Loss of CENP-I Impairs Homologous Recombination and Sensitizes Cells to PARP1 Inhibition
Abstract
Simple Summary
Abstract
1. Introduction
2. Results
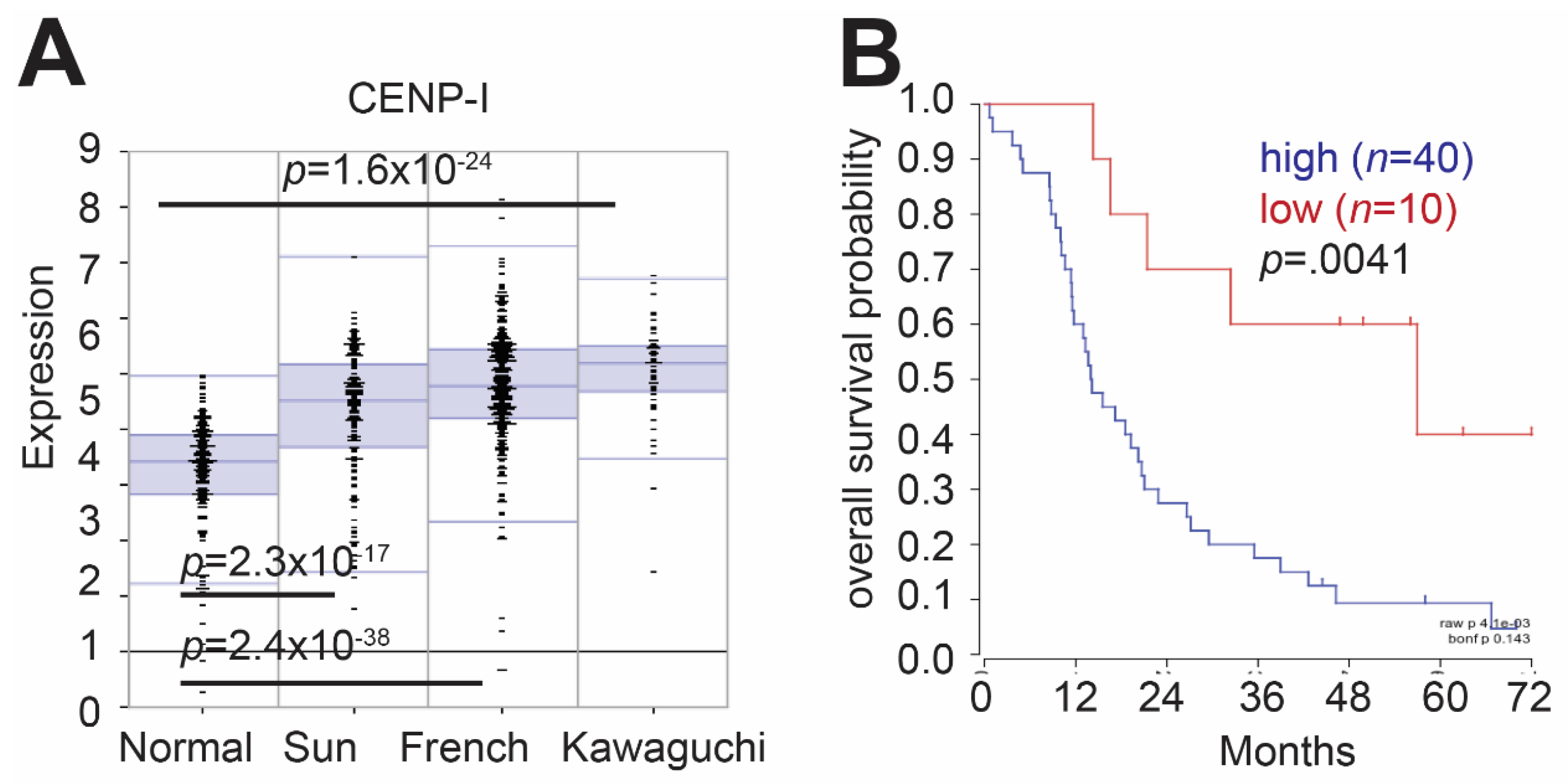
2.1. CENP-I Expression Is Elevated Gliomas as Compared to Normal Brain
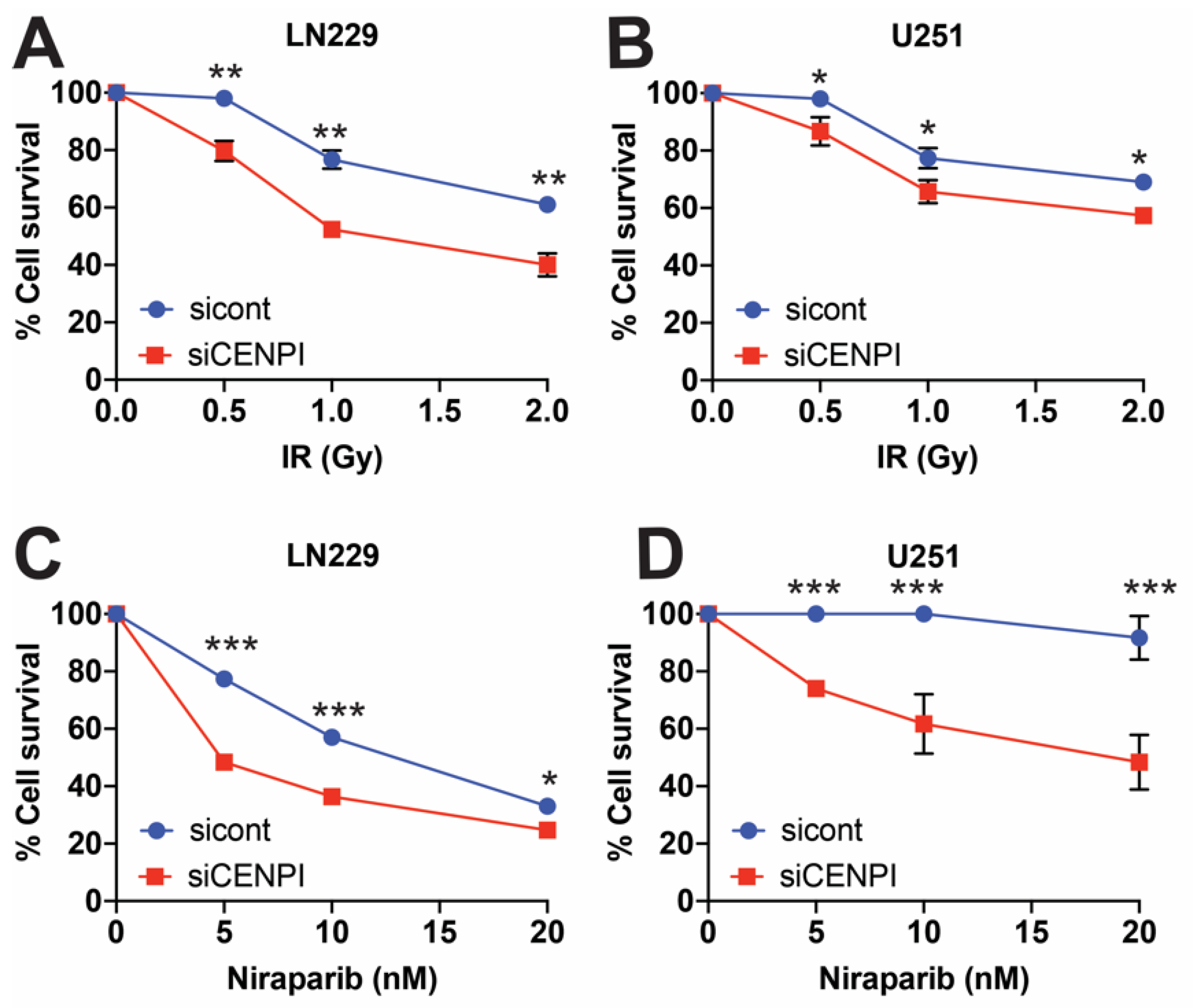
2.2. Loss of CENP-I Sensitizes Cells to Genotoxic Stress
2.3. Loss of CENP-I Results in Increased 53BP1 Foci and Delayed Repair Kinetics
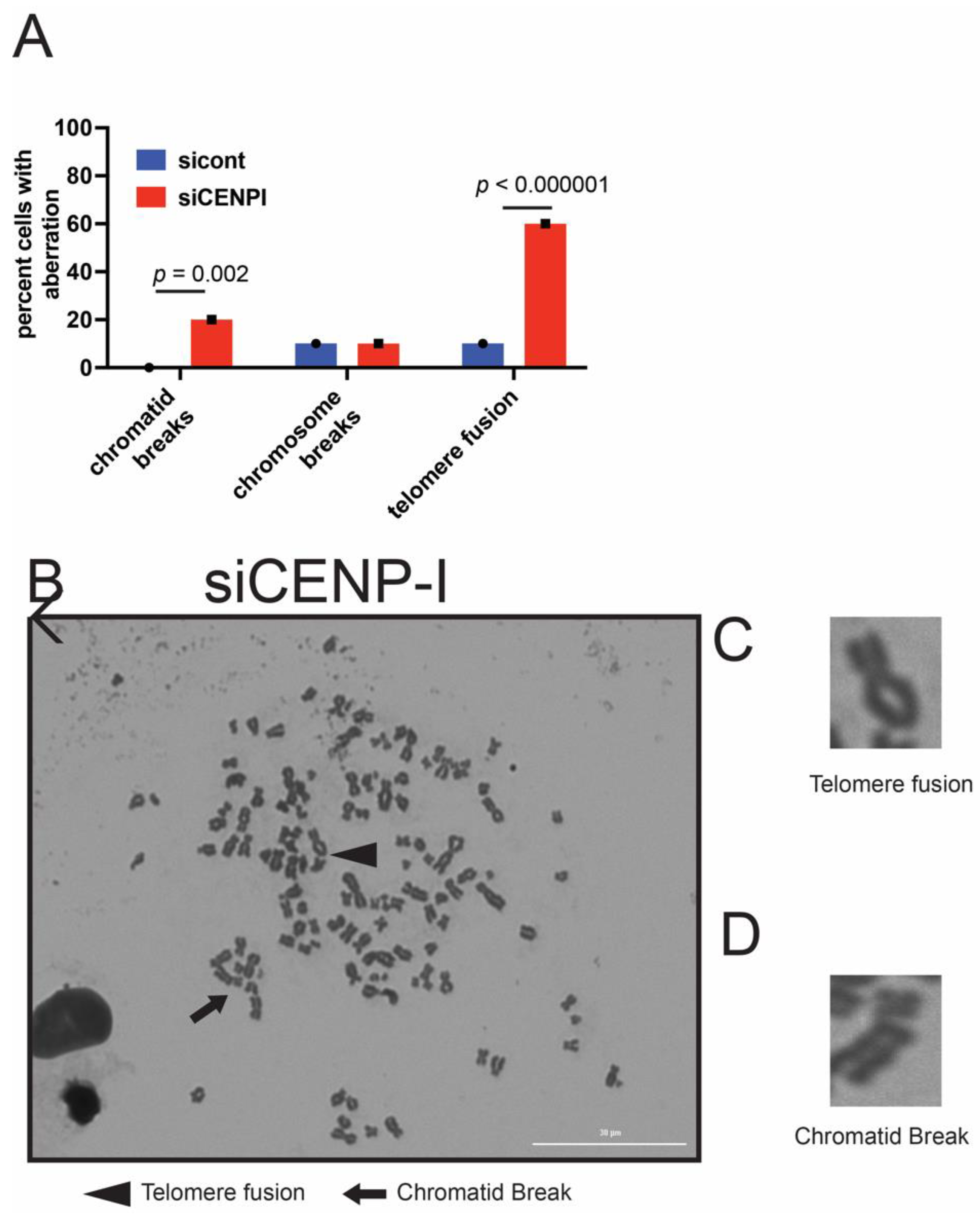
2.4. Loss of CENP-I Leads to Increased Chromosomal Aberrations
2.5. Loss of CENP-I Inhibits HR, But Is Dispensable for NHEJ DSB Repair
2.6. Loss of CENP-I Results in Increased R-Loop Formation
3. Discussion
3.1. Involvement of CENP-I and Homologous Recombination Repair
3.2. Therapeutic Potential of CENP-I Targeting
4. Materials and Methods
- siCENPI-01: CAAACCAUUUCGUGUGAGA
- siCENPI-02: CCAUGAACUCUGUGUCUAA
- siCENPI-03: GAAUUAAAUCACUAUUGA
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bakhoum, S.F.; Cantley, L.C. The Multifaceted Role of Chromosomal Instability in Cancer and Its Microenvironment. Cell 2018, 174, 1347–1360. [Google Scholar] [CrossRef] [PubMed]
- Sinclair, W.K.; Morton, R.A. X-ray sensitivity during the cell generation cycle of cultured Chinese hamster cells. Radiat. Res. 1966, 29, 450. [Google Scholar] [CrossRef] [PubMed]
- Burrell, R.A.; McClelland, S.E.; Endesfelder, D.; Groth, P.; Weller, M.-C.; Shaikh, N.; Domingo, E.; Kanu, N.; Dewhurst, S.M.; Gronroos, E.; et al. Replication stress links structural and numerical cancer chromosomal instability. Nat. Cell Biol. 2013, 494, 492–496. [Google Scholar] [CrossRef]
- Deckbar, D.; Birraux, J.; Krempler, A.; Tchouandong, L.; Beucher, A.; Walker, S.; Stiff, T.; Jeggo, P.; Löbrich, M. Chromosome breakage after G2 checkpoint release. J. Cell Biol. 2007, 176, 749–755. [Google Scholar] [CrossRef]
- Cann, K.L.; Dellaire, G. Heterochromatin and the DNA damage response: The need to relaxThis paper is one of a selection of papers in a Special Issue entitled 31st Annual International Asilomar Chromatin and Chromosomes Conference, and has undergone the Journal’s usual peer review process. Biochem. Cell Biol. 2011, 89, 45–60. [Google Scholar] [CrossRef]
- Ceccaldi, R.; Rondinelli, B.; D’Andrea, A.D. Repair Pathway Choices and Consequences at the Double-Strand Break. Trends Cell Biol. 2016, 26, 52–64. [Google Scholar] [CrossRef]
- Giunta, S.; Belotserkovskaya, R.; Jackson, S.P. DNA damage signaling in response to double-strand breaks during mitosis. J. Cell Biol. 2010, 190, 197–207. [Google Scholar] [CrossRef]
- Bakhoum, S.F.; Kabeche, L.; Compton, D.A.; Powell, S.N.; Bastians, H. Mitotic DNA Damage Response: At the Crossroads of Structural and Numerical Cancer Chromosome Instabilities. Trends Cancer 2017, 3, 225–234. [Google Scholar] [CrossRef] [PubMed]
- Bakhoum, S.F.; Kabeche, L.; Murnane, J.P.; Zaki, B.I.; Compton, D.A. DNA-Damage Response during Mitosis Induces Whole-Chromosome Missegregation. Cancer Discov. 2014, 4, 1281–1289. [Google Scholar] [CrossRef]
- Cheeseman, I.M.; Hori, T.; Fukagawa, T.; Desai, A. KNL1 and the CENP-H/I/K Complex Coordinately Direct Kinetochore Assembly in Vertebrates. Mol. Biol. Cell 2008, 19, 587–594. [Google Scholar] [CrossRef] [PubMed]
- Kabeche, L.; Nguyen, H.D.; Buisson, R.; Zou, L. A mitosis-specific and R loop–driven ATR pathway promotes faithful chromosome segregation. Science 2017, 359, 108–114. [Google Scholar] [CrossRef]
- Zeitlin, S.G.; Baker, N.M.; Chapados, B.R.; Soutoglou, E.; Wang, J.Y.J.; Berns, M.W.; Cleveland, D.W. Double-strand DNA breaks recruit the centromeric histone CENP-A. Proc. Natl. Acad. Sci. USA 2009, 106, 15762–15767. [Google Scholar] [CrossRef]
- Helfricht, A.; Wiegant, W.W.; Thijssen, P.E.; Vertegaal, A.; Luijsterburg, M.S.; Van Attikum, H. Remodeling and spacing factor 1 (RSF1) deposits centromere proteins at DNA double-strand breaks to promote non-homologous end-joining. Cell Cycle 2013, 12, 3070–3082. [Google Scholar] [CrossRef]
- Giunta, S.; Hervé, S.; White, R.R.; Wilhelm, T.; Dumont, M.; Scelfo, A.; Gamba, R.; Wong, C.K.; Rancati, G.; Smogorzewska, A.; et al. CENP-A chromatin prevents replication stress at centromeres to avoid structural aneuploidy. Proc. Natl. Acad. Sci. USA 2021, 118. [Google Scholar] [CrossRef] [PubMed]
- Kawaguchi, A.; Yajima, N.; Tsuchiya, N.; Homma, J.; Sano, M.; Natsumeda, M.; Takahashi, H.; Fujii, Y.; Kakuma, T.; Yamanaka, R. Gene expression signature-based prognostic risk score in patients with glioblastoma. Cancer Sci. 2013, 104, 1205–1210. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Hui, A.-M.; Su, Q.; Vortmeyer, A.; Kotliarov, Y.; Pastorino, S.; Passaniti, A.; Menon, J.; Walling, J.; Bailey, R.; et al. Neuronal and glioma-derived stem cell factor induces angiogenesis within the brain. Cancer Cell 2006, 9, 287–300. [Google Scholar] [CrossRef]
- Gravendeel, L.A.M.; Kouwenhoven, M.; Gevaert, O.; De Rooi, J.J.; Stubbs, A.P.; Duijm, J.E.; Daemen, A.; Bleeker, F.E.; Bralten, L.B.C.; Kloosterhof, N.K.; et al. Intrinsic Gene Expression Profiles of Gliomas Are a Better Predictor of Survival than Histology. Cancer Res. 2009, 69, 9065–9072. [Google Scholar] [CrossRef]
- Jones, P.; Altamura, S.; Boueres, J.; Ferrigno, F.; Fonsi, M.; Giomini, C.; Lamartina, S.; Monteagudo, E.; Ontoria, J.M.; Orsale, M.V.; et al. Discovery of 2-{4-[(3S)-Piperidin-3-yl]phenyl}-2H-indazole-7-carboxamide (MK-4827): A Novel Oral Poly(ADP-ribose)polymerase (PARP) Inhibitor Efficacious in BRCA-1 and -2 Mutant Tumors. J. Med. Chem. 2009, 52, 7170–7185. [Google Scholar] [CrossRef]
- Li, H.-L.; Wang, C.-Y.; Fu, J.; Yang, X.-J.; Sun, Y.; Shao, Y.-H.; Zhang, L.-H.; Yang, X.-M.; Zhang, X.-L.; Lin, J. PTEN expression in U251 glioma cells enhances their sensititivy to ionizing radiation by suppressing DNA repair capacity. Eur. Rev. Med. Pharmacol. Sci. 2019, 23. [Google Scholar]
- McCormick, A.; Donoghue, P.; Dixon, M.; O’Sullivan, R.; O’Donnell, R.L.; Murray, J.; Kaufmann, A.; Curtin, N.J.; Edmondson, R.J. Ovarian Cancers Harbor Defects in Nonhomologous End Joining Resulting in Resistance to Rucaparib. Clin. Cancer Res. 2016, 23, 2050–2060. [Google Scholar] [CrossRef] [PubMed]
- Morales, J.C.; Xia, Z.; Lu, T.; Aldrich, M.B.; Wang, B.; Rosales, C.; Kellems, R.E.; Hittelman, W.N.; Elledge, S.J.; Carpenter, P.B. Role for the BRCA1 C-terminal Repeats (BRCT) Protein 53BP1 in Maintaining Genomic Stability. J. Biol. Chem. 2003, 278, 14971–14977. [Google Scholar] [CrossRef] [PubMed]
- Bassing, C.H.; Chua, K.F.; Sekiguchi, J.; Suh, H.; Whitlow, S.R.; Fleming, J.C.; Monroe, B.C.; Ciccone, D.N.; Yan, C.; Vlasakova, K.; et al. Increased ionizing radiation sensitivity and genomic instability in the absence of histone H2AX. Proc. Natl. Acad. Sci. USA 2002, 99, 8173–8178. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Jin, S.; Gao, Y.; Weaver, D.T.; Alt, F.W. Ku70-deficient embryonic stem cells have increased ionizing radiosensitivity, defective DNA end-binding activity, and inability to support V(D)J recombination. Proc. Natl. Acad. Sci. USA 1997, 94, 8076–8081. [Google Scholar] [CrossRef] [PubMed]
- Morales, M.; Theunissen, J.-W.F.; Kim, C.F.B.; Kitagawa, R.; Kastan, M.B.; Petrini, J.H. The Rad50S allele promotes ATM-dependent DNA damage responses and suppresses ATM deficiency: Implications for the Mre11 complex as a DNA damage sensor. Genes Dev. 2005, 19, 3043–3054. [Google Scholar] [CrossRef]
- Bryant, H.E.; Schultz, N.; Thomas, H.D.; Parker, K.M.; Flower, D.; Lopez, E.; Kyle, S.; Meuth, M.; Curtin, N.J.; Helleday, T. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 2005, 434, 913–917. [Google Scholar] [CrossRef]
- Popp, H.D.; Brendel, S.; Hofmann, W.-K.; Fabarius, A. Immunofluorescence Microscopy of γH2AX and 53BP1 for Analyzing the Formation and Repair of DNA Double-strand Breaks. J. Vis. Exp. 2017, e56617. [Google Scholar] [CrossRef]
- Sakaue-Sawano, A.; Kurokawa, H.; Morimura, T.; Hanyu, A.; Hama, H.; Osawa, H.; Kashiwagi, S.; Fukami, K.; Miyata, T.; Miyoshi, H.; et al. Visualizing Spatiotemporal Dynamics of Multicellular Cell-Cycle Progression. Cell 2008, 132, 487–498. [Google Scholar] [CrossRef]
- Gunn, A.; Stark, J.M. I-SceI-Based Assays to Examine Distinct Repair Outcomes of Mammalian Chromosomal Double Strand Breaks. Meth. Molec. Biol. 2012, 920, 379–391. [Google Scholar] [CrossRef]
- Morales, J.C.; Richard, P.; Rommel, A.; Fattah, F.J.; Motea, E.A.; Patidar, P.L.; Xiao, L.; Leskov, K.; Wu, S.-Y.; Hittelman, W.N.; et al. Kub5-Hera, the human Rtt103 homolog, plays dual functional roles in transcription termination and DNA repair. Nucleic Acids Res. 2014, 42, 4996–5006. [Google Scholar] [CrossRef] [PubMed]
- Dang, T.T.; Morales, J.C. XRN2 Links RNA:DNA Hybrid Resolution to Double Strand Break Repair Pathway Choice. Cancers 2020, 12, 1821. [Google Scholar] [CrossRef]
- Morales, J.C.; Richard, P.; Patidar, P.L.; Motea, E.A.; Dang, T.T.; Manley, J.; Boothman, D.A. XRN2 Links Transcription Termination to DNA Damage and Replication Stress. PLoS Genet. 2016, 12, e1006107. [Google Scholar] [CrossRef]
- Smolka, J.A.; Sanz, L.A.; Hartono, S.R.; Chédin, F. Recognition of RNA by the S9.6 antibody creates pervasive artifacts when imaging RNA:DNA hybrids. J. Cell Biol. 2021, 220. [Google Scholar] [CrossRef]
- Ziv, Y.; Bar-Shira, A.; Pecker, I.; Russell, P.; Jorgensen, T.J.; Tsarfaty, I.; Shiloh, Y. Recombinant ATM protein complements the cellular A-T phenotype. Oncogene 1997, 15, 159–167. [Google Scholar] [CrossRef] [PubMed]
- Gil Del Alcazar, C.R.; Todorova, P.K.; Habib, A.A.; Mukherjee, B.; Burma, S. Augmented HR Repair Mediates Acquired Temozolomide Resistance in Glioblastoma. Mol. Cancer Res. 2016, 14, 928–940. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.-B.S.; Elledge, S.J. The DNA damage response: Putting checkpoints in perspective. Nat. Cell Biol. 2000, 408, 433–439. [Google Scholar] [CrossRef] [PubMed]
- Ohle, C.; Tesorero, R.; Schermann, G.; Dobrev, N.; Sinning, I.; Fischer, T. Transient RNA-DNA Hybrids Are Required for Efficient Double-Strand Break Repair. Cell 2016, 167, 1001–1013. [Google Scholar] [CrossRef]
- Shono, N.; Ohzeki, J.-I.; Otake, K.; Martins, N.; Nagase, T.; Kimura, H.; Larionov, V.; Earnshaw, W.C.; Masumoto, H. CENP-C and CENP-I are key connecting factors for kinetochore and CENP-A assembly. J. Cell Sci. 2015, 128, 4572–4587. [Google Scholar] [CrossRef]
- Tykocki, T.; Eltayeb, M. Ten-year survival in glioblastoma. A systematic review. J. Clin. Neurosci. 2018, 54, 7–13. [Google Scholar] [CrossRef]
- Batash, R.; Asna, N.; Schaffer, P.; Francis, N.; Schaffer, M. Glioblastoma Multiforme, Diagnosis and Treatment; Recent Literature Review. Curr. Med. Chem. 2017, 24, 3002–3009. [Google Scholar] [CrossRef]
- Roy, S.; Lahiri, D.; Maji, T.; Biswas, J. Recurrent Glioblastoma: Where we stand. South. Asian J. Cancer 2015, 4, 163. [Google Scholar] [CrossRef]
- Wen, P.Y.; Kesari, S. Malignant Gliomas in Adults. N Engl. J. Med. 2008, 359, 492–507. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, Y.P.; Weatherbee, J.L.; Wheelhouse, R.T.; Ross, A.H. Glioblastoma Multiforme Therapy and Mechanisms of Resistance. Pharm. 2013, 6, 1475–1506. [Google Scholar] [CrossRef] [PubMed]
- Erasimus, H.; Gobin, M.; Niclou, S.; Van Dyck, E. DNA repair mechanisms and their clinical impact in glioblastoma. Mutat. Res. Mutat. Res. 2016, 769, 19–35. [Google Scholar] [CrossRef] [PubMed]
- Aguilera, A.; Garcia-Muse, T. R Loops: From Transcription Byproducts to Threats to Genome Stability. Mol. Cell 2012, 46, 115–124. [Google Scholar] [CrossRef] [PubMed]
- Gunn, A.; Stark, J.M. I-SceI-Based Assays to Examine Distinct Repair Outcomes of Mammalian Chromosomal Double Strand Breaks. In DNA Repair Protocols; Bjergbæk, L., Ed.; Humana Press: Totowa, NJ, USA, 2012; pp. 379–391. [Google Scholar]
- Sughrue, M.E.; Rutkowski, M.J.; Kane, A.J.; Parsa, A.T. Human glioma demonstrates cell line specific results with ATP-based chemiluminescent cellular proliferation assays. J. Clin. Neurosci. 2010, 17, 1573–1577. [Google Scholar] [CrossRef][Green Version]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dang, T.T.; Morales, J.C. Loss of CENP-I Impairs Homologous Recombination and Sensitizes Cells to PARP1 Inhibition. Cancers 2021, 13, 3202. https://doi.org/10.3390/cancers13133202
Dang TT, Morales JC. Loss of CENP-I Impairs Homologous Recombination and Sensitizes Cells to PARP1 Inhibition. Cancers. 2021; 13(13):3202. https://doi.org/10.3390/cancers13133202
Chicago/Turabian StyleDang, Tuyen T., and Julio C. Morales. 2021. "Loss of CENP-I Impairs Homologous Recombination and Sensitizes Cells to PARP1 Inhibition" Cancers 13, no. 13: 3202. https://doi.org/10.3390/cancers13133202
APA StyleDang, T. T., & Morales, J. C. (2021). Loss of CENP-I Impairs Homologous Recombination and Sensitizes Cells to PARP1 Inhibition. Cancers, 13(13), 3202. https://doi.org/10.3390/cancers13133202






