NRF2 Regulation by Noncoding RNAs in Cancers: The Present Knowledge and the Way Forward
Abstract
Simple Summary
Abstract
1. Introduction
2. Brief Overview of the NRF2 Signaling Pathway: Negative and Positive Regulators and ARE Genes’ Targets
3. NRF2 Deregulation in Cancer: Focus on the Epigenetic Modifications
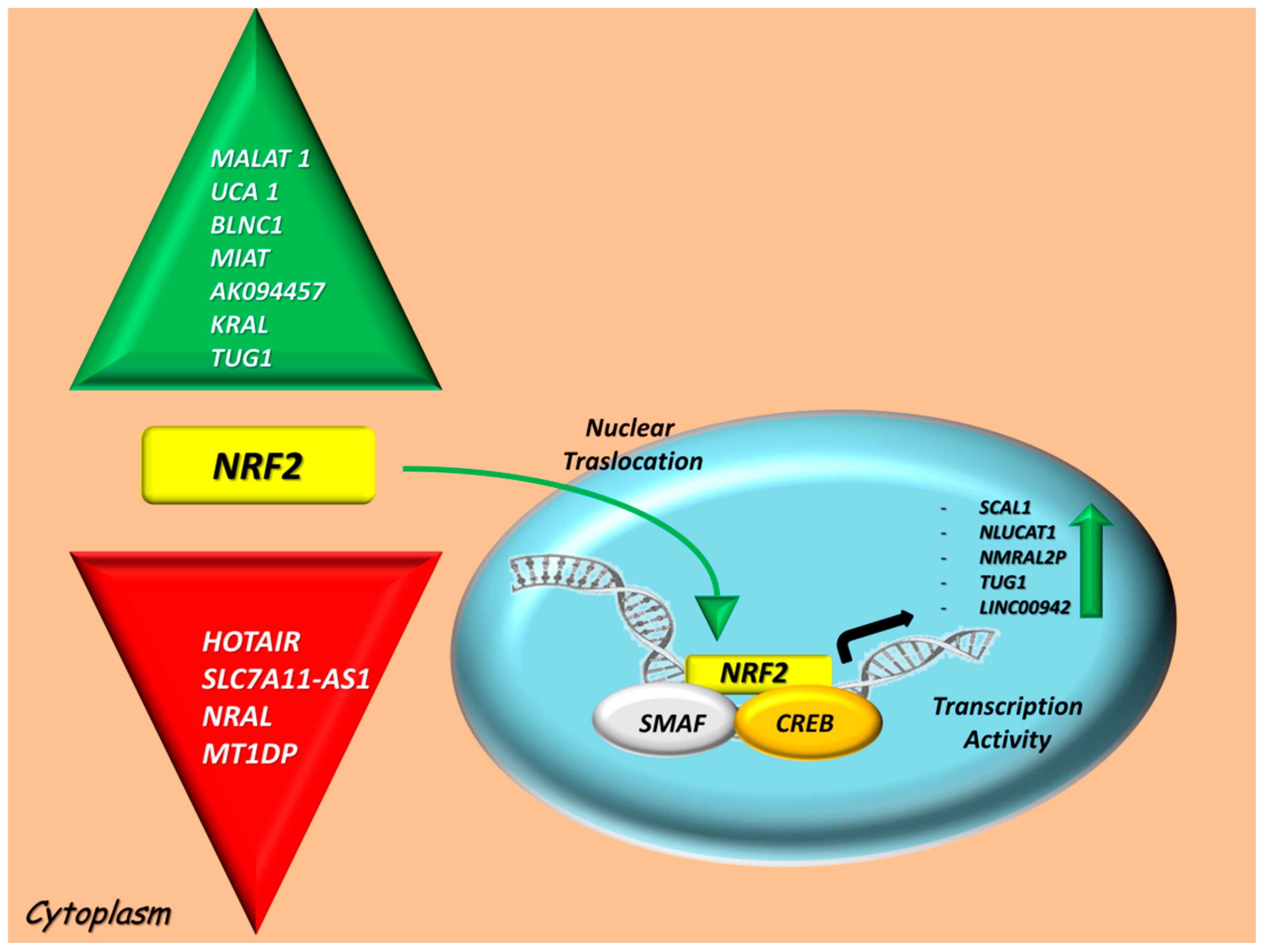
4. LncRNAs Intercepting the NRF2 Axis
4.1. LncRNAs Modulated by NRF2 Signaling
4.1.1. SCAL1 (LUCAT1)
4.1.2. NLUCAT1
4.1.3. NMRAL2P
4.1.4. TUG1
4.1.5. LINC00942
4.2. Positive and Negative LncRNA Regulators of the NRF2 Activity
4.2.1. HOTAIR
4.2.2. MALAT1
4.2.3. UCA1
4.2.4. BLNC1
4.2.5. MIAT and AK094457
4.2.6. SLC7A11-AS1
4.2.7. KRAL and NRAL
4.2.8. MT1DP
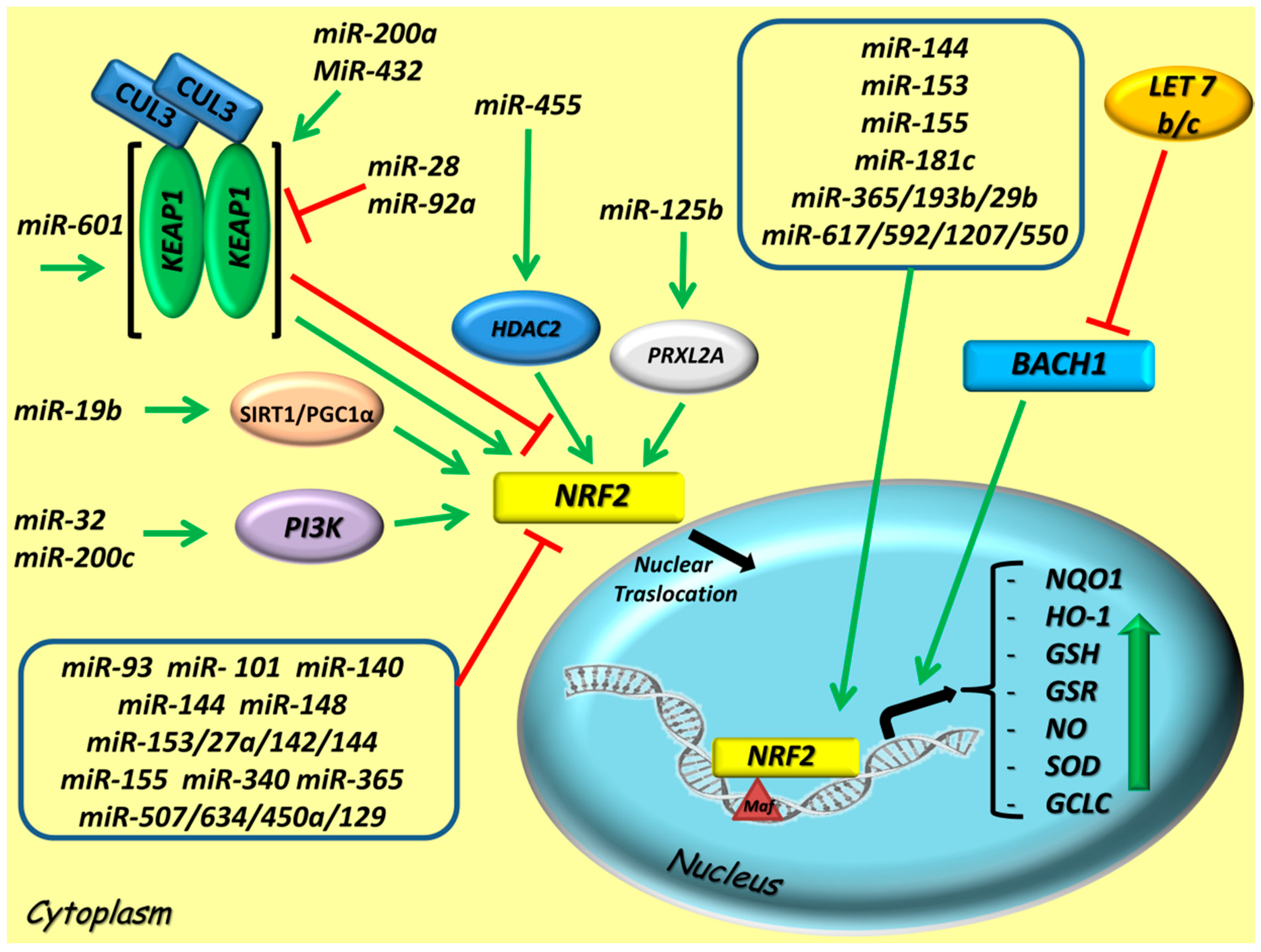
5. miRNAs Intercepting the NRF2 Axis
5.1. Positive miRNA Regulators of NRF2 Activity
5.2. Negative miRNA Regulators of NRF2 Activity
6. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jaramillo, M.C.; Zhang, D.D. The emerging role of the Nrf2-Keap1 signaling pathway in cancer. Genes Dev. 2013, 27, 2179–2191. [Google Scholar] [CrossRef] [PubMed]
- Tonelli, C.; Chio, I.I.C.; Tuveson, D.A. Transcriptional regulation by Nrf2. Antioxid. Redox Signal. 2018, 29, 1727–1745. [Google Scholar] [CrossRef] [PubMed]
- Levings, D.C.; Wang, X.; Kohlhase, D.; Bell, D.A.; Slattery, M. A distinct class of antioxidant response elements is consistently activated in tumors with NRF2 mutations. Redox Biol. 2018, 19, 235–249. [Google Scholar] [CrossRef] [PubMed]
- Kaspar, J.W.; Jaiswal, A.K. An autoregulatory loop between Nrf2 and Cul3-Rbx1 controls their cellular abundance. J. Biol. Chem. 2010, 285, 21349–21358. [Google Scholar] [CrossRef]
- Li, W.; Yu, S.; Liu, T.; Kim, J.H.; Blank, V.; Li, H.; Kong, A.N. Heterodimerization with small Maf proteins enhances nuclear retention of Nrf2 via masking the NESzip motif. Biochim. Biophys. Acta 2008, 1783, 1847–1856. [Google Scholar] [CrossRef]
- Otsuki, A.; Yamamoto, M. Cis-element architecture of Nrf2-sMaf heterodimer binding sites and its relation to diseases. Arch. Pharm. Res. 2020, 43, 275–285. [Google Scholar] [CrossRef]
- Yamamoto, T.; Suzuki, T.; Kobayashi, A.; Wakabayashi, J.; Maher, J.; Motohashi, H.; Yamamoto, M. Physiological significance of reactive cysteine residues of Keap1 in determining Nrf2 activity. Mol. Cell. Biol. 2008, 28, 2758–2770. [Google Scholar] [CrossRef]
- Canning, P.; Sorrell, F.J.; Bullock, A.N. Structural basis of Keap1 interactions with Nrf2. Free Radic. Biol. Med. 2015, 88, 101–107. [Google Scholar] [CrossRef]
- Villeneuve, N.F.; Lau, A.; Zhang, D.D. Regulation of the Nrf2-Keap1 antioxidant response by the ubiquitin proteasome system: An insight into cullin-ring ubiquitin ligases. Antioxid. Redox Signal. 2010, 13, 1699–1712. [Google Scholar] [CrossRef]
- McMahon, M.; Itoh, K.; Yamamoto, M.; Hayes, J.D. Keap1-dependent proteasomal degradation of transcription factor Nrf2 contributes to the negative regulation of antioxidant response element-driven gene expression. J. Biol. Chem. 2003, 278, 21592–21600. [Google Scholar] [CrossRef]
- Bryan, H.K.; Olayanju, A.; Goldring, C.E.; Park, B.K. The Nrf2 cell defence pathway: Keap1-dependent and -independent mechanisms of regulation. Biochem. Pharmacol. 2013, 85, 705–717. [Google Scholar] [CrossRef] [PubMed]
- de la Rojo Vega, M.; Chapman, E.; Zhang, D.D. NRF2 and the Hallmarks of Cancer. Cancer Cell. 2018, 34, 21–43. [Google Scholar] [CrossRef] [PubMed]
- Menegon, S.; Columbano, A.; Giordano, S. The dual roles of NRF2 in cancer. Trends Mol. Med. 2016, 22, 578–593. [Google Scholar] [CrossRef] [PubMed]
- Fabrizio, F.P.; Sparaneo, A.; Trombetta, D.; Muscarella, L.A. Epigenetic versus genetic deregulation of the KEAP1/NRF2 axis in solid tumors: Focus on methylation and noncoding RNAs. Oxid. Med. Cell. Longev. 2018, 2018, 2492063. [Google Scholar] [CrossRef] [PubMed]
- Cloer, E.W.; Goldfarb, D.; Schrank, T.P.; Weissman, B.E.; Major, M.B. NRF2 activation in cancer: From DNA to protein. Cancer Res. 2019, 79, 889–898. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, J.; Acuña, S.M.; Aoki, J.I.; Floeter-Winter, L.M.; Muxel, S.M. Long non-coding RNAs in the regulation of gene expression: Physiology and disease. Noncoding RNA 2019, 5, 17. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Lu, H.; Bai, Y. Nrf2 in cancers: A double-edged sword. Cancer Med. 2019, 8, 2252–2267. [Google Scholar] [CrossRef]
- Moi, P.; Chan, K.; Asunis, I.; Cao, A.; Kan, Y.W. Isolation of NF-E2-related factor 2 (Nrf2), a NF-E2-like basic leucine zipper transcriptional activator that binds to the tandem NF-E2/AP1 repeat of the beta-globin locus control region. Proc. Natl. Acad. Sci. USA 1994, 91, 9926–9930. [Google Scholar] [CrossRef]
- Ma, Q. Role of nrf2 in oxidative stress and toxicity. Annu. Rev. Pharmacol. Toxicol. 2013, 53, 401–426. [Google Scholar] [CrossRef]
- Kobayashi, A.; Kang, M.I.; Okawa, H.; Ohtsuji, M.; Zenke, Y.; Chiba, T.; Igarashi, K.; Yamamoto, M. Oxidative stress sensor Keap1 functions as an adaptor for Cul3-based E3 ligase to regulate proteasomal degradation of Nrf2. Mol. Cell. Biol. 2004, 24, 7130–7139. [Google Scholar] [CrossRef]
- Baird, L.; Yamamoto, M. The molecular mechanisms regulating the KEAP1-NRF2 pathway. Mol. Cell. Biol. 2020, 40, e00099-20. [Google Scholar] [CrossRef] [PubMed]
- Hayes, J.D. Regulation of the CNC-bZIP transcription factor Nrf2 by Keap1 and the axis between GSK-3 and β-TrCP. Curr. Opin. Toxicol. 2016, 1, 92–103. [Google Scholar] [CrossRef]
- Zhang, D.D.; Lo, S.C.; Cross, J.V.; Templeton, D.J.; Hannink, M. Keap1 is a redox-regulated substrate adaptor protein for a Cul3-dependent ubiquitin ligase complex. Mol. Cell. Biol. 2004, 24, 10941–10953. [Google Scholar] [CrossRef] [PubMed]
- Tebay, L.E.; Robertson, H.; Durant, S.T.; Vitale, S.R.; Penning, T.M.; Dinkova-Kostova, A.T.; Hayes, J.D. Mechanisms of activation of the transcription factor Nrf2 by redox stressors, nutrient cues, and energy status and the pathways through which it attenuates degenerative disease. Free Radic. Biol. Med. 2015, 88, 108–146. [Google Scholar] [CrossRef]
- Hayes, J.D.; Chowdhry, S.; Dinkova-Kostova, A.T.; Sutherland, C. Dual regulation of transcription factor Nrf2 by Keap1 and by the combined actions of beta-TrCP and GSK-3. Biochem. Soc. Trans. 2015, 43, 611–620. [Google Scholar] [CrossRef]
- Chen, W.; Sun, Z.; Wang, X.J.; Jiang, T.; Huang, Z.; Fang, D.; Zhang, D.D. Direct interaction between Nrf2 and p21(Cip1/WAF1) upregulates the Nrf2-mediated antioxidant response. Mol. Cell 2009, 34, 663–673. [Google Scholar] [CrossRef]
- Gorrini, C.; Baniasadi, P.S.; Harris, I.S.; Silvester, J.; Inoue, S.; Snow, B.; Joshi, P.A.; Wakeham, A.; Molyneux, S.D.; Martin, B.; et al. BRCA1 interacts with Nrf2 to regulate antioxidant signaling and cell survival. J. Exp. Med. 2013, 210, 1529–1544. [Google Scholar] [CrossRef]
- Ichimura, Y.; Komatsu, M. Activation of p62/SQSTM1-Keap1-nuclear factor erythroid 2-related factor 2 pathway in cancer. Front. Oncol. 2018, 8, 210. [Google Scholar] [CrossRef]
- Jain, A.; Lamark, T.; Sjøttem, E.; Larsen, K.B.; Awuh, J.A.; Øvervatn, A.; McMahon, M.; Hayes, J.D.; Johansen, T. p62/SQSTM1 is a target gene for transcription factor NRF2 and creates a positive feedback loop by inducing antioxidant response element-driven gene transcription. J. Biol. Chem. 2010, 285, 22576–22591. [Google Scholar] [CrossRef]
- Perdigoto, C.N. Epigenetic cancer evolution, one cell at a time. Nat. Rev. Genet. 2019, 20, 434–435. [Google Scholar] [CrossRef]
- Arbour, K.C.; Jordan, E.; Kim, H.R.; Dienstag, J.; Yu, H.A.; Sanchez-Vega, F.; Lito, P.; Berger, M.; Solit, D.B.; Hellmann, M.; et al. Effects of co-occurring genomic alterations on outcomes in patients with KRAS-mutant non-small cell lung cancer. Clin. Cancer Res. 2018, 24, 334–340. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Kim, J.H.; Ko, E.; Kim, J.Y.; Park, M.J.; Kim, M.J.; Seo, H.; Li, S.; Lee, J.Y. Resistance to gefitinib and cross-resistance to irreversible EGFR-TKIs mediated by disruption of the Keap1-Nrf2 pathway in human lung cancer cells. FASEB J. 2018, 32, 5862–5873. [Google Scholar] [CrossRef] [PubMed]
- Frank, R.; Scheffler, M.; Merkelbach-Bruse, S.; Ihle, M.A.; Kron, A.; Rauer, M.; Ueckeroth, F.; König, K.; Michels, S.; Fischer, R.; et al. Clinical and pathological characteristics of KEAP1- and NFE2L2-mutated Non-Small Cell Lung Carcinoma (NSCLC). Clin. Cancer Res. 2018, 24, 3087–3096. [Google Scholar] [CrossRef] [PubMed]
- Hellyer, J.A.; Stehr, H.; Das, M.; Padda, S.K.; Ramchandran, K.; Neal, J.W.; Diehn, M.; Wakelee, H.A. Impact of KEAP1/NFE2L2/CUL3 mutations on duration of response to EGFR tyrosine kinase inhibitors in EGFR mutated non-small cell lung cancer. Lung Cancer 2019, 134, 42–45. [Google Scholar] [CrossRef]
- Riely GJ, J.E.; Kim, H.R.; Yu, H.A.; Berger, M.F.; Solit, D.B. Association of outcomes and co-occurring genomic alterations in patients with KRAS-mutant non-small cell lung cancer. J. Clin. Oncol. 2016, 34, 90192016. [Google Scholar] [CrossRef]
- Papagiannakopoulos, T.; Warren, L. The pleiotropic role of the KEAP1/NRF2 pathway in cancer. Annu. Rev. Cancer Biol. 2020, 4, 413–435. [Google Scholar]
- Singh, A.; Misra, V.; Thimmulappa, R.K.; Lee, H.; Ames, S.; Hoque, M.O.; Herman, J.G.; Baylin, S.B.; Sidransky, D.; Gabrielson, E.; et al. Dysfunctional KEAP1-NRF2 interaction in non-small-cell lung cancer. PLoS Med. 2006, 3, e420. [Google Scholar] [CrossRef]
- Ciriello, G.; Miller, M.L.; Aksoy, B.A.; Senbabaoglu, Y.; Schultz, N.; Sander, C. Emerging landscape of oncogenic signatures across human cancers. Nat. Genet. 2013, 45, 1127–1133. [Google Scholar] [CrossRef]
- Weinstein, J.N.; Collisson, E.A.; Mills, G.B.; Shaw, K.R.; Ozenberger, B.A.; Ellrott, K.; Shmulevich, I.; Sander, C.; Stuart, J.M. The cancer genome atlas pan-cancer analysis project. Nat. Genet. 2013, 45, 1113–1120. [Google Scholar] [CrossRef]
- Shibata, T.; Ohta, T.; Tong, K.I.; Kokubu, A.; Odogawa, R.; Tsuta, K.; Asamura, H.; Yamamoto, M.; Hirohashi, S. Cancer related mutations in NRF2 impair its recognition by Keap1-Cul3 E3 ligase and promote malignancy. Proc. Natl. Acad. Sci. USA 2008, 105, 13568–13573. [Google Scholar] [CrossRef]
- Shibata, T.; Kokubu, A.; Saito, S.; Narisawa-Saito, M.; Sasaki, H.; Aoyagi, K.; Yoshimatsu, Y.; Tachimori, Y.; Kushima, R.; Kiyono, T.; et al. NRF2 mutation confers malignant potential and resistance to chemoradiation therapy in advanced esophageal squamous cancer. Neoplasia 2011, 13, 864–873. [Google Scholar] [CrossRef] [PubMed]
- Guichard, C.; Amaddeo, G.; Imbeaud, S.; Ladeiro, Y.; Pelletier, L.; Maad, I.B.; Calderaro, J.; Bioulac-Sage, P.; Letexier, M.; Degos, F. Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nat. Genet. 2012, 44, 694–698. [Google Scholar] [CrossRef] [PubMed]
- Ooi, A.; Dykema, K.; Ansari, A.; Petillo, D.; Snider, J.; Kahnoski, R.; Anema, J.; Craig, D.; Carpten, J.; Teh, B.T.; et al. CUL3 and NRF2 mutations confer an NRF2 activation phenotype in a sporadic form of papillary renal cell carcinoma. Cancer Res. 2013, 73, 2044–2051. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.R.; Oh, J.E.; Kim, M.S.; Kang, M.R.; Park, S.W.; Han, J.Y.; Eom, H.S.; Yoo, N.J.; Lee, S.H. Oncogenic NRF2 mutations in squamous cell carcinomas of oesophagus and skin. J. Pathol. 2010, 220, 446–451. [Google Scholar] [CrossRef]
- Goldstein, L.D.; Lee, J.; Gnad, F.; Klijn, C.; Schaub, A.; Reeder, J.; Daemen, A.; Bakalarski, C.E.; Holcomb, T.; Shames, D.S.; et al. Recurrent loss of NFE2L2 exon 2 is a mechanism for Nrf2 pathway activation in human cancers. Cell Rep. 2016, 16, 2605–2617. [Google Scholar] [CrossRef]
- Fabrizio, F.P.; Mazza, T.; Castellana, S.; Sparaneo, A.; Muscarella, L.A. Epigenetic scanning of KEAP1 CpG sites uncovers new molecular-driven patterns in lung adeno and squamous cell carcinomas. Antioxidants 2020, 9, 904. [Google Scholar] [CrossRef]
- Fabrizio, F.P.; Sparaneo, A.; Centra, F.; Trombetta, D.; Storlazzi, C.T.; Graziano, P.; Maiello, E.; Fazio, V.M.; Muscarella, L.A. Methylation density pattern of KEAP1 gene in lung cancer cell lines detected by quantitative methylation specific PCR and pyrosequencing. Int. J. Mol. Sci. 2019, 20, 2697. [Google Scholar] [CrossRef]
- Guo, Y.; Yu, S.; Zhang, C.; Kong, A.N. Epigenetic regulation of Keap1-Nrf2 signaling. Free Radic. Biol. Med. 2015, 88, 337–349. [Google Scholar] [CrossRef]
- Rinn, J.L.; Chang, H.Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 2012, 81, 145–166. [Google Scholar] [CrossRef]
- Tay, Y.; Rinn, J.; Pandolfi, P.P. The multilayered complexity of ceRNA crosstalk and competition. Nature 2012, 505, 344–352. [Google Scholar] [CrossRef]
- Bhan, A.; Soleimani, M.; Mandal, S.S. Long noncoding RNA and cancer: A new paradigm. Cancer Res. 2017, 77, 3965–3981. [Google Scholar] [CrossRef] [PubMed]
- Moran, V.A.; Perera, R.J.; Khalil, A.M. Emerging functional and mechanistic paradigms of mammalian long non-coding RNAs. Nucleic Acids Res. 2012, 40, 6391–6400. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Shen, C.; Zhu, J.; Shen, G.; Li, Z.; Dong, J. Long noncoding RNAs in the regulation of oxidative stress. Oxid. Med. Cell. Longev. 2019, 2019, 1318795. [Google Scholar] [CrossRef] [PubMed]
- Thai, P.; Statt, S.; Chen, C.H.; Liang, E.; Campbell, C.; Wu, R. Characterization of a novel long noncoding RNA, SCAL1, induced by cigarette smoke and elevated in lung cancer cell lines. Am. J. Respir. Cell. Mol. Biol. 2013, 49, 204–211. [Google Scholar] [CrossRef]
- White, N.M.; Cabanski, C.R.; Silva-Fisher, J.M.; Dang, H.X.; Govindan, R.; Maher, C.A. Transcriptome sequencing reveals altered long intergenic non-coding RNAs in lung cancer. Genome Biol. 2014, 15, 429. [Google Scholar] [CrossRef]
- Moreno Leon, L.; Gautier, M.; Allan, R.; Ilié, M.; Nottet, N.; Pons, N.; Paquet, A.; Lebrigand, K.; Truchi, M.; Fassy, J.; et al. The nuclear hypoxia-regulated NLUCAT1 long non-coding RNA contributes to an aggressive phenotype in lung adenocarcinoma through regulation of oxidative stress. Oncogene 2019, 38, 7146–7165. [Google Scholar] [CrossRef]
- Johnson, G.S.; Li, J.; Beaver, L.M.; Dashwood, W.M.; Sun, D.; Rajendran, P.; Williams, D.E.; Ho, E.; Dashwood, R.H. A functional pseudogene, NMRAL2P, is regulated by Nrf2 and serves as a coactivator of NQO1 in sulforaphane-treated colon cancer cells. Mol. Nutr. Food Res. 2017, 61. [Google Scholar] [CrossRef]
- Zhou, H.; Sun, L.; Wan, F. Molecular mechanisms of TUG1 in the proliferation, apoptosis, migration and invasion of cancer cells. Oncol. Lett. 2019, 18, 4393–4402. [Google Scholar] [CrossRef]
- Sun, Z.; Huang, G.; Cheng, H. Transcription factor Nrf2 induces the up-regulation of lncRNA TUG1 to promote progression and Adriamycin resistance in urothelial carcinoma of the bladder. Cancer Manag. Res. 2019, 11, 6079–6090. [Google Scholar] [CrossRef]
- Ashouri, A.; Sayin, V.I.; Van den Eynden, J.; Singh, S.X.; Papagiannakopoulos, T.; Larsson, E. Pan-cancer transcriptomic analysis associates long non-coding RNAs with key mutational driver events. Nat. Commun. 2016, 7, 13197. [Google Scholar] [CrossRef]
- Zhang, L.; Liu, Z.; Li, X.; Zhang, P.; Wang, J.; Zhu, D.; Chen, X.; Ye, L. Low long non-coding RNA HOTAIR expression is associated with down-regulation of Nrf2 in the spermatozoa of patients with asthenozoospermia or oligoasthenozoospermia. Int. J. Clin. Exp. Pathol. 2015, 8, 14198–14205. [Google Scholar] [PubMed]
- Zhang, X.; Wang, T.; Yang, Y.; Li, R.; Chen, Y.; Li, R.; Jiang, X.; Wang, L. Tanshinone IIA attenuates acetaminophen-induced hepatotoxicity through HOTAIR-Nrf2-MRP2/4 signaling pathway. Biomed. Pharmacother. 2020, 130, 110547. [Google Scholar] [CrossRef] [PubMed]
- Zeng, R.; Zhang, R.; Song, X.; Ni, L.; Lai, Z.; Liu, C.; Ye, W. The long non-coding RNA MALAT1 activates Nrf2 signaling to protect human umbilical vein endothelial cells from hydrogen peroxide. Biochem. Biophys. Res. Commun. 2018, 495, 2532–2538. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Ke, S.; Zhong, L.; Wu, J.; Tseng, A.; Morpurgo, B.; Golovko, A.; Wang, G.; Cai, J.J.; Ma, X.; et al. Long noncoding RNA MALAT1 regulates generation of reactive oxygen species and the insulin responses in male mice. Biochem. Pharmacol. 2018, 152, 94–103. [Google Scholar] [CrossRef]
- Lu, Y.; Liu, W.G.; Lu, J.H.; Liu, Z.J.; Li, H.B.; Liu, G.J.; She, H.Y.; Li, G.Y.; Shi, X.H. LncRNA UCA1 promotes renal cell carcinoma proliferation through epigenetically repressing p21 expression and negatively regulating miR-495. Tumour. Biol. 2017, 39, 1010428317701632. [Google Scholar] [CrossRef]
- Li, C.; Fan, K.; Qu, Y.; Zhai, W.; Huang, A.; Sun, X.; Xing, S. Deregulation of UCA1 expression may be involved in the development of chemoresistance to cisplatin in the treatment of non-small-cell lung cancer via regulating the signaling pathway of microRNA-495/NRF2. J. Cell Physiol. 2020, 235, 3721–3730. [Google Scholar] [CrossRef]
- Geng, J.F.; Liu, X.; Zhao, H.B.; Fan, W.F.; Geng, J.J.; Liu, X.Z. LncRNA UCA1 inhibits epilepsy and seizure-induced brain injury by regulating miR-495/Nrf2-ARE signal pathway. Int. J. Biochem. Cell Biol. 2018, 99, 133–139. [Google Scholar] [CrossRef]
- Zhou, L.; Xu, D.Y.; Sha, W.G.; Shen, L.; Lu, G.Y.; Yin, X. Long non-coding MIAT mediates high glucose-induced renal tubular epithelial injury. Biochem. Biophys. Res. Commun. 2015, 468, 726–732. [Google Scholar] [CrossRef]
- Liu, M.; Song, Y.; Han, Z. Study on the effect of LncRNA AK094457 on OX-LDL induced vascular smooth muscle cells. Am. J. Transl. Res. 2019, 11, 5623–5633. [Google Scholar]
- Yang, Q.; Li, K.; Huang, X.; Zhao, C.; Mei, Y.; Li, X.; Jiao, L.; Yang, H. lncRNA SLC7A11-AS1 promotes chemoresistance by blocking SCF (beta-TRCP)-mediated degradation of NRF2 in pancreatic cancer. Mol. Ther. Nucleic Acids 2020, 19, 974–985. [Google Scholar] [CrossRef]
- Wu, L.; Pan, C.; Wei, X.; Shi, Y.; Zheng, J.; Lin, X.; Shi, L. lncRNA KRAL reverses 5-fluorouracil resistance in hepatocellular carcinoma cells by acting as a ceRNA against miR-141. Cell Commun. Signal. 2018, 16, 47. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.L.; Cai, W.P.; Lei, X.; Shi, K.Q.; Lin, X.Y.; Shi, L. NRAL mediates cisplatin resistance in hepatocellular carcinoma via miR-340-5p/Nrf2 axis. J. Cell. Commun. Signal. 2019, 13, 99–112. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Xiong, R.; Li, C.; Xu, M.; Guo, M. LncRNA TUG1 promotes cisplatin resistance in esophageal squamous cell carcinoma cells by regulating Nrf2. Acta Biochim. Biophys. Sin. 2019, 51, 826–833. [Google Scholar] [CrossRef]
- Gao, M.; Li, C.; Xu, M.; Liu, Y.; Cong, M.; Liu, S. LncRNA MT1DP aggravates cadmium-induced oxidative stress by repressing the function of Nrf2 and is dependent on interaction with miR-365. Adv. Sci. (Weinh) 2018, 5, 1800087. [Google Scholar] [CrossRef] [PubMed]
- Gai, C.; Liu, C.; Wu, X.; Yu, M.; Zheng, J.; Zhang, W.; Lv, S.; Li, W. MT1DP loaded by folate-modified liposomes sensitizes erastin-induced ferroptosis via regulating miR-365a-3p/NRF2 axis in non-small cell lung cancer cells. Cell Death Dis. 2020, 11, 751. [Google Scholar] [CrossRef]
- Yang, G.; Yin, H.; Lin, F.; Gao, S.; Zhan, K.; Tong, H.; Tang, X.; Pan, Q.; Gou, X. Long noncoding RNA TUG1 regulates prostate cancer cell proliferation, invasion and migration via the Nrf2 signaling axis. Pathol. Res. Pract. 2020, 216, 152851. [Google Scholar] [CrossRef]
- Khalil, A.M.; Guttman, M.; Huarte, M.; Garber, M.; Raj, A.; Rivea Morales, D.; Thomas, K.; Presser, A.; Bernstein, B.E.; van Oudenaarden, A.; et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl. Acad. Sci. USA 2009, 106, 11667–11672. [Google Scholar] [CrossRef]
- Dai, Q.; Li, J.; Zhou, K.; Liang, T. Competing endogenous RNA: A novel posttranscriptional regulatory dimension associated with the progression of cancer. Oncol. Lett. 2015, 10, 2683–2690. [Google Scholar] [CrossRef]
- Kornienko, A.E.; Guenzl, P.M.; Barlow, D.P.; Pauler, F.M. Gene regulation by the act of long non-coding RNA transcription. BMC Biol. 2013, 11, 59. [Google Scholar] [CrossRef]
- Li, Z.; Xu, L.; Tang, N.; Xu, Y.; Ye, X.; Shen, S.; Niu, X.; Lu, S.; Chen, Z. The polycomb group protein EZH2 inhibits lung cancer cell growth by repressing the transcription factor Nrf2. FEBS Lett. 2014, 588, 3000–3007. [Google Scholar] [CrossRef]
- Feng, X.; Zhao, J.; Ding, J.; Shen, X.; Zhou, J.; Xu, Z. LncRNA Blnc1 expression and its effect on renal fibrosis in diabetic nephropathy. Am. J. Transl. Res. 2019, 11, 5664–5672. [Google Scholar] [PubMed]
- Bartel, D.P. Metazoan microRNAs. Cell 2018, 173, 20–51. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Croce, C.M. The role of MicroRNAs in human cancer. Signal Transduct. Target Ther. 2016, 1, 15004. [Google Scholar] [CrossRef] [PubMed]
- Shah, N.M.; Rushworth, S.A.; Murray, M.Y.; Bowles, K.M.; MacEwan, D.J. Understanding the role of NRF2-regulated miRNAs in human malignancies. Oncotarget 2013, 4, 1130–1142. [Google Scholar] [CrossRef]
- Ashrafizadeh, M.; Ahmadi, Z.; Samarghandian, S.; Mohammadinejad, R.; Yaribeygi, H.; Sathyapalan, T.; Sahebkar, A. MicroRNA-mediated regulation of Nrf2 signaling pathway: Implications in disease therapy and protection against oxidative stress. Life Sci. 2020, 244, 117329. [Google Scholar] [CrossRef]
- Reichard, J.F.; Motz, G.T.; Puga, A. Heme oxygenase-1 induction by NRF2 requires inactivation of the transcriptional repressor BACH1. Nucleic Acids Res. 2007, 35, 7074–7086. [Google Scholar] [CrossRef]
- Hou, W.; Tian, Q.; Steuerwald, N.M.; Schrum, L.W.; Bonkovsky, H.L. The let-7 microRNA enhances heme oxygenase-1 by suppressing Bach1 and attenuates oxidant injury in human hepatocytes. Biochim. Biophys. Acta 2012, 1819, 1113–1122. [Google Scholar] [CrossRef]
- Liu, X.; Zhao, H.; Luo, C.; Du, D.; Huang, J.; Ming, Q.; Jin, F.; Wang, D.; Huang, W. Acetaminophen responsive miR-19b modulates SIRT1/Nrf2 signaling pathway in drug-induced hepatotoxicity. Toxicol. Sci. 2019, 170, 476–488. [Google Scholar] [CrossRef]
- Wang, L.; Chen, Y.; Sternberg, P.; Cai, J. Essential roles of the PI3 kinase/Akt pathway in regulating Nrf2-dependent antioxidant functions in the RPE. Invest. Ophthalmol. Vis. Sci. 2008, 49, 1671–1678. [Google Scholar] [CrossRef]
- Chen, Y.F.; Wei, Y.Y.; Yang, C.C.; Liu, C.J.; Yeh, L.Y.; Chou, C.H.; Chang, K.W.; Lin, S.C. miR-125b suppresses oral oncogenicity by targeting the anti-oxidative gene PRXL2A. Redox Biol. 2019, 22, 101140. [Google Scholar] [CrossRef]
- Zhou, C.; Zhao, L.; Zheng, J.; Wang, K.; Deng, H.; Liu, P.; Chen, L.; Mu, H. MicroRNA-144 modulates oxidative stress tolerance in SH-SY5Y cells by regulating nuclear factor erythroid 2-related factor 2-glutathione axis. Neurosci. Lett. 2016, 655, 21–27. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Teng, Y.; Liu, Q. MicroRNA-153 regulates NRF2 expression and is associated with breast carcinogenesis. Clin. Lab. 2016, 62, 39–47. [Google Scholar] [CrossRef] [PubMed]
- Gu, S.; Lai, Y.; Chen, H.; Liu, Y.; Zhang, Z. miR-155 mediates arsenic trioxide resistance by activating Nrf2 and suppressing apoptosis in lung cancer cells. Sci. Rep. 2017, 7, 12155. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Mao, L.; Wang, S.G.; Chen, F.L.; Ji, F.; Fei, H.D. MicroRNA-200a activates Nrf2 signaling to protect osteoblasts from dexamethasone. Oncotarget 2017, 8, 104867–104876. [Google Scholar] [CrossRef]
- Kim, W.D.; Kim, Y.W.; Cho, I.J.; Lee, C.H.; Kim, S.G. E-cadherin inhibits nuclear accumulation of Nrf2: Implications for chemoresistance of cancer cells. J. Cell. Sci. 2012, 125, 1284–1295. [Google Scholar] [CrossRef]
- Chorley, B.N.; Campbell, M.R.; Wang, X.; Karaca, M.; Sambandan, D.; Bangura, F.; Xue, P.; Pi, J.; Kleeberger, S.R.; Bell, D.A. Identification of novel NRF2-regulated genes by ChIP-Seq: Influence on retinoid X receptor alpha. Nucleic Acids Res. 2012, 40, 7416–7429. [Google Scholar] [CrossRef]
- Luna, C.; Li, G.; Qiu, J.; Epstein, D.L.; Gonzalez, P. Role of miR-29b on the regulation of the extracellular matrix in human trabecular meshwork cells under chronic oxidative stress. Mol. Vis. 2009, 15, 2488–2497. [Google Scholar]
- Chan, K.; Lu, R.; Chang, J.C.; Kan, Y.W. NRF2, a member of the NFE2 family of transcription factors, is not essential for murine erythropoiesis, growth, and development. Proc. Natl. Acad. Sci. USA 1996, 93, 13943–13948. [Google Scholar] [CrossRef]
- Amodio, N.; Di Martino, M.T.; Foresta, U.; Leone, E.; Lionetti, M.; Leotta, M.; Gullà, A.M.; Pitari, M.R.; Conforti, F.; Rossi, M.; et al. miR-29b sensitizes multiple myeloma cells to bortezomib-induced apoptosis through the activation of a feedback loop with the transcription factor Sp1. Cell Death Dis. 2012, 3, e436. [Google Scholar] [CrossRef]
- Akdemir, B.; Nakajima, Y.; Inazawa, J.; Inoue, J. miR-432 Induces NRF2 stabilization by directly targeting KEAP1. Mol. Cancer Res. 2017, 15, 1570–1578. [Google Scholar] [CrossRef]
- Zhang, S.; Wu, W.; Jiao, G.; Li, C.; Liu, H. MiR-455-3p activates Nrf2/ARE signaling via HDAC2 and protects osteoblasts from oxidative stress. Int. J. Biol. Macromol. 2018, 107, 2094–2101. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Zhu, H.; Wang, C.; Zhu, X.; Liu, G.; Chen, C.; Cui, Z. microRNA-455 targets cullin 3 to activate Nrf2 signaling and protect human osteoblasts from hydrogen peroxide. Oncotarget 2017, 8, 59225–59234. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chen, Z.J.; Rong, L.; Huang, D.; Jiang, Q. Targeting cullin 3 by miR-601 activates Nrf2 signaling to protect retinal pigment epithelium cells from hydrogen peroxide. Biochem. Biophys. Res. Commun. 2019, 515, 679–687. [Google Scholar] [CrossRef] [PubMed]
- Akhdar, H.; Loyer, P.; Rauch, C.; Corlu, A.; Guillouzo, A.; Morel, F. Involvement of Nrf2 activation in resistance to 5-fluorouracil in human colon cancer HT-29 cells. Eur. J. Cancer 2009, 45, 2219–2227. [Google Scholar] [CrossRef]
- Inami, Y.; Waguri, S.; Sakamoto, A.; Kouno, T.; Nakada, K.; Hino, O.; Watanabe, S.; Ando, J.; Iwadate, M.; Yamamoto, M.; et al. Persistent activation of Nrf2 through p62 in hepatocellular carcinoma cells. J. Cell Biol. 2011, 193, 275–284. [Google Scholar] [CrossRef]
- Moltzahn, F.; Olshen, A.B.; Baehner, L.; Peek, A.; Fong, L.; Stöppler, H.; Simko, J.; Hilton, J.F.; Carroll, P.; Blelloch, R. Microfluidic-based multiplex qRT-PCR identifies diagnostic and prognostic microRNA signatures in the sera of prostate cancer patients. Cancer Res. 2011, 71, 550–560. [Google Scholar] [CrossRef]
- Wang, C.; Su, Z.; Sanai, N.; Xue, X.; Lu, L.; Chen, Y.; Wu, J.; Zheng, W.; Zhuge, Q.; Wu, Z.B. microRNA expression profile and differentially-expressed genes in prolactinomas following bromocriptine treatment. Oncol. Rep. 2012, 27, 1312–1320. [Google Scholar]
- Jung, K.A.; Lee, S.; Kwak, M.K. NFE2L2/NRF2 activity is linked to mitochondria and AMP-activated protein kinase signaling in cancers through miR-181c/mitochondria-encoded cytochrome c oxidase regulation. Antioxid. Redox Signal. 2017, 27, 945–961. [Google Scholar] [CrossRef]
- Yang, M.; Yao, Y.; Eades, G.; Zhang, Y.; Zhou, Q. MiR-28 regulates Nrf2 expression through a Keap1-independent mechanism. Breast Cancer Res. Treat. 2011, 129, 983–991. [Google Scholar] [CrossRef]
- Liu, H.; Wu, H.Y.; Wang, W.Y.; Zhao, Z.L.; Liu, X.Y.; Wang, L.Y. Regulation of miR-92a on vascular endothelial aging via mediating Nrf2-KEAP1-ARE signal pathway. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 2734–2742. [Google Scholar]
- Singh, B.; Ronghe, A.M.; Chatterjee, A.; Bhat, N.K.; Bhat, H.K. MicroRNA-93 regulates NRF2 expression and is associated with breast carcinogenesis. Carcinogenesis 2013, 34, 1165–1172. [Google Scholar] [CrossRef] [PubMed]
- Yi, J.; Huang, W.Z.; Wen, Y.Q.; Yi, Y.C. Effect of miR-101 on proliferation and oxidative stress-induced apoptosis of breast cancer cells via Nrf2 signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 8931–8939. [Google Scholar] [PubMed]
- Duru, N.; Gernapudi, R.; Zhang, Y.; Yao, Y.; Lo, P.K.; Wolfson, B.; Zhou, Q. NRF2/miR-140 signaling confers radioprotection to human lung fibroblasts. Cancer Lett. 2015, 369, 184–191. [Google Scholar] [CrossRef]
- Duru, N.; Zhang, Y.; Gernapudi, R.; Wolfson, B.; Lo, P.K.; Yao, Y.; Zhou, Q. Loss of miR-140 is a key risk factor for radiation-induced lung fibrosis through reprogramming fibroblasts and macrophages. Sci. Rep. 2017, 6, 39572. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Liu, D.; Xue, Y.; Hu, X. Enforced miR-144-3p expression as a non-invasive biomarker for the acute myeloid leukemia patients mainly by targeting NRF2. Clin. Lab. 2017, 63, 679–687. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Ye, W.; Zhang, Y.; Yu, D.; Shao, Q.; Liang, J.; Zhang, M. miR-144 reverses chemoresistance of hepatocellular carcinoma cell lines by targeting Nrf2-dependent antioxidant pathway. Am. J. Transl. Res. 2010, 8, 2992–3002. [Google Scholar]
- Qu, J.; Zhang, L.; Li, L.; Su, Y. miR-148b functions as a tumor suppressor by targeting endoplasmic reticulum Metallo protease 1 in human endometrial cancer cells. Oncol. Res. 2018, 27, 81–88. [Google Scholar] [CrossRef]
- Narasimhan, M.; Patel, D.; Vedpathak, D.; Rathinam, M.; Henderson, G.; Mahimainathan, L. Identification of novel microRNAs in post-transcriptional control of Nrf2 expression and redox homeostasis in neuronal, SH-SY5Y cells. PLoS ONE 2012, 7, e51111. [Google Scholar] [CrossRef]
- Chen, C.; Jiang, X.; Gu, S.; Zhang, Z. MicroRNA-155 regulates arsenite-induced malignant transformation by targeting Nrf2-mediated oxidative damage in human bronchial epithelial cells. Toxicol. Lett. 2017, 278, 38–47. [Google Scholar] [CrossRef]
- Pottier, N.; Maurin, T.; Chevalier, B.; Puisségur, M.P.; Lebrigand, K.; Robbe-Sermesant, K.; Bertero, T.; Lino Cardenas, C.L.; Courcot, E.; Rios, G.; et al. Identification of keratinocyte growth factor as a target of microRNA-155 in lung fibroblasts: Implication in epithelial-mesenchymal interactions. PLoS ONE 2009, 4, e6718. [Google Scholar] [CrossRef]
- Zhao, M.W.; Yang, P.; Zhao, L.L. Chlorpyrifos activates cell pyroptosis and increases susceptibility on oxidative stress-induced toxicity by miR-181/SIRT1/PGC-1alpha/Nrf2 signaling pathway in human neuroblastoma SH-SY5Y cells: Implication for association between chlorpyrifos and Parkinson's disease. Environ. Toxicol. 2019, 34, 699–707. [Google Scholar] [PubMed]
- Shi, L.; Chen, Z.G.; Wu, L.L.; Zheng, J.J.; Yang, J.R.; Chen, X.F.; Chen, Z.Q.; Liu, C.L.; Chi, S.Y.; Zheng, J.Y.; et al. miR-340 reverses cisplatin resistance of hepatocellular carcinoma cell lines by targeting Nrf2-dependent antioxidant pathway. Asian Pac. J. Cancer Prev. 2014, 15, 10439–10444. [Google Scholar] [CrossRef] [PubMed]
- Qaisiya, M.; Coda Zabetta, C.D.; Bellarosa, C.; Tiribelli, C. Bilirubin mediated oxidative stress involves antioxidant response activation via Nrf2 pathway. Cell Signal. 2014, 26, 512–520. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, S.; Inoue, J.; Kawano, T.; Kozaki, K.; Omura, K.; Inazawa, J. The impact of miRNA-based molecular diagnostics and treatment of NRF2-stabilized tumors. Mol. Cancer Res. 2014, 12, 58–68. [Google Scholar] [CrossRef] [PubMed]
| (A) | ||||||
| LncRNA ID | Chr Location * | Role | NRF2 Levels | NRF2-Related Effects | Cell Cultures/Validation Models | Refs. |
| SCAL1 (LUCAT1) | chr5: 91,285,160–91,314,516 | Transcriptional activity | Upregulated | Downstream mediator of NRF2 against ROS-induced oxidative stress, having a pro-tumorigenic role | HBE cells; Lung ADC and SqCC | [54,55] |
| NLUCAT1 | 5q14.3 | Transcriptional activity | Upregulated | It induces a positive loop on the NRF2 network, correlates with hypoxic stress and contributes to tumor invasion and proliferation | A549 ADC cells | [56] |
| NMRAL2P | chr3: 185,959,943–185,980,872 | Transcriptional activity | Upregulated | Downstream regulator of NRF2-dependent NQO1 activation. It exerts an oncogenic action by promoting EMT | Colon cancer cells | [57] |
| TUG1 | chr22: 30,970,182–30,978,847 | Transcriptional activity | Upregulated | Oncogenic roles in apoptosis, proliferation and chemoresistance by modulating the NRF2 signaling | UCB cells; prostate cancer cells | [58,59] |
| LINC00942 | chr12: 1,500,525–1,504,424 | Transcriptional activity | Upregulated | It concurs on NRF2 antioxidant pathway activation | 19 cancer types from TCGA | [60] |
| (B) | ||||||
| LncRNA ID | Chr Locations * | Role | NRF2-Related Effects | Cell Cultures/Validation Models | Refs. | |
| HOTAIR | chr12: 53,962,308–53,974,956 | Negative regulator | Mediates histone H4 acetylation at the NFE2L2 gene promoter and is also involved in the epigenetic suppression via NRF2-MRP2/4 pathway | GC-1 spg; human hepatocytes L02 cells | [61,62] | |
| MALAT1 | chr11: 65,497,688–65,506,431 | Positive regulator | Participates in MALAT1-mediated HUVEC protection from hydrogen peroxide and it is negatively regulated by KEAP1 | HUVEC cells; mouse primary hepatocytes | [63,64] | |
| UCA1 | chr19: 15,828,206–15,836,136 | Positive regulator | It has an oncogenic role promotes proliferation through p21 repression, it enhances chemoresistance to cisplatin via UCA1/NRF2 crosstalk signaling by sponging miR-495 and promoting the inhibition of apoptosis | Human RCC cells; A549 lung ADC cells; HEK-293 cells | [65,66,67] | |
| BLNC1 | chr9: 95,559,657–95,568,023 | Positive regulator | Participates in the hyperactivation of HO-1/NRF2 and crosstalks with NF-κB pathway by attenuating renal fibrosis, inflammation and oxidative stress | Renal HK-2 cells | [62] | |
| MIAT | chr22: 26,657,520–26,676,475 | Positive regulator | Targets NRF2 by mediating high glucose- induced tubular injury | renal HK-2 epithelial cells | [68] | |
| AK094457 | chr10: 124,745,200–124,748,551 | Positive regulator | Inversely correlates with the upregulation of NRF2 and HO-1 | VSMC cells | [69] | |
| SLC7A11-AS1 | chr4: 138,089,014–138,178,177 | Negative regulator | Promotes resistance to gemcitabine by repressing SCFβ-TRCP-mediated degradation of NRF2 | PDAC cells | [70] | |
| KRAL | 19q10.14 | Positive regulator | Increases sensitivity to fluorouracil by targeting miR-141/KEAP1 axis as ceRNA | HCC SMMC-7721 and HepG2 cells | [71] | |
| NRAL | 19q10.14 | Negative regulator | Enhances resistance to cisplatin for NRF2 by binding to miR-340–5p | HCC SMMC-7721 and HepG2 cells | [72] | |
| TUG1 | chr22: 30,970,182–30,978,847 | Positive regulator | Promotes cisplatin resistance by regulating and stabilizing the NRF2 protein | human EC TE-1 cells | [73] | |
| MT1DP | chr16: 56,643,705–56,644,786 | Negative regulator | Enhances the sensitivity of NRF2 overexpression to erastin-induced ferroptosis by stabilizing miR-365a-3p | HCC HepG2 cells; NSCLC A549 and H1299 cells | [74,75] | |
| miRNA ID | Direct or Indirect Interaction/ Targets | NRF2-Related Effects | Cell Cultures/Validation Models | Refs. |
|---|---|---|---|---|
| Let 7b/c | Indirect; BACH1, HO-1 | Participates in negative regulation by suppressing BACH1 and thus stimulating the overexpression of HO-1 via NRF2 | HaCaT human keratinocytes cells; HCC (Huh-7 and HepG2) cells | [86,87] |
| MiR-19b | Indirect; SIRT-1 | Stimulates the activation of SIRT1 by activating NRF2 cascade and its downstream target genes; enhancing drug hepatotoxicity | Human liver LO2 cells | [88] |
| MiR-32 | Indirect; PI3K | Exerts an oncogenic role by inducing the upregulation of NRF2and promoting cells survival | Human RPE cells | [89] |
| MiR-125b | Indirect; PRXL2A | Promotes PRXL2A activation by positive feedback loops involving NRF2 signaling pathway | OSCC (SAS, OECM1, HSC3, FaDu, OC3) cells, HEK-293 and primary NOK cells | [90] |
| MiR-144 | Direct | Regulates the ROS scavenging via NRF2 and stimulates the GSH synthesis | Neuroblastoma SH-SY5Y cells | [91] |
| MiR-153 | Direct | Acts as an oncogene by promoting cell migration and invasion via NRF2 signaling | CRC cells; OSCC (CAL 27) cells and tissues | [92] |
| MiR-155 | Direct | Upregulates NRF2 and its downstream targets (HO-1 and NQO1) by inhibiting apoptosis and mediating ATO resistance | Human lung ADC A549 cells | [93] |
| MiR-181c | Direct | Enhances the activation of NRF2 and other multiple pro-survival pathways | Human CRC (HT29) cells | [108] |
| MiR-200a | Indirect; KEAP1 | Stimulates NRF2 signaling pathway by suppressing KEAP1 and decreasing ROS concentration | OB-6 human osteoblastic cells | [94] |
| MiR-200c | Indirect; PI3K/AKT | Its downregulation provides a feedback mechanism by which NRF2 indirectly regulates E-cadherin and metastasis via PI3K/AKT activation | RPE (ARPE-19) cells | [89] |
| MiR-365/193b/29b | Direct | Enforces NRF2 and its downstream targets activation | Lymphoblast cells | [96] |
| MiR-432 | Indirect; KEAP1 | Targets KEAP1 and decreases the sensitivity to cisplatin (CDDP) drug via activation and stabilization of NRF2 | ESCC (KYSE170, KYSE770, and KYSE2270) cells | [100] |
| MiR-455 | Indirect; HDAC2 | Promotes NRF2 expression via its upstream mediator, HDAC2, by ensuring more protection against oxidative injury and activating ARE-related genes (NQO1, HO-1 and GCLC) | MC3T3-E1 and hFOB1.19 osteoblastic cells | [101,102] |
| MiR-601 | Indirect; CUL3/KEAP1 | Targets CUL3 and concurs into a dependent mechanism of CUL3-KEAP1-NRF2 activation against oxidative damage | RPE (ARPE-19) cells | [103] |
| MiR-617/592/1207/550 | Direct | Exert a pro-oncogenic actions on the NRF2 upregulation; promote cell growth and proliferation by activating antioxidant defense system | ESCC cells; colorectal and liver carcinoma cells; prostate cancer cells | [41,104,105,106,107] |
| miRNA ID | Direct or Indirect Interaction/Targets | NRF2-Related Effects | Cell Cultures/Validation Models | Refs. |
|---|---|---|---|---|
| MiR-28 | Indirect; KEAP1 | Exerts a negative effect by repressing NRF2 and inhibiting tumor cell growth independently from KEAP1, without any changes at protein level | Human breast MCF-7 and HEK-293 cells | [109] |
| MiR-92a | Indirect; KEAP1/ARE | Its inhibition leads to cell proliferation by decreasing apoptosis-inducing factors and KEAP1 expression with a consequent upregulation of NRF2 and its target genes | HUVECs | [110] |
| MiR-93 | Direct | Acts as a oncogenic regulator by downregulating NRF2 gene and by targeting specific 3′UTR sites with a strong impact on decreasing apoptosis, increasing colony formation and cell migration | Human breast epithelial MCF-10A and neoplastic T47D cells | [111] |
| MiR-101 | Direct | Affects the binding to 3′UTR of NRF2, thus promoting NRF2 overexpression and inhibiting cell proliferation | Human breast MCF-7 cells | [112] |
| MiR-140 | Direct | Targets NRF2 gene by ensuring more radioprotection against TGF-β1-mediated inflammation and fibroblasts differentiation | Human normal lung fibroblast (CCD-19Lu) and mammary epithelial MCF10A cells | [113,114] |
| MiR-144 | Direct | Exerts an oncogenic role by inhibiting NRF2 signaling pathway as well as promotes cell viability, suppresses apoptosis and finally reverses chemoresistance NRF2-related | HL-60 AML and Bel-7402 HCC cells | [115,116] |
| MiR-148b | Direct | Suppresses cell proliferation and regulates the oxidative stress response by downregulating HIF-1 and NRF2 and inhibiting ERMP1 | RL95-2 human endometrial cancer cells | [117] |
| MiR-153/27a/142/144 | Direct | Act as oncogenic miRNAs which decrease apoptosis and successively reinforce cells proliferation via repressing NRF2 and its downstream target genes (GCLC, GSR) | Breast cancer and CRC cell lines; SH-SY5Y neuroblastoma cells | [92,118] |
| MiR-155 | Direct | Accelerates cell malignant transformation by targeting NRF2-mediated oxidative damage and repressing NRF2 and its related gene expressions (GSH, NO and SOD) | 16-HBE cells | [119] |
| MiR-181 | Indirect; SIRT1/PGC-1α | Suppresses SIRT1/PGC-1α/NRF2 signaling pathway by activating cell apoptosis and oxidative stress | SH-SY5Y neuroblastoma cells | [121] |
| MiR-340 | Direct | Inhibits NRF2-dependent antioxidant pathway and enhancing cells sensitivity to cisplatin | HCC SMMC-7721 and HepG2 cells | [122] |
| MiR-365 | Indirect; MT1DP | Represses NRF2 activity via direct binding to its 3′UTR and induce an aggravation of oxidative stress by activating MT1DP | HCC HepG2 cells | [74] |
| MiR-507/634/450a/129 | Direct | Negatively affect NRF2 oncogenic regulation by directly targeting NRF2, thus increasing cisplatin sensitivity and suppressing cell growth | A549 ADC cells | [123,124] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fabrizio, F.P.; Sparaneo, A.; Muscarella, L.A. NRF2 Regulation by Noncoding RNAs in Cancers: The Present Knowledge and the Way Forward. Cancers 2020, 12, 3621. https://doi.org/10.3390/cancers12123621
Fabrizio FP, Sparaneo A, Muscarella LA. NRF2 Regulation by Noncoding RNAs in Cancers: The Present Knowledge and the Way Forward. Cancers. 2020; 12(12):3621. https://doi.org/10.3390/cancers12123621
Chicago/Turabian StyleFabrizio, Federico Pio, Angelo Sparaneo, and Lucia Anna Muscarella. 2020. "NRF2 Regulation by Noncoding RNAs in Cancers: The Present Knowledge and the Way Forward" Cancers 12, no. 12: 3621. https://doi.org/10.3390/cancers12123621
APA StyleFabrizio, F. P., Sparaneo, A., & Muscarella, L. A. (2020). NRF2 Regulation by Noncoding RNAs in Cancers: The Present Knowledge and the Way Forward. Cancers, 12(12), 3621. https://doi.org/10.3390/cancers12123621





