A Nationwide Study on the Impact of Routine Testing for EGFR Mutations in Advanced NSCLC Reveals Distinct Survival Patterns Based on EGFR Mutation Subclasses
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Patient Selection
2.2. Data Extraction and Handling
2.3. Statistical Analysis
3. Results
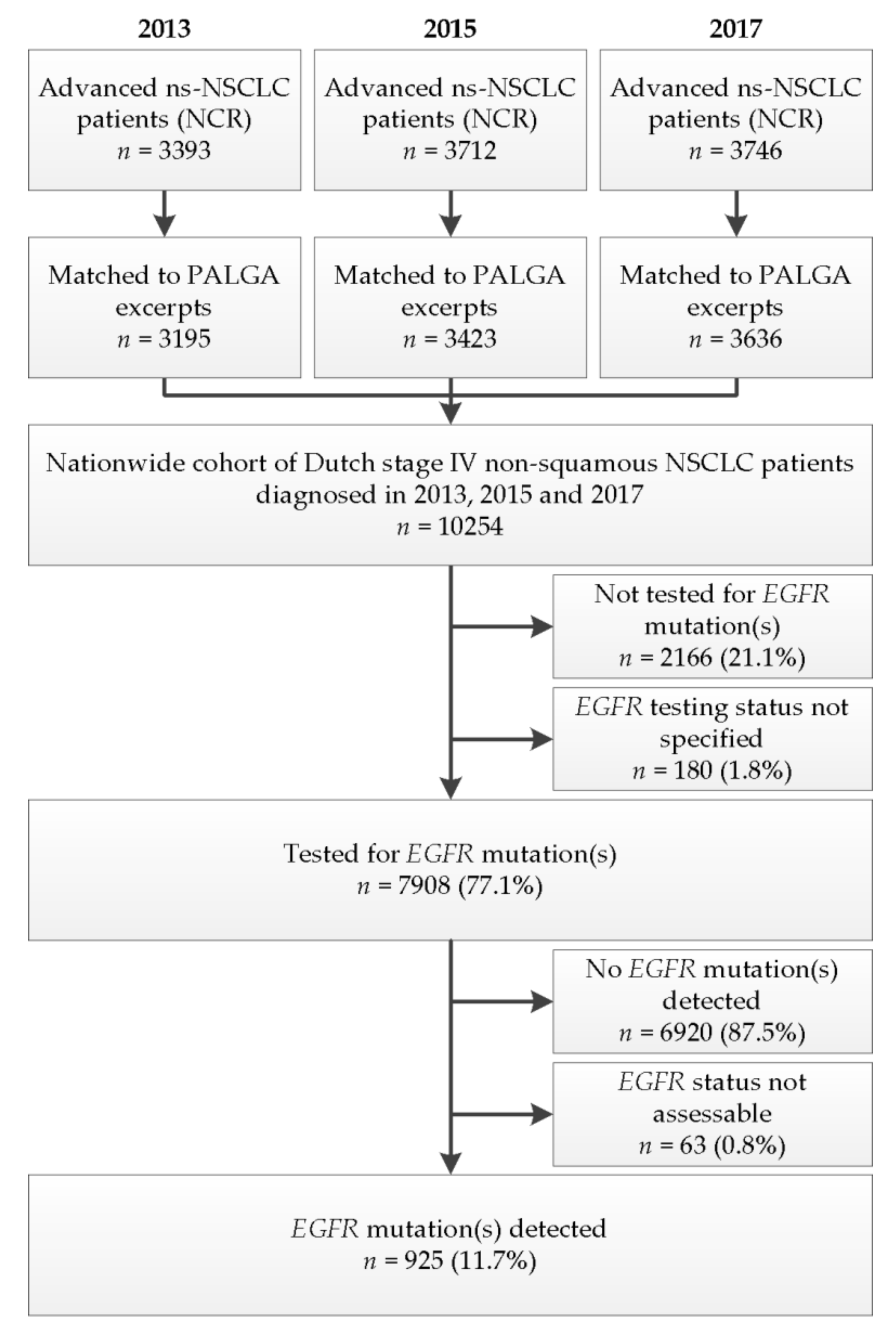
3.1. Patients Included in the Analysis
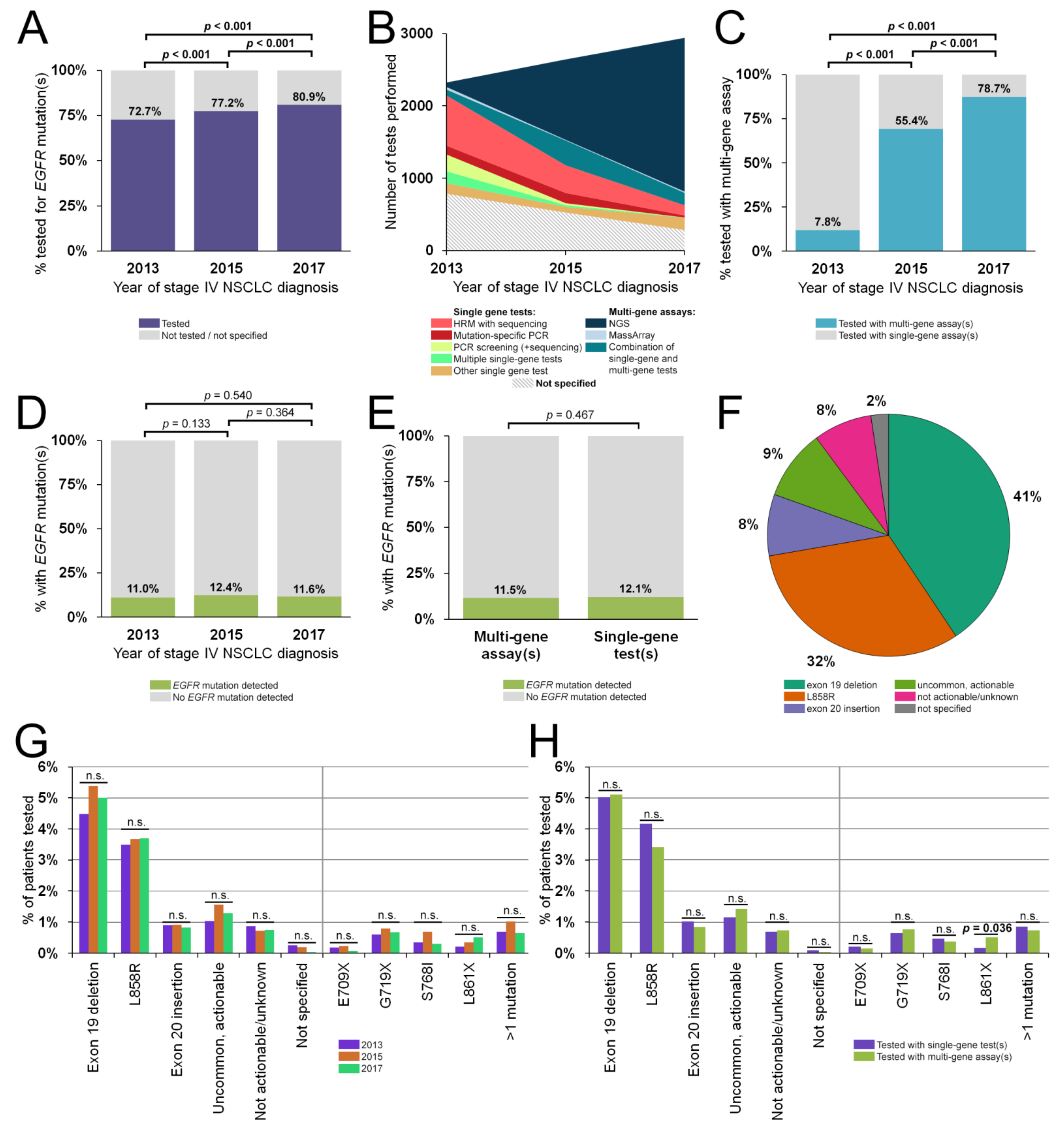
3.2. EGFR Testing Rates, Inter-Pathology Department Variance and Assays Used
3.3. Landscape of EGFR Mutations in Untreated Dutch NSCLC Patients
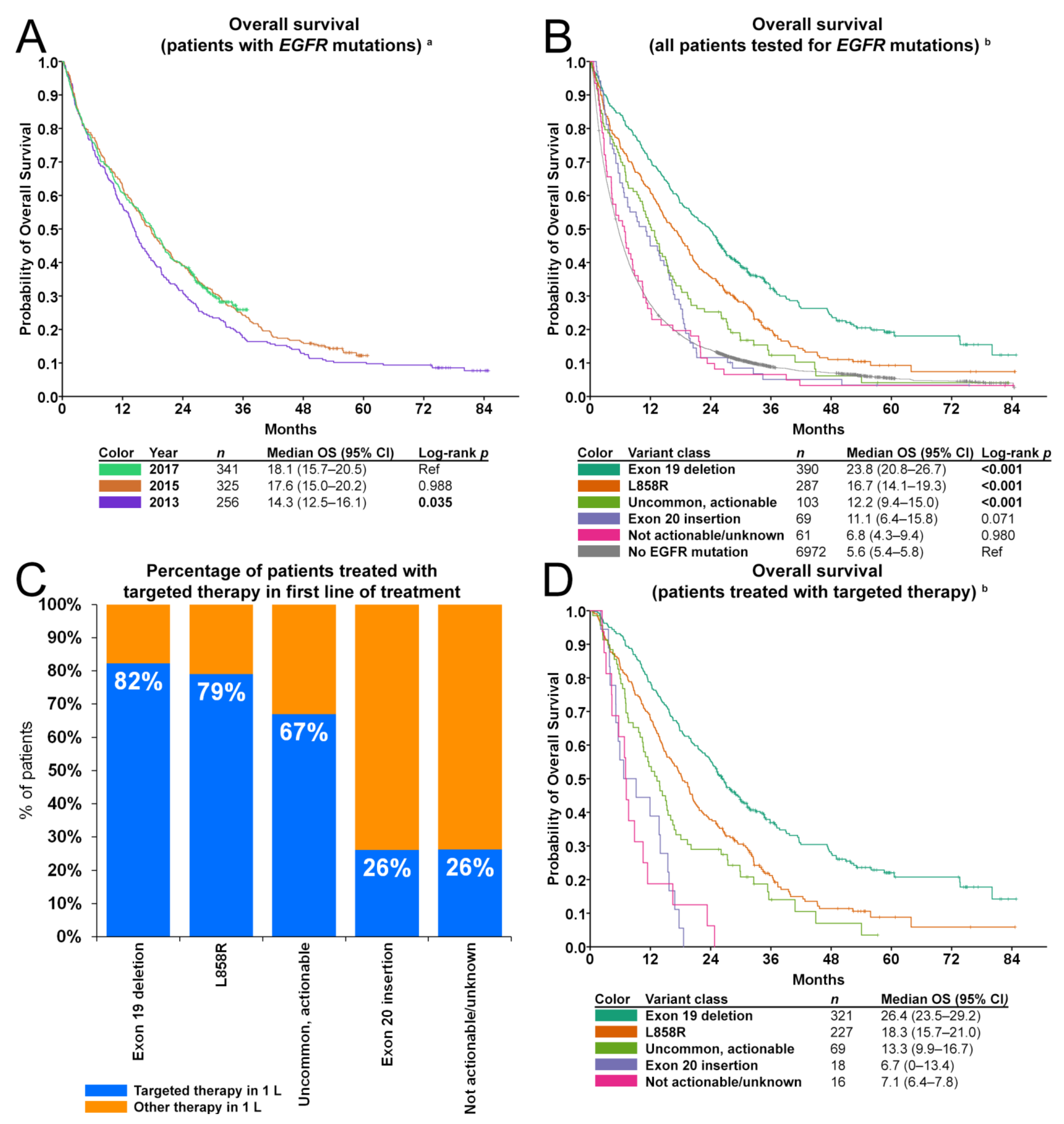
3.4. Impact of EGFR Testing on First-Line Targeted Therapy and Subsequent Overall Survival
3.4.1. Overall Survival of EGFR-Mutant Patients Versus Those without EGFR Mutations
3.4.2. Overall Survival in Patients Treated with First-Line Targeted Therapy
4. Discussion
4.1. Impact of Routine EGFR Testings on Detection of EGFR Mutations and Overall Survival
4.2. Beyond EGFR
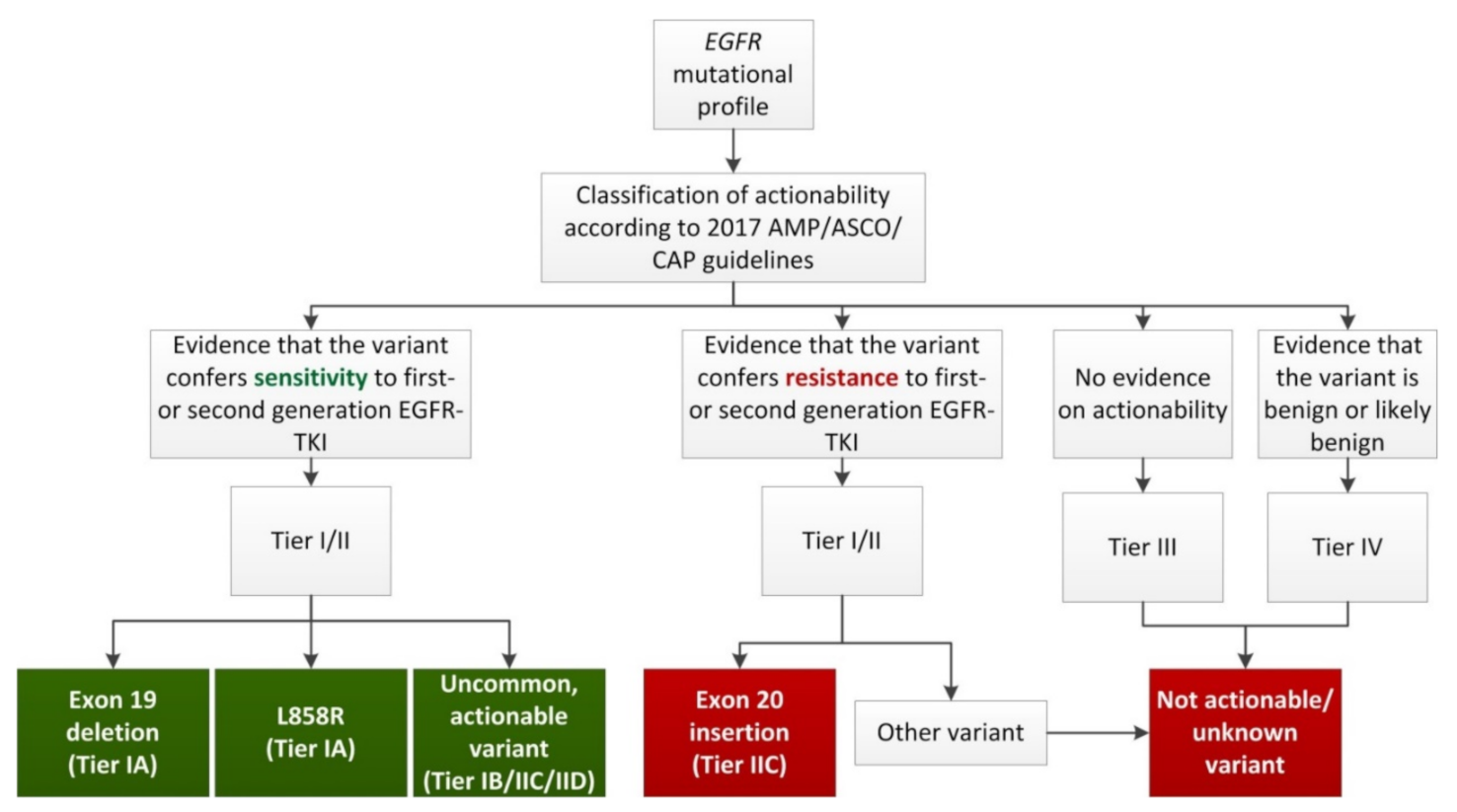
4.3. A Clinical Evidence-Driven Reclassification of EGFR Mutations
4.4. Limitations of the Study
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- AACR Project GENIE Consortium AACR Project GENIE: Powering precision medicine through an international consortium. Cancer Discov. 2017, 7, 818–831. [CrossRef] [PubMed] [Green Version]
- Sequist, L.V.; Martins, R.G.; Spigel, D.; Grunberg, S.M.; Spira, A.; Jänne, P.A.; Joshi, V.A.; McCollum, D.; Evans, T.L.; Muzikansky, A.; et al. First-line gefitinib in patients with advanced non-small-cell lung cancer harboring somatic EGFR mutations. J. Clin. Oncol. 2008, 26, 2442–2449. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.-L.; Zhou, C.; Liam, C.-K.; Wu, G.; Liu, X.; Zhong, Z.; Lu, S.; Cheng, Y.; Han, B.; Chen, L.; et al. First-line erlotinib versus gemcitabine/cisplatin in patients with advanced EGFR mutation-positive non-small-cell lung cancer: Analyses from the phase III, randomized, open-label, ENSURE study. Ann. Oncol. 2015, 26, 1883–1889. [Google Scholar] [CrossRef] [PubMed]
- Park, K.; Tan, E.-H.; O’Byrne, K.; Zhang, L.; Boyer, M.; Mok, T.; Hirsh, V.; Yang, J.C.-H.; Lee, K.H.; Lu, S.; et al. Afatinib versus gefitinib as first-line treatment of patients with EGFR mutation-positive non-small-cell lung cancer (LUX-Lung 7): A phase 2B, open-label, randomised controlled trial. Lancet Oncol. 2016, 17, 577–589. [Google Scholar] [CrossRef]
- Wu, Y.-L.; Cheng, Y.; Zhou, X.; Lee, K.H.; Nakagawa, K.; Niho, S.; Tsuji, F.; Linke, R.; Rosell, R.; Corral, J.; et al. Dacomitinib versus gefitinib as first-line treatment for patients with EGFR-mutation-positive non-small-cell lung cancer (ARCHER 1050): A randomised, open-label, phase 3 trial. Lancet Oncol. 2017, 18, 1454–1466. [Google Scholar] [CrossRef]
- Soria, J.-C.; Ohe, Y.; Vansteenkiste, J.; Reungwetwattana, T.; Chewaskulyong, B.; Lee, K.H.; Dechaphunkul, A.; Imamura, F.; Nogami, N.; Kurata, T.; et al. Osimertinib in untreated EGFR-mutated advanced non-small-cell lung cancer. N. Engl. J. Med. 2018, 378, 113–125. [Google Scholar] [CrossRef]
- D’Addario, G.; Früh, M.; Reck, M.; Baumann, P.; Klepetko, W.; Felip, E.; ESMO Guidelines Working Group. Metastatic non-small-cell lung cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2010, 21 (Suppl. 5), v116–v119. [Google Scholar] [CrossRef]
- Planchard, D.; Popat, S.; Kerr, K.; Novello, S.; Smit, E.F.; Faivre-Finn, C.; Mok, T.S.; Reck, M.; van Schil, P.E.; Hellmann, M.D.; et al. Metastatic non-small cell lung cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2018, 29, iv192–iv237. [Google Scholar] [CrossRef] [PubMed]
- Hendriks, L.E.L.; Dingemans, A.-M.C.; de Ruysscher, D.K.M.; Aarts, M.J.; Barberio, L.; Cornelissen, R.; Hartemink, K.J.; van den Heuvel, M.; Schuuring, E.; Smit, H.J.M.; et al. Lung cancer in the Netherlands. J. Thorac. Oncol. 2021, 16, 355–365. [Google Scholar] [CrossRef]
- Goodwin, S.; McPherson, J.D.; McCombie, W.R. Coming of age: Ten years of next-generation sequencing technologies. Nat. Rev. Genet. 2016, 17, 333–351. [Google Scholar] [CrossRef]
- Lin, M.-T.; Mosier, S.L.; Thiess, M.; Beierl, K.F.; Debeljak, M.; Tseng, L.-H.; Chen, G.; Yegnasubramanian, S.; Ho, H.; Cope, L.; et al. Clinical validation of KRAS, BRAF, and EGFR mutation detection using next-generation sequencing. Am. J. Clin. Pathol. 2014, 141, 856–866. [Google Scholar] [CrossRef] [Green Version]
- Planchard, D.; Besse, B.; Groen, H.J.M.; Souquet, P.-J.; Quoix, E.; Baik, C.S.; Barlesi, F.; Kim, T.M.; Mazieres, J.; Novello, S.; et al. Dabrafenib plus trametinib in patients with previously treated BRAF(V600E)-mutant metastatic non-small cell lung cancer: An open-label, multicentre phase 2 trial. Lancet Oncol. 2016, 17, 984–993. [Google Scholar] [CrossRef] [Green Version]
- Wolf, J.; Seto, T.; Han, J.-Y.; Reguart, N.; Garon, E.B.; Groen, H.J.M.; Tan, D.S.W.; Hida, T.; de Jonge, M.; Orlov, S.V.; et al. Capmatinib in MET Exon 14-mutated or MET-amplified non-small-cell lung cancer. N. Engl. J. Med. 2020, 383, 944–957. [Google Scholar] [CrossRef] [PubMed]
- Li, B.T.; Shen, R.; Buonocore, D.; Olah, Z.T.; Ni, A.; Ginsberg, M.S.; Ulaner, G.A.; Offin, M.; Feldman, D.; Hembrough, T.; et al. Ado-trastuzumab emtansine for patients with HER2-mutant lung cancers: Results from a phase II basket trial. J. Clin. Oncol. 2018, 36, 2532–2537. [Google Scholar] [CrossRef] [PubMed]
- UMCG Pathologie en Medische Biologie Moleculaire Diagnostiek. Available online: https://www.umcg.nl/NL/UMCG/Afdelingen/Pathologie/Professionals/moleculaire-diagnostiek/Paginas/default.aspx (accessed on 19 May 2020).
- Casparie, M.; Tiebosch, A.T.M.G.; Burger, G.; Blauwgeers, H.; van de Pol, A.; van Krieken, J.H.J.M.; Meijer, G.A. Pathology databanking and biobanking in The Netherlands, a central role for PALGA, the nationwide histopathology and cytopathology data network and archive. Cell. Oncol. 2007, 29, 19–24. [Google Scholar] [CrossRef]
- NVALT Niet-Kleincellig Longcarcinoom: Landelijke Richtlijn. Available online: https://richtlijnendatabase.nl/richtlijn/niet_kleincellig_longcarcinoom/startpagina_-_niet-kleincelling_longcarcinoom.html (accessed on 8 July 2021).
- Den Dunnen, J.T.; Dalgleish, R.; Maglott, D.R.; Hart, R.K.; Greenblatt, M.S.; McGowan-Jordan, J.; Roux, A.-F.; Smith, T.; Antonarakis, S.E.; Taschner, P.E.M. HGVS Recommendations for the description of sequence variants: 2016 update. Hum. Mutat. 2016, 37, 564–569. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, M.M.; Datto, M.; Duncavage, E.J.; Kulkarni, S.; Lindeman, N.I.; Roy, S.; Tsimberidou, A.M.; Vnencak-Jones, C.L.; Wolff, D.J.; Younes, A.; et al. Standards and guidelines for the interpretation and reporting of sequence variants in cancer: A joint consensus recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J. Mol. Diagn. 2017, 19, 4–23. [Google Scholar] [CrossRef] [Green Version]
- Heigener, D.F.; Schumann, C.; Sebastian, M.; Sadjadian, P.; Stehle, I.; Märten, A.; Lüers, A.; Griesinger, F.; Scheffler, M.; Abdollahi, A.; et al. Afatinib in non-small cell lung cancer harboring uncommon EGFR mutations pretreated with reversible EGFR inhibitors. Oncologist 2015, 20, 1167–1174. [Google Scholar] [CrossRef] [Green Version]
- Deans, Z.C.; Costa, J.L.; Cree, I.; Dequeker, E.; Edsjö, A.; Henderson, S.; Hummel, M.; Ligtenberg, M.J.; Loddo, M.; Machado, J.C.; et al. Integration of next-generation sequencing in clinical diagnostic molecular pathology laboratories for analysis of solid tumours; an expert opinion on behalf of IQN Path ASBL. Virchows Arch. 2017, 470, 5–20. [Google Scholar] [CrossRef] [Green Version]
- Boonstra, P.A.; ter Elst, A.; Tibbesma, M.; Bosman, L.J.; Mathijssen, R.; Atrafi, F.; van Coevorden, F.; Steeghs, N.; Farag, S.; Gelderblom, H.; et al. A single digital droplet PCR assay to detect multiple KIT exon 11 mutations in tumor and plasma from patients with gastrointestinal stromal tumors. Oncotarget 2018, 9, 13870–13883. [Google Scholar] [CrossRef] [Green Version]
- Sie, D.; Snijders, P.J.F.; Meijer, G.A.; Doeleman, M.W.; van Moorsel, M.I.H.; van Essen, H.F.; Eijk, P.P.; Grünberg, K.; van Grieken, N.C.T.; Thunnissen, E.; et al. Performance of amplicon-based next generation DNA sequencing for diagnostic gene mutation profiling in oncopathology. Cell. Oncol. 2014, 37, 353–361. [Google Scholar] [CrossRef]
- Steeghs, E.M.P.; Kroeze, L.I.; Tops, B.B.J.; van Kempen, L.C.; ter Elst, A.; Kastner-van Raaij, A.W.M.; Hendriks-Cornelissen, S.J.B.; Hermsen, M.J.W.; Jansen, E.A.M.; Nederlof, P.M.; et al. Comprehensive routine diagnostic screening to identify predictive mutations, gene amplifications, and microsatellite instability in FFPE tumor material. BMC Cancer 2020, 20, 291. [Google Scholar] [CrossRef]
- Gijtenbeek, R.G.P.; Damhuis, R.A.M.; Groen, H.J.M.; van der Wekken, A.J.; van Geffen, W.H. Nationwide real-world cohort study of first-line tyrosine kinase inhibitor treatment in epidermal growth factor receptor-mutated non-small-cell lung cancer. Clin. Lung Cancer 2020, 21, e647–e653. [Google Scholar] [CrossRef]
- Peters, S.; Adjei, A.A.; Gridelli, C.; Reck, M.; Kerr, K.; Felip, E.; ESMO Guidelines Working Group. Metastatic non-small-cell lung cancer (NSCLC): ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2012, 23 (Suppl. 7), vii56–vii64. [Google Scholar] [CrossRef]
- Smits, A.J.J.; Kummer, J.A.; Hinrichs, J.W.J.; Herder, G.J.M.; Scheidel-Jacobse, K.C.; Jiwa, N.M.; Ruijter, T.E.G.; Nooijen, P.T.G.A.; Looijen-Salamon, M.G.; Ligtenberg, M.J.L.; et al. EGFR and KRAS mutations in lung carcinomas in the Dutch population: Increased EGFR mutation frequency in malignant pleural effusion of lung adenocarcinoma. Cell. Oncol. 2012, 35, 189–196. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.-L.; Yuan, J.-Q.; Wang, K.-F.; Fu, X.-H.; Han, X.-R.; Threapleton, D.; Yang, Z.-Y.; Mao, C.; Tang, J.-L. The prevalence of EGFR mutation in patients with non-small cell lung cancer: A systematic review and meta-analysis. Oncotarget 2016, 7, 78985–78993. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brindel, A.; Althakfi, W.; Barritault, M.; Bringuier, P.-P.; Watkin, E.; Maury, J.-M.; Girard, N.; Brevet, M. Uncommon EGFR mutations in lung adenocarcinomas: Clinical features and response to tyrosine kinase inhibitors. Ann. Oncol. 2018, 29, viii747–viii748. [Google Scholar] [CrossRef]
- Leduc, C.; Merlio, J.P.; Besse, B.; Blons, H.; Debieuvre, D.; Bringuier, P.P.; Monnet, I.; Rouquette, I.; Fraboulet-Moreau, S.; Lemoine, A.; et al. Clinical and molecular characteristics of non-small-cell lung cancer (NSCLC) harboring EGFR mutation: Results of the nationwide French Cooperative Thoracic Intergroup (IFCT) program. Ann. Oncol. 2017, 28, 2715–2724. [Google Scholar] [CrossRef]
- Yun, C.-H.; Mengwasser, K.E.; Toms, A.V.; Woo, M.S.; Greulich, H.; Wong, K.-K.; Meyerson, M.; Eck, M.J. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc. Natl. Acad. Sci. USA 2008, 105, 2070–2075. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cross, D.A.E.; Ashton, S.E.; Ghiorghiu, S.; Eberlein, C.; Nebhan, C.A.; Spitzler, P.J.; Orme, J.P.; Finlay, M.R.V.; Ward, R.A.; Mellor, M.J.; et al. AZD9291, an irreversible EGFR TKI, overcomes T790M-mediated resistance to EGFR inhibitors in lung cancer. Cancer Discov. 2014, 4, 1046–1061. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reck, M.; Rodríguez-Abreu, D.; Robinson, A.G.; Hui, R.; Csőszi, T.; Fülöp, A.; Gottfried, M.; Peled, N.; Tafreshi, A.; Cuffe, S.; et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N. Engl. J. Med. 2016, 375, 1823–1833. [Google Scholar] [CrossRef] [Green Version]
- Lee, C.K.; Man, J.; Lord, S.; Links, M.; Gebski, V.; Mok, T.; Yang, J.C.-H. Checkpoint inhibitors in metastatic EGFR-mutated non-small cell lung cancer—A meta-analysis. J. Thorac. Oncol. 2017, 12, 403–407. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kwak, E.L.; Bang, Y.-J.; Camidge, D.R.; Shaw, A.T.; Solomon, B.; Maki, R.G.; Ou, S.-H.I.; Dezube, B.J.; Jänne, P.A.; Costa, D.B.; et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N. Engl. J. Med. 2010, 363, 1693–1703. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Drilon, A.; Siena, S.; Ou, S.-H.I.; Patel, M.; Ahn, M.J.; Lee, J.; Bauer, T.M.; Farago, A.F.; Wheler, J.J.; Liu, S.V.; et al. Safety and antitumor activity of the multitargeted pan-TRK, ROS1, and ALK inhibitor entrectinib: Combined results from two phase I trials (ALKA-372-001 and STARTRK-1). Cancer Discov. 2017, 7, 400–409. [Google Scholar] [CrossRef] [Green Version]
- Drilon, A.; Oxnard, G.R.; Tan, D.S.W.; Loong, H.H.F.; Johnson, M.; Gainor, J.; McCoach, C.E.; Gautschi, O.; Besse, B.; Cho, B.C.; et al. Efficacy of selpercatinib in RET fusion-positive non-small-cell lung cancer. N. Engl. J. Med. 2020, 383, 813–824. [Google Scholar] [CrossRef]
- Shaw, A.T.; Riely, G.J.; Bang, Y.-J.; Kim, D.-W.; Camidge, D.R.; Solomon, B.J.; Varella-Garcia, M.; Iafrate, A.J.; Shapiro, G.I.; Usari, T.; et al. Crizotinib in ROS1-rearranged advanced non-small-cell lung cancer (NSCLC): Updated results, including overall survival, from PROFILE 1001. Ann. Oncol. 2019. [Google Scholar] [CrossRef]
- Burns, T.F.; Borghaei, H.; Ramalingam, S.S.; Mok, T.S.; Peters, S. Targeting KRAS-mutant non-small-cell lung cancer: One mutation at a time, with a focus on KRAS G12C mutations. J. Clin. Oncol. 2020, 38, 4208–4218. [Google Scholar] [CrossRef] [PubMed]
- Gristina, V.; la Mantia, M.; Galvano, A.; Cutaia, S.; Barraco, N.; Castiglia, M.; Perez, A.; Bono, M.; Iacono, F.; Greco, M.; et al. Non-small cell lung cancer harboring concurrent EGFR genomic alterations: A systematic review and critical appraisal of the double dilemma. J. Mol. Pathol. 2021, 2, 173–196. [Google Scholar] [CrossRef]
- Canale, M.; Petracci, E.; Delmonte, A.; Chiadini, E.; Dazzi, C.; Papi, M.; Capelli, L.; Casanova, C.; de Luigi, N.; Mariotti, M.; et al. Impact of TP53 mutations on outcome in EGFR-mutated patients treated with first-line tyrosine kinase inhibitors. Clin. Cancer Res. 2017, 23, 2195–2202. [Google Scholar] [CrossRef] [Green Version]
- Horinouchi, H. To combine or not to combine: Anti-vascular endothelial growth factor therapies in EGFR mutation positive non-small cell lung cancer. Ann. Transl. Med. 2020, 8, 554. [Google Scholar] [CrossRef]
- Nakagawa, K.; Garon, E.B.; Seto, T.; Nishio, M.; Ponce Aix, S.; Paz-Ares, L.; Chiu, C.-H.; Park, K.; Novello, S.; Nadal, E.; et al. Ramucirumab plus erlotinib in patients with untreated, EGFR-mutated, advanced non-small-cell lung cancer (RELAY): A randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol. 2019, 20, 1655–1669. [Google Scholar] [CrossRef] [Green Version]
- Zhao, H.; Yao, W.; Min, X.; Gu, K.; Yu, G.; Zhang, Z.; Cui, J.; Miao, L.; Zhang, L.; Yuan, X.; et al. Apatinib plus gefitinib as first-line treatment in advanced EGFR-mutant NSCLC: The phase III ACTIVE study (CTONG1706). J. Thorac. Oncol. 2021. [Google Scholar] [CrossRef]
- Suda, K.; Mitsudomi, T.; Shintani, Y.; Okami, J.; Ito, H.; Ohtsuka, T.; Toyooka, S.; Mori, T.; Watanabe, S.-I.; Asamura, H.; et al. Clinical impacts of EGFR mutation status: Analysis of 5780 surgically resected lung cancer cases. Ann. Thorac. Surg. 2021, 111, 269–276. [Google Scholar] [CrossRef]
- Bergqvist, M.; Christensen, H.N.; Wiklund, F.; Bergström, S. Real world utilization of EGFR TKIs and prognostic factors for survival in NSCLC during 2010–2016 in Sweden: A nationwide observational study. Int. J. Cancer 2020, 146, 2510–2517. [Google Scholar] [CrossRef]
- Aye, P.S.; McKeage, M.J.; Tin Tin, S.; Khwaounjoo, P.; Elwood, J.M. Factors associated with overall survival in a population-based cohort of non- squamous NSCLC patients from northern New Zealand: A comparative analysis by EGFR mutation status. Cancer Epidemiol. 2020, 69, 101847. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.C.-H.; Sequist, L.V.; Geater, S.L.; Tsai, C.-M.; Mok, T.S.K.; Schuler, M.; Yamamoto, N.; Yu, C.-J.; Ou, S.-H.I.; Zhou, C.; et al. Clinical activity of afatinib in patients with advanced non-small-cell lung cancer harbouring uncommon EGFR mutations: A combined post-hoc analysis of LUX-Lung 2, LUX-Lung 3, and LUX-Lung 6. Lancet Oncol. 2015, 16, 830–838. [Google Scholar] [CrossRef]
- Passaro, A.; Prelaj, A.; Bonanno, L.; Tiseo, M.; Tuzi, A.; Proto, C.; Chiari, R.; Rocco, D.; Genova, C.; Sini, C.; et al. Activity of EGFR TKIs in Caucasian patients with NSCLC harboring potentially sensitive uncommon EGFR mutations. Clin. Lung Cancer 2019, 20, e186–e194. [Google Scholar] [CrossRef] [PubMed]
- Pilotto, S.; Rossi, A.; Vavalà, T.; Follador, A.; Tiseo, M.; Galetta, D.; Morabito, A.; di Maio, M.; Martelli, O.; Caffo, O.; et al. Outcomes of first-generation EGFR-TKIs against non-small-cell lung cancer harboring uncommon EGFR mutations: A post hoc analysis of the BE-POSITIVE study. Clin. Lung Cancer 2018, 19, 93–104. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, H.; Park, E.; Yun, C.-H.; Sng, N.J.; Lucena-Araujo, A.R.; Yeo, W.-L.; Huberman, M.S.; Cohen, D.W.; Nakayama, S.; Ishioka, K.; et al. Structural, biochemical, and clinical characterization of epidermal growth factor receptor (EGFR) exon 20 insertion mutations in lung cancer. Sci. Transl. Med. 2013, 5, 216ra177. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arrieta, O.; Cardona, A.F.; Corrales, L.; Campos-Parra, A.D.; Sánchez-Reyes, R.; Amieva-Rivera, E.; Rodríguez, J.; Vargas, C.; Carranza, H.; Otero, J.; et al. The impact of common and rare EGFR mutations in response to EGFR tyrosine kinase inhibitors and platinum-based chemotherapy in patients with non-small cell lung cancer. Lung Cancer 2015, 87, 169–175. [Google Scholar] [CrossRef]
- Chang, L.-C.; Lim, C.-K.; Chang, L.-Y.; Chen, K.-Y.; Shih, J.-Y.; Yu, C.-J. Non-small cell lung cancer harbouring non-resistant uncommon EGFR mutations: Mutation patterns, effectiveness of epidermal growth factor receptor-tyrosine kinase inhibitors and prognostic factors. Eur. J. Cancer 2019, 119, 77–86. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Yang, H.; Jiang, T.; Li, X.; Zhao, C.; Zhang, L.; Zhao, S.; Liu, X.; Jia, Y.; Wang, Y.; et al. Uncommon EGFR mutations in a cohort of Chinese NSCLC patients and outcomes of first-line EGFR-TKIs and platinum-based chemotherapy. Chin. J. Cancer Res. 2017, 29, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Jackman, D.M.; Yeap, B.Y.; Sequist, L.V.; Lindeman, N.; Holmes, A.J.; Joshi, V.A.; Bell, D.W.; Huberman, M.S.; Halmos, B.; Rabin, M.S.; et al. Exon 19 deletion mutations of epidermal growth factor receptor are associated with prolonged survival in non-small cell lung cancer patients treated with gefitinib or erlotinib. Clin. Cancer Res. 2006, 12, 3908–3914. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- IASCLC. Deeper Understanding of EGFR Mutation Subgroups Will Further Personalized Treatment for NSCLC. Available online: https://www.iaslc.org/iaslc-news/ilcn/deeper-understanding-egfr-mutation-subgroups-will-further-personalize-treatment (accessed on 16 February 2021).
- Gristina, V.; Malapelle, U.; Galvano, A.; Pisapia, P.; Pepe, F.; Rolfo, C.; Tortorici, S.; Bazan, V.; Troncone, G.; Russo, A. The significance of epidermal growth factor receptor uncommon mutations in non-small cell lung cancer: A systematic review and critical appraisal. Cancer Treat. Rev. 2020, 85, 101994. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.C.; Vivanco, I.; Beroukhim, R.; Huang, J.H.Y.; Feng, W.L.; DeBiasi, R.M.; Yoshimoto, K.; King, J.C.; Nghiemphu, P.; Yuza, Y.; et al. Epidermal growth factor receptor activation in glioblastoma through novel missense mutations in the extracellular domain. PLoS Med. 2006, 3, e485. [Google Scholar] [CrossRef]
- Ackerman, A.; Goldstein, M.A.; Kobayashi, S.; Costa, D.B. EGFR delE709_T710insD: A rare but potentially EGFR inhibitor responsive mutation in non–small-cell lung cancer. J. Thorac. Oncol. 2012, 7, e19–e20. [Google Scholar] [CrossRef] [Green Version]
- Koopman, B.; van der Wekken, A.J.; ter Elst, A.; Hiltermann, T.J.N.; Vilacha, J.F.; Groves, M.R.; van den Berg, A.; Hiddinga, B.I.; Hijmering-Kappelle, L.B.M.; Stigt, J.A.; et al. Relevance and effectiveness of molecular tumor board recommendations for patients with non–small-cell lung cancer with rare or complex mutational profiles. JCO Precis. Oncol. 2020, 4, 393–410. [Google Scholar] [CrossRef]
- Cappuzzo, F.; Soo, R.; Hochmair, M.; Schuler, M.; Lam, K.C.; Stehle, G.; Cseh, A.; Lorence, R.M.; Linden, S.; Forman, N.D.; et al. Global named patient use program of afatinib in advanced non-small-cell lung carcinoma patients who progressed following prior therapies. Future Oncol. 2018, 14, 1477–1486. [Google Scholar] [CrossRef] [Green Version]
- Sehgal, K.; Rangachari, D.; VanderLaan, P.A.; Kobayashi, S.S.; Costa, D.B. clinical benefit of tyrosine kinase inhibitors in advanced lung cancer with EGFR -G719A and other uncommon EGFR mutations. Oncologist 2021, 26, 281–287. [Google Scholar] [CrossRef]
- Tanaka, I.; Morise, M.; Kodama, Y.; Matsui, A.; Ozawa, N.; Ozone, S.; Goto, D.; Miyazawa, A.; Hase, T.; Hashimoto, N.; et al. Potential for afatinib as an optimal treatment for advanced non-small cell lung carcinoma in patients with uncommon EGFR mutations. Lung Cancer 2019, 127, 169–171. [Google Scholar] [CrossRef]
- Liang, S.-K.; Hsieh, M.-S.; Lee, M.-R.; Keng, L.-T.; Ko, J.-C.; Shih, J.-Y. Real-world experience of afatinib as a first-line therapy for advanced EGFR mutation-positive lung adenocarcinoma. Oncotarget 2017, 8, 90430–90443. [Google Scholar] [CrossRef] [Green Version]
- Brueckl, W.M.; Reck, M.; Schäfer, H.; Kortsik, C.; Gaska, T.; Rawluk, J.; Krüger, S.; Kokowski, K.; Budweiser, S.; Hoffmann, C.; et al. Efficacy of afatinib in the clinical practice: Final results of the GIDEON study in EGFR mutated non-small cell lung cancer (NSCLC) in Germany. J. Clin. Oncol. 2020, 38, e21636. [Google Scholar] [CrossRef]
- Caliman, E.; Petreni, P.; Brugia, M.; Antonuzzo, L.; Mazzoni, F. In regard to “Activity of EGFR TKIs in Caucasian Patients with NSCLC harbouring potentially sensitive uncommon EGFR mutations”. Clin. Lung Cancer 2020, 21, e363–e365. [Google Scholar] [CrossRef]
- Zhang, C.; Lin, L.; Zuo, R.; Wang, Y.; Chen, P. Response to tyrosine kinase inhibitors in lung adenocarcinoma with the rare epidermal growth factor receptor mutation S768I and G724S: A case report and literature review. Thorac. Cancer 2020, 11, 2743–2748. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Jiang, Q.; Xu, H.; Liu, A.; Huang, L. Two patients having NSCLC with novel duplication mutation in their EGFR gene (p.I740_K745dupIPVAIK) and their response to osimertinib. J. Thorac. Oncol. 2020, 15, e49–e51. [Google Scholar] [CrossRef] [PubMed]
- Liang, S.-K.; Ko, J.-C.; Yang, J.C.-H.; Shih, J.-Y. Afatinib is effective in the treatment of lung adenocarcinoma with uncommon EGFR p.L747P and p.L747S mutations. Lung Cancer 2019, 133, 103–109. [Google Scholar] [CrossRef]
- Hellmann, M.D.; Hayashi, T.; Reva, B.; Yu, H.A.; Riely, G.J.; Adusumilli, P.S.; Travis, W.D.; Wilkins, O.; Bramletta, N.; Chandramohan, R.; et al. Identification and functional characterization of EGFR V769M, a novel germline variant associated with multiple lung adenocarcinomas. JCO Precis. Oncol. 2017, 1–10. [Google Scholar] [CrossRef]
- Huo, R.; Li, J.; Li, X.; Shi, J.; Wang, K.; Jiao, J.; Shang, Y. Significant benefits of osimertinib against adenosquamous carcinoma harboring germline T790M mutation. Oncologist 2020, 25, 826–832. [Google Scholar] [CrossRef]
- Long, X.; Qin, T.; Lin, J. Great efficacy of afatinib in a patient with lung adenocarcinoma harboring EGFR L833V/H835L mutations: A case report. OncoTargets Ther. 2020, 13, 10689–10692. [Google Scholar] [CrossRef] [PubMed]
- Le Maignan, L.; Mirebeau-Prunier, D.; Vervueren, L.; Jeanfaivre, T.; Urban, T.; Hureaux, J. First case of A859T epidermal growth factor receptor mutation responding to erlotinib. J. Thorac. Oncol. 2011, 6, 639–640. [Google Scholar] [CrossRef] [Green Version]
- Wu, J.-Y.; Yu, C.-J.; Chang, Y.-C.; Yang, C.-H.; Shih, J.-Y.; Yang, P.-C. Effectiveness of Tyrosine kinase inhibitors on “uncommon” epidermal growth factor receptor mutations of unknown clinical significance in non–small cell lung cancer. Clin. Cancer Res. 2011, 17, 3812–3821. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alföldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 2020, 581, 434–443. [Google Scholar] [CrossRef]
- Cho, J.H.; Lim, S.H.; An, H.J.; Kim, K.H.; Park, K.U.; Kang, E.J.; Choi, Y.H.; Ahn, M.S.; Lee, M.H.; Sun, J.-M.; et al. Osimertinib for patients with non-small-cell lung cancer harboring uncommon EGFR mutations: A multicenter, open-label, phase II trial (KCSG-LU15-09). J. Clin. Oncol. 2020, 38, 488–495. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, H.; Kobayashi, S.; Costa, D.B. EGFR exon 20 insertion mutations in non-small-cell lung cancer: Preclinical data and clinical implications. Lancet Oncol. 2012, 13, e23–e31. [Google Scholar] [CrossRef]
- Yun, J.; Lee, S.-H.; Kim, S.-Y.; Jeong, S.-Y.; Kim, J.-H.; Pyo, K.-H.; Park, C.-W.; Heo, S.G.; Yun, M.R.; Lim, S.; et al. Antitumor activity of amivantamab (JNJ-61186372), an EGFR-MET bispecific antibody, in diverse models of EGFR exon 20 insertion-driven NSCLC. Cancer Discov. 2020, 10, 1194–1209. [Google Scholar] [CrossRef]
- Robichaux, J.P.; Elamin, Y.Y.; Tan, Z.; Carter, B.W.; Zhang, S.; Liu, S.; Li, S.; Chen, T.; Poteete, A.; Estrada-Bernal, A.; et al. Mechanisms and clinical activity of an EGFR and HER2 exon 20-selective kinase inhibitor in non-small cell lung cancer. Nat. Med. 2018, 24, 638–646. [Google Scholar] [CrossRef]
- Riely, G.J.; Neal, J.W.; Camidge, D.R.; Spira, A.I.; Piotrowska, Z.; Costa, D.B.; Tsao, A.S.; Patel, J.D.; Gadgeel, S.M.; Bazhenova, L.; et al. Activity and safety of mobocertinib (TAK-788) in previously treated non-small cell lung cancer with EGFR exon 20 insertion mutations from a phase 1/2 trial. Cancer Discov. 2021. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, H.; Yasuda, H.; Hamamoto, J.; Masuzawa, K.; Tani, T.; Nukaga, S.; Hirano, T.; Kobayashi, K.; Manabe, T.; Terai, H.; et al. Efficacy of afatinib or osimertinib plus cetuximab combination therapy for non-small-cell lung cancer with EGFR exon 20 insertion mutations. Lung Cancer 2019, 127, 146–152. [Google Scholar] [CrossRef]
- Koopman, B.; Groen, H.J.M.; Ligtenberg, M.J.L.; Grünberg, K.; Monkhorst, K.; Langen, A.J.; Boelens, M.C.; Paats, M.S.; Thüsen, J.H.; Dinjens, W.N.M.; et al. Multicenter comparison of molecular tumor boards in The Netherlands: Definition, composition, methods, and targeted therapy recommendations. Oncologist 2020. [Google Scholar] [CrossRef]
| Characteristic | Total | 2013 | 2015 | 2017 |
|---|---|---|---|---|
| Cases, n (%) | 10,254 (100) | 3195 (100) | 3423 (100) | 3636 (100) |
| Sex | ||||
| Male | 5498 (53.6) | 1730 (54.1) | 1820 (53.2) | 1947 (53.5) |
| Female | 4757 (46.4) | 1465 (45.9) | 1603 (46.8) | 1689 (46.5) |
| Age | ||||
| Mean (range) | 66.9 (20–101) | 66.2 (23–98) | 67.0 (24–97) | 67.4 (20–101) |
| <65 years | 4133 (40.3) | 1386 (43.4) | 1355 (39.6) | 1392 (38.3) |
| ≥65 years | 6121 (59.7) | 1809 (56.6) | 2068 (60.4) | 2244 (61.7) |
| Tumor histology | ||||
| Adenocarcinoma | 8845 (86.3) | 2724 (85.3) | 2961 (86.5) | 3160 (86.9) |
| Adenosquamous carcinoma | 62 (0.6) | 19 (0.6) | 25 (0.7) | 18 (0.5) |
| NSCLC, NOS | 1347 (13.1) | 452 (14.1) | 437 (12.8) | 458 (12.6) |
| EGFR mutation testing | ||||
| Tested | 7908 (77.1) | 2324 (72.7) | 2643 (77.2) | 2941 (80.9) |
| Not tested a | 2166 (21.1) | 765 (24.0) | 766 (22.4) | 635 (17.5) |
| Testing status not specified | 180 (1.8) | 106 (3.3) | 14 (0.4) | 60 (1.6) |
| EGFR test performed at initial diagnosis (% of n tested) | ||||
| Single-gene test | 2350 (29.7) | 1356 (58.3) | 653 (24.7) | 341 (11.6) |
| Fragment analysis | 8 (0.1) | 8 (0.3) | – | – |
| HRM (±sequencing) | 1215 (15.4) | 693 (29.8) | 382 (14.5) | 140 (4.8) |
| Immunohistochemistry | 20 (0.3) | 20 (0.9) | – | – |
| Mutation-specific PCR | 290 (3.7) | 121 (5.2) | 141 (5.3) | 28 (1.0) |
| PCR screening (±sequencing) b | 259 (3.3) | 232 (10.0) | 27 (1.0) | – |
| Pyrosequencing | 109 (1.4) | 2 (0.1) | 12 (0.5) | 95 (3.2) |
| Sanger sequencing | 257 (3.2) | 115 (4.9) | 67 (2.5) | 75 (2.6) |
| Combination of different single-gene tests | 192 (2.4) | 165 (7.1) | 24 (0.9) | 3 (0.1) |
| Multi-gene assay | 3958 (50.1) | 181 (7.8) | 1463 (55.4) | 2314 (78.7) |
| MassARRAY | 59 (0.7) | 35 (1.5) | 8 (0.3) | 16 (0.5) |
| NGS | 3299 (41.7) | 63 (2.7) | 1111 (42.0) | 2125 (72.3) |
| Combination of single- and multi-gene assays | 600 (7.6) | 83 (2.6) | 344 (13.0) | 176 (5.9) |
| Test not specified | 1600 (20.2) | 787 (33.9) | 527 (19.9) | 286 (9.7) |
| EGFR mutation status (% of n tested) | ||||
| EGFR mutation(s) reported | 925 (11.7) | 256 (11.0) | 328 (12.4) | 341 (11.6) |
| No EGFR mutation(s) | 6920 (87.5) | 2046 (88.0) | 2297 (86.9) | 2577 (87.6) |
| EGFR testing failed | 63 (0.8) | 22 (1.0) | 18 (0.7) | 23 (0.8) |
| A | By Year of Advanced NSCLC Diagnosis | ||||||
| Mutation(s) | Total | 2013 | 2015 | 2017 | p a | ||
| Tested for Mutations, n (%) | 7908 (100) | 2324 (100) | 2643 (100) | 2941 (100) | |||
| Any EGFR mutation | 925 (11.7) | 256 (11.0) | 328 (12.4) | 341 (11.6) | 0.303 | ||
| Distribution of predefined classes of sensitivity | |||||||
| Exon 19 deletion | 393 (5.0) | 104 (4.5) | 142 (5.4) | 147 (5.0) | 0.347 | ||
| L858R | 287 (3.6) | 81 (3.5) | 97 (3.7) | 109 (3.7) | 0.905 | ||
| Exon 20 insertion | 69 (0.9) | 21 (0.9) | 24 (0.9) | 24 (0.8) | 0.917 | ||
| Uncommon, actionable | 103 (1.3) | 24 (1.0) | 41 (1.6) | 38 (1.3) | 0.274 | ||
| Not actionable/unknown | 61 (0.8) | 20 (0.9) | 19 (0.7) | 22 (0.7) | 0.836 | ||
| Not specified | 12 (0.2) | 6 (0.3) | 5 (0.2) | 1 (0.0) | 0.097 | ||
| Distribution of uncommon EGFR mutation(s) | |||||||
| >1 EGFR mutation | 62 (0.8) | 16 (0.7) | 27 (1.0) | 19 (0.6) | 0.233 | ||
| E709X | 12 (0.2) | 4 (0.2) | 6 (0.2) | 2 (0.1) | 0.299 | ||
| G719X | 55 (0.7) | 14 (0.6) | 21 (0.8) | 20 (0.7) | 0.713 | ||
| S768I | 35 (0.4) | 8 (0.3) | 18 (0.7) | 9 (0.3) | 0.076 | ||
| L861X | 29 (0.4) | 5 (0.2) | 9 (0.3) | 15 (0.5) | 0.206 | ||
| B | By Molecular Diagnostic Modality | ||||||
| Mutation(s) | Total | Single-Gene Test(s) | Multi-Gene Assay | p c | |||
| Cases, n (%) | 6308 (100) b | 2350 (100) | 3958 (100) | ||||
| Distribution of predefined classes of sensitivity | |||||||
| Exon 19 deletion | 320 (5.1) | 118 (5.0) | 202 (5.1) | 0.906 | |||
| L858R | 233 (3.7) | 98 (4.2) | 135 (3.4) | 0.129 | |||
| Exon 20 insertion | 57 (0.9) | 24 (1.0) | 33 (0.8) | 0.492 | |||
| Uncommon, actionable | 83 (1.3) | 27 (1.1) | 56 (1.4) | 0.424 | |||
| Not actionable/unknown | 45 (0.7) | 16 (0.7) | 29 (0.7) | 0.878 | |||
| Not specified | 3 (0.0) | 2 (0.1) | 1 (0.0) | 0.560 | |||
| Distribution of individual uncommon EGFR mutation(s) | |||||||
| >1 EGFR mutation | 49 (0.8) | 20 (0.9) | 29 (0.7) | 0.657 | |||
| E709X | 11 (0.2) | 5 (0.2) | 6 (0.2) | 0.551 | |||
| G719X | 45 (0.7) | 15 (0.6) | 30 (0.8) | 0.645 | |||
| S768I | 26 (0.4) | 11 (0.5) | 15 (0.4) | 0.685 | |||
| L861X | 24 (0.4) | 4 (0.2) | 20 (0.5) | 0.036 | |||
| Factor | Univariate Analysis | Multivariate Analysis a | |||||
|---|---|---|---|---|---|---|---|
| n | HR | 95% CI | p | HR | 95% CI | p | |
| Age (cont.) | 651 | 1.02 | 1.01–1.03 | <0.001 | |||
| Age (in years) | |||||||
| <65 years | 258 | Ref | Ref | Ref | |||
| ≥65 years | 393 | 1.26 | 1.06–1.51 | 0.011 | |||
| Sex | |||||||
| Male | 223 | Ref | Ref | Ref | |||
| Female | 428 | 0.78 | 0.65–0.94 | 0.008 | |||
| Tumor histology | |||||||
| Adenocarcinoma | 630 | Ref | Ref | Ref | |||
| NSCLC, NOS | 19 | 1.45 | 0.88–2.39 | 0.142 | |||
| Adenosquamous carcinoma | 2 | 2.23 | 0.56–8.96 | 0.258 | |||
| Year of diagnosis | |||||||
| 2013 | 175 | 1.32 | 1.07–1.63 | 0.011 | |||
| 2015 | 223 | Ref | Ref | Ref | |||
| 2017 | 253 | 1.01 | 0.82–1.26 | 0.902 | |||
| EGFR mutation class (Ref: Exon 19 deletion) | |||||||
| Exon 19 deletion | 321 | Ref | Ref | Ref | Ref | Ref | Ref |
| L858R | 227 | 1.58 | 1.30–1.92 | <0.001 | 1.57 | 1.29–1.90 | <0.001 |
| Uncommon, actionable variant | 69 | 2.13 | 1.60–2.84 | <0.001 | 2.05 | 1.54–2.74 | <0.001 |
| Resistant/unknown variants b | 34 | 5.05 | 3.48–7.33 | <0.001 | 4.67 | 3.21–6.80 | <0.001 |
| EGFR mutation class (Ref: L858R) | |||||||
| L858R | 227 | Ref | Ref | Ref | Ref | Ref | Ref |
| Uncommon, actionable variant | 69 | 1.35 | 1.01–1.80 | 0.046 | 1.31 | 0.98–1.76 | 0.069 |
| Resistant/unknown variants b | 34 | 3.19 | 2.19–4.63 | <0.001 | 2.98 | 2.05–4.34 | <0.001 |
| EGFR mutation class (Ref: uncommon, actionable variant) | |||||||
| Uncommon, actionable variant | 69 | Ref | Ref | Ref | Ref | Ref | Ref |
| Resistant/unknown variants b | 34 | 2.37 | 1.55–3.63 | <0.001 | 2.27 | 1.48–3.49 | <0.001 |
| Variant(s) | n (%) a | Rationale | Prediction | LoE | Group |
|---|---|---|---|---|---|
| R108K | 1 (0.11%) | Known gain of function in brain tumors [59], but actionability unknown in NSCLC | no benefit | III | not actionable/unknown |
| C595F | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| Q701K | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| L703F | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| R705S | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| E709_T710delinsD | 3 (0.32%) | Clinical sensitivity to EGFR-TKI reported [60,61] | sensitive | IID | uncommon, actionable |
| E709A/G719A | 4 (0.43%) | Clinical sensitivity to EGFR-TKI reported [21] | sensitive | IIC | uncommon, actionable |
| E709A/G719R | 1 (0.11%) | Considered comparable to E709A/G719A | sensitive | IIC | uncommon, actionable |
| E709A/G719S | 1 (0.11%) | Considered comparable to E709A/G719A | sensitive | IIC | uncommon, actionable |
| E709D | 1 (0.11%) | No evidence on pathogenicity or actionability. Similar amino acid properties between Glu (E) and Asp (D), thus no effect expected | no benefit | III | not actionable/unknown |
| E709K/G719S | 1 (0.11%) | Clinical sensitivity to EGFR-TKI reported [62] | sensitive | IIC | uncommon, actionable |
| I715fs* | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| G719_S720delinsAF | 1 (0.11%) | Considered comparable to G719A | sensitive | IID | uncommon, actionable |
| G719A | 18 (1.95%) | Clinical sensitivity to EGFR-TKI reported [63] | sensitive | IIC | uncommon, actionable |
| G719A/D761Y | 1 (0.11%) | Considered comparable to G719A | sensitive | IIC | uncommon, actionable |
| G719A + Exon 20 insertion, NOS | 1 (0.11%) | No EGFR-TKI sensitivity expected due to exon 20 insertion, grouped accordingly | no benefit | IIC | exon 20 insertion |
| G719A/L861Q | 3 (0.32%) | Clinical sensitivity to EGFR-TKI reported [64] | sensitive | IIC | uncommon, actionable |
| G719A/R776H | 1 (0.11%) | Considered comparable to G719A | sensitive | IIC | uncommon, actionable |
| G719A/S768I | 5 (0.54%) | Clinical sensitivity to EGFR-TKI reported [65] | sensitive | IIC | uncommon, actionable |
| G719A/V769M | 1 (0.11%) | Considered comparable to G719A | sensitive | IIC | uncommon, actionable |
| G719C | 2 (0.22%) | Clinical sensitivity to EGFR-TKI reported [66] | sensitive | IIC | uncommon, actionable |
| G719C/S768I | 6 (0.65%) | Clinical sensitivity to EGFR-TKI reported [67] | sensitive | IIC | uncommon, actionable |
| G719S | 1 (0.11%) | Clinical sensitivity to EGFR-TKI reported [66] | sensitive | IIC | uncommon, actionable |
| G719S/L747S | 1 (0.11%) | Considered comparable to G719S | sensitive | IIC | uncommon, actionable |
| G719S/L861Q | 1 (0.11%) | Considered comparable to G719A/L861Q | sensitive | IIC | uncommon, actionable |
| G719S/S768I | 1 (0.11%) | Considered comparable to G719A/S768I and G719C/S768I | sensitive | IIC | uncommon, actionable |
| G719X, NOS | 2 (0.22%) | Similar grouping as other G719 substitutions | sensitive | IIC | uncommon, actionable |
| G719X, NOS/S768I | 3 (0.32%) | Similar grouping as other G719X/S768I variants | sensitive | IIC | uncommon, actionable |
| G724A/S768I | 1 (0.11%) | Considered comparable to G724S/S768I | sensitive | IIC | uncommon, actionable |
| G724S/S768I | 1 (0.11%) | Clinical sensitivity to EGFR-TKI reported [68] | sensitive | IID | uncommon, actionable |
| c.2184 + 19G > A | 1 (0.11%) | Likely a SNP due to high allele frequency in general population (3.5%; GnomAD) b | no benefit | IV | not actionable/unknown |
| L730fs*1 | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| I740_K745dup | 2 (0.22%) | Clinical sensitivity to osimertinib reported [69], but no evidence on first- or second-generation TKI | no benefit | III | not actionable/unknown |
| A743S | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| I744_P753delinsSNISG | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_A750del | 210 (22.7%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_A750del/G873E | 1 (0.11%) | Considered comparable to E746_A750del | sensitive | IA | exon 19 deletion |
| E746_A750del/K754Q | 1 (0.11%) | Considered comparable to E746_A750del | sensitive | IA | exon 19 deletion |
| E746_A750del/P848L | 1 (0.11%) | Considered comparable to E746_A750del | sensitive | IA | exon 19 deletion |
| E746_A750del/V765M | 1 (0.11%) | Considered comparable to E746_A750del | sensitive | IA | exon 19 deletion |
| E746_A750delinsEP | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_A750delinsIP | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_A750dup | 1 (0.11%) | Considered comparable to I740_K745dup | no benefit | III | not actionable/unknown |
| E746_K754delinsVSR | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_L747delinsIP | 1 (0.11%) | Considered comparable to L747P | sensitive | IID | uncommon, actionable |
| E746_P753delinsANKE | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_P753delinsIS | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_P753delinsVS | 2 (0.22%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_S752delinsI | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_S752delinsV | 20 (2.16%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_T751del | 2 (0.22%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_T751delinsA | 3 (0.32%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_T751delinsAA | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_T751delinsI | 2 (0.22%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_T751delinsK | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_T751delinsL | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_T751delinsP | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_T751delinsS | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| E746_T751delinsVP | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_A750delinsP | 10 (1.08%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_A750delinsP/V845L | 1 (0.11%) | Considered comparable to L747_A750delinsP | sensitive | IA | exon 19 deletion |
| L747_A755delinsSKD | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_E749del | 3 (0.32%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_K754delinsATSPE | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_K754delinsG | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_K754delinsQPN | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_P753delinsQ | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_P753delinsS | 25 (2.70%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_S752del | 7 (0.76%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_S752del/K754R | 1 (0.11%) | Considered comparable to L747_S752del | sensitive | IA | exon 19 deletion |
| L747_S752del/L777Q | 1 (0.11%) | Considered comparable to L747_S752del | sensitive | IA | exon 19 deletion |
| L747_S752delinsQ | 2 (0.22%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_T751del | 16 (1.73%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747_T751del/S768I | 1 (0.11%) | Considered comparable to L747_T751del | sensitive | IA | exon 19 deletion |
| L747_T751delinsP | 6 (0.65%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| L747P | 4 (0.43%) | Clinical sensitivity to EGFR-TKI reported [70] | sensitive | IID | uncommon, actionable |
| E749_A755delinsD | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| A750_E758delinsP | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| T751_I759delinsN | 1 (0.11%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| S752_I759del | 4 (0.43%) | Net loss of amino acids on exon 19 (deletion) | sensitive | IA | exon 19 deletion |
| P753L | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| K757R | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| A763S | 2 (0.22%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| A763_Y764insFQEA | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| V765M | 2 (0.22%) | Only reported in combination with other variants, no evidence on individual variant | no benefit | III | not actionable/unknown |
| A767_V769dup | 16 (1.73%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| A767T | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| S768_D770dup | 9 (0.97%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| S768_V769delinsIL | 4 (0.43%) | Considered comparable to S768I | sensitive | IIC | uncommon, actionable |
| S768I | 5 (0.54%) | Clinical sensitivity to EGFR-TKI reported [49] | sensitive | IIC | uncommon, actionable |
| S768I/L861Q | 2 (0.22%) | Clinical sensitivity to EGFR-TKI reported [65] | sensitive | IIC | uncommon, actionable |
| S768I/V774M | 2 (0.22%) | Considered comparable to S768I | sensitive | IIC | uncommon, actionable |
| V769L | 2 (0.22%) | Only reported in combination with other variants, no evidence on individual variant | no benefit | III | not actionable/unknown |
| V769M | 2 (0.22%) | Possible germline variant [71], no evidence sensitivity to EGFR-TKI | no benefit | III | not actionable/unknown |
| D770_H773dup | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| D770_N771insG | 3 (0.32%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| D770_N771insGF | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| D770_N771insSVA | 2 (0.22%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| D770_N771insT | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| D770_N771insY | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| D770_P772dup | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| D770delinsEQPP | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| D770delinsGY | 3 (0.32%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| D770Y | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| N771_H773dup | 2 (0.22%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| N771_P772insH | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| N771_P772insR | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| N771delinsGF | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| N771delinsGY | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| N771delinsKG | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| N771delinsTH | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| N771L | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| P772_C775dup | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| P772_H773dup | 3 (0.32%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| P772_H773insANP | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| H773_V774dup | 2 (0.22%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| H773_V774insAH | 2 (0.22%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| H773delinsYNPY | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| H773dup | 5 (0.54%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| V774delinsHC | 1 (0.11%) | Net gain of amino acids on exon 20 (insertion) | no benefit | IIC | exon 20 insertion |
| V774M | 3 (0.32%) | Only reported in combination with other variants, no evidence on individual variant | sensitive | III | not actionable/unknown |
| V774M/L861R | 1 (0.11%) | Considered comparable to L861R | sensitive | IIC | uncommon, actionable |
| C775F | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| R776H | 1 (0.11%) | Only reported in combination with other variants, no evidence on individual variant | no benefit | III | not actionable/unknown |
| R776L | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| G779F | 3 (0.32%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| G779S | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| C781F | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| Q787E | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| T790M | 1 (0.11%) | Known resistance-inducing mutation, but not transforming without a driver mutationg [72] | no benefit | III | not actionable/unknown |
| G796C | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| L799M | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| D830Y | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| L832T | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| L833V/H835L | 2 (0.22%) | Clinical sensitivity to EGFR-TKI reported [73] | sensitive | IID | uncommon, actionable |
| R836L | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| R836R | 4 (0.43%) | Likely a SNP due to high allele frequency in general population (1.7%; GnomAD) b | no benefit | IV | not actionable/unknown |
| A840T | 4 (0.43%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| P848L | 5 (0.54%) | No evidence on pathogenicity or actionability, may be a rare SNP (AF 0.03; GnomAD) b | no benefit | IV | not actionable/unknown |
| L858_K860delinsRTI | 1 (0.11%) | Considered comparable to L858R | sensitive | IID | L858R |
| L858R | 274 (29.6%) | Classical, well known activating mutation sensitive to first- and second-generation EGFR-TKI | sensitive | IA | L858R |
| L858R/A871E | 1 (0.11%) | Considered comparable to L858R | sensitive | IA | L858R |
| L858R/A871G | 1 (0.11%) | Considered comparable to L858R | sensitive | IA | L858R |
| L858R/E709G | 1 (0.11%) | Considered comparable to L858R | sensitive | IA | L858R |
| L858R/L718M | 1 (0.11%) | Considered comparable to L858R | sensitive | IA | L858R |
| L858R/L747V | 1 (0.11%) | Considered comparable to L858R | sensitive | IA | L858R |
| L858R/R776H | 2 (0.22%) | Considered comparable to L858R | sensitive | IA | L858R |
| L858R/S768I | 4 (0.43%) | Considered comparable to L858R | sensitive | IA | L858R |
| L858R/V834L | 1 (0.11%) | Considered comparable to L858R | sensitive | IA | L858R |
| A859T | 1 (0.11%) | Clinical sensitivity to EGFR-TKI reported [74] | sensitive | IID | uncommon, actionable |
| L861Q | 19 (2.05%) | Clinical sensitivity to EGFR-TKI reported [75] | sensitive | IIC | uncommon, actionable |
| L861R | 3 (0.32%) | Clinical sensitivity to EGFR-TKI reported [75] | sensitive | IIC | uncommon, actionable |
| G863D | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| A864T | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| A864V | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| E866K | 1 (0.11%) | No evidence on pathogenicity or actionability | no benefit | III | not actionable/unknown |
| EGFR variant, NOS | 4 (0.43%) | Exact variant and exon not reported | unknown | NA | unspecified |
| Exon 18 variant, NOS + Exon 20 variant, NOS | 2 (0.22%) | Exact variant not reported | unknown | NA | unspecified |
| Exon 19 deletion, NOS | 52 (5.62%) | Exact variant not reported but confirmed net loss of amino acids on exon 19 (deletion) | unknown | NA | exon 19 deletion |
| Exon 19 variant, NOS | 3 (0.32%) | Exact variant not reported (could be a deletion or a single-nucleotide variant) | sensitive | IA | unspecified |
| Exon 20 insertion, NOS | 3 (0.32%) | Exact variant not reported but confirmed net gain of amino acids on exon 21 (insertion) | unknown | NA | exon 20 insertion |
| Exon 20 SNV (silent) | 1 (0.11%) | Silent mutation, no change in amino acids | no benefit | IIC | not actionable/unknown |
| Exon 21 variant, NOS | 3 (0.32%) | Exact variant not reported (could be L858R or a different, non-classical mutation) | no benefit | IV | unspecified |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Koopman, B.; Cajiao Garcia, B.N.; Kuijpers, C.C.H.J.; Damhuis, R.A.M.; van der Wekken, A.J.; Groen, H.J.M.; Schuuring, E.; Willems, S.M.; van Kempen, L.C. A Nationwide Study on the Impact of Routine Testing for EGFR Mutations in Advanced NSCLC Reveals Distinct Survival Patterns Based on EGFR Mutation Subclasses. Cancers 2021, 13, 3641. https://doi.org/10.3390/cancers13143641
Koopman B, Cajiao Garcia BN, Kuijpers CCHJ, Damhuis RAM, van der Wekken AJ, Groen HJM, Schuuring E, Willems SM, van Kempen LC. A Nationwide Study on the Impact of Routine Testing for EGFR Mutations in Advanced NSCLC Reveals Distinct Survival Patterns Based on EGFR Mutation Subclasses. Cancers. 2021; 13(14):3641. https://doi.org/10.3390/cancers13143641
Chicago/Turabian StyleKoopman, Bart, Betzabel N. Cajiao Garcia, Chantal C. H. J. Kuijpers, Ronald A. M. Damhuis, Anthonie J. van der Wekken, Harry J. M. Groen, Ed Schuuring, Stefan M. Willems, and Léon C. van Kempen. 2021. "A Nationwide Study on the Impact of Routine Testing for EGFR Mutations in Advanced NSCLC Reveals Distinct Survival Patterns Based on EGFR Mutation Subclasses" Cancers 13, no. 14: 3641. https://doi.org/10.3390/cancers13143641
APA StyleKoopman, B., Cajiao Garcia, B. N., Kuijpers, C. C. H. J., Damhuis, R. A. M., van der Wekken, A. J., Groen, H. J. M., Schuuring, E., Willems, S. M., & van Kempen, L. C. (2021). A Nationwide Study on the Impact of Routine Testing for EGFR Mutations in Advanced NSCLC Reveals Distinct Survival Patterns Based on EGFR Mutation Subclasses. Cancers, 13(14), 3641. https://doi.org/10.3390/cancers13143641







