Mitochondrial Respiration in KRAS and BRAF Mutated Colorectal Tumors and Polyps
Abstract
1. Introduction
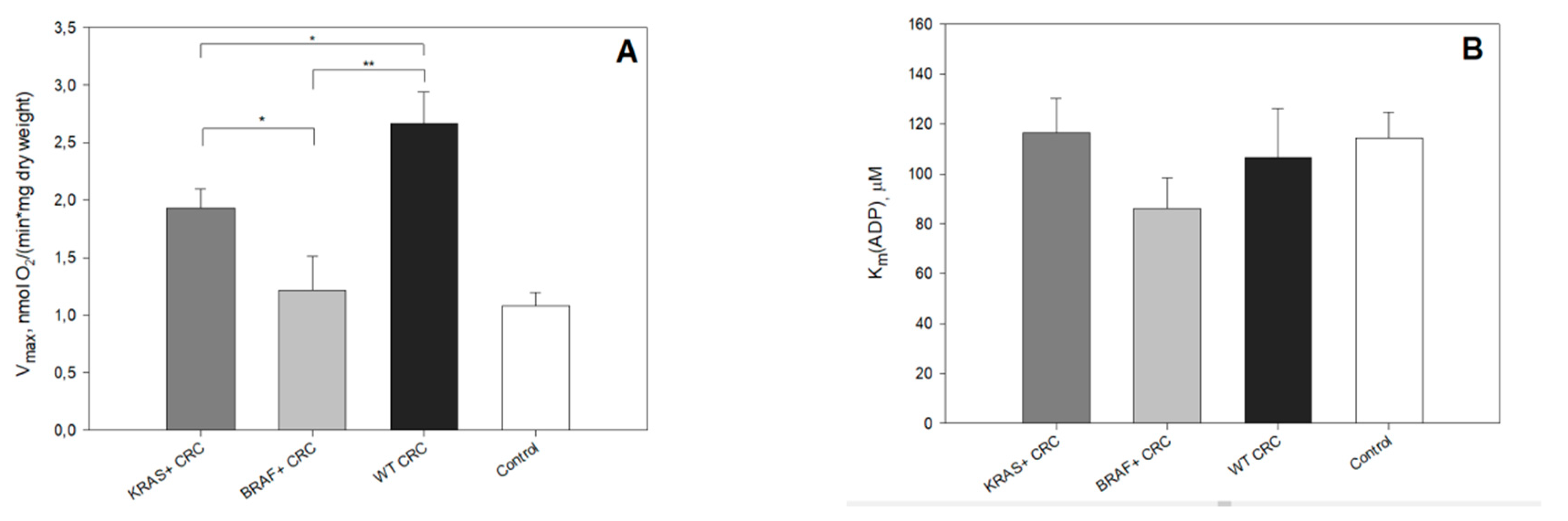
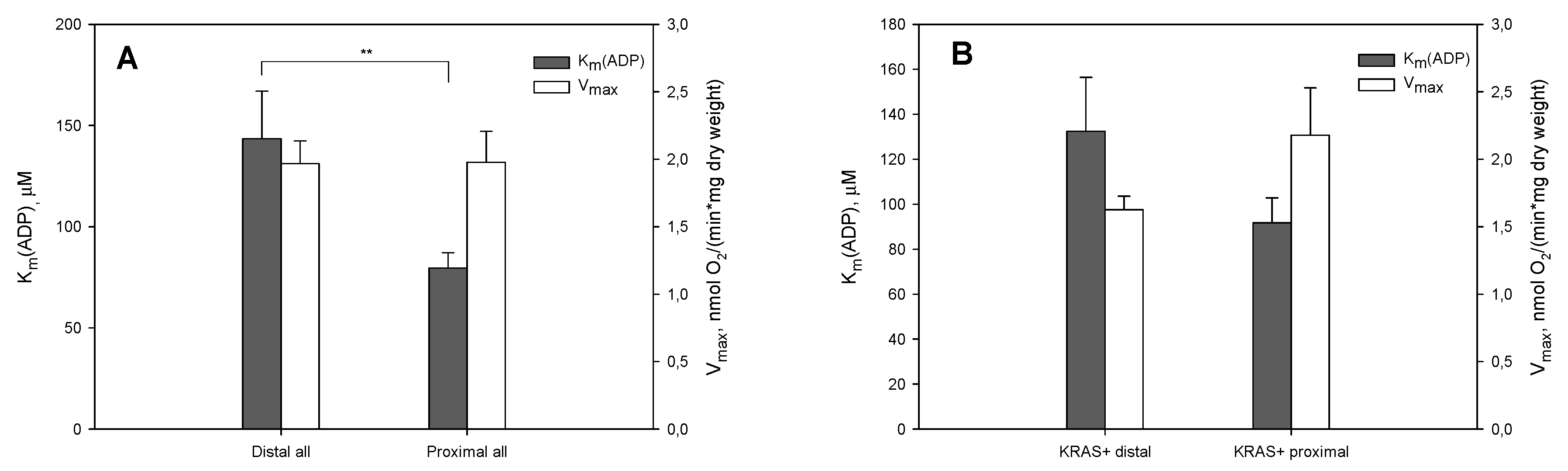
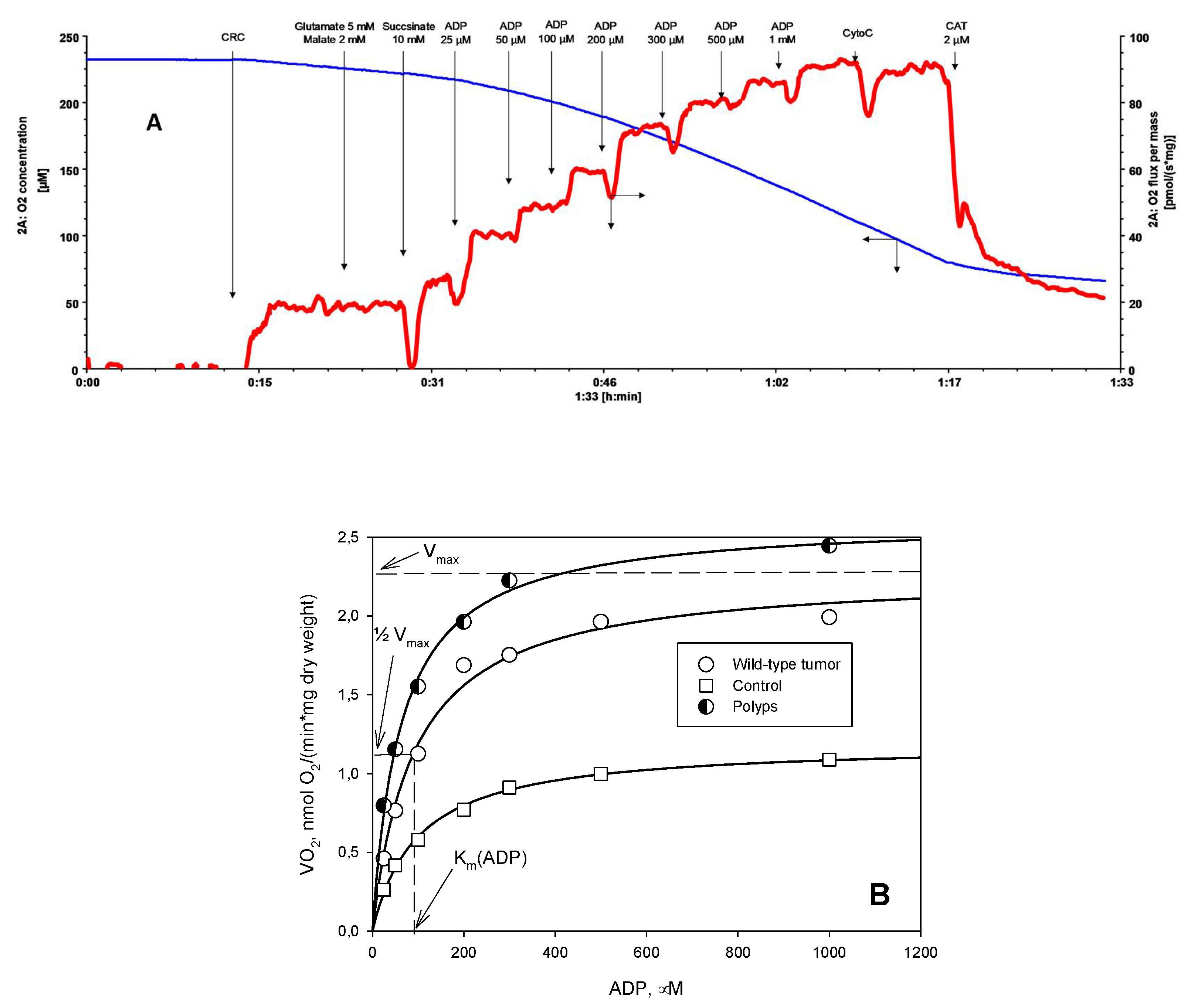
2. Results and Discussion
3. Materials and Methods
3.1. Reagents
3.2. Clinical Material
3.3. Preparation of Skinned Tumor Fibers and Permeabilization Procedure
3.4. Oxygraphic Measurements
3.5. DNA Extraction
3.6. KRAS and BRAF Mutation Analysis
3.7. Data Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ADP | adenosine diphosphate |
| CMS | consensus molecular subtype |
| CRC | colorectal cancer |
| Km | Michaelis–Menten constant |
| Km(ADP) | apparent affinity of mitochondria for exogenous ADP |
| OXPHOS | oxidative phosphorylation |
| MOM | outer mitochondrial membrane |
| VDAC | voltage-dependent anion channel |
| Vmax | maximal-ADP-activated respiration rate |
References
- Vogelstein, B.; Kinzler, K.W. Cancer genes and the pathways they control. Nat. Med. 2004, 10, 789–799. [Google Scholar] [CrossRef]
- Boutin, A.T.; Liao, W.T.; Wang, M.; Hwang, S.S.; Karpinets, T.V.; Cheung, H.; Chu, G.C.; Jiang, S.; Hu, J.; Chang, K.; et al. Oncogenic kras drives invasion and maintains metastases in colorectal cancer. Genes Dev. 2017, 31, 370–382. [Google Scholar] [CrossRef]
- Hao, Y.; Samuels, Y.; Li, Q.; Krokowski, D.; Guan, B.J.; Wang, C.; Jin, Z.; Dong, B.; Cao, B.; Feng, X.; et al. Oncogenic pik3ca mutations reprogram glutamine metabolism in colorectal cancer. Nat. Commun. 2016, 7, 11971. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Schwitalla, S. Tumor cell plasticity: The challenge to catch a moving target. J. Gastroenterol. 2014, 49, 618–627. [Google Scholar] [CrossRef]
- Kaldma, A.; Klepinin, A.; Chekulayev, V.; Mado, K.; Shevchuk, I.; Timohhina, N.; Tepp, K.; Kandashvili, M.; Varikmaa, M.; Koit, A.; et al. An in situ study of bioenergetic properties of human colorectal cancer: The regulation of mitochondrial respiration and distribution of flux control among the components of atp synthasome. Int. J. Biochem. Cell Biol. 2014, 55, 171–186. [Google Scholar] [CrossRef]
- Guinney, J.; Dienstmann, R.; Wang, X.; de Reynies, A.; Schlicker, A.; Soneson, C.; Marisa, L.; Roepman, P.; Nyamundanda, G.; Angelino, P.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350–1356. [Google Scholar] [CrossRef]
- Hutton, J.E.; Wang, X.; Zimmerman, L.J.; Slebos, R.J.; Trenary, I.A.; Young, J.D.; Li, M.; Liebler, D.C. Oncogenic kras and braf drive metabolic reprogramming in colorectal cancer. Mol. Cell. Proteom. 2016, 15, 2924–2938. [Google Scholar] [CrossRef]
- Corazao-Rozas, P.; Guerreschi, P.; Andre, F.; Gabert, P.E.; Lancel, S.; Dekiouk, S.; Fontaine, D.; Tardivel, M.; Savina, A.; Quesnel, B.; et al. Mitochondrial oxidative phosphorylation controls cancer cell’s life and death decisions upon exposure to mapk inhibitors. Oncotarget 2016, 7, 39473–39485. [Google Scholar] [CrossRef]
- Ruocco, M.R.; Avagliano, A.; Granato, G.; Vigliar, E.; Masone, S.; Montagnani, S.; Arcucci, A. Metabolic flexibility in melanoma: A potential therapeutic target. Semin. Cancer Biol. 2019, 59, 187–207. [Google Scholar] [CrossRef]
- Pupo, E.; Avanzato, D.; Middonti, E.; Bussolino, F.; Lanzetti, L. Kras-driven metabolic rewiring reveals novel actionable targets in cancer. Front. Oncol. 2019, 9, 848. [Google Scholar] [CrossRef] [PubMed]
- Yokota, T.; Ura, T.; Shibata, N.; Takahari, D.; Shitara, K.; Nomura, M.; Kondo, C.; Mizota, A.; Utsunomiya, S.; Muro, K.; et al. Braf mutation is a powerful prognostic factor in advanced and recurrent colorectal cancer. Br. J. Cancer 2011, 104, 856–862. [Google Scholar] [CrossRef] [PubMed]
- Phipps, A.I.; Buchanan, D.D.; Makar, K.W.; Win, A.K.; Baron, J.A.; Lindor, N.M.; Potter, J.D.; Newcomb, P.A. Kras-mutation status in relation to colorectal cancer survival: The joint impact of correlated tumour markers. Br. J. Cancer 2013, 108, 1757–1764. [Google Scholar] [CrossRef]
- De Roock, W.; Claes, B.; Bernasconi, D.; De Schutter, J.; Biesmans, B.; Fountzilas, G.; Kalogeras, K.T.; Kotoula, V.; Papamichael, D.; Laurent-Puig, P.; et al. Effects of kras, braf, nras, and pik3ca mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: A retrospective consortium analysis. Lancet Oncol. 2010, 11, 753–762. [Google Scholar] [CrossRef]
- Warburg, O. On the origin of cancer cells. Science 1956, 123, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Paudel, B.B.; Quaranta, V. Metabolic plasticity meets gene regulation. Proc. Natl. Acad. Sci. USA 2019, 116, 3370–3372. [Google Scholar] [CrossRef]
- Jia, D.; Lu, M.; Jung, K.H.; Park, J.H.; Yu, L.; Onuchic, J.N.; Kaipparettu, B.A.; Levine, H. Elucidating cancer metabolic plasticity by coupling gene regulation with metabolic pathways. Proc. Natl. Acad. Sci. USA 2019, 116, 3909–3918. [Google Scholar] [CrossRef]
- Yu, L.; Lu, M.; Jia, D.; Ma, J.; Ben-Jacob, E.; Levine, H.; Kaipparettu, B.A.; Onuchic, J.N. Modeling the genetic regulation of cancer metabolism: Interplay between glycolysis and oxidative phosphorylation. Cancer Res. 2017, 77, 1564–1574. [Google Scholar] [CrossRef]
- Cruz, M.D.; Ledbetter, S.; Chowdhury, S.; Tiwari, A.K.; Momi, N.; Wali, R.K.; Bliss, C.; Huang, C.; Lichtenstein, D.; Bhattacharya, S.; et al. Metabolic reprogramming of the premalignant colonic mucosa is an early event in carcinogenesis. Oncotarget 2017, 8, 20543–20557. [Google Scholar] [CrossRef]
- Grassetto, G.; Capirci, C.; Marzola, M.C.; Rampin, L.; Chondrogiannis, S.; Musto, A.; Crepaldi, G.; Minicozzi, A.M.; Massaro, A.; Rubello, D. Colorectal cancer: Prognostic role of 18f-fdg-pet/ct. Abdom. Imaging 2012, 37, 575–579. [Google Scholar] [CrossRef]
- Graziano, F.; Ruzzo, A.; Giacomini, E.; Ricciardi, T.; Aprile, G.; Loupakis, F.; Lorenzini, P.; Ongaro, E.; Zoratto, F.; Catalano, V.; et al. Glycolysis gene expression analysis and selective metabolic advantage in the clinical progression of colorectal cancer. Pharm. J. 2017, 17, 258–264. [Google Scholar] [CrossRef] [PubMed]
- Chekulayev, V.; Mado, K.; Shevchuk, I.; Koit, A.; Kaldma, A.; Klepinin, A.; Timohhina, N.; Tepp, K.; Kandashvili, M.; Ounpuu, L.; et al. Metabolic remodeling in human colorectal cancer and surrounding tissues: Alterations in regulation of mitochondrial respiration and metabolic fluxes. Biochem. Biophys. Rep. 2015, 4, 111–125. [Google Scholar] [CrossRef]
- Ou, J.; Miao, H.; Ma, Y.; Guo, F.; Deng, J.; Wei, X.; Zhou, J.; Xie, G.; Shi, H.; Xue, B.; et al. Loss of abhd5 promotes colorectal tumor development and progression by inducing aerobic glycolysis and epithelial-mesenchymal transition. Cell Rep. 2014, 9, 1798–1811. [Google Scholar] [CrossRef] [PubMed]
- Ounpuu, L.; Truu, L.; Shevchuk, I.; Chekulayev, V.; Klepinin, A.; Koit, A.; Tepp, K.; Puurand, M.; Rebane-Klemm, E.; Kaambre, T. Comparative analysis of the bioenergetics of human adenocarcinoma caco-2 cell line and postoperative tissue samples from colorectal cancer patients. Biochim. Biol. Cell. 2018, 96, 808–817. [Google Scholar] [CrossRef] [PubMed]
- Koit, A.; Shevchuk, I.; Ounpuu, L.; Klepinin, A.; Chekulayev, V.; Timohhina, N.; Tepp, K.; Puurand, M.; Truu, L.; Heck, K.; et al. Mitochondrial respiration in human colorectal and breast cancer clinical material is regulated differently. Oxid. Med. Cell. Longev. 2017, 2017, 1372640. [Google Scholar] [CrossRef]
- Colombini, M. The vdac channel: Molecular basis for selectivity. Biochim. Biophys. Acta (BBA) Mol. Cell Res. 2016, 1863, 2498–2502. [Google Scholar] [CrossRef]
- Noskov, S.Y.; Rostovtseva, T.K.; Chamberlin, A.C.; Teijido, O.; Jiang, W.; Bezrukov, S.M. Current state of theoretical and experimental studies of the voltage-dependent anion channel (vdac). Biochim. Biophys. Acta 2016, 1858, 1778–1790. [Google Scholar] [CrossRef]
- Klepinin, A.; Ounpuu, L.; Mado, K.; Truu, L.; Chekulayev, V.; Puurand, M.; Shevchuk, I.; Tepp, K.; Planken, A.; Kaambre, T. The complexity of mitochondrial outer membrane permeability and vdac regulation by associated proteins. J. Bioenerg. Biomembr. 2018, 50, 339–354. [Google Scholar] [CrossRef]
- Puurand, M.; Tepp, K.; Timohhina, N.; Aid, J.; Shevchuk, I.; Chekulayev, V.; Kaambre, T. Tubulin betaii and betaiii isoforms as the regulators of vdac channel permeability in health and disease. Cells 2019, 8, 239. [Google Scholar] [CrossRef]
- Maldonado, E.N.; Sheldon, K.L.; DeHart, D.N.; Patnaik, J.; Manevich, Y.; Townsend, D.M.; Bezrukov, S.M.; Rostovtseva, T.K.; Lemasters, J.J. Voltage-dependent anion channels modulate mitochondrial metabolism in cancer cells: Regulation by free tubulin and erastin. J. Biol. Chem. 2013, 288, 11920–11929. [Google Scholar] [CrossRef]
- Varikmaa, M.; Bagur, R.; Kaambre, T.; Grichine, A.; Timohhina, N.; Tepp, K.; Shevchuk, I.; Chekulayev, V.; Metsis, M.; Boucher, F.; et al. Role of mitochondria-cytoskeleton interactions in respiration regulation and mitochondrial organization in striated muscles. Biochim. Biophys. Acta 2014, 1837, 232–245. [Google Scholar] [CrossRef] [PubMed]
- Anmann, T.; Guzun, R.; Beraud, N.; Pelloux, S.; Kuznetsov, A.V.; Kogerman, L.; Kaambre, T.; Sikk, P.; Paju, K.; Peet, N.; et al. Different kinetics of the regulation of respiration in permeabilized cardiomyocytes and in hl-1 cardiac cells. Importance of cell structure/organization for respiration regulation. Biochim. Biophys. Acta 2006, 1757, 1597–1606. [Google Scholar] [CrossRef] [PubMed]
- Monge, C.; Beraud, N.; Tepp, K.; Pelloux, S.; Chahboun, S.; Kaambre, T.; Kadaja, L.; Roosimaa, M.; Piirsoo, A.; Tourneur, Y.; et al. Comparative analysis of the bioenergetics of adult cardiomyocytes and nonbeating hl-1 cells: Respiratory chain activities, glycolytic enzyme profiles, and metabolic fluxes. Can. J. Physiol. Pharmacol. 2009, 87, 318–326. [Google Scholar] [CrossRef] [PubMed]
- Klepinin, A.; Chekulayev, V.; Timohhina, N.; Shevchuk, I.; Tepp, K.; Kaldma, A.; Koit, A.; Saks, V.; Kaambre, T. Comparative analysis of some aspects of mitochondrial metabolism in differentiated and undifferentiated neuroblastoma cells. J. Bioenerg. Biomembr. 2014, 46, 17–31. [Google Scholar] [CrossRef]
- Kawada, K.; Toda, K.; Sakai, Y. Targeting metabolic reprogramming in kras-driven cancers. Int. J. Clin. Oncol. 2017, 22, 651–659. [Google Scholar] [CrossRef]
- Oikonomou, E.; Koustas, E.; Goulielmaki, M.; Pintzas, A. Braf vs ras oncogenes: Are mutations of the same pathway equal? Differential signalling and therapeutic implications. Oncotarget 2014, 5, 11752–11777. [Google Scholar] [CrossRef]
- Vaughn, C.P.; Zobell, S.D.; Furtado, L.V.; Baker, C.L.; Samowitz, W.S. Frequency of kras, braf, and nras mutations in colorectal cancer. Genes Chromosomes Cancer 2011, 50, 307–312. [Google Scholar] [CrossRef]
- Lai, E.; Pretta, A.; Impera, V.; Mariani, S.; Giampieri, R.; Casula, L.; Pusceddu, V.; Coni, P.; Fanni, D.; Puzzoni, M.; et al. Braf-mutant colorectal cancer, a different breed evolving. Expert Rev. Mol. Diagn. 2018, 18, 499–512. [Google Scholar] [CrossRef]
- Kaambre, T.; Chekulayev, V.; Shevchuk, I.; Karu-Varikmaa, M.; Timohhina, N.; Tepp, K.; Bogovskaja, J.; Kutner, R.; Valvere, V.; Saks, V. Metabolic control analysis of cellular respiration in situ in intraoperational samples of human breast cancer. J. Bioenerg. Biomembr. 2012, 44, 539–558. [Google Scholar] [CrossRef]
- Kaambre, T.; Chekulayev, V.; Shevchuk, I.; Tepp, K.; Timohhina, N.; Varikmaa, M.; Bagur, R.; Klepinin, A.; Anmann, T.; Koit, A.; et al. Metabolic control analysis of respiration in human cancer tissue. Front. Physiol. 2013, 4, 151. [Google Scholar] [CrossRef]
- Yun, J.; Rago, C.; Cheong, I.; Pagliarini, R.; Angenendt, P.; Rajagopalan, H.; Schmidt, K.; Willson, J.K.; Markowitz, S.; Zhou, S.; et al. Glucose deprivation contributes to the development of kras pathway mutations in tumor cells. Science 2009, 325, 1555–1559. [Google Scholar] [CrossRef]
- Hubackova, S.; Magalhaes Novais, S.; Davidova, E.; Neuzil, J.; Rohlena, J. Mitochondria-driven elimination of cancer and senescent cells. Biol. Chem. 2019, 400, 141–148. [Google Scholar] [CrossRef]
- Lu, J.; Tan, M.; Cai, Q. The warburg effect in tumor progression: Mitochondrial oxidative metabolism as an anti-metastasis mechanism. Cancer Lett. 2015, 356, 156–164. [Google Scholar] [CrossRef]
- Gnaiger, E.; Kemp, R.B. Anaerobic metabolism in aerobic mammalian cells: Information from the ratio of calorimetric heat flux and respirometric oxygen flux. Biochim. Biophys. Acta 1990, 1016, 328–332. [Google Scholar] [CrossRef]
- Gstraunthaler, G.; Seppi, T.; Pfaller, W. Impact of culture conditions, culture media volumes, and glucose content on metabolic properties of renal epithelial cell cultures. Are renal cells in tissue culture hypoxic? Cell Physiol. Biochem. 1999, 9, 150–172. [Google Scholar] [CrossRef]
- Sherr, C.J.; DePinho, R.A. Cellular senescence: Mitotic clock or culture shock? Cell 2000, 102, 407–410. [Google Scholar] [CrossRef]
- Jose, C.; Rossignol, R. Rationale for mitochondria-targeting strategies in cancer bioenergetic therapies. Int. J. Biochem. Cell Biol. 2013, 45, 123–129. [Google Scholar] [CrossRef]
- Swerdlow, R.H.; Lezi, E.; Aires, D.; Lu, J. Glycolysis-respiration relationships in a neuroblastoma cell line. Biochim. Biophys. Acta 2013, 1830, 2891–2898. [Google Scholar] [CrossRef]
- Chance, B.; Williams, G.R. Respiratory enzymes in oxidative phosphorylation. Ii. Difference spectra. J. Biol. Chem. 1955, 217, 395–407. [Google Scholar]
- Chance, B.; Williams, G.R. Respiratory enzymes in oxidative phosphorylation. Vi. The effects of adenosine diphosphate on azide-treated mitochondria. J. Biol. Chem. 1956, 221, 477–489. [Google Scholar]
- Saks, V.A.; Veksler, V.I.; Kuznetsov, A.V.; Kay, L.; Sikk, P.; Tiivel, T.; Tranqui, L.; Olivares, J.; Winkler, K.; Wiedemann, F.; et al. Permeabilized cell and skinned fiber techniques in studies of mitochondrial function in vivo. Mol. Cell. Biochem. 1998, 184, 81–100. [Google Scholar] [CrossRef]
- Mathupala, S.P.; Ko, Y.H.; Pedersen, P.L. Hexokinase-2 bound to mitochondria: Cancer’s stygian link to the “warburg effect” and a pivotal target for effective therapy. Semin. Cancer Biol. 2009, 19, 17–24. [Google Scholar] [CrossRef]
- Pedersen, P.L. Warburg, me and hexokinase 2: Multiple discoveries of key molecular events underlying one of cancers’ most common phenotypes, the “warburg effect”, i.E., elevated glycolysis in the presence of oxygen. J. Bioenerg. Biomembr. 2007, 39, 211–222. [Google Scholar] [CrossRef]
- Maldonado, E.N.; Patnaik, J.; Mullins, M.R.; Lemasters, J.J. Free tubulin modulates mitochondrial membrane potential in cancer cells. Cancer Res. 2010, 70, 10192–10201. [Google Scholar] [CrossRef]
- Draberova, E.; Sulimenko, V.; Vinopal, S.; Sulimenko, T.; Sladkova, V.; D’Agostino, L.; Sobol, M.; Hozak, P.; Kren, L.; Katsetos, C.D.; et al. Differential expression of human gamma-tubulin isotypes during neuronal development and oxidative stress points to a gamma-tubulin-2 prosurvival function. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2017, 31, 1828–1846. [Google Scholar] [CrossRef]
- Lindstrom, L.; Li, T.; Malycheva, D.; Kancharla, A.; Nilsson, H.; Vishnu, N.; Mulder, H.; Johansson, M.; Rossello, C.A.; Alvarado-Kristensson, M. The gtpase domain of gamma-tubulin is required for normal mitochondrial function and spatial organization. Commun. Biol. 2018, 1, 37. [Google Scholar] [CrossRef]
- Shoshan-Barmatz, V.; Krelin, Y.; Shteinfer-Kuzmine, A.; Arif, T. Voltage-dependent anion channel 1 as an emerging drug target for novel anti-cancer therapeutics. Front. Oncol. 2017, 7, 154. [Google Scholar] [CrossRef]
- Shoshan-Barmatz, V.; Maldonado, E.N.; Krelin, Y. Vdac1 at the crossroads of cell metabolism, apoptosis and cell stress. Cell Stress 2017, 1, 11–36. [Google Scholar] [CrossRef]
- Missiaglia, E.; Jacobs, B.; D’Ario, G.; Di Narzo, A.F.; Soneson, C.; Budinska, E.; Popovici, V.; Vecchione, L.; Gerster, S.; Yan, P.; et al. Distal and proximal colon cancers differ in terms of molecular, pathological, and clinical features. Ann. Oncol. 2014, 25, 1995–2001. [Google Scholar] [CrossRef]
- Drewes, J.L.; Housseau, F.; Sears, C.L. Sporadic colorectal cancer: Microbial contributors to disease prevention, development and therapy. Br. J. Cancer 2016, 115, 273–280. [Google Scholar] [CrossRef]
- De Palma, F.D.E.; D’Argenio, V.; Pol, J.; Kroemer, G.; Maiuri, M.C.; Salvatore, F. The molecular hallmarks of the serrated pathway in colorectal cancer. Cancers (Basel) 2019, 11, 1017. [Google Scholar] [CrossRef]
- Fang, S.; Fang, X. Advances in glucose metabolism research in colorectal cancer. Biomed. Rep. 2016, 5, 289–295. [Google Scholar] [CrossRef]
- Iwamoto, M.; Kawada, K.; Nakamoto, Y.; Itatani, Y.; Inamoto, S.; Toda, K.; Kimura, H.; Sasazuki, T.; Shirasawa, S.; Okuyama, H.; et al. Regulation of 18f-fdg accumulation in colorectal cancer cells with mutated kras. J. Nucl. Med. 2014, 55, 2038–2044. [Google Scholar] [CrossRef]
- Wang, P.; Song, M.; Zeng, Z.L.; Zhu, C.F.; Lu, W.H.; Yang, J.; Ma, M.Z.; Huang, A.M.; Hu, Y.; Huang, P. Identification of ndufaf1 in mediating k-ras induced mitochondrial dysfunction by a proteomic screening approach. Oncotarget 2015, 6, 3947–3962. [Google Scholar] [CrossRef]
- Ralph, S.J.; Low, P.; Dong, L.; Lawen, A.; Neuzil, J. Mitocans: Mitochondrial targeted anti-cancer drugs as improved therapies and related patent documents. Recent Pat. Anticancer Drug Discov. 2006, 1, 327–346. [Google Scholar] [CrossRef]
- Kuznetsov, A.V.; Veksler, V.; Gellerich, F.N.; Saks, V.; Margreiter, R.; Kunz, W.S. Analysis of mitochondrial function in situ in permeabilized muscle fibers, tissues and cells. Nat. Protoc. 2008, 3, 965–976. [Google Scholar] [CrossRef]
- Kuznetsov, A.V.; Tiivel, T.; Sikk, P.; Kaambre, T.; Kay, L.; Daneshrad, Z.; Rossi, A.; Kadaja, L.; Peet, N.; Seppet, E.; et al. Striking differences between the kinetics of regulation of respiration by adp in slow-twitch and fast-twitch muscles in vivo. Eur. J. Biochem. 1996, 241, 909–915. [Google Scholar] [CrossRef]
- Gnaiger, E. Oxygen solubility in experimental media. OROBOROS Bioenerg. News 2001, 6, 1–6. [Google Scholar]
- Puurand, M.; Tepp, K.; Klepinin, A.; Klepinina, L.; Shevchuk, I.; Kaambre, T. Intracellular energy-transfer networks and high-resolution respirometry: A convenient approach for studying their function. Int. J. Mol. Sci. 2018, 19, 2933. [Google Scholar] [CrossRef]
- Timohhina, N.; Guzun, R.; Tepp, K.; Monge, C.; Varikmaa, M.; Vija, H.; Sikk, P.; Kaambre, T.; Sackett, D.; Saks, V. Direct measurement of energy fluxes from mitochondria into cytoplasm in permeabilized cardiac cells in situ: Some evidence for mitochondrial interactosome. J. Bioenerg. Biomembr. 2009, 41, 259–275. [Google Scholar] [CrossRef]
| Sample | % of Low Oxidative Capacity of Mitochondrion |
|---|---|
| KRAS tumors | 28.1 |
| KRAS polyps | 65.9 |
| BRAF tumors | 43.0 |
| BRAF polyps | 68.6 |
| Wild-type tumors | 32.4 |
| Wild-type polyps | 61.7 |
| All controls | 29.0 |
| Characteristics | n |
|---|---|
| Total patients | 48 |
| Females | 19 |
| Males | 29 |
| Age at diagnosis | |
| Mean | 72 |
| Median | 74 |
| Range | 38–91 |
| Stage of tumor | |
| I-II | 15 |
| III-IV | 9 |
| Unknown | 9 |
| Molecular subtype of tumor | |
| KRAS mutated | 13 |
| BRAF mutated | 6 |
| KRAS and BRAF wild-type | 14 |
| Molecular subtypes of polyps | |
| KRAS mutated | 4 |
| BRAF mutated | 2 |
| KRAS and BRAF wild-type | 9 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rebane-Klemm, E.; Truu, L.; Reinsalu, L.; Puurand, M.; Shevchuk, I.; Chekulayev, V.; Timohhina, N.; Tepp, K.; Bogovskaja, J.; Afanasjev, V.; et al. Mitochondrial Respiration in KRAS and BRAF Mutated Colorectal Tumors and Polyps. Cancers 2020, 12, 815. https://doi.org/10.3390/cancers12040815
Rebane-Klemm E, Truu L, Reinsalu L, Puurand M, Shevchuk I, Chekulayev V, Timohhina N, Tepp K, Bogovskaja J, Afanasjev V, et al. Mitochondrial Respiration in KRAS and BRAF Mutated Colorectal Tumors and Polyps. Cancers. 2020; 12(4):815. https://doi.org/10.3390/cancers12040815
Chicago/Turabian StyleRebane-Klemm, Egle, Laura Truu, Leenu Reinsalu, Marju Puurand, Igor Shevchuk, Vladimir Chekulayev, Natalja Timohhina, Kersti Tepp, Jelena Bogovskaja, Vladimir Afanasjev, and et al. 2020. "Mitochondrial Respiration in KRAS and BRAF Mutated Colorectal Tumors and Polyps" Cancers 12, no. 4: 815. https://doi.org/10.3390/cancers12040815
APA StyleRebane-Klemm, E., Truu, L., Reinsalu, L., Puurand, M., Shevchuk, I., Chekulayev, V., Timohhina, N., Tepp, K., Bogovskaja, J., Afanasjev, V., Suurmaa, K., Valvere, V., & Kaambre, T. (2020). Mitochondrial Respiration in KRAS and BRAF Mutated Colorectal Tumors and Polyps. Cancers, 12(4), 815. https://doi.org/10.3390/cancers12040815





