Automated Classification of Significant Prostate Cancer on MRI: A Systematic Review on the Performance of Machine Learning Applications
Abstract
1. Introduction
2. Results
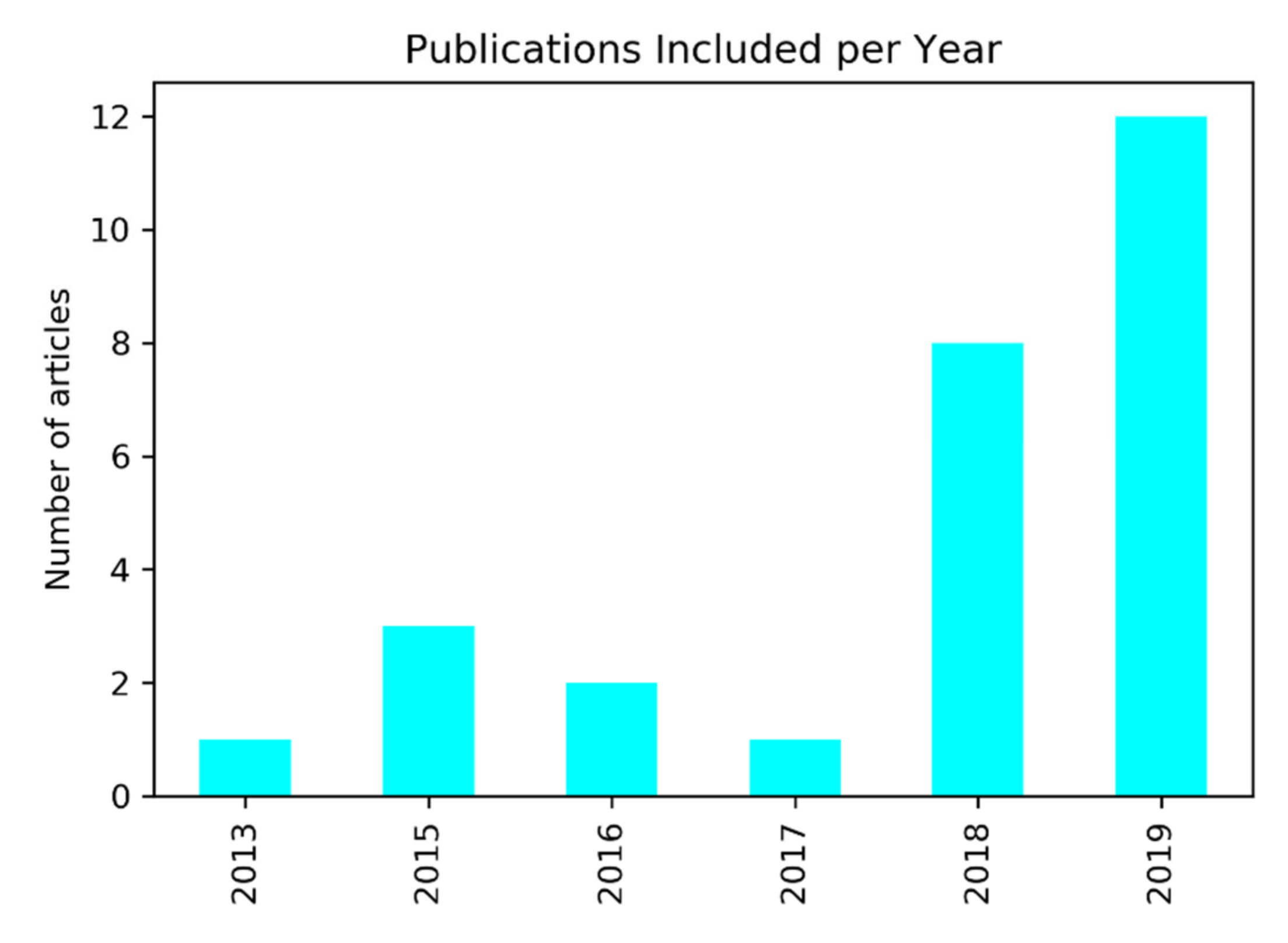
2.1. Search Results and Statistical Description
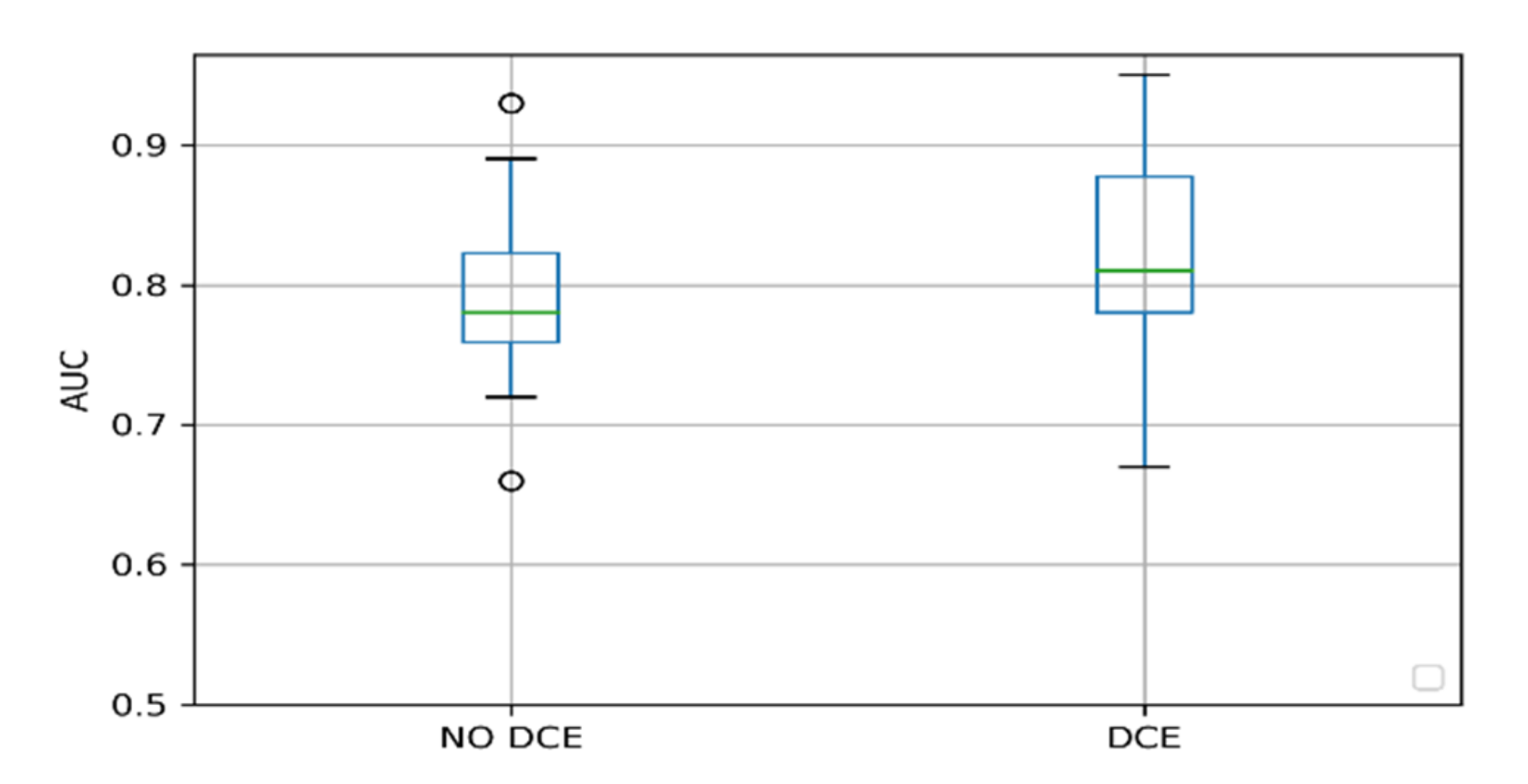
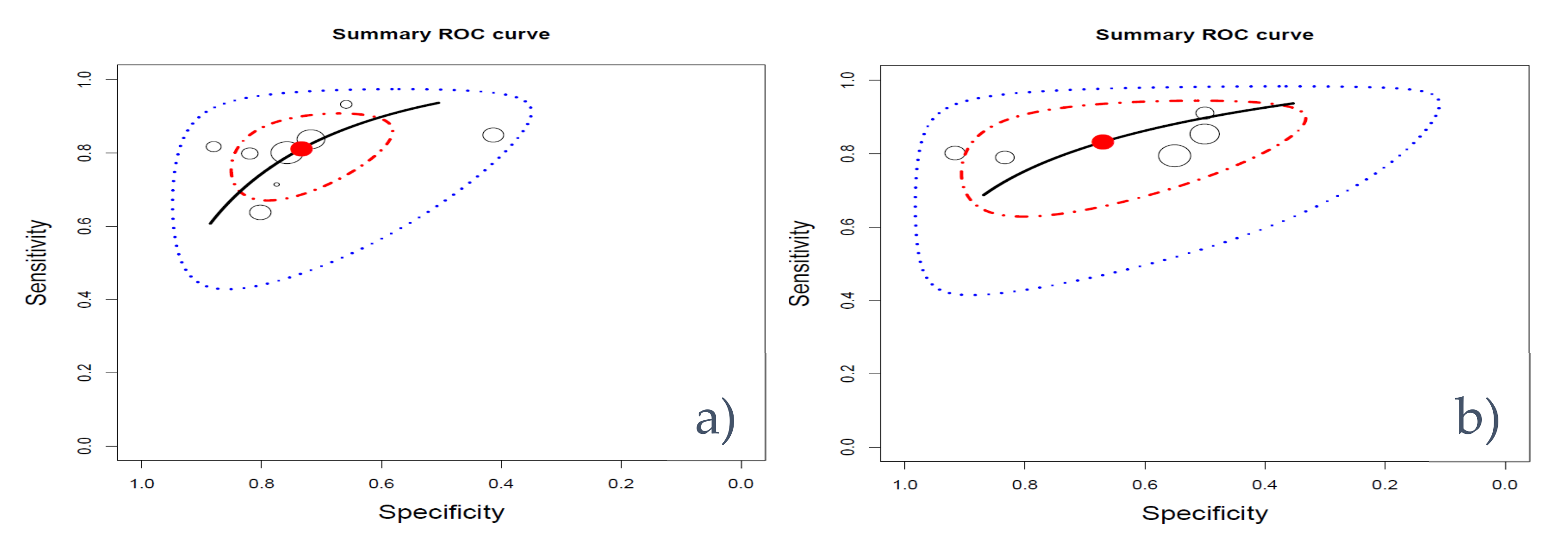
2.2. Factors Influencing the Performance
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Rawla, P. Epidemiology of Prostate Cancer. Rev. World J. Oncol. 2019, 10, 63–89. [Google Scholar] [CrossRef] [PubMed]
- Pinthus, J.H.; Witkos, M.; Fleshner, N.E.; Sweet, J.; Evans, A.; Jewett, M.A.; Krahn, M.; Alibhai, S.; Trachtenberg, J. Prostate Cancers Scored as Gleason 6 on Prostate Biopsy Are Frequently Gleason 7 Tumors at Radical Prostatectomy: Implication on Outcome. J. Urol. 2006, 176, 979–984. [Google Scholar] [CrossRef] [PubMed]
- Mottet, N.; van den Bergh, R.C.N.; Briers, E.; Cornford, P.; De Santis, M.; Fanti, S.; Gillessen, S.; Grummet, J.; Henry, A.M.; Lam, T.B.; et al. European Association of Urology: Prostate Cancer Guidelines. Available online: https://uroweb.org/wp-content/uploads/Prostate-Cancer-2018-pocket.pdf (accessed on 7 April 2020).
- Ahmed, H.U.; El-Shater Bosaily, A.; Brown, L.C.; Gabe, R.; Kaplan, R.; Parmar, M.K.; Collaco-Moraes, Y.; Ward, K.; Hindley, R.G.; Freeman, A.; et al. Diagnostic Accuracy of Multi-Parametric MRI and TRUS Biopsy in Prostate Cancer (PROMIS): A Paired Validating Confirmatory Study. Lancet 2017, 389, 815–822. [Google Scholar] [CrossRef]
- Abraham, B.; Nair, M.S. Computer-Aided Classification of Prostate Cancer Grade Groups from MRI Images Using Texture Features and Stacked Sparse Autoencoder. Comput. Med. Imaging Graph. 2018, 69, 60–68. [Google Scholar] [CrossRef] [PubMed]
- Abraham, B.; Nair, M.S. Automated Grading of Prostate Cancer Using Convolutional Neural Network and Ordinal Class Classifier. Inform. Med. Unlocked 2019, 17, 100256. [Google Scholar] [CrossRef]
- Hoang Dinh, A.; Melodelima, C.; Souchon, R.; Lehaire, J.; Bratan, F.; Mège-Lechevallier, F.; Ruffion, A.; Crouzet, S.; Colombel, M.; Rouvière, O. Quantitative Analysis of Prostate Multiparametric MR Images for Detection of Aggressive Prostate Cancer in the Peripheral Zone: A Multiple Imager Study. Radiology 2016, 280, 117–127. [Google Scholar] [CrossRef]
- Fehr, D.; Veeraraghavan, H.; Wibmer, A.; Gondo, T.; Matsumoto, K.; Vargas, H.A.; Sala, E.; Hricak, H.; Deasy, J.O. Automatic Classification of Prostate Cancer Gleason Scores from Multiparametric Magnetic Resonance Images. Proc. Natl. Acad. Sci. USA 2015, 112, E6265–E6273. [Google Scholar] [CrossRef]
- Jensen, C.; Carl, J.; Boesen, L.; Langkilde, N.C.; Østergaard, L.R. Assessment of Prostate Cancer Prognostic Gleason Grade Group Using Zonal-Specific Features Extracted from Biparametric MRI Using a KNN Classifier. J. Appl. Clin. Med. Phys. 2019, 20, 146–153. [Google Scholar] [CrossRef]
- Kwon, D.; Reis, I.M.; Breto, A.L.; Tschudi, Y.; Gautney, N.; Zavala-Romero, O.; Lopez, C.; Ford, J.C.; Punnen, S.; Pollack, A.; et al. Classification of Suspicious Lesions on Prostate Multiparametric MRI Using Machine Learning. J. Med. Imaging 2018, 5, 1. [Google Scholar] [CrossRef]
- Li, J.; Weng, Z.; Xu, H.; Zhang, Z.; Miao, H.; Chen, W.; Liu, Z.; Zhang, X.; Wang, M.; Xu, X.; et al. Support Vector Machines (SVM) Classification of Prostate Cancer Gleason Score in Central Gland Using Multiparametric Magnetic Resonance Images: A Cross-Validated Study. Eur. J. Radiol. 2018, 98, 61–67. [Google Scholar] [CrossRef]
- Li, Q.; Lu, H.; Choi, J.; Gage, K.; Feuerlein, S.; Pow-Sang, J.M.; Gillies, R.; Balagurunathan, Y. Radiological Semantics Discriminate Clinically Significant Grade Prostate Cancer. Cancer Imaging 2019, 19. [Google Scholar] [CrossRef] [PubMed]
- Maas, M.C.; Litjens, G.J.S.; Wright, A.J.; Attenberger, U.I.; Haider, M.A.; Helbich, T.H.; Kiefer, B.; Macura, K.J.; Margolis, D.J.A.; Padhani, A.R.; et al. A Single-Arm, Multicenter Validation Study of Prostate Cancer Localization and Aggressiveness With a Quantitative Multiparametric Magnetic Resonance Imaging Approach. Invest. Radiol. 2019, 54, 437–447. [Google Scholar] [CrossRef]
- Mehrtash, A.; Sedghi, A.; Ghafoorian, M.; Taghipour, M.; Tempany, C.M.; Wells, W.M.; Kapur, T.; Mousavi, P.; Abolmaesumi, P.; Fedorov, A. Classification of Clinical Significance of MRI Prostate Findings Using 3D Convolutional Neural Networks. In Medical Imaging 2017: Computer-Aided Diagnosis; SPIE: Bellingham, WA, USA, 2017; Volume 10134, p. 101342A. [Google Scholar] [CrossRef]
- Peng, Y.; Jiang, Y.; Yang, C.; Brown, J.B.; Antic, T.; Sethi, I.; Schmid-Tannwald, C.; Giger, M.L.; Eggener, S.E.; Oto, A. Quantitative Analysis of Multiparametric Prostate MR Images: Differentiation between Prostate Cancer and Normal Tissue and Correlation with Gleason Score-A Computer-Aided Diagnosis Development Study. Radiology 2013, 267, 787–796. [Google Scholar] [CrossRef] [PubMed]
- Penzias, G.; Singanamalli, A.; Elliott, R.; Gollamudi, J.; Shih, N.; Feldman, M.; Stricker, P.D.; Delprado, W.; Tiwari, S.; Böhm, M.; et al. Identifying the Morphologic Basis for Radiomic Features in Distinguishing Different Gleason Grades of Prostate Cancer on MRI: Preliminary Findings. PLoS ONE 2018, 13, e0200730. [Google Scholar] [CrossRef] [PubMed]
- Antonelli, M.; Johnston, E.W.; Dikaios, N.; Cheung, K.K.; Sidhu, H.S.; Appayya, M.B.; Giganti, F.; Simmons, L.A.M.; Freeman, A.; Allen, C.; et al. Machine Learning Classifiers Can Predict Gleason Pattern 4 Prostate Cancer with Greater Accuracy than Experienced Radiologists. Eur. Radiol. 2019, 29, 4754–4764. [Google Scholar] [CrossRef]
- Rozenberg, R.; Thornhill, R.E.; Flood, T.A.; Hakim, S.W.; Lim, C.; Schieda, N. Whole-Tumor Quantitative Apparent Diffusion Coefficient Histogram and Texture Analysis to Predict Gleason Score Upgrading in Intermediate-Risk 3 + 4 = 7 Prostate Cancer. Am. J. Roentgenol. 2016, 206, 775–782. [Google Scholar] [CrossRef]
- Schelb, P.; Kohl, S.; Radtke, J.P.; Wiesenfarth, M.; Kickingereder, P.; Bickelhaupt, S.; Kuder, T.A.; Stenzinger, A.; Hohenfellner, M.; Schlemmer, H.-P.; et al. Classification of Cancer at Prostate MRI: Deep Learning versus Clinical PI-RADS Assessment. Radiology 2019, 293, 607–617. [Google Scholar] [CrossRef]
- Toivonen, J.; Montoya Perez, I.; Movahedi, P.; Merisaari, H.; Pesola, M.; Taimen, P.; Boström, P.J.; Pohjankukka, J.; Kiviniemi, A.; Pahikkala, T.; et al. Radiomics and Machine Learning of Multisequence Multiparametric Prostate MRI: Towards Improved Non-Invasive Prostate Cancer Characterization. PLoS ONE 2019, 14, e0217702. [Google Scholar] [CrossRef]
- Transin, S.; Souchon, R.; Gonindard-Melodelima, C.; de Rozario, R.; Walker, P.; Funes de la Vega, M.; Loffroy, R.; Cormier, L.; Rouvière, O. Computer-Aided Diagnosis System for Characterizing ISUP Grade ≥ 2 Prostate Cancers at Multiparametric MRI: A Cross-Vendor Evaluation. Diagn. Interv. Imaging 2019, 100, 801–811. [Google Scholar] [CrossRef]
- Min, X.; Li, M.; Dong, D.; Feng, Z.; Zhang, P.; Ke, Z.; You, H.; Han, F.; Ma, H.; Tian, J.; et al. Multi-Parametric MRI-Based Radiomics Signature for Discriminating between Clinically Significant and Insignificant Prostate Cancer: Cross-Validation of a Machine Learning Method. Eur. J. Radiol. 2019, 115, 16–21. [Google Scholar] [CrossRef]
- Zhong, X.; Cao, R.; Shakeri, S.; Scalzo, F.; Lee, Y.; Enzmann, D.R.; Wu, H.H.; Raman, S.S.; Sung, K. Deep Transfer Learning-Based Prostate Cancer Classification Using 3 Tesla Multi-Parametric MRI. Abdom. Radiol. 2019, 44, 2030–2039. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Gao, G.; Liu, Y.; Han, C.; Liu, J.; Zhang, X.; Wang, X. Feasibility of Integrating Computer-Aided Diagnosis with Structured Reports of Prostate Multiparametric MRI. Clin. Imaging 2020, 60, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Chaddad, A.; Niazi, T.; Probst, S.; Bladou, F.; Anidjar, M.; Bahoric, B. Predicting Gleason Score of Prostate Cancer Patients Using Radiomic Analysis. Front. Oncol. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Chaddad, A.; Kucharczyk, M.J.; Niazi, T. Multimodal Radiomic Features for the Predicting Gleason Score of Prostate Cancer. Cancers 2018, 10, 249. [Google Scholar] [CrossRef]
- Chen, Q.; Hu, S.; Long, P.; Lu, F.; Shi, Y.; Li, Y. A Transfer Learning Approach for Malignant Prostate Lesion Detection on Multiparametric MRI. Technol. Cancer Res. Treat. 2019, 18. [Google Scholar] [CrossRef]
- Chen, T.; Li, M.; Gu, Y.; Zhang, Y.; Yang, S.; Wei, C.; Wu, J.; Li, X.; Zhao, W.; Shen, J. Prostate Cancer Differentiation and Aggressiveness: Assessment With a Radiomic-Based Model vs. PI-RADS V2. J. Magn. Reson. Imaging 2019, 49, 875–884. [Google Scholar] [CrossRef]
- Dikaios, N.; Alkalbani, J.; Sidhu, H.S.; Fujiwara, T.; Abd-Alazeez, M.; Kirkham, A.; Allen, C.; Ahmed, H.; Emberton, M.; Freeman, A.; et al. Logistic Regression Model for Diagnosis of Transition Zone Prostate Cancer on Multi-Parametric MRI. Eur. Radiol. 2015, 25, 523–532. [Google Scholar] [CrossRef]
- Dikaios, N.; Alkalbani, J.; Abd-Alazeez, M.; Sidhu, H.S.; Kirkham, A.; Ahmed, H.U.; Emberton, M.; Freeman, A.; Halligan, S.; Taylor, S.; et al. Zone-Specific Logistic Regression Models Improve Classification of Prostate Cancer on Multi-Parametric MRI. Eur. Radiol. 2015, 25, 2727–2737. [Google Scholar] [CrossRef]
- Dinh, A.H.; Melodelima, C.; Souchon, R.; Moldovan, P.C.; Bratan, F.; Pagnoux, G.; Mège-Lechevallier, F.; Ruffion, A.; Crouzet, S.; Colombel, M.; et al. Characterization of Prostate Cancer with Gleason Score of at Least 7 by Using Quantitative Multiparametric MR Imaging: Validation of a Computer-Aided Diagnosis System in Patients Referred for Prostate Biopsy. Radiology 2018, 287, 525–533. [Google Scholar] [CrossRef]
- McGauran, N.; Wieseler, B.; Kreis, J.; Schüler, Y.B.; Kölsch, H.; Kaiser, T. Reporting Bias in Medical Research -A Narrative Review. Trials. BioMed. Central 2010, 37. [Google Scholar] [CrossRef]
- Schoots, I.G.; Osses, D.F.; Drost, F.J.H.; Verbeek, J.F.M.; Remmers, S.; van Leenders, G.J.L.H.; Bangma, C.H.; Roobol, M.J. Reduction of MRI-Targeted Biopsies in Men with Low-Risk Prostate Cancer on Active Surveillance by Stratifying to PI-RADS and PSAdensity, with Different Thresholds for Significant Disease. Transl. Androl. Urol. 2018, 7, 132–144. [Google Scholar] [CrossRef] [PubMed]
- Capobianco, E.; Dominietto, M. From Medical Imaging to Radiomics: Role of Data Science for Advancing Precision Health. J. Pers. Med. 2020, 10, 15. [Google Scholar] [CrossRef] [PubMed]
- Buch, K.; Kuno, H.; Qureshi, M.M.; Li, B.; Sakai, O. Quantitative Variations in Texture Analysis Features Dependent on MRI Scanning Parameters: A Phantom Model. J. Appl. Clin. Med. Phys. 2018, 19, 253–264. [Google Scholar] [CrossRef] [PubMed]
- Padhani, A.R.; Schoots, I.; Villeirs, G. Contrast Medium or No Contrast Medium for Prostate Cancer Diagnosis. That Is the Question. J. Magn. Reson. Imaging 2020. [Google Scholar] [CrossRef]
- Wolters, T.; Roobol, M.J.; van Leeuwen, P.J.; van den Bergh, R.C.N.; Hoedemaeker, R.F.; van Leenders, G.J.L.H.; Schröder, F.H.; van der Kwast, T.H. A Critical Analysis of the Tumor Volume Threshold for Clinically Insignificant Prostate Cancer Using a Data Set of a Randomized Screening Trial. J. Urol. 2011, 185, 121–125. [Google Scholar] [CrossRef]
- Starmans, M.P.A.; van der Voort, S.R.; Tovar, J.M.C.; Veenland, J.F.; Klein, S.; Niessen, W.J. Radiomics. In Handbook of Medical Image Computing and Computer Assisted Intervention; Elsevier: Amsterdam, The Netherlands, 2019; pp. 429–456. [Google Scholar] [CrossRef]
- Santos, M.S.; Soares, J.P.; Abreu, P.H.; Araujo, H.; Santos, J. Cross-Validation for Imbalanced Datasets: Avoiding Overoptimistic and Overfitting Approaches [Research Frontier]. IEEE Comput. Intell. Mag. 2018, 13, 59–76. [Google Scholar] [CrossRef]
- Ursprung, S.; Beer, L.; Bruining, A.; Woitek, R.; Stewart, G.D.; Gallagher, F.A.; Sala, E. Radiomics of Computed Tomography and Magnetic Resonance Imaging in Renal Cell Carcinoma—A Systematic Review and Meta-Analysis. Eur. Radiol. 2020, 1–9. [Google Scholar] [CrossRef]
- Nieboer, D.; Van Ploeg, T.D.; Steyerberg, E.W. Assessing Discriminative Performance at External Validation of Clinical Prediction Models. PLoS ONE 2016, 11, e0148820. [Google Scholar] [CrossRef]
- Xu, N.; Wu, Y.P.; Li, X.D.; Lin, M.Y.; Zheng, Q.S.; Chen, S.H.; Li, J.F.; Wei, Y.; Xue, X.Y. Risk of Upgrading from Prostate Biopsy to Radical Prostatectomy Pathology: Is Magnetic Resonance Imaging-Guided Biopsy More Accurate? J. Cancer 2018, 9, 3634–3639. [Google Scholar] [CrossRef]
- Moher, D.; Shamseer, L.; Clarke, M.; Ghersi, D.; Liberati, A.; Petticrew, M.; Shekelle, P.; Stewart, L.A.; Estarli, M.; Barrera, E.S.A.; et al. Preferred Reporting Items for Systematic Review and Meta-Analysis Protocols (PRISMA-P) 2015 Statement. Rev. Esp. Nutr. Humana y Diet. 2016, 20, 148–160. [Google Scholar] [CrossRef]
- Lambin, P.; Leijenaar, R.T.H.; Deist, T.M.; Peerlings, J.; De Jong, E.E.C.; Van Timmeren, J.; Sanduleanu, S.; Larue, R.T.H.M.; Even, A.J.G.; Jochems, A.; et al. Radiomics: The Bridge between Medical Imaging and Personalized Medicine. In Nature Reviews Clinical Oncology; Nature Publishing Group: Berlin, Germany, 2017; pp. 749–762. [Google Scholar] [CrossRef]
- Whiting, P.F. QUADAS-2: A Revised Tool for the Quality Assessment of Diagnostic Accuracy Studies. Ann. Intern. Med. 2011, 155, 529. [Google Scholar] [CrossRef] [PubMed]
- Takwoingi, Y. Meta-Analysis of Test Accuracy Studies in R: A Summary of User-Written Programs and Step-by-Step Guide to Using Glmer; 2016; pp 1–30. Cochrane Methods Screening and Diagnostic Tests. Available online: https://methods.cochrane.org/sdt/sites/methods.cochrane.org.sdt/files/public/uploads/R%20DTA%20meta-analysis%20v1.0.zip (accessed on 3 April 2020).


| Method | Median (AUC) | IQR | N | |
|---|---|---|---|---|
| SVM | 0.91 | 0.84 | 0.92 | 3 |
| Linear mix model * | 0.89 | 0.84 | 0.92 | 3 |
| k-nearest neighbor | 0.87 | 0.87 | 0.87 | 1 |
| Neural Network | 0.81 | 0.76 | 0.84 | 2 |
| CNN | 0.80 | 0.78 | 0.83 | 6 |
| Random Forest * | 0.80 | 0.75 | 0.82 | 4 |
| Logistic regression | 0.79 | 0.77 | 0.79 | 6 |
| Linear discriminant analysis | 0.74 | 0.72 | 0.76 | 2 |
| Criteria | Points | Min | Max |
|---|---|---|---|
| Bias due to population selection | Patient selection introduces bias (−5), patient selection might introduce bias 0, patient selection does not introduce bias (+5). | −5 | 5 |
| MRI protocol description | MRI protocols are well documented (+1). | 0 | 1 |
| Multiple segmentations | Study includes segmentations from several physicians/algorithms/software. Study analyzes method robustness to segmentation variabilities (+1). | 0 | 1 |
| Multivariable analysis with non-radiomics features | Study includes multivariable analysis with non-radiomics features, for instance, age and prostate-specific antigen (+1). | 0 | 1 |
| Discrimination statistics | The study reports discrimination statistics (for example, receiver operating curve, AUC) (+1). The study reports the validation method (for example, cross-validation, bootstrapping) and confidence intervals (+1). | 0 | 2 |
| Prospective study | The study has a prospective design (+7). | 0 | 7 |
| Validation | The study does not report a validation method (−5), a validation performed on a dataset from the same institute (+2), a validation on a dataset from another institute (+3), a validation on a dataset from two distinct institutes (+4), a validation on a dataset from three different institutes (+5). *Validation set size should be of comparable size and representative of the training set. | −5 | 5 |
| Comparison to gold standard | The study assesses the extent to which the model agrees with/is superior to the current “gold standard” (+2). | 0 | 2 |
| Report potential clinical utility | The study reports potential clinical utility and potential application of the model in a clinical setting (+2). | 0 | 2 |
| Open science and data | Study scans are open source (+1), region of interest (ROI) segmentations are open source (+1). The classification model with parameter settings is publicly available (+1). | 0 | 3 |
| Score | (number of points × 100)/29 | 0% | 100% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Castillo T., J.M.; Arif, M.; Niessen, W.J.; Schoots, I.G.; Veenland, J.F. Automated Classification of Significant Prostate Cancer on MRI: A Systematic Review on the Performance of Machine Learning Applications. Cancers 2020, 12, 1606. https://doi.org/10.3390/cancers12061606
Castillo T. JM, Arif M, Niessen WJ, Schoots IG, Veenland JF. Automated Classification of Significant Prostate Cancer on MRI: A Systematic Review on the Performance of Machine Learning Applications. Cancers. 2020; 12(6):1606. https://doi.org/10.3390/cancers12061606
Chicago/Turabian StyleCastillo T., Jose M., Muhammad Arif, Wiro J. Niessen, Ivo G. Schoots, and Jifke F. Veenland. 2020. "Automated Classification of Significant Prostate Cancer on MRI: A Systematic Review on the Performance of Machine Learning Applications" Cancers 12, no. 6: 1606. https://doi.org/10.3390/cancers12061606
APA StyleCastillo T., J. M., Arif, M., Niessen, W. J., Schoots, I. G., & Veenland, J. F. (2020). Automated Classification of Significant Prostate Cancer on MRI: A Systematic Review on the Performance of Machine Learning Applications. Cancers, 12(6), 1606. https://doi.org/10.3390/cancers12061606





