Characterization of a PERK Kinase Inhibitor with Anti-Myeloma Activity
Simple Summary
Abstract
1. Introduction
2. Results
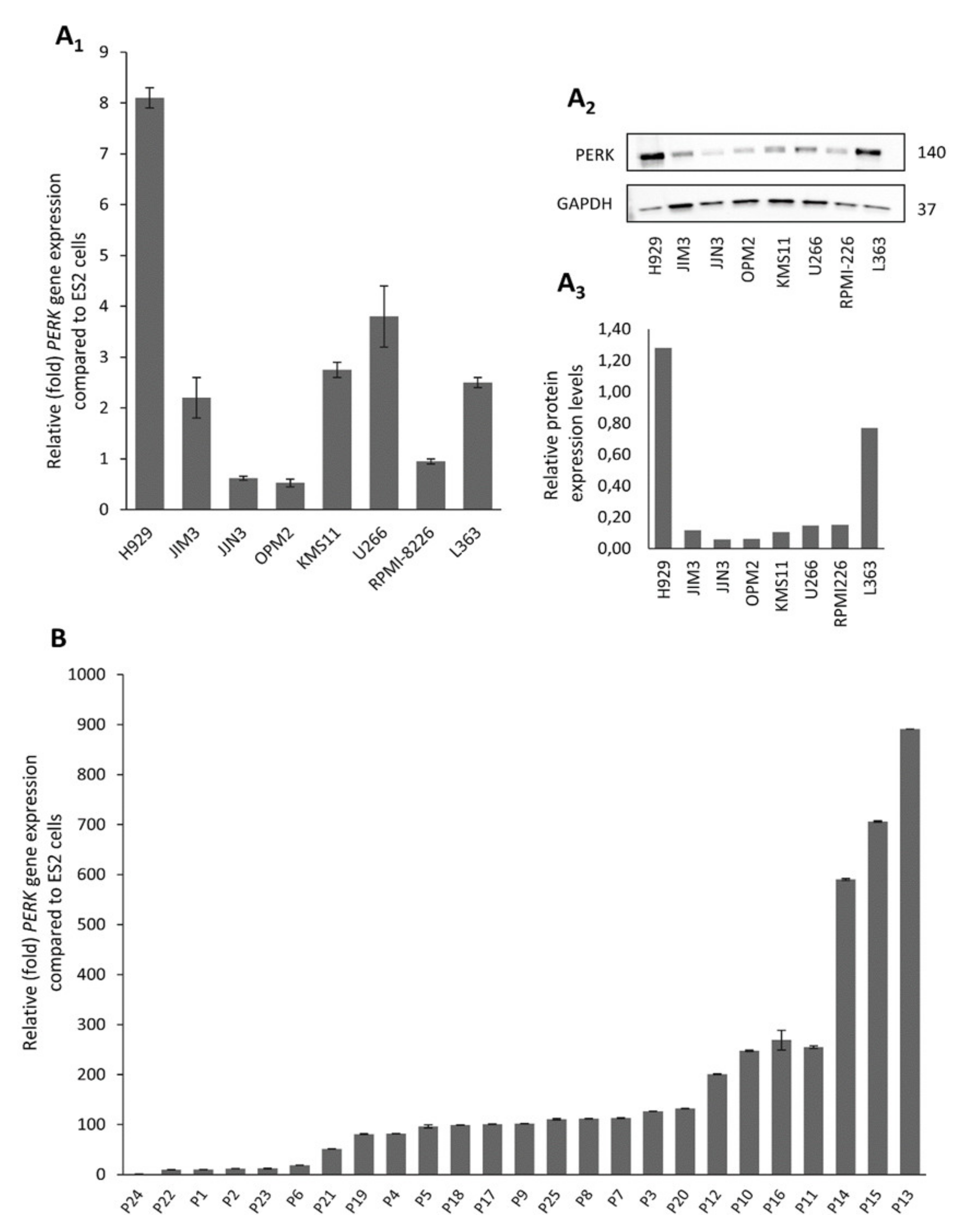
2.1. Human Myeloma Cell Lines and CD138+ Myeloma Cells from MM Patients Express High Levels of PERK
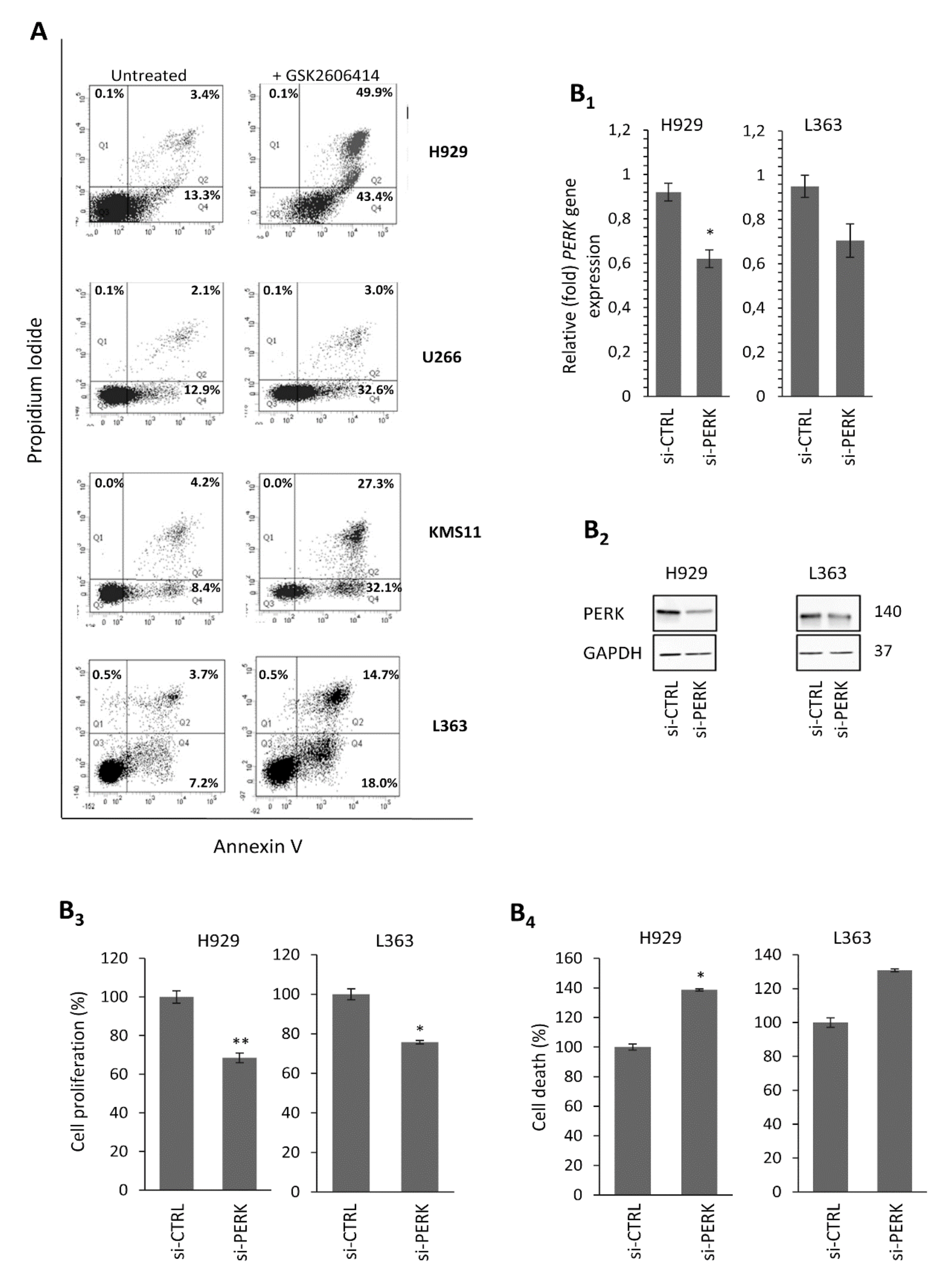
2.2. The PERK Specific Inhibitor GSK2606414 Decreases MM Cell Survival and Induces MM Cell Apoptosis
2.3. PERK Downregulation Reduces Cell Survival and Induces Apoptosis of MM Cells
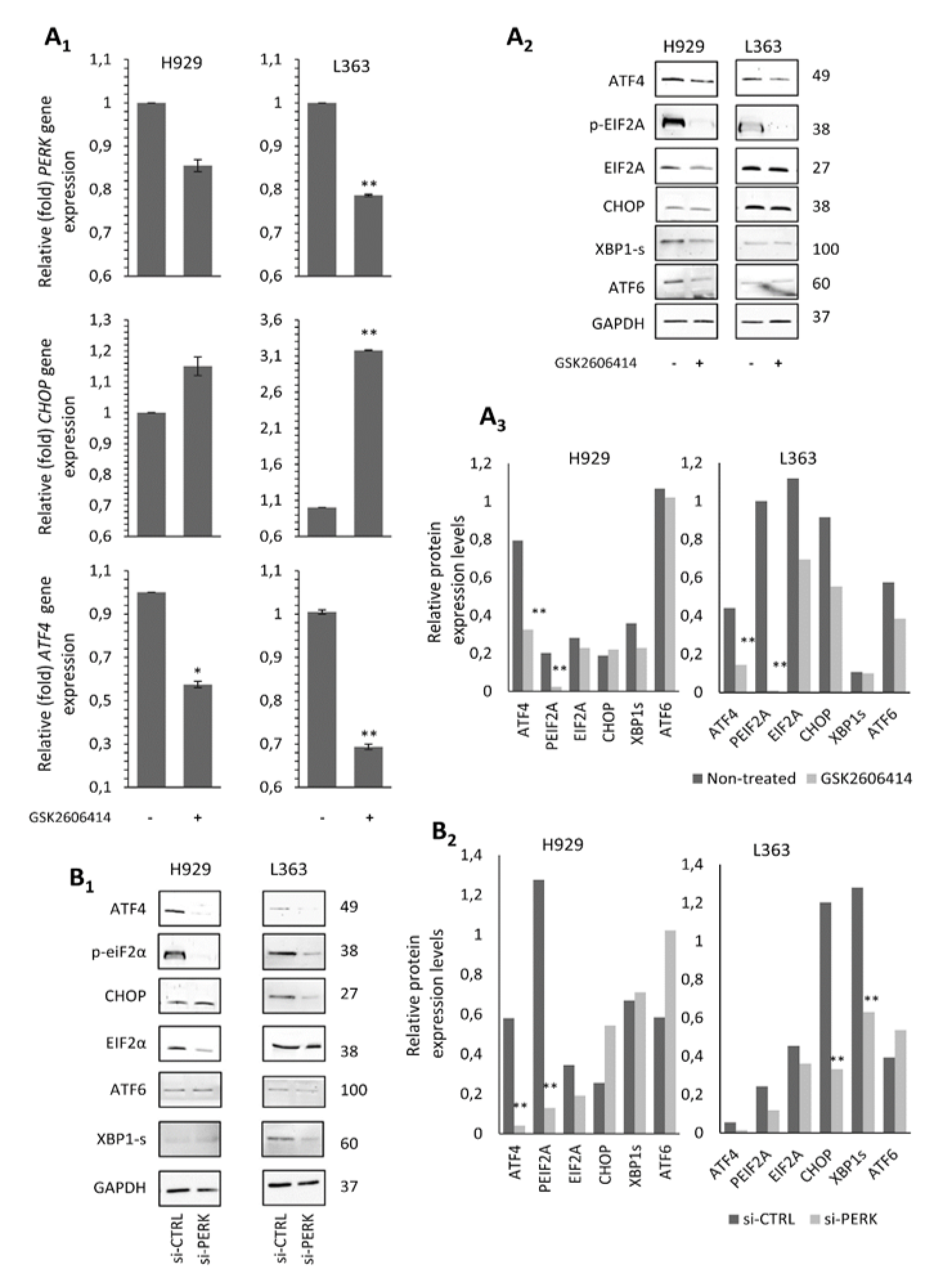
2.4. The GSK2606414 Inhibitor Suppresses the PERK Signaling Axis in MM Cells
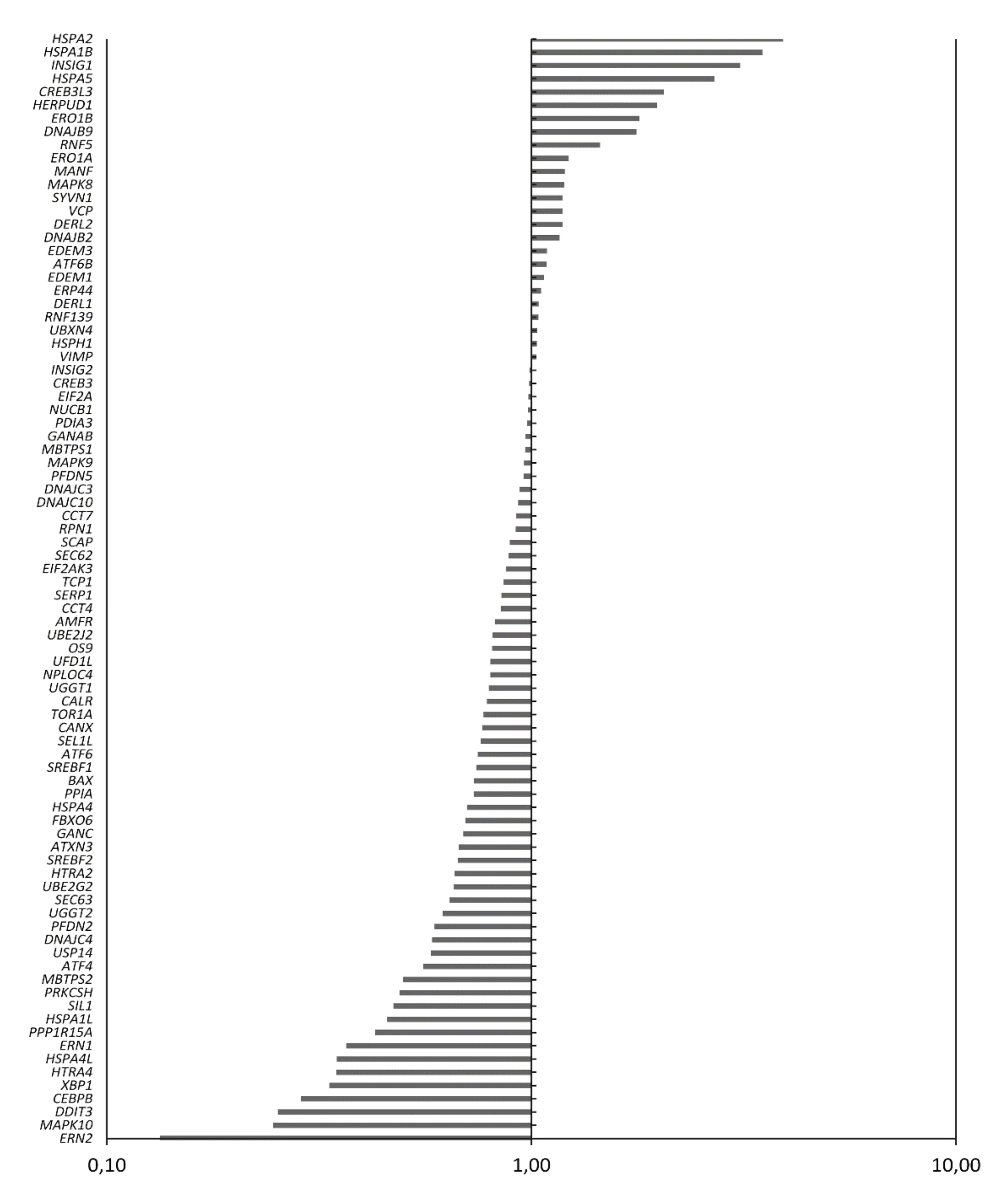
2.5. Treatment of MM Cells with GSK2606414 Differentially Regulates UPR-Related Genes
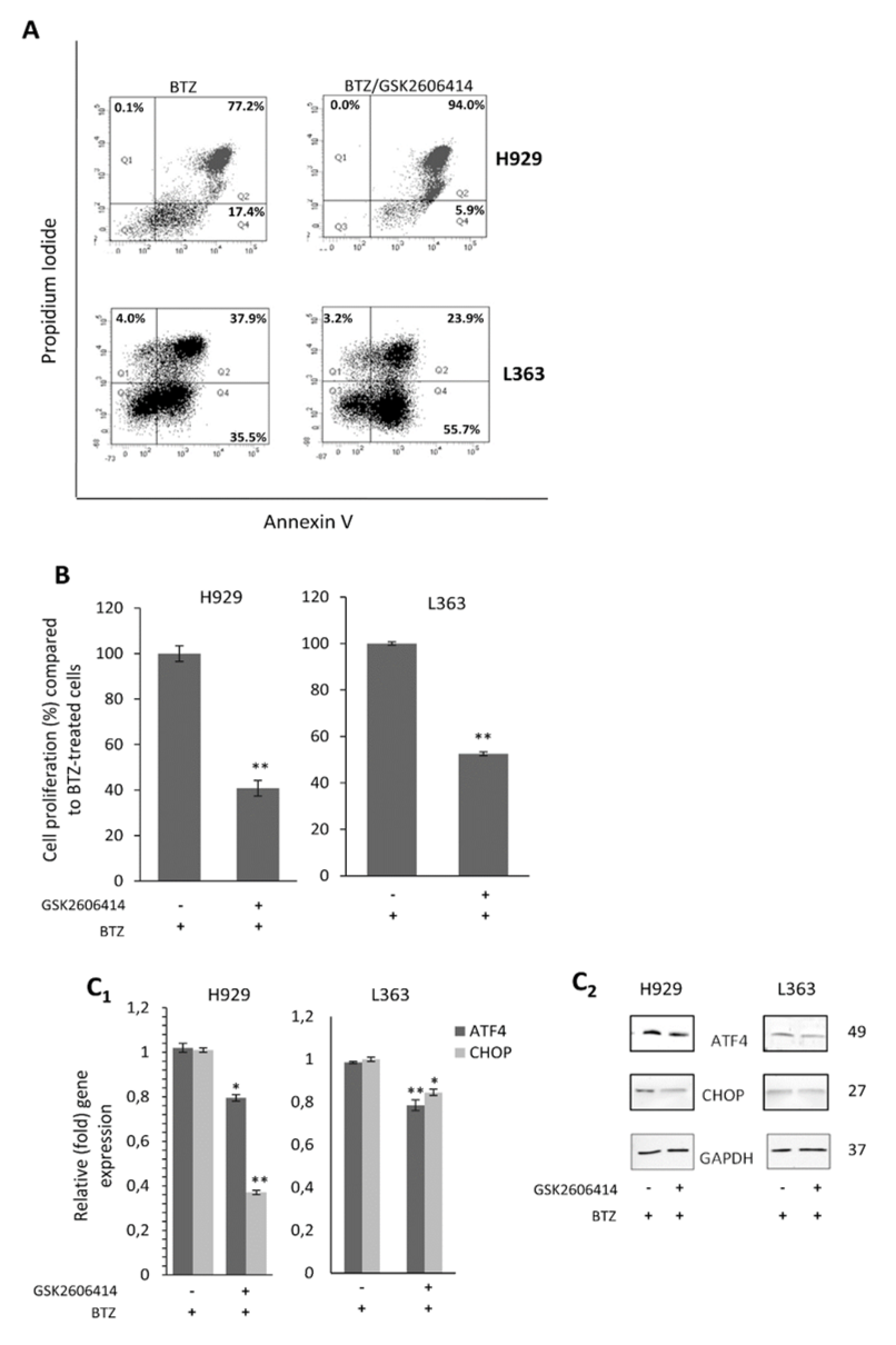
2.6. GSK2606414 Exerts Synergistic to BTZ Effects in MM Cells
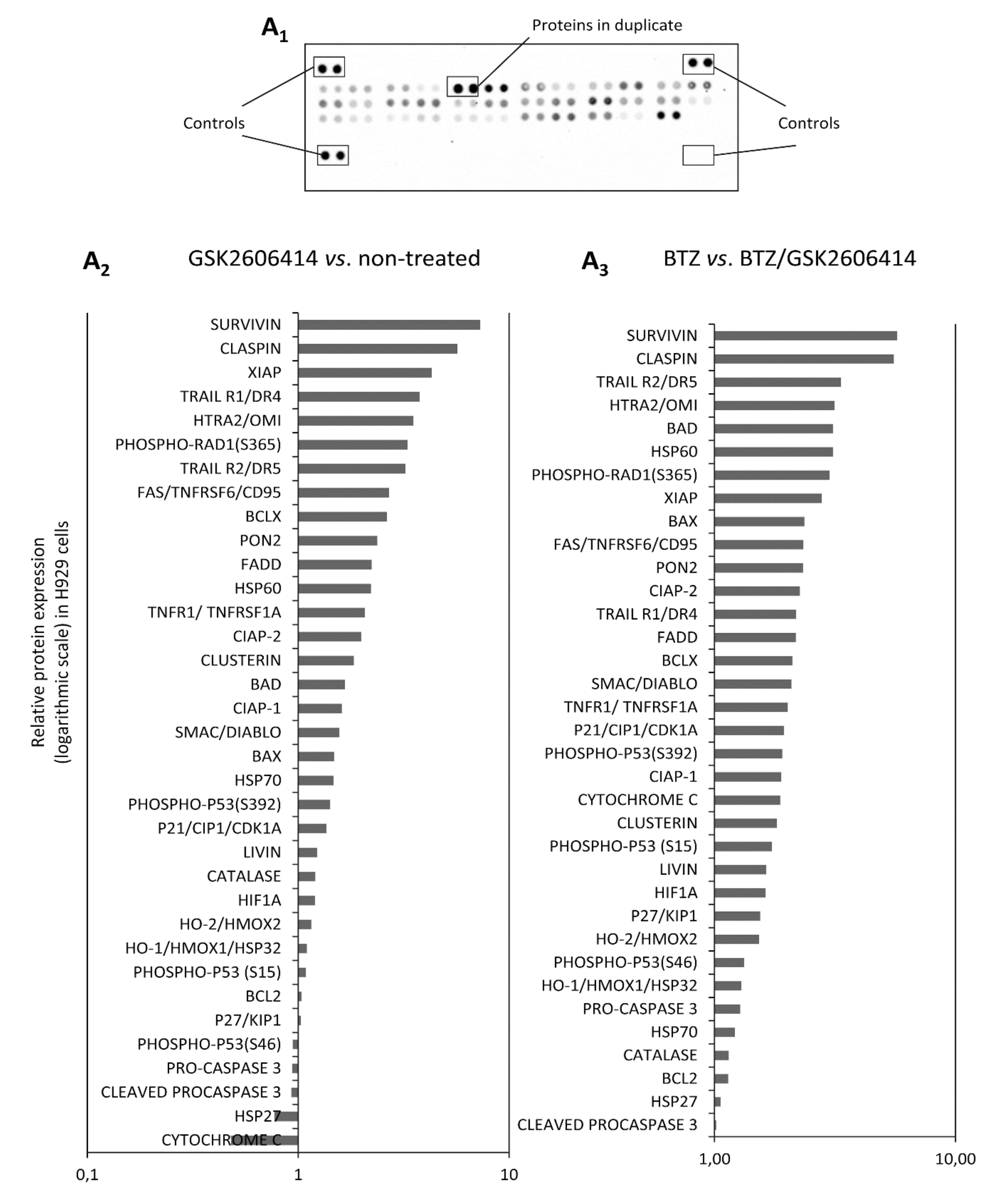
2.7. GSK2606414 Differentially Regulates Apoptotic Pathways in MM cells
3. Discussion
4. Materials and Methods
4.1. CD138+ Plasma Cell and Human MM Cell Lines
4.2. Reagents
4.3. Cell Survival/Viability Assay
4.4. Apoptosis Assay
4.5. RNA Extraction, Quantification and Amplification
4.6. Real-Time Polymerase Chain Reaction
4.7. Polymerase Chain Reaction (PCR)
4.8. siRNA Transfection
4.9. Immunoblotting Analysis
4.10. UPR Gene Expression Profiling
4.11. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Davenport, E.L.; Moore, H.E.; Dunlop, A.S.; Sharp, S.Y.; Workman, P.; Morgan, G.J.; Davies, F.E. Heat shock protein inhibition is associated with activation of the unfolded protein response pathway in myeloma plasma cells. Blood 2007, 110, 2641–2649. [Google Scholar] [CrossRef] [PubMed]
- Schröder, M.; Kaufman, R.J. ER stress and the unfolded protein response. Mutat. Res. Mol. Mech. Mutagen. 2005, 569, 29–63. [Google Scholar] [CrossRef] [PubMed]
- Bertolotti, A.; Zhang, Y.; Hendershot, L.M.; Harding, H.P.; Ron, D. Dynamic interaction of BiP and ER stress transducers in the unfolded-protein response. Nat. Cell Biol. 2000, 2, 326–332. [Google Scholar] [CrossRef] [PubMed]
- Urano, F.; Wang, X.; Bertolotti, A.; Zhang, Y.; Chung, P.; Harding, H.P.; Ron, D. Coupling of Stress in the ER to Activation of JNK Protein Kinases by Transmembrane Protein Kinase IRE1. Science 2000, 287, 664–666. [Google Scholar] [CrossRef] [PubMed]
- Kimata, Y.; Kohno, K. How is ER stress sensed? Tanpakushitsu kakusan koso. Protein Nucleic Acid Enzyme 2004, 49, 998–1001. [Google Scholar]
- Xu, C.; Bailly-Maitre, B.; Reed, J.C. Endoplasmic reticulum stress: Cell life and death decisions. J. Clin. Investig. 2005, 115, 2656–2664. [Google Scholar] [CrossRef]
- Harding, H.P.; Zhang, Y.; Bertolotti, A.; Zeng, H.; Ron, D. Perk Is Essential for Translational Regulation and Cell Survival during the Unfolded Protein Response. Mol. Cell 2000, 5, 897–904. [Google Scholar] [CrossRef]
- Harding, H.P.; Zhang, Y.; Ron, D. Protein translation and folding are coupled by an endoplasmic-reticulum-resident kinase. Nature 1999, 397, 271–274. [Google Scholar] [CrossRef]
- Scheuner, D.; Song, B.; McEwen, E.; Liu, C.; Laybutt, R.; Gillespie, P.; Saunders, T.L.; Bonner-Weir, S.; Kaufman, R.J. Translational Control Is Required for the Unfolded Protein Response and In Vivo Glucose Homeostasis. Mol. Cell 2001, 7, 1165–1176. [Google Scholar] [CrossRef]
- Ron, D. Translational control in the endoplasmic reticulum stress response. J. Clin. Invest. 2002, 110, 1383–1388. [Google Scholar] [CrossRef]
- Harding, H.P.; Zhang, Y.; Zeng, H.; Novoa, I.; Lu, P.D.; Calfon, M.; Sadri, N.; Yun, C.; Popko, B.; Paules, R.S.; et al. An Integrated Stress Response Regulates Amino Acid Metabolism and Resistance to Oxidative Stress. Mol. Cell 2003, 11, 619–633. [Google Scholar] [CrossRef]
- Zinszner, H.; Kuroda, M.; Wang, X.; Batchvarova, N.; Lightfoot, R.T.; Remotti, H.E.; Stevens, J.L.; Ron, D. CHOP is implicated in programmed cell death in response to impaired function of the endoplasmic reticulum. Genes Dev. 1998, 12, 982–995. [Google Scholar] [CrossRef] [PubMed]
- Harding, H.P.; Novoa, I.; Zhang, Y.; Zeng, H.; Wek, R.; Schapira, M.; Ron, D. Regulated translation initiation controls stress-induced gene expression in mammalian cells. Mol. Cell 2000, 6, 1099–1108. [Google Scholar] [CrossRef]
- Ma, Y.; Brewer, J.W.; Diehl, J.A.; Hendershot, L.M. Two Distinct Stress Signaling Pathways Converge Upon the CHOP Promoter During the Mammalian Unfolded Protein Response. J. Mol. Biol. 2002, 318, 1351–1365. [Google Scholar] [CrossRef]
- Ranganathan, A.C.; Ojha, S.; Kourtidis, A.; Conklin, D.S.; Aguirre-Ghiso, J.A. Dual function of pancreatic endoplasmic reticulum kinase in tumor cell growth arrest and survival. Cancer Res. 2008, 68, 3260–3268. [Google Scholar] [CrossRef]
- Estone, S.; Ho, Y.; Li, X.; Jamison, S.; Harding, H.P.; Ron, D.; Lin, W. Dual role of the integrated stress response in medulloblastoma tumorigenesis. Oncotarget 2016, 7, 64124–64135. [Google Scholar] [CrossRef]
- Ho, Y.; Li, X.; Jamison, S.; Harding, H.P.; McKinnon, P.J.; Ron, D.; Lin, W. PERK Activation Promotes Medulloblastoma Tumorigenesis by Attenuating Premalignant Granule Cell Precursor Apoptosis. Am. J. Pathol. 2016, 186, 1939–1951. [Google Scholar] [CrossRef]
- Kraus, M.; Malenke, E.; Gogel, J.; Müller, H.; Overkleeft, H.S.; Ovaa, H.; Koscielniak, E.; Hartmann, J.T.; Driessen, C.; Rückrich, T. Ritonavir induces endoplasmic reticulum stress and sensitizes sarcoma cells toward bortezomib-induced apoptosis. Mol. Cancer Ther. 2008, 7, 1940–1948. [Google Scholar] [CrossRef]
- Lust, S.; Vanhoecke, B.; Van Gele, M.; Boelens, J.; Van Melckebeke, H.; Kaileh, M.; Berghe, W.V.; Haegeman, G.; Philippé, J.; Bracke, M.; et al. Xanthohumol activates the proapoptotic arm of the unfolded protein response in chronic lymphocytic leukemia. Anticancer Res. 2009, 29, 3797–3805. [Google Scholar]
- Yan, Y.; Gao, Y.Y.; Liu, B.Q.; Niu, X.F.; Zhuang, Y.; Wang, H.Q. Resveratrol-induced cytotoxicity in human Burkitt’s lymphoma cells is coupled to the unfolded protein response. BMC Cancer 2010, 10, 445. [Google Scholar] [CrossRef]
- Fribley, A.; Cruz, P.G.; Miller, J.R.; Callaghan, M.U.; Cai, P.; Narula, N.; Neubig, R.R.; Showalter, H.D.; Larsen, S.D.; Kirchhoff, P.D.; et al. Complementary cell-based high-throughput screens identify novel modulators of the unfolded protein response. J. Biomol. Screen 2011, 16, 825–835. [Google Scholar] [CrossRef] [PubMed]
- Qiao, S.; Cabello, C.M.; Lamore, S.D.; Lesson, J.L.; Wondrak, G.T. D-Penicillamine targets metastatic melanoma cells with induction of the unfolded protein response (UPR) and Noxa (PMAIP1)-dependent mitochondrial apoptosis. Apoptosis 2012, 17, 1079–1094. [Google Scholar] [CrossRef] [PubMed]
- Sailaja, G.S.; Bhoopathi, P.; Gorantla, B.; Chetty, C.; Gogineni, V.R.; Velpula, K.K.; Gondi, C.S.; Rao, J.S. The secreted protein acidic and rich in cysteine (SPARC) induces endoplasmic reticulum stress leading to autophagy-mediated apoptosis in neuroblastoma. Int. J. Oncol. 2012, 42, 188–196. [Google Scholar] [CrossRef] [PubMed]
- White-Gilbertson, S.; Hua, Y.; Liu, B. The role of endoplasmic reticulum stress in maintaining and targeting multiple myeloma: A double-edged sword of adaptation and apoptosis. Front. Genet. 2013, 4, 109. [Google Scholar] [CrossRef] [PubMed]
- Michallet, A.-S.; Mondière, P.; Taillardet, M.; Leverrier, Y.; Genestier, L.; Defrance, T. Compromising the Unfolded Protein Response Induces Autophagy-Mediated Cell Death in Multiple Myeloma Cells. PLoS ONE 2011, 6, e25820. [Google Scholar] [CrossRef] [PubMed]
- Schewe, D.M.; Aguirre-Ghiso, J.A. Inhibition of eIF2alpha dephosphorylation maximizes bortezomib efficiency and eliminates quiescent multiple myeloma cells surviving proteasome inhibitor therapy. Cancer Res. 2009, 69, 1545–1552. [Google Scholar] [CrossRef] [PubMed]
- Obeng, E.A.; Carlson, L.M.; Gutman, D.M.; Harrington, W.J.; Lee, K.P.; Boise, L.H. Proteasome inhibitors induce a terminal unfolded protein response in multiple myeloma cells. Blood 2006, 107, 4907–4916. [Google Scholar] [CrossRef]
- Adams, J. The development of proteasome inhibitors as anticancer drugs. Cancer Cell 2004, 5, 417–421. [Google Scholar] [CrossRef]
- Dong, H.; Chen, L.; Chen, X.-Q.; Gu, H.; Gao, G.; Gao, Y.; Dong, B. Dysregulation of unfolded protein response partially underlies proapoptotic activity of bortezomib in multiple myeloma cells. Leuk. Lymphoma 2009, 50, 974–984. [Google Scholar] [CrossRef]
- Hu, J.; Dang, N.; Menu, E.; De Bryune, E.; Xu, D.; Van Camp, B.; Van Valckenborgh, E.; Vanderkerken, K. Activation of ATF4 mediates unwanted Mcl-1 accumulation by proteasome inhibition. Blood 2012, 119, 826–837. [Google Scholar] [CrossRef]
- Tagoug, I.; Jordheim, L.P.; Herveau, S.; Matera, E.-L.; Huber, A.-L.; Chettab, K.; Manié, S.; Dumontet, C. Therapeutic Enhancement of ER Stress by Insulin-Like Growth Factor I Sensitizes Myeloma Cells to Proteasomal Inhibitors. Clin. Cancer Res. 2013, 19, 3556–3566. [Google Scholar] [CrossRef] [PubMed]
- Ri, M. Endoplasmic-reticulum stress pathway-associated mechanisms of action of proteasome inhibitors in multiple myeloma. Int. J. Hematol. 2016, 104, 273–280. [Google Scholar] [CrossRef] [PubMed]
- Rivas, A.; Vidal, R.L.; Hetz, C. Targeting the unfolded protein response for disease intervention. Expert Opin. Ther. Targets 2015, 19, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Hetz, C.; Axten, J.M.; Patterson, J.B. Pharmacological targeting of the unfolded protein response for disease intervention. Nat. Methods 2019, 15, 764–775. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Vattem, K.M.; Sood, R.; An, J.; Liang, J.; Stramm, L.; Wek, R.C. Identification and Characterization of Pancreatic Eukaryotic Initiation Factor 2 α-Subunit Kinase, PEK, Involved in Translational Control. Mol. Cell. Biol. 1998, 18, 7499–7509. [Google Scholar] [CrossRef]
- Mahameed, M.; Wilhelm, T.; Darawshi, O.; Obiedat, A.; Tommy, W.-S.; Chintha, C.; Schubert, T.; Samali, A.; Chevet, E.; Eriksson, L.A.; et al. The unfolded protein response modulators GSK2606414 and KIRA6 are potent KIT inhibitors. Cell Death Dis. 2019, 10, 300. [Google Scholar] [CrossRef]
- Axten, J.M.; Medina, J.R.; Feng, Y.; Shu, A.; Romeril, S.P.; Grant, S.W.; Li, W.H.H.; Heerding, D.A.; Minthorn, E.; Mencken, T.; et al. Discovery of 7-Methyl-5-(1-{[3-(trifluoromethyl)phenyl]acetyl}-2,3-dihydro-1H-indol-5-yl)-7H-pyrrolo[2,3-d]pyrimidin-4-amine (GSK2606414), a Potent and Selective First-in-Class Inhibitor of Protein Kinase R (PKR)-like Endoplasmic Reticulum Kinase (PERK). J. Med. Chem. 2012, 55, 7193–7207. [Google Scholar] [CrossRef]
- Bagratuni, T.; Wu, P.; De Castro, D.G.; Davenport, E.L.; Dickens, N.J.; Walker, B.A.; Boyd, K.; Johnson, D.C.; Gregory, W.; Morgan, G.J.; et al. XBP1s levels are implicated in the biology and outcome of myeloma mediating different clinical outcomes to thalidomide-based treatments. Blood 2010, 116, 250–253. [Google Scholar] [CrossRef]
- Harnoss, J.M.; Le Thomas, A.; Shemorry, A.; Marsters, S.A.; Lawrence, D.A.; Lu, M.; Chen, Y.-C.A.; Qing, J.; Totpal, K.; Kan, D.; et al. Disruption of IRE1α through its kinase domain attenuates multiple myeloma. Proc. Natl. Acad. Sci. USA 2019, 116, 16420–16429. [Google Scholar] [CrossRef]
- Pal, R.; Janz, M.; Galson, D.L.; Gries, M.; Li, S.; Jöhrens, K.; Anagnostopoulos, I.; Dörken, B.; Mapara, M.Y.; Borghesi, L.; et al. C/EBPbeta regulates transcription factors critical for proliferation and survival of multiple myeloma cells. Blood 2009, 114, 3890–3898. [Google Scholar] [CrossRef]
- Novoa, I.; Zeng, H.; Harding, H.P.; Ron, D. Feedback Inhibition of the Unfolded Protein Response by GADD34-Mediated Dephosphorylation of eIF2α. J. Cell Biol. 2001, 153, 1011–1022. [Google Scholar] [CrossRef] [PubMed]
- Klimczak, M.; Biecek, P.; Zylicz, A.; Zylicz, M. Heat shock proteins create a signature to predict the clinical outcome in breast cancer. Sci. Rep. 2019, 9, 7507. [Google Scholar] [CrossRef] [PubMed]
- Segawa, T.; Nau, M.E.; Xu, L.L.; Chilukuri, R.N.; Makarem, M.; Zhang, W.; Petrovics, G.; Sesterhenn, I.A.; McLeod, D.G.; Moul, J.W.; et al. Androgen-induced expression of endoplasmic reticulum (ER) stress response genes in prostate cancer cells. Oncogene 2002, 21, 8749–8758. [Google Scholar] [CrossRef]
- Ma, Y.; Hendershot, L.M. Herp Is Dually Regulated by Both the Endoplasmic Reticulum Stress-specific Branch of the Unfolded Protein Response and a Branch That Is Shared with Other Cellular Stress Pathways. J. Biol. Chem. 2004, 279, 13792–13799. [Google Scholar] [CrossRef]
- Hsu, S.-K.; Chiu, C.-C.; Dahms, H.-U.; Chou, C.-K.; Cheng, C.M.; Chang, W.-T.; Cheng, K.-C.; Wang, H.-M.D.; Lin, I.-L. Unfolded Protein Response (UPR) in Survival, Dormancy, Immunosuppression, Metastasis, and Treatments of Cancer Cells. Int. J. Mol. Sci. 2019, 20, 2518. [Google Scholar] [CrossRef] [PubMed]
- Gundamaraju, R.; Vemuri, R.; Chong, W.C.; Myers, S.A.; Norouzi, S.; Shastri, M.D.; Eri, R. Interplay between Endoplasmic Reticular Stress and Survivin in Colonic Epithelial Cells. Cells 2018, 7, 171. [Google Scholar] [CrossRef]
- Cabrera, E.; Hernández-Pérez, S.; Koundrioukoff, S.; Debatisse, M.; Kim, D.; Smolka, M.B.; Freire, R.; Gillespie, D.A. PERK inhibits DNA replication during the Unfolded Protein Response via Claspin and Chk1. Oncogene 2016, 36, 678–686. [Google Scholar] [CrossRef]
- Zhu, F.; Jiang, D.; Zhang, M.; Zhao, B. 2,4-Dihydroxy-3′-methoxy-4′-ethoxychalcone suppresses cell proliferation and induces apoptosis of multiple myeloma via the PI3K/akt/mTOR signaling pathway. Pharm. Biol. 2019, 57, 641–648. [Google Scholar] [CrossRef]
- Paíno, T.; Garcia-Gomez, A.; González-Méndez, L.; Segundo, L.S.; Hernández-García, S.; López-Iglesias, A.-A.; Algarín, E.M.; Corchete, L.A.; Corbacho-González, D.; Ortiz-De-Solórzano, C.; et al. The novel pan-PIM kinase inhibitor, PIM447, displays dual anti-myeloma and bone protective effects, and potently synergizes with current standards of care. Clin. Cancer Res. 2016, 23, 225–238. [Google Scholar] [CrossRef]
- Wang, J.; Li, Y.; Sun, W.; Liu, J.; Chen, W. Synergistic effects of rmhTRAIL and 17-AAG on the proliferation and apoptosis of multiple myeloma cells. Hematology 2018, 23, 620–625. [Google Scholar] [CrossRef]
- Mai, E.K.; Haas, E.-M.; Lücke, S.; Löpprich, M.; Kunz, C.; Pritsch, M.; Knaup-Gregori, P.; Raab, M.S.; Schlenzka, J.; Bertsch, U.; et al. A systematic classification of death causes in multiple myeloma. Blood Cancer J. 2018, 8, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Fandy, T.E.; Shankar, S.; Ross, D.D.; Sausville, E.; Srivastava, R.K. Interactive Effects of HDAC Inhibitors and TRAIL on Apoptosis Are Associated with Changes in Mitochondrial Functions and Expressions of Cell Cycle Regulatory Genes in Multiple Myeloma. Neoplasia 2005, 7, 646–657. [Google Scholar] [CrossRef] [PubMed]
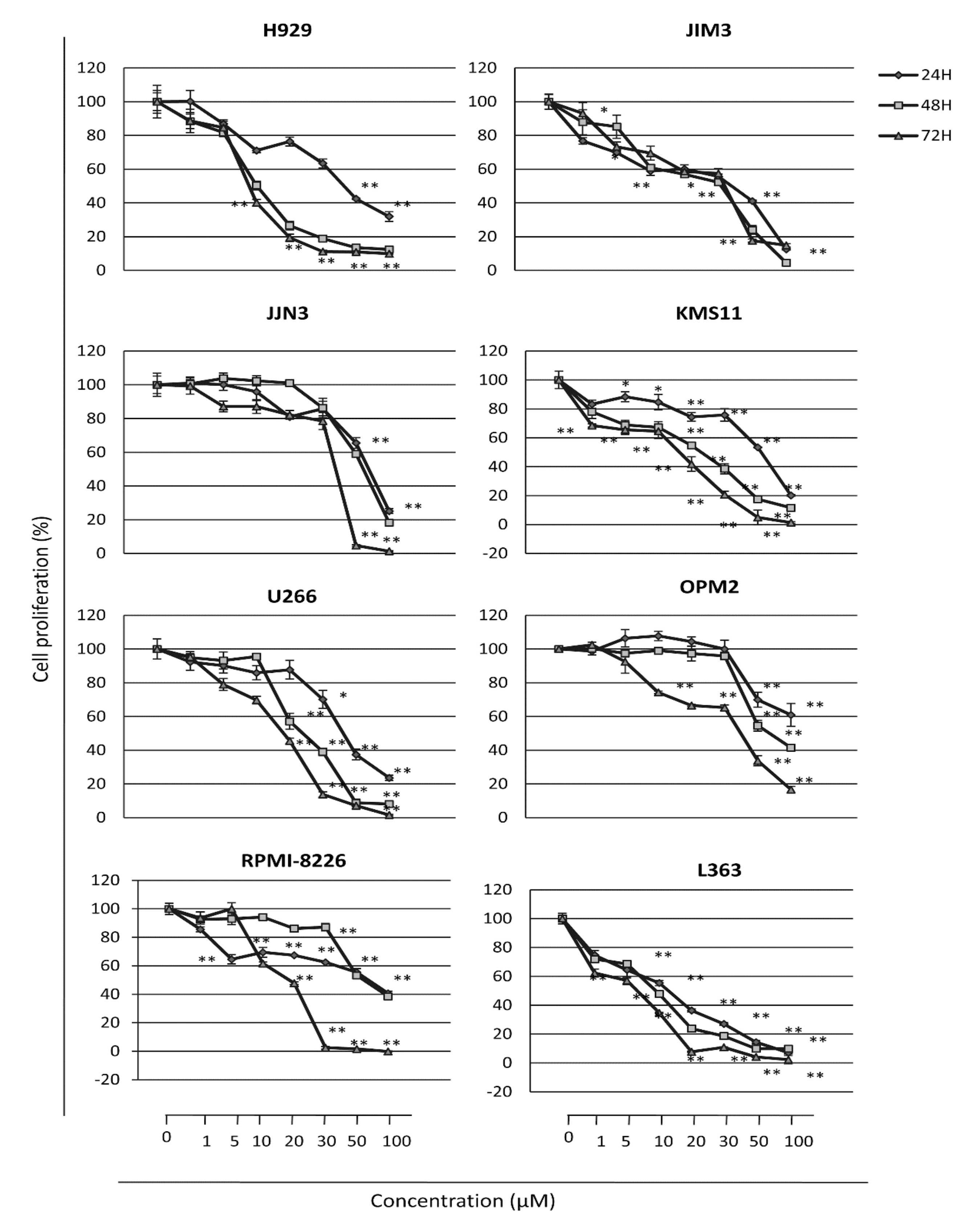
| Cell Lines | IC50 Values (μM) * |
|---|---|
| H929 | 10 ± 0.04 |
| JIM3 | 29± 0.01 |
| JJN3 | 75± 0.2 |
| KMS11 | 22± 0.035 |
| L363 | 9.5± 0.251 |
| OPM2 | 75± 0.641 |
| RPMI-8226 | 61± 0.56 |
| U266 | 21± 0.08 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bagratuni, T.; Patseas, D.; Mavrianou-Koutsoukou, N.; Liacos, C.I.; Sklirou, A.D.; Rousakis, P.; Gavriatopoulou, M.; Terpos, E.; Tsitsilonis, O.E.; Trougakos, I.P.; et al. Characterization of a PERK Kinase Inhibitor with Anti-Myeloma Activity. Cancers 2020, 12, 2864. https://doi.org/10.3390/cancers12102864
Bagratuni T, Patseas D, Mavrianou-Koutsoukou N, Liacos CI, Sklirou AD, Rousakis P, Gavriatopoulou M, Terpos E, Tsitsilonis OE, Trougakos IP, et al. Characterization of a PERK Kinase Inhibitor with Anti-Myeloma Activity. Cancers. 2020; 12(10):2864. https://doi.org/10.3390/cancers12102864
Chicago/Turabian StyleBagratuni, Tina, Dimitrios Patseas, Nefeli Mavrianou-Koutsoukou, Christine Ivy Liacos, Aimilia D. Sklirou, Pantelis Rousakis, Maria Gavriatopoulou, Evangelos Terpos, Ourania E. Tsitsilonis, Ioannis P. Trougakos, and et al. 2020. "Characterization of a PERK Kinase Inhibitor with Anti-Myeloma Activity" Cancers 12, no. 10: 2864. https://doi.org/10.3390/cancers12102864
APA StyleBagratuni, T., Patseas, D., Mavrianou-Koutsoukou, N., Liacos, C. I., Sklirou, A. D., Rousakis, P., Gavriatopoulou, M., Terpos, E., Tsitsilonis, O. E., Trougakos, I. P., Kastritis, E., & Dimopoulos, M. A. (2020). Characterization of a PERK Kinase Inhibitor with Anti-Myeloma Activity. Cancers, 12(10), 2864. https://doi.org/10.3390/cancers12102864








