The Identification of RNA-Binding Proteins Functionally Associated with Tumor Progression in Gastrointestinal Cancer
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. siRNA and Transfection
2.3. SRB Assays
2.4. MTT Assays
2.5. Western Blotting
2.6. cDNA Analyses
2.7. Data Corrections
2.8. Statistical Assessments
3. Results
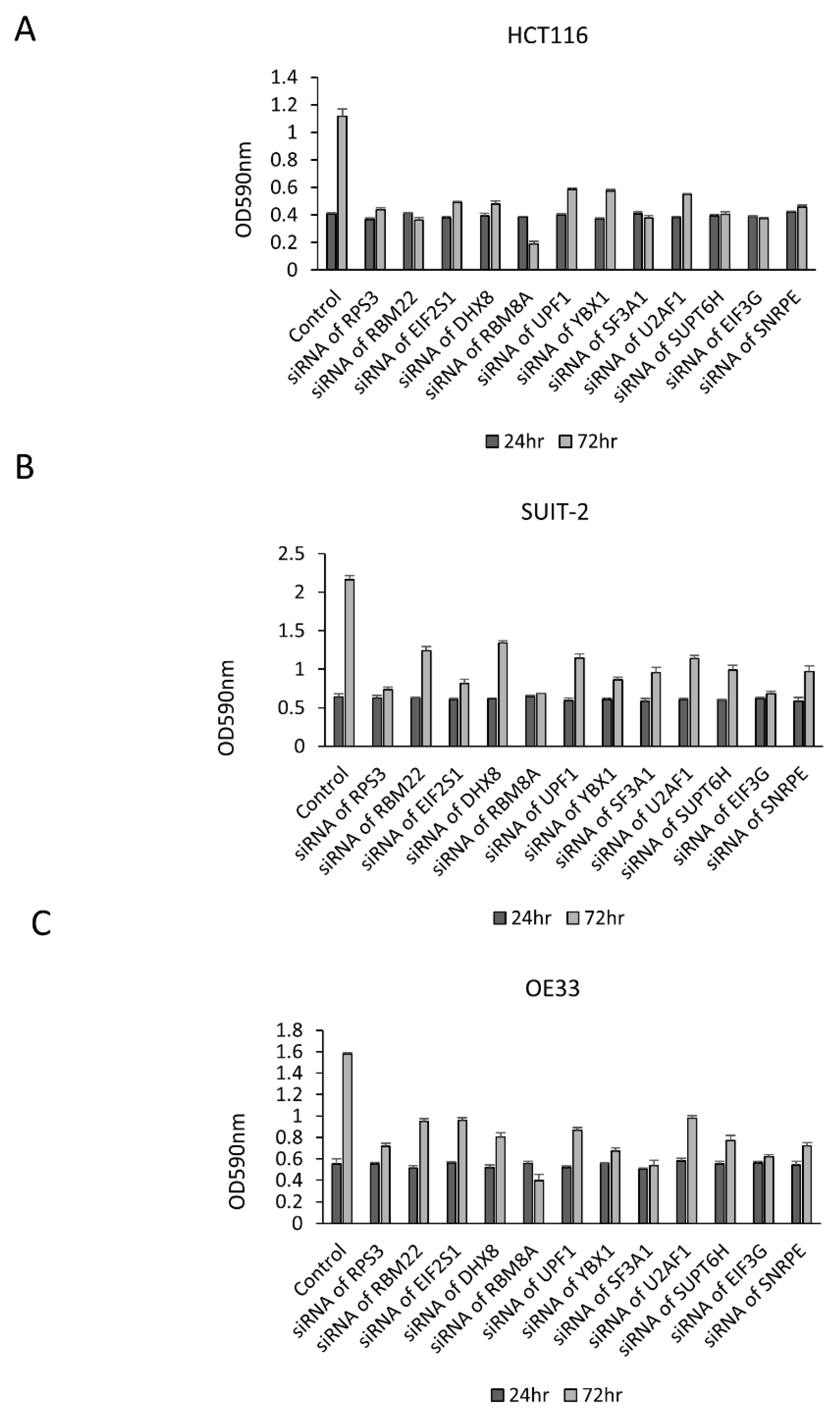
3.1. Growth Change by the Knockdown of RBPs in Gastrointestinal Cancer Cells
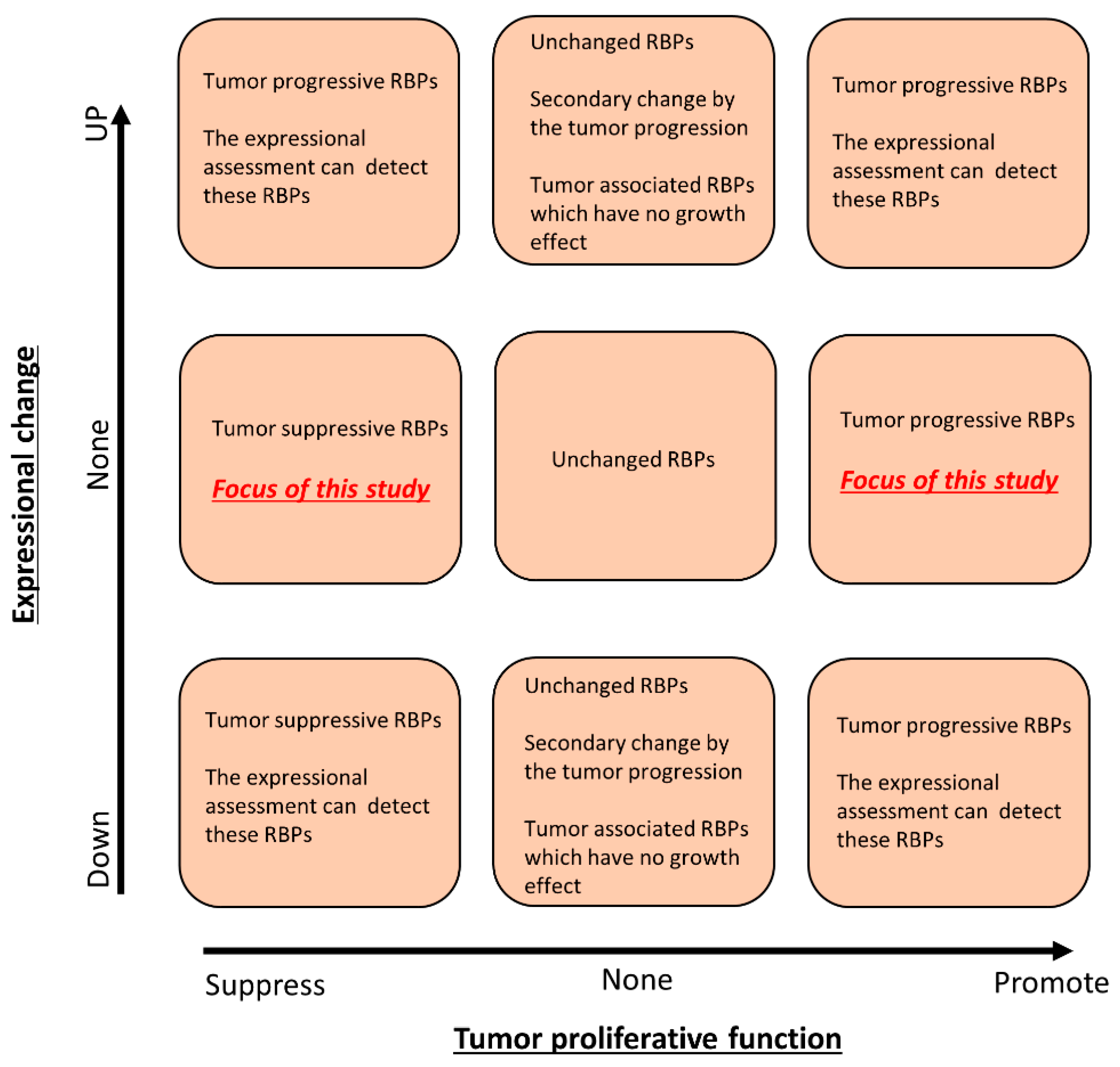
3.2. The Association between Expressional Changes and Tumor Growth Promotive Functions of RBPs in Gastrointestinal Cancer Cells
3.3. Identification of Tumor Growth-Associated RBPs without Any Significant Expressional Changes in Gastrointestinal Cancer Cells
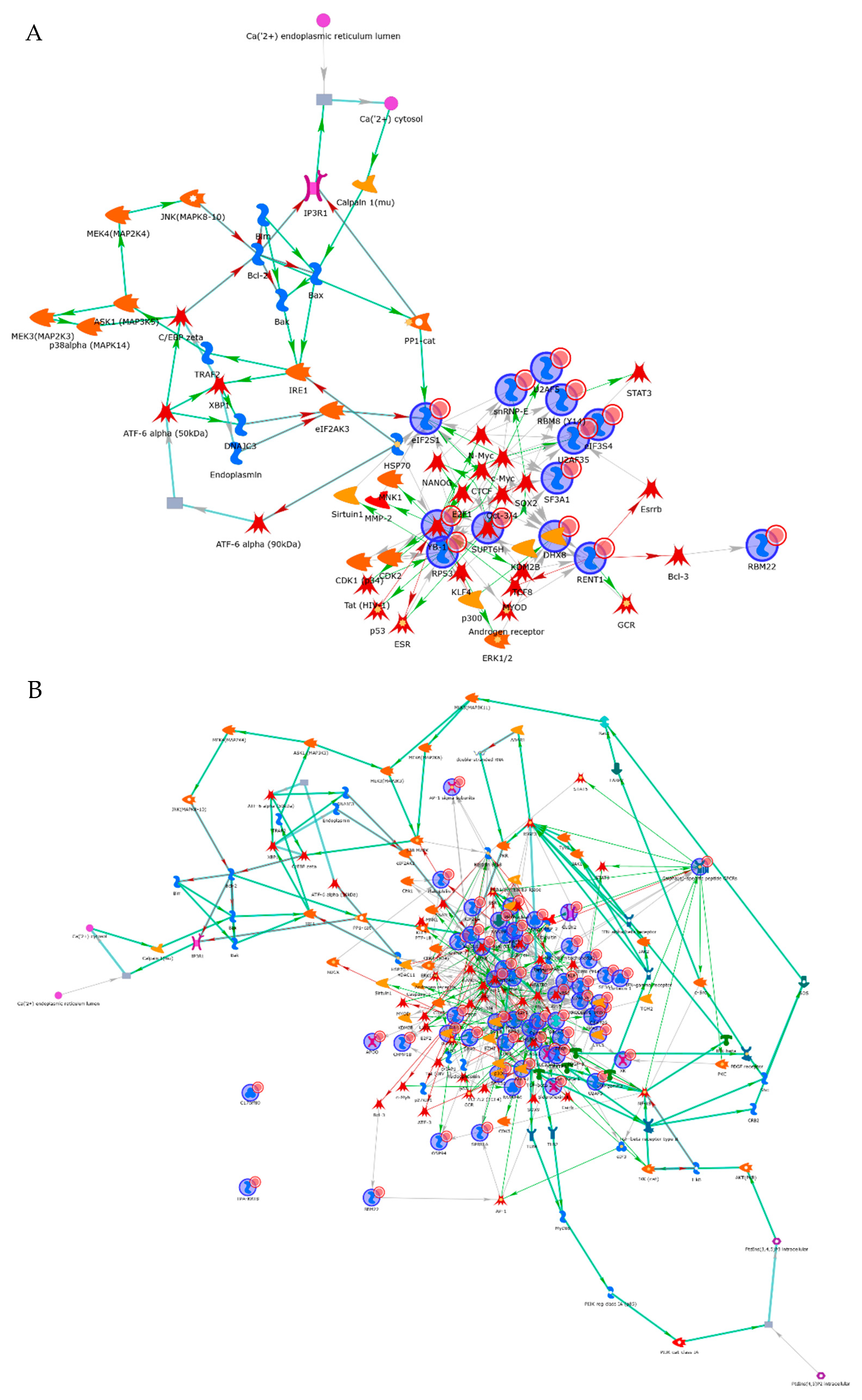
3.4. Interactome Analysis of RBPs in Cancer Cells
3.5. Identification of Organ-Specific RBPs Exhibiting the Tumor-Promotive Function
3.6. Identification of Tumor Growth Associated RBPs without Genetic Mutations in Gastrointestinal Cancer Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mohibi, S.; Chen, X.; Zhang, J. Cancer the ‘RBP’ eutics-RNA-binding proteins as therapeutic targets for cancer. Pharmacol. Ther. 2019, 203, 107390. [Google Scholar] [CrossRef] [PubMed]
- Nilsen, T.W.; Graveley, B.R. Expansion of the eukaryotic proteome by alternative splicing. Nature 2010, 463, 457–463. [Google Scholar] [CrossRef] [Green Version]
- Gerstberger, S.; Hafner, M.; Tuschl, T. A census of human RNA-binding proteins. Nat. Rev. Genet. 2014, 15, 829–845. [Google Scholar] [CrossRef]
- Chatterji, P.; Rustgi, A.K. RNA Binding Proteins in Intestinal Epithelial Biology and Colorectal Cancer. Trends Mol. Med. 2018, 24, 490–506. [Google Scholar] [CrossRef] [PubMed]
- Kudinov, A.E.; Karanicolas, J.; Golemis, E.A.; Boumber, Y. Musashi RNA-Binding Proteins as Cancer Drivers and Novel Therapeutic Targets. Clin. Cancer Res. 2017, 23, 2143–2153. [Google Scholar] [CrossRef] [Green Version]
- Mitobe, Y.; Iino, K.; Takayama, K.; Ikeda, K.; Suzuki, T.; Aogi, K.; Kawabata, H.; Suzuki, Y.; Horie-Inoue, K.; Inoue, S. PSF Promotes ER-Positive Breast Cancer Progression via Posttranscriptional Regulation of ESR1 and SCFD2. Cancer Res. 2020, 80, 2230–2242. [Google Scholar] [CrossRef] [Green Version]
- Denkert, C.; Weichert, W.; Pest, S.; Koch, I.; Licht, D.; Köbel, M.; Reles, A.; Sehouli, J.; Dietel, M.; Hauptmann, S. Overexpression of the embryonic-lethal abnormal vision-like protein HuR in ovarian carcinoma is a prognostic factor and is associated with increased cyclooxygenase 2 expression. Cancer Res. 2004, 64, 189–195. [Google Scholar] [CrossRef] [Green Version]
- Kang, H.; Heo, S.; Shin, J.J.; Ji, E.; Tak, H.; Ahn, S.; Lee, K.J.; Lee, E.K.; Kim, W. A miR-194/PTBP1/CCND3 axis regulates tumor growth in human hepatocellular carcinoma. J. Pathol. 2019, 249, 395–408. [Google Scholar] [CrossRef]
- Franses, J.W.; Philipp, J.; Missios, P.; Bhan, I.; Liu, A.; Yashaswini, C.; Tai, E.; Zhu, H.; Ligorio, M.; Nicholson, B.; et al. Pancreatic circulating tumor cell profiling identifies LIN28B as a metastasis driver and drug target. Nat. Commun. 2020, 11, 3303. [Google Scholar] [CrossRef]
- Wang, Z.L.; Li, B.; Luo, Y.X.; Lin, Q.; Liu, S.R.; Zhang, X.Q.; Zhou, H.; Yang, J.H.; Qu, L.H. Comprehensive Genomic Characterization of RNA-Binding Proteins across Human Cancers. Cell Rep. 2018, 22, 286–298. [Google Scholar] [CrossRef] [Green Version]
- Fujiya, M.; Konishi, H.; Mohamed Kamel, M.K.; Ueno, N.; Inaba, Y.; Moriichi, K.; Tanabe, H.; Ikuta, K.; Ohtake, T.; Kohgo, Y. microRNA-18a induces apoptosis in colon cancer cells via the autophagolysosomal degradation of oncogenic heterogeneous nuclear ribonucleoprotein A1. Oncogene 2014, 33, 4847–4856. [Google Scholar] [CrossRef]
- Konishi, H.; Fujiya, M.; Kashima, S.; Sakatani, A.; Dokoshi, T.; Ando, K.; Ueno, N.; Iwama, T.; Moriichi, K.; Tanaka, H.; et al. A tumor-specific modulation of heterogeneous ribonucleoprotein A0 promotes excessive mitosis and growth in colorectal cancer cells. Cell Death Dis. 2020, 11, 245. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Frost, P.; Hoang, B.; Benavides, A.; Gera, J.; Lichtenstein, A. IL-6-induced enhancement of c-Myc translation in multiple myeloma cells: Critical role of cytoplasmic localization of the rna-binding protein hnRNP A1. J. Biol. Chem. 2011, 286, 67–78. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Konishi, H.; Fujiya, M.; Ueno, N.; Moriichi, K.; Sasajima, J.; Ikuta, K.; Tanabe, H.; Tanaka, H.; Kohgo, Y. Microrna-26a and -584 inhibit the colorectal cancer progression through inhibition of the binding of hnRNP A1-CDK6 mRNA. Biochem. Biophys. Res. Commun. 2015, 467, 847–852. [Google Scholar] [CrossRef] [PubMed]
- Forrester, K.; Almoguera, C.; Han, K.; Grizzle, W.E.; Perucho, M. Detection of high incidence of K-ras oncogenes during human colon tumorigenesis. Nature 1987, 327, 298–303. [Google Scholar] [CrossRef] [Green Version]
- Dachs, G.U.; Patterson, A.V.; Firth, J.D.; Ratcliffe, P.J.; Townsend, K.M.; Stratford, I.J.; Harris, A.L. Targeting gene expression to hypoxic tumor cells. Nat. Med. 1997, 3, 515–520. [Google Scholar] [CrossRef]
- Eliyahu, D.; Raz, A.; Gruss, P.; Givol, D.; Oren, M. Participation of p53 cellular tumour antigen in transformation of normal embryonic cells. Nature 1984, 312, 646–649. [Google Scholar] [CrossRef]
- Chen, T.; Zhang, B.; Ziegenhals, T.; Prusty, A.B.; Fröhler, S.; Grimm, C.; Hu, Y.; Schaefke, B.; Fang, L.; Zhang, M.; et al. A missense mutation in SNRPE linked to non-syndromal microcephaly interferes with U snRNP assembly and pre-mRNA splicing. PLoS Genet. 2019, 15, e1008460. [Google Scholar] [CrossRef]
- Sharma, S.; Wongpalee, S.P.; Vashisht, A.; Wohlschlegel, J.A.; Black, D.L. Stem-loop 4 of U1 snRNA is essential for splicing and interacts with the U2 snRNP-specific SF3A1 protein during spliceosome assembly. Genes Dev. 2014, 28, 2518–2531. [Google Scholar] [CrossRef] [Green Version]
- Endoh, M.; Zhu, W.; Hasegawa, J.; Watanabe, H.; Kim, D.K.; Aida, M.; Inukai, N.; Narita, T.; Yamada, T.; Furuya, A.; et al. Human Spt6 stimulates transcription elongation by RNA polymerase II in vitro. Mol. Cell Biol. 2004, 24, 3324–3336. [Google Scholar] [CrossRef] [Green Version]
- Obara, E.A.A.; Aguilar-Morante, D.; Rasmussen, R.D.; Frias, A.; Vitting-Serup, K.; Lim, Y.C.; Elbæk, K.J.; Pedersen, H.; Vardouli, L.; Jensen, K.E.; et al. SPT6-driven error-free DNA repair safeguards genomic stability of glioblastoma cancer stem-like cells. Nat. Commun. 2020, 11, 4709. [Google Scholar] [CrossRef]
- Bedi, U.; Scheel, A.H.; Hennion, M.; Begus-Nahrmann, Y.; Rüschoff, J.; Johnsen, S.A. SUPT6H controls estrogen receptor activity and cellular differentiation by multiple epigenomic mechanisms. Oncogene 2015, 34, 465–473. [Google Scholar] [CrossRef]
- Chen, Y.; Zhang, L.; Jones, K.A. SKIP counteracts p53-mediated apoptosis via selective regulation of p21Cip1 mRNA splicing. Genes Dev. 2011, 25, 701–716. [Google Scholar] [CrossRef] [Green Version]
- Felisberto-Rodrigues, C.; Thomas, J.C.; McAndrew, C.; Le Bihan, Y.V.; Burke, R.; Workman, P.; van Montfort, R.L.M. Structural and functional characterisation of human RNA helicase DHX8 provides insights into the mechanism of RNA-stimulated ADP release. Biochem. J. 2019, 476, 2521–2543. [Google Scholar] [CrossRef] [Green Version]
- Choe, J.; Oh, N.; Park, S.; Lee, Y.K.; Song, O.K.; Locker, N.; Chi, S.G.; Kim, Y.K. Translation initiation on mRNAs bound by nuclear cap-binding protein complex CBP80/20 requires interaction between CBP80/20-dependent translation initiation factor and eukaryotic translation initiation factor 3g. J. Biol. Chem. 2012, 287, 18500–18509. [Google Scholar] [CrossRef] [Green Version]
- Rasche, N.; Dybkov, O.; Schmitzová, J.; Akyildiz, B.; Fabrizio, P.; Lührmann, R. Cwc2 and its human homologue RBM22 promote an active conformation of the spliceosome catalytic centre. EMBO J. 2012, 31, 1591–1604. [Google Scholar] [CrossRef] [Green Version]
- Wu, T.; Fu, X.D. Genomic functions of U2AF in constitutive and regulated splicing. RNA Biol. 2015, 12, 479–485. [Google Scholar] [CrossRef] [Green Version]
- Bhattacharya, A.; Czaplinski, K.; Trifillis, P.; He, F.; Jacobson, A.; Peltz, S.W. Characterization of the biochemical properties of the human Upf1 gene product that is involved in nonsense-mediated mRNA decay. RNA 2000, 6, 1226–1235. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; Geng, Y.; Feng, R.; Zhu, Q.; Miao, B.; Cao, J.; Fei, S. The Human RNA Surveillance Factor UPF1 Modulates Gastric Cancer Progression by Targeting Long Non-Coding RNA MALAT1. Cell Physiol. Biochem. 2017, 42, 2194–2206. [Google Scholar] [CrossRef] [Green Version]
- Hasegawa, S.L.; Doetsch, P.W.; Hamilton, K.K.; Martin, A.M.; Okenquist, S.A.; Lenz, J.; Boss, J.M. DNA binding properties of YB-1 and dbpA: Binding to double-stranded, single-stranded, and abasic site containing DNAs. Nucleic Acids Res. 1991, 19, 4915–4920. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.Y.; Gherzi, R.; Andersen, J.S.; Gaietta, G.; Jürchott, K.; Royer, H.D.; Mann, M.; Karin, M. Nucleolin and YB-1 are required for JNK-mediated interleukin-2 mRNA stabilization during T-cell activation. Genes Dev. 2000, 14, 1236–1248. [Google Scholar]
- Raffetseder, U.; Frye, B.; Rauen, T.; Jürchott, K.; Royer, H.D.; Jansen, P.L.; Mertens, P.R. Splicing factor SRp30c interaction with Y-box protein-1 confers nuclear YB-1 shuttling and alternative splice site selection. J. Biol. Chem. 2003, 278, 18241–18248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, X.; Su, L.; Liu, X. Loss of CDH1 up-regulates epidermal growth factor receptor via phosphorylation of YBX1 in non-small cell lung cancer cells. FEBS Lett. 2013, 587, 3995–4000. [Google Scholar] [CrossRef] [Green Version]
- Wan, F.; Lenardo, M.J. The nuclear signaling of NF-kappaB: Current knowledge, new insights, and future perspectives. Cell Res. 2010, 20, 24–33. [Google Scholar] [CrossRef] [Green Version]
- Scheuner, D.; Song, B.; McEwen, E.; Liu, C.; Laybutt, R.; Gillespie, P.; Saunders, T.; Bonner-Weir, S.; Kaufman, R.J. Translational control is required for the unfolded protein response and in vivo glucose homeostasis. Mol. Cell 2001, 7, 1165–1176. [Google Scholar] [CrossRef]
- Mao, H.; McMahon, J.J.; Tsai, Y.H.; Wang, Z.; Silver, D.L. Haploinsufficiency for Core Exon Junction Complex Components Disrupts Embryonic Neurogenesis and Causes p53-Mediated Microcephaly. PLoS Genet. 2016, 12, e1006282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chong, C.R.; Jänne, P.A. The quest to overcome resistance to EGFR-targeted therapies in cancer. Nat. Med. 2013, 19, 1389–1400. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Groden, J.; Thliveris, A.; Samowitz, W.; Carlson, M.; Gelbert, L.; Albertsen, H.; Joslyn, G.; Stevens, J.; Spirio, L.; Robertson, M.; et al. Identification and characterization of the familial adenomatous polyposis coli gene. Cell 1991, 66, 589–600. [Google Scholar] [CrossRef]
- Powell, S.M.; Zilz, N.; Beazer-Barclay, Y.; Bryan, T.M.; Hamilton, S.R.; Thibodeau, S.N.; Vogelstein, B.; Kinzler, K.W. APC mutations occur early during colorectal tumorigenesis. Nature 1992, 359, 235–237. [Google Scholar] [CrossRef] [PubMed]



| Name | Characteristics | Organs | Morphology | Supplier |
|---|---|---|---|---|
| HCT116 | Colorectal cancer cells | Colon | Epithelial | American Type Culture Collection (ATCC) |
| SW480 | Colorectal cancer cells | Colon | Epithelial | American Type Culture Collection (ATCC) |
| HCEC-1CT | Immortalized colorectal epithelial cells | Colon | Epithelial | Evercyte |
| SUIT2 | Pancreatic cancer cells | Pancreas | Epithelial | Health Science Research Resources Bank |
| PANC-1 | Pancreatic cancer cells | Pancreas | Epithelial | American Type Culture Collection (ATCC) |
| HPPEC | Primary pancreatic epithelial cells | Pancreas | Epithelial | Cell Biologics, Inc. |
| OE33 | Esophageal cancer cells | Esophagus | Epithelial | European Collection of Authenticated Cell Cultures (ECACC) |
| KYSE70 | Esophageal cancer cells | Esophagus | Epithelial | European Collection of Authenticated Cell Cultures (ECACC) |
| Het-1A | Immortalized esophageal epithelial cells | Esophagus | Epithelial | American Type Culture Collection (ATCC) |
| Gene Symbol | siRNA ID | Growth Inhibition Effect | Array ID | Fold Change | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| HCT 116 | SW 480 | SUIT -2 | PANC -1 | OE 33 | KYSE 70 | HCT 116 | SW 480 | SUIT -2 | PANC −1 | OE 33 | KYSE 70 | |||
| DHX8 | s4018 | 94.6% | 10.7% | 64.5% | 39.5% | 88.8% | 20.8% | TC1700012263.hg.1 | 1.28 | 1.10 | −1.02 | −2.79 | −1.43 | −2.39 |
| s4017 | 91.5% | 41.8% | 58.4% | 61.3% | 109.1% | 47.3% | ||||||||
| s4016 | 80.8% | 10.3% | 60.7% | 53.5% | 75.8% | 36.9% | ||||||||
| EIF2S1 | s4557 | 92.0% | 10.5% | 79.1% | 24.9% | 63.0% | 42.5% | TC1400007494.hg.1 | 1.29 | 1.39 | −1.45 | 1.01 | −1.04 | 1.24 |
| s4556 | 90.1% | 4.2% | 80.1% | 30.4% | 65.1% | 42.5% | ||||||||
| s4555 | 87.0% | 31.0% | 92.7% | 44.1% | 66.7% | 48.9% | ||||||||
| EIF3G | s225017 | 89.6% | 26.7% | 88.0% | 72.9% | 84.2% | 67.6% | TC1900009608.hg.1 | 1.34 | 1.01 | −1.80 | −2.10 | −1.42 | 1.02 |
| s16505 | 88.8% | 39.2% | 84.5% | 46.7% | 77.1% | 70.8% | ||||||||
| s16504 | 83.2% | 47.8% | 90.7% | 67.3% | 85.1% | 79.0% | ||||||||
| RBM22 | s31272 | 95.3% | 44.3% | 54.7% | 49.5% | 61.3% | 53.3% | TC0500012485.hg.1 | 1.71 | 1.78 | 1.05 | 1.77 | −1.82 | −1.66 |
| s31273 | 54.6% | 15.5% | 53.9% | 43.9% | 55.4% | 28.3% | ||||||||
| s31274 | 50.8% | 1.0% | 41.0% | 14.7% | 37.8% | 29.6% | ||||||||
| RBM8A | s532199 | 121.7% | 116.8% | 112.3% | 75.7% | 106.9% | 49.2% | TC0100018477.hg.1 | 1.48 | 1.22 | 1.55 | 1.81 | −1.08 | −1.11 |
| s532200 | 113.3% | 53.9% | 108.5% | 74.2% | 115.4% | 36.7% | ||||||||
| s19292 | 112.1% | 73.5% | 97.3% | 62.7% | 112.4% | 64.9% | ||||||||
| RPS3 | s12255 | 96.2% | 48.9% | 88.6% | 60.3% | 79.8% | 71.9% | TC1100008462.hg.1 | 1.26 | 1.43 | 1.15 | −1.21 | 1.08 | −1.34 |
| s12257 | 91.3% | 37.6% | 83.5% | 80.1% | 78.2% | 85.0% | ||||||||
| s12256 | 90.5% | 66.0% | 91.8% | 52.6% | 74.5% | 85.3% | ||||||||
| SF3A1 | s20116 | 105.1% | 38.7% | 105.7% | 57.3% | 89.0% | 60.7% | TC2200008434.hg.1 | 1.62 | 1.26 | 1.99 | 1.88 | −1.33 | −1.00 |
| s20118 | 96.7% | 11.5% | 79.0% | 59.2% | 63.7% | 54.1% | ||||||||
| s20117 | 88.1% | 49.7% | 85.1% | 43.5% | 79.3% | 51.1% | ||||||||
| SNRPE | s13238 | 102.4% | 56.8% | 89.4% | 77.9% | 75.8% | 74.4% | TC0100011276.hg.1 | 1.85 | 2.00 | 1.42 | 1.36 | 1.57 | 1.52 |
| s13239 | 92.4% | 27.6% | 75.3% | 55.8% | 81.8% | 64.3% | ||||||||
| SUPT6H | s13636 | 87.3% | 24.6% | 70.8% | 55.5% | 88.3% | 71.7% | TC1700007381.hg.1 | −1.05 | −1.32 | 1.07 | −1.05 | 1.18 | −2.53 |
| s13634 | 84.9% | 64.5% | 64.2% | 48.9% | 76.9% | 54.5% | ||||||||
| s13635 | 71.7% | 40.4% | 62.9% | 48.6% | 61.6% | 72.7% | ||||||||
| U2AF1 | s14555 | 60.2% | 15.4% | 77.6% | 51.2% | 68.8% | 30.7% | TC2100008286.hg.1 | 1.92 | 2.82 | 1.80 | 1.28 | 1.21 | 3.73 |
| s14553 | 57.8% | 38.4% | 92.4% | 30.1% | 85.3% | 20.6% | TC2100007492.hg.1 | 1.85 | 2.68 | 1.73 | 1.67 | 1.34 | 2.85 | |
| s14554 | 50.3% | 14.0% | 104.1% | 18.6% | 68.2% | 10.9% | ||||||||
| UPF1 | s11926 | 83.0% | 11.9% | 98.0% | 32.4% | 64.6% | 43.3% | TC1900007410.hg.1 | 1.41 | 1.04 | −1.36 | 3.15 | −1.08 | −1.49 |
| s11928 | 54.9% | 7.4% | 90.3% | 44.1% | 80.6% | 34.9% | ||||||||
| s11927 | 10.6% | −0.2% | −13.6% | 32.3% | 34.2% | 9.8% | ||||||||
| YBX1 | s9733 | 99.2% | 14.0% | 78.9% | 28.9% | 100.2% | 10.8% | TC0100008026.hg.1 | 1.25 | 1.42 | 1.12 | 1.03 | −1.29 | 1.11 |
| s9731 | 60.8% | 18.9% | 68.5% | 75.6% | 97.5% | 36.2% | TC0100008025.hg.1 | 1.09 | 1.32 | −1.25 | −1.14 | −4.14 | −3.16 | |
| s9732 | 41.7% | 15.7% | 41.4% | 19.9% | 37.2% | 14.1% | ||||||||
| Gene Symbol | Growth Inhibition Effect | ||
|---|---|---|---|
| HCEC-1CT | HPPEC | Het1A | |
| DHX8 | 27.0% | 11.3% | −14.9% |
| EIF2S1 | 52.9% | 21.8% | 6.4% |
| EIF3G | 20.8% | 18.0% | 19.9% |
| RBM22 | 11.0% | 29.0% | −2.1% |
| RBM8A | 53.4% | 69.3% | −8.5% |
| RPS3 | 58.1% | 46.2% | 2.1% |
| SF3A1 | 24.9% | −6.7% | −10.6% |
| SNRPE | 34.4% | 37.8% | 17.0% |
| SUPT6H | 35.5% | 22.8% | 49.0% |
| U2AF1 | 8.4% | 0.9% | 39.8% |
| UPF1 | 31.9% | 31.9% | 17.0% |
| YBX1 | 29.9% | 31.9% | −2.1% |
| From | To | Effect | Mechanism |
|---|---|---|---|
| Bcl-3 | RBM22 | Transcription regulation | Unspecified |
| c-Myc | RBM8 (Y14) | Transcription regulation | Unspecified |
| N-Myc | RBM8 (Y14) | Transcription regulation | Unspecified |
| SOX2 | RBM8 (Y14) | Transcription regulation | Unspecified |
| Androgen receptor | RENT1 | Transcription regulation | Unspecified |
| KDM2B | RENT1 | Co-regulation of transcription | Unspecified |
| Oct-3/4 | RENT1 | Transcription regulation | Unspecified |
| TCF8 | RENT1 | Transcription regulation | Unspecified |
| Beta-catenin | snRNP-E | Co-regulation of transcription | Unspecified |
| c-Myc | snRNP-E | Transcription regulation | Unspecified |
| NANOG | snRNP-E | Transcription regulation | Unspecified |
| N-Myc | snRNP-E | Transcription regulation | Unspecified |
| SOX2 | snRNP-E | Transcription regulation | Activation |
| SOX9 | snRNP-E | Transcription regulation | Unspecified |
| CDK2 | SUPT6H | Phosphorylation | Unspecified |
| c-Myc | SUPT6H | Transcription regulation | Unspecified |
| NANOG | SUPT6H | Transcription regulation | Unspecified |
| SOX2 | SUPT6H | Transcription regulation | Unspecified |
| TCF8 | SUPT6H | Transcription regulation | Unspecified |
| c-Myc | U2AF35 | Transcription regulation | Activation |
| CTCF | U2AF35 | Transcription regulation | Unspecified |
| Esrrb | U2AF35 | Transcription regulation | Unspecified |
| N-Myc | U2AF35 | Transcription regulation | Unspecified |
| Oct-3/4 | U2AF35 | Transcription regulation | Unspecified |
| STAT3 | U2AF35 | Transcription regulation | Unspecified |
| c-Myc | YB-1 | Transcription regulation | Activation |
| E2F1 | YB-1 | Transcription regulation | Activation |
| ERK1/2 | YB-1 | Phosphorylation | Inhibition |
| ERK2 (MAPK1) | YB-1 | Phosphorylation | Activation |
| HSP60 | YB-1 | Binding | Inhibition |
| HSP70 | YB-1 | Binding | Activation |
| MYOD | YB-1 | Transcription regulation | Activation |
| NANOG | YB-1 | Transcription regulation | Unspecified |
| Oct-3/4 | YB-1 | Transcription regulation | Unspecified |
| p53 | YB-1 | Binding | Inhibition |
| STAT1 | YB-1 | Transcription regulation | Unspecified |
| Ubiquitin | YB-1 | Binding | Inhibition |
| YB-1 | Androgen receptor | Transcription regulation | Unspecified |
| RENT1 | ATF-3 | Binding | Inhibition |
| RENT1 | Bcl-3 | Binding | Inhibition |
| YB-1 | CDK1 (p34) | Transcription regulation | Unspecified |
| SUPT6H | CDK2 | Co-regulation of transcription | Unspecified |
| YB-1 | CDK2 | Transcription regulation | Activation |
| RBM22 | c-Fos | Co-regulation of transcription | Unspecified |
| YB-1 | c-IAP1 | Transcription regulation | Activation |
| eIF3 | c-Jun | Binding | Activation |
| RENT1 | c-Myb | Binding | Inhibition |
| SUPT6H | c-Myc | Co-regulation of transcription | Unspecified |
| YB-1 | c-Myc | Binding | Activation |
| YB-1 | CTCF | Binding | Activation |
| RPS3 | E2F1 | Binding | Activation |
| YB-1 | E2F2 | Transcription regulation | Activation |
| YB-1 | ERK2 (MAPK1) | Binding | Activation |
| YB-1 | ESR | Binding | Inhibition |
| SUPT6H | ESR1 (nuclear) | Binding | Activation |
| YB-1 | ESR1 (nuclear) | Binding | Inhibition |
| RENT1 | Esrrb | Binding | Inhibition |
| YB-1 | EZH2 | Transcription regulation | Unspecified |
| RENT1 | GCR | Binding | Activation |
| YB-1 | GRP78 | Binding | Inhibition |
| YB-1 | HDAC11 | Transcription regulation | Unspecified |
| snRNP-E | HSPA4 | Co-regulation of transcription | Unspecified |
| YB-1 | KDM2B | Transcription regulation | Unspecified |
| RENT1 | Keratin 8 | Binding | Inhibition |
| YB-1 | KLF4 | Transcription regulation | Activation |
| YB-1 | KLF5 | Transcription regulation | Unspecified |
| YB-1 | MMP-2 | Transcription regulation | Activation |
| YB-1 | MNK1 | Transcription regulation | Activation |
| RENT1 | MYOD | Ubiquitination | Inhibition |
| SUPT6H | NANOG | Co-regulation of transcription | Activation |
| YB-1 | NANOG | Binding | Activation |
| RPS3 | NF-kB1 (p50) | Binding | Activation |
| YB-1 | N-Myc | Binding | Activation |
| YB-1 | Nucleophosmin | Transcription regulation | Unspecified |
| SUPT6H | Oct-3/4 | Co-regulation of transcription | Activation |
| YB-1 | p300 | Binding | Inhibition |
| RPS3 | p53 | Binding | Activation |
| YB-1 | p53 | Transcription regulation | Inhibition |
| YB-1 | p63 | Transcription regulation | Unspecified |
| YB-1 | PDGF-B | Transcription regulation | Unspecified |
| SUPT6H | PR (nuclear) | Co-regulation of transcription | Activation |
| YB-1 | PTP-1B | Transcription regulation | Activation |
| RPS3 | RelA (p65 NF-kB subunit) | Binding | Activation |
| RPS3 | Sirtuin1 | Binding | Activation |
| YB-1 | SNAIL1 | Binding | Activation |
| snRNP-E | snRNP-F | Binding | Activation |
| SUPT6H | SOX2 | Co-regulation of transcription | Activation |
| YB-1 | SOX2 | Transcription regulation | Inhibition |
| RBM8 (Y14) | STAT3 | Binding | Activation |
| YB-1 | Tat (HIV-1) | Binding | Activation |
| YB-1 | TCF8 | Transcription regulation | Activation |
| YB-1 | TGF-beta 1 | Binding | Inhibition |
| YB-1 | TGF-beta 2 | Transcription regulation | Unspecified |
| Cell Lines | Gene List | |||||
|---|---|---|---|---|---|---|
| HCT116 | CPEB4 | MKI67IP | RPS3 | SNRPB | YBX1 | |
| DHX8 | NOL8 | RRP7A | SNRPB2 | ZC3H12C | ||
| EIF2S1 | PABPN1 | SAMD4A | SNRPC | |||
| EIF3G | PNO1 | SF3A1 | SNRPE | |||
| HNRNPA1L2 | RAVER1 | SF3B14 | SRRM2 | |||
| HNRNPAB | RBM22 | SF3B4 | SRSF5 | |||
| HNRNPCL1 | RBM8A | SFRS13A | SUPT6H | |||
| LSM2 | RNF113A | SFRS3 | TIA1 | |||
| LSM5 | RNPS1 | SFRS7 | TLR2 | |||
| LSM8 | ROD1 | SLBP | U2AF1 | |||
| SW480 | RBM8A | |||||
| SF3B14 | ||||||
| SNRPB | ||||||
| SUIT-2 | CHERP | LARP6 | RBM11 | RPS3 | SNRPE | UPF3B |
| CPSF4 | LIN28B | RBM18 | SF3A1 | SNRPG | YBX1 | |
| CPSF4L | LSM6 | RBM22 | SF3B14 | SRRM2 | YTHDC1 | |
| DAZL | LSM8 | RBM43 | SFRS13A | SUPT6H | ZC3H12D | |
| DHX8 | MKI67IP | RBM45 | SFRS3 | TDRD10 | ZC3H15 | |
| EIF2S1 | NCL | RBM4B | SFRS6 | TIA1 | ZC3H18 | |
| EIF3B | NONO | RBM7 | SNRPC | TUT1 | ZC3HAV1 | |
| EIF3G | PABPN1L | RBM8A | SNRPD1 | U2AF1 | ZC3HAV1L | |
| HNRNPAB | POLR2G | RBMY1A1 | SNRPD2 | U2SURP | ZFP36L1 | |
| HNRNPK | PUF60 | RC3H1 | SNRPD3 | UPF1 | ||
| PANC-1 | RBM8A | SF3B14 | SNRPD3 | |||
| RPS3 | SNRPD1 | SNRPE | ||||
| SF3A1 | SNRPD2 | SNRPG | ||||
| OE33 | CARHSP1 | LARP1 | PABPC1L2A | RBM8A | SRBD1 | ZC3H18 |
| CHERP | LARP6 | PABPC1L2B | RPS3 | SUPT6H | ZC3H4 | |
| CSDC2 | LEMD3 | PABPC5 | RRP7A | U2AF1 | ZC3H7B | |
| DAZL | LIN28B | PARP10 | SF1 | U2SURP | ZC3HAV1L | |
| DHX8 | LSM2 | PCBP2 | SF3A1 | UPF1 | ZFP36L1 | |
| EIF2S1 | LSM3 | PRPF3 | SFSWAP | UPF3B | ZFP36L2 | |
| HNRNPA1L2 | MOV10L1 | RBM14 | SNRPD2 | YTHDC1 | ZGPAT | |
| HNRNPC | NONO | RBM22 | SNRPD3 | ZC3H11A | ZNF74 | |
| HNRNPCL1 | NUPL2 | RBM25 | SNRPE | ZC3H12B | ZRANB2 | |
| HNRNPR | PABPC1 | RBM39 | SNRPG | ZC3H12D | ZRSR2 | |
| KYSE-70 | EIF3G | PUF60 | SNRPD2 | |||
| LARP1 | RPS3 | SNRPE | ||||
| NONO | SF3B4 | SRRM1 | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Konishi, H.; Kashima, S.; Goto, T.; Ando, K.; Sakatani, A.; Tanaka, H.; Ueno, N.; Moriichi, K.; Okumura, T.; Fujiya, M. The Identification of RNA-Binding Proteins Functionally Associated with Tumor Progression in Gastrointestinal Cancer. Cancers 2021, 13, 3165. https://doi.org/10.3390/cancers13133165
Konishi H, Kashima S, Goto T, Ando K, Sakatani A, Tanaka H, Ueno N, Moriichi K, Okumura T, Fujiya M. The Identification of RNA-Binding Proteins Functionally Associated with Tumor Progression in Gastrointestinal Cancer. Cancers. 2021; 13(13):3165. https://doi.org/10.3390/cancers13133165
Chicago/Turabian StyleKonishi, Hiroaki, Shin Kashima, Takuma Goto, Katsuyoshi Ando, Aki Sakatani, Hiroki Tanaka, Nobuhiro Ueno, Kentaro Moriichi, Toshikatsu Okumura, and Mikihiro Fujiya. 2021. "The Identification of RNA-Binding Proteins Functionally Associated with Tumor Progression in Gastrointestinal Cancer" Cancers 13, no. 13: 3165. https://doi.org/10.3390/cancers13133165
APA StyleKonishi, H., Kashima, S., Goto, T., Ando, K., Sakatani, A., Tanaka, H., Ueno, N., Moriichi, K., Okumura, T., & Fujiya, M. (2021). The Identification of RNA-Binding Proteins Functionally Associated with Tumor Progression in Gastrointestinal Cancer. Cancers, 13(13), 3165. https://doi.org/10.3390/cancers13133165






