Investigation of the Effect of KIR–HLA Pairs on Hepatocellular Carcinoma in Hepatitis C Virus Cirrhotic Patients
Abstract
:Simple Summary
Abstract
1. Introduction
2. Results
2.1. Patient Characteristics
2.2. HLA and KIR Genotyping in Patients with HCV-Induced Cirrhosis
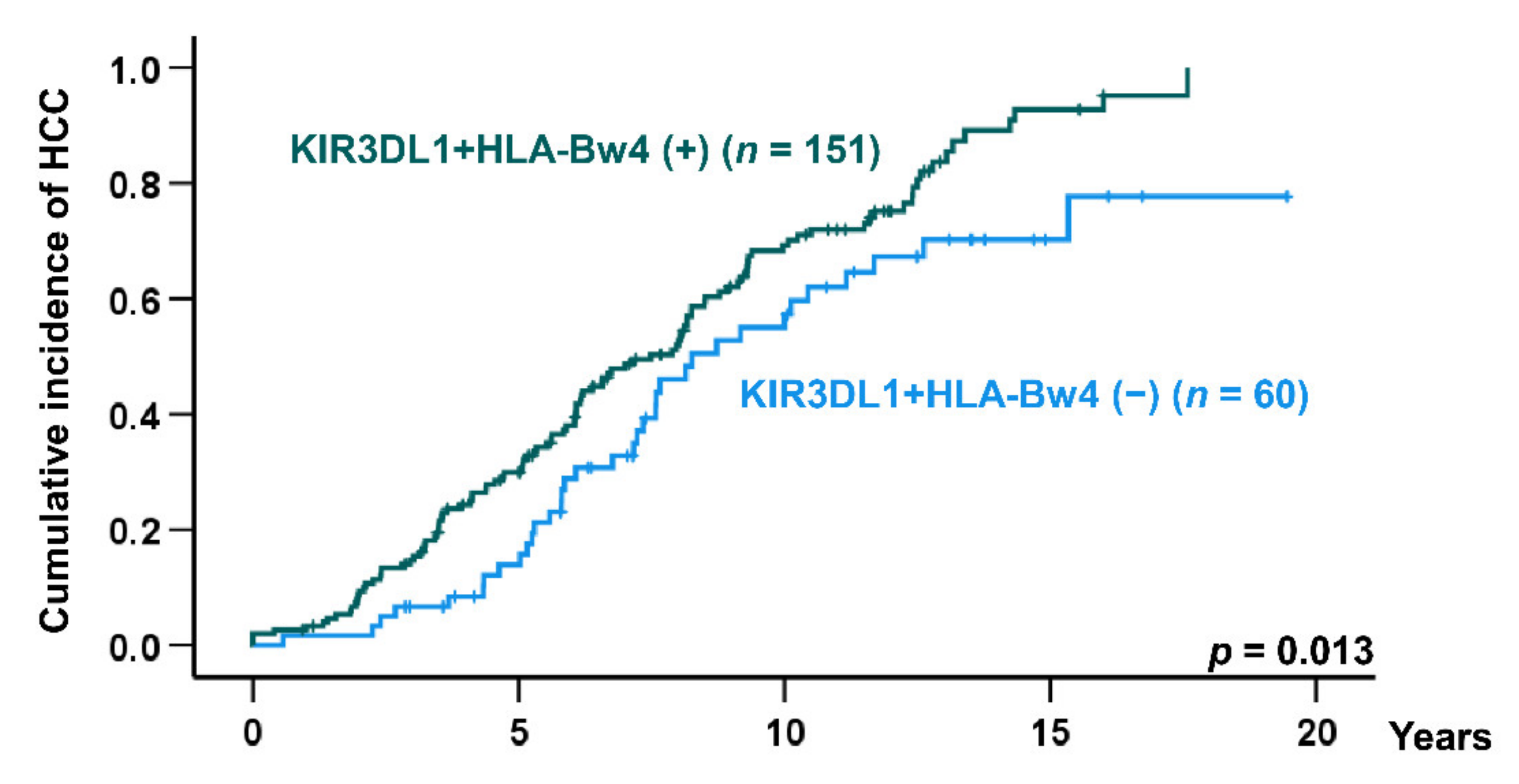
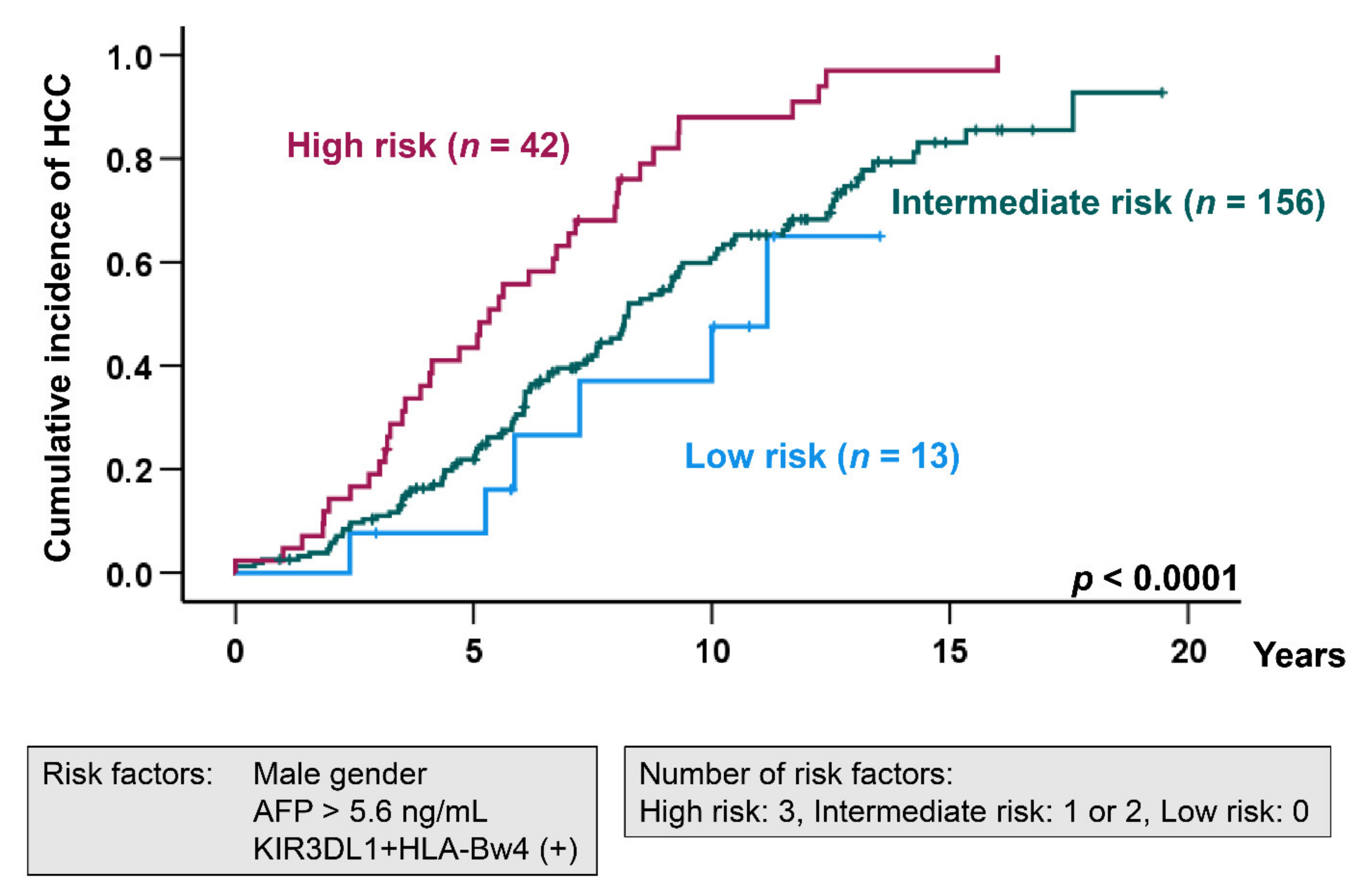
2.3. Risk Factors for HCC Development in HCV-Induced Cirrhosis Patients
3. Discussion
4. Materials and Methods
4.1. Study Population and HCC Surveillance
4.2. HLA Class I and KIR Typing
4.3. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kiyosawa, K.; Sodeyama, T.; Tanaka, E.; Gibo, Y.; Yoshizawa, K.; Nakano, Y.; Furuta, S.; Akahane, Y.; Nishioka, K.; Purcell, R.H.; et al. Interrelationship of blood transfusion, non-A, non-B hepatitis and hepatocellular carcinoma: Analysis by detection of antibody to hepatitis C virus. Hepatology 1990, 12, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Villanueva, A. Hepatocellular Carcinoma. New Engl. J. Med. 2019, 380, 1450–1462. [Google Scholar] [CrossRef] [Green Version]
- Umemura, T.; Ichijo, T.; Yoshizawa, K.; Tanaka, E.; Kiyosawa, K. Epidemiology of hepatocellular carcinoma in Japan. J. Gastroenterol. 2009, 44 (Suppl. 19), 102–107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singal, A.G.; Lampertico, P.; Nahon, P. Epidemiology and surveillance for hepatocellular carcinoma: New trends. J. Hepatol. 2020, 72, 250–261. [Google Scholar] [CrossRef] [Green Version]
- Tateishi, R.; Uchino, K.; Fujiwara, N.; Takehara, T.; Okanoue, T.; Seike, M.; Yoshiji, H.; Yatsuhashi, H.; Shimizu, M.; Torimura, T.; et al. A nationwide survey on non-B, non-C hepatocellular carcinoma in Japan: 2011-2015 update. J. Gastroenterol. 2019, 54, 367–376. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muller, M.; Bird, T.G.; Nault, J.C. The landscape of gene mutations in cirrhosis and hepatocellular carcinoma. J. Hepatol. 2020, 72, 990–1002. [Google Scholar] [CrossRef] [Green Version]
- Caruso, S.; O’Brien, D.R.; Cleary, S.P.; Roberts, L.R.; Zucman-Rossi, J. Genetics of Hepatocellular Carcinoma: Approaches to Explore Molecular Diversity. Hepatology. 2021, 73 (Suppl. S1), 14–26. [Google Scholar] [CrossRef]
- Vivier, E.; Tomasello, E.; Baratin, M.; Walzer, T.; Ugolini, S. Functions of natural killer cells. Nat. Immunol. 2008, 9, 503–510. [Google Scholar] [CrossRef]
- Martinet, L.; Smyth, M.J. Balancing natural killer cell activation through paired receptors. Nat. Rev. Immunol. 2015, 15, 243–254. [Google Scholar] [CrossRef]
- Parham, P. MHC class I molecules and KIRs in human history, health and survival. Nat. Rev. Immunol. 2005, 5, 201–214. [Google Scholar] [CrossRef]
- Colonna, M.; Borsellino, G.; Falco, M.; Ferrara, G.B.; Strominger, J.L. HLA-C is the inhibitory ligand that determines dominant resistance to lysis by NK1- and NK2-specific natural killer cells. Proc. Natl. Acad. Sci. USA 1993, 90, 12000–12004. [Google Scholar] [CrossRef] [Green Version]
- Cella, M.; Longo, A.; Ferrara, G.B.; Strominger, J.L.; Colonna, M. NK3-specific natural killer cells are selectively inhibited by Bw4-positive HLA alleles with isoleucine 80. J. Exp. Med. 1994, 180, 1235–1242. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Naumova, E.; Mihaylova, A.; Stoitchkov, K.; Ivanova, M.; Quin, L.; Toneva, M. Genetic polymorphism of NK receptors and their ligands in melanoma patients: Prevalence of inhibitory over activating signals. Cancer Immunol. Immunother. 2005, 54, 172–178. [Google Scholar] [CrossRef] [PubMed]
- Naumova, E.; Mihaylova, A.; Ivanova, M.; Mihailova, S. Impact of KIR/HLA ligand combinations on immune responses in malignant melanoma. Cancer Immunol. Immunother. 2007, 56, 95–100. [Google Scholar] [CrossRef] [PubMed]
- Saito, H.; Umemura, T.; Joshita, S.; Yamazaki, T.; Fujimori, N.; Kimura, T.; Komatsu, M.; Matsumoto, A.; Tanaka, E.; Ota, M. KIR2DL2 combined with HLA-C1 confers risk of hepatitis C virus-related hepatocellular carcinoma in younger patients. Oncotarge. 2018, 9, 19650–19661. [Google Scholar] [CrossRef]
- Lopez-Vazquez, A.; Rodrigo, L.; Martinez-Borra, J.; Perez, R.; Rodriguez, M.; Fdez-Morera, J.L.; Fuentes, D.; Rodriguez-Rodero, S.; Gonzaez, S.; Lopez-Larrea, C. Protective effect of the HLA-Bw4I80 epitope and the killer cell immunoglobulin-like receptor 3DS1 gene against the development of hepatocellular carcinoma in patients with hepatitis C virus infection. J. Infect. Dis. 2005, 192, 162–165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Re, V.; Caggiari, L.; de Zorzi, M.; Repetto, O.; Zignego, A.L.; Izzo, F.; Tornesello, M.L.; Buonaguro, F.M.; Mangia, A.; Sansonno, D.; et al. Genetic diversity of the KIR/HLA system and susceptibility to hepatitis C virus-related diseases. PLoS ONE 2015, 10, e0117420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mele, D.; Pasi, A.; Cacciatore, R.; Mantovani, S.; Oliviero, B.; Mondelli, M.U.; Varchetta, S. Decreased interferon-gamma production by NK cells from KIR haplotype B carriers in hepatitis C virus infection. Liver Int. 2019, 39, 1237–1245. [Google Scholar] [CrossRef]
- Sawai, H.; Nishida, N.; Khor, S.S.; Honda, M.; Sugiyama, M.; Baba, N.; Yamada, K.; Sawada, N.; Tsugane, S.; Koike, K.; et al. Genome-wide association study identified new susceptible genetic variants in HLA class I region for hepatitis B virus-related hepatocellular carcinoma. Sci Rep. 2018, 8, 7958. [Google Scholar] [CrossRef]
- Matsuura, K.; Tanaka, Y. Host genetic variants influencing the clinical course of hepatitis C virus infection. J. Med. Virol. 2016, 88, 185–195. [Google Scholar] [CrossRef]
- Tseng, K.C.; Tseng, C.W.; Hsieh, Y.H.; Chang, C.K.; Lai, N.S.; Hung, T.H.; Chang, T.T. Effect of human leukocyte antigen class I and II alleles on hepatitis C viral load among chronic hepatitis C patients in Southern Taiwan. Hum. Immunol. 2013, 74, 978–982. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Kaul, R.; Kaul, A.; Khan, K. A comparative review of HLA associations with hepatitis B and C viral infections across global populations. World J. Gastroenterol. 2007, 13, 1770–1787. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Golubovic, G.; Stajic, M.; Stolic, I.; Nikolic, J.A.; Neskovic, A.N.; Pandey, L. Histocompatibility antigens in patients with hepatocellular carcinoma. Z Gastroenterol. 1996, 34, 15–20. [Google Scholar] [PubMed]
- Hofmann, M.; Tauber, C.; Hensel, N.; Thimme, R. CD8+ T Cell Responses during HCV Infection and HCC. J. Clinical Med. 2021, 10, 991. [Google Scholar] [CrossRef]
- Khakoo, S.I.; Thio, C.L.; Martin, M.P.; Brooks, C.R.; Gao, X.; Astemborski, J.; Cheng, J.; Goedert, J.J.; Vlahov, D.; Hilgartner, M.; et al. HLA and NK cell inhibitory receptor genes in resolving hepatitis C virus infection. Science 2004, 305, 872–874. [Google Scholar] [CrossRef]
- Knapp, S.; Warshow, U.; Hegazy, D.; Brackenbury, L.; Guha, I.N.; Fowell, A.; Little, A.M.; Alexander, G.J.; Rosenberg, W.M.; Cramp, M.E.; et al. Consistent beneficial effects of killer cell immunoglobulin-like receptor 2DL3 and group 1 human leukocyte antigen-C following exposure to hepatitis C virus. Hepatology 2010, 51, 1168–1175. [Google Scholar] [CrossRef]
- Vidal-Castineira, J.R.; Lopez-Vazquez, A.; Diaz-Pena, R.; Alonso-Arias, R.; Martinez-Borra, J.; Perez, R.; Fernandez-Suarez, J.; Melon, S.; Prieto, J.; Rodrigo, L.; et al. Effect of killer immunoglobulin-like receptors in the response to combined treatment in patients with chronic hepatitis C virus infection. J. Virol. 2010, 84, 475–481. [Google Scholar] [CrossRef] [Green Version]
- Suppiah, V.; Gaudieri, S.; Armstrong, N.J.; O’Connor, K.S.; Berg, T.; Weltman, M.; Abate, M.L.; Spengler, U.; Bassendine, M.; Dore, G.J.; et al. IL28B, HLA-C, and KIR variants additively predict response to therapy in chronic hepatitis C virus infection in a European Cohort: A cross-sectional study. PLoS Med. 2011, 8, e1001092. [Google Scholar] [CrossRef]
- Dring, M.M.; Morrison, M.H.; McSharry, B.P.; Guinan, K.J.; Hagan, R.; O’Farrelly, C.; Gardiner, C.M. Innate immune genes synergize to predict increased risk of chronic disease in hepatitis C virus infection. Proc. Natl. Acad. Sci. USA 2011, 108, 5736–5741. [Google Scholar] [CrossRef] [Green Version]
- Nozawa, Y.; Umemura, T.; Joshita, S.; Katsuyama, Y.; Shibata, S.; Kimura, T.; Morita, S.; Komatsu, M.; Matsumoto, A.; Tanaka, E.; et al. KIR, HLA, and IL28B Variant Predict Response to Antiviral Therapy in Genotype 1 Chronic Hepatitis C Patients in Japan. PLoS ONE 2013, 8, e83381. [Google Scholar] [CrossRef] [Green Version]
- Fitzmaurice, K.; Hurst, J.; Dring, M.; Rauch, A.; McLaren, P.J.; Gunthard, H.F.; Gardiner, C.; Klenerman, P.; Irish HCV Research Consortium; Swiss HIV Cohort Study. Additive effects of HLA alleles and innate immune genes determine viral outcome in HCV infection. Gut 2015, 64, 813–819. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thons, C.; Senff, T.; Hydes, T.J.; Manser, A.R.; Heinemann, F.M.; Heinold, A.; Heilmann, M.; Kim, A.Y.; Uhrberg, M.; Scherbaum, N.; et al. HLA-Bw4 80(T) and multiple HLA-Bw4 copies combined with KIR3DL1 associate with spontaneous clearance of HCV infection in people who inject drugs. J. Hepatol. 2017, 67, 462–470. [Google Scholar] [CrossRef] [Green Version]
- Umemura, T.; Joshita, S.; Saito, H.; Yoshizawa, K.; Norman, G.L.; Tanaka, E.; Ota, M. KIR/HLA genotypes confer susceptibility and progression in patients with autoimmune hepatitis. JHEP Rep. 2019, 1, 353–360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sugiura, A.; Joshita, S.; Umemura, T.; Yamazaki, T.; Fujimori, N.; Kimura, T.; Matsumoto, A.; Igarashi, K.; Usami, Y.; Wada, S.; et al. Past history of hepatocellular carcinoma is an independent risk factor of treatment failure in patients with chronic hepatitis C virus infection receiving direct-acting antivirals. J. Viral Hepat. 2018, 25, 1462–1471. [Google Scholar] [CrossRef] [PubMed]
- Sugiura, A.; Joshita, S.; Yamashita, Y.; Yamazaki, T.; Fujimori, N.; Kimura, T.; Matsumoto, A.; Wada, S.; Mori, H.; Shibata, S.; et al. Effectiveness of Glecaprevir/Pibrentasvir for Hepatitis C: Real-World Experience and Clinical Features of Retreatment Cases. Biomedicines 2020, 8, 74. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iio, E.; Matsuura, K.; Shimada, N.; Atsukawa, M.; Itokawa, N.; Abe, H.; Kato, K.; Takaguchi, K.; Senoh, T.; Eguchi, Y.; et al. TLL1 variant associated with development of hepatocellular carcinoma after eradication of hepatitis C virus by interferon-free therapy. J. Gastroenterol. 2019, 54, 339–346. [Google Scholar] [CrossRef]
- Miki, D.; Akita, T.; Kurisu, A.; Kawaoka, T.; Nakajima, T.; Hige, S.; Karino, Y.; Toyoda, H.; Kumada, T.; Tsuge, M.; et al. PNPLA3 and HLA-DQB1 polymorphisms are associated with hepatocellular carcinoma after hepatitis C virus eradication. J. Gastroenterol. 2020, 55, 1162–1170. [Google Scholar] [CrossRef] [PubMed]
- Gardiner, C.M.; Guethlein, L.A.; Shilling, H.G.; Pando, M.; Carr, W.H.; Rajalingam, R.; Vilches, C.; Parham, P. Different NK cell surface phenotypes defined by the DX9 antibody are due to KIR3DL1 gene polymorphism. J. Immunol. 2001, 166, 2992–3001. [Google Scholar] [CrossRef] [Green Version]
- Yawata, M.; Yawata, N.; Draghi, M.; Little, A.M.; Partheniou, F.; Parham, P. Roles for HLA and KIR polymorphisms in natural killer cell repertoire selection and modulation of effector function. J. Exp. Med. 2006, 203, 633–645. [Google Scholar] [CrossRef] [Green Version]
- Trundley, A.; Frebel, H.; Jones, D.; Chang, C.; Trowsdale, J. Allelic expression patterns of KIR3DS1 and 3DL1 using the Z27 and DX9 antibodies. Eur. J. Immunol. 2007, 37, 780–787. [Google Scholar] [CrossRef]
- Tajik, N.; Shahsavar, F.; Nasiri, M.; Radjabzadeh, M.F. Compound KIR-HLA genotype analyses in the Iranian population by a novel PCR-SSP assay. Int. J. Immunogenet. 2010, 37, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Vilches, C.; Castano, J.; Gomez-Lozano, N.; Estefania, E. Facilitation of KIR genotyping by a PCR-SSP method that amplifies short DNA fragments. Tissue Antigens 2007, 70, 415–422. [Google Scholar] [CrossRef] [PubMed]
- Cooley, S.; Trachtenberg, E.; Bergemann, T.L.; Saeteurn, K.; Klein, J.; Le, C.T.; Marsh, S.G.; Guethlein, L.A.; Parham, P.; Miller, J.S.; et al. Donors with group B KIR haplotypes improve relapse-free survival after unrelated hematopoietic cell transplantation for acute myelogenous leukemia. Blood 2009, 113, 726–732. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Characteristic | Total (n = 211) | Patients with Hepatocellular Carcinoma (HCC) (n = 147) | Patients without HCC (n = 64) | p-Value |
|---|---|---|---|---|
| Age, y | 64 (46–78) | 63 (46–77) | 65 (45–80) | 0.110 |
| Male, n (%) | 110 (52.1) | 84 (57.1) | 26 (40.6) | 0.027 |
| Body Mass Index | 22.3 (17.6–29.8) | 22.2 (17.7–29.7) | 22.6 (17.2–33.5) | 0.417 |
| Albumin, g/dL | 3.8 (3.0–4.5) | 3.8 (2.9–4.5) | 4.1 (3.1–4.5) | 0.009 |
| Bilirubin, mg/dL | 0.9 (0.5–1.8) | 0.9 (0.5–1.8) | 0.9 (0.4–1.9) | 0.532 |
| ALT, IU/L | 55 (20–200) | 56 (21–209) | 54 (17–201) | 0.177 |
| Platelet count, ×109/L | 10.6 (4.8–21.6) | 10.2 (4.8–21.1) | 12.0 (5.0–22.4) | 0.033 |
| Prothrombin time % | 84.7 (62.3–108.0) | 84.7 (58.4–104.9) | 84.8 (67.9–113.1) | 0.031 |
| α-fetoprotein, ng/mL | 6.9 (1.5–41.8) | 9.2 (1.8–44.3) | 3.7 (1.2–30.8) | <0.001 |
| Des-gamma-carboxy prothrombin, mAU/mL | 18.0 (9.5–40.0) | 19.0 (9.2–40.9) | 13.0 (8.2–37.6) | 0.015 |
| HCV RNA, log IU/mL | 6.5 (3.4–7.3) | 6.5 (3.3–7.3) | 6.4 (4.0–7.4) | 0.838 |
| HCV genotype 1, n (%) | 180 (85.3) | 125 (85.0) | 55 (85.9) | 0.865 |
| Interferon treatment, n (%) | 94 (44.5) | 67 (45.6) | 27 (42.2) | 0.649 |
| Genetic Factor | Patients with HCC (n = 147) | Patients without HCC (n = 64) | Odds Ratio | 95% Confidence Interval | p-Value |
|---|---|---|---|---|---|
| HLA-Bw4 | 116 (78.9) | 41 (64.1) | 2.10 | 1.10–4.01 | 0.023 |
| HLA-Bw4-80I | 97 (66.0) | 33 (51.6) | 1.82 | 1.00–3.31 | 0.048 |
| HLA-Bw4-80T | 31 (21.1) | 12 (18.8) | 1.16 | 0.55–2.43 | 0.698 |
| HLA-Bw6 | 132 (89.8) | 56 (87.5) | 1.26 | 0.50–3.13 | 0.623 |
| HLA-C1 | 145 (98.6) | 64 (100.0) | - | - | 0.869 |
| HLA-C2 | 19 (12.9) | 8 (12.5) | 1.04 | 0.43–2.51 | 0.932 |
| KIR2DL1 | 147 (100.0) | 64 (100.0) | - | - | - |
| KIR2DL2 | 25 (17.0) | 7 (10.9) | 1.67 | 0.68–4.08 | 0.259 |
| KIR2DL3 | 147 (100.0) | 64 (100.0) | - | - | - |
| KIR2DL4 | 147 (100.0) | 64 (100.0) | - | - | - |
| KIR2DL5 | 58 (39.5) | 29 (45.3) | 0.79 | 0.43–1.42 | 0.427 |
| KIR2DS1 | 61 (41.5) | 28 (43.8) | 0.91 | 0.50–1.65 | 0.761 |
| KIR2DS2 | 23 (15.6) | 8 (12.5) | 1.30 | 0.55–3.08 | 0.553 |
| KIR2DS3 | 20 (13.6) | 12 (18.8) | 0.68 | 0.31–1.50 | 0.338 |
| KIR2DS4 | 138 (93.9) | 58 (90.6) | 1.59 | 0.54–4.66 | 0.398 |
| KIR2DS5 | 41 (27.9) | 19 (29.7) | 0.92 | 0.48–1.75 | 0.790 |
| KIR3DL1 | 138 (93.9) | 61 (95.3) | 0.75 | 0.20–2.88 | 0.679 |
| KIR3DL2 | 147 (100.0) | 64 (100.0) | - | - | - |
| KIR3DL3 | 147 (100.0) | 64 (100.0) | - | - | - |
| KIR3DS1 | 60 (40.8) | 29 (45.3) | 0.83 | 0.46–1.50 | 0.543 |
| Genotype A/A | 72 (49.0) | 27 (42.2) | 1.32 | 0.73–2.38 | 0.363 |
| Genotype B/x | 75 (51.0) | 37 (57.8) | 0.76 | 0.42–1.37 | 0.363 |
| Genetic Factor | Patients with HCC (n = 147) | Patients without HCC (n = 64) | Odds Ratio | 95% Confidence Interval | p-Value |
|---|---|---|---|---|---|
| KIR2DL1 + HLA-C2 | 19 (12.9) | 8 (12.5) | 1.04 | 0.43–2.51 | 0.932 |
| KIR2DS1 + HLA-C2 | 9 (6.1) | 5 (7.8) | 0.77 | 0.25–2.39 | 0.650 |
| KIR2DL2 + HLA-C1 | 24 (16.3) | 7 (10.9) | 1.59 | 0.65–3.90 | 0.309 |
| KIR2DL3 + HLA-C1 | 145 (98.6) | 64 (100.0) | - | - | 0.869 |
| KIR2DS2 + HLA-C1 | 22 (15.0) | 8 (12.5) | 1.23 | 0.52–2.94 | 0.637 |
| KIR3DL1 + HLA-Bw4 | 112 (76.2) | 39 (60.9) | 2.05 | 1.09–3.85 | 0.024 |
| KIR3DL1 + HLA-Bw4-80I | 94 (63.9) | 31 (48.4) | 1.89 | 1.04–3.42 | 0.035 |
| KIR3DL1 + HLA-Bw4-80T | 30 (20.4) | 12 (18.8) | 1.11 | 0.53–2.34 | 0.782 |
| Factor | HR | 95% CI | p-Value |
|---|---|---|---|
| Sex, female vs. male | 1.56 | 1.12–2.17 | 0.009 |
| AFP (ng/mL), ≤5.6 vs. > 5.6 | 1.56 | 1.10–2.20 | 0.011 |
| KIR3DL1 + HLA-Bw4, negative vs. positive | 1.69 | 1.15–2.48 | 0.007 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Umemura, T.; Joshita, S.; Saito, H.; Wakabayashi, S.-i.; Kobayashi, H.; Yamashita, Y.; Sugiura, A.; Yamazaki, T.; Ota, M. Investigation of the Effect of KIR–HLA Pairs on Hepatocellular Carcinoma in Hepatitis C Virus Cirrhotic Patients. Cancers 2021, 13, 3267. https://doi.org/10.3390/cancers13133267
Umemura T, Joshita S, Saito H, Wakabayashi S-i, Kobayashi H, Yamashita Y, Sugiura A, Yamazaki T, Ota M. Investigation of the Effect of KIR–HLA Pairs on Hepatocellular Carcinoma in Hepatitis C Virus Cirrhotic Patients. Cancers. 2021; 13(13):3267. https://doi.org/10.3390/cancers13133267
Chicago/Turabian StyleUmemura, Takeji, Satoru Joshita, Hiromi Saito, Shun-ichi Wakabayashi, Hiroyuki Kobayashi, Yuki Yamashita, Ayumi Sugiura, Tomoo Yamazaki, and Masao Ota. 2021. "Investigation of the Effect of KIR–HLA Pairs on Hepatocellular Carcinoma in Hepatitis C Virus Cirrhotic Patients" Cancers 13, no. 13: 3267. https://doi.org/10.3390/cancers13133267
APA StyleUmemura, T., Joshita, S., Saito, H., Wakabayashi, S.-i., Kobayashi, H., Yamashita, Y., Sugiura, A., Yamazaki, T., & Ota, M. (2021). Investigation of the Effect of KIR–HLA Pairs on Hepatocellular Carcinoma in Hepatitis C Virus Cirrhotic Patients. Cancers, 13(13), 3267. https://doi.org/10.3390/cancers13133267






