MiR-378a-3p Is Critical for Burkitt Lymphoma Cell Growth
Abstract
:Simple Summary
Abstract
1. Introduction
2. Results
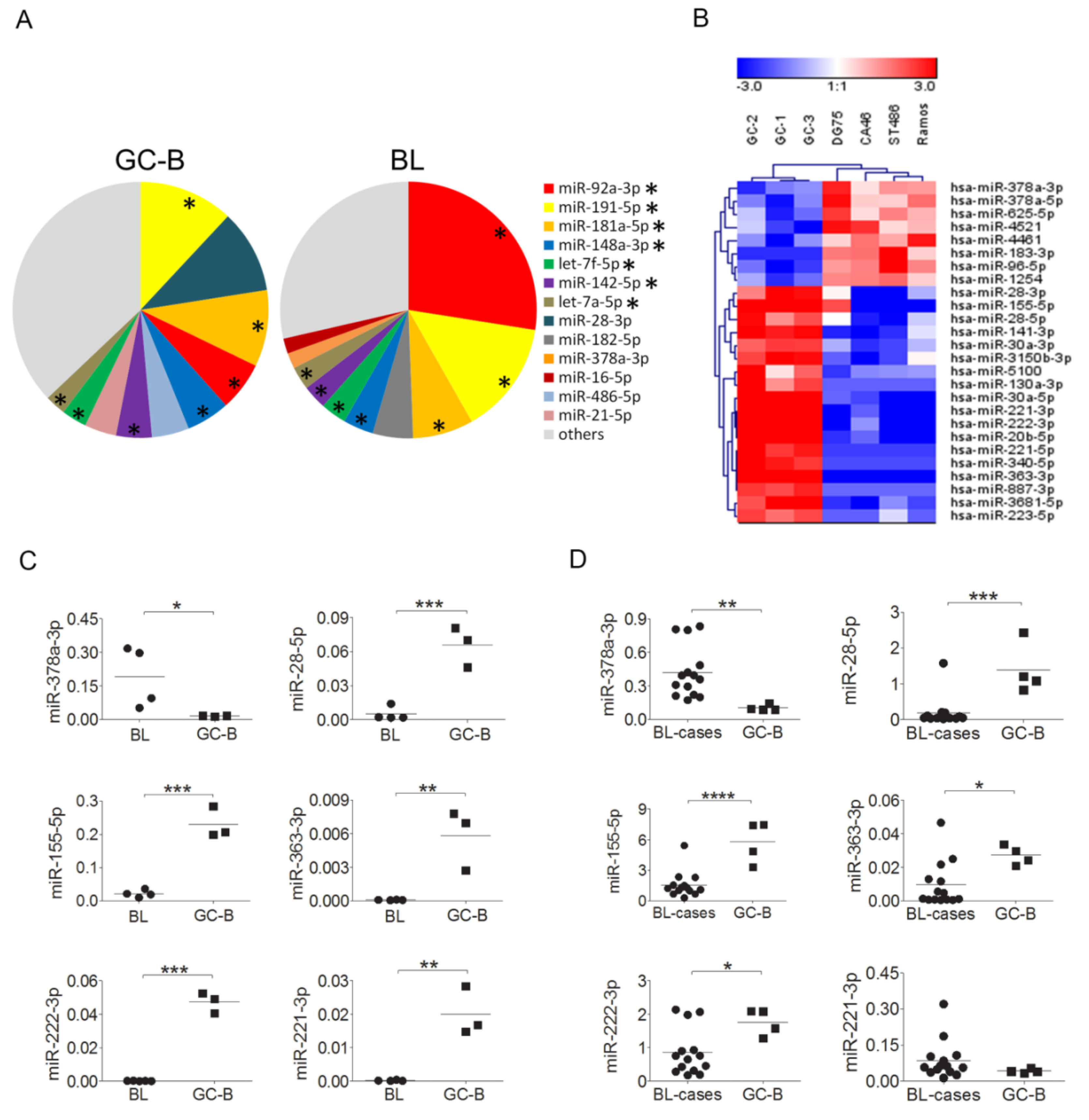
2.1. MiRNA Expression Profiling in BL and GC-B Cells
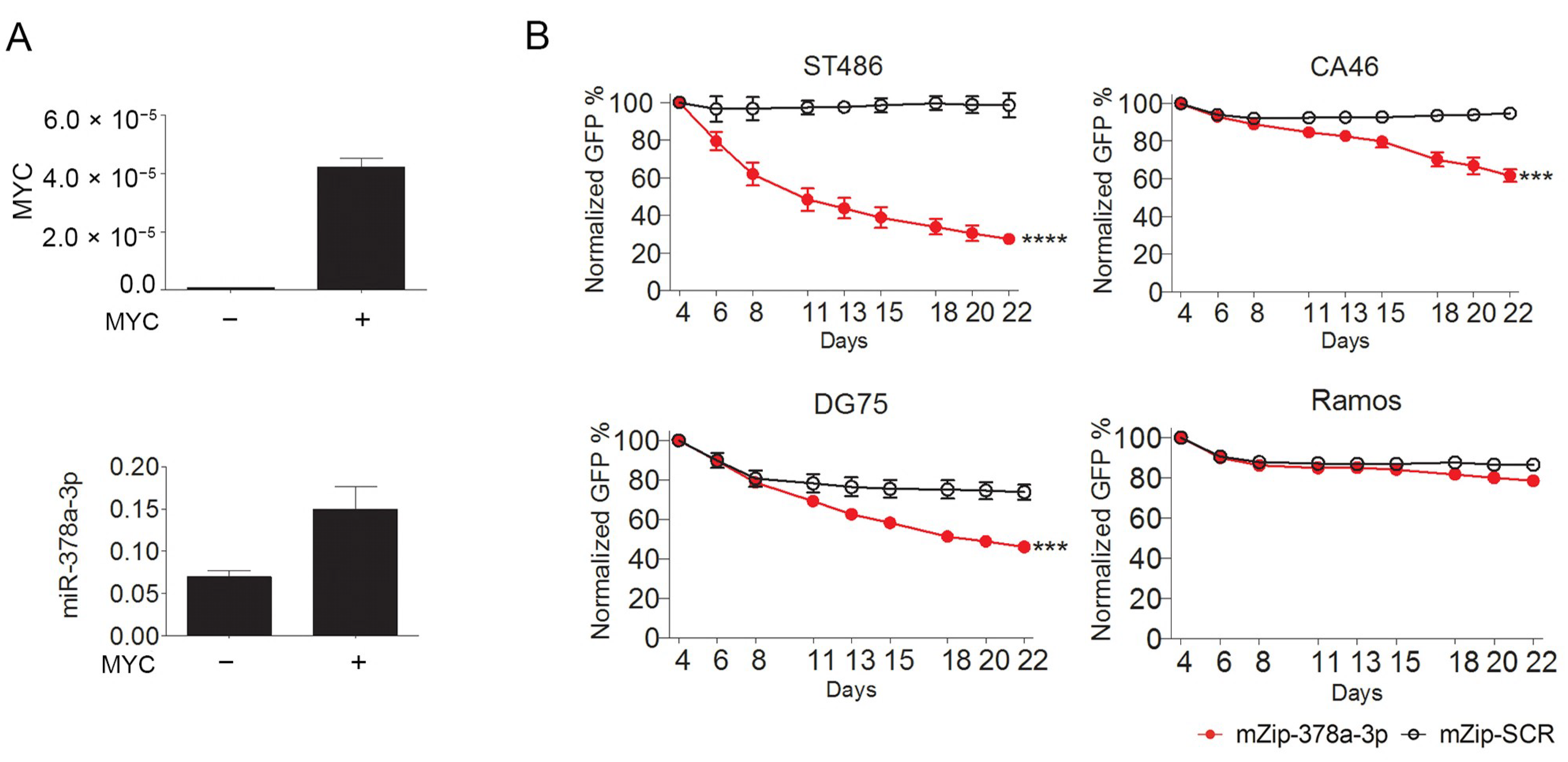
2.2. MYC-Induced miR-378a-3p Controls BL Cell Growth
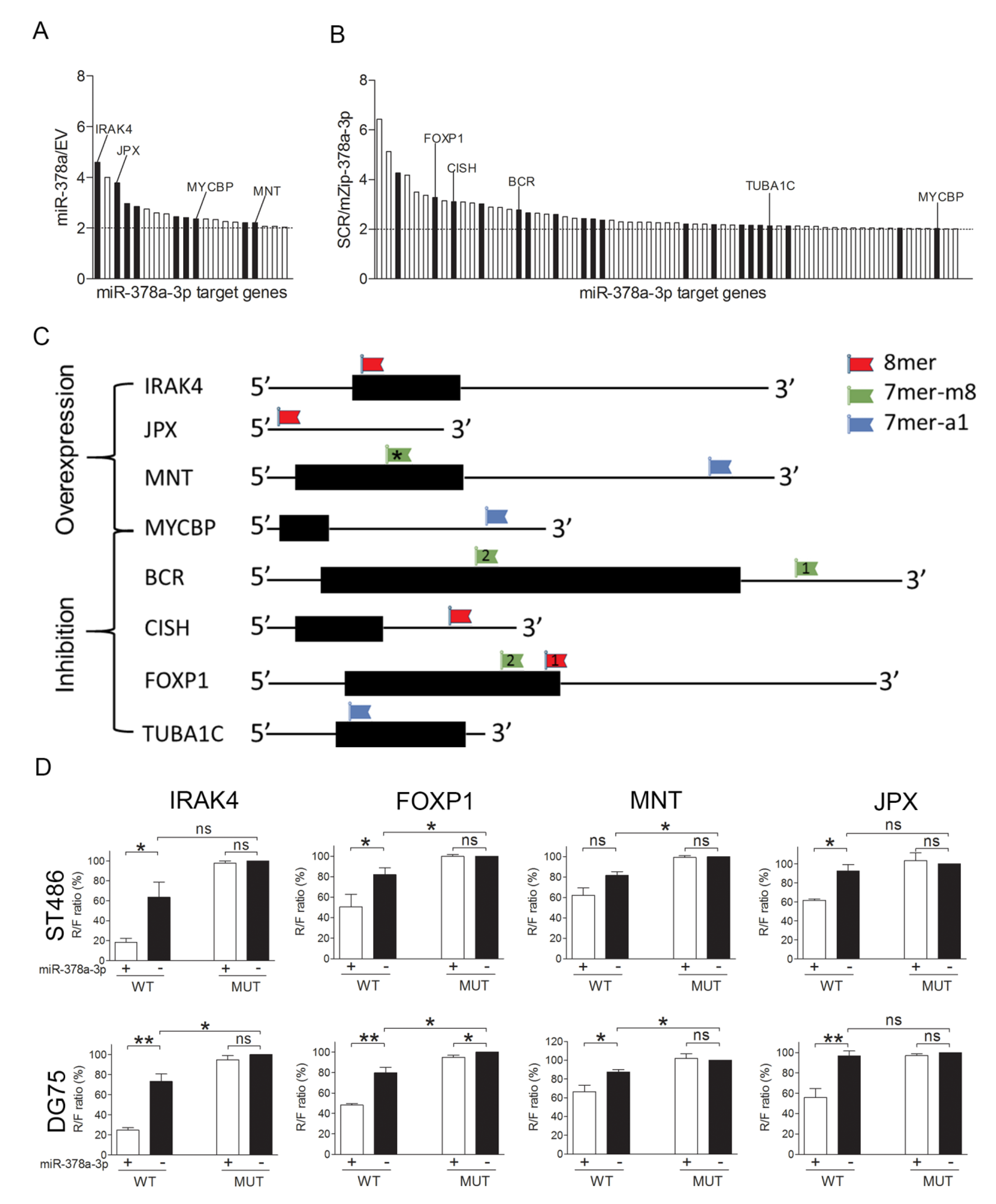
2.3. Identification of miR-378a-3p Targets
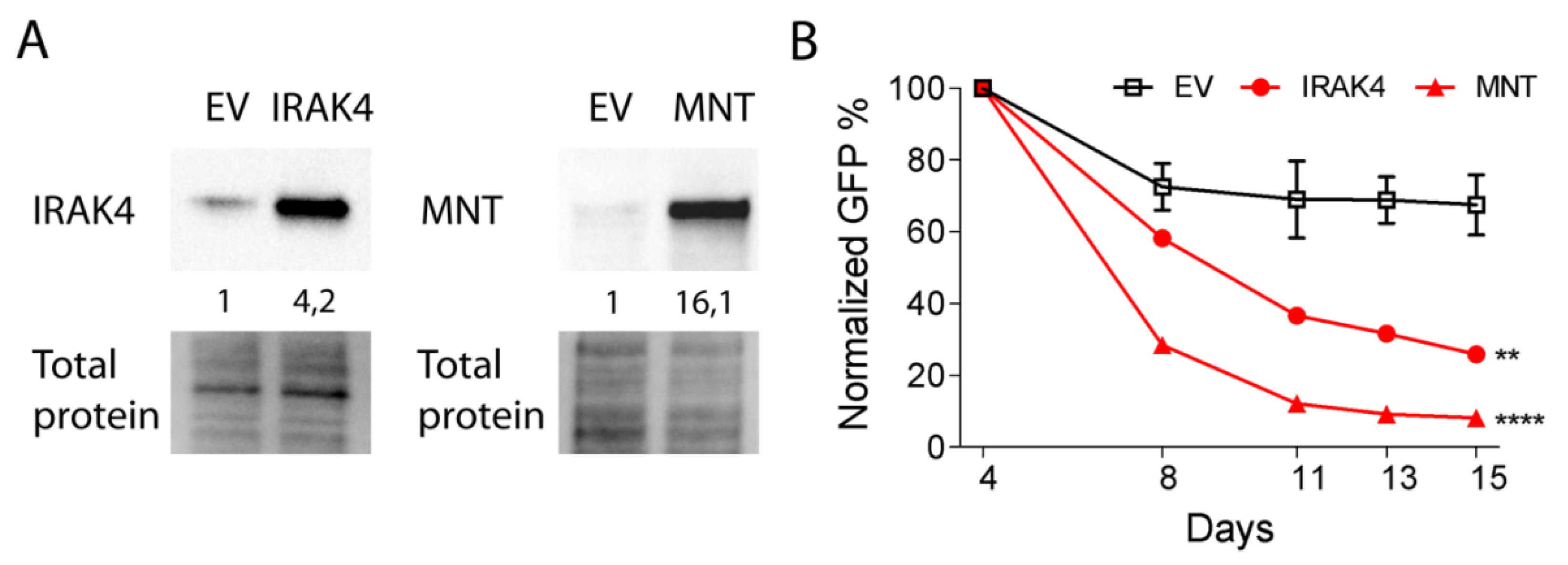
2.4. IRAK4 and MNT Are Involved in the Function of miR-378a-3p
3. Discussion
4. Materials and Methods
4.1. BL Cell Lines, Germinal Center (GC) B Cells, and BL Patient Material
4.2. RNA Isolation
4.3. Small RNA Library Preparation and Sequencing
4.4. qRT-PCR
4.5. Lentiviral Constructs, Transduction, and GFP Competition Assay
4.6. AGO2-IP Procedure
4.7. Western Blotting
4.8. Microarray Analysis
4.9. Identification of miR-378a-3p Seed Sites and Gene Ontology (GO) Terms
4.10. Cloning of miRNA Binding Sites and Luciferase Reporter Assay
4.11. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Burkitt, D.P. Etiology of Burkitt’s lymphoma—An alternative hypothesis to a vectored virus. J. Natl. Cancer Inst. 1969, 42, 19–28. [Google Scholar] [PubMed]
- Zech, L.; Haglund, U.; Nilsson, K.; Klein, G. Characteristic chromosomal abnormalities in biopsies and lymphoid-cell lines from patients with Burkitt and non-Burkitt lymphomas. Int. J. Cancer J. Int. Cancer 1976, 17, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Victora, G.D.; Dominguez-Sola, D.; Holmes, A.B.; Deroubaix, S.; Dalla-Favera, R.; Nussenzweig, M.C. Identification of human germinal center light and dark zone cells and their relationship to human B-cell lymphomas. Blood 2012, 120, 2240–2248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Selbach, M.; Schwanhausser, B.; Thierfelder, N.; Fang, Z.; Khanin, R.; Rajewsky, N. Widespread changes in protein synthesis induced by microRNAs. Nature 2008, 455, 58–63. [Google Scholar] [CrossRef]
- Baek, D.; Villen, J.; Shin, C.; Camargo, F.D.; Gygi, S.P.; Bartel, D.P. The impact of microRNAs on protein output. Nature 2008, 455, 64–71. [Google Scholar] [CrossRef] [Green Version]
- Ameres, S.L.; Zamore, P.D. Diversifying microRNA sequence and function. Nat. Rev. Mol. Cell Biol. 2013, 14, 475–488. [Google Scholar] [CrossRef]
- Lawrie, C.H. MicroRNAs in hematological malignancies. Blood Rev. 2013, 27, 143–154. [Google Scholar] [CrossRef]
- Oduor, C.I.; Kaymaz, Y.; Chelimo, K.; Otieno, J.A.; Ong’echa, J.M.; Moormann, A.M.; Bailey, J.A. Integrative microRNA and mRNA deep-sequencing expression profiling in endemic Burkitt lymphoma. BMC Cancer 2017, 17, 761. [Google Scholar] [CrossRef]
- Robertus, J.L.; Kluiver, J.; Weggemans, C.; Harms, G.; Reijmers, R.M.; Swart, Y.; Kok, K.; Rosati, S.; Schuuring, E.; van Imhoff, G.; et al. MiRNA profiling in B non-Hodgkin lymphoma: A MYC-related miRNA profile characterizes Burkitt lymphoma. Br. J. Haematol. 2010, 149, 896–899. [Google Scholar] [CrossRef]
- Lenze, D.; Leoncini, L.; Hummel, M.; Volinia, S.; Liu, C.G.; Amato, T.; De Falco, G.; Githanga, J.; Horn, H.; Nyagol, J.; et al. The different epidemiologic subtypes of Burkitt lymphoma share a homogenous micro RNA profile distinct from diffuse large B-cell lymphoma. Leukemia 2011, 25, 1869–1876. [Google Scholar] [CrossRef] [Green Version]
- Hezaveh, K.; Kloetgen, A.; Bernhart, S.H.; Mahapatra, K.D.; Lenze, D.; Richter, J.; Haake, A.; Bergmann, A.K.; Brors, B.; Burkhardt, B.; et al. Alterations of microRNA and microRNA-regulated messenger RNA expression in germinal center B-cell lymphomas determined by integrative sequencing analysis. Haematologica 2016, 101, 1380–1389. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Di Lisio, L.; Sanchez-Beato, M.; Gomez-Lopez, G.; Rodriguez, M.E.; Montes-Moreno, S.; Mollejo, M.; Menarguez, J.; Martinez, M.A.; Alves, F.J.; Pisano, D.G.; et al. MicroRNA signatures in B-cell lymphomas. Blood Cancer J. 2012, 2, e57. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bartolome-Izquierdo, N.; de Yebenes, V.G.; Alvarez-Prado, A.F.; Mur, S.M.; Lopez Del Olmo, J.A.; Roa, S.; Vazquez, J.; Ramiro, A.R. miR-28 regulates the germinal center reaction and blocks tumor growth in preclinical models of non-Hodgkin lymphoma. Blood 2017, 129, 2408–2419. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Wang, Z.; Dai, X.; Pan, J.; Ge, J.; Han, X.; Wu, Z.; Zhou, X.; Zhao, T. Re-expression of microRNA-150 induces EBV-positive Burkitt lymphoma differentiation by modulating c-Myb in vitro. Cancer Sci. 2013, 104, 826–834. [Google Scholar] [CrossRef] [PubMed]
- Dzikiewicz-Krawczyk, A.; Kok, K.; Slezak-Prochazka, I.; Robertus, J.L.; Bruining, J.; Tayari, M.M.; Rutgers, B.; de Jong, D.; Koerts, J.; Seitz, A.; et al. ZDHHC11 and ZDHHC11B are critical novel components of the oncogenic MYC-miR-150-MYB network in Burkitt lymphoma. Leukemia 2017, 31, 1470–1473. [Google Scholar] [CrossRef] [Green Version]
- Kluiver, J.; Haralambieva, E.; de Jong, D.; Blokzijl, T.; Jacobs, S.; Kroesen, B.J.; Poppema, S.; van den Berg, A. Lack of BIC and microRNA miR-155 expression in primary cases of Burkitt lymphoma. Genes Chromosom. Cancer 2006, 45, 147–153. [Google Scholar] [CrossRef]
- Sandhu, S.K.; Fassan, M.; Volinia, S.; Lovat, F.; Balatti, V.; Pekarsky, Y.; Croce, C.M. B-cell malignancies in microRNA Emu-miR-17~92 transgenic mice. Proc. Natl. Acad. Sci. USA 2013, 110, 18208–18213. [Google Scholar] [CrossRef] [Green Version]
- Teng, G.; Hakimpour, P.; Landgraf, P.; Rice, A.; Tuschl, T.; Casellas, R.; Papavasiliou, F.N. MicroRNA-155 is a negative regulator of activation-induced cytidine deaminase. Immunity 2008, 28, 621–629. [Google Scholar] [CrossRef] [Green Version]
- Dorsett, Y.; McBride, K.M.; Jankovic, M.; Gazumyan, A.; Thai, T.H.; Robbiani, D.F.; Di Virgilio, M.; San-Martin, B.R.; Heidkamp, G.; Schwickert, T.A.; et al. MicroRNA-155 suppresses activation-induced cytidine deaminase-mediated Myc-Igh translocation. Immunity 2008, 28, 630–638. [Google Scholar] [CrossRef] [Green Version]
- Feng, M.; Li, Z.; Aau, M.; Wong, C.H.; Yang, X.; Yu, Q. Myc/miR-378/TOB2/cyclin D1 functional module regulates oncogenic transformation. Oncogene 2011, 30, 2242–2251. [Google Scholar] [CrossRef] [Green Version]
- Pajic, A.; Spitkovsky, D.; Christoph, B.; Kempkes, B.; Schuhmacher, M.; Staege, M.S.; Brielmeier, M.; Ellwart, J.; Kohlhuber, F.; Bornkamm, G.W.; et al. Cell cycle activation by c-myc in a burkitt lymphoma model cell line. Int. J. Cancer 2000, 87, 787–793. [Google Scholar] [CrossRef]
- Li, H.; Dai, S.; Zhen, T.; Shi, H.; Zhang, F.; Yang, Y.; Kang, L.; Liang, Y.; Han, A. Clinical and biological significance of miR-378a-3p and miR-378a-5p in colorectal cancer. Eur. J. Cancer 2014, 50, 1207–1221. [Google Scholar] [CrossRef] [PubMed]
- Megiorni, F.; Cialfi, S.; McDowell, H.P.; Felsani, A.; Camero, S.; Guffanti, A.; Pizer, B.; Clerico, A.; De Grazia, A.; Pizzuti, A.; et al. Deep Sequencing the microRNA profile in rhabdomyosarcoma reveals down-regulation of miR-378 family members. BMC Cancer 2014, 14, 880. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, M.; Sun, X.; Yang, Y.; Jiao, W. Long non-coding RNA OIP5-AS1 promotes proliferation of lung cancer cells and leads to poor prognosis by targeting miR-378a-3p. Thorac. Cancer 2018, 9, 939–949. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.G.; Zhou, W.; Han, T.; Du, S.Q.; Li, Z.H.; Zhang, Z.; Shan, G.Y.; Kong, C.Z. MiR-378 suppresses prostate cancer cell growth through downregulation of MAPK1 in vitro and in vivo. Tumor Biol. 2016, 37, 2095–2103. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.H.; Yao, T.Z.; Liu, W. miR-378a-3p sensitizes ovarian cancer cells to cisplatin through targeting MAPK1/GRB2. Biomed. Pharmacother. 2018, 107, 1410–1417. [Google Scholar] [CrossRef]
- Liu, H.; Zhu, L.; Liu, B.; Yang, L.; Meng, X.; Zhang, W.; Ma, Y.; Xiao, H. Genome-wide microRNA profiles identify miR-378 as a serum biomarker for early detection of gastric cancer. Cancer Lett. 2012, 316, 196–203. [Google Scholar] [CrossRef]
- Yu, B.L.; Peng, X.H.; Zhao, F.P.; Liu, X.; Lu, J.; Wang, L.; Li, G.; Chen, H.H.; Li, X.P. MicroRNA-378 functions as an onco- miR in nasopharyngeal carcinoma by repressing TOB2 expression. Int. J. Oncol. 2014, 44, 1215–1222. [Google Scholar] [CrossRef] [Green Version]
- Qian, J.; Lin, J.; Qian, W.; Ma, J.C.; Qian, S.X.; Li, Y.; Yang, J.; Li, J.Y.; Wang, C.Z.; Chai, H.Y.; et al. Overexpression of miR-378 is frequent and may affect treatment outcomes in patients with acute myeloid leukemia. Leuk. Res 2013, 37, 765–768. [Google Scholar] [CrossRef]
- Tanaka, H.; Hazama, S.; Iida, M.; Tsunedomi, R.; Takenouchi, H.; Nakajima, M.; Tokumitsu, Y.; Kanekiyo, S.; Shindo, Y.; Tomochika, S.; et al. miR-125b-1 and miR-378a are predictive biomarkers for the efficacy of vaccine treatment against colorectal cancer. Cancer Sci. 2017, 108, 2229–2238. [Google Scholar] [CrossRef] [Green Version]
- Vasilevsky, N.A.; Ruby, C.E.; Hurlin, P.J.; Weinberg, A.D. OX40 engagement stabilizes Mxd4 and Mnt protein levels in antigen-stimulated T cells leading to an increase in cell survival. Eur. J. Immunol. 2011, 41, 1024–1034. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Campbell, K.J.; Vandenberg, C.J.; Anstee, N.S.; Hurlin, P.J.; Cory, S. Mnt modulates Myc-driven lymphomagenesis. Cell Death Differ. 2017, 24, 2117–2126. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, G.; Hurlin, P.J. MNT and Emerging Concepts of MNT-MYC Antagonism. Genes 2017, 8, 83. [Google Scholar] [CrossRef]
- Hooker, C.W.; Hurlin, P.J. Of Myc and Mnt. J. Cell Sci. 2006, 119, 208–216. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hurlin, P.J.; Zhou, Z.Q.; Toyo-oka, K.; Ota, S.; Walker, W.L.; Hirotsune, S.; Wynshaw-Boris, A. Deletion of Mnt leads to disrupted cell cycle control and tumorigenesis. EMBO J. 2003, 22, 4584–4596. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nilsson, J.A.; Maclean, K.H.; Keller, U.B.; Pendeville, H.; Baudino, T.A.; Cleveland, J.L. Mnt loss triggers Myc transcription targets, proliferation, apoptosis, and transformation. Mol. Cell Biol. 2004, 24, 1560–1569. [Google Scholar] [CrossRef] [Green Version]
- Walker, W.; Zhou, Z.Q.; Ota, S.; Wynshaw-Boris, A.; Hurlin, P.J. Mnt-Max to Myc-Max complex switching regulates cell cycle entry. J. Cell Biol. 2005, 169, 405–413. [Google Scholar] [CrossRef]
- Kim, T.W.; Staschke, K.; Bulek, K.; Yao, J.; Peters, K.; Oh, K.H.; Vandenburg, Y.; Xiao, H.; Qian, W.; Hamilton, T.; et al. A critical role for IRAK4 kinase activity in Toll-like receptor-mediated innate immunity. J. Exp. Med. 2007, 204, 1025–1036. [Google Scholar] [CrossRef] [Green Version]
- Kuppers, R. IRAK4 inhibition to shut down TLR signaling in autoimmunity and MyD88-dependent lymphomas. J. Exp. Med. 2015, 212, 2184. [Google Scholar] [CrossRef] [Green Version]
- Akhter, A.; Street, L.; Ghosh, S.; Burns, B.F.; Elyamany, G.; Shabani-Rad, M.T.; Stewart, D.A.; Mansoor, A. Concomitant high expression of Toll-like receptor (TLR) and B-cell receptor (BCR) signalling molecules has clinical implications in mantle cell lymphoma. Hematol. Oncol. 2017, 35, 79–86. [Google Scholar] [CrossRef]
- Akhter, A.; Masir, N.; Elyamany, G.; Phang, K.C.; Mahe, E.; Al-Zahrani, A.M.; Shabani-Rad, M.T.; Stewart, D.A.; Mansoor, A. Differential expression of Toll-like receptor (TLR) and B cell receptor (BCR) signaling molecules in primary diffuse large B-cell lymphoma of the central nervous system. J. Neuro-Oncol. 2015, 121, 289–296. [Google Scholar] [CrossRef] [PubMed]
- Ngo, V.N.; Young, R.M.; Schmitz, R.; Jhavar, S.; Xiao, W.; Lim, K.H.; Kohlhammer, H.; Xu, W.; Yang, Y.; Zhao, H.; et al. Oncogenically active MYD88 mutations in human lymphoma. Nature 2011, 470, 115–119. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kelly, P.N.; Romero, D.L.; Yang, Y.B.; Shaffer, A.L.; Chaudhary, D.; Robinson, S.; Miao, W.Y.; Rui, L.X.; Westlin, W.F.; Kapeller, R.; et al. Selective interleukin-1 receptor-associated kinase 4 inhibitors for the treatment of autoimmune disorders and lymphoid malignancy. J. Exp. Med. 2015, 212, 2189–2201. [Google Scholar] [CrossRef] [PubMed]
- Klapproth, K.; Sander, S.; Marinkovic, D.; Baumann, B.; Wirth, T. The IKK2/NF-{kappa}B pathway suppresses MYC-induced lymphomagenesis. Blood 2009, 114, 2448–2458. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gehringer, F.; Weissinger, S.E.; Moller, P.; Wirth, T.; Ushmorov, A. Physiological levels of the PTEN-PI3K-AKT axis activity are required for maintenance of Burkitt lymphoma. Leukemia 2019. [Google Scholar] [CrossRef]
- Hu, H.; Wang, B.; Borde, M.; Nardone, J.; Maika, S.; Allred, L.; Tucker, P.W.; Rao, A. Foxp1 is an essential transcriptional regulator of B cell development. Nat. Immunol. 2006, 7, 819–826. [Google Scholar] [CrossRef]
- Koon, H.B.; Ippolito, G.C.; Banham, A.H.; Tucker, P.W. FOXP1: A potential therapeutic target in cancer. Expert Opin. Ther. Targets 2007, 11, 955–965. [Google Scholar] [CrossRef]
- Sagardoy, A.; Martinez-Ferrandis, J.I.; Roa, S.; Bunting, K.L.; Aznar, M.A.; Elemento, O.; Shaknovich, R.; Fontan, L.; Fresquet, V.; Perez-Roger, I.; et al. Downregulation of FOXP1 is required during germinal center B-cell function. Blood 2013, 121, 4311–4320. [Google Scholar] [CrossRef] [Green Version]
- van Keimpema, M.; Gruneberg, L.J.; Mokry, M.; van Boxtel, R.; Koster, J.; Coffer, P.J.; Pals, S.T.; Spaargaren, M. FOXP1 directly represses transcription of proapoptotic genes and cooperates with NF-kappaB to promote survival of human B cells. Blood 2014, 124, 3431–3440. [Google Scholar] [CrossRef] [Green Version]
- Tian, D.; Sun, S.; Lee, J.T. The Long Noncoding RNA, Jpx, Is a Molecular Switch for X Chromosome Inactivation. Cell 2010, 143, 390–403. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Feng, L.; Tian, C.; Tang, Y.L.; Tang, Y.; Hu, F.Q. Long noncoding RNA-JPX predicts the poor prognosis of ovarian cancer patients and promotes tumor cell proliferation, invasion and migration by the PI3K/Akt/mTOR signaling pathway. Eur. Rev. Med. Pharmacol. 2018, 22, 8135–8144. [Google Scholar]
- Jin, M.; Ren, J.; Luo, M.; You, Z.; Fang, Y.; Han, Y.; Li, G.; Liu, H. Long noncoding RNA JPX correlates with poor prognosis and tumor progression in non-small cell lung cancer by interacting with miR-145-5p and CCND2. Carcinogenesis 2019. [Google Scholar] [CrossRef]
- Lin, X.Q.; Huang, Z.M.; Chen, X.; Wu, F.; Wu, W. XIST Induced by JPX Suppresses Hepatocellular Carcinoma by Sponging miR-155-5p. Yonsei Med. J. 2018, 59, 816–826. [Google Scholar] [CrossRef] [PubMed]
- Winkle, M.; van den Berg, A.; Tayari, M.; Sietzema, J.; Terpstra, M.; Kortman, G.; de Jong, D.; Visser, L.; Diepstra, A.; Kok, K.; et al. Long noncoding RNAs as a novel component of the Myc transcriptional network. FASEB J. 2015, 29, 2338–2346. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuan, Y.; Kluiver, J.; Koerts, J.; de Jong, D.; Rutgers, B.; Razak, F.R.A.; Terpstra, M.; Plaat, B.E.; Nolte, I.M.; Diepstra, A.; et al. miR-24-3p Is Overexpressed in Hodgkin Lymphoma and Protects Hodgkin and Reed-Sternberg Cells from Apoptosis. Am. J. Pathol. 2017, 187, 1343–1355. [Google Scholar] [CrossRef] [Green Version]
- Friedlander, M.R.; Chen, W.; Adamidi, C.; Maaskola, J.; Einspanier, R.; Knespel, S.; Rajewsky, N. Discovering microRNAs from deep sequencing data using miRDeep. Nat. Biotechnol. 2008, 26, 407–415. [Google Scholar] [CrossRef]
- Griffiths-Jones, S.; Grocock, R.J.; van Dongen, S.; Bateman, A.; Enright, A.J. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 2006, 34, D140–D144. [Google Scholar] [CrossRef]
- Kluiver, J.; Gibcus, J.H.; Hettinga, C.; Adema, A.; Richter, M.K.; Halsema, N.; Slezak-Prochazka, I.; Ding, Y.; Kroesen, B.J.; van den Berg, A. Rapid generation of microRNA sponges for microRNA inhibition. PLoS ONE 2012, 7, e29275. [Google Scholar] [CrossRef] [Green Version]
- Tan, L.P.; Seinen, E.; Duns, G.; de Jong, D.; Sibon, O.C.M.; Poppema, S.; Kroesen, B.J.; Kok, K.; van den Berg, A. A high throughput experimental approach to identify miRNA targets in human cells. Nucleic Acids Res. 2009, 37, e137. [Google Scholar] [CrossRef]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. eLife 2015, 4. [Google Scholar] [CrossRef]

| IP/T Ratio | pCDH (n = 9233) | mZip (n = 8944) | ||||
|---|---|---|---|---|---|---|
| EV | 378a | 378a/EV | SCR | mZip-378a-3p | SCR/mZip-378a-3p | |
| ≥2 | 611 | 586 | 20 | 741 | 878 | 63 |
| ≥4 | 117 | 117 | 2 | 171 | 196 | 4 |
| ≥8 | 25 | 18 | 0 | 53 | 34 | 0 |
| Gene | Transcript ID | IP/T Ratio | miR-378a-3p Binding Site | Growth-Related GO | ||||
|---|---|---|---|---|---|---|---|---|
| EV * | 378a | FC | 5′UTR | CDS | 3′UTR | |||
| IRAK4 | ENST00000613694 | 1.0 | 4.6 | 4.6 | 8m | yes | ||
| CDKN2A | ENST00000304494 | 1.0 | 4.0 | 4.0 | yes | |||
| JPX | ENST00000415215 | 1.5 | 5.8 | 3.9 | 8m ** | |||
| PLGRKT | ENST00000223864 | 1.0 | 3.0 | 3.0 | 8m | |||
| TMEM245 | ENST00000374586 | 1.0 | 2.9 | 2.9 | 7m8/8m | |||
| TOMM6 | ENST00000398884 | 1.5 | 4.1 | 2.7 | ||||
| CDK1 | ENST00000395284 | 1.0 | 2.6 | 2.6 | yes | |||
| FAM117A | ENST00000240364 | 1.4 | 3.6 | 2.6 | ||||
| WDR83OS | ENST00000596731 | 1.3 | 3.2 | 2.5 | 7m8 | |||
| CCNK | ENST00000389879 | 1.0 | 2.4 | 2.4 | 8m | yes | ||
| MYCBP | ENST00000397572 | 2.3 | 5.4 | 2.3 | 7mA1 | |||
| RNF34 | ENST00000392465 | 1.3 | 3.0 | 2.3 | yes | |||
| UBC | ENST00000339647 | 1.0 | 2.3 | 2.3 | yes | |||
| POP4 | ENST00000585603 | 1.0 | 2.3 | 2.3 | ||||
| MNT | ENST00000174618 | 1.2 | 2.7 | 2.3 | 7m8 | 7mA1 | yes | |
| HSPB1 | ENST00000248553 | 1.8 | 3.9 | 2.2 | 7mA1 | yes | ||
| INAFM1 | ENST00000552360 | 1.3 | 2.8 | 2.2 | ||||
| PCNA | ENST00000379160 | 1.0 | 2.1 | 2.1 | yes | |||
| SP100 | ENST00000264052 | 1.2 | 2.5 | 2.1 | ||||
| RPP25L | ENST00000297613 | 1.2 | 2.4 | 2.0 | ||||
| Gene | Transcript ID | IP/T Ratio | miR-378a-3p Binding Site | Growth-Related GO | ||||
|---|---|---|---|---|---|---|---|---|
| mZip-378a-3p * | SCR | FC | 5′UTR | CDS | 3′UTR | |||
| DYNLRB1 | ENST00000357156 | 1.0 | 6.4 | 6.4 | ||||
| VPS18 | ENST00000220509 | 8.5 | 43.4 | 5.1 | ||||
| NAPA-AS1 | ENST00000594367 | 1.2 | 5.2 | 4.3 | 7mA1 ** | |||
| C11orf95 | ENST00000433688 | 1.0 | 4.2 | 4.2 | ||||
| HOMEZ | ENST00000357460 | 1.2 | 4.3 | 3.5 | ||||
| TMEM79 | ENST00000405535 | 1.7 | 5.7 | 3.4 | ||||
| FOXP1 | ENST00000493089 | 4.5 | 14.9 | 3.3 | 7m8/8m | yes | ||
| PCIF1 | ENST00000372409 | 1.0 | 3.2 | 3.2 | 7m8 | |||
| ATP6V0C | ENST00000330398 | 1.0 | 3.1 | 3.1 | ||||
| CISH | ENST00000348721 | 2.1 | 6.6 | 3.1 | 8m | yes | ||
| lnc-FOXB1-8 | lnc-FOXB1-8:1 | 1.0 | 3.3 | 3.1 | ||||
| MT1B | ENST00000334346 | 1.1 | 3.5 | 3.1 | yes | |||
| lnc-EGLN1-1 | lnc-EGLN1-1:6 | 1.0 | 2.9 | 2.9 | ||||
| NELFA | ENST00000382882 | 1.0 | 3.0 | 2.9 | ||||
| BCR | ENST00000305877 | 1.0 | 2.8 | 2.8 | 7m8 | 7m8 | yes | |
| MT1L | ENST00000565768 | 1.2 | 3.2 | 2.8 | ||||
| FAT3 | ENST00000409404 | 1.0 | 2.7 | 2.7 | 8m | 7mA1 | ||
| TRAF3IP2-AS1 | ENST00000525151 | 1.0 | 2.7 | 2.7 | ||||
| C22orf39 | ENST00000611555 | 1.0 | 2.6 | 2.6 | ||||
| KRTCAP2 | ENST00000295682 | 1.6 | 4.1 | 2.6 | ||||
| LINC01122 | ENST00000427421 | 11.2 | 29 | 2.6 | 7mA1 ** | |||
| ACTG1P20 | ENSG00000241547 | 1.3 | 3.2 | 2.5 | ||||
| lnc-KRTAP5-10-1 | lnc-KRTAP5-10-1:1 | 1.5 | 3.5 | 2.4 | 7m8 ** | |||
| MT1E | ENST00000306061 | 1.1 | 2.8 | 2.4 | yes | |||
| NUDT19 | ENST00000397061 | 1.0 | 2.5 | 2.4 | 7mA1 | |||
| PRDX4 | ENST00000379341 | 1.0 | 2.4 | 2.4 | yes | |||
| TMEM258 | ENST00000537328 | 1.1 | 2.6 | 2.4 | ||||
| XLOC_l2_005952 | TCONS_l2_00011050 | 6.6 | 15.6 | 2.4 | 7m8 | |||
| BTG3 | ENST00000629582 | 1.6 | 3.7 | 2.3 | yes | |||
| EVI5L | ENST00000270530 | 1.3 | 3.0 | 2.3 | ||||
| LINC01534 | ENST00000433232 | 1.0 | 2.3 | 2.3 | ||||
| lnc-ADA-1 | lnc-ADA-1:2 | 1.0 | 2.3 | 2.3 | ||||
| lnc-ZNF431-4 | lnc-ZNF431-4:1 | 1.1 | 2.6 | 2.3 | ||||
| MT1A | ENST00000290705 | 2.5 | 5.6 | 2.3 | yes | |||
| CSRP2 | ENST00000311083 | 2.7 | 5.9 | 2.2 | ||||
| FYCO1 | ENST00000296137 | 1.0 | 2.2 | 2.2 | 7m8 | |||
| HYAL3 | ENST00000336307 | 2.0 | 4.3 | 2.2 | ||||
| KCNQ1 | ENST00000632153 | 1.0 | 2.2 | 2.2 | ||||
| lnc-RP11-158I9.5.1-2 | TCONS_00019776 | 1.5 | 3.3 | 2.2 | ||||
| PCNX | ENST00000304743 | 3.8 | 8.3 | 2.2 | 7mA1/8m | |||
| PRSS36 | ENST00000268281 | 1.4 | 2.8 | 2.2 | ||||
| SMARCA4 | ENST00000344626 | 1.0 | 2.2 | 2.2 | 7m8 | yes | ||
| TNRC6C | ENST00000335749 | 8.5 | 18.3 | 2.2 | 7m8 | |||
| TOLLIP | ENST00000317204 | 1.3 | 2.8 | 2.2 | 7mA1 | |||
| ARF4 | ENST00000303436 | 1.0 | 2.1 | 2.1 | yes | |||
| ATG4D | ENST00000309469 | 1.4 | 2.8 | 2.1 | yes | |||
| CSE1L | ENST00000262982 | 1.2 | 2.5 | 2.1 | yes | |||
| lnc-PCF11-1 | lnc-PCF11-1:12 | 1.5 | 3.2 | 2.1 | ||||
| MARS | ENST00000262027 | 1.7 | 3.6 | 2.1 | ||||
| NANS | ENST00000210444 | 1.0 | 2.1 | 2.1 | ||||
| ORMDL2 | ENST00000243045 | 1.1 | 2.3 | 2.1 | ||||
| PFKFB2 | ENST00000367080 | 1.4 | 3.0 | 2.1 | 7m8 | 7mA1 | ||
| PGM2L1 | ENST00000298198 | 1.8 | 3.7 | 2.1 | ||||
| PTPN23 | ENST00000265562 | 1.5 | 3.1 | 2.1 | ||||
| TSACC | ENST00000368255 | 1.5 | 3.1 | 2.1 | ||||
| TUBA1C | ENST00000301072 | 1.1 | 2.4 | 2.1 | 7mA1 | yes | ||
| XLOC_l2_013031 | TCONS_l2_00024809 | 6.5 | 13.7 | 2.1 | ||||
| C15orf61 | ENST00000342683 | 1.3 | 2.6 | 2.0 | 8m | |||
| LAT | ENST00000360872 | 1.0 | 2.0 | 2.0 | ||||
| LOC101929494 | N/A | 1.6 | 3.2 | 2.0 | ||||
| MYCBP | ENST00000397572 | 1.5 | 3.1 | 2.0 | 7mA1 | |||
| NDRG4 | ENST00000394279 | 1.1 | 2.2 | 2.0 | yes | |||
| TUBE1 | ENST00000368662 | 1.1 | 2.2 | 2.0 | yes | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Niu, F.; Dzikiewicz-Krawczyk, A.; Koerts, J.; de Jong, D.; Wijenberg, L.; Fernandez Hernandez, M.; Slezak-Prochazka, I.; Winkle, M.; Kooistra, W.; van der Sluis, T.; et al. MiR-378a-3p Is Critical for Burkitt Lymphoma Cell Growth. Cancers 2020, 12, 3546. https://doi.org/10.3390/cancers12123546
Niu F, Dzikiewicz-Krawczyk A, Koerts J, de Jong D, Wijenberg L, Fernandez Hernandez M, Slezak-Prochazka I, Winkle M, Kooistra W, van der Sluis T, et al. MiR-378a-3p Is Critical for Burkitt Lymphoma Cell Growth. Cancers. 2020; 12(12):3546. https://doi.org/10.3390/cancers12123546
Chicago/Turabian StyleNiu, Fubiao, Agnieszka Dzikiewicz-Krawczyk, Jasper Koerts, Debora de Jong, Laura Wijenberg, Margot Fernandez Hernandez, Izabella Slezak-Prochazka, Melanie Winkle, Wierd Kooistra, Tineke van der Sluis, and et al. 2020. "MiR-378a-3p Is Critical for Burkitt Lymphoma Cell Growth" Cancers 12, no. 12: 3546. https://doi.org/10.3390/cancers12123546
APA StyleNiu, F., Dzikiewicz-Krawczyk, A., Koerts, J., de Jong, D., Wijenberg, L., Fernandez Hernandez, M., Slezak-Prochazka, I., Winkle, M., Kooistra, W., van der Sluis, T., Rutgers, B., Terpstra, M. M., Kok, K., Kluiver, J., & van den Berg, A. (2020). MiR-378a-3p Is Critical for Burkitt Lymphoma Cell Growth. Cancers, 12(12), 3546. https://doi.org/10.3390/cancers12123546








