Liquid Biopsy from Bile-Circulating Tumor DNA in Patients with Biliary Tract Cancer
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Bile Collection
2.3. Chemicals and Reagents
2.4. Preparation of Blood Collection
2.5. DNA Extraction from Bile, Plasma, and FFPE Samples
2.6. Droplet Digital PCR
2.7. Direct Extraction of Single-Stranded RNA
2.8. SMARTer smRNA Library
2.9. mRNA-Seq Data Analysis
2.10. Statistical Analysis
3. Results
3.1. Patient Characteristics
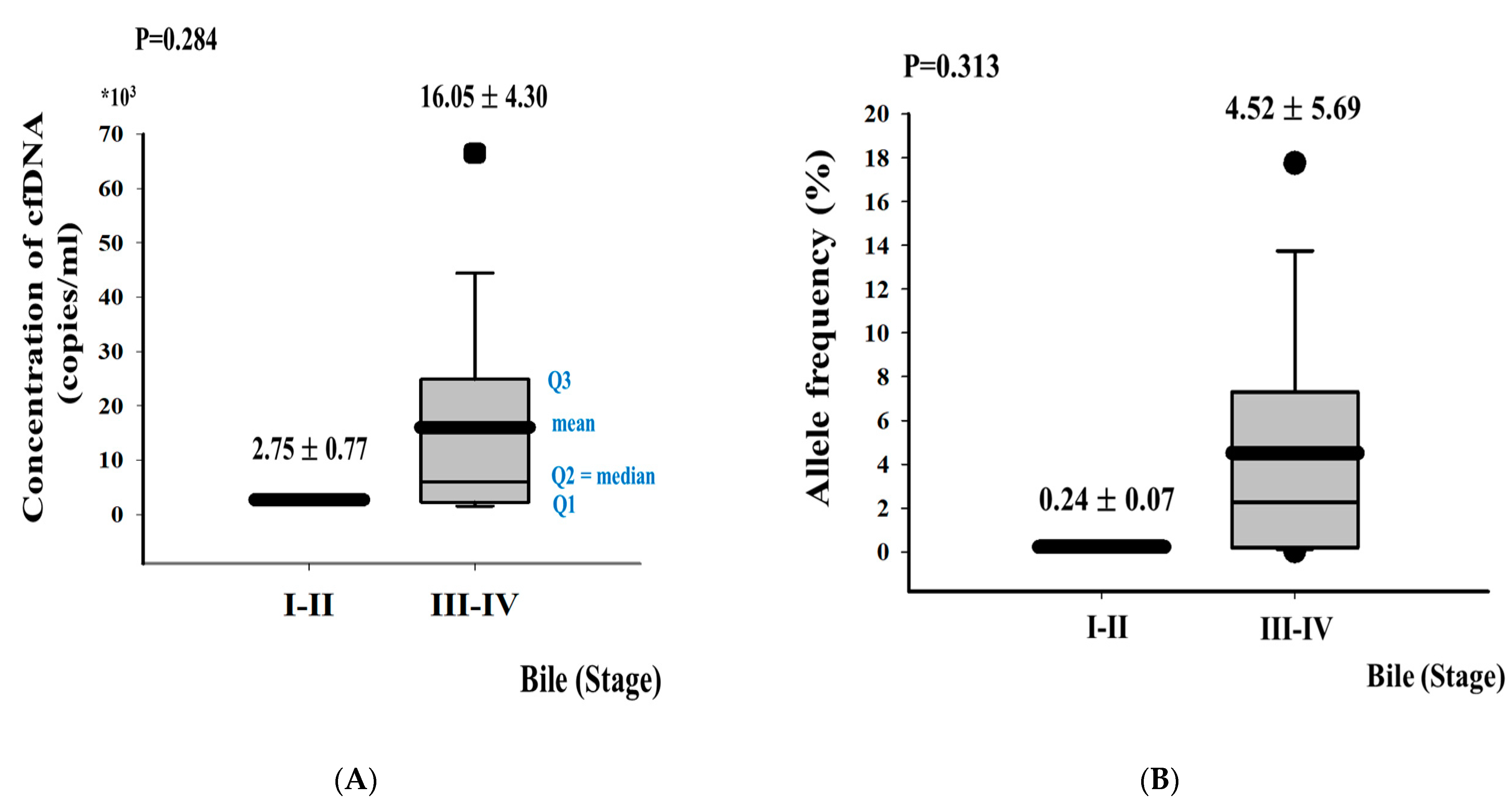
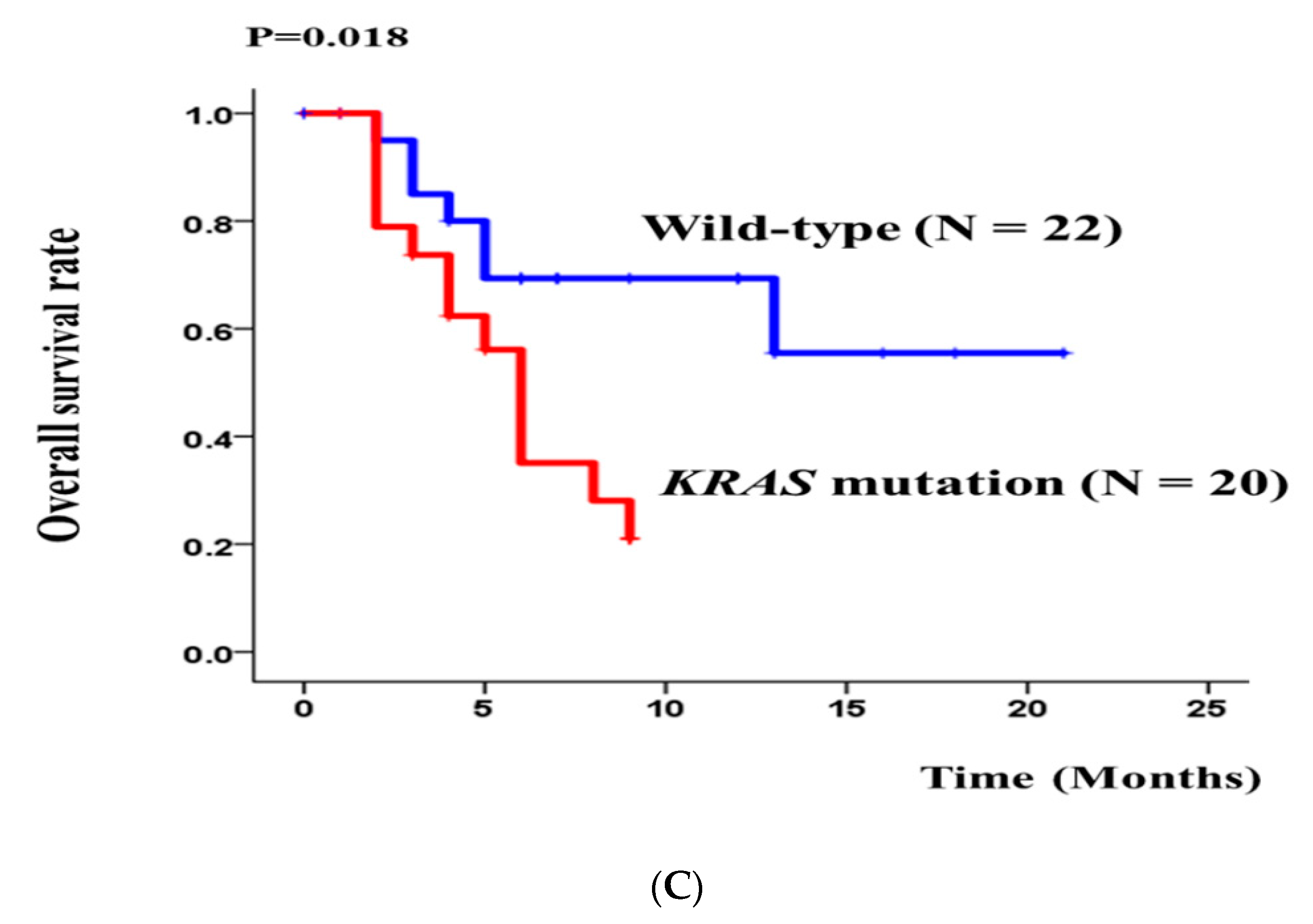
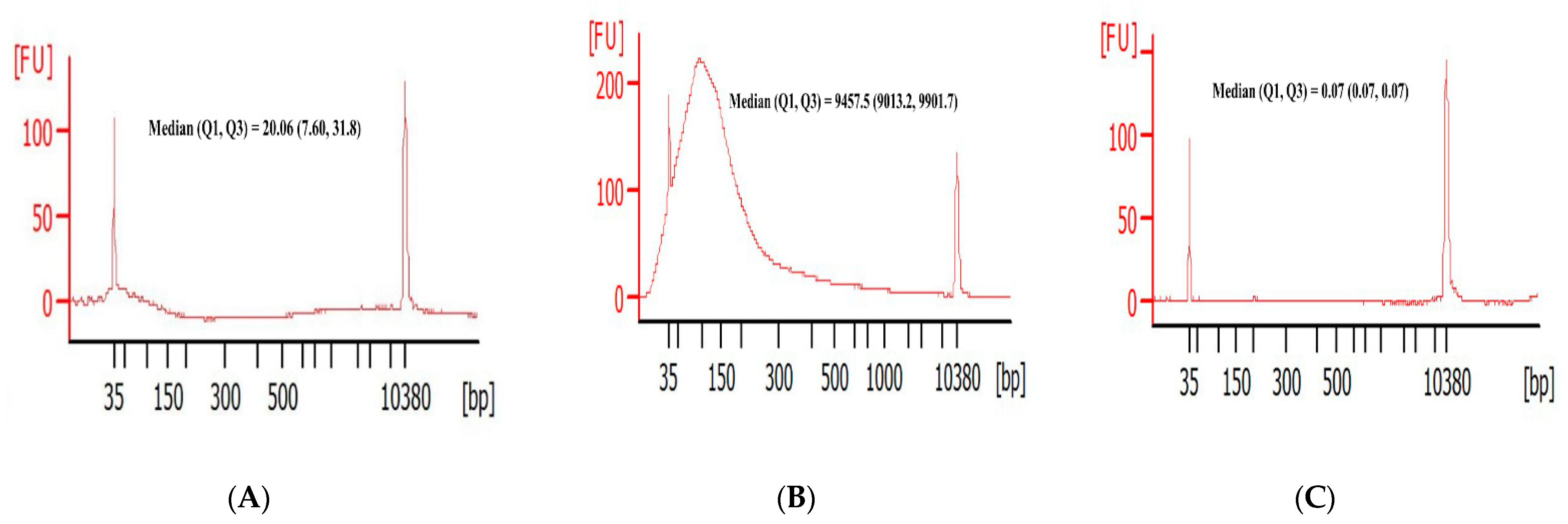
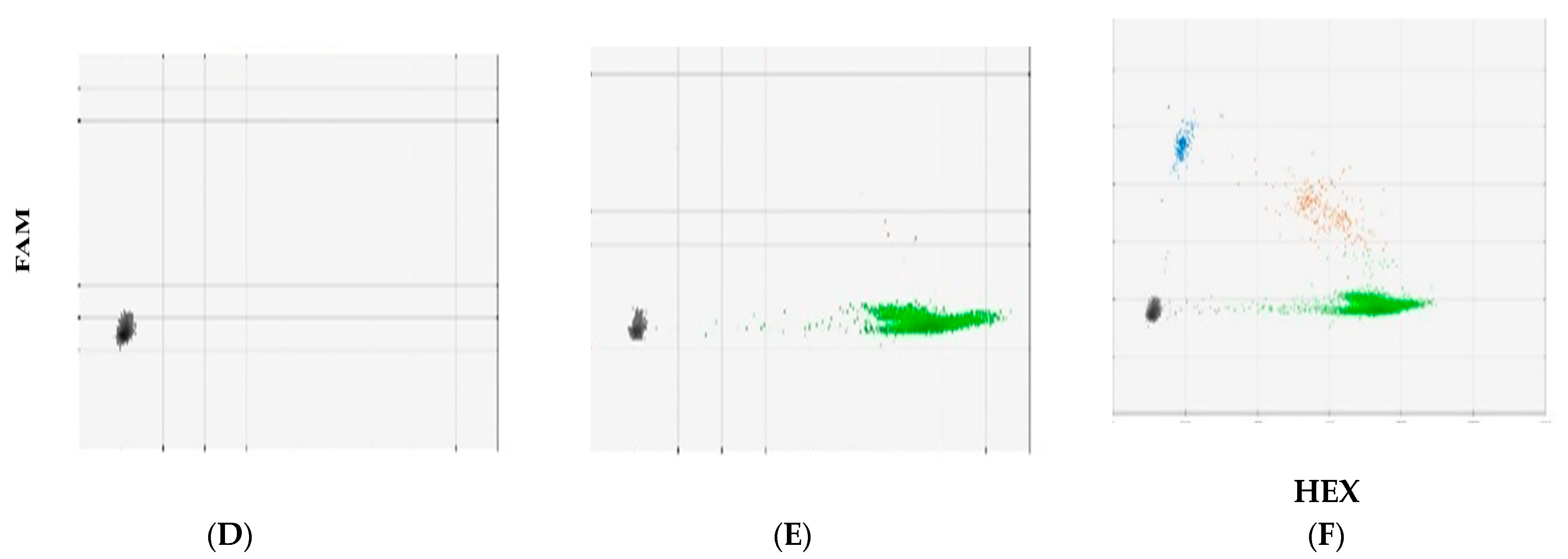
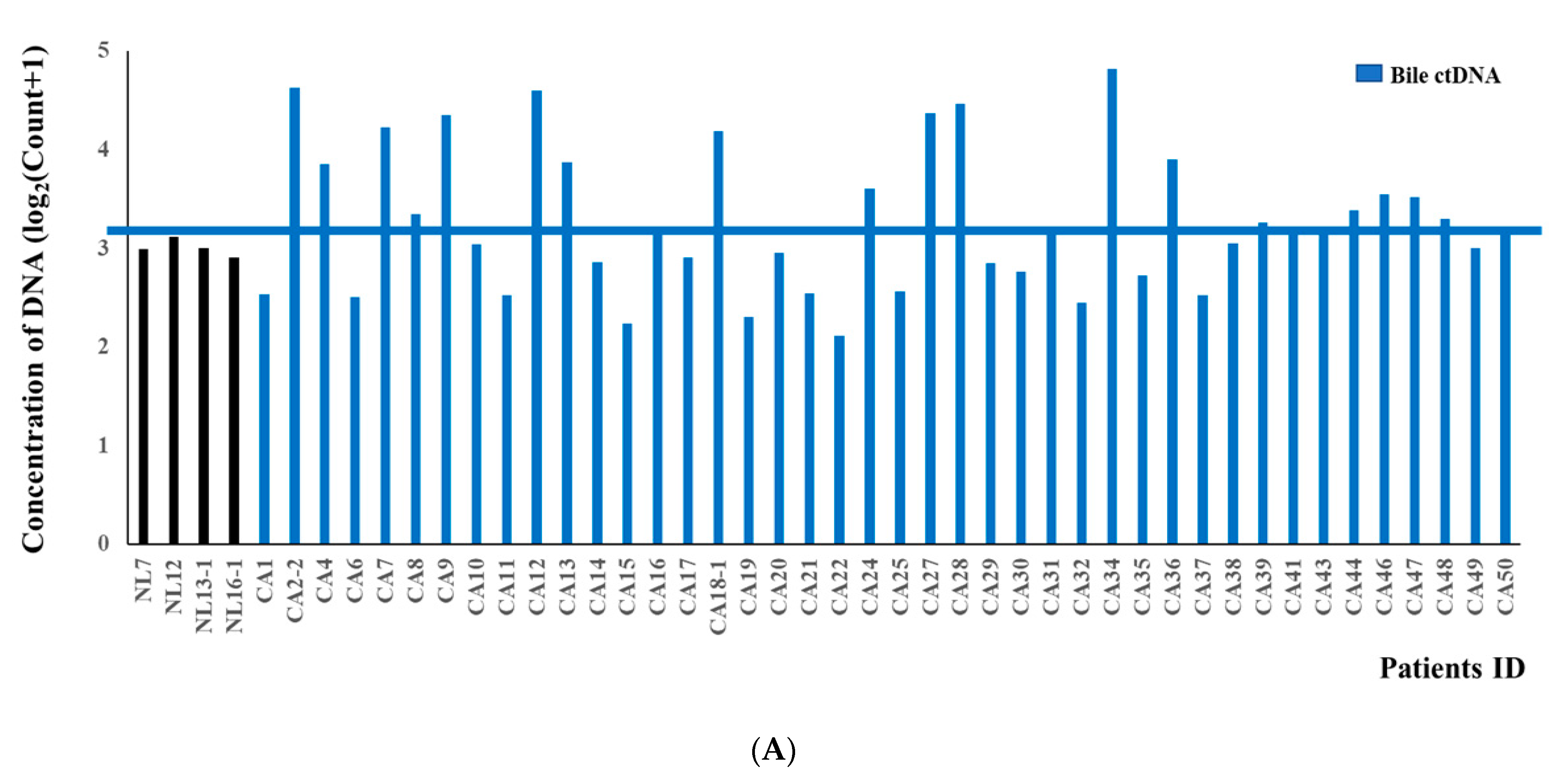
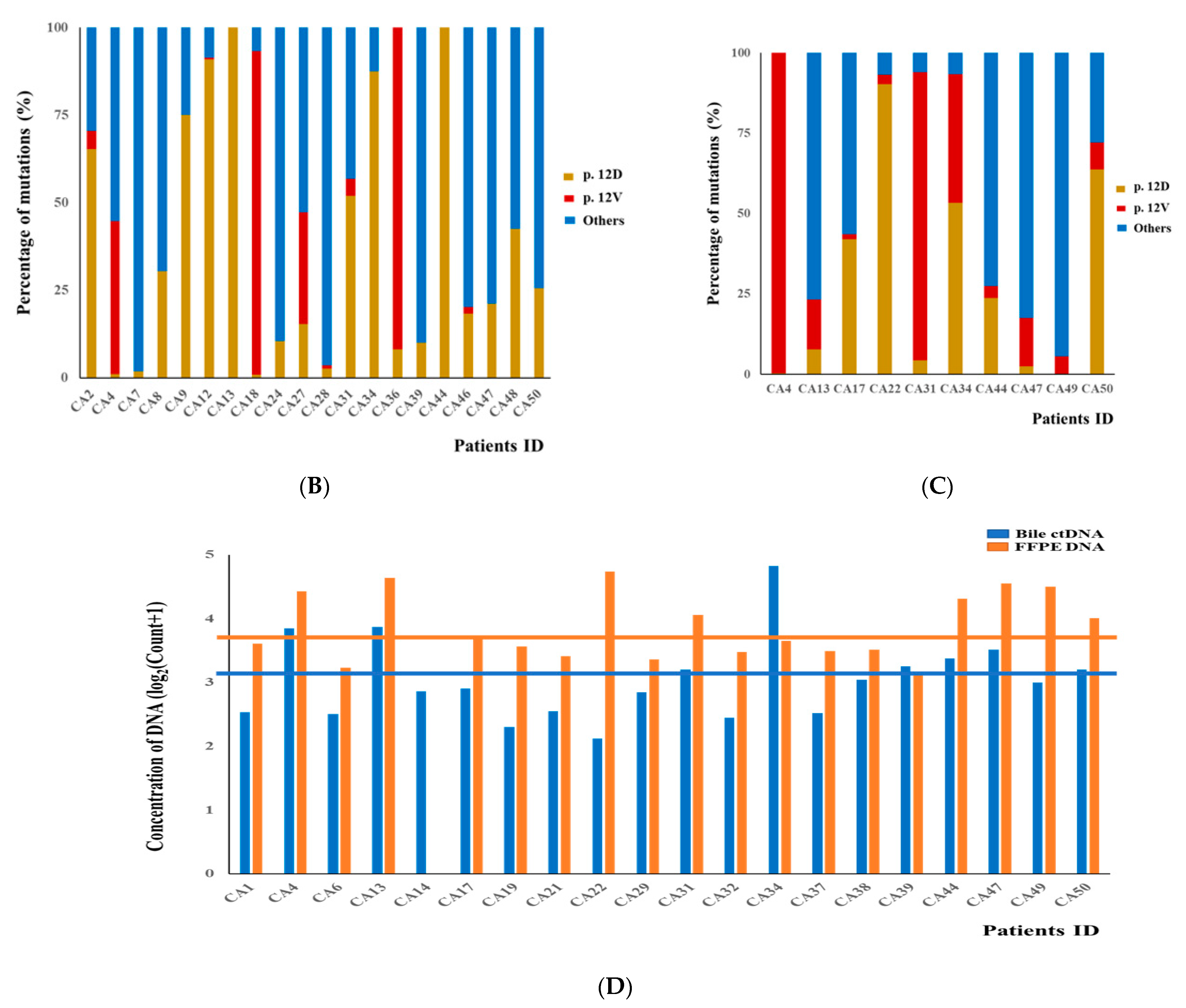
3.2. DNA Fragment Size Distributions and Histograms of KRAS Mutations in Bile ctDNA, FFPE, and Plasma Samples
3.3. Concordance of KRAS Mutations among ctDNA in Bile, Plasma, and FFPE DNA Samples from BTC Patients
3.4. Transcriptomic Sequencing in Bile, and Paired Plasma and Tissue
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Javle, M.; Bekaii-Saab, T.; Jain, A.; Wang, Y.; Kelley, R.K.; Wang, K.; Kang, H.C.; Catenacci, D.; Ali, S.; Krishnan, S.; et al. Biliary cancer: Utility of next-generation sequencing for clinical management. Cancer 2016, 122, 3838–3847. [Google Scholar] [CrossRef]
- de Groen, P.C.; Gores, G.J.; LaRusso, N.F.; Gunderson, L.L.; Nagorney, D.M. Biliary tract cancers. N. Engl. J. Med. 1999, 341, 1368–1378. [Google Scholar] [CrossRef]
- Valle, J.W. Advances in the treatment of metastatic or unresectable biliary tract cancer. Ann. Oncol. 2010, 21, vii345–vii348. [Google Scholar] [CrossRef]
- Malats, N.; Porta, M.; Pinol, J.L.; Corominas, J.M.; Rifa, J.; Real, F.X. Ki-ras mutations as a prognostic factor in extrahepatic bile system cancer. PANK-ras I Project Investigators. J. Clin. Oncol 1995, 13, 1679–1686. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.; Kim, R. Adjuvant therapy for resected extrahepatic cholangiocarcinoma: A review of the literature and future directions. Cancer Treat. Rev. 2009, 35, 322–327. [Google Scholar] [CrossRef]
- DeOliveira, M.L.; Cunningham, S.C.; Cameron, J.L.; Kamangar, F.; Winter, J.M.; Lillemoe, K.D.; Choti, M.A.; Yeo, C.J.; Schulick, R.D. Cholangiocarcinoma: Thirty-one-year experience with 564 patients at a single institution. Ann. Surg. 2007, 245, 755–762. [Google Scholar] [CrossRef] [PubMed]
- Diaz, L.A., Jr.; Bardelli, A. Liquid biopsies: Genotyping circulating tumor DNA. J. Clin. Oncol. 2014, 32, 579–586. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Bi, J.; Bao, L. Genetic profiling of cancer with circulating tumor DNA analysis. J. Genet. Genom. 2018, 45, 79–85. [Google Scholar] [CrossRef]
- Mouliere, F.; El Messaoudi, S.; Pang, D.; Dritschilo, A.; Thierry, A.R. Multi-marker analysis of circulating cell-free DNA toward personalized medicine for colorectal cancer. Mol. Oncol. 2014, 8, 927–941. [Google Scholar] [CrossRef]
- Mouliere, F.; Robert, B.; Arnau Peyrotte, E.; Del Rio, M.; Ychou, M.; Molina, F.; Gongora, C.; Thierry, A.R. High fragmentation characterizes tumour-derived circulating DNA. PLoS ONE 2011, 6, e23418. [Google Scholar] [CrossRef]
- Said, R.; Guibert, N.; Oxnard, G.R.; Tsimberidou, A.M. Circulating tumor DNA analysis in the era of precision oncology. Oncotarget 2020, 11, 188–211. [Google Scholar] [CrossRef] [PubMed]
- Davidson, B.R.; Gurusamy, K. Is preoperative histological diagnosis necessary for cholangiocarcinoma? HPB 2008, 10, 94–97. [Google Scholar] [CrossRef][Green Version]
- Molina-Vila, M.A.; de-Las-Casas, C.M.; Bertran-Alamillo, J.; Jordana-Ariza, N.; Gonzalez-Cao, M.; Rosell, R. cfDNA analysis from blood in melanoma. Ann. Transl. Med. 2015, 3, 309. [Google Scholar] [PubMed]
- Hadano, N.; Murakami, Y.; Uemura, K.; Hashimoto, Y.; Kondo, N.; Nakagawa, N.; Sueda, T.; Hiyama, E. Prognostic value of circulating tumour DNA in patients undergoing curative resection for pancreatic cancer. Br. J. Cancer 2016, 115, 59–65. [Google Scholar] [CrossRef]
- Han, J.Y.; Ahn, K.S.; Baek, W.K.; Suh, S.I.; Kim, Y.H.; Kim, T.S.; Kang, K.J. Usefulness of bile as a biomarker via ferroptosis and cysteine prenylation in cholangiocarcinoma; role of diagnosis and differentiation from benign biliary disease. Surg. Oncol. 2020, 34, 174–181. [Google Scholar] [CrossRef] [PubMed]
- Kinugasa, H.; Nouso, K.; Ako, S.; Dohi, C.; Matsushita, H.; Matsumoto, K.; Kato, H.; Okada, H. Liquid biopsy of bile for the molecular diagnosis of gallbladder cancer. Cancer Biol. Ther. 2018, 19, 934–938. [Google Scholar] [CrossRef]
- Valle, J.W.; Lamarca, A.; Goyal, L.; Barriuso, J.; Zhu, A.X. New Horizons for Precision Medicine in Biliary Tract Cancers. Cancer Discov. 2017, 7, 943–962. [Google Scholar] [CrossRef]
- Matsubara, T.; Sakurai, Y.; Sasayama, Y.; Hori, H.; Ochiai, M.; Funabiki, T.; Matsumoto, K.; Hirono, I. K-ras point mutations in cancerous and noncancerous biliary epithelium in patients with pancreaticobiliary maljunction. Cancer 1996, 77, 1752–1757. [Google Scholar] [CrossRef]
- Churi, C.R.; Shroff, R.; Wang, Y.; Rashid, A.; Kang, H.C.; Weatherly, J.; Zuo, M.; Zinner, R.; Hong, D.; Meric-Bernstam, F.; et al. Mutation profiling in cholangiocarcinoma: Prognostic and therapeutic implications. PLoS ONE 2014, 9, e115383. [Google Scholar]
- Chen, T.C.; Jan, Y.Y.; Yeh, T.S. K-ras mutation is strongly associated with perineural invasion and represents an independent prognostic factor of intrahepatic cholangiocarcinoma after hepatectomy. Ann. Surg. Oncol. 2012, 19, S675–S681. [Google Scholar] [CrossRef]
- Stolze, B.; Reinhart, S.; Bulllinger, L.; Frohling, S.; Scholl, C. Comparative analysis of KRAS codon 12, 13, 18, 61, and 117 mutations using human MCF10A isogenic cell lines. Sci. Rep. 2015, 5, 8535. [Google Scholar] [CrossRef]
- Misale, S.; Di Nicolantonio, F.; Sartore-Bianchi, A.; Siena, S.; Bardelli, A. Resistance to anti-EGFR therapy in colorectal cancer: From heterogeneity to convergent evolution. Cancer Discov. 2014, 4, 1269–1280. [Google Scholar] [CrossRef] [PubMed]
- Mao, C.; Qiu, L.X.; Liao, R.Y.; Du, F.B.; Ding, H.; Yang, W.C.; Li, J.; Chen, Q. KRAS mutations and resistance to EGFR-TKIs treatment in patients with non-small cell lung cancer: A meta-analysis of 22 studies. Lung Cancer 2010, 69, 272–278. [Google Scholar] [CrossRef] [PubMed]
- Shao, F.; Qi, W.; Meng, F.T.; Qiu, L.; Huang, Q. Role of palliative radiotherapy in unresectable intrahepatic cholangiocarcinoma: Population-based analysis with propensity score matching. Cancer Manag. Res. 2018, 10, 1497–1506. [Google Scholar] [CrossRef] [PubMed]
- Zabron, A.A.; Horneffer-van der Sluis, V.M.; Wadsworth, C.A.; Laird, F.; Gierula, M.; Thillainayagam, A.V.; Vlavianos, P.; Westaby, D.; Taylor-Robinson, S.D.; Edwards, R.J.; et al. Elevated levels of neutrophil gelatinase-associated lipocalin in bile from patients with malignant pancreatobiliary disease. Am. J. Gastroenterol. 2011, 106, 1711–1717. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Pertea, M.; Kim, D.; Pertea, G.M.; Leek, J.T.; Salzberg, S.L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 2016, 11, 1650–1667. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Gonda, T.A.; Glick, M.P.; Sethi, A.; Poneros, J.M.; Palmas, W.; Iqbal, S.; Gonzalez, S.; Nandula, S.V.; Emond, J.C.; Brown, R.S.; et al. Polysomy and p16 deletion by fluorescence in situ hybridization in the diagnosis of indeterminate biliary strictures. Gastrointest. Endosc. 2012, 75, 74–79. [Google Scholar] [CrossRef]
- Khan, S.A.; Davidson, B.R.; Goldin, R.D.; Heaton, N.; Karani, J.; Pereira, S.P.; Rosenberg, W.M.; Tait, P.; Taylor-Robinson, S.D.; Thillainayagam, A.V.; et al. Guidelines for the diagnosis and treatment of cholangiocarcinoma: An update. Gut 2012, 61, 1657–1669. [Google Scholar] [CrossRef]
- Miyazaki, M.; Yoshitomi, H.; Miyakawa, S.; Uesaka, K.; Unno, M.; Endo, I.; Ota, T.; Ohtsuka, M.; Kinoshita, H.; Shimada, K.; et al. Clinical practice guidelines for the management of biliary tract cancers 2015: The 2nd English edition. J. Hepato-Biliary-Pancreat. Sci. 2015, 22, 249–273. [Google Scholar] [CrossRef] [PubMed]
- Crowley, E.; Di Nicolantonio, F.; Loupakis, F.; Bardelli, A. Liquid biopsy: Monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 2013, 10, 472–484. [Google Scholar] [CrossRef] [PubMed]
- Shen, N.N.; Zhang, C.; Li, Z.; Kong, L.C.; Wang, X.H.; Gu, Z.C.; Wang, J.L. MicroRNA expression signatures of atrial fibrillation: The critical systematic review and bioinformatics analysis. Exp. Biol Med. 2020, 245, 42–53. [Google Scholar] [CrossRef]
- Driescher, C.; Fuchs, K.; Haeberle, L.; Wolfgang, G.; Lisa, F.; Opitz, F.V.; Haeussinger, D.; Knoefel, W.T.; Keitel, V.; Esposito, I. Bile-Based Cell-Free DNA Analysis Is a Reliable Diagnostic Tool in Pancreatobiliary Cancer. Cancers 2021, 13, 39. [Google Scholar] [CrossRef]
- Yang, N.; Li, Y.; Liu, Z.; Qin, H.; Du, D.; Cao, X.; Cao, X.; Li, J.; Li, D.; Jiang, B.; et al. The characteristics of ctDNA reveal the high complexity in matching the corresponding tumor tissues. BMC Cancer 2018, 18, 319. [Google Scholar] [CrossRef]
- Okada, R.; Ito, T.; Nomura, F.; Kirimura, S.; Cho, Y.; Sekine, M.; Tateishi, Y.; Ariizumi, Y.; Asakage, T. The quantitative analysis of the human papillomavirus DNA load in submandibular gland lesions with droplet digital polymerase chain reaction. Acta Otolaryngol. 2019, 139, 105–110. [Google Scholar] [CrossRef]
- Shen, N.; Zhang, D.; Yin, L.; Qiu, Y.; Liu, J.; Yu, W.; Fu, X.; Zhu, B.; Xu, X.; Duan, A.; et al. Bile cellfree DNA as a novel and powerful liquid biopsy for detecting somatic variants in biliary tract cancer. Oncol. Rep. 2019, 42, 549–560. [Google Scholar]
- Pishvaian, M.J.; Joseph Bender, R.; Matrisian, L.M.; Rahib, L.; Hendifar, A.; Hoos, W.A.; Mikhail, S.; Chung, V.; Picozzi, V.; Heartwell, C.; et al. A pilot study evaluating concordance between blood-based and patient-matched tumor molecular testing within pancreatic cancer patients participating in the Know Your Tumor (KYT) initiative. Oncotarget 2017, 8, 83446–83456. [Google Scholar] [CrossRef] [PubMed]

| Bile (N = 42) | FFPE (N = 20) | Plasma (N = 16) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Wild Type (N = 22) | KRAS Mutant (N = 20) | p | Wild Type (N = 10) | KRAS Mutant (N = 10) | p | Wild Type (N = 13) | KRAS Mutant (N = 3) | p | |
| Age (Mean, SD) | 74.4 ± 9.6 | 71.9 ± 9.5 | 0.408 | 75.3 ± 8.3 | 65.7 ± 8.8 | 0.022 | 73.1 ± 8.3 | 74.3 ± 1.5 | 0.802 |
| Sex (Male, %) | 10 (45.5) | 13 (65.0) | 0.337 | 6 (60.0) | 7 (70.0) | 0.774 | 7 (53.8) | 2 (66.7) | 1.000 |
| AJCC stage (I, II/III, IV) | 5 (22.7)/ 17 (77.3) | 3 (15.0)/ 17 (85.0) | 0.808 | 3 (30.0)/ 7 (70.0) | 5 (50.0)/ 5 (50.0) | 0.478 | 7 (53.8)/ 6 (46.2) | 0 (0.0)/ 3 (100.0) | 0.238 |
| Type of cancer | 0.544 | 0.589 | 0.809 | ||||||
| eCCA | 19 (86.4) | 17 (85.0) | 9 (90.0)) | 8 (80.0) | 12 (92.3) | 3 (100.0) | |||
| GBC | 3 (13.6) | 2 (10.0) | 1 (10.0) | 1(10.0) | 0 (0.0) | 0 (0.0) | |||
| AC | 0 (0.0) | 1 (5.0) | 0 (0.0) | 1 (10.0) | 1 (7.7) | 0 (0.0) | |||
| Treatment | 0.476 | 1.000 | 0.809 | ||||||
| Surgery | 10 (45.5) | 6 (30.0) | 7 (70.0) | 8 (80.0) | 12 (92.3) | 2 (66.7) | |||
| Palliative care | 12 (54.5) | 14 (70.0) | 3 (30.0) | 2 (20.0) | 1 (7.7) | 1 (33.3) | |||
| Bile (N = 42) | FFPE (N = 20) | Plasma (N = 16) | ||||
|---|---|---|---|---|---|---|
| N (%) | Concentration of DNA (Copies/mL) | N (%) | Concentration of DNA (Copies/mL) | N (%) | Concentration of DNA(Copies/mL) | |
| KRAS multiplex mutation | 20 (47.6) | 14,995.0 ± 17,602.6 | 10 (50.0) | 24,515.0 ± 17,128.9 | 3 (18.7) | 12.5 ± 26.9 |
| KRASpoint mutation | ||||||
| p. G12D | 20 (100.0) | 7917 ± 15,436.7 | 9 (90.0) | 6982.0 ± 14,951.6 | 1 (33.3) | 4.3 ± 17.5 |
| p. G12V | 9 (45.0) | 1775.5 ± 3782.3 | 10 (100.0) | 5647.0 ± 8185.9 | 3 (100.0) | 11.8 ± 25.6 |
| p. G12D + p. G12V | 20 (100.0) | 9692.5 ± 15,287.9 | 10 (100.0) | 12,632.0 ± 15,207.5 | 3 (100.0) | 16.2 ± 37.5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Han, J.-Y.; Ahn, K.S.; Kim, T.-S.; Kim, Y.H.; Cho, K.B.; Shin, D.W.; Baek, W.-K.; Suh, S.-I.; Jang, B.-C.; Kang, K.J. Liquid Biopsy from Bile-Circulating Tumor DNA in Patients with Biliary Tract Cancer. Cancers 2021, 13, 4581. https://doi.org/10.3390/cancers13184581
Han J-Y, Ahn KS, Kim T-S, Kim YH, Cho KB, Shin DW, Baek W-K, Suh S-I, Jang B-C, Kang KJ. Liquid Biopsy from Bile-Circulating Tumor DNA in Patients with Biliary Tract Cancer. Cancers. 2021; 13(18):4581. https://doi.org/10.3390/cancers13184581
Chicago/Turabian StyleHan, Jin-Yi, Keun Soo Ahn, Tae-Seok Kim, Yong Hoon Kim, Kwang Bum Cho, Dong Woo Shin, Won-Ki Baek, Seong-Il Suh, Byeong-Churl Jang, and Koo Jeong Kang. 2021. "Liquid Biopsy from Bile-Circulating Tumor DNA in Patients with Biliary Tract Cancer" Cancers 13, no. 18: 4581. https://doi.org/10.3390/cancers13184581
APA StyleHan, J.-Y., Ahn, K. S., Kim, T.-S., Kim, Y. H., Cho, K. B., Shin, D. W., Baek, W.-K., Suh, S.-I., Jang, B.-C., & Kang, K. J. (2021). Liquid Biopsy from Bile-Circulating Tumor DNA in Patients with Biliary Tract Cancer. Cancers, 13(18), 4581. https://doi.org/10.3390/cancers13184581







