Recent Advances in the Delivery Carriers and Chemical Conjugation Strategies for Nucleic Acid Drugs
Abstract
:Simple Summary
Abstract
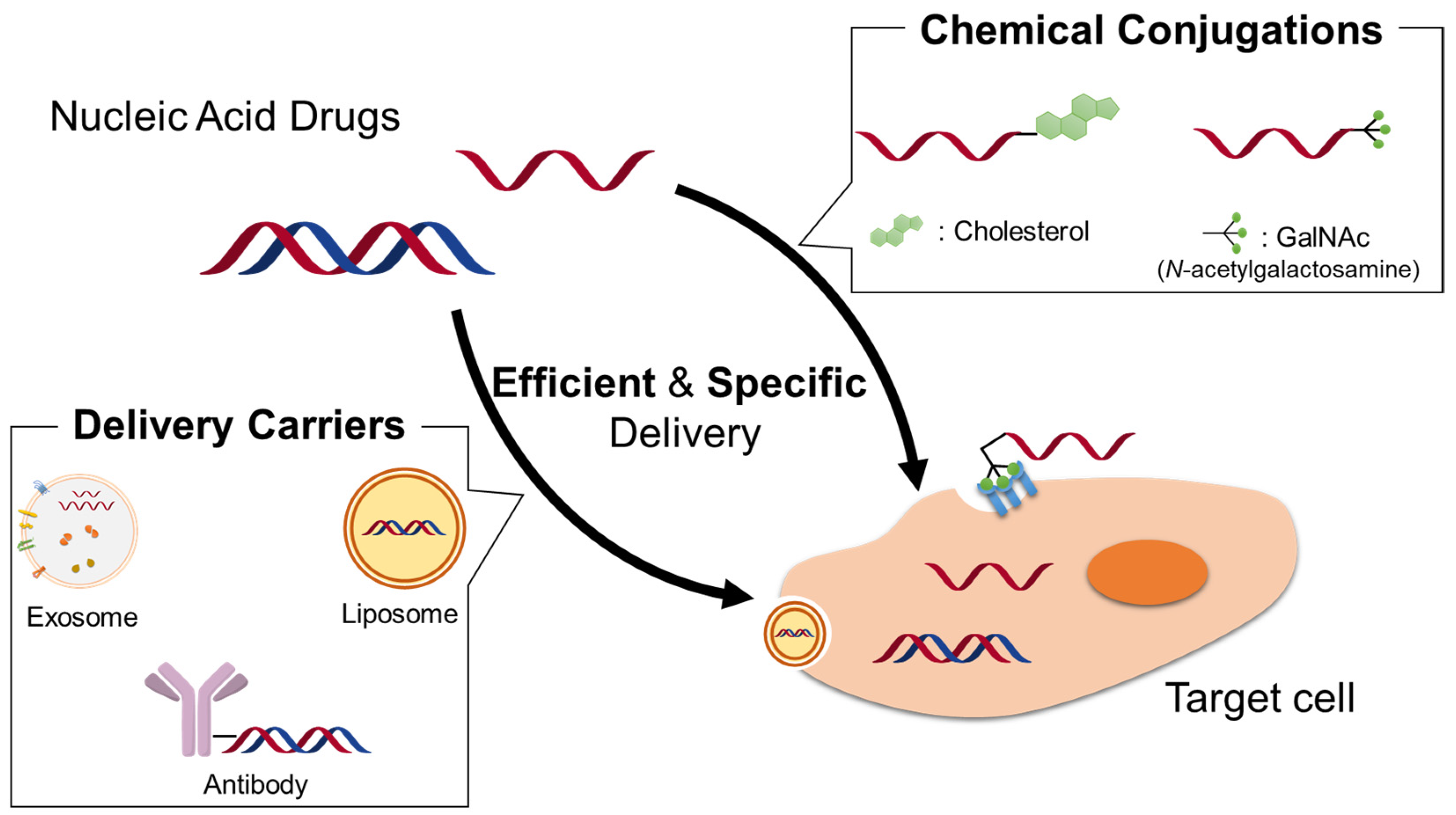
1. Introduction
2. Nonviral Drug Delivery Systems for Nucleic Acid Drugs
2.1. Cationic Vectors for Nucleic Acid Delivery
2.2. Liposomes and Lipid Nanoparticles
3. Conjugation of Functional Molecules to Therapeutic Oligonucleotides
3.1. Cholesterol
3.2. GalNAc
3.3. Folic Acid
3.4. Cell Penetrating Peptides
4. Antibody-Oligonucleotide Conjugates
4.1. Basic Composition and Functions of AOCs
4.2. Therapeutic Applications of AOCs
5. Exosome-Hijacking DDS
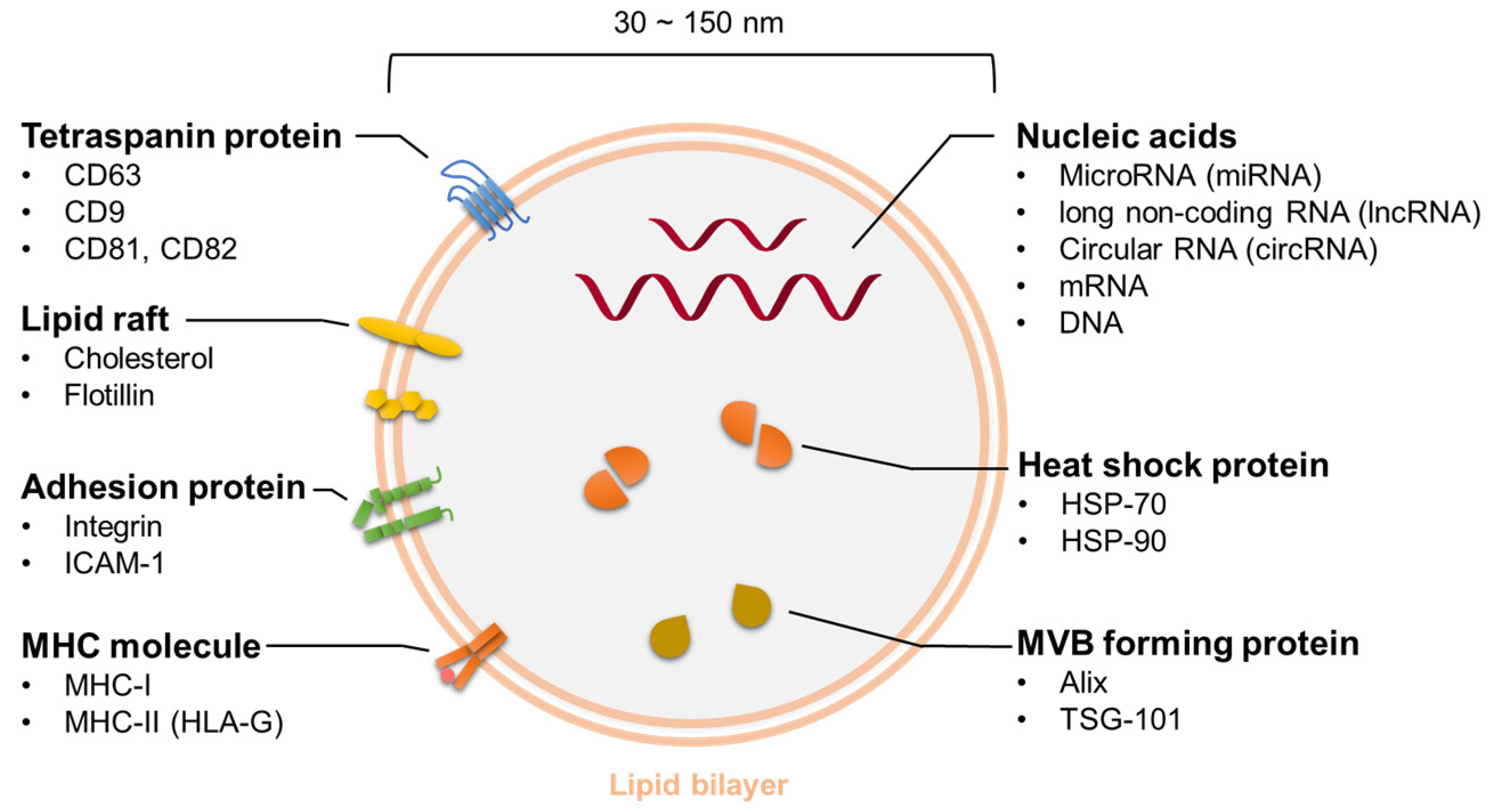
5.1. Properties of Exosomes
5.1.1. Biogenesis and Cellular Uptake
5.1.2. Contents of Exosomes
5.2. Exosomes Used for the Delivery of Nucleic Acid Drugs
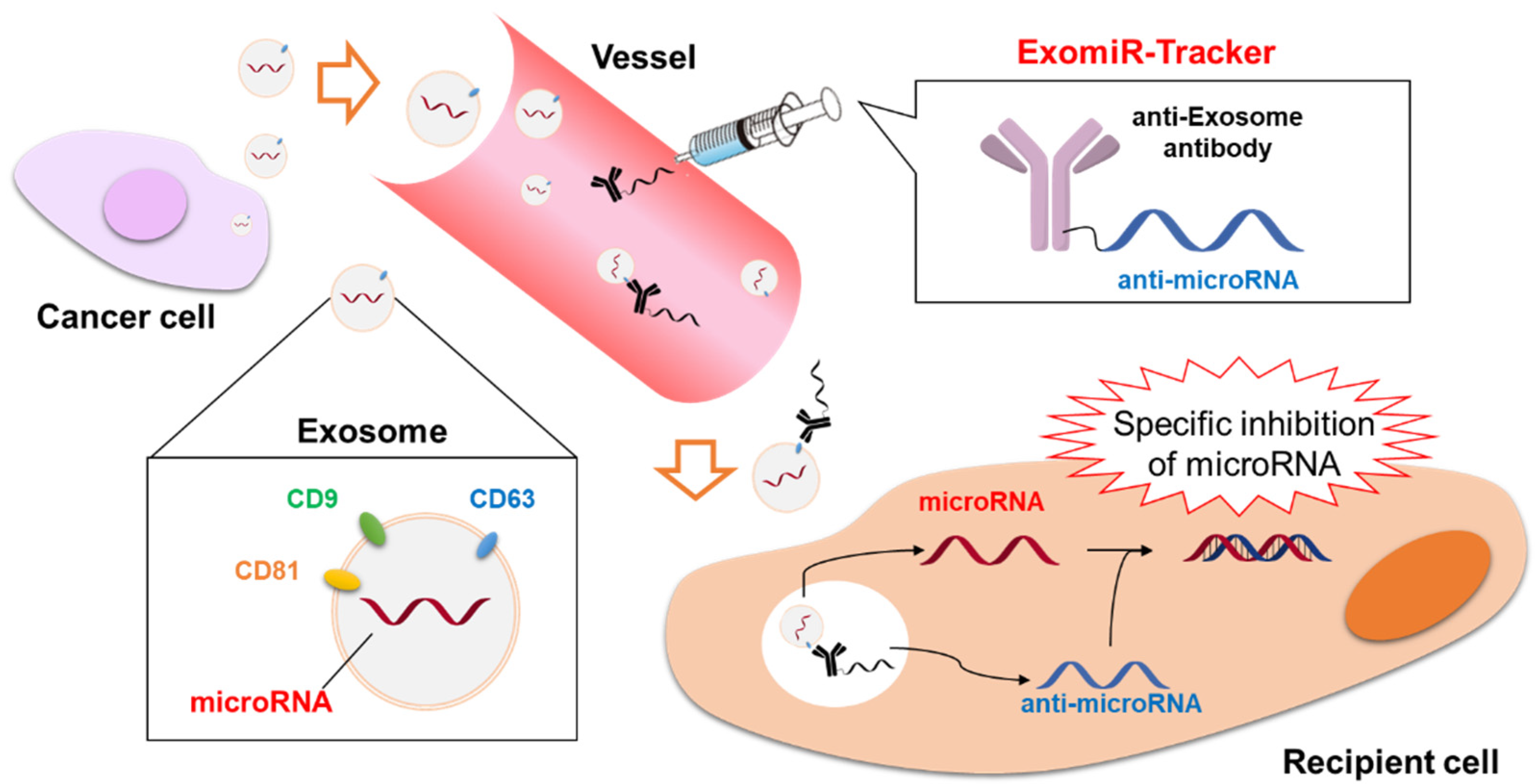
5.3. Antibody-Oligonucleotide Conjugates Targeting microRNAs in Exosomes: ExomiR-Tracker
6. Application to Cancer Treatments Using Surface Molecules on Exosome Membranes
6.1. Phosphatidylserine
6.2. Tetraspanin Proteins
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| Abbreviation | Definition |
| ADCC | Antibody-dependent cell-mediated cytotoxicity |
| ADCs | Antibody-drug conjugates |
| AIDS | Acquired immunodeficiency syndrome |
| ALAS1 | 5-Aminolevulinate synthase 1 |
| AOCs | Antibody-oligonucleotide conjugates |
| ApoB | Apolipoprotein B |
| ASOs | Antisense oligonucleotides |
| ASGPR | Asialoglycoprotein receptor |
| CBSA | Cationic bovine serum albumin |
| CPPs | Cell penetrating peptides |
| DDS | Drug delivery systems |
| DNA | Deoxyribonucleic acid |
| dsAON | Double stranded antisense oligonucleotide |
| DsiRNAs | Dicer substrate siRNAs |
| EDV | EnGeneIC delivery vehicle |
| EGFR | Epidermal growth factor receptor |
| EPM | Exosomes and PEI matrix |
| ESCRT | Endosomal sorting complex required for transport |
| FA | Folic acid |
| FDA | Food and Drug Administration |
| GalNAc | N-Acetylgalactosamine |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase |
| GSCs | Glioblastoma stem cells |
| GSTP | Glutathione-S-transferase P |
| HAO1 | Hydroxyacid Oxidase 1 |
| hATTR | Hereditary transthyretin amyloidosis |
| HIV-1 | Human immunodeficiency virus-1 |
| HLA-G | Human leukocyte antigen-G |
| hsiRNA | Hydrophobically modified siRNA |
| HSPs | Heat shock proteins |
| ICAM-1 | Intercellular adhesion molecule-1 |
| IgG | Immunoglobulin G |
| ILVs | Intraluminal vesicles |
| Lamp | Lysosomal-associated membrane protein |
| LNAs | Locked nucleic acids |
| LNPs | Lipid nanoparticles |
| LODER | Local Drug Eluter |
| miRNA | MicroRNA |
| mRNA | Messenger RNA |
| MSC | Mesenchymal Stem Cell |
| MVBs | Multivesicular bodies |
| PC | Phosphatidylcholine |
| PDAC | Pancreatic ductal adenocarcinoma |
| PEA | Phosphatidylethanolamine |
| PEG | Poly(ethylene glycol) |
| PEI | Polyethyleneimine |
| PK | Pharmacokinetics |
| PKN3 | Protein Kinase N3 |
| PMO | Phosphorodiamidate morpholino oligomer |
| PS | Phosphatidylserine |
| PSLs | PEG-stabilized liposomes |
| RIG-1 | Retinoic acid-inducible gene-1 |
| RNA | Ribonucleic acid |
| RNAi | RNA interference |
| RVG | Rabies virus glycoprotein |
| siRNA | Small interfering RNA |
| SM | Sphingomyelin |
| SNALP | Stable nucleic acid lipid particle |
| STAT3 | Signal transducer and activator of transcription 3 |
| TEM | Tetraspanin-enriched microdomain |
| TNBC | Triple-negative breast cancer |
| TTR | Transthyretin |
| VEGF | Vascular endothelial growth factor |
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 2018, 68, 7–30. [Google Scholar] [CrossRef]
- Lameire, N. Nephrotoxicity of recent anti-cancer agents. Clin. Kidney J. 2014, 7, 11–22. [Google Scholar] [CrossRef] [Green Version]
- Suter, T.M.; Ewer, M.S. Cancer drugs and the heart: Importance and management Eur. Heart J. 2013, 34, 1102–1111. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Levin, A.A. Treating disease at the RNA level with oligonucleotides. N. Engl. J. Med. 2019, 380, 57–70. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Zuroske, T.; Watts, J.K. RNA therapeutics on the rise. Nat. Rev. Drug Discov. 2020, 19, 441–442. [Google Scholar] [CrossRef] [PubMed]
- Craig, K.; Abrams, M.; Amiji, M. Recent preclinical and clinical advances in oligonucleotide conjugates. Expert Opin. Drug Deliv. 2018, 15, 629–640. [Google Scholar] [CrossRef]
- Quemener, A.M.; Bachelot, L.; Forestier, A.; Donnou-Fournet, E.; Gilot, D.; Galibert, M.D. The powerful world of antisense oligonucleotides: From bench to bedside. Wiley Interdiscip. Rev. RNA 2020, 11, e1594. [Google Scholar] [CrossRef] [PubMed]
- Mukalel, A.J.; Riley, R.S.; Zhang, R.; Mitchell, M.J. Nanoparticles for nucleic acid delivery: Applications in cancer immunotherapy. Cancer Lett. 2019, 458, 102–112. [Google Scholar] [CrossRef] [PubMed]
- Fumoto, S.; Yamamoto, T.; Okami, K.; Maemura, Y.; Terada, C.; Yamayoshi, A.; Nishida, K. Understanding In Vivo Fate of Nucleic Acid and Gene Medicines for the Rational Design of Drugs. Pharmaceutics 2021, 13, 159. [Google Scholar] [CrossRef]
- Judge, A.D.; Robbins, M.; Tavakoli, I.; Levi, J.; Hu, L.; Fronda, A.; Ambegia, E.; McClintock, K.; MacLachlan, I. Confirming the RNAi-mediated mechanism of action of siRNA-based cancer therapeutics in mice. J. Clin. Investig. 2009, 119, 661–673. [Google Scholar] [CrossRef] [Green Version]
- Nair, J.K.; Willoughby, J.L.; Chan, A.; Charisse, K.; Alam, M.R.; Wang, Q.; Hoekstra, M.; Kandasamy, P.; Kel’in, A.V.; Milstein, S.; et al. Multivalent N-acetylgalactosamine-conjugated siRNA localizes in hepatocytes and elicits robust RNAi-mediated gene silencing. J. Am. Chem. Soc. 2014, 136, 16958–16961. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khvorova, A.; Watts, J.K. The chemical evolution of oligonucleotide therapies of clinical utility. Nat. Biotechnol. 2017, 35, 238–248. [Google Scholar] [CrossRef]
- Garrelfs, S.F.; Frishberg, Y.; Hulton, S.A.; Koren, M.J.; O’Riordan, W.D.; Cochat, P.; Deschênes, G.; Shasha-Lavsky, H.; Saland, J.M.; Van’t Hoff, W.G.; et al. Lumasiran, an RNAi Therapeutic for Primary Hyperoxaluria Type 1. N. Engl. J. Med. 2021, 384, 1216–1226. [Google Scholar] [CrossRef]
- Santel, A.; Aleku, M.; Keil, O.; Endruschat, J.; Esche, V.; Durieux, B.; Löffler, K.; Fechtner, M.; Röhl, T.; Fisch, G.; et al. RNA interference in the mouse vascular endothelium by systemic administration of siRNA-lipoplexes for cancer therapy. Gene Ther. 2006, 13, 1360–1370. [Google Scholar] [CrossRef] [PubMed]
- Schultheis, B.; Strumberg, D.; Kuhlmann, J.; Wolf, M.; Link, K.; Seufferlein, T.; Kaufmann, J.; Feist, M.; Gebhardt, F.; Khan, M.; et al. Safety, Efficacy and Pharcacokinetics of Targeted Therapy with The Liposomal RNA Interference Therapeutic Atu027 Combined with Gemcitabine in Patients with Pancreatic Adenocarcinoma. A Randomized Phase Ib/IIa Study. Cancers 2020, 12, 3130. [Google Scholar] [CrossRef]
- Atu027 Plus Gemcitabine in Advanced or Metastatic Pancreatic Cancer (Atu027-I-02) (Atu027-I-02). ClinicalTrials.gov, Identifier: NCT01808638 ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/show/NCT01808638 (accessed on 7 July 2021).
- Golan, T.; Khvalevsky, E.Z.; Hubert, A.; Gabai, R.M.; Hen, N.; Segal, A.; Domb, A.; Harari, G.; David, E.B.; Raskin, S.; et al. RNAi therapy targeting KRAS in combination with chemotherapy for locally advanced pancreatic cancer patients. Oncotarget 2015, 6, 24560–24570. [Google Scholar] [CrossRef] [Green Version]
- Shemi, A.; Khvalevsky, E.Z.; Gabai, R.M.; Gabai, R.M.; Hen, N.; Segal, A.; Domb, A.; Harari, G.; David, E.B.; Raskin, S.; et al. Multistep, effective drug distribution within solid tumors. Oncotarget 2015, 6, 39564–39577. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- A Phase 2 Study of siG12D LODER in Combination with Chemotherapy in Patients with Locally Advanced Pancreatic Cancer (PROTACT). ClinicalTrials.gov, Identifier: NCT01676259 ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/show/NCT01676259 (accessed on 7 July 2021).
- Merkel, O.M.; Beyerle, A.; Librizzi, D.; Pfestroff, A.; Behr, T.M.; Sproat, B.; Barth, P.J.; Kissel, T. Nonviral siRNA delivery to the lung: Investigation of PEG-PEI polyplexes and their in vivo performance. Mol. Pharm. 2009, 6, 1246–1260. [Google Scholar] [CrossRef]
- Hussain, M.; Shchepinov, M.; Sohail, M.; Benter, I.F.; Hollins, A.J.; Southern, E.M.; Akhtar, S. A novel anionic dendrimer for improved cellular delivery of antisense oligonucleotides. J. Control. Release 2004, 99, 139–155. [Google Scholar] [CrossRef]
- Ko, Y.T.; Bhattacharya, R.; Bickel, U. Liposome encapsulated polyethylenimine/ODN polyplexes for brain targeting. J. Control. Release 2009, 133, 230–237. [Google Scholar] [CrossRef]
- Hangeland, J.J.; Levis, J.T.; Lee, Y.C.; Ts’O, P.O. Cell-type specific and ligand specific enhancement of cellular uptake of oligodeoxynucleoside methylphosphonates covalently linked with a neoglycopeptide, YEE(ah-GalNAc)3. Bioconjug. Chem. 1995, 6, 695–701. [Google Scholar] [CrossRef]
- Prakash, T.P.; Graham, M.J.; Yu, J.; Carty, R.; Low, A.; Chappell, A.; Schmidt, K.; Zhao, C.; Aghajan, M.; Murray, H.F.; et al. Targeted delivery of antisense oligonucleotides to hepatocytes using triantennary N-acetyl galactosamine improves potency 10-fold in mice. Nucleic Acids Res. 2014, 42, 8796–8807. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rajeev, K.G.; Nair, J.K.; Jayaraman, M.; Charisse, K.; Taneja, N.; O’Shea, J.; Willoughby, J.L.; Yucius, K.; Nguyen, T.; Shulga-Morskaya, S.; et al. Hepatocyte-specific delivery of siRNAs conjugated to novel non-nucleosidic trivalent N-acetylgalactosamine elicits robust gene silencing in vivo. Chembiochem 2015, 16, 903–908. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, T.; Sawamura, M.; Wada, F.; Harada-Shiba, M.; Obika, S. Serial incorporation of a monovalent GalNAc phosphoramidite unit into hepatocyte-targeting antisense oligonucleotides. Bioorg. Med. Chem. 2016, 24, 26–32. [Google Scholar] [CrossRef] [PubMed]
- Yamayoshi, A.; Oyama, S.; Kishimoto, Y.; Konishi, R.; Yamamoto, T.; Kobori, A.; Harada, H.; Ashihara, E.; Sugiyama, H.; Murakami, A. Development of Antibody–Oligonucleotide Complexes for Targeting Exosomal MicroRNA. Pharmaceutics 2020, 12, 545. [Google Scholar] [CrossRef]
- Balwani, M.; Sardh, E.; Ventura, P.; Peiró, P.A.; Rees, D.C.; Stölzel, U.; Bissell, M.; Bonkovsky, H.L.; Windyga, J.; Anderson, K.E.; et al. Phase 3 Trial of RNAi Therapeutic Givosiran for Acute Intermittent Porphyria. N. Engl. J. Med. 2020, 382, 2289–2301. [Google Scholar] [CrossRef]
- Ray, K.K.; Wright, R.S.; Kallend, D.; Koenig, W.; Leiter, L.A.; Raal, F.J.; Bisch, J.A.; Richardson, T.; Jaros, M.; Wijngaard, P.L.J.; et al. Two Phase 3 Trials of Inclisiran in Patients with Elevated LDL Cholesterol. N. Engl. J. Med. 2020, 382, 1507–1519. [Google Scholar] [CrossRef]
- Alvarez-Erviti, L.; Seow, Y.; Yin, H.; Betts, C.; Lakhal, S.; Wood, M.J.A. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat. Biotechnol. 2011, 29, 341–345. [Google Scholar] [CrossRef] [PubMed]
- Kooijmans, S.A.A.; Stremersch, S.; Braeckmans, K.; de Smedt, S.C.; Hendrix, A.; Wood, M.J.A.; Schiffelers, R.M.; Raemdonck, K.; Vader, P. Electroporation-induced siRNA precipitation obscures the efficiency of siRNA loading into extracellular vesicles. J. Control. Release 2013, 172, 229–238. [Google Scholar] [CrossRef]
- Didiot, M.C.; Hall, L.M.; Coles, A.H.; Haraszti, R.A.; Godinho, B.M.D.C.; Chase, K.; Sapp, E.; Ly, S.; Alterman, J.F.; Hassler, M.R. Exosome-mediated Delivery of Hydrophobically Modified siRNA for Huntingtin mRNA Silencing. Mol. Ther. 2016, 24, 1836–1847. [Google Scholar] [CrossRef] [Green Version]
- Zhao, L.; Gu, C.; Gan, Y.; Shao, L.; Chen, H.; Zhu, H. Exosome-mediated siRNA delivery to suppress postoperative breast cancer metastasis. J. Control. Release 2020, 318, 1–15. [Google Scholar] [CrossRef]
- Munagala, R.; Aqil, F.; Jeyabalan, J.; Kandimalla, R.; Wallena, M.; Tyagi, N.; Wilcher, S.; Yan, J.; Schultz, D.J.; Spencer, W.; et al. Exosome-mediated delivery of RNA and DNA for gene therapy. Cancer Lett. 2021, 505, 58–72. [Google Scholar] [CrossRef] [PubMed]
- Kamerkar, S.; LeBleu, V.S.; Sugimoto, H.; Yang, S.; Ruivo, C.F.; Melo, S.A.; Lee, J.J.; Kalluri, R. Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature 2017, 546, 498–503. [Google Scholar] [CrossRef]
- iExosomes in Treating Participants with Metastatic Pancreas Cancer With KrasG12D Mutation. ClinicalTrials.gov, Identifier: NCT03608631 ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/show/NCT03608631 (accessed on 7 July 2021).
- Semple, S.C.; Klimuk, S.K.; Harasym, T.O.; Dos Santos, N.; Ansell, S.M.; Wong, K.F.; Maurer, N.; Stark, H.; Cullis, P.R.; Hope, M.J.; et al. Efficient encapsulation of antisense oligonucleotides in lipid vesicles using ionizable aminolipids: Formation of novel small multilamellar vesicle structures. Biochim. Biophys. Acta 2001, 1510, 152–166. [Google Scholar] [CrossRef] [Green Version]
- Morrissey, D.V.; Lockridge, J.A.; Shaw, L.; Blanchard, K.; Jensen, K.; Breen, W.; Hartsough, K.; Machemer, L.; Radka, S.; Jadhav, V.; et al. Potent and persistent in vivo anti-HBV activity of chemically modified siRNAs. Nat. Biotechnol. 2005, 23, 1002–1007. [Google Scholar] [CrossRef]
- Judge, A.D.; Bola, G.; Lee, A.C.; MacLachlan, I. Design of noninflammatory synthetic siRNA mediating potent gene silencing in vivo. Mol. Ther. 2006, 13, 494–505. [Google Scholar] [CrossRef]
- A Study of NBF-006 in Non-Small Cell Lung, Pancreatic, or Colorectal Cancer. ClinicalTrials.gov, Identifier: NCT03819387 ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/show/NCT03819387 (accessed on 7 July 2021).
- Sahin, U.; Oehm, P.; Derhovanessian, E.; Jabulowsky, R.A.; Vormehr, M.; Gold, M.; Maurus, D.; Schwarck-Kokarakis, D.; Kuhn, A.N.; Omokoko, T.; et al. An RNA vaccine drives immunity in checkpoint-inhibitor-treated melanoma. Nature 2020, 585, 107–112. [Google Scholar] [CrossRef]
- Trial With BNT111 and Cemiplimab in Combination or as Single Agents in Patients with Anti-PD1-refractory/Relapsed, Unresectable Stage III or IV Melanoma. ClinicalTrials.gov, Identifier: NCT04526899 ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/show/NCT04526899 (accessed on 7 July 2021).
- Song, E.; Zhu, P.; Lee, S.K.; Chowdhury, D.; Kussman, S.; Dykxhoorn, D.M.; Feng, Y.; Palliser, D.; Weiner, D.B.; Shankar, P.; et al. Antibody mediated in vivo delivery of small interfering RNAs via cell-surface receptors. Nat. Biotechnol. 2005, 23, 709–717. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Kowolik, C.M.; Swiderski, P.M.; Kortylewski, M.; Yu, H.; Horne, D.A.; Jove, R.; Caballero, O.L.; Simpson, A.J.G.; Lee, F.-T.; et al. Humanized Lewis-Y specific antibody based delivery of STAT3 siRNA. ACS Chem. Biol. 2011, 6, 962–970. [Google Scholar] [CrossRef] [Green Version]
- Xia, C.F.; Zhang, Y.; Zhang, Y.; Boado, R.J.; Pardridge, W.M. Intravenous siRNA of brain cancer with receptor targeting and avidin-biotin technology. Pharm. Res. 2007, 24, 2309–2316. [Google Scholar] [CrossRef] [PubMed]
- Arnold, A.E.; Malek-Adamian, E.; Le, P.U.; Meng, A.; Martínez-Montero, S.; Petrecca, K.; Damha, M.J.; Shoichet, M.S. Antibody-Antisense Oligonucleotide Conjugate Downregulates a Key Gene in Glioblastoma Stem Cells. Mol. Ther. Nucleic Acids 2018, 11, 518–527. [Google Scholar] [CrossRef] [Green Version]
- Sugo, T.; Terada, M.; Oikawa, T.; Miyata, K.; Nishimura, S.; Kenjo, E.; Ogasawara-Shimizu, M.; Makita, Y.; Imaichi, S.; Murata, S.; et al. Development of antibody-siRNA conjugate targeted to cardiac and skeletal muscles. J. Control. Release 2016, 237, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Fishman, J.B.; Rubin, J.B.; Handrahan, J.V.; Connor, J.R.; Fine, R.E. Receptor-mediated transcytosis of transferrin across the blood-brain barrier. J. Neurosci. Res. 1987, 18, 299–304. [Google Scholar] [CrossRef]
- Junghans, M.; Kreuter, J.; Zimmer, A. Antisense delivery using protamine-oligonucleotide particles. Nucleic Acids Res. 2000, 28, E45. [Google Scholar] [CrossRef] [PubMed]
- Dinauer, N.; Lochmann, D.; Demirhan, I.; Bouazzaoui, A.; Zimmer, A.; Chandrac, A.; JörgKreuter; Briesena, H. Intracellular tracking of protamine/antisense oligonucleotide nanoparticles and their inhibitory effect on HIV-1 transactivation. J. Control. Release 2004, 96, 497–507. [Google Scholar] [CrossRef] [PubMed]
- Meleshko, A.N.; Petrovskaya, N.A.; Savelyeva, N.; Vashkevich, K.P.; Doronina, S.N.; Sacivko, N.V. Phase I clinical trial of idiotypic DNA vaccine administered as a complex with polyethylenimine to patients with B-cell lymphoma. Hum. Vaccin. Immunother. 2017, 13, 1–6. [Google Scholar] [CrossRef] [Green Version]
- Intratumoral/Intralesional Administration of MK-4621/JetPEI™ with or without Pembrolizumab in Participants with Advanced/Metastatic or Recurrent Solid Tumors (MK-4621–002). ClinicalTrials.gov, Identifier: NCT03739138 ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/show/NCT03739138 (accessed on 7 July 2021).
- Reid, G.; Pel, M.E.; Kirschner, M.B.; Cheng, Y.Y.; Mugridge, N.; Weiss, J.; Williams, M.; Wright, C.; Edelman, J.J.B.; Vallely, M.P.; et al. Restoring expression of miR-16: A novel approach to therapy for malignant pleural mesothelioma. Ann. Oncol. 2013, 24, 3128–3135. [Google Scholar] [CrossRef] [PubMed]
- MesomiR 1: A Phase I Study of TargomiRs as 2nd or 3rd Line Treatment for Patients with Recurrent MPM and NSCLC. ClinicalTrials.gov, Identifier: NCT02369198 ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/show/NCT02369198 (accessed on 7 July 2021).
- Wolfrum, C.; Shi, S.; Jayaprakash, K.N.; Jayaraman, M.; Wang, G.; Pandey, R.K.; Rajeev, K.G.; Nakayama, T.; Charrise, K.; Ndungo, E.M.; et al. Mechanisms and optimization of in vivo delivery of lipophilic siRNAs. Nat. Biotechnol. 2007, 25, 1149–1157. [Google Scholar] [CrossRef]
- Wong, S.C.; Klein, J.J.; Hamilton, H.L.; Chu, Q.; Frey, C.L.; Truvetskoy, V.S.; Hegge, J.; Wakefield, D.; Rozema, D.B.; Lewis, D.L. Co-injection of a targeted, reversibly masked endosomolytic polymer dramatically improves the efficacy of cholesterol-conjugated small interfering RNAs in vivo. Nucleic Acid Ther. 2012, 22, 380–390. [Google Scholar] [CrossRef] [Green Version]
- Orellana, O.; Tenneti, S.; Rangasamy, L.; Lyle, T.; Low, P.S.; Kasinski, A.L. FolamiRs: Ligand-targeted, vehicle-free delivery of microRNAs for the treatment of cancer. Sci. Transl. Med. 2017, 9, eaam9327. [Google Scholar] [CrossRef] [Green Version]
- Amantana, A.; Moulton, H.M.; Cate, M.L.; Reddy, M.T.; Whitehead, T.; Hassinger, J.N.; Youngblood, D.S.; Iversen, P.L. Pharmacokinetics, biodistribution, stability and toxicity of a cell-penetrating peptide-morpholino oligomer conjugate. Bioconjug. Chem. 2007, 18, 1325–1331. [Google Scholar] [CrossRef] [PubMed]
- Boussif, O.; Lezoualc’h, F.; Zanta, M.A.; Mergny, M.D.; Scherman, D.; Demeneix, B.; Behr, J.P. A versatile vector for gene and oligonucleotide transfer into cells in culture and in vivo: Polyethylenimine. Proc. Natl. Acad. Sci. USA 1995, 92, 7297–7301. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taranejoo, S.; Liu, J.; Verma, P.; Hourigan, K. A review of the developments of characteristics of PEI derivatives for gene delivery applications. J. Appl. Polym. Sci. 2015, 132, 42096. [Google Scholar] [CrossRef]
- Akinc, A.; Thomas, M.; Klibanov, A.M.; Langer, R. Exploring polyethylenimine-mediated DNA transfection and the proton sponge hypothesis. J. Gene Med. 2005, 7, 657–663. [Google Scholar] [CrossRef] [PubMed]
- Guiner, C.L.; Stieger, K.; Snyder, R.O.; Rolling, F.; Moullier, P. Immune responses to gene product of inducible promoters. Curr. Gene Ther. 2007, 7, 334–346. [Google Scholar] [CrossRef]
- Juliano, R.L.; Ming, X.; Nakagawa, O. Cellular uptake and intracellular trafficking of antisense and siRNA oligonucleotides. Bioconjug. Chem. 2012, 23, 147–157. [Google Scholar] [CrossRef] [Green Version]
- Low, P.S.; Henne, W.A.; Doornereerd, D.D. Discovery and Development of Folic-Acid-Based Receptor Targeting for Imaging and Therapy of Cancer and Inflammatory Diseases. Acc. Chem. Res. 2008, 41, 120–129. [Google Scholar] [CrossRef]
- Zhou, W.; Yuan, X.; Wilson, A.; Yang, L.; Mokotoff, M.; Pitt, B.; Li, S. Efficient intracellular delivery of oligonucleotides formulated in folate receptor-targeted lipid vesicles. Bioconjug. Chem. 2002, 13, 1220–1225. [Google Scholar] [CrossRef]
- Wang, M.; Hu, H.; Sun, Y.; Qiua, L.; Zhang, J.; Guan, G.; Zhao, X.; Qiao, M.; Cheng, L.; Cheng, L.; et al. A pH-sensitive gene delivery system based on folic acid-PEG-chitosan—PAMAM-plasmid DNA complexes for cancer cell targeting. Biomaterials 2013, 34, 10120–10132. [Google Scholar] [CrossRef]
- Dohmen, C.; Fröhlich, T.; Lächelt, U.; Röhl, I.; Vornlocher, H.-P.; Hadwiger, P.; Wagner, E. Defined Folate-PEG-siRNA Conjugates for Receptor-specific Gene Silencing. Mol. Ther. Nucleic Acids 2012, 1, e7. [Google Scholar] [CrossRef]
- Frankel, A.D.; Pabo, C.O. Cellular uptake of the tat protein from human immunodeficiency virus. Cell 1988, 55, 1189–1193. [Google Scholar] [CrossRef]
- Derossi, D.; Joliot, A.H.; Chassaing, G.; Prochiantz, A. The third helix of the Antennapedia homeodomain translocates through biological membranes. J. Biol. Chem. 1994, 269, 10444–10450. [Google Scholar] [CrossRef]
- Pooga, M.; Hällbrink, M.; Zorko, M.; Langel, U. Cell penetration by transportan. FASEB J. 1998, 12, 67–77. [Google Scholar] [CrossRef] [Green Version]
- Futaki, S.; Suzuki, T.; Ohashi, W.; Yagami, T.; Tanaka, S.; Ueda, K.; Sugiura, Y. Arginine-rich peptides. An abundant source of membrane-permeable peptides having potential as carriers for intracellular protein delivery. J. Biol. Chem. 2001, 276, 5836–5840. [Google Scholar] [CrossRef] [Green Version]
- Boisguérin, P.; Deshayes, S.; Gait, M.J.; O’Donovan, L.; Godfrey, C.; Betts, C.A.; Wood, M.J.; Lebleu, B. Delivery of therapeutic oligonucleotides with cell penetrating peptides. Adv. Drug Deliv. Rev. 2015, 87, 52–67. [Google Scholar] [CrossRef]
- De Taeye, S.W.; Rispens, T.; Vidarsson, G. The Ligands for Human IgG and Their Effector Functions. Antibodies 2019, 8, 30. [Google Scholar] [CrossRef] [Green Version]
- Wang, W.; Erbe, A.K.; Hank, J.A.; Morris, Z.S.; Sondel, P.M. NK Cell-Mediated Antibody-Dependent Cellular Cytotoxicity in Cancer Immunotherapy. Front. Immunol. 2015, 6, 368. [Google Scholar] [CrossRef] [Green Version]
- Idusogie, E.E.; Presta, L.G.; Gazzano-Santoro, H.; Totpal, K.; Wong, P.Y.; Ultsch, M.; Meng, Y.G.; Mulkerrin, M.G. Mapping of the C1q binding site on rituxan, a chimeric antibody with a human IgG1 Fc. J. Immunol. 2000, 164, 4178–4184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dovgan, I.; Koniev, O.; Kolodych, S.; Wagner, A. Antibody-Oligonucleotide Conjugates as Therapeutic, Imaging, and Detection Agents. Bioconjug. Chem. 2019, 30, 2483–2501. [Google Scholar] [CrossRef] [PubMed]
- Alley, S.C.; Okeley, N.M.; Senter, P.D. Antibody-drug conjugates: Targeted drug delivery for cancer. Curr. Opin. Chem. Biol. 2010, 14, 529–537. [Google Scholar] [CrossRef] [PubMed]
- Beck, A.; Goetsch, L.; Dumontet, C.; Corvaïa, N. Strategies and challenges for the next generation of antibody-drug conjugates. Nat. Rev. Drug Discov. 2017, 16, 315–337. [Google Scholar] [CrossRef] [PubMed]
- Hu, Q.-Y.; Berti, F.; Adamo, R. Towards the next generation of biomedicines by site-selective conjugation. Chem. Soc. Rev. 2016, 45, 1691–1719. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsuchikama, K.; An, Z. Antibody-drug conjugates: Recent advances in conjugation and linker chemistries. Protein Cell 2018, 9, 33–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tagawa, H.; Murayama, K.; Sasaki, K.; Konoue, N.; Kishimura, A.; Kanai, M.; Mori, T.; Oisaki, K.; Katayama, Y. Induction of ADCC by a folic acid–mAb conjugate prepared by tryptophan-selective reaction toward folate-receptor-positive cancer cells. RSC Adv. 2020, 10, 16727–16731. [Google Scholar] [CrossRef]
- Kishimoto, S.; Nakashimada, Y.; Yokota, R.; Hatanaka, T.; Adachi, M.; Ito, Y. Site-Specific Chemical Conjugation of Antibodies by Using Affinity Peptide for the Development of Therapeutic Antibody Format. Bioconjug. Chem. 2019, 30, 698–702. [Google Scholar] [CrossRef]
- Théry, C. Exosomes: Secreted vesicles and intercellular communications. F1000 Biol. Rep. 2011, 3, 15. [Google Scholar] [CrossRef]
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ohno, S.; Takanashi, M.; Sudo, K.; Ueda, S.; Ishikawa, A.; Matsuyama, N.; Fujita, K.; Mizutani, T.; Ohgi, T.; Ochiya, T.; et al. Systemically Injected Exosomes Targeted to EGFR Deliver Antitumor MicroRNA to Breast Cancer Cells. Mol. Ther. 2013, 21, 185–191. [Google Scholar] [CrossRef] [Green Version]
- Kalra, H.; Simpson, R.J.; Ji, H.; Aikawa, E.; Altevogt, P.; Askenase, P.; Bond, V.C.; Borràs, F.E.; Breakefield, X.; Budnik, V.; et al. Vesiclepedia: A Compendium for Extracellular Vesicles with Continuous Community Annotation. PLoS Biol. 2012, 10, e1001450. [Google Scholar] [CrossRef] [Green Version]
- Yu, X.; Odenthal, M.; Fries, J.W. Exosomes as miRNA Carriers: Formation–Function–Future. Int. J. Mol. Sci. 2016, 17, 2028. [Google Scholar] [CrossRef]
- Harding, C.; Heuser, J.; Stahl, P. Endocytosis and intracellular processing of transferrin and colloidal gold-transferrin in rat reticulocytes: Demonstration of a pathway for receptor shedding. Eur. J. Cell Biol. 1984, 35, 256–263. [Google Scholar] [PubMed]
- Larios, J.; Mercier, V.; Roux, A.; Gruenberg, J. ALIX- and ESCRT-III-dependent sorting of tetraspanins to exosomes. J. Cell Biol. 2020, 219, e201904113. [Google Scholar] [CrossRef] [Green Version]
- Morita, E.; Sandrin, V.; Chung, H.Y.; Morham, S.G.; Gygi, S.P.; Rodesch, C.K.; Sundquist, W.I. Human ESCRT and ALIX proteins interact with proteins of the midbody and function in cytokinesis. EMBO J. 2007, 26, 4215–4227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Babst, M. MVB vesicle formation: ESCRT-dependent, ESCRT-independent and everything in between. Curr. Opin. Cell Biol. 2011, 23, 452–457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Frankel, E.B.; Audhya, A. ESCRT-dependent cargo sorting at multivesicular endosomes. Semin. Cell Dev. Biol. 2018, 74, 4–10. [Google Scholar] [CrossRef] [PubMed]
- Wollert, T.; Hurley, J.H. Molecular mechanism of multivesicular body biogenesis by ESCRT complexes. Nature 2010, 8, 464, 864–869. [Google Scholar] [CrossRef] [Green Version]
- Kenific, C.M.; Zhang, H.; Lyden, D. An exosome pathway without an ESCRT. Cell Res. 2021, 31, 105–106. [Google Scholar] [CrossRef]
- Wei, D.; Zhan, W.; Gao, Y.; Huang, L.; Gong, R.; Wang, W.; Zhang, R.; Wu, Y.; Gao, S.; Kang, T. RAB31 marks and controls an ESCRT-independent exosome pathway. Cell Res. 2021, 31, 157–177. [Google Scholar] [CrossRef]
- Tian, T.; Zhu, Y.L.; Zhou, Y.Y.; Liang, G.F.; Wang, Y.Y.; Hu, F.H.; Xiao, Z.D. Exosome uptake through clathrin-mediated endocytosis and macropinocytosis and mediating miR-21 delivery. J. Biol. Chem. 2014, 289, 22258–22267. [Google Scholar] [CrossRef] [Green Version]
- Nanbo, A.; Kawanishi, E.; Yoshida, R.; Yoshiyama, H. Exosomes derived from Epstein-Barr virus-infected cells are internalized via caveola-dependent endocytosis and promote phenotypic modulation in target cells. J. Virol. 2013, 87, 10334–10347. [Google Scholar] [CrossRef] [Green Version]
- Costa Verdera, H.; Gitz-Francois, J.J.; Schiffelers, R.M.; Vader, P. Cellular uptake of extracellular vesicles is mediated by clathrin-independent endocytosis and macropinocytosis. J. Control. Release 2017, 266, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Feng, D.; Zhao, W.L.; Ye, Y.Y.; Bai, X.C.; Liu, R.Q.; Chang, L.F.; Zhou, Q.; Sui, S.F. Cellular internalization of exosomes occurs through phagocytosis. Traffic 2010, 11, 675–687. [Google Scholar] [CrossRef] [PubMed]
- Svensson, K.J.; Christianson, H.C.; Wittrup, A.; Bourseau-Guilmain, E.; Lindqvist, E.; Svensson, L.M.; Mörgelin, M.; Belting, M. Exosome uptake depends on ERK1/2-heat shock protein 27 signaling and lipid Raft-mediated endocytosis negatively regulated by caveolin-1. J. Biol. Chem. 2013, 288, 17713–17724. [Google Scholar] [CrossRef] [Green Version]
- Pols, M.S.; Klumperman, J. Trafficking and function of the tetraspanin CD63. Exp. Cell Res. 2009, 315, 1584–1592. [Google Scholar] [CrossRef] [PubMed]
- Malla, R.R.; Pandrangi, S.; Kumari, S.; Gavara, M.M.; Badana, A.K. Exosomal tetraspanins as regulators of cancer progression and metastasis and novel diagnostic markers. Asia Pac. J. Clin. Oncol. 2018, 14, 383–391. [Google Scholar] [CrossRef] [Green Version]
- Andreu, Z.; Yáñez-Mó, M. Tetraspanins in extracellular vesicle formation and function. Front. Immunol. 2014, 5, 442. [Google Scholar] [CrossRef] [Green Version]
- Hemler, M.E. Tetraspanin proteins mediate cellular penetration, invasion, and fusion events and define a novel type of membrane microdomain. Annu. Rev. Cell Dev. Biol. 2003, 19, 397–422. [Google Scholar] [CrossRef] [PubMed]
- Shimaoka, M.; Kawamoto, E.; Gaowa, A.; Okamoto, T.; Park, E.J. Connexins and Integrins in Exosomes. Cancers 2019, 11, 106. [Google Scholar] [CrossRef] [Green Version]
- Rebmann, V.; König, L.; Nardi Fda, S.; Wagner, B.; Manvailer, L.F.; Horn, P.A. The Potential of HLA-G-Bearing Extracellular Vesicles as a Future Element in HLA-G Immune Biology. Front. Immunol. 2016, 7, 173. [Google Scholar] [CrossRef] [Green Version]
- Taha, E.A.; Ono, K.; Eguchi, T. Roles of Extracellular HSPs as Biomarkers in Immune Surveillance and Immune Evasion. Int. J. Mol. Sci. 2019, 20, 4588. [Google Scholar] [CrossRef] [Green Version]
- Reddy, V.S.; Madala, S.K.; Trinath, J.; Reddy, G.B. Extracellular small heat shock proteins: Exosomal biogenesis and function. Cell Stress Chaperones 2018, 23, 441–454. [Google Scholar] [CrossRef] [PubMed]
- Bissig, C.; Gruenberg, J. ALIX and the multivesicular endosome: ALIX in Wonderland. Trends Cell Biol. 2014, 24, 19–25. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Li, S.; Li, L.; Li, M.; Guo, C.; Yao, J.; Mi, S. Exosome and exosomal microRNA: Trafficking, sorting, and function. Genom. Proteom. Bioinform. 2015, 13, 17–24. [Google Scholar] [CrossRef] [Green Version]
- Sun, Z.; Yang, S.; Zhou, Q.; Wang, G.; Song, J.; Li, Z.; Zhang, Z.; Xu, J.; Xia, K.; Chang, Y.; et al. Emerging role of exosome-derived long non-coding RNAs in tumor microenvironment. Mol. Cancer 2018, 17, 82. [Google Scholar] [CrossRef]
- Fanale, D.; Taverna, S.; Russo, A.; Bazan, V. Circular RNA in Exosomes. Adv. Exp. Med. Biol. 2018, 1087, 109–117. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, J.; Ma, J.; Sun, T.; Zhou, Q.; Wang, W.; Wang, G.; Wu, P.; Wang, H.; Jiang, L.; et al. Exosomal circRNAs: Biogenesis, effect and application in human diseases. Mol. Cancer 2019, 18, 116. [Google Scholar] [CrossRef]
- Zhang, H.; Freitas, D.; Kim, H.S.; Fabijanic, K.; Li, Z.; Chen, H.; Mark, M.T.; Molina, H.; Martin, A.B.; Bojmar, L.; et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat. Cell Biol. 2018, 20, 332–343. [Google Scholar] [CrossRef]
- Lim, L.P.; Lau, N.C.; Garrett-Engele, P.; Grimson, A.; Schelter, J.M.; Castle, J.; Bartel, D.P.; Linsley, P.S.; Johnson, J.M. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 2005, 433, 769–773. [Google Scholar] [CrossRef] [PubMed]
- Griffiths-Jones, S.; Grocock, R.J.; van Dongen, S.; Bateman, A.; Enright, A.J. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 2006, 34, D140–D144. [Google Scholar] [CrossRef]
- Friedman, R.C.; Farh, K.K.; Burge, C.B.; Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009, 19, 92–105. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Liu, D.; Chen, X.; Li, J.; Li, L.; Bian, Z.; Sun, F.; Lu, J.; Yin, Y.; Cai, X.; et al. Secreted monocytic miR-150 enhances targeted endothelial cell migration. Mol. Cell 2010, 39, 133–144. [Google Scholar] [CrossRef] [Green Version]
- Peinado, H.; Zhang, H.; Matei, I.R.; Costa-Silva, B.; Hoshino, A.; Rodrigues, G.; Psaila, B.; Kaplan, R.N.; Bromberg, J.F.; Kang, Y.; et al. Pre-metastatic niches: Organ-specific homes for metastases. Nat. Rev. Cancer 2017, 17, 302–317. [Google Scholar] [CrossRef]
- Hoshino, A.; Costa-Silva, B.; Shen, T.L.; Rodrigues, G.; Hashimoto, A.; Mark, M.T.; Molina, H.; Kohsaka, S.; di Giannatale, A.; Ceder, S.; et al. Tumour exosome integrins determine organotropic metastasis. Nature 2015, 527, 329–335. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, L.; Thomson, J.M.; Hemann, M.T.; Hernando-Monge, E.; Mu, D.; Goodson, S.; Powers, S.; Cordon-Cardo, C.; Lowe, S.W.; Hannon, G.J.; et al. A microRNA polycistron as a potential human oncogene. Nature 2005, 435, 828–833. [Google Scholar] [CrossRef] [PubMed]
- Obika, S.; Nanbu, D.; Hari, Y.; Andoh, J.; Morio, K.; Doi, T.; Imanishi, T. Stability and structural features of the duplexes containing nucleoside analogues with a fixed N-type conformation, 2′-O,4′-C-methyleneribonucleosides. Tetrahedron Lett. 1998, 39, 5401–5404. [Google Scholar] [CrossRef]
- Van der Ree, M.H.; van der Meer, A.J.; van Nuenen, A.C.; de Bruijne, J.; Ottosen, S.; Janssen, H.L.; Kootstra, N.A.; Reesink, H.W. Miravirsen dosing in chronic hepatitis C patients results in decreased microRNA-122 levels without affecting other microRNAs in plasma. Aliment. Pharmacol. Ther. 2016, 43, 102–113. [Google Scholar] [CrossRef] [Green Version]
- Piccin, A.; Murphy, W.G.; Smith, O.P. Circulating microparticles: Pathophysiology and clinical implications. Blood Rev. 2007, 21, 157–171. [Google Scholar] [CrossRef]
- Llorente, A.; Skotland, T.; Sylvänne, T.; Kauhanen, D.; Róg, T.; Orłowski, A.; Vattulainen, I.; Ekroos, K.; Sandvig, K. Molecular lipidomics of exosomes released by PC-3 prostate cancer cells. Biochim. Biophys. Acta 2013, 1831, 1302–1309. [Google Scholar] [CrossRef]
- Segawa, K.; Nagata, S. An Apoptotic ‘Eat Me’ Signal: Phosphatidylserine Exposure. Trends Cell Biol. 2015, 25, 639–650. [Google Scholar] [CrossRef]
- Kooijmans, S.A.A.; Gitz-Francois, J.J.M.; Schiffelers, R.M.; Vader, P. Recombinant phosphatidylserine-binding nanobodies for targeting of extracellular vesicles to tumor cells: A plug-and-play approach. Nanoscale 2018, 10, 2413–2426. [Google Scholar] [CrossRef] [Green Version]
- Wong, W. A New Way to Cluster. Sci. Signal. 2008, 1, ec381. [Google Scholar] [CrossRef]
- Huang, C.; Fu, C.; Wren, J.D.; Wang, X.; Zhang, F.; Zhang, Y.H.; Connel, S.A.; Chen, T.; Zhang, X.A. Tetraspanin-enriched microdomains regulate digitation junctions. Cell Mol. Life Sci. 2018, 75, 3423–3439. [Google Scholar] [CrossRef]
- Van Niel, G.; Charrin, S.; Simoes, S.; Romao, M.; Rochin, L.; Saftig, P.; Marks, M.S.; Rubinstein, E.; Raposo, G. The tetraspanin CD63 regulates ESCRT-independent and -dependent endosomal sorting during melanogenesis. Dev. Cell 2011, 21, 708–721. [Google Scholar] [CrossRef] [Green Version]
- Little, K.D.; Hemler, M.E.; Stipp, C.S. Dynamic regulation of a GPCR-tetraspanin-G protein complex on intact cells: Central role of CD81 in facilitating GPR56-Galpha q/11 association. Mol. Biol. Cell 2004, 15, 2375–2387. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, M.S.; Tung, K.S.; Coonrod, S.A.; Takahashi, Y.; Bigler, D.; Chang, A.; Yamashita, Y.; Kincade, P.W.; Herr, J.C.; White, J.M. Role of the integrin-associated protein CD9 in binding between sperm ADAM 2 and the egg integrin alpha6beta1: Implications for murine fertilization. Proc. Natl. Acad. Sci. USA 1999, 96, 11830–11835. [Google Scholar] [CrossRef] [Green Version]
- Powner, D.; Kopp, P.M.; Monkley, S.J.; Critchley, D.R.; Berditchevski, F. Tetraspanin CD9 in cell migration. Biochem. Soc. Trans. 2011, 39, 563–567. [Google Scholar] [CrossRef]
- Zöller, M. Tetraspanins: Push and pull in suppressing and promoting metastasis. Nat. Rev. Cancer 2009, 9, 40–55. [Google Scholar] [CrossRef] [PubMed]
- Lu, W.; Fei, A.; Jiang, Y.; Chen, L.; Wang, Y. Tetraspanin CD9 interacts with α-secretase to enhance its oncogenic function in pancreatic cancer. Am. J. Transl. Res. 2020, 12, 5525–5537. [Google Scholar] [PubMed]
- Reyes, R.; Cardeñes, B.; Machado-Pineda, Y.; Cabañas, C. Tetraspanin CD9: A Key Regulator of Cell Adhesion in the Immune System. Front. Immunol. 2018, 9, 863. [Google Scholar] [CrossRef] [PubMed]
- Murayama, Y.; Oritani, K.; Tsutsui, S. Novel CD9-targeted therapies in gastric cancer. World J. Gastroenterol. 2015, 21, 3206–3213. [Google Scholar] [CrossRef] [PubMed]
- Yoshioka, Y.; Kosaka, N.; Konishi, Y.; Ohta, H.; Okamoto, H.; Sonoda, H.; Nonaka, R.; Yamamoto, H.; Ishii, H.; Mori, M.; et al. Ultra-sensitive liquid biopsy of circulating extracellular vesicles using ExoScreen. Nat. Commun. 2014, 5, 3591. [Google Scholar] [CrossRef] [PubMed]
| Carrier | Molecule | Target | Disease | Approved | Clinical Trial | References | |
|---|---|---|---|---|---|---|---|
| Phase (Approved) | ClinicalTrials.gov Identifier | ||||||
| Protamine Protamine | ASO ASO | c-myc HIV-1 | Histiocytic lymphoma cell HIV-AIDS | - - | - - | - - | [49] [50] |
| PEI PEI | DNA vaccine RGT100 | - RIG-1 | B-cell non-Hodgkin’s lymphoma Advanced metastatic solid tumor | - - | I/ongoing I/completed | ISRCTN31090206 NCT03739138 | [51] [52] |
| Anionic dendrimer | ASO | EGFR | Epidermoid carcinoma | - | - | - | [21] |
| LNP Lipoplex Lipoplex Liposome PSL | siRNA siRNA mRNA vaccine siRNA DNA | TTR PKN3 - GSTP - | hATTR amyloidosis advanced pancreatic cancer (cancer vaccine) Non-small cell lung cancer (brain targeting) | Yes, patisiran - - - - | (2018) II/completed II/recruiting I/recruiting - | - NCT01808638 NCT04526899 NCT03819387 - | [10,38,39] [14,15,16] [41,42] [40] [22] |
| EDV | miR-16 mimic | EGFR | Malignant pleural mesothelioma Non-small cell lung cancer | - | I/completed | NCT02369198 | [53,54] |
| LODER™ | siRNA | K-ras G12D | Pancreatic cancer | - | II/recruitig | NCT01676259 | [17,18,19] |
| GalNAc GalNAc Cholesterol Folic acid (FA) CPP | siRNA siRNA siRNA microRNA PMO | ALAS1 HAO1 ApoB - c-myc | Acute hepatic porphyria Primary hyperoxaluria type 1 (liver targeting) Breast cancer (enhance PMO’s PK) | Yes, givosiran Yes, lumasiran - - - | (2019) (2020) - - - | - - - - - | [11] [13] [55,56] [57] [58] |
| Anti-body Antibody Fab’ + protamine | siRNA dsASO siRNA | STAT3 DRR/FAM107A c-myc, VEGF | Lewis-y positive cancer cell Glioblastoma stem cell HIV/AIDS | - - - | - - - | - - - | [44] [46] [43] |
| Antibody + exosome | ASO | miR-21 | Adenosquamous carcinoma | - | - | - | [27] |
| Exosome Exosome Exosome MSC-derived exosome Exosome + FA + PEI | siRNA hsiRNA siRNA siRNA siRNA/plasmid | GAPDH Huntintin S100A4 K-ras G12D K-ras/p53 | Alzheimer’s disease Huntington’s disease TNBC Pancreatic cancer Lung cancer | - - - - - | - - - I/recruiting - | - - - NCT03608631 - | [30] [32] [33] [35,36] [34] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oyama, S.; Yamamoto, T.; Yamayoshi, A. Recent Advances in the Delivery Carriers and Chemical Conjugation Strategies for Nucleic Acid Drugs. Cancers 2021, 13, 3881. https://doi.org/10.3390/cancers13153881
Oyama S, Yamamoto T, Yamayoshi A. Recent Advances in the Delivery Carriers and Chemical Conjugation Strategies for Nucleic Acid Drugs. Cancers. 2021; 13(15):3881. https://doi.org/10.3390/cancers13153881
Chicago/Turabian StyleOyama, Shota, Tsuyoshi Yamamoto, and Asako Yamayoshi. 2021. "Recent Advances in the Delivery Carriers and Chemical Conjugation Strategies for Nucleic Acid Drugs" Cancers 13, no. 15: 3881. https://doi.org/10.3390/cancers13153881
APA StyleOyama, S., Yamamoto, T., & Yamayoshi, A. (2021). Recent Advances in the Delivery Carriers and Chemical Conjugation Strategies for Nucleic Acid Drugs. Cancers, 13(15), 3881. https://doi.org/10.3390/cancers13153881







