.L.; Badve, S.; Fitzgibbons, P.L.; Francis, G.; Goldstein, N.S.; Hayes, M.; et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer (unabridged version). Arch. Pathol. Lab. Med. 2010, 134, e48–e72. [Google Scholar] [CrossRef]
- Allison, K.H.; Hammond, M.E.H.; Dowsett, M.; McKernin, S.E.; Carey, L.A.; Fitzgibbons, P.L.; Hayes, D.F.; Lakhani, S.R.; Chavez-MacGregor, M.; Perlmutter, J.; et al. Estrogen and Progesterone Receptor Testing in Breast Cancer: ASCO/CAP Guideline Update. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2020, 38, 1346–1366. [Google Scholar] [CrossRef]
- Wolff, A.C.; Hammond, M.E.; Schwartz, J.N.; Hagerty, K.L.; Allred, D.C.; Cote, R.J.; Dowsett, M.; Fitzgibbons, P.L.; Hanna, W.M.; Langer, A.; et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. Arch. Pathol. Lab. Med. 2007, 131, 18–43. [Google Scholar] [CrossRef]
- Wolff, A.C.; Hammond, M.E.; Hicks, D.G.; Dowsett, M.; McShane, L.M.; Allison, K.H.; Allred, D.C.; Bartlett, J.M.; Bilous, M.; Fitzgibbons, P.; et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2013, 31, 3997–4013. [Google Scholar] [CrossRef] [PubMed]
- Wolff, A.C.; Hammond, M.E.H.; Allison, K.H.; Harvey, B.E.; Mangu, P.B.; Bartlett, J.M.S.; Bilous, M.; Ellis, I.O.; Fitzgibbons, P.; Hanna, W.; et al. Human Epidermal Growth Factor Receptor 2 Testing in Breast Cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. Arch. Pathol. Lab. Med. 2018, 142, 1364–1382. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Viale, G.; Giobbie-Hurder, A.; Regan, M.M.; Coates, A.S.; Mastropasqua, M.G.; Dell’Orto, P.; Maiorano, E.; MacGrogan, G.; Braye, S.G.; Ohlschlegel, C.; et al. Prognostic and predictive value of centrally reviewed Ki-67 labeling index in postmenopausal women with endocrine-responsive breast cancer: Results from Breast International Group Trial 1-98 comparing adjuvant tamoxifen with letrozole. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2008, 26, 5569–5575. [Google Scholar] [CrossRef] [PubMed]
- Erber, R.; Gluz, O.; Brunner, N.; Kreipe, H.H.; Pelz, E.; Kates, R.; Bartels, A.; Huober, J.; Mohrmann, S.; Moustafa, Z.; et al. Predictive role of HER2/neu, topoisomerase-II-alpha, and tissue inhibitor of metalloproteinases (TIMP-1) for response to adjuvant taxane-based chemotherapy in patients with intermediate-risk breast cancer: Results from the WSG-AGO EC-Doc trial. Breast Cancer Res. Treat. 2015, 150, 279–288. [Google Scholar] [CrossRef] [PubMed]
- Polley, M.Y.; Leung, S.C.; McShane, L.M.; Gao, D.; Hugh, J.C.; Mastropasqua, M.G.; Viale, G.; Zabaglo, L.A.; Penault-Llorca, F.; Bartlett, J.M.; et al. An international Ki67 reproducibility study. J. Natl. Cancer Inst. 2013, 105, 1897–1906. [Google Scholar] [CrossRef] [PubMed]
- Hammond, M.E.; Hayes, D.F.; Dowsett, M.; Allred, D.C.; Hagerty, K.L.; Badve, S.; Fitzgibbons, P.L.; Francis, G.; Goldstein, N.S.; Hayes, M.; et al. American Society of Clinical Oncology/College Of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2010, 28, 2784–2795. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goldhirsch, A.; Wood, W.C.; Gelber, R.D.; Coates, A.S.; Thurlimann, B.; Senn, H.J. Meeting highlights: Updated international expert consensus on the primary therapy of early breast cancer. J. Clin. Oncol. 2003, 21, 3357–3365. [Google Scholar] [CrossRef]
- Goldhirsch, A.; Glick, J.H.; Gelber, R.D.; Coates, A.S.; Thurlimann, B.; Senn, H.J. Meeting highlights: International expert consensus on the primary therapy of early breast cancer 2005. Ann. Oncol. Off. J. Eur. Soc. Med Oncol. 2005, 16, 1569–1583. [Google Scholar] [CrossRef]
- Cheang, M.C.U.; Chia, S.K.; Voduc, D.; Gao, D.; Leung, S.; Snider, J.; Watson, M.; Davies, S.; Bernard, P.S.; Parker, J.S.; et al. Ki67 index, HER2 status, and prognosis of patients with luminal B breast cancer. J. Natl. Cancer Inst. 2009, 101, 736–750. [Google Scholar] [CrossRef] [Green Version]
- Landmann, A.; Farrugia, D.J.; Zhu, L.; Diego, E.J.; Johnson, R.R.; Soran, A.; Dabbs, D.J.; Clark, B.Z.; Puhalla, S.L.; Jankowitz, R.C.; et al. Low Estrogen Receptor (ER)-Positive Breast Cancer and Neoadjuvant Systemic Chemotherapy: Is Response Similar to Typical ER-Positive or ER-Negative Disease? Am. J. Clin. Pathol. 2018, 150, 34–42. [Google Scholar] [CrossRef]
- Chen, T.; Zhang, N.; Moran, M.S.; Su, P.; Haffty, B.G.; Yang, Q. Borderline ER-Positive Primary Breast Cancer Gains No Significant Survival Benefit From Endocrine Therapy: A Systematic Review and Meta-Analysis. Clin. Breast Cancer 2018, 18, 1–8. [Google Scholar] [CrossRef]
- AGOEV. Guidelines Breast Version 2020 1D. Available online: https://www.ago-online.de/fileadmin/ago-online/downloads/_leitlinien/kommission_mamma/2020/PDF_DE/2020D%2004_Pathologie_mit%20Literatur.pdf (accessed on 21 February 2021).
- Nielsen, T.O.; Leung, S.C.Y.; Rimm, D.L.; Dodson, A.; Acs, B.; Badve, S.; Denkert, C.; Ellis, M.J.; Fineberg, S.; Flowers, M.; et al. Assessment of Ki67 in Breast Cancer: Updated Recommendations from the International Ki67 in Breast Cancer Working Group. J. Natl. Cancer Inst. 2021, 113, 808–819. [Google Scholar] [CrossRef] [PubMed]
- Laible, M.; Schlombs, K.; Kaiser, K.; Veltrup, E.; Herlein, S.; Lakis, S.; Stöhr, R.; Eidt, S.; Hartmann, A.; Wirtz, R.M.; et al. Technical validation of an RT-qPCR in vitro diagnostic test system for the determination of breast cancer molecular subtypes by quantification of ERBB2, ESR1, PGR and MKI67 mRNA levels from formalin-fixed paraffin-embedded breast tumor specimens. BMC Cancer 2016, 16, 398. [Google Scholar] [CrossRef] [Green Version]
- Gupta, S.; Mani, N.R.; Carvajal-Hausdorf, D.E.; Bossuyt, V.; Ho, K.; Weidler, J.; Wong, W.; Rhees, B.; Bates, M.; Rimm, D.L. Macrodissection prior to closed system RT-qPCR is not necessary for estrogen receptor and HER2 concordance with IHC/FISH in breast cancer. Lab. Investig. 2018, 98, 1076–1083. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, N.C.; Wong, W.; Ho, K.E.; Chu, V.C.; Rizo, A.; Davenport, S.; Kelly, D.; Makar, R.; Jassem, J.; Duchnowska, R.; et al. Comparison of central laboratory assessments of ER, PR, HER2, and Ki67 by IHC/FISH and the corresponding mRNAs (ESR1, PGR, ERBB2, and MKi67) by RT-qPCR on an automated, broadly deployed diagnostic platform. Breast Cancer Res. Treat. 2018, 172, 327–338. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Filipits, M.; Rudas, M.; Singer, C.; Bago-Horvath, Z.; Greil, R.; Balic, M.; Lax, S.; Wu, N.; Zhao, S.; Weidler, J.; et al. mRNA Expression of ER, PR, HER2 and Ki67 are Concordant with Central IHC and Predict Clinical Outcome: A Validation Study from the ABCSG-6 Biomarker Cohort. Ann. Oncol. 2018, 29 (Suppl. 8), 1339. [Google Scholar] [CrossRef]
- Remmele, W.; Stegner, H.E. Recommendation for uniform definition of an immunoreactive score (IRS) for immunohistochemical estrogen receptor detection (ER-ICA) in breast cancer tissue. Pathologe 1987, 8, 138–140. [Google Scholar] [PubMed]
- Bethune, G.C.; Veldhuijzen van Zanten, D.; MacIntosh, R.F.; Rayson, D.; Younis, T.; Thompson, K.; Barnes, P.J. Impact of the 2013 American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 (HER2) testing of invasive breast carcinoma: A focus on tumours assessed as ‘equivocal’ for HER2 gene amplification by fluorescence in-situ hybridization. Histopathology 2015, 67, 880–887. [Google Scholar] [CrossRef] [PubMed]
- Viale, G.; On the behalf of the International Breast Cancer Study Group; Regan, M.M.; Mastropasqua, M.G.; Maffini, F.; Maiorano, E. Predictive Value of Tumor Ki-67 Expression in Two Randomized Trials of Adjuvant Chemoendocrine Therapy for Node-Negative Breast Cancer. JNCI J. Natl. Cancer Inst. 2008, 100, 207–212. [Google Scholar] [CrossRef] [Green Version]
- R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria, 2019. Available online: https://www.R-project.org/ (accessed on 21 February 2021).
- Xpert Breast Cancer STRAT4 (CE-IVD) package insert. 301-4981, Rev. C March 2020.
- Wirtz, R.M.; Sihto, H.; Isola, J.; Heikkilä, P.; Kellokumpu-Lehtinen, P.L.; Auvinen, P.; Turpeenniemi-Hujanen, T.; Jyrkkiö, S.; Lakis, S.; Schlombs, K.; et al. Biological subtyping of early breast cancer: A study comparing RT-qPCR with immunohistochemistry. Breast Cancer Res. Treat. 2016, 157, 437–446. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harris, L.; Fritsche, H.; Mennel, R.; Norton, L.; Ravdin, P.; Taube, S.; Somerfield, M.R.; Hayes, D.F.; Bast, R.C., Jr. American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2007, 25, 5287–5312. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kirchner, M.; Glade, J.; Lehmann, U.; Merkelbach-Bruse, S.; Hummel, M.; Lehmann, A.; Trautmann, M.; Kumbrink, J.; Jung, A.; Dietmaier, W.; et al. NTRK testing: First results of the QuiP-EQA scheme and a comprehensive map of NTRK fusion variants and their diagnostic coverage by targeted RNA-based NGS assays. Genes Chromosomes Cancer 2020, 59, 445–453. [Google Scholar] [CrossRef]
- Jöhrens, K.; von Wasielewski, R.; Kreipe, H.H.; Forberger, A.; Jurmeister, P.; Dietel, M.; Stenzinger, A.; Fischer, J. Quality assurance in diagnostic in situ hybridization-experience of QuIP. Pathologe 2020, 41, 614–620. [Google Scholar] [CrossRef]
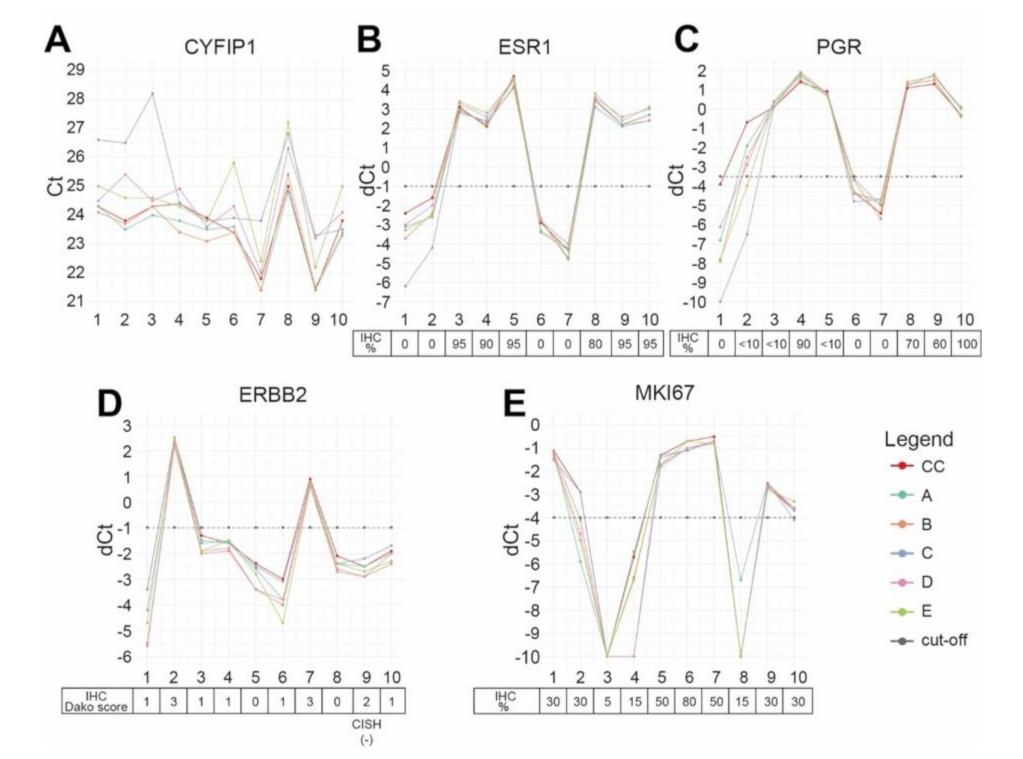
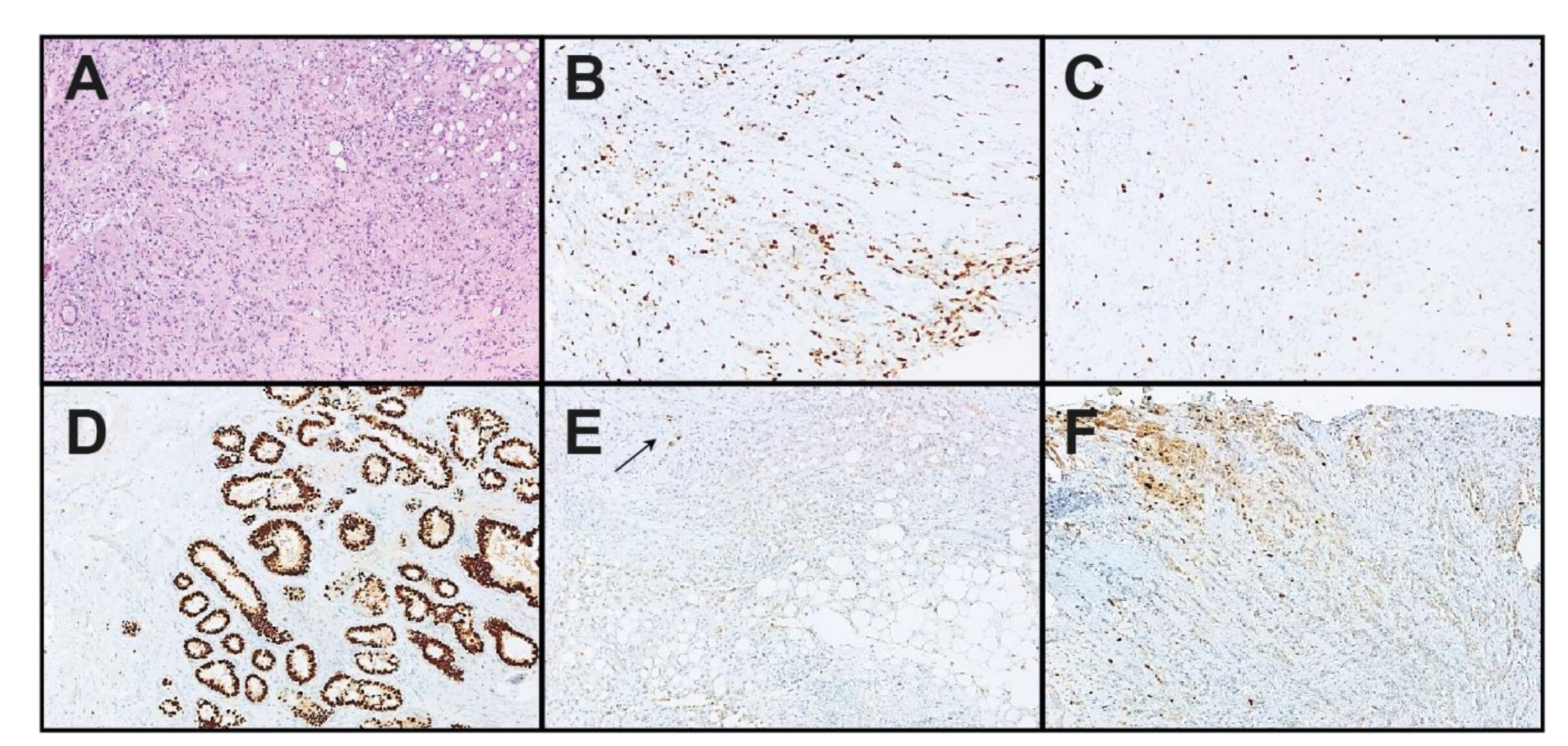
| Sample Information | ER IHC | PgR IHC | HER2 IHC/CISH | Ki-67 IHC | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EQA Sample ID | Subtype by IHC | ER IHC Status | ER IHC Percentage | ER IHC IRS | PgR IHC Status | PgR IHC Percentage | PgR IHC IRS | HER2 IHC/CISH Status | HER2 Dako Score (IHC) | HER2 CISH | Ki-67 IHC Status | Ki-67 IHC Percentage (%) |
| 1 | TNBC | negative | 0 | 0 | negative | 0 | 0 | negative | 1 | n.p. | positive | 30 |
| 2 | HER2+ | negative | 0 | 0 | positive (low) | <10 | 3 | positive | 3 | n.p. | positive | 30 |
| 3 | HR+, Ki-67 low | positive | 95 | 12 | positive (low) | <10 | 1 | negative | 1 | n.p. | negative | 5 |
| 4 | HR+, Ki-67 low | positive | 90 | 12 | positive | 90 | 12 | negative | 1 | n.p. | negative | 15 |
| 5 | HR+, Ki-67 high | positive | 95 | 12 | positive (low) | <10 | 3 | negative | 0 | n.p. | positive | 50 |
| 6 | TNBC | negative | 0 | 0 | negative | 0 | 0 | negative | 1 | n.p. | positive | 80 |
| 7 | HER2+ | negative | 0 | 0 | negative | 0 | 0 | positive | 3 | n.p. | positive | 50 |
| 8 | HR+, Ki-67 low | positive | 80 | 12 | positive | 70 | 6 | negative | 0 | n.p. | negative | 15 |
| 9 | HR+, Ki-67 high | positive | 95 | 12 | positive | 60 | 9 | negative | 2 | Negative (HER2/ CEN17 ratio = 0.87) | positive | 30 |
| 10 | HR+, Ki-67 high | positive | 95 | 12 | positive | 100 | 12 | negative | 1 | n.p. | positive | 30 |
| Performance Index | ER | PgR | HER2 | Ki-67 |
|---|---|---|---|---|
| Sensitivity | 1 (1–1) | 0.94 (0.87–1) | 1 (1–1) | 0.86 (0.76–0.95) |
| Specificity | 1 (1–1) | 1 (1–1) | 1 (1–1) | 1 (1–1) |
| Precision (PPV) | 1 (1–1) | 1 (1–1) | 1 (1–1) | 1 (1–1) |
| Accuracy | 1 (1–1) | 0.96 (0.91–1) | 1 (1–1) | 0.9 (0.83–0.97) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Erber, R.; Hartmann, A.; Fasching, P.A.; Ruebner, M.; Stöhr, R.; Beckmann, M.W.; Zentgraf, M.; Popp, V.; Weidler, J.; Simon, I.; et al. Reproducibility of mRNA-Based Testing of ESR1, PGR, ERBB2, and MKI67 Expression in Invasive Breast Cancer—A Europe-Wide External Quality Assessment. Cancers 2021, 13, 4718. https://doi.org/10.3390/cancers13184718
Erber R, Hartmann A, Fasching PA, Ruebner M, Stöhr R, Beckmann MW, Zentgraf M, Popp V, Weidler J, Simon I, et al. Reproducibility of mRNA-Based Testing of ESR1, PGR, ERBB2, and MKI67 Expression in Invasive Breast Cancer—A Europe-Wide External Quality Assessment. Cancers. 2021; 13(18):4718. https://doi.org/10.3390/cancers13184718
Chicago/Turabian StyleErber, Ramona, Arndt Hartmann, Peter Andreas Fasching, Matthias Ruebner, Robert Stöhr, Matthias Wilhelm Beckmann, Miriam Zentgraf, Verena Popp, Jodi Weidler, Iris Simon, and et al. 2021. "Reproducibility of mRNA-Based Testing of ESR1, PGR, ERBB2, and MKI67 Expression in Invasive Breast Cancer—A Europe-Wide External Quality Assessment" Cancers 13, no. 18: 4718. https://doi.org/10.3390/cancers13184718
APA StyleErber, R., Hartmann, A., Fasching, P. A., Ruebner, M., Stöhr, R., Beckmann, M. W., Zentgraf, M., Popp, V., Weidler, J., Simon, I., Becker, S., Huebner, H., Fischer, J., Guerini Rocco, E., Viale, G., Cayre, A., Penault-Llorca, F., Caniego Casas, T., Pérez-Miés, B., ... Ferrazzi, F. (2021). Reproducibility of mRNA-Based Testing of ESR1, PGR, ERBB2, and MKI67 Expression in Invasive Breast Cancer—A Europe-Wide External Quality Assessment. Cancers, 13(18), 4718. https://doi.org/10.3390/cancers13184718









